- Istituto Nazionale Genetica Molecolare “Romeo ed Enrica Invernizzi,” Milan, Italy
Induction of the adaptive immune system is evaluated mostly by assessment of serum antibody titers and T lymphocyte responses in peripheral blood, although T and B cell activation occurs in lymphoid tissues. In recent years, the release of microRNAs (miRNAs) in the extra-cellular environment has been exploited to assess cell functions at distance via measurement of serum miRNAs. Activated lymphocytes release a large amount of nano-sized vesicles (exosomes), containing miRNA, however there are insufficient data to determine whether this phenomenon is reflected in modulation of serum miRNAs. Interestingly, miRNA signatures of CD4+ T cell-derived exosomes are substantially different from intracellular miRNA signatures of the same cells. We have recently identified serum circulating miR-150 as a sensor of general lymphocyte activation and we strongly believe that miRNAs differentially released by specific CD4+ effector T cell subsets (Th1, Th2, Th17, and Treg) may serve as serum biomarkers of their elicitation in lymphoid tissues but also in damaged tissues, potentially providing clinically relevant information about the nature of immune responses in health and disease.
Blood-Circulating miRNAs as Biomarkers in Health and Disease
MicroRNAs (miRNAs) are small single-stranded RNA molecules (18–25 nt), that once loaded into the Argonaute protein of the silencing complex, pair with messenger RNAs, directly effecting post-transcriptional regulation (1). In recent years, it has been found that most cells release miRNAs in the extra-cellular environment, predominantly in association with either vesicles or protein complexes that protect them from RNAses (2–4). This release can be a passive phenomenon that results from tissue damage, or an active process as in the case of miRNAs actively secreted into the surrounding environment by healthy cells, where once outside, miRNAs can reach the bloodstream and constitute what it is now referred to as the “blood-circulating extra-cellular miRNome.” Extra-cellular miRNAs can be extracted from serum, plasma, and other body fluids and profiled through microarray, real time quantitative PCR or sequencing. This possibility has been exploited to assess cell functions at distance via measurement of serum miRNAs and nowadays blood-circulating miRNAs are regarded among the most promising clinical biomarkers for the diagnosis, prognosis, and therapeutic options of a variety of pathological conditions such as cancer (5–7), cardiovascular diseases (8, 9), diabetes (10), liver pathologies (11, 12), and sepsis (13, 14), among others [reviewed in Ref. (15)].
Circulating miRNAs as clinical biomarkers are not without some technical challenges. First, dilution effects in blood limit the amount of RNA per volume of starting material. Second, cellular detritus, hemolysis, and the presence of contaminating components constitute pre-analytical challenges, potentially impacting reproducibility and sensitivity. Finally, as miRNAs are released by virtually all cells in the body and most of the blood miRNAs are released by large organs as well as highly dividing cells, specificity is impacted by high background. However, the fact that serum miRNAs circulate in different compartments might provide an advantage. miRNAs circulate in association with vesicles of nanometric size (20–100 nm) called exosomes, that are formed by the inward budding and subsequent fusion to the plasma membrane of multivesicular endosomes (16); vesicles of larger size (0.2–1 μm) that bud directly from the plasma membrane, are called microvesicles and comprise also apoptotic and senescent bodies (17); in association with Argonaute protein in a vesicle-free form (3); or linked to high-density lipoproteins (18). This compartmentalization facilitates the purification of specific isolates, enriching for biomarkers of interest. Moreover, the possibility of isolating cell lineage-specific exosomes based upon their parental protein expression patterns, could in principle represent a further advantage in the identification and validation of biomarkers in lymphocyte activation.
