- Department of Chemical and Systems Biology, Stanford University, Stanford, CA, United States
Understanding the mammalian energy balance can pave the way for future therapeutics that enhance energy expenditure as an anti-obesity and anti-diabetic strategy. Several studies showed that brown adipose tissue activity increases daily energy expenditure. However, the size and activity of brown adipose tissue is reduced in individuals with obesity and type two diabetes. Humans have an abundance of functionally similar beige adipocytes that have the potential to contribute to increased energy expenditure. This makes beige adipocytes a promising target for metabolic disease therapies. While brown adipocytes tend to be stable, beige adipocytes have a high level of plasticity that allows for the rapid and dynamic induction of thermogenesis by external stimuli such as low environmental temperatures. This means that after browning stimuli have been withdrawn beige adipocytes quickly transition back to their white adipose state. The detailed molecular mechanisms regulating beige adipocytes development, function, and reversibility are not fully understood. The goal of this review is to give a comprehensive overview of beige fat development and origins, along with the transcriptional and epigenetic programs that lead to beige fat formation, and subsequent thermogenesis in humans. An improved understanding of the molecular pathways of beige adipocyte plasticity will enable us to selectively manipulate beige cells to induce and maintain their thermogenic output thus improving the whole-body energy homeostasis.
Introduction
Prolonged periods of excess energy storage lead to weight gain and obesity. This excess energy is stored in white adipose tissue (WAT), which is the major fat storage depot and is linked to metabolic disease states. Contrarily, brown adipose tissues (BAT) dissipates energy as heat and consequently modulates daily energy expenditure (1). As the amount of metabolically active BAT is limited in patients with obesity and type two diabetes (T2D), alternatives are required to increase energetic expenditure via thermogenesis (2–6). In addition to classic brown adipocytes, human adults have inducible brown adipocytes (named as brown-in-white or beige) with unique characteristics that differentiate them from both white and brown adipocytes (1, 7–10). Inducing beige adipocytes formation in WAT (browning) potentially decreases the negative effects of excess WAT and improves overall metabolic health (11). In response to cold exposure, inducible BAT greatly increases mitochondria and uncoupling protein 1 (UCP1) abundance. Additionally, after browning stimuli are removed, there is a rapid decrease of the thermogenic gene expression. This dynamic response in beige fat can be contrasted with classic BAT where the levels of UCP1 and mitochondria are constitutively high (12, 13). Although numerous studies have identified several regulators of browning and thermogenesis, the molecular basis underlying beige adipocyte reversibility is yet to be understood (14–20).
Development and Origin of Brown and Beige Adipocytes
Beige adipocytes mainly reside in subcutaneous white adipose tissue (scWAT) depots (8). The scWAT depots in humans include cranial, facial, abdominal, femoral, and gluteal depots. In rodents, scWAT includes the anterior subcutaneous white adipose tissues (ascWAT) and the posterior subcutaneous adipose tissue (pscWAT) which itself includes inguinal, gluteal, and dorso-lumbal WAT (7, 21). BAT depots are distributed in the thoracic (mediastinal) and scapulae (interscapular, cervical, and axillary) areas of mice and rats (22). In humans, BAT was initially thought to exist only in the neck and shoulder of infants (23). However, later studies found active BAT in the paracervical and supraclavicular as well as in the anterior neck regions of adult humans (23–27).
In mammals, BAT is formed earlier during embryogenesis as compared to WAT. In human fetuses BAT formation begins early in the second trimester primarily in the head and neck regions and later in development forms in the trunk as well as in upper and lower limbs. The development of subcutaneous white adipose tissue is completed prenatally (28). In rodents, functional thermogenic BAT is formed 2 days before birth (E18–19) (29–32) and the development of scWAT continues postnatally (33–35).
Both white and brown adipose tissues are known to have mesodermal origins including the intermediate and lateral plate as well as the axial, and paraxial mesoderm. The paraxial mesoderm gives rise to BAT (36), and though the origin of scWAT is still debated, the progenitors of scWAT are known to originate from both the mesoderm and neuroectoderm (37–40). Furthermore, each fat depot includes numerous distinct progenitor fields that vary with age, gender, and environmental conditions. Additionally, scWAT depots are mainly derived from paired related homeobox 1 (PRX1) expressing progenitors (41–45). Despite the previous view that myogenic factor 5 (MYF5), paired box 7 (PAX7), and paired box 3 (PAX3) expressing progenitors only give rise to BAT, it is now believed that the scWAT depots of the dorsal–anterior body region originated partly from those progenitors (9, 39, 46–49). While ex vivo studies reported the presence of both PDGFRα and PDGFRβ in adipocyte stem cells (ASCc) (50), in adult mouse progenitors are heterogeneous and either express PDGFRα or PDGFRβ (51, 52). A recent study by Gao et al. suggested that the balance between PDGFRα and PDGFRβ determines whether progenitors will commit to beige (PDGFRα) or white (PDGFRβ) adipocytes (53). Some satellite cell-derived myoblasts in skeletal muscle and fibro-adipogenic progenitors (FAPs) also give rise to beige fat with higher rate of glycolysis and hence, named as glycolytic beige adipocyte (54).
Transcriptional Regulation of Brown and Beige Adipocytes
In brown and beige adipocytes, the adipogenic differentiation program and the thermogenic program are regulated by different pathways each involving a complex network of transcription factors (TFs) and epigenetic factors. The main parts of transcriptional machinery regulating fat cell differentiation is common among various types of fat cells and has been extensively discussed elsewhere (8, 55–57). Here, several TFs which are the main signatures of brown and beige fat are briefly described.
EBF2
Early beta Cell factor 2 is a marker of committed brown adipocytes in rodents and humans (29, 57, 58). EBF2 promotes brown adipocytes differentiation by recruiting peroxisome proliferator-activated receptor gamma (PPARγ) to its brown-specific chromatin regions (58). Overexpression of EBF2 in myoblast induces brown adipogenesis and inhibits myogenesis (30). In the absence of EBF2, the brown fat specific features of BAT are abolished (58). Moreover, overexpression of EBF2 in WAT induces browning and thermogenesis (59).
PRDM16
PR domain zinc finger 16 promotes brown and beige adipocyte differentiation and inhibits myogenic and WAT gene expression in mice and in vitro in human fibroblasts (46, 60–63). PRDM16 is also important in brown fat maintenance by binding to specific enhancer regions along with a mediator complex to establish enhancer-promoter looping, leading to the expression of PPARα and PPARγ co-activator 1A (PGC1α). Also, PRDM16 interacts with PGC1α (described below) and increases its transcription (60, 64, 65). PRDM16 also inhibits the signaling of repressor type 1 interferon response genes thereby preventing mitochondrial dysfunction and a decrease in UCP1 levels (66). PRDM16 overexpression increases beige adipogenesis and thermogenesis in WAT and its deficiency inhibits beige adipocyte formation (67, 68).
PGC1α
PPARγ co-activator 1A is known to directly interact with PRDM16 and PPARγ in brown adipocytes (60, 69). In differentiated brown and beige adipocytes, PGC1α plays a crucial role in cold-induced thermogenesis and thermogenic maintenance in mice and human WAT (69, 70). Interacting with other transcriptional regulators, PGC1α activates the transcription of UCP1 and several mitochondrial genes (65, 71, 72). PGC1α overexpression induces thermogenesis in adipocytes and myocytes (73, 74). Brown adipocytes lacking PGC1α express lower levels of UCP1 in response to adrenergic stimuli (75, 76). PGC1α is also required for the browning of WAT (77).
IRF4
Interferon regulatory factor 4 interacts with PGC1α upon cold stimuli and regulates UCP1 expression by binding to its chromatin regulatory regions in BAT of mice and humans (78).
ZFP516
Zinc finger protein 516 also increases brown adipogenesis and thermogenesis upon cold induction. ZFP516 interacts with PRDM16 and activates UCP1 and PGC1α expression (79).
CREB-ATF2
Phosphorylation and activation of activating transcription factor 2 (ATF2) and cAMP-responsive element-binding (CREB) downstream of cold-induced adrenergic signaling results in activation of UCP1 and PGC1α gene expression (80).
KLF11
Kruppel-like factor 11 expression is induced in vitro in human white adipocytes in response to the PPARγ agonist rosiglitazone and promotes beige adipocyte-selective gene expression via increasing PPARγ binding to beige-selective super-enhancers (81).
FOXC2
Forkhead box protein C2 expression increases beige adipocyte formation by promoting protein kinase A (PKA) activity; a main kinase activated downstream of the adrenergic pathway upon cold induction (82).
