- 1Department of Medical Genetics, Shiraz University of Medical Sciences, Shiraz, Iran
- 2Academic Unit of Bone Biology, Department of Oncology and Metabolism, The Mellanby Centre for Bone Research, The University of Sheffield, Sheffield, UK
- 3Transgenic Technology Research Center, Shiraz University of Medical Sciences, Shiraz, Iran
- 4Endocrine and Metabolic Research Center, Namazi Hospital, Shiraz University of Medical Sciences, Shiraz, Iran
- 5Design Trials and Statistics, School of Health and Related Research (ScHARR), The University of Sheffield, Sheffield, UK
The genetic epidemiology of variation in bone mineral density (BMD) and osteoporosis is not well studied in Iranian populations and needs more research. We report a candidate gene association study of BMD variation in a healthy cross-sectional study of 501 males and females sampled from the Iranian Multi-Centre Osteoporosis Study, Shiraz, Iran. We selected to study the association with 21 single nucleotide polymorphisms (SNPs) located in the 7 candidate genes LRP5, RANK, RANKL, OPG, P2RX7, VDR, and ESR1. BMD was measured at the three sites L2–L4, neck of femur, and total hip. Association between BMD and each SNP was assessed using multiple linear regression assuming an allele dose (additive effect) on BMD (adjusted for age and sex). Statistically significant (at the unadjusted 5% level) associations were seen with seven SNPs in five of the candidate genes. Two SNPs showed statistically significant association with more than one BMD site. Significant association was seen between BMD at all the three sites with the VDR SNP rs731246 (L2–L4 p = 0.038; neck of femur p = 0.001; and total hip p < 0.001). The T allele was consistently associated with lower BMD than the C allele. Significant association was also seen for the P2RX7 SNP rs3751143, where the G allele was consistently associated with lower BMD than the T allele (L2–L4 p = 0.069; neck of femur p = 0.024; and total hip p = 0.045).
Introduction
Osteoporosis is a skeletal disorder characterized by a reduction in the strength of bone with a consequent increase in bone fragility and susceptibility to fracture. Low bone mass is associated with micro-architectural deterioration of bone tissue (1). The loss of strength and bone mass can be as a result of a multitude of factors including the inability to reach optimal peak bone mass in early adulthood, increased resorption of bone having reached peak mass, or dysfunction of bone formation during bone remodeling (2). In this regard, hip fractures are the most important complication associated with osteoporosis and are linked to morbidity and mortality. While the cause of death is often due to other chronic disease, 25% of deaths are directly linked to the hip fracture or complications arising from the fracture such as pressure sores, infection, and bronchopneumonia (3). The 12-month survival rate post-hip fracture in the UK is 63% for men and 75% for women. In addition, one in two women and one in five men who are aged over 50 years will have suffered from an osteoporotic fracture in their remaining lifetime (4).
It is expected that by 2050, the world population will increase; as a result, it is also thought that the population of elderly people will grow (4). In the UK, by the year 2020, the over 85 years old age group is estimated to rise to 2.1 million, whereas this population was 1.2 million in 2002 (5). This increase in the aging population in the UK is predicted to cause double the number of osteoporotic fractures over the next 50 years (6). On this subject, genetic factors are important in the pathogenesis of osteoporosis, with links to a reduction in bone mineral density (BMD) and an increased susceptibility to osteoporotic fractures (7). Although genetic effects on BMD are mediated by several genetic factors with mild influences, more studies are needed for determining the relation between genetic factors and bone loss (8). In this regard, genome-wide association studies (GWAS) and candidate gene association studies are the two robust approaches to investigate correlation between genetic variation and disease risk to identify genome regions or candidate genes that contribute to osteoporosis disease (9, 10). The data obtained from these two approaches are growing with some suggestion of ethnic differences in the genetic associations with BMD (11).
The genetic epidemiology of variation in BMD is not well studied in Iranian populations and needs more research. To this end, we set out to examine the relative contributions of seven major candidate genes to BMD variation in a healthy cross-sectional study of males and females sampled from Shiraz, Iran.
Materials and Methods
Study Participants and Phenotypes
Subjects for the study were selected from The Shiraz Centre of the Iranian Multi-Centre Osteoporosis Study (IMOS), which is an ongoing study to determine reference values for bone densitometry in the female and male Iranian population from 2005. In the Shiraz Centre, 540 healthy subjects aged 20–75 years were selected randomly from 1080 subjects who were registered in the IMOS cohort and invited to participate in this study. Five hundred and one responded to the invitation and consented to participate (This sample of subjects is referred to as IMOS-AD throughout this report.). The study protocol was approved by the Research Ethics Committee of the Shiraz Medical University of the Iranian Ministry of Health and Medical Education. All the participants provided written informed consent after being fully informed of the nature of the study (in accordance with Helsinki Declaration).