Release of Exosome-Associated miRNAs upon Activation of Lymphocytes: Biological Aspects
The presence of RNA, such as messengers and regulatory RNAs (among which miRNAs) within exosomes was initially described in 2007 (19) but the role of exosomes in conveying intercellular communication had already been extensively investigated in the immune system [reviewed in Ref. (20, 21)]. In 1996, it was first demonstrated that B lymphocytes release exosomes with a significantly different overall surface protein profile from that of the plasma membrane and that they do contain MHC class II and are able to induce antigen-specific MHC class II-restricted T cell responses, demonstrating a specific role for exosomes in antigen presentation in vivo (22). Later it was also shown that upon TCR triggering, T lymphocytes produce a large amount of exosomes that bear TCR from the pool of activated complexes. It was suggested that exosomes may work as powerful vehicles to specifically deliver signals to cells with a specific combination of peptide/MHC complexes (23). Furthermore, T lymphocytes have been observed killing target cells by CD95 engagement through membrane CD95L containing exosomes (24). Exosomes can be viewed as communication modules between cells of the immune system, and are regarded by some authors as important to the microenvironment as the release of cytokines and chemokines (25). While the role of intracellular miRNAs has long been recognized as a key level of gene-expression regulation in cells of the immune system (26, 27), its role in exosomes has only begun to be explored. Initial data indicate an exosome-mediated transfer of RNA between T cells and antigen presenting cells during antigen recognition, as well as the capacity of miRNAs transferred during immune synapse to modulate gene-expression in recipient cells (28, 29). To fully elucidate the role of exosome-associated miRNAs, it will be necessary to better characterize the fundamental processes of extra-cellular RNA biogenesis, distribution, uptake, and how they contribute to overall function. In principle, since exosomes contain myriad miRNAs in varying numbers they possess the potential to regulate the expression of multiple genes leading to very effective paracrine control over neighboring cells (30).
Lymphocyte Signatures of Exosome-Associated miRNAs: A New Landscape to be Unveiled
The release of RNA through exosomes is not a passive phenomenon, but actually a regulated, active process demonstrated by the fact that exosomal RNA content is not at all a mere reflection of that found within the intracellular milieu. By next-generation sequencing of small RNA species present in vesicles released in co-culture of T lymphocytes and dendritic cells, it has been described that distinct RNA categories, such as small ribosomal RNA and specific tRNA fragments, long interspersed elements (LINEs), and long terminal repeats (LTRs) are conveyed in, and released by vesicles in significantly greater numbers than other RNA types, such as lincRNAs (31). A quantitative analysis shows the selective enrichment of some miRNAs in purified exosomes compared to cells. The differential rate of release for various RNA molecules was further demonstrated by our finding that the intracellular miRNome of CD4+ T lymphocytes is more similar to the intracellular miRNome of B lymphocytes than to their own exosomal miRNome (Figure 1A) (32). Indeed, we have identified a discrete set of miRNAs whose intracellular concentrations, compared to that found within their cognate lymphocyte-derived exosomes, was significantly different (32).
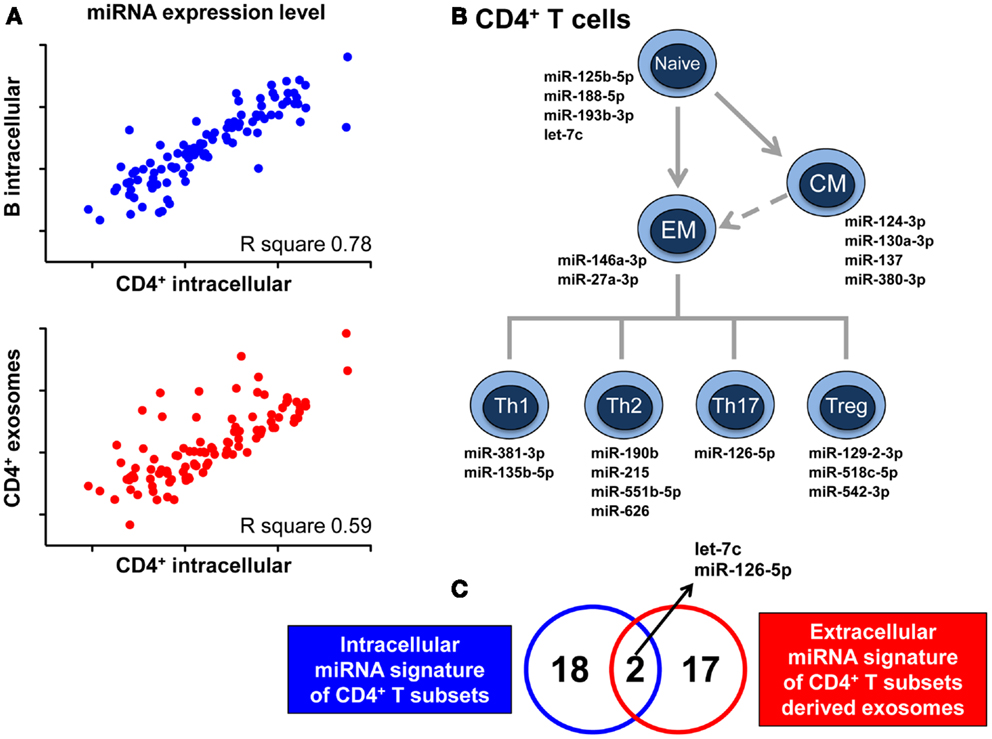