FOXP1
Foxhead P1 expression is highly enriched in vWAT and acts as a transcriptional repressor that directly represses β3-AR transcription. FOXP1 deletion increases brown fat activation and the browning of WAT and its overexpression is inhibitory to browning and thermogenesis in mice and humans (83).
GABPα
GA-binding protein α is expressed in myoblasts and inhibits myogenesis and promotes adipogenesis and beige fat development. The interaction between PGC1α and GABPα is also shown to stimulate mitochondria biogenesis and oxidative phosphorylation (84–86). GABPα expressing beige adipocytes are unlike other beige adipocytes with higher glucose oxidation rate than fatty acid oxidation (54).
EBF2 and GABPα mainly function as commitment factors. EBF2 marks committed brown adipocytes and GABPα inhibits myogenic development thereby promoting brown/beige fat formation. During brown fat differentiation, EBF2, PRDM16, and ZFP516 together with other common adipogenic regulators such as C/EBPβ and PPARγ regulate the induction of brown fat specific genes with PRDM16 acting mainly as a coactivator. Upon cold exposure and activation of adrenergic signaling, the concentration of cyclic AMP (cAMP) and PKA activity increase. PKA activity, which is further increased by FOXC2, results in p38 MAPK phosphorylation and activation. Phosphorylated p38 then phosphorylates and activates both ATF2 and PGC1α. IRF4 recruits phosphorylated PGC1α to the chromatin which will then coactivate PPARγ, thyroid receptor (TR), and retinoid X receptor alpha (RXRα) to increase the expression of thermogenic genes.
Several other transcriptional regulators and nuclear receptors are known to be involved in activating or repressing brown and beige fat specific programs (87). The roles of long non-coding RNA and microRNA in brown/beige fat regulation are comprehensively discussed elsewhere (88, 89).
Epigenetic Regulation of Brown and Beige Adipocytes
The chromatin landscape which plays a critical role in brown/ beige fat identity, differentiation, and activation is modulated via tight cooperation between TFs and epigenetic modifiers. ChIP-seq analysis showed that PPARγ bind to 55% similar regions in BAT, scWAT, and vWAT. However, only 10% of PPARγ binding sites are BAT specific (90). In a separate study in human adipocytes, PPARγ ChIP-seq before and after rosiglitazone induced browning identified only 10% of the binding sites were different, indicating that the beige and BAT-selective characteristics are derived from a small subset of genomic sites (81, 90). Nuclear isolation and chromatin analysis of specific cell types from heterogeneous adipose tissue using UCP1-NuTRAP mice demonstrated the stability of the chromatin landscape in BAT upon temperature changes while extreme plasticity of chromatin landscape was observed in beige adipocytes with temperature changes (91).
Enhancers in BAT, but not WAT are enriched in active histone marks such as H3K4me1/2 and H3K27ac (92). The UCP1 promoter in BAT is enriched for H3K4me3 and in WAT is enriched for H3K27me3 (93). Several histone-modifying enzymes have been identified that regulate the chromatin landscape in brown fat (94, 95). Some examples are ubiquitously transcribed tetratricopeptide repeat X chromosome (UTX), lysine-specific histone demethylase 1 (LSD1), jumonji domain containing 3 (JMJD3), and jumonji domain containing 1A/lysine demethylase 3A (JMJD1A/KDM3A) demethylate H3K27me3 at the promoters of UCP1 which leads to the upregulation of BAT selective genes in response to acute cold exposure (93, 96–98). Deletion of myeloid/lymphoid or mixed-lineage leukemia 4/lysine demethylase 2D (MLL4/KMT2D) in brown precursors increases the level of repressor H3K27me3 marks and impairs brown adipogenesis in mice and humans (99). Ablation of euchromatic histone-lysine N-methyltransferase 1 (EHMT1) decreases activator H3K4me3 marks and impairs brown adipocyte differentiation thereby activating myogenesis (62). The depletion of histone deacetylase 3 (HDAC3) in mice decreases thermogenesis in cold temperatures (100). In general, cold induction in brown adipocytes increases the expression of activator H3K27ac marks and removes the repressive H3K27me3 marks (101).
Browning: Paths and Players
The increase of UCP1 positive, multilocular, thermogenic beige adipocytes within WAT (browning) is a potential therapeutic approach to increase insulin sensitivity and combat metabolic diseases such as obesity (102, 103). In mammals all browning features can be achieved by adrenergic stimulation, the main signaling pathway of thermogenic BAT which is induced by cold temperatures. In addition, several alternatives to this canonical pathway have been reported to regulate browning of WAT through interorgan crosstalk (104). Several browning agents have been reported that are extensively reviewed elsewhere (20, 105–107) (Figure 1). For example, numerous pharmacological small molecules, dietary compounds, and nutritional agents are known to increase WAT browning (108–111). Additionally, various organs respond to environmental challenges such as cold, fasting, feeding, and exercise by secreting several factors and hormones that contribute to the browning (3, 112). Gut microbiota as well as immune cells and macrophages influence WAT browning process and have been well-discussed by others (113–115). In mammals, increased energy expenditure and browning of WAT after gastric bypass surgery have been reported (116–118). The link between WAT browning and thermogenesis is supported by generic mouse models of UCP1 knockout and BAT paucity, both leading to compensatory browning of WAT (119, 120).
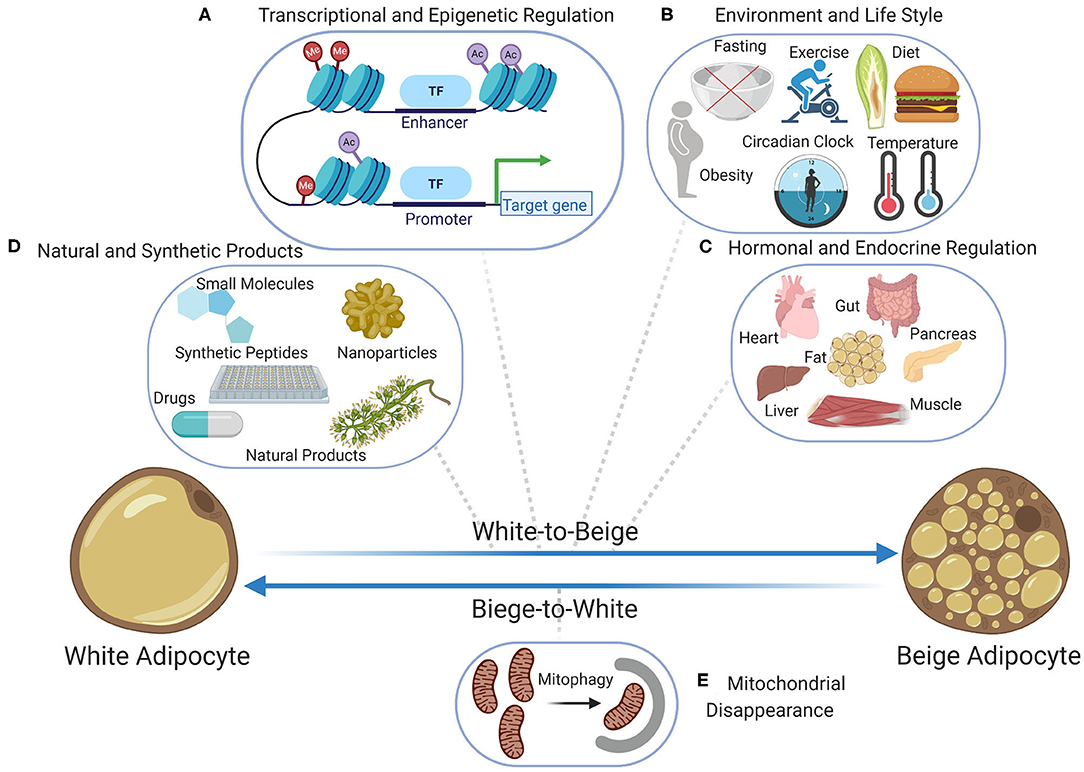
Figure 1. Bidirectional transition between beige and white adipocytes; the beige-to-white transition (browning) has been extensively studied at the levels of (A) transcriptional and epigenetic regulation including the chromatin landscape, transcriptional regulators, and epigenetic modifiers, (B) the role of lifestyle and environment including diet, fasting, obesity, exercise, temperature, and circadian rhythm, (C) the role of endocrine factors and hormones secreted by various organs including pancreas, muscle, liver, heart, gut, and fat when adapting to environmental challenges, (D) the role of natural products and plant extracts as well as the role of synthetic chemical products including small molecules, nanoparticles, synthetic peptides, and drug. Contrarily, the beige-to-white transition which is the immediate result of stimuli removal is poorly investigated and so far, (E) mitochondrial disappearance (mitophagy) is known to be the main contributor. Figure created with ©BioRender.io.
