- Department of Psychosomatic Medicine, Graduate School of Medical Sciences, Kyushu University, Fukuoka, Japan
There is increasing interest in the interactions among the gut microbiota, gut, and brain, which is often referred to as the “microbiota-gut-brain” axis. Biogenic amines including dopamine, norepinephrine, serotonin, and histamines are all generated by commensal gut microorganisms and are suggested to play roles as signaling molecules mediating the function of the “microbiota-gut-brain” axis. In addition, such amines generated in the gut have attracted attention in terms of possible clues into the etiologies of depression, anxiety, and even psychosis. This review covers the latest research related to the potential role of microbe-derived amines such as catecholamine, serotonin, histamine, as well as other trace amines, in modulating not only gut physiology but also brain function of the host. Further attention in this field can offer not only insight into expanding the fundamental roles and impacts of the human microbiome, but also further offer new therapeutic strategies for psychological disorders based on regulating the balance of resident bacteria.
Introduction
Humans co-exist with a vast number of microorganisms (1), and there has been increased research interest in uncovering the role of gut bacteria in maintaining health of the host. Accumulating evidence points to a major role of the gut microbiota in not only normal gut function but also in brain development and function (2–4). The recognition of such interactions between gut microorganisms and the brain has led to a new research field commonly referred to as the “microbiota-gut-brain” axis (5–9).
Biogenic amines are biogenic substances containing one or more amine groups (10, 11). Five of these amines were found to function as neurotransmitters including dopamine, norepinephrine, epinephrine, histamine, and serotonin. In general, these amines are utilized in the central and peripheral nervous systems, and participate in various types of physiological functions such as regulating cognitive abilities, mood, and gut motility (12–14). In addition to these well-established roles as neurotransmitters, accumulating evidence suggests that these amines might act as important signal molecules between commensal microbiota and the host in the gut (15–17). Here, the latest research progress on the relationship between gut microbiota and biogenic amines is reviewed, with a particular focus on interactions occurring in the gut lumen, in which gut bacteria are presumed to closely associate with biogenic amines.
Catecholamines: Norepinephrine and Dopamine
Catecholamines as Inter-Kingdom Signals in the Gut Lumen
Bacteria can communicate with each other through hormone-like signals in a process known as quorum sensing (18). Recent evidence has revealed that quorum sensing is not only restricted to bacterial cell–cell communication but also facilitates communication between microorganisms and their hosts. This type of bidirectional communication is often referred to as “microbial endocrinology” (15, 16) or “inter-kingdom signaling” (17, 19, 20), which thus mediates the symbiotic and pathogenic relationships between bacteria and their mammalian hosts.
Interestingly, catecholamines have emerged as potential inter-kingdom signaling molecules in the gut, in addition to their well-established roles as neurotransmitters in the central and peripheral nervous systems. In their pioneering studies conducted in the 1990s, Lyte et al. (21, 22) demonstrated that some pathogenic species could recognize exogenous catecholamines in vitro, which enhanced the bacterial proliferative capacity. Subsequently, Sperandio et al. (19) showed that enterohemorrhagic Escherichia coli virulence is increased upon exposure to epinephrine and norepinephrine, and that epinephrine binds and signals through the QseC receptor, providing a molecular basis for catecholamines acting as an inter-kingdom signal.
Catecholamines Are Enriched in the Lumen of the Gut
Since a large number of bacteria inhabit areas in close proximity to the gastrointestinal tract of mammals, inter-kingdom signaling through catecholamines is presumed to continually and preferentially occur in the gut lumen. However, confirmation of this localized activity is a technical challenge because luminal samples generate substantial artifact peaks and contaminants, which hinder the precise evaluation of luminal catecholamine levels. In contrast, a clear and unique peak of 5-hydroxytryptamine (5-HT), or serotonin, was clearly identified without interference by contaminants (23). Therefore, to more reliably measure fecal catecholamine levels, we used a three-column high-performance liquid chromatography system with diphenylethylenediamine as a pre-column fluorescence derivatization reagent (24, 25), which was originally developed to evaluate plasma and urine free catecholamine levels. As a result, this analysis revealed considerable amounts of biologically active, free norepinephrine, and dopamine present in the gut lumen of specific pathogen free (SPF) mice, which harbor commensal gut microbiota that lacks specific pathogens (26).