Figure 1. MicroRNAs signature of CD4+ T cell-derived exosomes cannot be predicted by relevant intracellular miRNA signatures of CD4+ T cells development and differentiation. (A) Pearson correlation between miRNA expression level in CD4+ lymphocytes and either miRNA expression level in B lymphocytes (upper panel) or miRNA representation in CD4+ lymphocyte-derived exosomes (lower panel). R square values are reported. (B) Schematic view of CD4+ T lymphocytes with the relative intracellular miRNA signatures identified as relevant for the development and differentiation of these subsets (EM, effector memory; CM, central memory). (C) Venn diagram showing the partial overlap between the 20 miRNAs composing the intracellular signatures of CD4+ T subsets of panel B and the 19 miRNAs at later stage identified for being either exclusive or differentially up-represented in exosomes derived from CD4+ T subsets compared to exosomes derived from B lymphocytes. Data shown are adapted from previous published studies (32, 33).
As previously reported, we have identified 20 intracellular miRNAs that are able to distinguish between different subpopulations of CD4+ T cells, defining the development and differentiation of this lineage (Figure 1B) (33). Only two of these miRNAs were independently demonstrated to be differentially represented in Th cell-derived compared to B cell-derived exosomes (Figure 1C); further 17 miRNAs with a signature specifically associated to exosomes released by CD4+ T cells have been subsequently identified (32). These observations tell us that even a thorough analysis of the biological relevance of miRNAs at the intracellular level is not helpful in deciphering the spectrum of differences at the extra-cellular level and that only a high-throughput quantitative investigation of miRNA in exosomes will define lymphocyte-specific exosome-associated miRNA signatures. Consequently, this endeavor commenced by showing that CD4+ T and B lymphocytes display significantly different selective enrichment of specific extra-cellular miRNAs, and we are now in the process of fully elucidating the differential exosomal miRNomes of each CD4+ T subset.
These studies will potentially have two different and equally relevant impacts on the field. On the one side, the analysis of extra-cellular lymphocyte signatures may shed light on the role of miRNA disposal during activation. For example, as previously described, when primary CD4+ T cells are activated in vitro, they dramatically down-regulate intracellular miR-150 while accumulating it in extra-cellular vesicles, suggesting that this type of release may represent an additional layer of post-transcriptional regulation for miRNAs with very rapid effects on target genes of the discarded miRNAs (32). We thus believe that correlating data of intracellular modulation upon activation with data of extra-cellular disposal will tell us if the case of miR-150 is isolated or not, and if this type of regulation is specific for different lymphocyte subsets. Furthermore, should exosome-associated miRNAs display a paracrine control over neighboring cells, the full description of extra-cellular miRNAs differentially released by different subsets of effector cells will profoundly change the knowledge we have on how these cells impact on the extra-cellular environment.
On the other side, these same studies will pave the way for the identification and validation of potentially powerful biomarkers of lymphocyte activation. Indeed, upon the identification of miRNAs that are differentially released by various lymphocyte effector cells (e.g., Th1, Th2, Th17, and Treg), the assessment of their modulation in serum may render possible to mark the elicitation of these cells, which occurs in lymphoid tissues or damaged organs (34).