Height was read from a free-standing height with the nearest 0.5 cm. Subsequently, subjects were weighed and body mass index (BMI) was calculated (kilograms per square meter).
Identifying Candidate Genes
We aimed to study variants with established associations with BMD variation in other populations and/or those with relevance to the Iranian population. The genes with established associations were selected if evidence for association was reported and replicated through a candidate gene association study (CGAS), a meta-analysis of candidate gene association study (MA-C), a GWAS, and a meta-analysis of genome-wide association study (MA-G). Six key publications were used to select the genes with well-replicated associations with BMD (12–17). This led to the selection of LRP5, RANK, RANKL, OPG, and ESR1 as these associations were reported by CGAS, MA-C, GWAS, and MA-G studies. These genes are also involved in three biological pathways, the Wnt/-catenin signaling pathway, the estrogen endocrine pathway, and the RANKL/RANK/OPG pathway. The VDR gene was also selected as it is the candidate gene related to osteoporosis most studied in Iranian populations (18, 19). P2RX7 was also selected as it was a novel candidate gene (at the time of recruiting the subjects to the study) associated with BMD and osteoporotic fracture risk (20, 21).
SNP Selection
Genome-wide association studies literature review was used for the strategic selection of single nucleotide polymorphisms (SNPs) that were the most relevant in BMD. The catalog of published GWAS in the National Human Genome Research Institute (NHGRI) was used to select associated SNPs for the selected candidate genes (22). This resulted in a total of 22 SNPs for the candidate gene association analysis with BMD variation in the IMOS-AD study. The specific SNPs selected were rs599083 (LRP5), rs3736228 (LRP5), rs884205 (RANK), rs3018362 (RANK), rs9594738 (RANKL), rs9594759 (RANKL), rs1021188 (RANKL), rs6469804 (OPG), rs4355801 (OPG), rs2062375 (OPG), rs2062377 (OPG), rs11995824 (OPG), rs2504063 (ESR1), rs1999805 (ESR1), rs4870044 (ESR1), rs1038304 (ESR1), rs2941740 (ESR1), rs6929137 (ESR1), rs3751143 (P2RX7), rs28360457 (P2XR7), rs731236 (VDR), and rs2228570 (VDR).
Bone Mineral Density
Bone mineral densitometry was performed by dual-energy X-ray absorptiometry (DEXA) method (GE Lunar DPX-IQ Bone densitometer, GE Healthcare, Madison, WI, USA). It was used to measure appendicular and axial sites and measurements are reported in units of milligrams per square centimeter. Minimal radiation exposure was performed as produced in the method by Blake and Fogelman (23). Definitive data of BMD measurements of the total proximal femur and lumbar vertebrae (L2–L4) in the anterior–posterior position were obtained.
Blood Sampling and DNA Extraction
Blood sampling was performed by the brachial vein puncture method (24, 25), and 200 μl of peripheral blood was collected from each participant in an EDTA tube. Finally, there were 501 participants whose blood samples and BMD were available.
The collected peripheral blood samples were subjected to DNA extraction through application of QIAamp 96 DNA Blood Kit. Finally, the extracted DNA samples were assessed by spectrophotometric measurements. The quality and purity of DNA was checked using a NanoDrop-1000 spectrophometer (Thermo Scientific) with an OD 260/280 ratio between 1.8 and 2 used as the criteria for acceptable DNA purity.
Genotyping
Genotyping of the selected SNPs was performed by Competitive Allele-Specific Polymerase chain reaction (KASPar) assay (KBioscience services; United Kingdom). The KASPar assay SNP genotyping system from KBioscience is based on fluorescent resonance energy transfer (FRET) technology. Following completion of the reaction, the data reported were evaluated using the KlusterCaller 1.1 software (KBioscience). Hardy–Weinberg equilibrium analysis was conducted for quality control. One SNP (rs9594759) gave a p-value below the threshold of 0.001 (26) and was excluded from further analysis, so final results are presented for 21 SNPs only. All included SNPs had a call rate in excess of 95%.
All assays were tested on LGC Genomics’ in-house validation DNA prior to being run on the study samples. Assays were deemed to be working successfully when clusters were distinct and call rates consistently high. The data are automatically quality control checked on a per SNP basis. No template controls (NTCs) were included on each plate to enable the detection of contamination or non-specific amplification – these samples must not amplify during the reaction.
The number of genotypes that were callable must be greater than 90% and minor allele frequency should be greater than 2% unless the SNP is known to be a very low frequency.
