- 1INSERM, U892, Nantes, France
- 2Université de Nantes, Nantes, France
- 3CNRS, UMR 6299, Nantes, France
- 4Service d’Hématologie CHU de Nantes, Nantes, France
Multiple myeloma (MM) is a plasma-cell (PC) malignancy that is heterogeneous in its clinical presentation and prognosis. Monoclonal gammopathy of undetermined significance (MGUS) consistently preceded development of MM. The presence of primary IgH translocations and the universal overexpression of cyclin D genes led to a molecular classification of MM patients into different disease subtypes. Since Bcl-2 family proteins determine cell fate, we analyzed a publicly available Affymetrix gene expression of 44 MGUS and 414 newly diagnosed MM patients to investigate (1) the global change of Bcl-2 family members in MM versus MGUS (2) whether the four major subtypes defined as hyperdiploid, CyclinD1, MAF, and MMSET, display specific apoptotic machineries. We showed that among the main anti-apoptotic members (Bcl-2, Bcl-xL, and Mcl-1), Mcl-1 up-regulation discriminated MM from MGUS, in agreement with the prominent role of Mcl-1 in PC differentiation. Surprisingly, the expression of multi-domain pro-apoptotic Bak and Bax were increased during the progression of MGUS to MM. The combined profile of Bcl-2, Bcl-xL, and Mcl-1 was sufficient to distinguish MM molecular groups. While specific pro-apoptotic members expression was observed for each MM subtypes, CyclinD1 subgroup, was identified as a particular entity characterized by a low expression of BH3-only (Puma, Bik, and Bad) and multi-domain pro-apoptotic members (Bax and Bak). Our analysis supports the notion that MM heterogeneity is extended to the differential expression of the Bcl-2 family content in each MM subgroup. The influence of Bcl-2 family profile in the survival of the different patient groups will be further discussed to establish the potential consequences for therapeutic interventions. Finally, the use of distinct pro-survival members in the different steps of immune responses to antigen raises also the question of whether the different Bcl-2 anti-apoptotic profile could reflect a different origin of MM cells.
Introduction
Multiple myeloma (MM) is a plasma-cell (PC) malignancy that is heterogeneous in its clinical presentation and prognosis. Monoclonal gammopathy of undetermined significance (MGUS) consistently precedes the development of MM. The presence of primary IgH translocations and the universal overexpression of CCND (cyclinD) genes has led to a molecular classification of MM patients into different disease subtypes (1–3). The main translocations involve the immunoglobulin gene heavy chain locus on 14q32.33 with recurrent chromosome partners. These include t(11;14), t(4;14), t(14;16), and t(14;20) with an overexpression of CCND1, MMSET, c-MAF, and MAFB, respectively. Moreover, half of MM patients do not exhibit IgH translocation but present multiple trisomies involving chromosomes 3, 5, 7, 9, 11, 15, 19, and 21 and constitute the hyperdiploid subgroup of MM patients.
Impaired apoptosis is often associated with tumorigenesis and resistance to treatment. Apoptosis is controlled at multiple levels and members of the Bcl-2 family regulate the mitochondrial apoptosis pathway. They can be divided into three functional groups. The anti-apoptotic group comprises Bcl-2, Mcl-1, Bcl-xL, A1, and Bcl-w. These molecules contain four BH-2 homology domains. The pro-apoptotic multi-domain effectors, Bax and Bak, induce mitochondria damage upon activation and constitute a second group (4). Lastly, the BH3-only group, encompasses direct activators of Bax/Bak (Bid, Bim, and Puma) and sensitizers (Noxa, Bik, Bad, Hrk, and Bmf), which bind to anti-apoptotic relatives in order to induce the release of BH3 activators (5).
Individual BH3-only proteins exhibit differential affinities for their pro-survival counterparts. The activators Bim, Puma, and Bid bind all pro-survival members with high affinity, whereas BH3-only sensitizers display more selectivity. For instance, Bad binds with high affinity only to Bcl-2, Bcl-xL, and Bcl-w, and Noxa only to Mcl-1 and A1. Bax and Bak also differ in their interaction profile. Bak is tightly bound by Mcl-1 and Bcl-xL but weakly by Bcl-2 whereas Bax seems to be neutralized by all pro-survival members (6).
The cellular content of the Bcl-2 family molecules varies among the different cell types, however it is certain that the interaction between anti-apoptotic and pro-apoptotic Bcl-2 members dictates whether a cell should die or not (4).
Since Bcl-2 family proteins determine cell fate, we analyzed a publicly available library of Affymetrix gene expression levels from 44 MGUS and 414 newly diagnosed MM patients to investigate (a) the global change of Bcl-2 family members in MM versus MGUS and (b) whether the four major MM subtypes, defined as respectively hyperdiploid (HY) or IgH translocation in 11q13 (CCND1), 16q23 (MAF), and 4p16 (MMSET), display specific apoptotic machineries.