The Growing Need for New Biomarkers of Vaccination
Vaccinations are based on the activation of the adaptive immune system. Their efficacy is evaluated mostly by assessing serum antibody titers and lymphocyte responses in peripheral blood, despite T and B cell activation occurring within the lymphoid tissues. While the protective role of vaccination is primarily conferred by the generation of B cells-derived antigen-specific antibodies, T cells are fundamental for the induction of high-affinity antibodies and immune memory, and in certain types of vaccines, these cells must be regarded as prime effectors. Then, the identification of novel immune correlates of vaccine efficacy that take into account parameters different than antibody titers will become increasingly more important in the development of new adjuvants and the optimization of current vaccines (35). Moreover, as some subsets of effector T helper cells can trigger adverse events from allergy to autoimmunity, the ability to easily monitor the induction of these subsets by new vaccines is becoming critical concern in clinical development.
Toward these ends, a pilot study to address the feasibility of following the generation of an immune response through the profiling of serum-associated miRNAs has been undertaken (32).
Exosome-Associated miR-150 as a General Sensor of Vaccination
Over the past few years, one specific miRNA, namely miR-150, has been confirmed to play a critical role in the development of lymphoid and myeloid lineages in both mice and humans (36). In particular, miR-150 is expressed at a low level in B- and T-progenitor cells, but gets highly up-regulated in mature lymphocytes. Moreover, as naïve T cells differentiate into effector Th cells, the level of miR-150 is down-modulated again (37).
miR-150 has been also frequently observed to be dramatically dysregulated in several types of leukemias and lymphomas (36). When the level of intracellular miR-150 decreases, some critical targets get de-repressed, e.g., c-Myb, a transcription factor that promotes lymphocyte survival by inducing Bcl2 (38, 39). Another target of miR-150 action is Notch3, a member of the Notch family of receptors, which plays a pivotal role in T cell differentiation and leukemogenesis (40).
We recently described that when primary human CD4+ Th cells dramatically down-regulate intracellular miR-150 upon activation, it’s released by exosomes, suggesting that this process of extra-cellular miRNA “disposal” may represent an additional layer of post-transcriptional down-regulation for miRNAs with very rapid effects on target genes that critically control lymphocyte responses (32).
In parallel with the biological implications of these observations, we decided to investigate miRNA-150 as a potential candidate for the optimal biomarker of lymphocyte activation because: (i) it is highly expressed in both human lymphocyte cells and lymphocyte-derived exosomes; (ii) it is expressed specifically in spleen compared to other human tissues, supporting the idea that the major source of serum miR-150 are lymphoid cells and (iii) it is easily detectable and significantly enriched in exosomes circulating in human blood (32). Hence, our working hypothesis is based on the assumption that when the immune system is activated by vaccination, the lymphocytes that participate in the response will release an easily detectable number of exosomes into the bloodstream and consequently a readily measureable level of a lymphocyte-derived exosomal miRNA (as it is the case for miR-150) (Figure 2A). Before proceeding to analyze precious human samples of vaccinated individuals, we made use of the mouse model, and discovered that, as correctly hypothesized, serum miR-150 levels increase significantly in mice upon vaccination with adjuvant-OVA. Instead, miR-150 serum concentrations remain unchanged in immunized mice that are depleted of mature CD4+ T lymphocytes [major histocompatibility complex class II-deficient mice (41)] showing that serum miR-150 modulation is a specific phenomenon strictly dependent on adaptive immune responses elicited by effective vaccination (Figure 2B) (32).