Statistical Analysis
GraphPad Prism 6 and IBM® SPSS® were used to analyze the data. The comparison of means for two independent groups was analyzed using the Student’s t-test. The t-test was used for all two group comparisons as the sample size in each group was always in excess of 50; the variables were approximately normally distributed, and sample SDs were similar. The association between BMD and the number of copies of the minor allele was analyzed using linear regression with the BMD variable as the outcome adjusting for the covariates age and sex of participant. For each linear regression analysis, the SNP genotype was coded as 0, 1, or 2 in terms of how many copies of the minor allele they carried. We decided to adjust for these two common confounders as this adjustment has been done in other studies (15) and adjusting for more covariates would reduce the statistical power. The adjusted association is reported along with 95% confidence intervals (95% CI). All reported p-values are nominal and not adjusted for multiple testing.
The power of the patient sample size of 501 to detect true associations depends upon the frequency of the genotypes. If the frequency of the common homozygote and the heterozygote were similar (each around 40%), then the sample would have 90% power to detect a true association responsible for a difference of 0.23 of a SD in BMD. If the heterozygote is as rare as 16% and the homozygote has frequency of 84%, then 501 patients would have 90% power to detect a difference in BMD of 0.4 of a SD. These calculations assume a two-sided 5% type 1 error rate.
Results
The BMI of the Males and Females within a Random Selection of Subjects from an Iranian Population
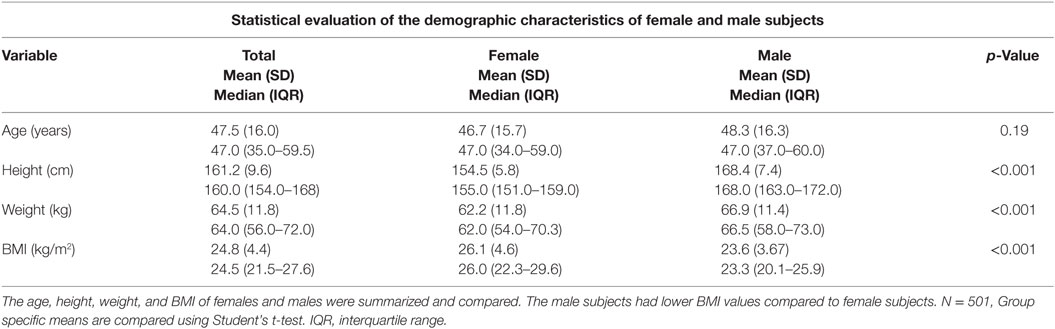
Each subject was supplied with a questionnaire from which demographic variables for this study including age, height, weight, and BMI were determined. The data were analyzed separately by gender as BMI is known to be higher in females than in males in the Iranian population (27). Menopausal status was recorded on the women, and 111 of the 248 women were postmenopausal. Statistical differences between genders were formally compared using a Student’s t-test. Statistical analysis showed that there were no statistically significant differences in age between the male and female subjects of our sample group, but the male subjects within our sample population had significantly higher weight and height metrics compared to the female subjects (p < 0.001). The calculated BMI was higher for females than for the male subjects within the selected Iranian population sample (p < 0.001) (Table 1).
The BMD of the Males and Females within a Random Selection of Subjects from an Iranian Population
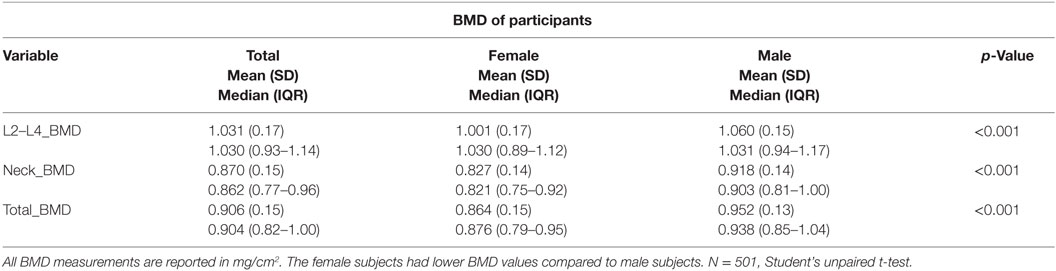
The L2–L4 (L2–L4_BMD), neck of femur (Neck_BMD), and total hip BMD (Total_BMD) of the IMOS-AD sample consisting of 501 individuals was assessed using the DXA Scanner. Data from the whole sample population were separated into male and female groups to examine any gender associations with BMD. The gender-specific group means were compared using a Student’s t-test.
Analysis of the Iranian sample population showed that the male subjects had statistically significantly higher BMD means associated with L2–L4_BMD, neck_BMD, and total_BMD compared to that of the female population (p < 0.001) (Table 2).
