- 1Molecular Immunology Division, Walter and Eliza Hall Institute of Medical Research, Melbourne, VIC, Australia
- 2Department of Medical Biology, The University of Melbourne, Melbourne, VIC, Australia
The Oct2 protein, encoded by the Pou2f2 gene, was originally predicted to act as a DNA binding transcriptional activator of immunoglobulin (Ig) in B lineage cells. This prediction flowed from the earlier observation that an 8-bp sequence, the “octamer motif,” was a highly conserved component of most Ig gene promoters and enhancers, and evidence from over-expression and reporter assays confirmed Oct2-mediated, octamer-dependent gene expression. Complexity was added to the story when Oct1, an independently encoded protein, ubiquitously expressed from the Pou2f1 gene, was characterized and found to bind to the octamer motif with almost identical specificity, and later, when the co-activator Obf1 (OCA-B, Bob.1), encoded by the Pou2af1 gene, was cloned. Obf1 joins Oct2 (and Oct1) on the DNA of a subset of octamer motifs to enhance their transactivation strength. While these proteins variously carried the mantle of determinants of Ig gene expression in B cells for many years, such a role has not been borne out for them by characterization of mice lacking functional copies of the genes, either as single or as compound mutants. Instead, we and others have shown that Oct2 and Obf1 are required for B cells to mature fully in vivo, for B cells to respond to the T cell cytokines IL5 and IL4, and for B cells to produce IL6 normally during a T cell dependent immune response. We show here that Oct2 affects Syk gene expression, thus influencing B cell receptor signaling, and that Oct2 loss blocks Slamf1 expression in vivo as a result of incomplete B cell maturation. Upon IL4 signaling, Stat6 up-regulates Obf1, indirectly via Xbp1, to enable plasma cell differentiation. Thus, Oct2 and Obf1 enable B cells to respond normally to antigen receptor signals, to express surface receptors that mediate physical interaction with T cells, or to produce and respond to cytokines that are critical drivers of B cell and T cell differentiation during a humoral immune response.
Introduction
Octamer binding protein 2, or Oct2, is encoded by the Pou2f2 gene. It was one of the first cell type-specific transcription factors identified and cloned (1). As indicated by its name, it is a founding member of a family of DNA binding proteins concurrently discovered, that share a conserved bipartite DNA binding domain comprising a homeobox-like domain and a second conserved sequence entitled the POU domain, for the Pit1, Oct1/Oct2, Unc86 proteins (2). Oct2 binds to a conserved consensus DNA sequence, the “octamer motif” found in the promoters and enhancers of many genes, including those encoding immunoglobulins (3, 4). The Obf1 protein encoded by the Pou2af1 gene, which is also known as OCA-B and Bob.1 was subsequently cloned using a yeast 1-hybrid screen for B cell proteins that physically interact with Oct1 or Oct2 (5–7). While Oct1/Oct2 and Obf1 share the capacity to bind to and activate genes adjacent to octamer motifs, they are selective in the genes to which they bind. The selectivity of target gene binding is determined, in part, by the sequence of the octamer motif, and whether it conforms to one of two classes of site, designated “PORE” and “MORE” motifs (8). Whether binding mediates activation or repression is also influenced by the participation of cofactors [reviewed by Tantin (9)], including Obf1, which can potentiate the transactivation potential of Oct1 and Oct2 (8, 10).
Oct2 is expressed primarily but not exclusively in the B cell lineage, where it increases with cellular activation (11). Neurons, macrophages, and T cells have also been shown to express Oct2 (12–18). Oct2 is required for post-natal survival (19), so must regulate critically important genes outside of the immune system. These will not be discussed here. The Oct2 gene is large, displays complex splicing patterns, and encodes protein isoforms with multiple essential activation domains (20–22). Oct2 is largely localized to the nucleus. Obf1 expression is mostly restricted to B lineage cells, where it is also highly induced upon activation (23). Zwilling et al. (24) have reported expression in T cells, but myeloid cells do not express Obf1 (15). A small protein of ~35 kDa, Obf1 is found in both the nucleus and cytoplasm, where a proportion may be tethered to the cell membrane after post-translational myristoylation (25), and a potential role for membrane-associated Obf1 in B cell receptor (BCR) signaling has been proposed (26).
A series of studies have shown that Oct2 and Obf1 are required for full functional and phenotypic maturation of B cells. In single knockout (KO) mice of each gene, peripheral B cells are numerically reduced and display some features of immature transitional cells (27, 28). The peritoneal B1 and splenic marginal zone (MZ) populations are missing in Oct2−/− mice (27, 29). Obf1−/− mice are viable and fertile, but show B cell developmental defects (30, 31), have an expanded B1 cell population (32). They also lack MZ B cells (33) and completely fail to produce germinal centers (GCs), the sites of cognate B cell:T cell interaction and expansion, upon immunization, or infection (34–37). Both Oct2- and Obf1-deficient splenic B cells display aberrant responses to BCR signaling and other characteristics of immature B cells (27, 34, 38). Oct2-deficient B cells also fail to respond to lipopolysaccharide (LPS), which signals through TLR4 (38). In vivo, serum immunoglobulin (Ig) levels in both mutants, particularly those that are T cell dependent, are strongly reduced (34–36, 38). Mice doubly deficient for Oct2 and Obf1 show a stronger humoral deficiency phenotype, reflecting the distinct activities of the two factors, but still express Ig genes (39). Thus, the two factors are not required, singly or in combination, for Ig gene expression by B cells.
Detailed functional studies on Oct2- and Obf1-deficient B cells in vitro and in vivo have identified a number of genes regulated by the two factors. Oct2 directly regulates the gene encoding CD36, a class B scavenger receptor family (40), but only in B cells, not in macrophages or dendritic cells (41–43). However, no role for CD36 in B cells has been determined (44). Oct2-deficent B cells have been shown to be defective in their responses to the T cell cytokine IL5 as a result of the direct regulation of the Cd125 gene encoding the IL5Rα chain (29). IL5 promotes antibody-secreting cell (ASC) differentiation in mouse B cells (45), and Oct2−/− B cells are defective in this process (29). In another study, it was shown that both Oct2 and Obf1 contribute to the regulation of IL6 production by activated B cells, through direct effects, at least by Oct2, on the Il6 gene (11). As Obf1 does not contact DNA (5), it is difficult using current procedures to prove direct interaction of Obf1 with putative target gene loci. IL6 is important during T follicular helper (Tfh) cell polarization (46). We have also shown, using the same quantitative tools that identified the role of Oct2 and IL5 in ASC differentiation, that Obf1 is required for T cell dependent ASC differentiation, but not isotype switching, both in vitro and in vivo (29).
In addition to these established roles for Oct2 and Obf1 in B cells, we include below data from studies on other genes that we have found to be differentially regulated in Oct2- and Obf1-deficient B cells. These include the genes encoding the Syk protein, which is an important transducer of BCR signals and Slamf1, an essential mediator of cell:cell contact, especially in the context of a developing GC. Expression of Syk and Slamf1 are sensitive to Oct2 loss, through different mechanisms. We also show that Obf1 is downstream of Stat6 in the IL4 signaling pathway of B cells, with Xbp1, another Stat6 target, its direct activator. We include these data to add to our understanding of the valuable roles that Oct2 and Obf1 play in B cell responses to antigen and to T cell help.