Beige adipocytes form via three main processes (121): (I) proliferation and de novo differentiation of beige adipocytes from the progenitor pool located in adipose vasculature mural cells and express smooth muscle actin (SMA), myosin-11 (MYH-11), and PDGFRα (122, 123). In addition to adipose tissue vasculature, smooth muscle cells are also proposed as a source of beige progenitors (124); (II) transdifferentiation of mature white adipocytes to beige adipocytes (12, 125) where adrenergic stimulation by cold and high-fat diet feeding increases de novo formation of beige adipocytes as well transdifferentiation of mature white cells into beige adipocytes (16, 51, 126, 127); (III) the activation of dormant beige adipocytes without the involvement of progenitors contributes to the formation of thermogenic beige fat (12, 128, 129). These competing hypotheses remain unresolved as current lineage tracing technologies are unable to distinguish between white-to-beige adipocyte transdifferentiation and the activation of dormant beige cells. The current strategies for lineage tracing are summarized by Sebo and Rodeheffer (129).
WAT Browning in Humans
Recent studies that carried out FDG-PET/CT imaging were able to identify BAT activity in supraclavicular, cervical, axillary, and less often in abdominal, mediastinal, and paraspinal fat depots. The UCP1 positive fat depots in adult humans detected as 18F-FDG positive contained both classical brown adipocytes as well as beige adipocytes (6, 11, 24, 26, 130–135). Beige and brown fat activity in humans is increased in response to cold exposure and is inversely associated with age, body mass index, and the levels of circulating lipid and glucose (24, 25, 136–141). Studies have shown that even in lean people with larger BAT depot, the cold induced thermogenic function of BAT does not significantly impact energy balance (142, 143). Hence, targeting the large scWAT for browning to increase thermogenesis has recently become a target for therapeutic approaches. So far, the browning of WAT in humans has only been reported under extreme conditions and its contribution to energy expenditure compared to BAT is minor (144–146). Ten days of cold exposure in humans was insufficient to induce WAT browning, despite increasing BAT activity. This indicates a requirement for higher levels of adrenergic stimuli (147, 148). The effects of prolonged physical training on human scWAT browning and increasing the levels of circulating adiponectin, apelin, irisin, and FGF21 are in line with improved metabolic health (145, 149). Additionally, severe weight loss in cancer patients as well as in obese patients going through weight loss surgeries leads to increased browning (150, 151). Prolonged elevations in norepinephrine levels as a consequence of burn injury also leads to increased scWAT browning and thermogenesis (152). Surgical trauma, not necessarily related to the incision, is also linked to local and distal WAT browning in humans (153).
Beige Reversibility and Maintenance
The thermogenic phenotype of beige adipocytes is reversible upon withdrawal of the external stimuli. Upon removal of adrenergic stimulation (for example in warm temperature), beige fat will gradually convert into cells with a unilocular lipid droplet and will progressively lose beige characteristics while increasing the white characteristics (e.g., reduced mitochondria and thermogenesis). This beige-to-white transformation is accompanied by reduced innervation, vasculature, and UCP1 expression and increased neural chemorepellent (semaphorin III) secretion and leptin expression (154–156). Although beige phenotype reversibility seems like a recent hallmark of adipocyte plasticity, it has been reported for decades (157–160). The phenotypic and morphological conversion found in beige fat upon withdrawal of stimuli is not observed in classical brown adipocytes (46). In 2013, Christian Wolfrum's research team used lineage tracing to validate beige-white interconversion. They showed that the cold-induced beige fat was reversed within 5 weeks of warm temperature and almost 75% of the whitened beige adipocyte could become beige again upon cold induction. Interestingly, after a second cold exposure, half of the beige adipocytes were formed from the former whitened beige adipocytes and the other half of the newly formed beige adipocytes seemed to come from a different source (12). Though the beige adipocytes lost their brown-like phenotype and acquired a white-like phenotype when the temperature was increased, they kept their epigenetic memory of the cold exposure which allowed them to activate browning genes as soon as they were exposed to cold temperatures (91). Interestingly, beige fat apoptosis and death was not found to be the cause of beige phenotype loss (12). Contrarily, BAT whitening was shown to increase cell death by increasing adipose inflammation, indicating a lack of plasticity in BAT (161). In 2015, Kozak and his research team reported much higher dynamics in UCP1 and mitochondrial turnover in beige fat when compared to BAT (162). In 2016, Kajimura and his research team elegantly linked the beige-to-white transition to mitochondrial disappearance (mitophagy). Mitophagy increased upon adrenergic stimuli withdrawal and was shown to be mediated by parkin (PARK2) recruitment to the mitochondria. Inhibition of autophagy via deletion of autophagy-related 5 (ATG5), autophagy-related 12 (ATG12), and PARK2 maintained the beige phenotype after stimuli removal. By monitoring single-cells, the same study also observed direct transdifferentiation from beige-to-WAT which did not involve an intermediate step (163). Recently, a natural and more stable beige adipose depot called thigh adipose tissue (tAT) was identified in mice (164). In contrast to classic beige adipocytes, tAT seems rather stable and maintains a beige fat phenotype in warm temperatures. However, high-fat diet (HFD) feeding and aging increased the white phenotypic features of tAT including the presence of unilocular adipocytes. Browning stimuli can increase brown adipocyte gene expression in tAT to a higher level than in iWAT. Furthermore, tAT has a higher rate of energy expenditure and lower expression of inflammatory genes relative to iWAT (164).
Future Directions and Perspectives
With the worldwide increase in obesity and its comorbidities, many studies have indicated the great potential of beige fat to increase energy expenditure. The main bottleneck in the use of beige adipocytes as a therapeutic target is the fact that the beige adipocytes transition to a white phenotypic state shortly after stimuli withdrawal. Studies in mice have shown that obesity increases the beige-to-white adipocyte transition (165, 166). The instability of beige adipocyte compared to classic BAT could be partly explained by differences in their developmental origins. Chromatin and epigenetic analysis during cold to warm thermal shifts uncovered that the chromatin landscape is stable in BAT while in beige adipocytes chromatin changes are transient (91). Currently mitophagy is thought to be the main contributor to beige-to-white transition (Figure 1), and a higher rate of mitochondrial biogenesis or a lower rate of mitophagy could explain BAT stability when compared to beige adipocytes. Understanding the underlying mechanism of thermogenic maintenance in BAT and tAT after external stimuli removal may elucidate the pathways that prevent the beige-to-white transition. In this regard, strategies to inhibit mitochondrial autophagy as well as to increase mitochondrial biogenesis are potential therapeutic prospects to prolong the thermogenic phenotype of beige adipocytes. Beige adipocyte maintenance has the potential to attenuate reoccurring weight gain after weight loss surgery, diet, and exercise.
Author Contributions
The author confirms being the sole contributor of this work and has approved it for publication.
Funding
AR was supported by NNF16OC0018642 from the Novo Nordisk Foundation and the Stanford Bio-X Program.
Conflict of Interest
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
AR would like to thank Kyle Murphy Kovary and Carl Takeshi Johnson (Stanford University) for proofreading this manuscript.
References
1. Seale P, Lazar MA. Brown fat in humans: turning up the heat on obesity. Diabetes. (2009) 58:1482–4. doi: 10.2337/db09-0622
2. Hanssen MJW, Van Der Lans AAJJ, Brans B, Hoeks J, Jardon KMC, Schaart G, et al. Short-term cold acclimation recruits brown adipose tissue in obese humans. Diabetes. (2016) 65:1179–89. doi: 10.2337/db15-1372
3. Bartelt A, Heeren J. Adipose tissue browning and metabolic health. Nat Rev Endocrinol. (2014) 10:24–36. doi: 10.1038/nrendo.2013.204
4. Trayhurn P. Recruiting brown adipose tissue in human obesity. Diabetes. (2016) 65:1158–60. doi: 10.2337/dbi16-0002
5. Devlin MJ. The “skinny” on brown fat, obesity, and bone. Am J Phys Anthropol. (2015) 156:98–115. doi: 10.1002/ajpa.22661
6. Leitner BP, Huang S, Brychta RJ, Duckworth CJ, Baskin AS, McGehee S, et al. Mapping of human brown adipose tissue in lean and obese young men. Proc Natl Acad Sci USA. (2017) 114:8649–54. doi: 10.1073/pnas.1705287114