Analysis of Genotyping Data of 21 SNPS in Seven Selected Candidate Genes and BMD in the Iranian Subjects
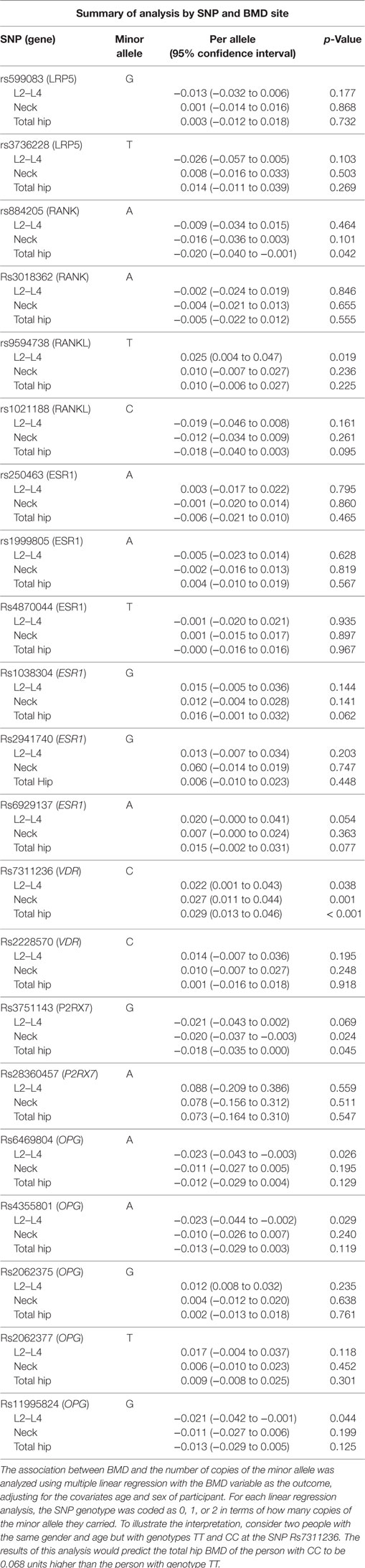
LRP5, RANK, RANKL, ESR1, VDR, P2XR7, and OPG were selected following the literature review of genes for consideration in an Iranian population. The genetic association analysis was conducted assuming an additive allelic effect on the outcome variable BMD. The analysis was repeated for each BMD variable and for each SNP (i.e., 3 × 21 linear regression analyses). Regression analyses were performed adjusting for age and sex. The results in Table 3 show the linear regression coefficient for the association between the BMD outcome and the number of copies of the risk allele.
Table 3 shows the association analyses grouped by SNP. Out of a total of 63 regression analyses of the data, 10 tests are statistically significant at the 5% level (three would be expected under the null hypotheses). However, the most notable result here is of the VDR polymorphism rs731236, which shows a statistically significant association at all the three BMD sites. The allele T is associated with lower BMD after adjusting for age and sex. The P2RX7 allele G for SNP rs3751143 is statistically significantly associated with lower BMD at two sites (the neck and total hip) and is marginally associated at the L2–L4 site.
Discussion
This study looked at whether the BMD of a random selection of subjects from an Iranian population was genetically associated with a selection of candidate genes known to be associated with BMD. A total of 21 SNPs in 7 selected genes were evaluated for this study. The descriptive analysis of our random selection of subjects from the Iranian population showed that there were no significant differences between the mean ages of the males and females within the group. Additionally, the weight and height of the males were higher than that of the females, as observed in similar subject groups of other similar Iranian studies (27). These data are an indication that as the demographic characteristics of the randomly selected sample group was comparable to samples within other studies; further genetic studies would also be comparable to the literature. It is of interest to note that these data showed that male subjects had a lower BMI range compared to the female subjects and these data were consistent with findings from Bahrami et al. (27).
Data collected from the BMD analysis of our Iranian population samples showed that the total hip BMD of the male subjects was higher than the female subjects. The average BMD of our male sample group was 0.95 mg/cm2, and the females had an average BMD value of 0.86 mg/cm2. These data were consistent with BMD data obtained from other Iranian groups (28, 29), European groups (30) and American groups (31). These data suggest that the BMD pattern observed within our sample group was comparable to that of others used in similar genetic association studies (32, 33).
As mentioned previously, a low BMD is associated with the risk of developing osteoporosis (34). The relationship between BMD and BMI, including its weight and height constituents, is widely reported (35–37). In the literature, BMD is reported to be lower in lean postmenopausal women in the majority of studies. However, some studies have also reported increased BMD in lean postmenopausal women (38).
It is expected that many different genetic variants contribute to the regulation of BMD. Most of the variants are assumed to have small effect size, but there is evidence that rare variants of large effect size also contribute in some individuals (39). In the past 10 years, there have been advances in the identification and validation of osteoporosis susceptibility loci via large-scale association studies, meta-analyses, and GWAS of SNPs. A thorough review of latter developments has revealed that more than 15 genes can be assigned as confirmed BMD genes linked to osteoporosis (40).