Results
Analysis of Bcl-2 Family Expression between MGUS and MM
We found that among anti-apoptotic members, only Mcl-1 was significantly up-regulated in MM compared to MGUS (p < 0.0001 Mann–Whitney test; Figure 1) whereas BCL2 and BCLXL remained unchanged or slightly decreased in MM versus MGUS (Figure 1). Of note, we excluded BCL2A1 from the study since it is largely expressed in B cells but lost during PC differentiation. The expression of all BH3-only proteins was not modified during the progression from MGUS to MM (Figure 2). In contrast, we observed that the expression of multi-domain pro-apoptotic BAX and BAK was significantly increased in MM (p = 0.055 and p < 0.0001 respectively Mann–Whitney test; Figure 1). Altogether, the major modifications of the Bcl-2 family gene expression during the progression of MGUS to MM mainly affected MCL1 and BAK gene expression (1.27 and 1.54 median fold change, respectively). These modifications between MGUS and MM should be interpreted with caution since PC populations in MGUS include both normal and malignant cells. Accordingly, it has been previously reported that normal PC in MGUS can represent up to 65% whereas in MM the percentage of normal PCs is <2% (7).
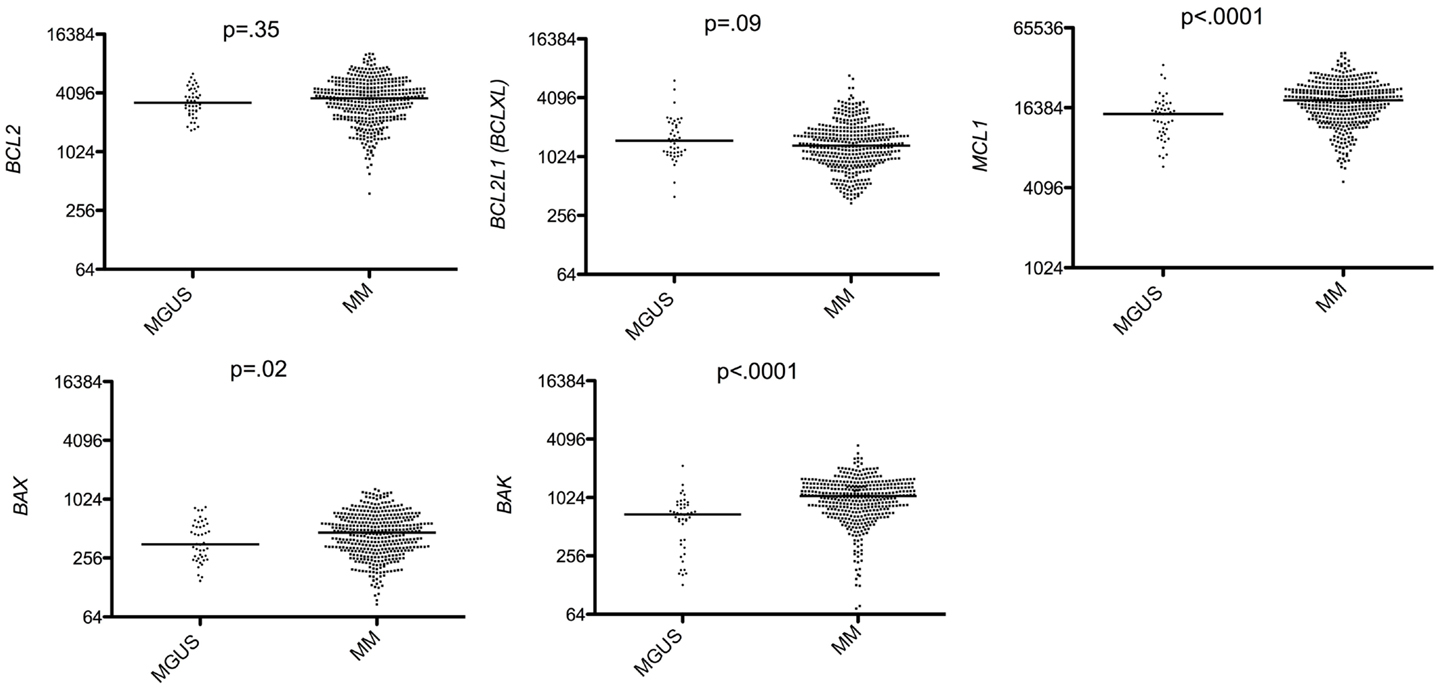
Figure 1. Affymetrix data from a cohort of 44 MGUS and 414 untreated patients from the Arkansas Cancer Research Center were analyzed for BCL2 (probe set 203685_at), BCL2L1 (probe set 212312_at), MCL1 (probe set 200797_s_at), BAX (probe set 208478_s_at), and BAK (probe set 203728_at) expression using the Amazonia database (http://amazonia.transcriptome.eu/). Median values were indicated. Statistical analyses were done using the Mann–Whitney test.