Results
Consistent with their distinct roles in vivo, Oct2 and Obf1 have quite distinct patterns of expression in peripheral B cells, as measured by RNAseq of sorted populations from naive C57BL/6 mice (Figure 1A). Oct2 levels are highest in B1 cells of the peritoneal cavity, and decline with terminal differentiation to ASCs. In contrast, Obf1 levels peak in GC B cells, which require Obf1 for their generation, and remain high in ASC. For contrast, expression of Syk and Slamf1 in the same populations are shown in Figure 1A, as these two genes are influenced directly and indirectly, respectively, by Oct2, and will be discussed below.
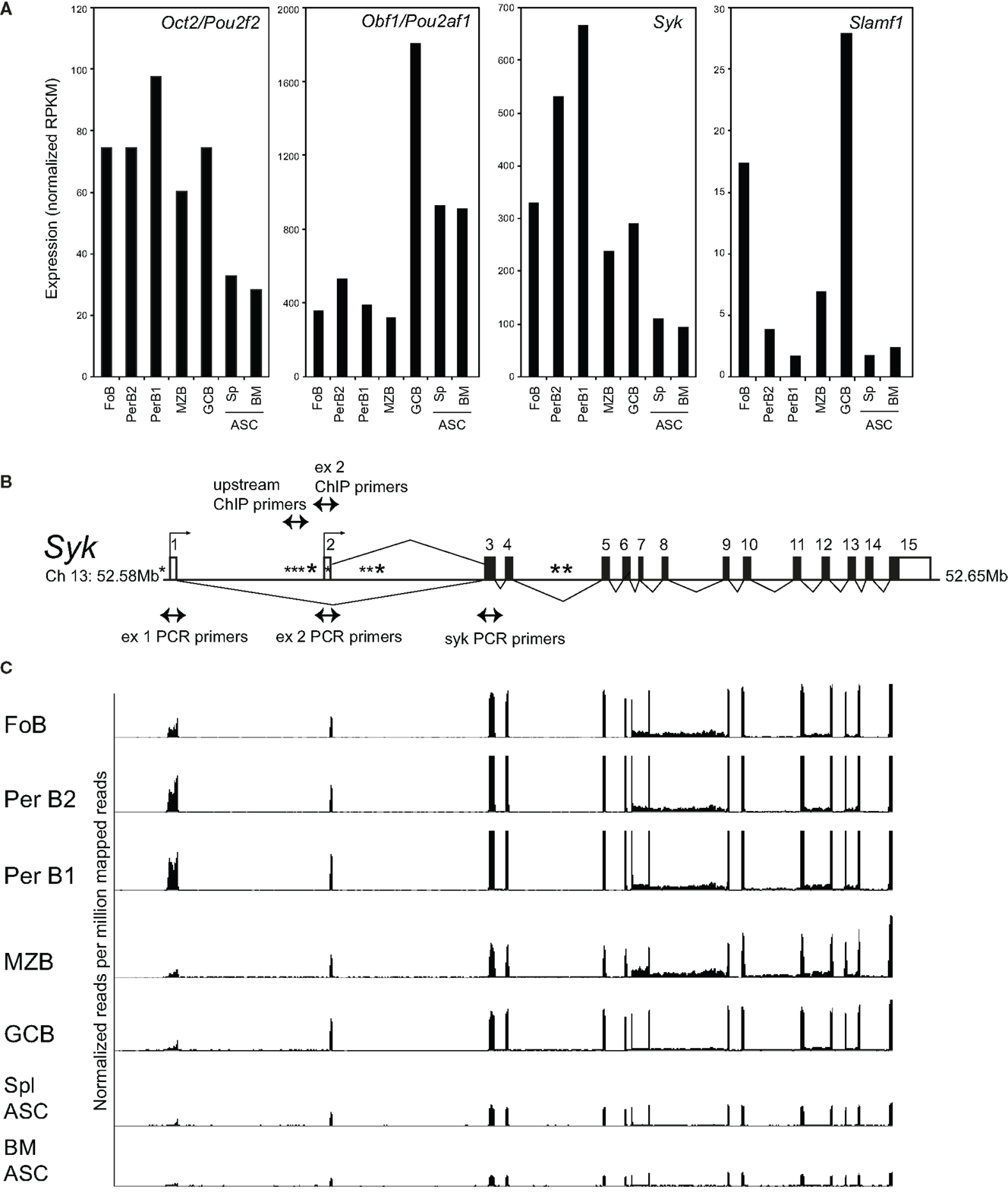
Figure 1. Expression of Oct2, Obf1, Syk, and Slamf1 in peripheral B cell populations. (A) RNAseq data measuring expression of Oct2/Pou2f2, Obf1/Pou2af1, Syk, and Slamf1 in B cell populations sorted ex vivo from naïve C57BL/6 mice. FoB, follicular B cells from spleen (small B220+, IgM+, IgD+) PerB1 and PerB2, B220+ cells from peritoneal lavages of naïve mice, stained with CD23 and Mac1. B1 cells were CD23− and Mac1lo and B2 cells were CD23+ and Mac1−; MZB, splenic marginal zone B cells, B220+, IgMhi, CD21hi; GCB, germinal center B cells (B220+, Fas+, GL7+) from spleens of mice immunized 8 days previously with SRBC; ASC, antibody-secreting cells sorted as syndecan1+, GFP+ cells from spleens (Spl), and bone marrows (BM) of mice carrying the Blimp-GFP reporter gene (47). Data were derived from at least two independent biological replicates in all cases. Because Ig sequences can represent >70% of the RNA from plasma cells (data not shown), the RNAseq data shown in the figure excludes all reads mapping to the Ig (heavy and light chain) loci as described in Section “Materials and Methods.” (B) Structure of the mouse Syk gene, showing exons, alternative transcriptional start sites (small arrows), the locations of a perfect consensus octamer motif (*) and the positions of PCR primers used here. Filled boxes indicate protein coding sequence, and open boxes, sequence comprising the 5′ and 3′ untranslated regions of Syk mRNA. (C) RNAseq tracks showing expression of the Syk gene exons in different sorted B cell populations, normalized to library size, and aligned with the gene structure of (B). Note that exon 1, as shown in this panel, is not included in the RefSeq (Mouse mm9, July 2007) map of Syk mRNA, but is represented in alternate Syk transcripts ENSMUST00000120135 and ENSMUST00000118756 in the Ensembl database.
Oct2 Modulates B Cell Receptor Signaling by Fine-Tuning Syk Expression
A microarray screen for Oct2-dependent genes identified the tyrosine kinase Syk as a potential target gene. Characterization of the murine Syk promoter using 5′ RACE identified two alternative transcriptional initiation sites (Figure 1B) that append alternative 5′ non-coding exons to Syk mRNAs in B cells. Both transcripts encode the same protein, as the start of Syk translation lies in an exon common to the two transcripts. RNAseq data for Syk in sorted B cell populations show that the usage of exons 1 or 2 varies subtly among them (Figure 1C).