The LRP5, RANK, RANKL, P2RX7, ESR1, OPG, and VDR genes selected for this study are known to have important roles in four well-defined biological pathways; the estrogen endocrine pathway, the Wnt/β-catenin signaling pathway, the RANKL/RANK/OPG pathway, and the vitamin D synthesis pathway (40). There are limited data related to effective genetic variants in osteoporosis, particularly in respect to the mentioned genes, in the Iranian population. The association analysis found 10 of the 63 tests resulted in p < 0.05 (Table 3). Under the null of no genetic association, we would only expect to see three of the results statistically significant. Hence, we do see some confirmation of associations with this set of genetic loci in the Iranian sample. However, the SNPs selected for analysis in this cohort are already established as risk SNPs, so in this evaluation, we are mainly interested to know if the overall pattern of association is similar to other populations or may be different. Such differences may be due to different population, alleles, or the impact of a distinct environment; hence, it is more the estimate and 95% CI that is of interest.
Out of the 21 selected SNPs, our genotyping data identified statistically significant associations between BMD and 7 of these SNPs: Rs731236, Rs3751143, Rs884205, Rs9594738, Rs4355801, rs6469804, and Rs11995824.
For the RANK gene, the present study showed that SNP rs884205 was associated with total hip BMD, and the same association was observed in studies by Styrkarsdottir et al. (30) within a European and East Asian population (15, 41). The present study further showed that RANKL SNP rs9594738 was also closely linked to spine BMD, similar to studies by Duncan et al. and Styrkarsdottir et al. in the European population (16, 42). The OPG SNPs rs2073804, rs1995824, and rs4355801 associations with spine BMD determined in the present study were also observed in other studies specific to spine and hip BMD in post/premenopausal women and the European population, respectively (33, 43). Although there are few published data on P2RX7, one study did correlate with ours in that SNP rs3751143 was associated with BMD in a similar number of subjects (44).
The most interesting result in our study is the association with the VDR SNP rs731246 (also known as TaqI), which is statistically significant over all the three measured BMD sites. Early work from Keen et al. (45) and Spector et al. (46) using UK Twin cohorts, as well as the work from Ferrari et al. (47), led to the identification of common polymorphisms in the VDR as being associated with BMD and an important target for the treatment of osteoporosis. However, there have been conflicting results (48), and more importantly meta-analyses including this genetic variant have not found consistent evidence for its association with BMD variation or osteoporotic fracture risk (8, 49–51). However, one Iranian study has shown a statistically significant association with the VDR SNP rs2228570 and BMD in females, though they did not examine the SNP rs731246.
While this study is a substantial contribution to the genetic epidemiology of variation in BMD in the Iranian population, it does have limitations. The sample size is modest, and this means the study is powered to detect larger effects but limits the potential for adjustment for additional covariates (such as menopausal status and BMI, height or weight). Over-adjustment for covariates impacts statistical power. When studying the association between the SNP genotypes and BMD, it is possible that for some SNPs, there was an imbalance of post- and premenopausal women in the genotype groups and this might lead to a spurious association. However, including age and gender as covariates would mitigate this to some extent. This study had no information on incidence of fractures hence we cannot be confident any of these associations would extend to increased facture risk.
In conclusion, our data demonstrate that the overall pattern of association of these established risk SNPs in the Iranian population are similar to those observed in other populations. From our data, we concur that BMD is genetically dependent (52), and that the VDR is statistically significantly associated with BMD in the Iranian population. Future prospective studies using this cohort would enable us to analyze fracture incidence, as well as the functional effects of these SNPs in bone cells.
Author Contributions
SD designed and carried out the full IMOS-AD research project, and collected the data and patient samples. He conducted the statistical analysis and drafted the research paper. MT, AG, ST, and GO advised supervised and guided the design and analysis of the research project. All four were involved in drafting and revising the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Funding
The research presented in this manuscript was conducted by SAD while registered as a self-funded joint location PhD student at the University of Sheffield and University of Shiraz. The project was supported by the Vice Chancellor Fund of Shiraz Medical University.