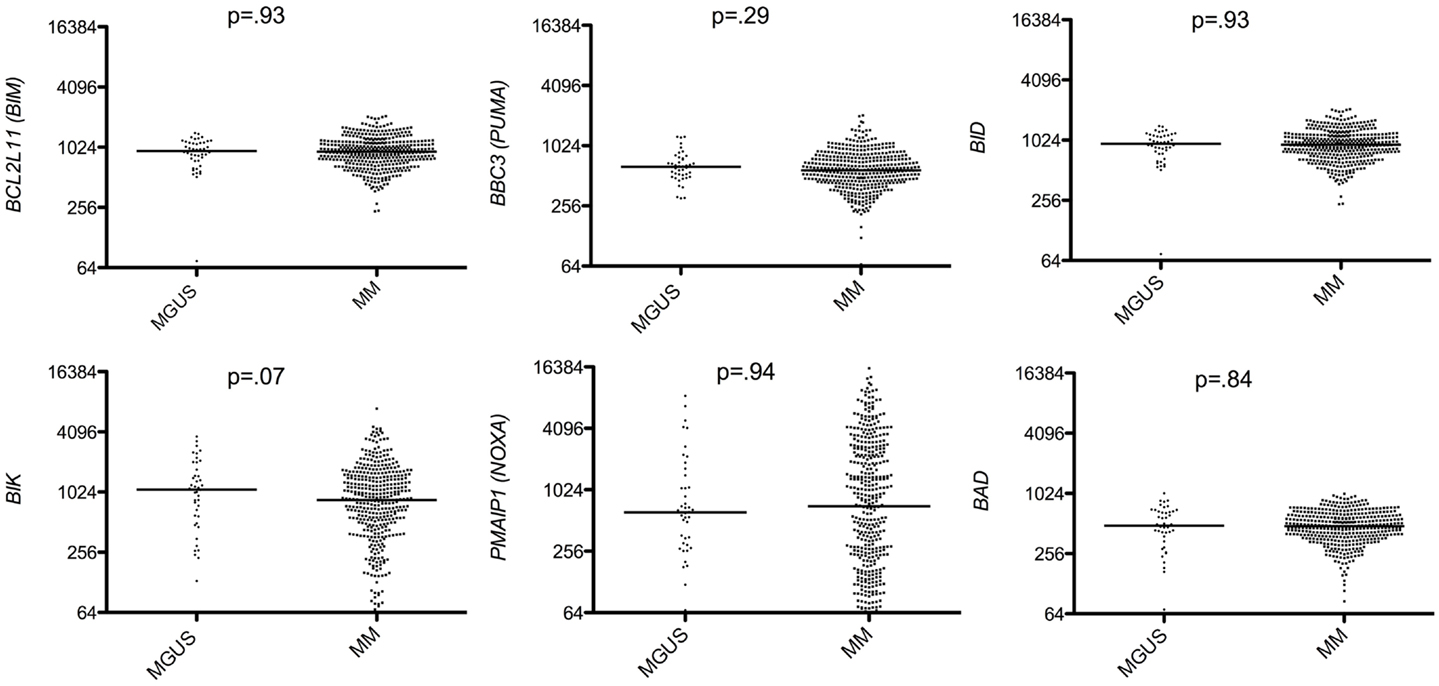
Figure 2. Affymetrix data from a cohort of 44 MGUS and 414 untreated patients from the Arkansas Cancer Research Center were analyzed for BIM (probe set 1553088_a_at), BBC3 (probe set 211692_s_at), BID (probe set 211725_s_at), BIK (probe set 205780_at), PMAIP1 (probe set 204285_s_at), and BAD (probe set 1861_at), expression using the Amazonia database. Median values were indicated. Statistical analyses were done using the Mann–Whitney test.