The promoter upstream of exon 2 is positively regulated by Oct2. B cells sorted from Oct2−/− mice (Figure 2A) have a lower level of Syk transcripts and protein than wild type (WT) mice (Figures 2B,C). Using qPCR to distinguish Syk transcripts derived from exon 1 or exon 2, we found that those derived from exon 2 were selectively reduced in Oct2−/− B cells (Figure 2D). To confirm the influence of Oct2 on the Syk exon 2 promoter, we stably introduced an estradiol-inducible form of Oct2 (29) into a cloned Oct2−/− B lymphoma cell line, OM1 (48). Upon treatment with estradiol, there was no effect on expression from exon 1 with either the vector only control or the inducible Oct2 construct. However, estradiol induction of Oct2 selectively enhanced Syk expression from exon 2 and culminated in markedly increased Syk mRNA levels (Figures 2E,F). A DNA sequence search revealed three perfect consensus octamer sequences in the Syk gene (Figure 1B). Chromatin immunoprecipitation (ChIP) of Oct2 on DNA from the WT B lymphoma WEHI231 showed strong enrichment of DNA from the promoter of the known Oct2 target gene Cd36 (Figure 2G). The DNA adjacent to the perfect octamer sequence upstream of exon 2 was also significantly enriched in the ChIP, but adjacent sequences in intron 1/2 were not. Thus, Oct2 can directly increase Syk levels in B cells, acting at one of two alternative promoters.
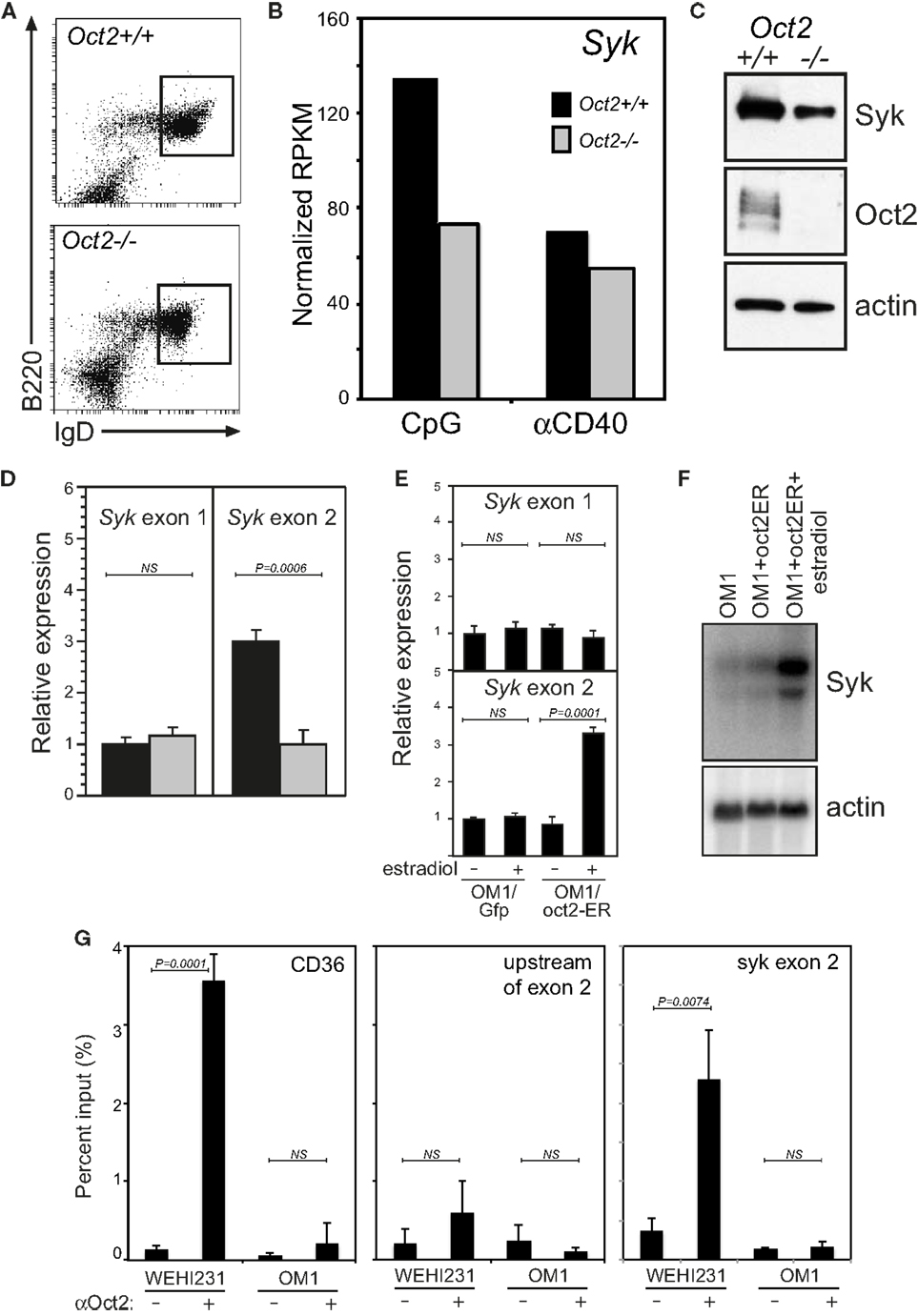
Figure 2. Oct2 directly and selectively activates transcription from Syk exon 2. (A) Splenic B cells from WT and Oct2 KO mice stained for B220 and IgD expression and sorted for phenotypically mature Fo B cells. (B) Syk RNAseq data from WT (black bars) and Oct2−/− (gray bars) B cells, sorted as in (A), activated for 48 h with CpG or anti-CD40. (C) Syk protein in sorted resting Fo B cells from Oct2+/+ or Oct2−/−mice. (D) qPCR of Syk mRNA distinguishing transcripts initiated at exons 1 or 2 in sorted splenic Fo B from WT and Oct2 KO mice. Expression is relative to that of the hmbs housekeeping gene. Values are means ± SD of triplicate assays. (E) Specific induction of transcription from Syk exon 2 upon Oct2 over-expression in OM1 cells, which are Oct2−/− (48), as shown by qPCR. Values are means ± SD of triplicates. (F) Northern blot for total Syk mRNA from a parallel experiment. (G) Quantitation of Oct2 chromatin immunoprecipitation (ChIP) qPCR data, showing enrichment of the Cd36 promoter [a known Oct2 target gene (41, 42)] and Syk sequences upstream of exon 2. Oct2:DNA complexes were precipitated from WEHI231 B lymphoma cells (which are Oct2+/+) and OM1 B lymphoma cells. Oct2 does not bind appreciatively to adjacent sequences in Syk intron 1/2. Values are means ± SD of triplicate assays. P values were calculated using the unpaired Student’s t-test; NS, not significant.