7. Cinti S. The adipose organ. In: Prostaglandins Leukotrienes and Essential Fatty Acids. (2005). p. 9–15.
8. Rosen ED, Spiegelman BM. What we talk about when we talk about fat. Cell. (2014) 156:20–44. doi: 10.1016/j.cell.2013.12.012
9. Peirce V, Carobbio S, Vidal-Puig A. The different shades of fat. Nature. (2014) 510:76–83. doi: 10.1038/nature13477
10. Wu J, Boström P, Sparks LM, Ye L, Choi JH, Giang AH, et al. Beige adipocytes are a distinct type of thermogenic fat cell in mouse and human. Cell. (2012) 150:366–76. doi: 10.1016/j.cell.2012.05.016
11. Lizcano F. The beige adipocyte as a therapy for metabolic diseases. Int J Mol Sci. (2019) 20:5058. doi: 10.3390/ijms20205058
12. Rosenwald M, Perdikari A, Rülicke T, Wolfrum C. Bi-directional interconversion of brite and white adipocytes. Nat Cell Biol. (2013) 15:659–67. doi: 10.1038/ncb2740
13. Klingenspor M, Fromme T, Hughes DA, Manzke L, Polymeropoulos E, Riemann T, et al. An ancient look at UCP1. Biochim Biophys Acta. (2008) 1777:637–41. doi: 10.1016/j.bbabio.2008.03.006
14. Cousin B, Cinti S, Morroni M, Raimbault S, Ricquier D, Penicaud L, et al. Occurrence of brown adipocytes in rat white adipose tissue: molecular and morphological characterization. J Cell Sci. (1992) 103:931–42.
15. Guerra C, Koza RA, Yamashita H, Walsh K, Kozak LP. Emergence of brown adipocytes in white fat in mice is under genetic control effects on body weight and adiposity. J Clin Invest. (1998) 102:412–20. doi: 10.1172/JCI3155
16. Himms-Hagen J, Melnyk A, Zingaretti MC, Ceresi E, Barbatelli G, Cinti S. Multilocular fat cells in WAT of CL-316243-treated rats derive directly from white adipocytes. Am J Physiol. (2000) 279:C670–81. doi: 10.1152/ajpcell.2000.279.3.C670
17. Chevalier C, Stojanović O, Colin DJ, Suarez-Zamorano N, Tarallo V, Veyrat-Durebex C, et al. Gut microbiota orchestrates energy homeostasis during cold. Cell. (2015) 163:1360–74. doi: 10.1016/j.cell.2015.11.004
18. Suárez-Zamorano N, Fabbiano S, Chevalier C, Stojanović O, Colin DJ, Stevanović A, et al. Microbiota depletion promotes browning of white adipose tissue and reduces obesity. Nat Med. (2015) 21:1497–501. doi: 10.1038/nm.3994
19. Fabbiano S, Suárez-Zamorano N, Rigo D, Veyrat-Durebex C, Stevanovic Dokic A, Colin DJ, et al. Caloric restriction leads to browning of white adipose tissue through type 2 immune signaling. Cell Metab. (2016) 24:434–46. doi: 10.1016/j.cmet.2016.07.023
20. Chondronikola M, Sidossis LS. Brown and beige fat: from molecules to physiology. Biochim Biophys Acta Mol Cell Biol Lipids. (2019) 1864:91–103. doi: 10.1016/j.bbalip.2018.05.014
21. Tran TT, Kahn CR. Transplantation of adipose tissue and stem cells: role in metabolism and disease. Nat Rev Endocrinol. (2010) 6:195–213. doi: 10.1038/nrendo.2010.20
22. de Jong JMA, Larsson O, Cannon B, Nedergaard J. A stringent validation of mouse adipose tissue identity markers. Am J Physiol. (2015) 308:E1085–105. doi: 10.1152/ajpendo.00023.2015
23. Enerbäck S. Brown adipose tissue in humans. Int J Obes. (2010) 34:S43–6. doi: 10.1038/ijo.2010.183
24. Virtanen KA, Lidell ME, Orava J, Heglind M, Westergren R, Niemi T, et al. Functional brown adipose tissue in healthy adults. N Engl J Med. (2009) 360:1518–25. doi: 10.1056/NEJMoa0808949
25. Van Marken Lichtenbelt WD, Vanhommerig JW, Smulders NM, Drossaerts JMAFL, Kemerink GJ, et al. Cold-activated brown adipose tissue in healthy men. N Engl J Med. (2009) 360:1500–8. doi: 10.1056/NEJMoa0808718
26. Cypess AM, Lehman S, Williams G, Tal I, Rodman D, Goldfine AB, et al. Identification and importance of brown adipose tissue in adult humans. N Engl J Med. (2009) 360:1509–17. doi: 10.1056/NEJMoa0810780
27. Richard MA, Pallubinsky H, Blondin DP. Functional characterization of human brown adipose tissue metabolism. Biochem J. (2020) 477:1261–86. doi: 10.1042/BCJ20190464
28. Poissonnet CM, Burdi AR, Garn SM. The chronology of adipose tissue appearance and distribution in the human fetus. Early Hum Dev. (1984) 10:1–11. doi: 10.1016/0378-3782(84)90106-3
29. Houštěk J, Kopecký J, Rychter Z, Soukup T. Uncoupling protein in embryonic brown adipose tissue - existence of nonthermogenic and thermogenic mitochondria. BBA Bioenerg. (1988) 935:19–25. doi: 10.1016/0005-2728(88)90103-X
30. Wang W, Kissig M, Rajakumari S, Huang L, Lim HW, Won KJ, et al. Ebf2 is a selective marker of brown and beige adipogenic precursor cells. Proc Natl Acad Sci USA. (2014) 111:14466–71. doi: 10.1073/pnas.1412685111
31. Giralt M, Martin I, Iglesias R, Viñas O, Villarroya F, Mampel T. Ontogeny and perinatal modulation of gene expression in rat brown adipose tissue: unaltered iodothyronine 5′-deiodinase activity is necessary for the response to environmental temperature at birth. Eur J Biochem. (1990) 193:297–302. doi: 10.1111/j.1432-1033.1990.tb19336.x
32. Birsoy K, Berry R, Wang T, Ceyhan O, Tavazoie S, Friedman JM, et al. Analysis of gene networks in white adipose tissue development reveals a role for ETS2 in adipogenesis. J Cell Sci. (2011) 124:e1. doi: 10.1242/jcs.101436
33. Berry R, Rodeheffer MS. Characterization of the adipocyte cellular lineage in vivo. Nat Cell Biol. (2013) 15:302–8. doi: 10.1038/ncb2696
34. Han J, Lee JE, Jin J, Lim JS, Oh N, Kim K, et al. The spatiotemporal development of adipose tissue. Development. (2011) 138:5027–37. doi: 10.1242/dev.067686
35. Wang QA, Tao C, Gupta RK, Scherer PE. Tracking adipogenesis during white adipose tissue development, expansion and regeneration. Nat Med. (2013) 19:1338–44. doi: 10.1038/nm.3324
36. Atit R, Sgaier SK, Mohamed OA, Taketo MM, Dufort D, Joyner AL, et al. β-Catenin activation is necessary and sufficient to specify the dorsal dermal fate in the mouse. Dev Biol. (2006) 296:164–76. doi: 10.1016/j.ydbio.2006.04.449
37. Billon N, Iannarelli P, Monteiro MC, Glaviuex-Pardanaud C, Richardson WD, Kessaris N, et al. The generation of adipocytes by the neural crest. Development. (2007) 134:2283–92. doi: 10.1242/dev.002642
38. Lemos DR, Paylor B, Chang C, Sampaio A, Underhill TM, Rossi FMV. Functionally convergent white adipogenic progenitors of different lineages participate in a diffused system supporting tissue regeneration. Stem Cells. (2012) 30:1152–62. doi: 10.1002/stem.1082
39. Sanchez-Gurmaches J, Guertin DA. Adipocytes arise from multiple lineages that are heterogeneously and dynamically distributed. Nat Commun. (2014) 5:4099. doi: 10.1038/ncomms5099
40. Le Lievre CS, Le Douarin NM. Mesenchymal derivatives of the neural crest: analysis of chimaeric quail and chick embryos. J Embryol Exp Morphol. (1975) 34:125–54.