References
1. Fauci AS. Harrison’s Principles of Internal Medicine. 17th ed. McGraw-Hill, Medical Publishing Division (2008).
2. Sattui SE, Saag KG. Fracture mortality: associations with epidemiology and osteoporosis treatment. Nat Rev Endocrinol (2014) 10(10):592–602. doi:10.1038/nrendo.2014.125
3. Cole ZA, Dennison EM, Cooper C. Osteoporosis epidemiology update. Curr Rheumatol Rep (2008) 10(2):92–6. doi:10.1007/s11926-008-0017-6
4. Kassim Javaid M, Chana J, Cooper C. Hip fracture as the tracer condition. Best Pract Res Clin Rheumatol (2013) 27(6):711–5. doi:10.1016/j.berh.2014.03.003
5. Clark S. Osteoporosis – the disease of the 21st century? Lancet (2002) 359(9319):1714. doi:10.1016/S0140-6736(02)08662-2
6. Kanis JA, Johnell O. Requirements for DXA for the management of osteoporosis in Europe. Osteoporos Int (2005) 16(3):229–38. doi:10.1007/s00198-004-1811-2
7. Cheung CL, Xiao SM, Kung AW. Genetic epidemiology of age-related osteoporosis and its clinical applications. Nat Rev Rheumatol (2010) 6(9):507–17. doi:10.1038/nrrheum.2010.106
8. Uitterlinden AG, Ralston SH, Brandi ML, Carey AH, Grinberg D, Langdahl BL, et al. The association between common vitamin D receptor gene variations and osteoporosis: a participant-level meta-analysis (vol 145, pg 255, 2006). Ann Intern Med (2006) 145(12):936–936. doi:10.7326/0003-4819-145-4-200608150-00005
9. Zmuda JM, Sheu YT, Moffett SP. Genetic epidemiology of osteoporosis: past, present, and future. Curr Osteoporos Rep (2005) 3(3):111–5. doi:10.1007/s11914-005-0019-5
10. Richards JB, Zheng HF, Spector TD. Genetics of osteoporosis from genome-wide association studies: advances and challenges. Nat Rev Genet (2012) 13(8):576–88. doi:10.1038/nrg3228
11. Zengin A, Prentice A, Ward KA. Ethnic differences in bone health. Front Endocrinol (2015) 6:24. doi:10.3389/fendo.2015.00024
12. Richards JB, Rivadeneira F, Inouye M, Pastinen TM, Soranzo N, Wilson SG, et al. Bone mineral density, osteoporosis, and osteoporotic fractures: a genome-wide association study. Lancet (2008) 371(9623):1505–12. doi:10.1016/S0140-6736(08)60599-1
13. Richards JB, Kavvoura FK, Rivadeneira F, Styrkársdóttir U, Estrada K, Halldórsson BV, et al. Collaborative meta-analysis: associations of 150 candidate genes with osteoporosis and osteoporotic fracture. Ann Intern Med (2009) 151(8):528–37. doi:10.7326/0003-4819-151-8-200910200-00006
14. Paternoster L, Lorentzon M, Vandenput L, Karlsson MK, Ljunggren O, Kindmark A, et al. Genome-wide association meta-analysis of cortical bone mineral density unravels allelic heterogeneity at the RANKL locus and potential pleiotropic effects on bone. PLoS Genet (2010) 6(11):e1001217. doi:10.1371/journal.pgen.1001217
15. Rivadeneira F, Styrkársdottir U, Estrada K, Halldórsson BV, Hsu YH, Richards JB, et al. Twenty bone-mineral-density loci identified by large-scale meta-analysis of genome-wide association studies. Nat Genet (2009) 41(11):1199–206. doi:10.1038/ng.446
16. Styrkarsdottir U, Halldorsson BV, Gretarsdottir S, Gudbjartsson DF, Walters GB, Ingvarsson T, et al. Multiple genetic loci for bone mineral density and fractures. N Engl J Med (2008) 358(22):2355–65. doi:10.1056/NEJMoa0801197
17. Styrkarsdottir U, Halldorsson BV, Gretarsdottir S, Gudbjartsson DF, Walters GB, Ingvarsson T, et al. New sequence variants associated with bone mineral density. Nat Genet (2009) 41(1):15–7. doi:10.1038/ng.284
18. Hossein-nezhad A, Varzaneh FN, Mirzaei K. Association of the FokI polymorphism in the vitamin D receptor gene with vertebral fracture in Iranian postmenopausal women. Endocr Pract (2011) 17(5):826–7. doi:10.4158/EP.17.5.826
19. Pouresmaeili F, Jamshidi J, Azargashb E, Samangouee S. Association between vitamin D receptor gene BsmI polymorphism and bone mineral density in a population of 146 Iranian women. Cell J (2013) 15(1):75–82.