As mentioned above, Oct2 is required for the full functional and phenotypic maturation of B cells, such that peripheral B cells in Oct2−/− mice are numerically reduced and display some features of immature transitional cells, and the B1 and MZ populations are missing (27, 29, 38). Like transitional B cells, Oct2-deficient B cells are killed rather that activated by BCR cross-linking [Ref. (38, 49); see also Figure 3A]. However, Oct2−/− B cells respond normally to the survival factor Baff/BlyS, and survive to expand normally when BCR signaling occurs in the presence of Baff (Figures 3A,B and data not shown). Oct2−/− mice expressing a Bcl2 transgene still lack B1 and MZ B cells (data not shown). Thus abnormal survival properties are not responsible for the lack of these two populations in the Oct2 mutant mice.
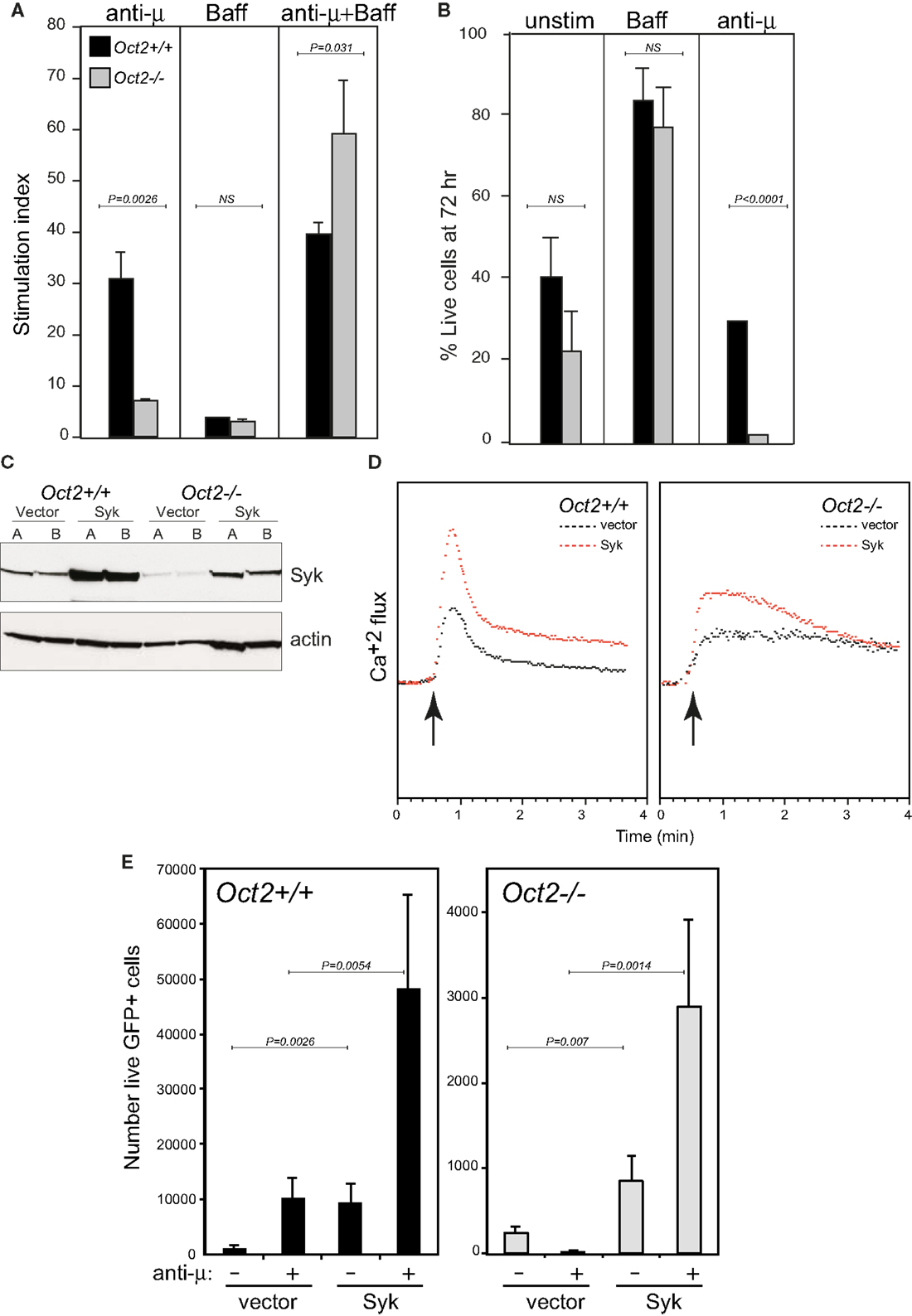
Figure 3. Oct2 is required for normal signaling from the B cell receptor, and ectopic Syk expression enhances the response in both WT and KO B cells. (A) B cell proliferation in response to BCR signaling, Baff, or both in combination after 3 days. Stimulation index is calculated as proliferation relative to unstimulated cells. Filled bars, Oct2+/+ gray bars, Oct2−/−. All values are the mean of triplicates ± SD. (B) Survival, assessed by propidium iodide exclusion, of B cells cultured with Baff or with anti-μ for 3 days. Filled bars, Oct2+/+ gray bars, Oct2−/−. All values are the mean of triplicates ± SD. (C) Syk protein levels in cloned B lymphoma cells transduced with a Syk-expressing retrovirus. The Oct2+/+ and Oct2−/−cell lines are BC1 and OM1, respectively (48). (D) Cytometric measurement of Ca2+ flux in clones of BC1 and OM1 cells transduced with vector only (black) or a Syk-expressing retrovirus (red). The arrow indicates the timing of addition of anti-μ to cross-link the BCR. (E) Primary splenic B cells from Oct2+/+ and Oct2−/−mice were activated, transduced (see Materials and Methods) and subsequently treated with anti-μ. The number of live, transduced (GFP+) cells in each culture after 48 h is shown. Values are means (n = 4) ± SD. Filled bars, Oct2+/+ gray bars, Oct2−/−.
We speculated that the reduced Syk levels in Oct2−/− B cells might contribute to their failure to mature in vivo and respond to BCR signals in vitro. We constructed a retroviral vector expressing Syk, and infected and cloned WT (BC1) and Oct2-deficient (OM1) lymphoma cells (Figure 3C). Boosting Syk protein levels enhanced the BCR response, as measured by calcium flux, in both WT and Oct2−/− cells (Figure 3D). Finally, using transduction of primary B cells (see Materials and Methods), we found that elevating Syk levels improved the proliferation of WT B cells, both unstimulated, and more strongly, upon BCR cross-linking, and that Oct2−/− B cells complemented with Syk retrovirus were activated to expand, rather than be killed by a BCR signal (Figure 3E). The rescue was not complete, as cell survival was still lower overall in the mutant cell cultures. This is likely to reflect technical limitations of the assay, including comparative infectivity of WT and mutant cells, and the correct timing of exogenous Syk expression in the context of the BCR signal. However, the results strongly suggest that Syk levels are limiting in Oct2−/− and, to a lesser extent, in WT B cells. We propose that Oct2 regulates Syk gene expression to enable positive selection through the BCR and therefore entrance to the mature follicular B cell pool, and it may similarly enable differentiation of B1 and MZ B cells, which are highly dependent on BCR signal strength.