41. Sanchez-Gurmaches J, Hsiao WY, Guertin DA. Highly selective in vivo labeling of subcutaneous white adipocyte precursors with Prx1-Cre. Stem Cell Rep. (2015) 4:541–50. doi: 10.1016/j.stemcr.2015.02.008
42. Bergwerff M, Gittenberger-de Groot AC, Wisse LJ, DeRuiter MC, Wessels A, Martin JF, et al. Loss of function of the Prx1 and Prx2 hameobox genes alters architecture of the great elastic arteries and ductus arteriosus. Virchows Arch. (2000) 436:12–9. doi: 10.1007/PL00008193
43. Kuratani S, Martin JF, Wawersik S, Lilly B, Eichele G, Olson EN. The expression pattern of the chick homeobox gene gMHox suggests a role in patterning of the limbs and face and in compartmentalization of somites. Dev Biol. (1994) 161:357–69. doi: 10.1006/dbio.1994.1037
44. Leussink B, Brouwer A, El Khattabi M, Poelmann RE, Gittenberger-de Groot AC, Meijlink F. Expression patterns of the paired-related homeobox genes MHox Prx1 and S8 Prx2 suggest roles in development of the heart and the forebrain. Mech Dev. (1995) 52:51–64. doi: 10.1016/0925-4773(95)00389-I
45. Shimozaki K, Clemenson GD Jr, Gage FH. Erratum to paired related homeobox protein 1 is a regulator of stemness in adult neural stem/progenitor cells. J Neurosci. (2015) 35:1335. doi: 10.1523/JNEUROSCI.4753-14.2015
46. Seale P, Bjork B, Yang W, Kajimura S, Chin S, Kuang S, et al. PRDM16 controls a brown fat/skeletal muscle switch. Nature. (2008) 454:961–7. doi: 10.1038/nature07182
47. Sanchez-Gurmaches J, Hung CM, Sparks CA, Tang Y, Li H, Guertin DA. PTEN loss in the Myf5 lineage redistributes body fat and reveals subsets of white adipocytes that arise from Myf5 precursors. Cell Metab. (2012) 16:348–62. doi: 10.1016/j.cmet.2012.08.003
48. Shan T, Liang X, Bi P, Zhang P, Liu W, Kuang S. Distinct populations of adipogenic and myogenic Myf5-lineage progenitors in white adipose tissues. J Lipid Res. (2013) 54:2214–24. doi: 10.1194/jlr.M038711
49. Sebo ZL, Jeffery E, Holtrup B, Rodeheffer MS. A mesodermal fate map for adipose tissue. Development. (2018) 145:dev166801. doi: 10.1242/dev.166801
50. Traktuev DO, Merfeld-Clauss S, Li J, Kolonin M, Arap W, Pasqualini R, et al. A population of multipotent CD34-positive adipose stromal cells share pericyte and mesenchymal surface markers, reside in a periendothelial location, and stabilize endothelial networks. Circ Res. (2008) 102:77–85. doi: 10.1161/CIRCRESAHA.107.159475
51. Lee YH, Petkova AP, Mottillo EP, Granneman JG. In vivo identification of bipotential adipocyte progenitors recruited by β3-adrenoceptor activation and high-fat feeding. Cell Metab. (2012) 15:480–91. doi: 10.1016/j.cmet.2012.03.009
52. Daquinag AC, Tseng C, Salameh A, Zhang Y, Amaya-Manzanares F, Dadbin A, et al. Depletion of white adipocyte progenitors induces beige adipocyte differentiation and suppresses obesity development. Cell Death Differ. (2015) 22:351–63. doi: 10.1038/cdd.2014.148
53. Gao Z, Daquinag AC, Su F, Snyder B, Kolonin MG. PDGFRα/PDGFRβ signaling balance modulates progenitor cell differentiation into white and beige adipocytes. Development. (2018) 145:dev155861. doi: 10.1242/dev.155861
54. Chen Y, Ikeda K, Yoneshiro T, Scaramozza A, Tajima K, Wang Q, et al. Thermal stress induces glycolytic beige fat formation via a myogenic state. Nature. (2019) 565:180–5. doi: 10.1038/s41586-018-0801-z
55. Farmer SR. Transcriptional control of adipocyte formation. Cell Metab. (2006) 4:263–73. doi: 10.1016/j.cmet.2006.07.001
56. Rangwala SM, Lazar MA. Transcriptional control of adipogenesis. Annu Rev Nutr. (2000) 20:535–59. doi: 10.1146/annurev.nutr.20.1.535
57. Siersbæk R, Mandrup S. Transcriptional networks controlling adipocyte differentiation. Cold Spring Harb Symp Quant Biol. (2011) 76:247–55. doi: 10.1101/sqb.2011.76.010512
58. Rajakumari S, Wu J, Ishibashi J, Lim HW, Giang AH, Won KJ, et al. EBF2 determines and maintains brown adipocyte identity. Cell Metab. (2013) 17:562–74. doi: 10.1016/j.cmet.2013.01.015
59. Stine RR, Shapira SN, Lim HW, Ishibashi J, Harms M, Won KJ, et al. EBF2 promotes the recruitment of beige adipocytes in white adipose tissue. Mol Metab. (2016) 5:57–65. doi: 10.1016/j.molmet.2015.11.001
60. Seale P, Kajimura S, Yang W, Chin S, Rohas LM, Uldry M, et al. Transcriptional Control of Brown Fat Determination by PRDM16. Cell Metab. (2007) 6:38–54. doi: 10.1016/j.cmet.2007.06.001
61. Kajimura S, Seale P, Kubota K, Lunsford E, Frangioni JV, Gygi SP, et al. Initiation of myoblast to brown fat switch by a PRDM16-C/EBP-β transcriptional complex. Nature. (2009) 460:1154–8. doi: 10.1038/nature08262
62. Ohno H, Shinoda K, Ohyama K, Sharp LZ, Kajimura S. EHMT1 controls brown adipose cell fate and thermogenesis through the PRDM16 complex. Nature. (2013) 504:163–7. doi: 10.1038/nature12652
63. Kajimura S, Seale P, Tomaru T, Erdjument-Bromage H, Cooper MP, Ruas JL, et al. Regulation of the brown and white fat gene programs through a PRDM16/CtBP transcriptional complex. Genes Dev. (2008) 22:1397–409. doi: 10.1101/gad.1666108
64. Harms MJ, Ishibashi J, Wang W, Lim HW, Goyama S, Sato T, et al. Prdm16 is required for the maintenance of brown adipocyte identity and function in adult mice. Cell Metab. (2014) 19:593–604. doi: 10.1016/j.cmet.2014.03.007
65. Iida S, Chen W, Nakadai T, Ohkuma Y, Roeder RG. PRDM16 enhances nuclear receptordependent transcription of the brown fat-specific Ucp1 gene through interactions with mediator subunit MED1. Genes Dev. (2015) 29:308–21. doi: 10.1101/gad.252809.114
66. Kissig M, Ishibashi J, Harms MJ, Lim H, Stine RR, Won K, et al. PRDM16 represses the type I interferon response in adipocytes to promote mitochondrial and thermogenic programing. EMBO J. (2017) 36:1528–42. doi: 10.15252/embj.201695588
67. Seale P, Conroe HM, Estall J, Kajimura S, Frontini A, Ishibashi J, et al. Prdm16 determines the thermogenic program of subcutaneous white adipose tissue in mice. J Clin Invest. (2011) 121:96–105. doi: 10.1172/JCI44271
68. Cohen P, Levy JD, Zhang Y, Frontini A, Kolodin DP, Svensson KJ, et al. Ablation of PRDM16 and beige adipose causes metabolic dysfunction and a subcutaneous to visceral fat switch. Cell. (2014) 156:304–16. doi: 10.1016/j.cell.2013.12.021
69. Puigserver P, Wu Z, Park CW, Graves R, Wright M, Spiegelman BM. A cold-inducible coactivator of nuclear receptors linked to adaptive thermogenesis. Cell. (1998) 92:829–39. doi: 10.1016/S0092-8674(00)81410-5
70. Tiraby C, Tavernier G, Lefort C, Larrouy D, Bouillaud F, Ricquier D, et al. Acquirement of brown fat cell features by human white adipocytes. J Biol Chem. (2003) 278:33370–6. doi: 10.1074/jbc.M305235200
71. Chen W, Yang Q, Roeder RG. Dynamic interactions and cooperative functions of PGC-1α and MED1 in TRα-mediated activation of the brown-fat-specific UCP-1 gene. Mol Cell. (2009) 35:755–68. doi: 10.1016/j.molcel.2009.09.015
72. Lin J, Handschin C, Spiegelman BM. Metabolic control through the PGC-1 family of transcription coactivators. Cell Metab. (2005) 1:361–70. doi: 10.1016/j.cmet.2005.05.004