20. Ohlendorff SD, Tofteng CL, Jensen JE, Petersen S, Civitelli R, Fenger M, et al. Single nucleotide polymorphisms in the P2X7 gene are associated to fracture risk and to effect estrogen treatment. Pharmacogenet Genomics (2007) 17(7):555–67. doi:10.1097/FPC.0b013e3280951625
21. Gartland A, Skarratt KK, Hocking LJ, Parsons C, Stokes L, Jørgensen NR, et al. Polymorphisms in the P2X7 receptor gene are associated with low lumbar spine bone mineral density and accelerated bone loss in post-menopausal women. Eur J Hum Genet (2012) 20(5):559–64. doi:10.1038/ejhg.2011.245
22. Hindorff LA, Sethupathy P, Junkins HA, Ramos EM, Mehta JP, Collins FS, et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc Natl Acad Sci U S A (2009) 106(23):9362–7. doi:10.1073/pnas.0903103106
23. Blake GM, Fogelman I. Technical principles of dual energy x-ray absorptiometry. Semin Nucl Med (1997) 27(3):210–28. doi:10.1016/S0001-2998(97)80025-6
24. Galena HJ. Complications occurring from diagnostic venipuncture. J Fam Pract (1992) 34(5):582–4.
25. Okeson GC, Wulbrecht PH. The safety of brachial artery puncture for arterial blood sampling. Chest (1998) 114(3):748–51. doi:10.1378/chest.114.3.748
26. Anderson CA, Pettersson FH, Clarke GM, Cardon LR, Morris AP, Zondervan KT. Data quality control in genetic case-control association studies. Nat Protoc (2010) 5(9):1564–73. doi:10.1038/nprot.2010.116
27. Bahrami H, Sadatsafavi M, Pourshams A, Kamangar F, Nouraei M, Semnani S, et al. Obesity and hypertension in an Iranian cohort study; Iranian women experience higher rates of obesity and hypertension than American women. BMC Public Health (2006) 6:158. doi:10.1186/1471-2458-6-158
28. Larijani B, Moayyeri A, Keshtkar AA, Hossein-Nezhad A, Soltani A, Bahrami A, et al. Peak bone mass of Iranian population: the Iranian Multicenter Osteoporosis Study. J Clin Densitom (2006) 9(3):367–74. doi:10.1016/j.jocd.2006.05.001
29. Omrani GR. Bone mineral density in the normal Iranian population: a comparison with American reference data. Arch Osteoporos (2006) 1(1–2):29–35. doi:10.1007/s11657-006-0005-2
30. Styrkarsdottir U, Halldorsson BV, Gudbjartsson DF, Tang NL, Koh JM, Xiao SM, et al. European bone mineral density loci are also associated with BMD in East-Asian populations. PLoS One (2010) 5(10):e13217. doi:10.1371/journal.pone.0013217
31. Mazess RB, Barden H. Bone density of the spine and femur in adult white females. Calcif Tissue Int (1999) 65(2):91–9. doi:10.1007/s002239900663
32. Kiel DP, Ferrari SL, Cupples LA, Karasik D, Manen D, Imamovic A, et al. Genetic variation at the low-density lipoprotein receptor-related protein 5 (LRP5) locus modulates Wnt signaling and the relationship of physical activity with bone mineral density in men. Bone (2007) 40(3):587–96. doi:10.1016/j.bone.2006.09.029
33. Shang M, Lin L, Cui H. Association of genetic polymorphisms of RANK, RANKL and OPG with bone mineral density in Chinese peri- and postmenopausal women. Clin Biochem (2013) 46(15):1493–501. doi:10.1016/j.clinbiochem.2013.03.011
34. Cummings SR, Nevitt MC, Browner WS, Stone K, Fox KM, Ensrud KE, et al. Risk factors for hip fracture in white women. Study of Osteoporotic Fractures Research Group. N Engl J Med (1995) 332(12):767–73. doi:10.1056/NEJM199503233321202
35. Felson DT, Zhang Y, Hannan MT, Anderson JJ. Effects of weight and body mass index on bone mineral density in men and women: the Framingham study. J Bone Miner Res (1993) 8(5):567–73. doi:10.1002/jbmr.5650080507
36. Ravn P, Cizza G, Bjarnason NH, Thompson D, Daley M, Wasnich RD, et al. Low body mass index is an important risk factor for low bone mass and increased bone loss in early postmenopausal women. Early Postmenopausal Intervention Cohort (EPIC) study group. J Bone Miner Res (1999) 14(9):1622–7. doi:10.1359/jbmr.1999.14.9.1622
37. Nguyen TV, Center JR, Eisman JA. Osteoporosis in elderly men and women: effects of dietary calcium, physical activity, and body mass index. J Bone Miner Res (2000) 15(2):322–31. doi:10.1359/jbmr.2000.15.2.322