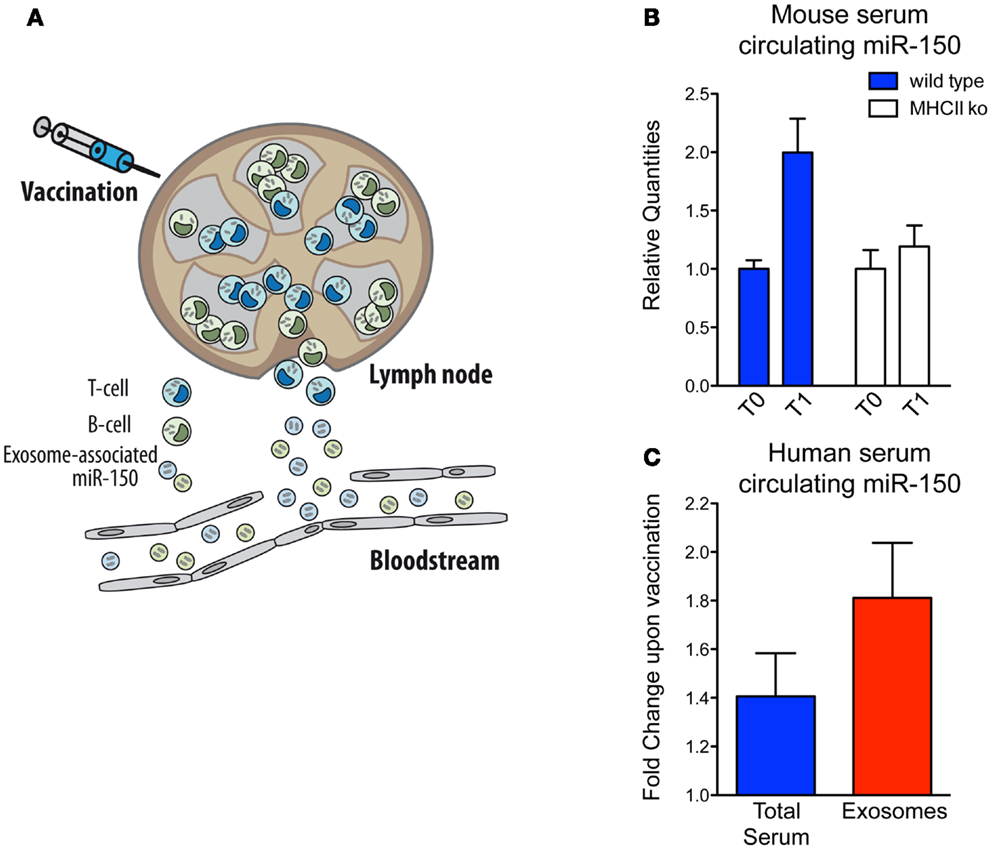
Figure 2. Serum circulating miR-150 increases in both human and mice upon vaccination. (A) Schematic view of exosome-associated miR-150 release upon induction of adaptive immune response in secondary lymphoid organs upon vaccination. (B) Mouse serum circulating miR-150 increase expressed as relative quantities at time T0 (pre-vaccination) and T1 (post-vaccination) in wild type compared to CD4+ cell-depleted MHCII knock out mice (four animals/group; the result shown is representative of two independent experiments). (C) Human circulating miR-150 increase expressed as fold change upon vaccination in total serum compared to serum purified exosomes (results come from 17 paired samples of sera collected at the time of vaccination and 4 weeks after that). Data shown are adapted from previous published studies (32).
Sera from adults or children vaccinated with the 2009 pandemic flu (H1N1) vaccine adjuvanted with MF-59 (42) was then assessed and, similarly to what was found in mice, the level of extra-cellular miR-150 increased significantly in human serum upon vaccination and this increase was significantly more evident upon purification of exosomes (Figure 2C) (32). This observation suggests that serum miR-150 modulation is specifically compartmentalized to lymphocyte-derived vesicles and that exosome purification strategies from blood may serve to increase lymphocyte biomarker sensitivity by enriching immune-related circulating extra-cellular RNAs, as suggested for other conditions (7, 43). Furthermore, in flu vaccinated individuals, miR-150 serum levels post-vaccination have been found to be significantly higher in people mounting higher antibody response, showing a quantitative correlation between the modulation of a circulating miRNA and the adaptive immune response (32).
In a recent study, reduced miR-150 serum concentrations have also been found to be associated with an unfavorable outcome in critically ill patients with sepsis. It has been hypothesized that lower circulating miR-150 levels might lead to de-repression of genes such as CXCR4 and c-Myb, both linked to immune response activation and poor prognosis (14).
Consistently, we do speculate that the significant uptick in circulating miR-150 levels that coincides with vaccination may play a role in down-modulating adaptive immune responses by carrying extra-cellular messages to other immune cells and consequently regulating miR-150 target genes.
Obviously, as miR-150 is ubiquitous across all lymphocyte populations, it may only serve as a generic lymphocyte activation sensor, devoid of more insights into the lymphocyte subsets involved. Nonetheless, this study has provided the first proof-of-concept that serum miRNAs can be readily detected, from a minimally invasive serum sample, toward their validation as sensitive and specific biomarkers of vaccination, and more generally of the adaptive immune response.