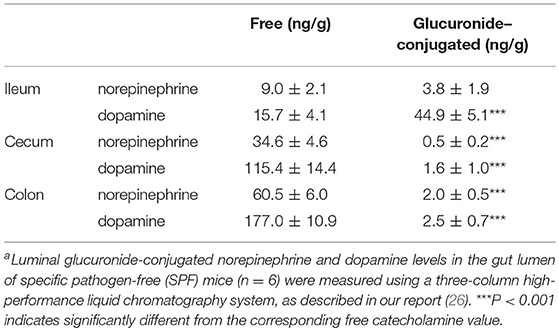
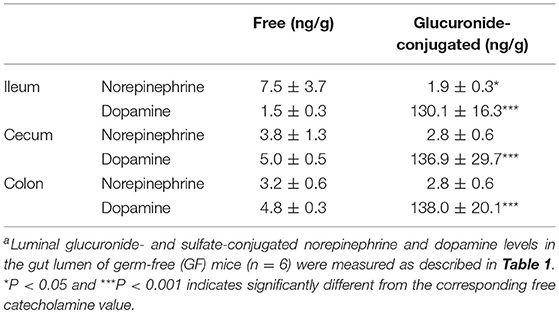
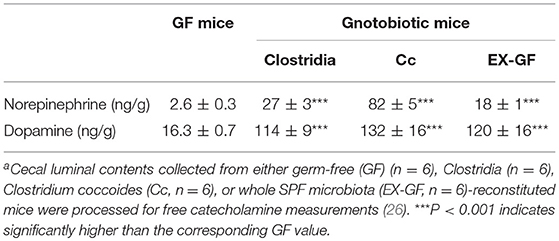
The intestinal tract contains substantial amounts of β-glucuronidase (GUS) in both the gut epithelium (tissue type) and gut bacteria (bacterial type). The optimum pH of bacterial GUS is 6.8, whereas that of tissue-type GUS is 4.5 (27, 28). Since the average pH in the intestinal lumen is 6.5–8.0, GUS enzymatic activity is mainly derived from gut bacteria. GUS can play an important role in gut–liver interactions such as the so-called enterohepatic circulation (29–31). For example, hormones such as thyroid hormone are glucurono-conjugated in the liver, and the resultant conjugated materials are released into the bile duct. After reaching the lower gastrointestinal tract, these hormones are converted into biologically active hormones by the action of bacterial GUS, and thereafter reabsorbed into the body (30). Therefore, we examined whether biologically active free catecholamines can be produced by the action of bacterial GUS. As summarized (26), almost all catecholamines detected in SPF mice with a normal gut microbiota were of the biologically active free type (Table 1). In contrast, more than 90% of the dopamine and approximately 20 to 40% of the norepinephrine detected in germ-free (GF) mice were of the glucurono-conjugated type (Table 2). However, when GF mice were provided either a mixture of Clostridia species, a single bacterial species of Clostridium coccoides, or complete SPF microbiota rich in GUS activity, they exhibited a dramatic increase in free norepinephrine and dopamine (Table 3).
These results clearly indicate that gut microbes play a critical role in the production of free catecholamines through bacterial GUS.
Can Commensal Bacteria Themselves Produce Catecholamines in vivo?
The next important question to address is whether gut microbes have the ability to generate catecholamines by themselves. Russian researchers (32) reported that some species of microorganisms could produce catecholamines in an in vitro culture system. Although this report is very interesting, the possibility that the medium used for bacterial culture contained catecholamines derived from blood, tissues, or other sources cannot be ruled out. To have the authentic capacity to synthesize catecholamines, microorganisms require tyrosine hydroxylase, a rate-limiting enzyme for catecholamine production. In fact, some bacteria were reported to possess enzymes that are similar to mammalian tyrosine hydroxylase (33, 34). In our study, the total norepinephrine levels in the cecal and colonic contents of SPF mice were substantially higher than those in GF mice (26). In addition, gut bacteria enriched from murine feces harbored substantial amounts of norepinephrine with a relatively lower amount of dopamine (35). These findings imply that gut microbes are an important source of luminal norepinephrine. However, some species of bacteria have a functional transporter for catecholamines such as the bacterial neurotransmitter sodium symporter family member, Leu T (36). Therefore, it remains to be clarified whether the norepinephrine and dopamine found in gut microbes originate from bacterial production via a tyrosine hydroxylase-like enzyme or if they are obtained from the gut lumen via a Leu T-like transporter (35). In this regard, Lyte and Brown (37) recently showed that Lactobacillus salivarius biofilms, but not Lactobacillus rhamnosus biofilms, might express both plasma membrane monoamine transporter- and organic cation transporter-like uptake systems, using specific fluorescence-based assays. These results indicate that bacteria residing in the gut lumen retain uptake systems that would act as a net sink for biogenic amines and neuroactive substances, and play an important role in preventing the excessive production of neurotoxic amines in the host.