Comparison of Bcl-2 Family Members in MM Subgroups
Analysis of anti-apoptotic gene expression within the four major subgroups provided evidence that the HY and CCND1 groups can be distinguished from MAF and MMSET groups by a high expression of BCL2 and a weak expression of MCL1 (1.3 and 0.7 median fold change, respectively), as already reported (8). Furthermore, BCLXL allowed to discriminate HY from CCND1 patients and also MAF from MMSET patients, since CCND1 expressed significantly less BCLXL than HY patients (p < 0.0001 Mann–Whitney test; Figure 3) and MMSET patients expressed significantly less BCLXL than MAF patients (p < 0.0001 Mann–Whitney test; Figure 3). Although BCLXL was heterogeneous among the four subtypes, its role in MM physiopathology remained elusive. While we have previously shown that silencing BCLXL did not alter the survival of myeloma cell lines (9), other studies have demonstrated that BCLXL played a role in chemoresistance (10). Of note, a high expression of MCL1 was found in the worse prognosis groups (MAF and MMSET) according to the essential function of Mcl-1 in MM cell survival (9, 11, 12). On the other hand, we may question whether the lowest MCL1 levels present in the CCDN1 group could influence its neutral outcome.
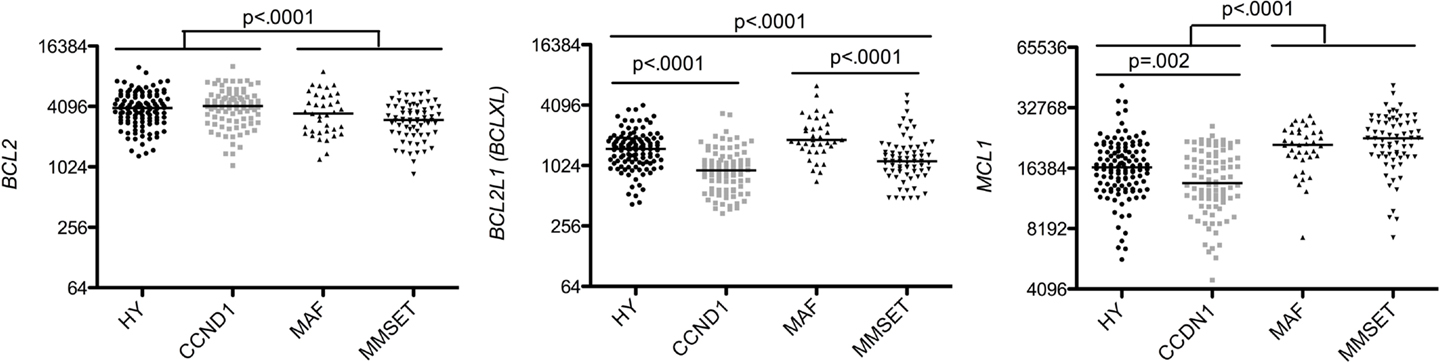
Figure 3. Affymetrix data from untreated patients were analyzed for BCL2, BCL2L1, and MCL1, expression using the Amazonia database in the different molecular groups classified as follow: hyperdiploid (HY), CCND1 (CD1 + CD2), MAF, and MMSET (3). Probe sets are indicated in Figure 1. Median values were indicated. Statistical analyses were conducted using the Kruskall–Wallis test when more than two groups were compared and Mann–Whitney test for the comparison between two groups.
BH3-mimetic small molecules that bind to the BH3 binding sites of anti-apoptotic proteins have been developed. Among them ABT-199 is the newest one, characterized by its high potency to inhibit specifically Bcl-2 (13). In agreement with the high expression of BCL2 in CCND1 patients, we have recently demonstrated in a small cohort of MM patients that ABT-199 sensitivity was restricted to t(11;14) patients (14). Further analysis of a larger cohort of MM patients for ABT-199 sensitivity could allow identifying HY patients able to respond to ABT-199.
Strikingly, analysis of multi-domain pro-apoptotic members showed that the worse prognosis groups MAF and MMSET displayed higher levels of BAX and BAK in contrast to the CCDN1 group, which expressed the lowest levels of both effectors (1.38 and 1.85 median fold change, respectively) (Figure 4). BH3-only activators (BIM, PUMA, BID) were constantly expressed in the four subtypes, suggesting that independently of the subtype, MM cells are primed for death, as already reported (15, 16) (Figure 5). Of note, Puma expression was weaker exclusively in CCDN1 group compared to the other subgroups (0.82 median fold change). The promiscuous binding of Bim and Puma to main pro-survival members, associated with the fact that knockout mice for them do not present a particular phenotype, suggest a complementary role for these two members (17). However, based on the constant endogenous expression of BIM and PUMA found in the four MM subgroups, it would be pertinent to address the question whether they have complementary roles or not in this pathology.