Oct2 Indirectly and Selectively Regulates Slamf1 Expression on B Cells
Slamf1 encodes CD150, a lymphocyte signaling and adhesion molecule (50) that is expressed in B cells and T cells and at very low levels in myeloid cells (see www.immgen.org). Our RNAseq analysis of activated B cells indicated that Slamf1 was expressed at abnormally low levels in Oct2 KO B compared to controls (Figure 4A). This was confirmed by flow cytometric analysis (Figure 4B). Resting WT and LPS-activated B cells expressed similar levels of Slamf1, but levels on Oct2-deficient B cells were much lower than controls under both conditions. Activated WT T cells up-regulated Slamf1 from resting levels. However, Slamf1 was not Oct2-dependent in resting or activated T cells, or in macrophages expanded from fetal liver, despite Oct2 normally being expressed in these cell types (15, 16, 18).
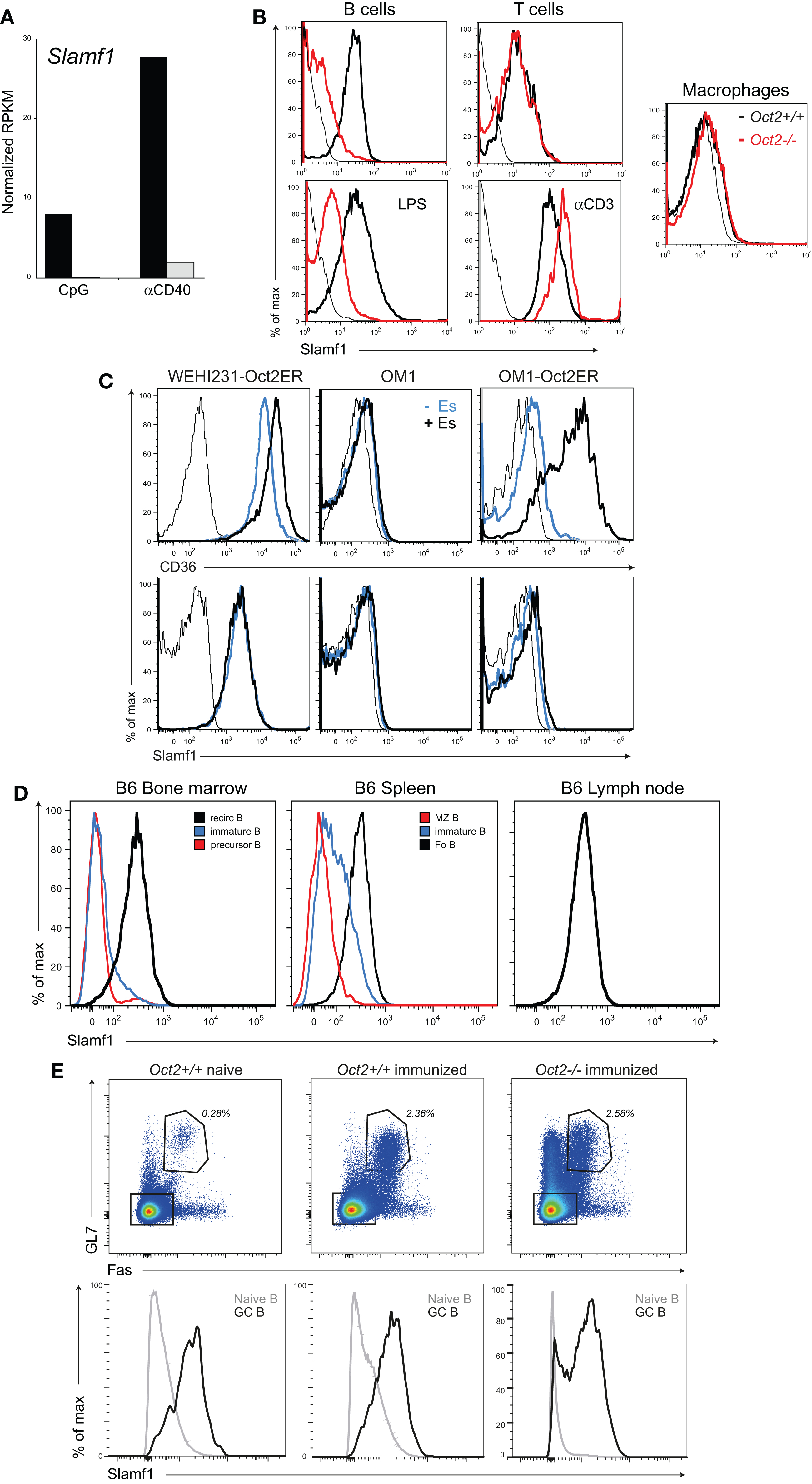
Figure 4. Oct2 influences and Slamf1 expression in B cells. (A) Analysis of Slamf1 expression in Oct2-null and Oct2+/+ B cells activated in vitro. Cells were cultured for 48 h in the presence of CpG or aCD40 before RNA was prepared for RNA sequencing. Filled bars, Oct2+/+ gray bars, Oct2−/−. (B) Slamf1/CD150 protein expression in cells of the indicated genotypes. Cells were assessed directly ex vivo (resting, top panels), or were activated in vitro for 48 h with either LPS (B cells) or anti-CD3 (T cells). Macrophages were expanded from fetal liver as described in Section “Materials and Methods.” In the histograms, WT cells are represented with heavy black lines and Oct2−/− as red. Unstained controls are indicated by thin black lines. (C) WEHI231 cells (Oct2+/+) and OM1 cells (Oct2−/−), either uninfected or transduced with an Oct-ER expression vector, were cultured in the presence or absence of estradiol (Es) for 24 h and CD36 and Slamf1 levels were determined by flow cytometry. Thin lines indicate background fluorescence of unstained controls, blue lines represent CD36 and Slamf1 levels in uninduced cultures, and heavy black lines, the levels after Es induction. (D) Slamf1 expression during B cell maturation. Colors indicate the populations represented in each histogram. For bone marrow, recirculating B cells were B220++ and IgM+, immature B were B220+ and IgM++, and precursor B were B220+ and IgM−. For spleen, immature B cells were IgMhi, IgDlo, MZ B cells were IgMhi, CD21hi, and Fo B cells were IgMlo, IgDhi, CD23+, and CD21+. Lymph node B cells were IgMlo, IgDhi, and CD23+. (E) Flow cytometric analysis of splenic B cells from naïve and immunized mice (9 days after SRBC immunization). Top panels show the percentage of GL7+Fas+ GC B cells (gated) among total B220+ cells in spleens from mice reconstituted with WT or Oct2−/− fetal liver. Bottom panels show the Slamf1 levels on non-GC (gray lines) and GC (black line) B cells for each animal, gated as shown in the upper panels.
We next asked whether Slamf1 was a direct Oct2 target using cloned WT (WEHI231) and KO (OM1) B lymphoma lines transduced with the inducible Oct2-ER vector, as described above. As expected, the Oct2 target Cd36 gene was expressed in WEHI231 but not OM1 (Figure 4C). Estradiol treatment enhanced Cd36 levels in WEHI231, and strongly induced Cd36 expression in OM1 cells. However, Slamf1 expression, while low in the Oct2−/− line, was not increased by Oct2 induction. Therefore, Oct2 is unlikely to directly regulate Slamf1 transcription in B cells.