Functional Aspects of Catecholamines in the Gut Lumen
To reveal the specific roles that catecholamines play in the gut lumen, we further examined the effects of dopamine on water absorption in a mouse colon loop in vitro model (38, 39). As we previously reported (26), dopamine injection into the loop resulted in a 30% increase in water absorption from the gut lumen compared to that with vehicle (saline) injection, indicating that luminal dopamine can contribute to gut physiology under normal conditions. In addition, a recent report indicated the possible involvement of catecholamines in inflammatory bowel disease (IBD). Inhibiting catecholamine-QseC signaling attenuated disease activities in multiple preclinical IBD models (40), suggesting the therapeutic potential of QseC blockade for intestinal inflammation. Taken together, gut luminal catecholamines might contribute to a variety of as-yet-unidentified physiological and pathological functions.
5-HT
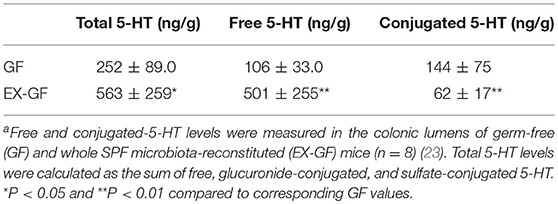
5-HT is abundant in the gastrointestinal tract, where it participates in various physiological functions such as peristalsis (41). The majority of 5-HT in the gut is stored in enterochromaffin cells (EC cells) and is secreted from the these cells into the gut lumen in response to various stimuli (42, 43). Intestinal bacteria are presumed to play a role in the secretion of 5-HT into the lumen; however, the precise underlying mechanism remains incompletely understood. Therefore, we examined kinetic 5-HT changes in the gut lumen after GF mice were provided SPF feces (23). As shown in Table 4, SPF fecal administration substantially elevated the cecal and colonic lumen 5-HT levels. In addition, approximately 50% of the 5-HT found in GF mice was in the conjugated form, whereas the majority of 5-HT found in EX-GF mice reconstituted with SPF feces was in the free form (Table 5). These results indicate that free 5-HT is not only released from EC stores in response to gut microbes but is also produced via the bacterial de-conjugation of conjugated 5-HT. Collectively, it can be concluded that free 5-HT in recolonized mice consists of the following three different components: (1) the first constitute is equivalent to free-5-HT in GF mice, which is likely released from EC cells in response to non-microbial stimuli; (2) the second is comparable to conjugated-5-HT in GF mice that is de-conjugated by microbial enzymes (GUS and sulfatase); (3) the third is mainly released from EC cells in response to microbial stimuli.
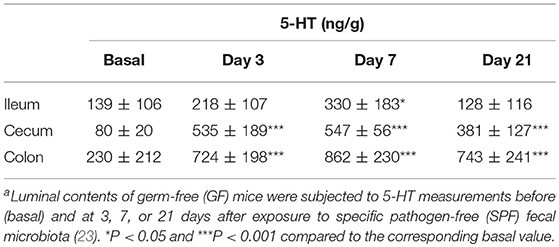
Table 4. Luminal 5-HT concentration in the gastrointestinal tract of GF mice after recolonization with SPF fecal microbiotaa.
Histamine and Other Biogenic Amines
Histamine is stored in mast cells and basophils and contributes to various types of patho-physiological processes. Histamine is best known as a mediator of the allergic reaction but also acts as a neurotransmitter for the brain (44). Histamine is not only synthesized in cells by the enzyme histidine decarboxylase but is also produced by the microbial decarboxylation of amino acids (45). This type of production is clinically important because bacteria-generated histamine often induces scombroid poisoning, which occurs after the consumption of food contaminated by amines (46). If an individual has a deficient capacity to detoxify biogenic amines due to genetic mutations or is taking antidepressants such as monoamine oxidase inhibitors (MAOIs) that slow the degradation of amines, they become more susceptible to histamine poisoning (46). Interestingly, commensal microorganisms in the gut can produce histamine and related compounds under physiological conditions (47, 48), suggesting the potential role of luminal histamine in gut immunoregulation. In fact, a recent elegant study demonstrated that histamine can exert an anti-inflammatory effects on the host by suppressing interleukin-18 production in the gut (49).
Moreover, researchers in this field have begun to turn their attention to the role of other biogenic amines such as tryptamine, phenethylamine, diamines (putrescine and cadaverine), and polyamines (spermine and spermidine) in mental health and psychiatric diseases (50, 51). In this regard, Gabastou et al. (52) offered an interesting case report. They measured time-course changes in fecal amines in a male patient who was admitted to a hospital due to severe brain damage incurred after a traffic accident at the age of 4 years. With a transient increase in these amines, the patient started to engage in self-injury and aggressive behaviors toward others. This case report indicates the possible involvement of bacteria-produced biogenic amines in the mental state. The authors suggested that antibiotics might be a suitable therapeutic option for psychomotor excitement that is refractory to commonly used treatments.
Thus, although the acute effects of histamine on the host, such as allergy and hypertension, are well established, the influence of histamine produced by indigenous microbes under physiological conditions remains unclear. This issue is especially relevant for not only individuals who are genetically susceptible to histamine exposure but also those who are regularly taking MAOIs for the treatment of depression.