Summary and Perspectives
Blood-circulating extra-cellular miRNAs have the potential to become highly valuable biomarkers in the near future. In particular, the identification of serum miRNA signatures able to directly report the differential activation state of clinically relevant lymphocytic subsets may become an innovative tool to provide pivotal information about the nature of the immune responses occurring in health (e.g., vaccination) and disease (e.g., auto-immune and immune-mediated disorders). We have recently described a significant increase of circulating miR-150 serum concentrations 1 month post-vaccination, and shown to be the release of exosome-associated miRNAs by lymphocytes activated in vivo by vaccines. This study gives support to the idea that profiling of serum miRNA levels may lead to the identification of new biomarkers of immune responses and that exosome purification represents a facile yet powerful step toward increased sensitivity, with possible increased specificity.
In conclusion, the compilation of a catalog of exosome-associated miRNAs derived from specific lymphocyte subsets should help confirm whether serum miRNAs are indeed able to report the activation of specific T cell subsets (e.g., Th1, Th2, Th17, Treg, etc.) occurring at distant sites. These lines of enquiry should benefit the assessment of pathogenic immune response during the course of auto-immune diseases and their therapies, as well as significantly contributing to the close, rapid monitoring of clinical trials with new immune-regulatory drugs, new vaccines, and/or adjuvants, particularly in the earlier stages of clinical development.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Riccardo Rossi, Chiara Zingaretti, and Giuseppe Matarese for helpful discussions, and Christopher Walentas for constructive review of the manuscript. Our research is supported by a CARIPLO grant (no. 2009–3603) http://www.fondazionecariplo.it; an ERC advanced grant to SA (no. 269022) http://erc.europa.eu; and by Fondazione Invernizzi.
References
1. Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell (2009) 136:215–33. doi:10.1016/j.cell.2009.01.002
2. Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K, et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res (2008) 18:997–1006. doi:10.1038/cr.2008.282
3. Arroyo JD, Chevillet JR, Kroh EM, Ruf IK, Pritchard CC, Gibson DF, et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc Natl Acad Sci U S A (2011) 108:5003–8. doi:10.1073/pnas.1019055108
4. Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A (2008) 105:10513–8. doi:10.1073/pnas.0804549105
5. Boeri M, Verri C, Conte D, Roz L, Modena P, Facchinetti F, et al. MicroRNA signatures in tissues and plasma predict development and prognosis of computed tomography detected lung cancer. Proc Natl Acad Sci U S A (2011) 108:3713–8. doi:10.1073/pnas.1100048108
6. Moussay E, Wang K, Cho JH, van Moer K, Pierson S, Paggetti J, et al. MicroRNA as biomarkers and regulators in B-cell chronic lymphocytic leukemia. Proc Natl Acad Sci U S A (2011) 108:6573–8. doi:10.1073/pnas.1019557108
7. Taylor DD, Gercel-Taylor C. MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol Oncol (2008) 110:13–21. doi:10.1016/j.ygyno.2008.04.033
8. D’Alessandra Y, Devanna P, Limana F, Straino S, Di Carlo A, Brambilla PG, et al. Circulating microRNAs are new and sensitive biomarkers of myocardial infarction. Eur Heart J (2010) 31:2765–73. doi:10.1093/eurheartj/ehq167
9. Goretti E, Vausort M, Wagner DR, Devaux Y. Association between circulating microRNAs, cardiovascular risk factors and outcome in patients with acute myocardial infarction. Int J Cardiol (2013) 168:4548–50. doi:10.1016/j.ijcard.2013.06.092