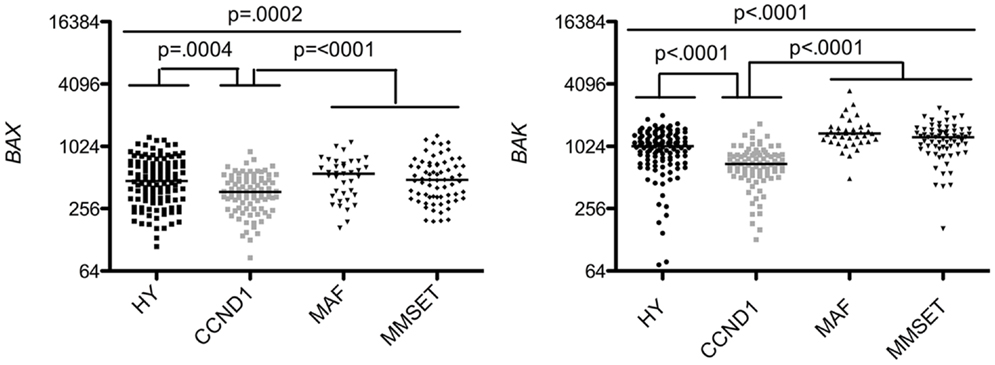
Figure 4. Affymetrix data from untreated patients were analyzed for BAX and BAK expression using the Amazonia database in the different molecular groups classified as above in Figure 3. Probe sets are indicated in Figure 1. Median values were indicated. Statistical analyses were conducted as in Figure 3.
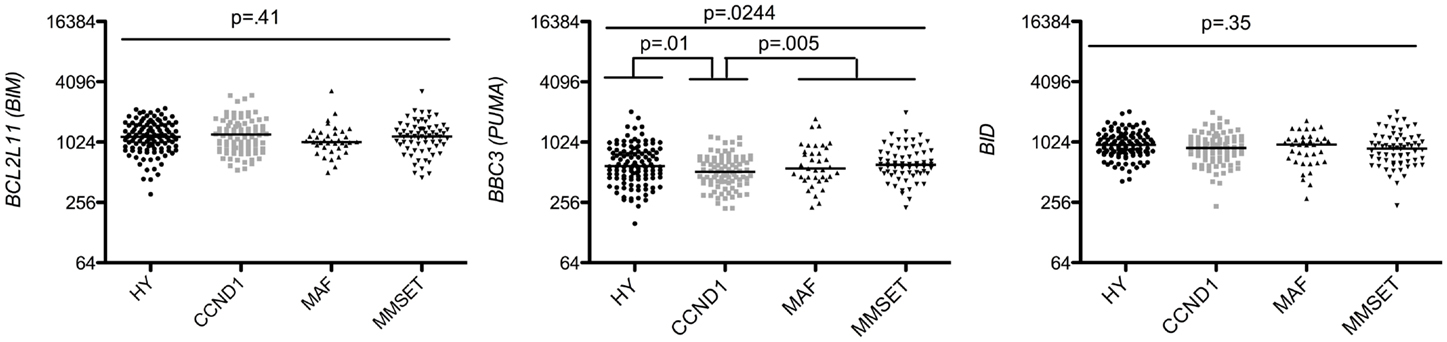
Figure 5. Affymetrix data from untreated patients were analyzed for BIM, BBC3, and BID, expression using the Amazonia database in the HY, CCND1, MAF, and MMSET molecular groups. Probe sets are indicated in Figure 2. Median values were indicated. Statistical analyses were conducted as in Figure 3.
In contrast to BH3-only activators, sensitizers (BIK, NOXA, BAD) were heterogeneously expressed in the different MM subtypes. However, each MM subgroup highly expressed at least one sensitizer (Figure 6). These results are consistent with the fact that sensitizer BH3-only proteins may have overlapping functions (18). In this respect, it was previously shown that knockout mice for either Bik or Noxa proteins do not develop spontaneous tumors (17, 19). Interestingly, we found that few patients in some subgroups lacked Bik, according to our previous finding showing that some MM cell lines do not express Bik at the protein level (20). Deletions and epigenetic alterations have been shown to contribute to the lack of Bik expression (21, 22). TEF, a PARbZIP transcription factor, was identified as a direct activator of BIK promoter (21). We have shown in MM cell lines, that Bik was expressed only in the presence of TEF mRNA (20). However, despite TEF expression, some cell lines did not express Bik. Altogether, these results suggest that lack of Bik might be the result of either an epigenetic alteration or a deletion, as frequently described in other cancers (22).