Biogenic Amine Receptors on Gut Epithelial Cells
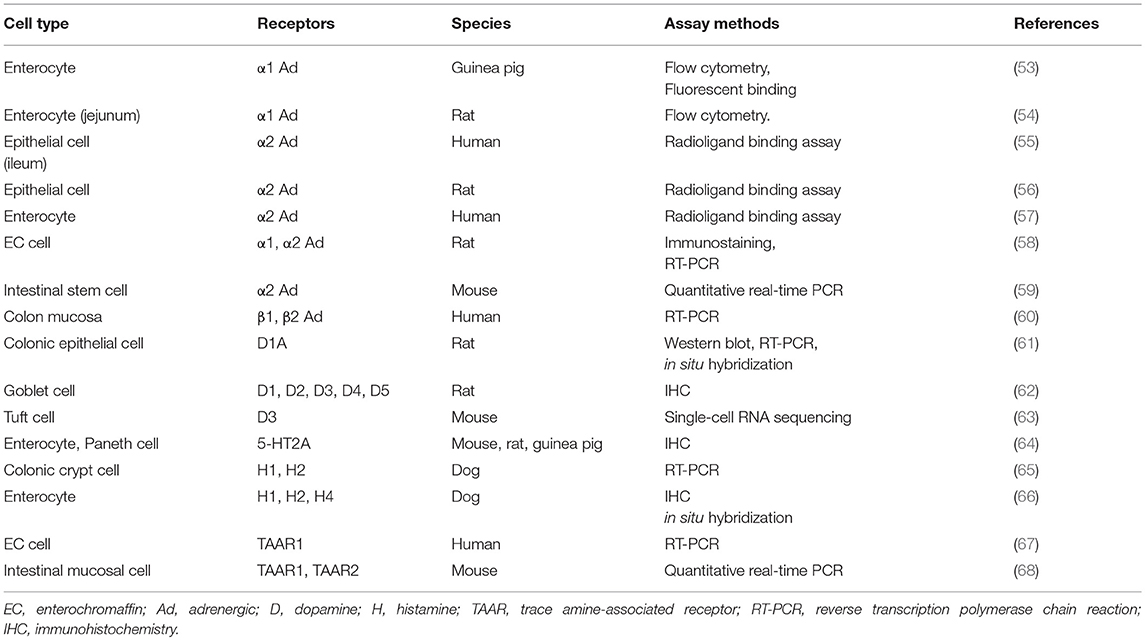
Gut microbe-derived amines are presumed to exert a direct effect on gut epithelial cells via specific receptors. Here, we provide a literature review of the expression of receptors of biogenic amines, especially focusing on intestinal epithelial cells. As summarized in Table 6, the expression of α1-adrenergic receptor on enterocytes was demonstrated by flow cytometry and binding assays (53, 54). Enterocytes from different species were also shown to harbor α2 adrenergic receptors, contributing to the net absorption of electrolytes and fluids (55–57). Interestingly, rat EC cells possess both α1 and α2 receptors, which was verified by immunostaining and RT-PCR (58). Recently, intestinal stem cells were also reported to retain not only muscarinic acetylcholine but also α2 adrenergic receptors (59). In humans, RT-PCR analysis showed that β1 and β2 adrenergic receptors are present in the colon mucosa (60). Regarding dopamine receptors, their subtypes were confirmed in rat colonic epithelia (61) and goblet cells (62). Moreover, an excellent report using an advanced single-cell RNA sequencing technique recently demonstrated that Tuft cells express dopamine receptor D3 genes (63).
Evidence as to whether the gut epithelia express receptors for other important monoamine molecules such as 5-HT and histamine is still scarce. One immunohistochemical analysis of different species demonstrated the presence of 5-HT2A receptors on enterocytes and Paneth cells (64). Histamine H1 and H2 receptors were also identified in canine enterocytes (65, 66); however, whether this expression is limited to dogs needs to be clarified. Finally, trace amine-associated receptors (TAARs) such as TAAR1 and TAAR2, for which ligands include tyramine, phenethylamine, and other biogenic amines (11), are also shown to exist in human EC cells (67) and the mouse intestinal mucosa (68). Since there is extensively increasing interest in trace amines derived from commensal bacteria, it is critically important to clarify the precise distribution and function of TAAR-related receptors in the gut. Finally, detailed information about the cellular localization of biogenic amine receptors, such as apical or basolateral membrane distributions, is still limited. This should be clarified by future studies.