10. Guay C, Regazzi R. Circulating microRNAs as novel biomarkers for diabetes mellitus. Nat Rev Endocrinol (2013) 9:513–21. doi:10.1038/nrendo.2013.86
11. Starkey Lewis PJ, Dear J, Platt V, Simpson KJ, Craig DG, Antoine DJ, et al. Circulating microRNAs as potential markers of human drug-induced liver injury. Hepatology (2011) 54:1767–76. doi:10.1002/hep.24538
12. Chen YJ, Zhu JM, Wu H, Fan J, Zhou J, Hu J, et al. Circulating microRNAs as a fingerprint for liver cirrhosis. PLoS One (2013) 8:e66577. doi:10.1371/journal.pone.0066577
13. Wang JF, Yu ML, Yu G, Bian JJ, Deng XM, Wan XJ, et al. Serum miR-146a and miR-223 as potential new biomarkers for sepsis. Biochem Biophys Res Commun (2010) 394:184–8. doi:10.1016/j.bbrc.2010.02.145
14. Roderburg C, Luedde M, Vargas Cardenas D, Vucur M, Scholten D, Frey N, et al. Circulating microRNA-150 serum levels predict survival in patients with critical illness and sepsis. PLoS One (2013) 8:e54612. doi:10.1371/journal.pone.0054612
15. Reid G, Kirschner MB, van Zandwijk N. Circulating microRNAs: association with disease and potential use as biomarkers. Crit Rev Oncol Hematol (2011) 80:193–208. doi:10.1016/j.critrevonc.2010.11.004
16. Stoorvogel W, Kleijmeer MJ, Geuze HJ, Raposo G. The biogenesis and functions of exosomes. Traffic (2002) 3:321–30. doi:10.1034/j.1600-0854.2002.30502.x
17. Raposo G, Stoorvogel W. Extracellular vesicles: exosomes, microvesicles, and friends. J Cell Biol (2013) 200:373–83. doi:10.1083/jcb.201211138
18. Vickers KC, Palmisano BT, Shoucri BM, Shamburek RD, Remaley AT. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat Cell Biol (2011) 13:423–33. doi:10.1038/ncb2210
19. Valadi H, Ekström K, Bossios A, Sjöstrand M, Lee JJ, Lötvall JO. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol (2007) 9:654–9. doi:10.1038/ncb1596
20. Thery C, Ostrowski M, Segura E. Membrane vesicles as conveyors of immune responses. Nat Rev Immunol (2009) 9:581–93. doi:10.1038/nri2567
21. Bobrie A, Colombo M, Raposo G, Thery C. Exosome secretion: molecular mechanisms and roles in immune responses. Traffic (2011) 12:1659–68. doi:10.1111/j.1600-0854.2011.01225.x
22. Raposo G, Nijman HW, Stoorvogel W, Liejendekker R, Harding CV, Melief CJ, et al. B lymphocytes secrete antigen-presenting vesicles. J Exp Med (1996) 183:1161–72. doi:10.1084/jem.183.3.1161
23. Blanchard N, Lankar D, Faure F, Regnault A, Dumont C, Raposo G, et al. TCR activation of human T cells induces the production of exosomes bearing the TCR/CD3/zeta complex. J Immunol (2002) 168:3235–41.
24. Taylor DD, Akyol S, Gercel-Taylor C. Pregnancy-associated exosomes and their modulation of T cell signaling. J Immunol (2006) 176:1534–42.
25. Simons M, Raposo G. Exosomes – vesicular carriers for intercellular communication. Curr Opin Cell Biol (2009) 21:575–81. doi:10.1016/j.ceb.2009.03.007
26. Xiao C, Rajewsky K. MicroRNA control in the immune system: basic principles. Cell (2009) 136:26–36. doi:10.1016/j.cell.2008.12.027
27. Bronevetsky Y, Ansel KM. Regulation of miRNA biogenesis and turnover in the immune system. Immunol Rev (2013) 253:304–16. doi:10.1111/imr.12059
28. Mittelbrunn M, Gutiérrez-Vázquez C, Villarroya-Beltri C, González S, Sánchez-Cabo F, González MÁ, et al. Unidirectional transfer of microRNA-loaded exosomes from T cells to antigen-presenting cells. Nat Commun (2011) 2:282. doi:10.1038/ncomms1285
29. Gutierrez-Vazquez C, Villarroya-Beltri C, Mittelbrunn M, Sanchez-Madrid F. Transfer of extracellular vesicles during immune cell-cell interactions. Immunol Rev (2013) 251:125–42. doi:10.1111/imr.12013