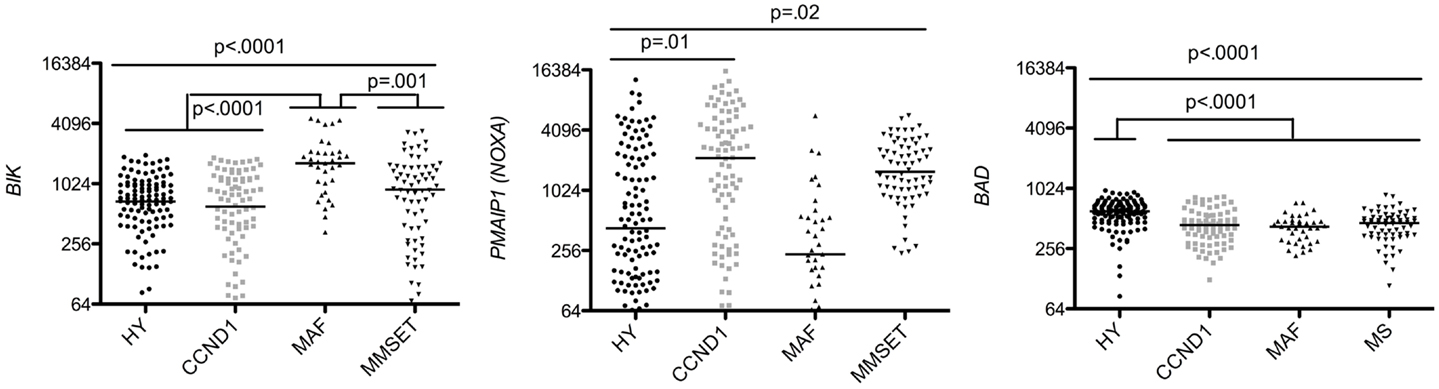
Figure 6. Affymetrix data from untreated patients were analyzed for BIK, PMAIP1, and BAD expression using the Amazonia database in the HY, CCND1, MAF, and MMSET molecular groups. Probe sets are indicated in Figure 2. Median values were indicated. Statistical analyses were conducted as in Figure 3.
In addition, NOXA allowed discriminating CCND1 from HY patients (5 median fold change) and also MMSET from MAF patients (6.68 median fold change) (Figure 6). Although the difference of NOXA expression was impressive among the four subtypes, its role in MM physiopathology remained to be determined.
Hyperdiploid patients expressed significantly higher levels of BAD than all other subgroups (1.35 median fold change) (Kruskall–Wallis p < 0.0001), which may be explained by the localization of BAD on chromosome 11q1 (Figure 6). Indeed, chromosome 11 trisomy is one the most frequent anomaly in the HY subgroup (23).
Concluding Remarks
Noteworthy, our analysis demonstrated that the combined profile of the three anti-apoptotic molecules (Bcl-2, Bclx-L, and Mcl-1) was sufficient to discriminate the different MM molecular groups. The CCDN1 subgroup was identified as a particular entity, characterized by a BCL2high MCL1low and a low expression of pro-apoptotic effectors and BH3-only (PUMA, BIK, and BAD) with the exception of high expression of NOXA. Since Noxa interacts only with Mcl-1, which is weak in this subtype, we can hypothesize that the anti-apoptotic function of Mcl-1 is totally neutralized by Noxa and that this subtype relies mainly on Bcl-2 for survival. Altogether, the apoptotic machinery of this myeloma subtype is very different from that of other subtypes, suggesting that specific therapeutic approaches should be investigated to target CCDN1 patients. We can also hypothesize that the specificity of this subgroup may reflect a specific origin of malignant PC immortalization. This is also supported by the fact that patients harboring a t(11;14) have a high prevalence of IgM isotype and represent a distinct biological and clinical subgroup (24). Furthermore, recurrent translocations also correlated with particular features (25). Indeed, t(11:14) translocations exhibited a mature lymphoplasmocytoid morphology with a higher incidence of non-secretory MM. In contrast, t(4;14) MM subtype revealed a morphology of immature plasma cells with a significant amount of plasmablasts (25, 26). The HY subgroup shares some similarities with CCND1 in the expression pattern of Bcl-2 family members, particularly high BCl2 and low MCL1 levels. However, HY patients expressed higher levels of pro-apoptotic members (BAX, BAK, and BAD) than those of the CCND1 subgroup (Figures 5 and 6). This could favor the apoptotic response to chemotherapy and therefore explain in part the better outcome of these patients. In contrast, MAF and MMSET subgroups differ from CCDN1 and HY in the expression of anti-apoptotic members and were characterized by low BCL2 and high MCL1 levels. Surprisingly, MAF and MMSET subgroups expressed high level of effectors, particularly Bak, suggesting their ability to trigger an effective drug response. It will be intriguing to define whether the high expression of effectors in MAF and MMSET subgroups may be related to the fact that bortezomib-containing regimens could overcome the poor prognosis associated with t(4;14) (27). Paradoxically, these subgroups have the poorest outcome, highlighting a potential role of Mcl-1 in chemoresistance. A potential and interesting approach to target efficiently MAF and MMSET patients would be to use pharmacological inhibitors of Mcl-1, which are currently under development (28).