Conclusion and Perspectives
From the 19th to the early 20th century, a faction of scientists postulated that psychiatric diseases might result from “autointoxication,” suggesting that waste products or toxins generated in the gut can lead to depression, anxiety, and even psychosis (7, 69, 70). Until recently, the concept of autointoxication was regarded as an “unscientific” theory and was largely neglected. However, this theory has reemerged as an attractive research area and is currently being extensively studied. As reviewed herein, biogenic amines are interesting candidates that could be important mediators of autointoxication. Further developments in this field could provide a strong rationale for the application of probiotics for the treatment of mental health and diseases.
Author Contributions
NS wrote the manuscript and contributed to all aspects of this paper.
Conflict of Interest Statement
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by a KAKENHI Grant-in-Aid for Scientific Research on Innovative Areas Will dynamics (JP16H06404: NS), Scientific Research (B) (JP16H05278: NS), Exploratory Research (JP16K15413: NS). The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
1. Qin J, Li R, Raes J, Arumugam M, Burgdorf KSS, Manichanh C, et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature. (2010) 464:59–65. doi: 10.1038/nature08821
2. Lozupone CA, Stombaugh JI, Gordon JI, Jansson JK, Knight R. Diversity, stability and resilience of the human gut microbiota. Nature. (2012) 489:220–30. doi: 10.1038/nature11550
3. Nicholson JK, Holmes E, Kinross J, Burcelin R, Gibson G, Jia W, et al. Host-gut microbiota metabolic interactions. Science. (2012) 336:1262–7. doi: 10.1126/science.1223813
4. Clemente JC, Ursell LK, Parfrey LW, Knight R. The impact of the gut microbiota on human health: an integrative view. Cell. (2012) 148:1258–70. doi: 10.1016/j.cell.2012.01.035
5. Forsythe P, Sudo N, Dinan T, Taylor VH, Bienenstock J. Mood and gut feelings. Brain Behav Immun. (2010) 24:9–16. doi: 10.1016/j.bbi.2009.05.058
6. Cryan JF, Dinan TG. Mind-altering microorganisms: the impact of the gut microbiota on brain and behaviour. Nat Rev Neurosci. (2012) 13:701–12. doi: 10.1038/nrn3346
8. Smith PA. Brain, meet gut: neuroscientists are probing the connections between intestinal microbes and brain development. Nature. (2015) 526:312–4. doi: 10.1038/526312a
9. Sampson TR, Mazmanian SK. Control of brain development, function, and behavior by the microbiome. Cell Host Microbe. (2015) 17:565–576. doi: 10.1016/j.chom.2015.04.011
10. Santos MHS. Biogenic amines: their importance in foods. Int J Food Microbiol. (1996) 29:213–31. doi: 10.1016/0168-1605(95)00032-1
11. Gainetdinov RR, Hoener MC, Berry MD. Trace amines and their receptors. Pharmacol Rev. (2018) 70:549–620. doi: 10.1124/pr.117.015305
12. Carrasco GA, Van de Kar LD. Neuroendocrine pharmacology of stress. Eur J Pharmacol. (2003) 463:235–72. doi: 10.1016/S0014-2999(03)01285-8
13. Esler M, Jennings G, Lambert G, Meredith I, Horne M, Eisenhofer G. Overflow of catecholamine neurotransmitters to the circulation: source, fate, and functions. Physiol Rev. (1990) 70:963–85. doi: 10.1152/physrev.1990.70.4.963
14. Brown RE, Stevens DR, Haas HL. The physiology of brain histamine. Prog Neurobiol. (2001) 63:637–72. doi: 10.1016/S0301-0082(00)00039-3
15. Lyte M. Microbial endocrinology and infectious disease in the 21st century. Trends Microbiol. (2004) 12:14–20. doi: 10.1016/j.tim.2003.11.004
16. Freestone PP, Sandrini SM, Haigh RD, Lyte M. Microbial endocrinology: how stress influences susceptibility to infection. Trends Microbiol. (2008) 16:55–64. doi: 10.1016/j.tim.2007.11.005
17. Hughes DT, Sperandio V. Inter-kingdom signalling: communication between bacteria and their hosts. Nat Rev Microbiol. (2008) 6:111–20. doi: 10.1038/nrmicro1836
18. Bassler BL. How bacteria talk to each other: regulation of gene expression by quorum sensing. Curr Opin Microbiol. (1999) 2:582–7. doi: 10.1016/S1369-5274(99)00025-9
19. Clarke MB, Hughes DT, Zhu C, Boedeker EC, Sperandio V. The QseC sensor kinase: a bacterial adrenergic receptor. Proc Natl Acad Sci USA. (2006) 103:10420–5. doi: 10.1073/pnas.0604343103
20. Baumler AJ, Sperandio V. Interactions between the microbiota and pathogenic bacteria in the gut. Nature. (2016) 535:85–93. doi: 10.1038/nature18849
21. Lyte M, Ernst S. Alpha and beta adrenergic receptor involvement in catecholamine-induced growth of gram-negative bacteria. Biochem Biophys Res Commun. (1993) 190:447–52. doi: 10.1006/bbrc.1993.1068
22. Lyte M, Ernst S. Catecholamine induced growth of gram negative bacteria. Life Sci. (1992) 50:203–12. doi: 10.1016/0024-3205(92)90273-R
23. Hata T, Asano Y, Yoshihara K, Kimura-Todani T, Miyata N, Zhang X-T, et al. Regulation of gut luminal serotonin by commensal microbiota in mice. PLoS ONE. (2017) 12:e0180745. doi: 10.1371/journal.pone.0180745
24. Claustre J, Serusclat P, Peyrin L. Glucuronide and sulfate catecholamine conjugates in rat and human plasma. J Neural Transm. (1983) 56:265–78. doi: 10.1007/BF01243495
25. Shibata O, Morooka H, Hara K, Nakamura H, Fukuzaki M, Gotoh Y, et al. Importance of the liver and the digestive tract in the formation of catecholamine conjugates. Masui. (1988) 37:428–33.