73. Wu Z, Puigserver P, Andersson U, Zhang C, Adelmant G, Mootha V, et al. Mechanisms controlling mitochondrial biogenesis and respiration through the thermogenic coactivator PGC-1. Cell. (1999) 98:115–24. doi: 10.1016/S0092-8674(00)80611-X
74. Lehman JJ, Barger PM, Kovacs A, Saffitz JE, Medeiros DM, Kelly DP. Peroxisome proliferator-activated receptor γ coactivator-1 promotes cardiac mitochondrial biogenesis. J Clin Invest. (2000) 106:847–56. doi: 10.1172/JCI10268
75. Lin J, Wu PH, Tarr PT, Lindenberg KS, St-Pierre J, Zhang CY, et al. Defects in adaptive energy metabolism with CNS-linked hyperactivity in PGC-1α null mice. Cell. (2004) 119:121–35. doi: 10.1016/j.cell.2004.09.013
76. Uldry M, Yang W, St-Pierre J, Lin J, Seale P, Spiegelman BM. Complementary action of the PGC-1 coactivators in mitochondrial biogenesis and brown fat differentiation. Cell Metab. (2006) 3:333–41. doi: 10.1016/j.cmet.2006.04.002
77. Kleiner S, Mepani RJ, Laznik D, Ye L, Jurczak MJ, Jornayvaz FR, et al. Development of insulin resistance in mice lacking PGC-1α in adipose tissues. Proc Natl Acad Sci USA. (2012) 109:9635–40. doi: 10.1073/pnas.1207287109
78. Kong X, Banks A, Liu T, Kazak L, Rao RR, Cohen P, et al. IRF4 is a key thermogenic transcriptional partner of PGC-1α. Cell. (2014) 158:69–83. doi: 10.1016/j.cell.2014.04.049
79. Dempersmier J, Sambeat A, Gulyaeva O, Paul SM, Hudak CSS, Raposo HF, et al. Cold-inducible Zfp516 activates UCP1 transcription to promote browning of white fat and development of brown fat. Mol Cell. (2015) 57:235–46. doi: 10.1016/j.molcel.2014.12.005
80. Cao W, Daniel KW, Robidoux J, Puigserver P, Medvedev AV, Bai X, et al. p38 mitogen-activated protein kinase is the central regulator of cyclic AMP-dependent transcription of the brown fat uncoupling protein 1 gene. Mol Cell Biol. (2004) 24:3057–67. doi: 10.1128/MCB.24.7.3057-3067.2004
81. Loft A, Forss I, Siersbæk MS, Schmidt SF, Larsen ASB, Madsen JGS, et al. Browning of human adipocytes requires KLF11 and reprogramming of PPARγ superenhancers. Genes Dev. (2015) 29:7–22. doi: 10.1101/gad.250829.114
82. Cederberg A, Gronning LM, Ahrén B, Taskén K, Carlsson P, Enerbäck S. FOXC2 is a winged helix gene that counteracts obesity, hypertriglyceridemia, and diet-induced insulin resistance. Cell. (2001) 106:563–73. doi: 10.1016/S0092-8674(01)00474-3
83. Liu P, Huang S, Ling S, Xu S, Wang F, Zhang W, et al. Foxp1 controls brown/beige adipocyte differentiation and thermogenesis through regulating β3-AR desensitization. Nat Commun. (2019) 10:5070. doi: 10.1038/s41467-019-12988-8
84. Yang Z-F, Drumea K, Mott S, Wang J, Rosmarin AG. GABP transcription factor (nuclear respiratory factor 2) is required for mitochondrial biogenesis. Mol Cell Biol. (2014) 34:3194–201. doi: 10.1128/MCB.00492-12
85. Mootha VK, Handschin C, Arlow D, Xie X, St. Pierre J, Sihag S, et al. Errα and Gabpa/b specify PGC-1α-dependent oxidative phosphorylation gene expression that is altered in diabetic muscle. Proc Natl Acad Sci USA. (2004) 101:6570–5. doi: 10.1073/pnas.0401401101
86. Gantner ML, Hazen BC, Eury E, Brown EL, Kralli A. Complementary roles of estrogen-related receptors in brown adipocyte thermogenic function. Endocrinology. (2016) 157:4770–81. doi: 10.1210/en.2016-1767
87. Shapira SN, Seale P. Transcriptional control of brown and beige fat development and function. Obesity. (2019) 27:13–21. doi: 10.1002/oby.22334
88. Xu S, Chen P, Sun L. Regulatory networks of non-coding RNAs in brown/beige adipogenesis. Biosci Rep. (2015) 35:e00262. doi: 10.1042/BSR20150155
89. Chen Y, Pan R, Pfeifer A. Regulation of brown and beige fat by microRNAs. Pharmacol Ther. (2017) 170:1–7. doi: 10.1016/j.pharmthera.2016.10.004
90. Siersbaek MS, Loft A, Aagaard MM, Nielsen R, Schmidt SF, Petrovic N, et al. Genome-wide profiling of peroxisome proliferator-activated receptor in primary epididymal, inguinal, and brown adipocytes reveals depot-selective binding correlated with gene expression. Mol Cell Biol. (2012) 32:3452–63. doi: 10.1128/MCB.00526-12
91. Roh HC, Tsai LTY, Shao M, Tenen D, Shen Y, Kumari M, et al. Warming induces significant reprogramming of beige, but not brown, adipocyte cellular identity. Cell Metab. (2018) 27:1121–37.e5. doi: 10.1016/j.cmet.2018.03.005
92. Lai B, Lee JE, Jang Y, Lifeng W, Peng W, Ge K. MLL3/MLL4 are required for CBP/p300 binding on enhancers and super-enhancer formation in brown adipogenesis. Nucleic Acids Res. (2017) 45:6388–403. doi: 10.1093/nar/gkx234
93. Pan D, Huang L, Zhu LJ, Zou T, Ou J, Zhou W, et al. Jmjd3-mediated H3K27me3 dynamics orchestrate brown fat development and regulate white fat plasticity. Dev Cell. (2015) 35:568–83. doi: 10.1016/j.devcel.2015.11.002
94. Nic-Can GI, Rodas-Junco BA, Carrillo-Cocom LM, Zepeda-Pedreguera A, Peñaloza-Cuevas R, Aguilar-Ayala FJ, et al. Epigenetic regulation of adipogenic differentiation by histone lysine demethylation. Int J Mol Sci. (2019) 20:3918. doi: 10.3390/ijms20163918
95. Tanimura K, Suzuki T, Vargas D, Shibata H, Inagaki T. Epigenetic regulation of beige adipocyte fate by histone methylation. Endocr J. (2019) 66:115–25. doi: 10.1507/endocrj.EJ18-0442
96. Zha L, Li F, Wu R, Artinian L, Rehder V, Yu L, et al. The histone demethylase UTX promotes brown adipocyte thermogenic program via coordinated regulation of H3K27 demethylation and acetylation. J Biol Chem. (2015) 290:25151–63. doi: 10.1074/jbc.M115.662650
97. Sambeat A, Gulyaeva O, Dempersmier J, Tharp KM, Stahl A, Paul SM, et al. LSD1 Interacts with Zfp516 to promote UCP1 transcription and brown fat program. Cell Rep. (2016) 15:2536–49. doi: 10.1016/j.celrep.2016.05.019
98. Abe Y, Rozqie R, Matsumura Y, Kawamura T, Nakaki R, Tsurutani Y, et al. JMJD1A is a signal-sensing scaffold that regulates acute chromatin dynamics via SWI/SNF association for thermogenesis. Nat Commun. (2015) 6:7052. doi: 10.1038/ncomms8052
99. Lee JE, Wang C, Xu S, Cho YW, Wang L, Feng X, et al. H3K4 mono- and di-methyltransferase MLL4 is required for enhancer activation during cell differentiation. Elife. (2013) 2:e01503. doi: 10.7554/eLife.01503.027
100. Emmett MJ, Lim HW, Jager J, Richter HJ, Adlanmerini M, Peed LC, et al. Histone deacetylase 3 prepares brown adipose tissue for acute thermogenic challenge. Nature. (2017) 546:544–8. doi: 10.1038/nature22819
101. Li F, Wu R, Cui X, Zha L, Yu L, Shi H, et al. Histone deacetylase 1 (HDAC1) negatively regulates thermogenic program in brown adipocytes via coordinated regulation of histone H3 lysine 27 (H3K27) deacetylation and methylation. J Biol Chem. (2016) 291:4523–36. doi: 10.1074/jbc.M115.677930
102. Stanford KI, Middelbeek RJW, Townsend KL, An D, Nygaard EB, Hitchcox KM, et al. Brown adipose tissue regulates glucose homeostasis and insulin sensitivity. J Clin Invest. (2013) 123:215–23. doi: 10.1172/JCI62308
103. Hanssen MJW, Hoeks J, Brans B, Van Der Lans AAJJ, Schaart G, Van Den Driessche JJ, et al. Short-term cold acclimation improves insulin sensitivity in patients with type 2 diabetes mellitus. Nat Med. (2015) 21:863–5. doi: 10.1038/nm.3891