To summarize, our analysis supports the notion that MM heterogeneity extends to the composition of the Bcl-2 family content in each MM subgroup, which should be taken into account for therapeutic intervention in the new approach of personalized therapies. Finally, a better knowledge of Bcl-2 expression patterns may be relevant to address the origin of malignant PC.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1. Bergsagel PL, Kuehl WM, Zhan F, Sawyer J, Barlogie B, Shaughnessy J Jr. Cyclin D dysregulation: an early and unifying pathogenic event in multiple myeloma. Blood (2005) 106(1):296–303. doi:10.1182/blood-2005-01-0034
2. Fonseca R, Bergsagel PL, Drach J, Shaughnessy J, Gutierrez N, Stewart AK, et al. International Myeloma Working Group molecular classification of multiple myeloma: spotlight review. Leukemia (2009) 23(12):2210–21. doi:10.1038/leu.2009.174
3. Zhan F, Huang Y, Colla S, Stewart JP, Hanamura I, Gupta S, et al. The molecular classification of multiple myeloma. Blood (2006) 108(6):2020–8. doi:10.1182/blood-2005-11-013458
4. Strasser A, Cory S, Adams JM. Deciphering the rules of programmed cell death to improve therapy of cancer and other diseases. EMBO J (2011) 30(18):3667–83. doi:10.1038/emboj.2011.307
5. Happo L, Strasser A, Cory S. BH3-only proteins in apoptosis at a glance. J Cell Sci (2012) 125(Pt:5):1081–7. doi:10.1242/jcs.090514
6. Willis SN, Fletcher JI, Kaufmann T, Van Delft MF, Chen L, Czobotar PE, et al. Apoptosis initiated when BH3 ligands engage multiple Bcl-2 homologs, not Bax or Bak. Science (2007) 315(5813):856–9. doi:10.1126/science.1133289
7. Schmidt-Hieber M, Gutiérrez ML, Pérez-Andrés M, Paiva B, Rasillo A, Tabernero MD, et al. Cytogenetic profiles in multiple myeloma and monoclonal gammopathy of undetermined significance: a study in highly purified aberrant plasma cells. Haematologica (2013) 98(2):279–87. doi:10.3324/haematol.2011.060632
8. Bodet L, Bougie-Gomez P, Touzeau C, Dousset C, Descamps G, Maiga S, et al. ABT-737 is highly effective against molecular subgroups of multiple myeloma. Blood (2011) 118(14):3901–10. doi:10.1182/blood-2010-11-317438
9. Derenne S, Monia B, Dean NM, Taylor JK, Rapp MJ, Harousseau JL, et al. Antisense strategy shows that Mcl-1 rather than Bcl-2 or Bcl-x(L) is an essential survival protein of human myeloma cells. Blood (2002) 100(1):194–9. doi:10.1182/blood.V100.1.194
10. Schwarze MM, Hawley RG. Prevention of myeloma cell apoptosis by ectopic bcl-2 expression or interleukin 6-mediated up-regulation of bcl-xL. Cancer Res (1995) 55(11):2262–5.
11. Jourdan M, Veyrune J-L, De Vos J, Redal N, Couderc G, Klein B. A major role for Mcl-1 antiapoptotic protein in the IL-6-induced survival of human myeloma cells. Oncogene (2003) 22(19):2950–9. doi:10.1038/sj.onc.1206423
12. Le Gouill S, Podar K, Harousseau JL, Anderson KC. Mcl-1 regulation and its role in multiple myeloma. Cell Cycle (2004) 3(10):1259–62. doi:10.4161/cc.3.10.1196
13. Souers AJ, Leverson JD, Boghaert ER, Ackler SL, Catron ND, Chen J, et al. ABT-199, a potent and selective BCL-2 inhibitor, achieves antitumor activity while sparing platelets. Nat Med (2013) 19(2):202–8. doi:10.1038/nm.3048
14. Touzeau C, Dousset C, Le Gouill S, Sampath D, Leverson JD, Souers AJ, et al. The Bcl-2 specific BH3 mimetic ABT-199: a promising targeted therapy for t(11;14) multiple myeloma. Leukemia (2013). doi:10.1038/leu.2013.216
15. Gomez-Bougie P, Bataille R, Amiot M. The imbalance between Bim and Mcl-1 expression controls the survival of human myeloma cells. Eur J Immunol (2004) 34(11):3156–64. doi:10.1002/eji.200424981