26. Asano Y, Hiramoto T, Nishino R, Aiba Y, Kimura T, Yoshihara K, et al. Critical role of gut microbiota in the production of biologically active, free catecholamines in the gut lumen of mice. Am J Physiol - Gastrointest Liver Physiol. (2012) 303:G1288–95. doi: 10.1152/ajpgi.00341.2012
27. Kim D-HH, Jin Y-HH, Jung E-AA, Han MJJ, Kobashi K. Purification and characterization of beta-glucuronidase from Escherichia coli HGU-3, a human intestinal bacterium. Biol Pharm Bull. (1995) 18:1184–8. doi: 10.1248/bpb.18.1184
28. Rod TO, Midtvedt T. Origin of intestinal beta-glucuronidase in germfree, mono-contaminated and conventional rats. Acta Pathol Microbiol Scand Sect B Microbiol. (1977) 85:271–6. doi: 10.1111/j.1699-0463.1977.tb01973.x
29. Graef V, Furuya E, Nishikaze O. Hydrolysis of steroid glucuronides with beta-glucuronidase preparations from bovine liver, Helix pomatia, and E. coli. Clin Chem. (1977) 23:532–5.
30. Hazenberg MP, de Herder WW, Visser TJ. Hydrolysis of iodothyronine conjugates by intestinal bacteria. FEMS Microbiol Rev. (1988) 4:9–16. doi: 10.1111/j.1574-6968.1988.tb02709.x-i1
31. Ilett KF, Tee LBG, Reeves PT, Minchin RF. Mebolism of drugs and other xenobiotics in the gut lumen and wall. Pharmacol Ther. (1990) 46:67–93. doi: 10.1016/0163-7258(90)90036-2
32. Tsavkelova EA, Botvinko IV, Kudrin VS, Oleskin AV. Detection of neurotransmitter amines in microorganisms with the use of high-performance liquid chromatography. Dokl Biochem. (2000) 372:115–7.
33. Solano F, García E, Pérez De Egea E, Sanchez-Amat A, Garcia E, Perez D, et al. Isolation and Characterization of Strain MMB-1 (CECT 4803), a Novel Melanogenic Marine Bacterium. Appl Environ Microbiol. (1997) 63:3499–506.
34. Hernández-Romero D, Sanchez-Amat A, Solano F, Hernandez-Romero D, Sanchez-Amat A, Solano F. A tyrosinase with an abnormally high tyrosine hydroxylase/dopa oxidase ratio. FEBS J. (2006) 273:257–70. doi: 10.1111/j.1742-4658.2005.05038.x
35. Sudo N. Microbiome, HPA axis and production of endocrine hormones in the gut. Adv Exp Med Biol. (2014) 817:177–94. doi: 10.1007/978-1-4939-0897-4_8
36. Singh SK, Yamashita A, Gouaux E. Antidepressant binding site in a bacterial homologue of neurotransmitter transporters. Nature. (2007) 448:952–6. doi: 10.1038/nature06038
37. Lyte M, Brown DR. Evidence for PMAT- and OCT-like biogenic amine transporters in a probiotic strain of Lactobacillus: Implications for interkingdom communication within the microbiota-gut-brain axis. PLoS ONE. (2018) 13:e0191037. doi: 10.1371/journal.pone.0191037
38. Lyte M, Ernst S, Vaughan CJ, Aherne AM, Lane E, Power O, et al. Electrogenic Na+/HCO3- co-transporter-1 is essential for the parathyroid hormone-stimulated intestinal HCO3- secretion. Biochem Biophys Res Commun. (2007) 18:289–90. doi: 10.1016/j.bbrc.2011.05.087
39. Lam RS, App EM, Nahirney D, Szkotak AJ, Vieira-Coelho MA, King M, et al. Regulation of Cl- secretion by alpha-adrenergic receptors in mouse colonic epithelium. J Physiol. (2003) 548:475–84. doi: 10.1113/jphysiol.2002.036806
40. Rooks MG, Veiga P, Reeve AZ, Lavoie S, Yasuda K, Asano Y, et al. QseC inhibition as an antivirulence approach for colitis-associated bacteria. Proc Natl Acad Sci USA. (2017) 114:142–7. doi: 10.1073/pnas.1612836114