104. Chouchani ET, Kazak L, Spiegelman BM. New advances in adaptive thermogenesis: UCP1 and beyond. Cell Metab. (2019) 29:27–37. doi: 10.1016/j.cmet.2018.11.002
105. Kaisanlahti A, Glumoff T. Browning of white fat: agents and implications for beige adipose tissue to type 2 diabetes. J Physiol Biochem. (2019) 75:1–10. doi: 10.1007/s13105-018-0658-5
106. Lo KA, Sun L. Turning WAT into BAT: a review on regulators controlling the browning of white adipocytes. Biosci Rep. (2013) 33:711–9. doi: 10.1042/BSR20130046
107. Nedergaard J, Cannon B. The browning of white adipose tissue: some burning issues. Cell Metab. (2014) 20:396–407. doi: 10.1016/j.cmet.2014.07.005
108. El Hadi H, Di Vincenzo A, Vettor R, Rossato M. Food ingredients involved in white-to-brown adipose tissue conversion and in calorie burning. Front Physiol. (2019) 10:1954. doi: 10.3389/fphys.2018.01954
109. Bonet ML, Oliver P, Palou A. Pharmacological and nutritional agents promoting browning of white adipose tissue. Biochim Biophys Acta Mol Cell Biol Lipids. (2013) 1831:969–85. doi: 10.1016/j.bbalip.2012.12.002
110. Song NJ, Chang SH, Li DY, Villanueva CJ, Park KW. Induction of thermogenic adipocytes: molecular targets and thermogenic small molecules. Exp Mol Med. (2017) 49:e353. doi: 10.1038/emm.2017.70
111. Abdullahi A, Jeschke MG. Taming the flames: targeting white adipose tissue browning in hypermetabolic conditions. Endocr Rev. (2017) 38:538–49. doi: 10.1210/er.2017-00163
112. Chondronikola M, Porter C, Malagaris I, Nella AA, Sidossis LS. Brown adipose tissue is associated with systemic concentrations of peptides secreted from the gastrointestinal system and involved in appetite regulation. Eur J Endocrinol. (2017) 177:33–40. doi: 10.1530/EJE-16-0958
113. Moreno-Navarrete JM, Fernandez-Real JM. The gut microbiota modulates both browning of white adipose tissue and the activity of brown adipose tissue. Rev Endocr Metab Disord. (2019) 20:387–97. doi: 10.1007/s11154-019-09523-x
114. Villarroya F, Cereijo R, Villarroya J, Gavaldà-Navarro A, Giralt M. Toward an understanding of how immune cells control brown and beige adipobiology. Cell Metab. (2018) 27:954–61. doi: 10.1016/j.cmet.2018.04.006
115. Nguyen KD, Qiu Y, Cui X, Goh YPS, Mwangi J, David T, et al. Alternatively activated macrophages produce catecholamines to sustain adaptive thermogenesis. Nature. (2011) 480:104–8. doi: 10.1038/nature10653
116. Hankir MK, Bronisch F, Hintschich C, Krügel U, Seyfried F, Fenske WK. Differential effects of Roux-en-Y gastric bypass surgery on brown and beige adipose tissue thermogenesis. Metabolism. (2015) 64:1240–9. doi: 10.1016/j.metabol.2015.06.010
117. Hu J, Wang M, Zhou Y, Zhang X, He B, Liu L, et al. Bariatric surgery in rats upregulates FSP27 expression in fat tissue to affect fat hydrolysis and metabolism. Biomed Res Int. (2019) 8:1–11. doi: 10.1155/2019/6415732
118. Liu L, Zhang T, Hu J, Ma R, He B, Wang M, et al. Adiponectin/SIRT1 axis induces white adipose browning after vertical sleeve gastrectomy of obese rats with type 2 diabetes. Obes Surg. (2020) 30:1392–403. doi: 10.1007/s11695-019-04295-4
119. Meyer CW, Willershäuser M, Jastroch M, Rourke BC, Fromme T, Oelkrug R, et al. Adaptive thermogenesis and thermal conductance in wild-type and UCP1-KO mice. Am J Physiol. (2010) 299:R1396–406. doi: 10.1152/ajpregu.00021.2009
120. Schulz TJ, Huang P, Huang TL, Xue R, McDougall LE, Townsend KL, et al. Brown-fat paucity due to impaired BMP signalling induces compensatory browning of white fat. Nature. (2013) 495:379–83. doi: 10.1038/nature11943
121. Jopling C, Boue S, Belmonte JCI. Dedifferentiation, transdifferentiation and reprogramming: three routes to regeneration. Nat Rev Mol Cell Biol. (2011) 12:79–89. doi: 10.1038/nrm3043
122. Min SY, Kady J, Nam M, Rojas-Rodriguez R, Berkenwald A, Kim JH, et al. Human “brite/beige” adipocytes develop from capillary networks, and their implantation improves metabolic homeostasis in mice. Nat Med. (2016) 22:312–8. doi: 10.1038/nm.4031
123. Tran Van K, Gealekman O, Frontini A, Zingaretti MC, Morroni M, Giordano A, et al. The vascular endothelium of the adipose tissue gives rise to both white and brown fat cells. Cell Metab. (2012) 15:222–9. doi: 10.1016/j.cmet.2012.01.008
124. Long JZ, Svensson KJ, Tsai L, Zeng X, Roh HC, Kong X, et al. A smooth muscle-like origin for beige adipocytes. Cell Metab. (2014) 19:810–20. doi: 10.1016/j.cmet.2014.03.025
125. Barbatelli G, Murano I, Madsen L, Hao Q, Jimenez M, Kristiansen K, et al. The emergence of cold-induced brown adipocytes in mouse white fat depots is determined predominantly by white to brown adipocyte transdifferentiation. Am J Physiol. (2010) 298:E1244–53. doi: 10.1152/ajpendo.00600.2009
126. Lee YH, Petkova AP, Konkar AA, Granneman JG. Cellular origins of cold-induced brown adipocytes in adult mice. FASEB J. (2015) 29:286–99. doi: 10.1096/fj.14-263038
127. Vishvanath L, Macpherson KA, Hepler C, Wang QA, Shao M, Spurgin SB, et al. Pdgfrβ+ mural preadipocytes contribute to adipocyte hyperplasia induced by high-fat-diet feeding and prolonged cold exposure in adult mice. Cell Metab. (2016) 23:350–9. doi: 10.1016/j.cmet.2015.10.018
128. Shao M, Wang QA, Song A, Vishvanath L, Busbuso NC, Scherer PE, et al. Cellular origins of beige fat cells revisited. Diabetes. (2019) 68:1874–85. doi: 10.2337/db19-0308
129. Sebo ZL, Rodeheffer MS. Assembling the adipose organ: adipocyte lineage segregation and adipogenesis in vivo. Development. (2019) 146:dev172098 . doi: 10.1242/dev.172098
130. Nedergaard J, Bengtsson T, Cannon B. Unexpected evidence for active brown adipose tissue in adult humans. Am J Physiol. (2007) 293:E444–52. doi: 10.1152/ajpendo.00691.2006
131. Zingaretti MC, Crosta F, Vitali A, Guerrieri M, Frontini A, Cannon B, et al. The presence of UCP1 demonstrates that metabolically active adipose tissue in the neck of adult humans truly represents brown adipose tissue. FASEB J. (2009) 23:3113–20. doi: 10.1096/fj.09-133546
132. Cypess AM, White AP, Vernochet C, Schulz TJ, Xue R, Sass CA, et al. Anatomical localization, gene expression profiling and functional characterization of adult human neck brown fat. Nat Med. (2013) 19:635–9. doi: 10.1038/nm.3112
133. Lidell ME, Betz MJ, Leinhard OD, Heglind M, Elander L, Slawik M, et al. Evidence for two types of brown adipose tissue in humans. Nat Med. (2013) 19:631–4. doi: 10.1038/nm.3017
134. Jespersen NZ, Larsen TJ, Peijs L, Daugaard S, Homøe P, Loft A, et al. A classical brown adipose tissue mrna signature partly overlaps with brite in the supraclavicular region of adult humans. Cell Metab. (2013) 17:798–805. doi: 10.1016/j.cmet.2013.04.011
135. Sharp LZ, Shinoda K, Ohno H, Scheel DW, Tomoda E, Ruiz L, et al. Human BAT possesses molecular signatures that resemble beige/brite cells. PLoS ONE. (2012) 7:e49452. doi: 10.1371/journal.pone.0049452
136. Yoneshiro T, Aita S, Matsushita M, Okamatsu-Ogura Y, Kameya T, Kawai Y, et al. Age-related decrease in cold-activated brown adipose tissue and accumulation of body fat in healthy humans. Obesity. (2011) 19:1755–60. doi: 10.1038/oby.2011.125