30. Lotvall J, Valadi H. Cell to cell signalling via exosomes through esRNA. Cell Adh Migr (2007) 1:156–8. doi:10.4161/cam.1.3.5114
31. Nolte-‘t Hoen EN, Buermans HP, Waasdorp M, Stoorvogel W, Wauben MH, ‘t Hoen PA. Deep sequencing of RNA from immune cell-derived vesicles uncovers the selective incorporation of small non-coding RNA biotypes with potential regulatory functions. Nucleic Acids Res (2012) 40:9272–85. doi:10.1093/nar/gks658
32. de Candia P, Torri A, Gorletta T, Fedeli M, Bulgheroni E, Cheroni C, et al. Intracellular modulation, extracellular disposal and serum increase of MiR-150 mark lymphocyte activation. PLoS One (2013) 8:e75348. doi:10.1371/journal.pone.0075348
33. Rossi RL, Rossetti G, Wenandy L, Curti S, Ripamonti A, Bonnal RJ, et al. Distinct microRNA signatures in human lymphocyte subsets and enforcement of the naive state in CD4+ T cells by the microRNA miR-125b. Nat Immunol (2011) 12:796–803. doi:10.1038/ni.2057
34. Pagani M, Rossetti G, Panzeri I, de Candia P, Bonnal RJ, Rossi RL, et al. Role of microRNAs and long-non-coding RNAs in CD4(+) T-cell differentiation. Immunol Rev (2013) 253:82–96. doi:10.1111/imr.12055
36. He Y, Jiang X, Chen J. The role of miR-150 in normal and malignant hematopoiesis. Oncogene (2013). doi:10.1038/onc.2013.346
37. Monticelli S, Ansel KM, Xiao C, Socci ND, Krichevsky AM, Thai TH, et al. MicroRNA profiling of the murine hematopoietic system. Genome Biol (2005) 6:R71. doi:10.1186/gb-2005-6-8-r71
38. Salomoni P, Perrotti D, Martinez R, Franceschi C, Calabretta B. Resistance to apoptosis in CTLL-2 cells constitutively expressing c-Myb is associated with induction of BCL-2 expression and Myb-dependent regulation of bcl-2 promoter activity. Proc Natl Acad Sci U S A (1997) 94:3296–301. doi:10.1073/pnas.94.7.3296
39. Xiao C, Calado DP, Galler G, Thai TH, Patterson HC, Wang J, et al. MiR-150 controls B cell differentiation by targeting the transcription factor c-Myb. Cell (2007) 131:146–59. doi:10.1016/j.cell.2007.07.021
40. Ghisi M, Corradin A, Basso K, Frasson C, Serafin V, Mukherjee S, et al. Modulation of microRNA expression in human T-cell development: targeting of NOTCH3 by miR-150. Blood (2011) 117:7053–62. doi:10.1182/blood-2010-12-326629
41. Grusby MJ, Johnson RS, Papaioannou VE, Glimcher LH. Depletion of CD4+ T cells in major histocompatibility complex class II-deficient mice. Science (1991) 253:1417–20. doi:10.1126/science.1910207
42. Vesikari T, Pellegrini M, Karvonen A, Groth N, Borkowski A, O’Hagan DT, et al. Enhanced immunogenicity of seasonal influenza vaccines in young children using MF59 adjuvant. Pediatr Infect Dis J (2009) 28:563–71. doi:10.1097/INF.0b013e31819d6394
Keywords: T lymphocytes, circulating microRNAs, exosomes, vaccination
Citation: de Candia P, Torri A, Pagani M and Abrignani S (2014) Serum microRNAs as biomarkers of human lymphocyte activation in health and disease. Front. Immunol. 5:43. doi: 10.3389/fimmu.2014.00043
Received: 16 November 2013; Accepted: 27 January 2014;
Published online: 10 February 2014.
Edited by:
Paola Zanovello, University of Padova, ItalyReviewed by:
Donna M. D’Agostino, University of Padova, ItalyFlavia Bazzoni, University of Verona, Italy
Copyright: © 2014 de Candia, Torri, Pagani and Abrignani. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Paola de Candia and Sergio Abrignani, Istituto Nazionale Genetica Molecolare “Romeo ed Enrica Invernizzi,” Via F. Sforza 35, Milan 20122, Italy e-mail:ZGVjYW5kaWFAaW5nbS5vcmc=;YWJyaWduYW5pQGluZ20ub3Jn
 Paola de Candia
Paola de Candia Anna Torri
Anna Torri Massimiliano Pagani
Massimiliano Pagani Sergio Abrignani
Sergio Abrignani