Examination of the pattern of expression of the Slamf1 gene during B cell development, using the Immgen database (http://www.immgen.org) and by flow cytometry of bone marrow (BM) and peripheral B cell populations indicated that Slamf1 is a marker of B cell maturation, appearing during the transition from immature (IgMhi/IgDlo) to mature (IgMlo/IgDhi) follicular B cells of the spleen (Figure 4D), with MZ B cells expressing intermediate Slamf1 levels. We conclude that loss of Oct2 blocks B cell maturation before the Slamf1+ stage, thereby indirectly regulating its expression.
It has been shown that B cells lacking Slamf1 cannot form the lasting interactions with Tfh cells that are required for GC formation (51), and yet Oct2 mice do form GC upon infection and immunization (11). We immunized WT and KO mice with SRBC and stained GC cells for Slamf1 9 days later. Oct2+/+ and Oct2−/−GC B cells expressed similar levels of Slamf1. This indicates that Oct2 is dispensable for Slamf1 expression, and that endogenous signals driving the GC response override the Oct2 maturation defect in vivo (Figure 4E). To explore the nature of these signals, we tested the capacity of a number of mitogens and cytokines to induce Slamf1 expression on Oct2-null B cells in vitro. Anti-μ, anti-CD40, Baff, and IL4, tested singly or in all possible combinations, failed to induce appreciable Slamf1 on Oct2−/− B cells, suggesting that other factors and cells contribute in vivo. Collectively these studies point to an Oct2-regulated differentiation step that enables efficient FoB cell maturation.
Obf1 Enables T Cell Dependent ASC Differentiation Driven by IL4
We have shown that both Oct2 and Obf1 affect a B cell’s capacity to differentiate to ASC in response to particular cytokines, with Oct2 regulating the response to IL5 (29), and Obf1 being essential for ASC differentiation driven by IL4 (52).
We determined that Obf1 lies downstream of Stat6 in the IL4 signaling cascade (Figures 5A–C). Interestingly, while the IL4/Stat6 axis drives both isotype switching and ASC differentiation (45), Obf1 is dispensable for IL4 driven switching (52). In order to learn how IL4 and Stat6 regulate Obf1 expression, we performed microarray analysis on the WEHI231 B lymphoma, an IL4 responsive line. Because Stat6 exists in a latent form in the cytoplasm, and is activated by phosphorylation to enter the nucleus and act as a transcription factor (53), direct Stat6 targets would be activated after IL4 signaling even in the absence of new protein synthesis. Cells therefore were treated with cyclohexamide (CHX) to inhibit translation, and 1 h later, IL4 was added. A parallel culture did not receive IL4. After another 4 h, RNA was prepared from both. Analysis using Illumina Sentrix Mouse v1.1 arrays identified a small number of genes whose expression rose two to fourfold during this short period of IL4 stimulation, genes likely to be regulated directly by Stat6. Among these were three transcriptions factors: Nfil3, Vdr, encoding the 1,25-dihydroxyvitamin D3 receptor, and Xbp1. Their induction under these conditions was validated by qPCR (Figure 5D). Interestingly, Obf1 transcription was not elevated by IL4 treatment in the presence of CHX, indicating that it is not a direct Stat6 target gene (Figure 5D).
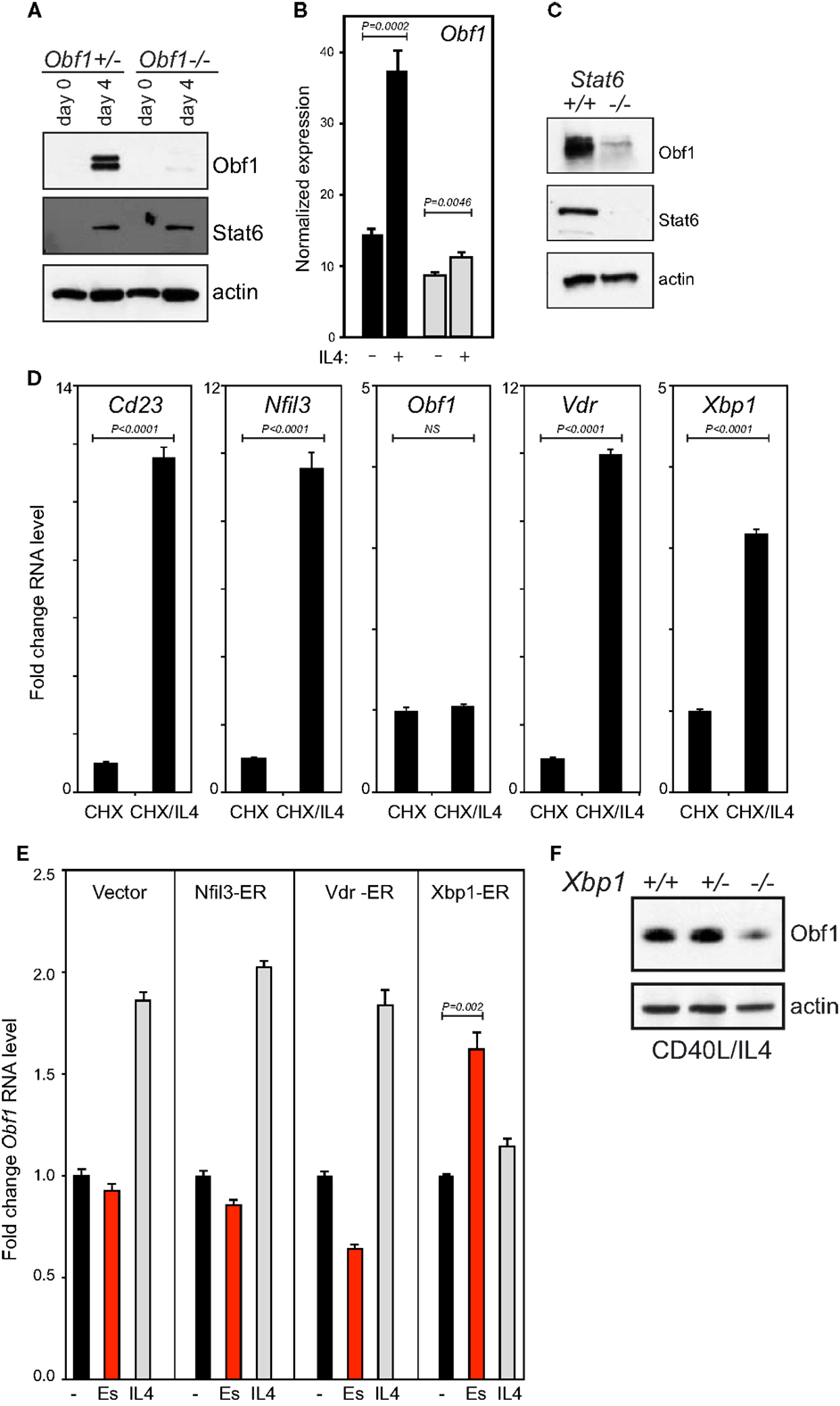
Figure 5. Obf1 is downstream of Stat6 in the IL4 signaling cascade. (A) Western blots to detect Obf1 and Stat6 proteins in purified B cells of the indicated genotypes, harvested 48 h after anti-CD40 ± IL4 treatment. Actin is included as a loading control. (B) RNA sequencing data showing Obf1 expression 48 h after anti-CD40 ± IL4 treatment in WT (black bars) and Stat6−/− B cells (gray bars). (C) Western blot on the same samples as in (B), to detect Stat6 and Obf1 proteins. (D) Induction of IL4 responsive genes in the presence of the translational inhibitor cycloheximide (CHX), as measured by qPCR. (E) Induction of Obf1 RNA levels in B lymphoma cell lines each expressing a ER fusion of each of the transcriptional regulators that are direct IL4 target genes: Nfil3, Vdr, and Xbp1. Cells were untreated or treated with either estradiol or IL4 for 24 h. In (B,D,E), values are means ± SD for triplicate samples. (F) Western blot to detect Obf1 protein in purified B cells from Xbp1+/+, heterozygous, or conditional KO mice. Cells were cultured in anti-CD40 plus IL4 for 48 h before analysis. In (B,D,E), means ± SD for triplicate measurements are shown, with P values determined using the unpaired Student’s t-test. NS, not significant.