137. Yoneshiro T, Aita S, Matsushita M, Kameya T, Nakada K, Kawai Y, et al. Brown adipose tissue, whole-body energy expenditure, and thermogenesis in healthy adult men. Obesity. (2011) 19:13–6. doi: 10.1038/oby.2010.105
138. Chen KY, Brychta RJ, Linderman JD, Smith S, Courville A, Dieckmann W, et al. Brown fat activation mediates cold-induced thermogenesis in adult humans in response to a mild decrease in ambient temperature. J Clin Endocrinol Metab. (2013) 98:E1218–23. doi: 10.1210/jc.2012-4213
139. Saito M, Okamatsu-Ogura Y, Matsushita M, Watanabe K, Yoneshiro T, Nio-Kobayashi J, et al. High incidence of metabolically active brown adipose tissue in healthy adult humans: effects of cold exposure and adiposity. Diabetes. (2009) 58:1526–31. doi: 10.2337/db09-0530
140. Matsushita M, Yoneshiro T, Aita S, Kameya T, Sugie H, Saito M. Impact of brown adipose tissue on body fatness and glucose metabolism in healthy humans. Int J Obes. (2014) 38:812–7. doi: 10.1038/ijo.2013.206
141. Chondronikola M, Volpi E, Børsheim E, Porter C, Annamalai P, Enerbäck S, et al. Brown adipose tissue improves whole-body glucose homeostasis and insulin sensitivity in humans. Diabetes. (2014) 63:4089–99. doi: 10.2337/db14-0746
142. Muzik O, Mangner TJ, Leonard WR, Kumar A, Janisse J, Granneman JG. 15O PET measurement of blood flow and oxygen consumption in cold-activated human brown fat. J Nucl Med. (2013) 54:523–31. doi: 10.2967/jnumed.112.111336
143. U Din M, Raiko J, Saari T, Kudomi N, Tolvanen T, Oikonen V, et al. Human brown adipose tissue [15O]O2 PET imaging in the presence and absence of cold stimulus. Eur J Nucl Med Mol Imaging. (2016) 43:1878–86. doi: 10.1007/s00259-016-3364-y
144. Kalinovich AV, de Jong JMA, Cannon B, Nedergaard J. UCP1 in adipose tissues: two steps to full browning. Biochimie. (2017) 134:127–37. doi: 10.1016/j.biochi.2017.01.007
145. Otero-Díaz B, Rodríguez-Flores M, Sánchez-Muñoz V, Monraz-Preciado F, Ordoñez-Ortega S, Becerril-Elias V, et al. Exercise induces white adipose tissue browning across the weight spectrum in humans. Front Physiol. (2018) 9:1781. doi: 10.3389/fphys.2018.01781
146. Patsouris D, Qi P, Abdullahi A, Stanojcic M, Chen P, Parousis A, et al. Burn induces browning of the subcutaneous white adipose tissue in mice and humans. Cell Rep. (2015) 13:1538–44. doi: 10.1016/j.celrep.2015.10.028
147. Van Der Lans AAJJ, Hoeks J, Brans B, Vijgen GHEJ, Visser MGW, Vosselman MJ, et al. Cold acclimation recruits human brown fat and increases nonshivering thermogenesis. J Clin Invest. (2013) 123:3395–403. doi: 10.1172/JCI68993
148. Chondronikola M, Volpi E, Børsheim E, Porter C, Saraf MK, Annamalai P, et al. Brown adipose tissue activation is linked to distinct systemic effects on lipid metabolism in humans. Cell Metab. (2016) 23:1200–6. doi: 10.1016/j.cmet.2016.04.029
149. Severinsen MCK, Schéele C, Pedersen BK. Exercise and browning of white adipose tissue – a translational perspective. Curr Opin Pharmacol. (2020) 52:18–24. doi: 10.1016/j.coph.2020.04.004
150. Petruzzelli M, Schweiger M, Schreiber R, Campos-Olivas R, Tsoli M, Allen J, et al. A switch from white to brown fat increases energy expenditure in cancer-associated cachexia. Cell Metab. (2014) 20:433–47. doi: 10.1016/j.cmet.2014.06.011
151. Hankir MK, Seyfried F. Do bariatric surgeries enhance brown/beige adipose tissue thermogenesis? Front Endocrinol. (2020) 11:275. doi: 10.3389/fendo.2020.00275
152. Sidossis LS, Porter C, Saraf MK, Børsheim E, Radhakrishnan RS, Chao T, et al. Browning of subcutaneous white adipose tissue in humans after severe adrenergic stress. Cell Metab. (2015) 22:219–27. doi: 10.1016/j.cmet.2015.06.022
153. Longchamp A, Tao M, Bartelt A, Ding K, Lynch L, Hine C, et al. Surgical injury induces local and distant adipose tissue browning. Adipocyte. (2016) 5:163–74. doi: 10.1080/21623945.2015.1111971
154. Giordano A, Coppari R, Castellucci M, Cinti S. Sema3a is produced by brown adipocytes and its secretion is reduced following cold acclimation. J Neurocytol. (2001) 30:5–10. doi: 10.1023/A:1011916822633
155. Cinti S, Frederich RC, Zingaretti MC, De Matteis R, Flier JS, Lowell BB. Immunohistochemical localization of leptin and uncoupling protein in white and brown adipose tissue. Endocrinology. (1997) 138:797–804. doi: 10.1210/endo.138.2.4908
156. Cancello R, Zingaretti MC, Sarzani R, Ricquier D, Cinti S. Leptin and UCP1 genes are reciprocally regulated in brown adipose tissue. Endocrinology. (1998) 139:4747–7. doi: 10.1210/endo.139.11.6434
157. Sbarbati A, Morroni M, Zancanaro C, Cinti S. Rat interscapular brown adipose tissue at different ages: a morphometric study. Int J Obes. (1991) 15:581–7.
158. Geloen A, Collet AJ, Bukowiecki LJ. Role of sympathetic innervation in brown adipocyte proliferation. Am J Physiol. (1992) 263:1176–81. doi: 10.1152/ajpregu.1992.263.6.R1176
159. Lončar D. Convertible adipose tissue in mice. Cell Tissue Res. (1991) 266:149–61. doi: 10.1007/BF00678721
160. Young P, Arch JRS, Ashwell M. Brown adipose tissue in the parametrial fat pad of the mouse. FEBS Lett. (1984) 167:10–4. doi: 10.1016/0014-5793(84)80822-4
161. Kotzbeck P, Giordano A, Mondini E, Murano I, Severi I, Venema W, et al. Brown adipose tissue whitening leads to brown adipocyte death and adipose tissue inflammation. J Lipid Res. (2018) 59:784–94. doi: 10.1194/jlr.M079665
162. Gospodarska E, Nowialis P, Kozak LP. Mitochondrial turnover: a phenotype distinguishing brown adipocytes from interscapular brown adipose tissue and white adipose tissue. J Biol Chem. (2015) 290:8243–55. doi: 10.1074/jbc.M115.637785
163. Altshuler-Keylin S, Shinoda K, Hasegawa Y, Ikeda K, Hong H, Kang Q, et al. Beige adipocyte maintenance is regulated by autophagy-induced mitochondrial clearance. Cell Metab. (2016) 24:402–19. doi: 10.1016/j.cmet.2016.08.002
164. Chan M, Lim YC, Yang J, Namwanje M, Liu L, Qiang L. Identification of a natural beige adipose depot in mice. J Biol Chem. (2019) 294:6751–61. doi: 10.1074/jbc.RA118.006838
165. Lee J, Choi J, Aja S, Scafidi S, Wolfgang MJ. Loss of adipose fatty acid oxidation does not potentiate obesity at thermoneutrality. Cell Rep. (2016) 14:1308–16. doi: 10.1016/j.celrep.2016.01.029
Keywords: brown fat, beige fat, development and origin, transcriptional regulation, epigenetic regulation, browning, thermogenesis, maintenance
Citation: Rabiee A (2020) Beige Fat Maintenance; Toward a Sustained Metabolic Health. Front. Endocrinol. 11:634. doi: 10.3389/fendo.2020.00634
Received: 14 May 2020; Accepted: 05 August 2020;
Published: 04 September 2020.
Edited by:
Maria Chondronikola, University of California, Davis, United StatesReviewed by:
Christian Wolfrum, ETH Zürich, SwitzerlandStefania Carobbio, Wellcome Sanger Institute (WT), United Kingdom
Copyright © 2020 Rabiee. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Atefeh Rabiee, YXJhYmllZUBzdGFuZm9yZC5lZHU=
 Atefeh Rabiee
Atefeh Rabiee