41. Tsukamoto K, Ariga H, Mantyh C, Pappas TNTN, Yanagi H, Yamamura T, et al. Luminally released serotonin stimulates colonic motility and accelerates colonic transit in rats. Am J Physiol Integr Comp Physiol. (2007) 293:R64–9. doi: 10.1152/ajpregu.00856.2006
42. Gershon MDD. 5-Hydroxytryptamine (serotonin) in the gastrointestinal tract. Curr Opin Endocrinol Diabetes Obes. (2013) 20:14–21. doi: 10.1097/MED.0b013e32835bc703
43. Bertrand PP, Bertrand RL. Serotonin release and uptake in the gastrointestinal tract. Auton Neurosci. (2010) 153:47–57. doi: 10.1016/j.autneu.2009.08.002
44. Thurmond RL, Gelfand EW, Dunford PJ. The role of histamine H1 and H4 receptors in allergic inflammation: the search for new antihistamines. Nat Rev Drug Discov. (2008) 7:41–53. doi: 10.1038/nrd2465
45. Stratton JE, Hutkins RW, Taylor SL. Biogenic amines in cheese and other fermented foods: a review. J Food Prot. (1991) 54:460–70. doi: 10.4315/0362-028X-54.6.460
46. Morrow JD, Margolies GR, Rowland J, Roberts LJ. Evidence that histamine is the causative toxin of scombroid-fish poisoning. N Engl J Med. (1991) 324:716–20. doi: 10.1056/NEJM199103143241102
47. Beaver MH, Wostmann BS. Histamine and 5-hydroxytryptamine in the intestinal tract of germ-free animals, animals harbouring one microbial species and conventional animals. Br J Pharmacol Chemother. (1962) 19:385–93. doi: 10.1111/j.1476-5381.1962.tb01443.x
48. Barcik W, Pugin B, Westermann P, Perez NR, Ferstl R, Wawrzyniak M, et al. Histamine-secreting microbes are increased in the gut of adult asthma patients. J Allergy Clin Immunol. (2016) 138:1491–4.e7. doi: 10.1016/j.jaci.2016.05.049
49. Levy M, Thaiss CA, Zeevi D, Dohnalová L, Zilberman-Schapira G, Mahdi JA, et al. Microbiota-modulated metabolites shape the intestinal microenvironment by regulating nlrp6 inflammasome signaling. Cell. (2015) 163:1428–43. doi: 10.1016/j.cell.2015.10.048
50. Matsumoto M, Kibe R, Ooga T, Aiba Y, Kurihara S, Sawaki E, et al. Impact of intestinal microbiota on intestinal luminal metabolome. Sci Rep. (2012) 2:233. doi: 10.1038/srep00233
51. Schretter CE, Vielmetter J, Bartos I, Marka Z, Marka S, Argade S, et al. A gut microbial factor modulates locomotor behaviour in Drosophila. Nature. (2018) 563:402–6. doi: 10.1038/s41586-018-0634-9
52. Gabastou JM, Nugon-Baudon L, Robert Y, Manuel C, Vaissade P, Bourgeon E, et al. Digestives amines of bacterial origin and behaviour. A case report. (Amines digestives d'origine bacterienne et troubles comportementaux. A propos d'une observation). Pathol Biol. (1996) 44:275–81.
53. Baglole CJ, Davison JS, Meddings JB. Epithelial distribution of neural receptors in the guinea pig small intestine. Can J Physiol Pharmacol. (2005) 83:389–95. doi: 10.1139/y05-024
54. Baglole CJ, Sigalet DL, Martin GR, Yao S, Meddings JB. Acute denervation alters the epithelial response to adrenoceptor activation through an increase in alpha1-adrenoceptor expression on villus enterocytes. Br J Pharmacol. (2006) 147:101–8. doi: 10.1038/sj.bjp.0706424
55. Valet P, Senard JM, Devedjian JC, Planat V, Salomon R, Voisin T, et al. Characterization and distribution of alpha 2-adrenergic receptors in the human intestinal mucosa. J Clin Invest. (1993) 91:2049–57. doi: 10.1172/JCI116427
56. Paris H, Voisin T, Remaury A, Rouyer-Fessard C, Daviaud D, Langin D, et al. Alpha-2 adrenoceptor in rat jejunum epithelial cells: characterization with [3H]RX821002 and distribution along the villus-crypt axis. J Pharmacol Exp Ther. (1990) 254:888–93.