As Obf1 RNA is elevated by IL4 treatment in B cells in the absence of CHX (Figure 5B), we hypothesized that one of the three transcription factors directly regulated by Stat6 might drive Obf1 expression. We therefore constructed ER fusion vectors for each to determine their effects on Obf1 expression. The Xbp1-ER expression vector contained the mature, processed, and active form of this factor (54). Clones of each line were cultured for 24 h unstimulated, with estradiol to induce each fusion protein, or with IL4. IL4 caused an increase in Obf1 RNA levels in all cases, as expected. However, estradiol induction only increased Obf1 levels significantly in the Xbp-ER expressing cell line (Figure 5E). Finally, primary splenic B cells from Xbp1+/+, heterozygous and conditional KO mice were stimulated for 48 h with CD40 ligand and IL4, and protein extracts prepared. Western blots showed that Xbp1-null B cells express markedly less Obf1 than the controls (Figure 5F). These data indicate that Nfil3 and Vdr do not influence Obf1 expression, but that Xbp1 has the capacity to directly activate the Obf1 gene in response to IL4/Stat6 signaling in B cells.
Discussion
The data we present here adds to a growing view of Oct2 and Obf1 as essential contributors to the sensing capacity of B cells (Figure 6). These two factors enhance the cell’s ability to deliver a BCR signal to drive maturation, or to sense a foreign antigen and become activated. We show here that Oct2 may do so by fine-tuning Syk levels. Since Oct2 loss blocks peripheral B cell maturation, such that MZ B cells are missing and mature FoB reduced, Oct2 may play its most important role prior to a divergence point of Fo and MZ B cells (55). The immature Slamf1−, BaffR+, CD23+, CD21+, IgMhi, Dlo, HSAhi phenotype of Oct2−/− B cells does not neatly fit into the phenotypic transition of immature to mature B cells (55), and may represent a normally transient phase of B cell maturation. Obf1 is required for normal B cell maturation and MZ B cell development (33), but as Obf1 does not influence Syk expression, it is likely that the two factors act through unique subsets of target genes for this aspect of B cell development. Indeed, it has been reported that Obf1 positively regulates SpiB gene expression (56), and that SpiB is required for normal B cell maturation and BCR signaling, through regulation of c-rel (57, 58).
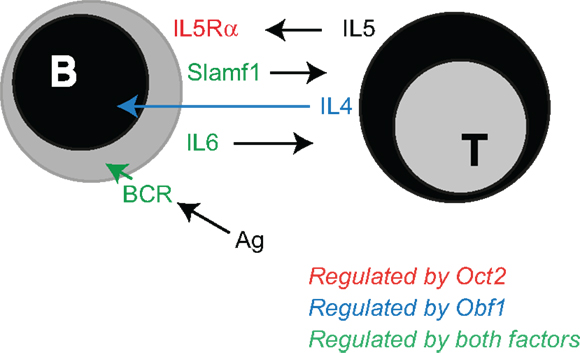
Figure 6. Oct2 and Obf1: finessing the B cell. Summary of published work and data discussed here, identifying the B cell responses that depend upon Oct2 and Obf1 for their optimal function. Oct2 (red arrow) selectively enables B cells to respond to IL5, a T cell-derived driver of ASC differentiation (45). Obf1 (blue arrow) selectively mediates IL4 driven ASC differentiation, downstream of Stat6. Both Oct2 and Obf1 (green arrows) enable BCR signaling, B cell maturation and indirectly, expression of Slamf1, an important mediator of B cell:T cell interactions during a humoral immune response.
T cell dependent antibody responses depend upon Oct2 and Obf1 in several ways. Both Oct2 and Obf1 are required for B cells to produce normal levels of IL6, a cytokine important in Tfh maturation in the context of a Slamf1-mediated B cell:Tfh cell interaction (11, 59, 60). These molecular interactions enable the initiation of a fruitful T cell-dependent humoral immune response. For example, IL6 produced by activated B cells early in the GC response reinforces the early dendritic cell signal that initiates Tfh differentiation, and IL21 produced by the nascent Tfh enhances both Tfh function and B cell differentiation into GC B cells and ASC (61–66). Slamf1, which is indirectly dependent upon Oct2, is required to prolong the B:Tfh interaction while these important signals are exchanged. Subsequently, Oct2 and Obf1 enable B cells to respond to other Th cell cytokines that drive ASC differentiation. Oct2 regulates Cd125 expression (29), and so expression of the high affinity receptor for IL5, an ASC differentiation factor (45). Obf1 is required for a normal B cell response to IL4, a growth, survival, isotype switching and differentiation factor for B and plasma cells [(45, 67) and this study].
Except for the specific case of Obf1 and a subset of VL genes (68), there is no evidence that Oct2 or Obf1 are required for Ig gene expression or for antibody secretion in ASC generated from single or double KO mice (29, 39, 52). Interestingly, however, we show here that Xbp1, which is highly expressed in ASC and known to be required for ASC function [(69–72) and our unpublished data], can directly activate Obf1. Shen and Hendershot (73) have also shown in ASC that Obf1 is a direct target of Xbp1. Accordingly, Obf1 expression is elevated in normal ASC compared to B cells, unlike Oct2 expression, which declines with differentiation (Figure 1). Xbp1, normally associated with the unfolded protein response in ASC, is an IL4 response gene in B cells. We confirm here that Obf1 is an Xbp1 target gene in FoB cells, and that IL4 and Stat6 directly induce Xbp1. Iwakoshi et al. (70) have earlier shown that, in the context of ASC differentiation, IL4 strongly induces Xbp1 expression and Stat6 is required. Thus, the selective ASC differentiation defect in Obf1 null B cells under T cell-dependent conditions may reflect an important Stat6-Xbp1-Obf1 axis.