38. Salamat MR, Salamat AH, Abedi I, Janghorbani M. Relationship between weight, body mass index, and bone mineral density in men referred for dual-energy X-ray absorptiometry scan in Isfahan, Iran. J Osteoporos (2013) 2013:205963. doi:10.1155/2013/205963
39. Ralston SH. Genetics of osteoporosis. Ann N Y Acad Sci (2010) 1192:181–9. doi:10.1111/j.1749-6632.2009.05317.x
40. Li WF, Hou SX, Yu B, Li MM, Férec C, Chen JM. Genetics of osteoporosis: accelerating pace in gene identification and validation. Hum Genet (2010) 127(3):249–85. doi:10.1007/s00439-009-0773-z
41. Estrada K, Styrkarsdottir U, Evangelou E, Hsu YH, Duncan EL, Ntzani EE, et al. Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nat Genet (2012) 44(5):491–501. doi:10.1038/ng.2249
42. Duncan EL, Danoy P, Kemp JP, Leo PJ, McCloskey E, Nicholson GC, et al. Genome-wide association study using extreme truncate selection identifies novel genes affecting bone mineral density and fracture risk. PLoS Genet (2011) 7(4):e1001372. doi:10.1371/journal.pgen.1001372
43. Mencej-Bedrač S, Preželj J, Marc J. TNFRSF11B gene polymorphisms 1181G > C and 245T > G as well as haplotype CT influence bone mineral density in postmenopausal women. Maturitas (2011) 69(3):263–7. doi:10.1016/j.maturitas.2011.02.010
44. Wesselius A, Bours MJ, Henriksen Z, Syberg S, Petersen S, Schwarz P, et al. Association of P2X7 receptor polymorphisms with bone mineral density and osteoporosis risk in a cohort of Dutch fracture patients. Osteoporos Int (2013) 24(4):1235–46. doi:10.1007/s00198-012-2059-x
45. Keen RW, Major PJ, Lanchbury JS, Spector TD. Vitamin-D-receptor-gene polymorphism and bone loss. Lancet (1995) 345(8955):990–1. doi:10.1016/S0140-6736(95)90740-8
46. Spector TD, Keen RW, Arden NK, Morrison NA, Major PJ, Nguyen TV, et al. Influence of vitamin D receptor genotype on bone mineral density in postmenopausal women: a twin study in Britain. BMJ (1995) 310(6991):1357–60. doi:10.1136/bmj.310.6991.1357
47. Ferrari S, Rizzoli R, Chevalley T, Slosman D, Eisman JA, Bonjour JP. Vitamin-D-receptor-gene polymorphisms and change in lumbar-spine bone mineral density. Lancet (1995) 345(8947):423–4. doi:10.1016/S0140-6736(95)90404-2
48. Peacock M, Hustmyer F, Hui S, Johnston CC, Christian J. Vitamin-D-receptor genotype and bone mineral density. Evidence conflicts on link. BMJ (1995) 311(7009):874–5. doi:10.1136/bmj.311.7009.874a
49. Fang Y, Rivadeneira F, van Meurs JB, Pols HA, Ioannidis JP, Uitterlinden AG. Vitamin D receptor gene BsmI and TaqI polymorphisms and fracture risk: a meta-analysis. Bone (2006) 39(4):938–45. doi:10.1016/j.bone.2006.04.016
50. Qin G, Dong Z, Zeng P, Liu M, Liao X. Association of vitamin D receptor BsmI gene polymorphism with risk of osteoporosis: a meta-analysis of 41 studies. Mol Biol Rep (2013) 40(1):497–506. doi:10.1007/s11033-012-2086-x
51. Ma C, Zhou QL, Deng YJ, Liang X, Wu DP, Dong YZ. Association of vitamin D receptor BsmI gene polymorphism with risk of low bone mineral density in post-menopausal women: a meta-analysis. Genet Mol Res (2014) 13(3):7791–9.
Keywords: VDR, P2RX7, candidate gene association, BMD, genetic epidemiology, Iranian population
Citation: Dastgheib SA, Gartland A, Tabei SMB, Omrani GR and Teare MD (2016) A Candidate Gene Association Study of Bone Mineral Density in an Iranian Population. Front. Endocrinol. 7:141. doi: 10.3389/fendo.2016.00141
Received: 11 April 2016; Accepted: 13 October 2016;
Published: 27 October 2016
Edited by:
Kate Anna Ward, Medical Research Council Human Nutrition Research, UKReviewed by:
Niklas Rye Jørgensen, Copenhagen University Hospital Glostruo, DenmarkAnders Eriksson, University of Cambridge, UK
Copyright: © 2016 Dastgheib, Gartland, Tabei, Omrani and Teare. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Seyed Alireza Dastgheib, ZGFzdGdoZWlic2FAZ21haWwuY29t;
Marion Dawn Teare, bS5kLnRlYXJlQHNoZWZmaWVsZC5hYy51aw==
 Seyed Alireza Dastgheib
Seyed Alireza Dastgheib Alison Gartland
Alison Gartland Seyed Mohammad Bagher Tabei1,3
Seyed Mohammad Bagher Tabei1,3 Marion Dawn Teare
Marion Dawn Teare