16. Ni Chonghaile T, Sarosiek KA, Vo TT, Ryan JA, Tammareddi A, Moore Vdel G, et al. Pretreatment mitochondrial priming correlates with clinical response to cytotoxic chemotherapy. Science (2011) 334(6059):1129–33. doi:10.1126/science.1206727
17. Fernandez-Luna JL. Regulation of pro-apoptotic BH3-only proteins and its contribution to cancer progression and chemoresistance. Cell Signal (2008) 20(11):1921–6. doi:10.1016/j.cellsig.2008.04.015
18. Coultas L, Bouillet P, Stanley EG, Brodnicki TC, Adams JM, Strasser A. Proapoptotic BH3-only Bcl-2 family member Bik/Blk/Nbk is expressed in hemopoietic and endothelial cells but is redundant for their programmed death. Mol Cell Biol (2004) 24(4):1570–81. doi:10.1128/MCB.24.4.1570-1581.2004
19. Villunger A, Michalak EM, Coultas L, Müllauer F, Böck G, Ausserlechner MJ, et al. p53- and drug-induced apoptotic responses mediated by BH3-only proteins puma and noxa. Science (2003) 302(5647):1036–8. doi:10.1126/science.1090072
20. Bodet L, Ménoret E, Descamps G, Pellat-Deceunynck C, Bataille R, Le Gouill S, et al. BH3-only protein Bik is involved in both apoptosis induction and sensitivity to oxidative stress in multiple myeloma. Br J Cancer (2010) 103(12):1808–14. doi:10.1038/sj.bjc.6605981
21. Ritchie A, Gutierrez O, Fernandez-Luna JL. PAR bZIP-bik is a novel transcriptional pathway that mediates oxidative stress-induced apoptosis in fibroblasts. Cell Death Differ (2009) 16(6):838–46. doi:10.1038/cdd.2009.13
22. Sturm I, Stephan C, Gillissen B, Siebert R, Janz M, Radetzki S, et al. Loss of the tissue-specific proapoptotic BH3-only protein Nbk/Bik is a unifying feature of renal cell carcinoma. Cell Death Differ (2006) 13(4):619–27. doi:10.1038/sj.cdd.4401782
23. Kumar S, Fonseca R, Ketterling RP, Dispenzieri A, Lacy MQ, Gertz MA, et al. Trisomies in multiple myeloma: impact on survival in patients with high-risk cytogenetics. Blood (2012) 119(9):2100–5. doi:10.1182/blood-2011-11-390658
24. Avet-Loiseau H, Garand R, Lode L, Harousseau JL, Bataille R. Translocation t(11;14)(q13;q32) is the hallmark of IgM, IgE, and nonsecretory multiple myeloma variants. Blood (2003) 101(4):1570–1. doi:10.1182/blood-2002-08-2436
25. Garand R, Avet-Loiseau H, Accard F, Moreau P, Harousseau JL, Bataille R. t(11;14) and t(4;14) translocations correlated with mature lymphoplasmacytoid and immature morphology, respectively, in multiple myeloma. Leukemia (2003) 17(10):2032–5. doi:10.1038/sj.leu.2403091
26. Greipp PR, Leong T, Bennett JM, Gaillard JP, Klein B, Stewart JA, et al. Plasmablastic morphology – an independent prognostic factor with clinical and laboratory correlates: Eastern Cooperative Oncology Group (ECOG) myeloma trial E9486 report by the ECOG Myeloma Laboratory Group. Blood (1998) 91(7):2501–7.
27. Rajkumar SV. Multiple myeloma: 2012 update on diagnosis, risk-stratification, and management. Am J Hematol (2012) 87(1):78–88. doi:10.1002/ajh.22237
Keywords: multiple myeloma, MGUS, Bcl-2 family, CCND1, MMSET, MAF
Citation: Gomez-Bougie P and Amiot M (2013) Apoptotic machinery diversity in multiple myeloma molecular subtypes. Front. Immunol. 4:467. doi: 10.3389/fimmu.2013.00467
Received: 30 September 2013; Accepted: 04 December 2013;
Published online: 23 December 2013.
Edited by:
Catherine Pellat-Deceunynck, Centre National de la Recherche Scientifique, FranceReviewed by:
To-Ha Thai, Beth Deaconess Israel Medical Center, USAChristian Billard, UMRS 872 INSERM – UPMC, France
Copyright: © 2013 Gomez-Bougie and Amiot. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Martine Amiot, CNRS, UMR 6299, INSERM, U892, Centre de Recherche en Cancérologie Nantes/Angers, 8 Quai Moncousu, 44007 Nantes, France e-mail:bWFydGluZS5hbWlvdEBpbnNlcm0uZnI=
 Patricia Gomez-Bougie1,2,3,4
Patricia Gomez-Bougie1,2,3,4 Martine Amiot
Martine Amiot