57. Lepor H, Rigaud G, Shapiro E, Baumann M, Kodner IJ, Fleshman JW. Muscarinic cholinergic and alpha 2-adrenergic receptors in the epithelium and muscularis of the human ileum. Surgery. (1990) 107:461–7.
58. Schäfermeyer A, Gratzl M, Rad R, Dossumbekova A, Sachs G, Prinz C. Isolation and receptor profiling of ileal enterochromaffin cells. Acta Physiol Scand. (2004) 182:53–62. doi: 10.1111/j.1365-201X.2004.01299.x
59. Davis EA, Zhou W, Dailey MJ. Evidence for a direct effect of the autonomic nervous system on intestinal epithelial stem cell proliferation. Physiol Rep. (2018) 6:e13745. doi: 10.14814/phy2.13745
60. Roberts SJ, Papaioannou M, Evans BA, Summers RJ. Functional and molecular evidence for beta 1-, beta 2- and beta 3- adrenoceptors in human colon. Br J Pharmacol. (1997) 120:1527–35. doi: 10.1038/sj.bjp.0701056
61. Vaughan CJ, Aherne AM, Lane E, Power O, Carey RM, O'Connell DP. Identification and regional distribution of the dopamine D 1A receptor in the gastrointestinal tract. Am J Physiol Integr Comp Physiol. (2000) 279:R599–609. doi: 10.1152/ajpregu.2000.279.2.R599
62. Li Y, Zhang Y, Zhang X-L, Feng X-Y, Liu C-Z, Zhang X-N, et al. Dopamine promotes colonic mucus secretion through dopamine D5 receptor in rats. Am J Physiol Cell Physiol. (2019) 316:C393–403. doi: 10.1152/ajpcell.00261.2017
63. Haber AL, Biton M, Rogel N, Herbst RH, Shekhar K, Smillie C, et al. A single-cell survey of the small intestinal epithelium. Nature. (2017) 551:333–9. doi: 10.1038/nature24489
64. Fiorica-Howells E, Hen R, Gingrich J, Li Z, Gershon MD. 5-HT 2A receptors: location and functional analysis in intestines of wild-type and 5-HT 2A knockout mice. Am J Physiol Liver Physiol. (2002) 282:G877–893. doi: 10.1152/ajpgi.00435.2001
65. Schwittlick U, Junginger J, Hahn K, Habierski A, Hewicker-Trautwein M. Histamine Receptor Expression in the Gastrointestinal Tract of Dogs. Anat Histol Embryol. (2017) 46:33–42. doi: 10.1111/ahe.12229
66. Khan I, Lapierre N, Rangachari PK. Messenger RNAs for neurokinin and histamine receptor subtypes in isolated canine colonic crypts. J Pharmacol Exp Ther. (1995) 272:1285–92.
67. Kidd M, Modlin IM, Gustafsson BI, Drozdov I, Hauso O, Pfragner R. Luminal regulation of normal and neoplastic human EC cell serotonin release is mediated by bile salts, amines, tastants, and olfactants. Am J Physiol Gastrointest Liver Physiol. (2008) 295:G260–72. doi: 10.1152/ajpgi.00056.2008
68. Ito J, Ito M, Nambu H, Fujikawa T, Tanaka K, Iwaasa H, et al. Anatomical and histological profiling of orphan G-protein-coupled receptor expression in gastrointestinal tract of C57BL/6J mice. Cell Tissue Res. (2009) 338:257–69. doi: 10.1007/s00441-009-0859-x
69. Bested AC, Logan AC, Selhub EM. Intestinal microbiota, probiotics and mental health: from Metchnikoff to modern advances: Part I – autointoxication revisited. Gut Pathog. (2013) 5:5. doi: 10.1186/1757-4749-5-5
Keywords: catecholamine, dopamine, gut microbiota, histamine, norepinephrine, serotonin
Citation: Sudo N (2019) Biogenic Amines: Signals Between Commensal Microbiota and Gut Physiology. Front. Endocrinol. 10:504. doi: 10.3389/fendo.2019.00504
Received: 01 May 2019; Accepted: 11 July 2019;
Published: 31 July 2019.
Edited by:
John Bienenstock, McMaster University, CanadaReviewed by:
Richard T. Premont, Harrington Discovery Institute, United StatesAlain Couvineau, Institut National de la Santé et de la Recherche Médicale (INSERM), France
Copyright © 2019 Sudo. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Nobuyuki Sudo, bm9idXl1a2kuc3Vkby45MzVAbS5tZWQua3l1c2h1LXUuYWMuanA=
 Nobuyuki Sudo
Nobuyuki Sudo