Instead of direct influences on Ig gene expression, poor humoral immune responses in Oct2 or Obf1 mutant mice more likely reflects a paucity of differentiated peripheral B cell populations and weak to absent influences of T cells on these B cells. Oct2 and Obf1 are most highly expressed in GC B cells, with Obf1 being essential for their differentiation, but Oct2 dispensable (11). In both mutants, the B cells defects are cell intrinsic. Outstanding questions remain: what are the genes that Oct2 and Obf1 regulate to ensure full peripheral maturation and competent BCR signaling? What are the Obf1 regulated genes that are essential for GC development under all circumstances tested so far, including immunization, infection, and autoimmunity (11, 32, 37)? While these critical Obf1 target(s) are not yet known, candidates such as Bcl6 (or its co-repressor MTA3), Bach2, Irf4 and CXCR5, all critical for normal GC formation or maintenance [see Ref. (74)], can be excluded, as they are not influenced by Obf1 loss (our unpublished data). Ongoing genome-wide RNA expression analysis, coupled with ChIPseq, will enable detailed characterization of the full Oct2 and Obf1 gene regulatory networks, and their shared and unique responsibilities in delivering effective B cell immunity.
Materials and Methods
Cell Lines, Cell Culture, and Retroviral Transduction
B lymphoma cell lines used here were all generated in house: WEHI231 (75), BC1, and OM1 (48). Primary splenic B and T cells were purified using anti-B220 or anti-CD4 microbeads (Miltenyi), as described (11). Oct2+/+ and Oct2−/−macrophages were expanded in vitro from E13 fetal liver using M-CSF, as described (15). Retroviral transduction in all cases used the pMX-pie vector and was preformed using spin infection as described (48, 76). For lymphoma lines, cells were cloned post-infection as single GFP+ cells in puromycin-supplemented medium.
For the retroviral complementation experiment of Figure 3E, primary splenic B220+ cells were stimulated for 24 h with CpG [1 μM oligonucleotide CpG 1668 (sequence 5′-TCCATGA CGTTCCTGATGCT-3′), fully phosphothioated GeneWorks] to promote cell cycling and enable retroviral infection. After overnight culture, cells were washed and resuspended in medium at 0.5 × 106 cells/ml, without CpG but containing anti-μ [10 μg/ml AffiniPure F(ab′)2 fragment, goat anti mouse Jackson Laboratories] and/or Baff (250 ng/ml a kind gift from Jürg Tschopp). Cell survival in transduced (GFP+) cells was assessed after a further 48 h by flow cytometry, propidium iodide exclusion, and cell counting using internal microbead controls.
Retroviruses expressing transcription factors fused to the human estrogen receptor (hER) dimerization domain were generated by amplification of each factor’s ORF, and sequencing each amplified product to ensure it was mutation-free and in frame with the hER. Production of ER fusion proteins of the correct size was confirmed by western blots of infected or transfected cells. Anti-CD40 (clone FGK4.5) was prepared in house and used at 10 μg/ml. β-Estradiol (Sigma) was used at 10 μM, cycloheximide at 50 μM.
Mice
Oct2−/−, Obf1−/−, and Stat6−/− mice have been described previously (19, 36) Xbp1fl/flCd19Cre/+mice, where Xbp1 is conditionally deleted in the B cell lineage, were generated by Hetz et al. (77) and further described in Taubenheim et al. (72). All mice were maintained on a C57BL/6 background, and all experiments conformed to the relevant regulatory standards of the Animal Ethics Committee of the Walter and Eliza Hall Institute of Medical Research (AEC Projects 2010.010 and 2013.014).
Antibodies
For westerns, antibodies used were specific for Oct2 (clone 9A2, in house), Obf1 (clone 6F10, in house) Stat6 (S-20), Syk (N-19), and actin (I-19) all from Santa Cruz and Xbp1 (Ab37152-100, Abcam). For flow cytometry, antibodies used were specific for B220 (RA3-6B2), IgD (11-26c.2a), fas/CD95 (Jo2), all from BD Pharmingen Slamf1/CD150 (TC15-12 F12.2, Biolegend), and GL7 (eBioscience). Calcium flux was assessed cytometrically as described (78).
Microarrays
Illumina Sentrix Mouse v1.1 arrays were probed with RNA prepared from independent B lymphoma cell lines: to identify Oct2-dependent genes, two independent clones of OM1 cells stably transduced with a control vector or one expressing an estradiol-inducible Oct-ER fusion protein were treated in vitro for 6 or 48 h with estradiol prior to RNA preparation. For Obf1 targets, two clones of the Obf1−/− BM1 lymphoma line (78), transduced with vector control or an Obf1-ER expression vector, were induced in a similar manner and RNA prepared and analyzed.
RNA Sequencing
Peripheral B cell populations were sorted from naïve or immunized C57Bl/6 mice as described in the legend to Figure 1. Two independent biological replicates were prepared for each population, except for spleen and BM ASC. Because of the paucity of ASC in these tissues, cells were pooled from three to four individuals before sorting. Two such pools were processed independently for sequencing. Normalized expression levels are shown in the graphs. As more than 70% of reads in RNA from ASC map to the Ig loci, all Ig reads were excluded from the data, for all populations, before normalization to generate the values shown in Figure 1A. Specifically, all reads from the IgH locus on chromosome 12, NC_000078.6 (positions 113,258,768–116,009,954), all reads from the Igλ locus on chromosome 16, NC_000082.6 (positions 19,026,858–19,260,844), and all reads from the Igκ locus on chromosome 6, NC_000072.6 (positions 67,555,636–70,726,754) were excluded. For Figure 2B, Oct2+/+ and Oct2−/−B cells were sorted, as in Figure 2A, from spleen of two independent mice of each genotype and activated for 48 h before RNA extraction and preparation for sequencing. As these conditions do not induce ASC differentiation, Ig sequence reads were not excluded from analysis of these samples.
Primers
The starts of Syk transcription in primary B cells were determined directly using FirstChoice® RLM-RACE (Ambion) and nested Syk gene specific primers specific for the first Syk coding exon: 5′-GTAGGTCAGGTGGTTGGCGCTGTCCACAGC-3′ and 5′-CCCGCCATGTCTGCACCCCTTCAGAGTTC-3′.
For exon-specific Syk qPCR, these primers were used:
5′Syk exon 1: 5′-CAGTGACTGCGGCTGAGCGCGGACC-3′
5′Syk exon 2: 5′-CAGCAGGAAACCTCCACTTGCTCTCC-3′
Common 3′ Syk primer: 5′-CCATGTCTGCACCCCTTCAGAGTTC-3′.
For Syk ChIP PCR, these primers were used:
Exon 2: fwd 5′-GCCTAGGCCACGATGGTCAAAGGAGG-3′ and
rev 5′-GGAGAGCAAGTGGAGGTTTCCTGCTG-3′.
Upstream of exon 2: fwd 5′-CCATTGGTGGGCCCTCAGCTTGGTTC-3′ and
rev 5′-GACCAGAGAAGAAATGGCCTCAGAAGACAGG-3′.
Chromatin immunoprecipitation was conducted as previously described (29), using high titer, polyclonal anti-Oct2 rabbit serum generated in house.
Primers for other qPCR were:
Stat6: fwd 5′-CTGCTGGGCCGAGGCTTCACATTT-3′
rev 5′-TCAGGGGCCATTCCAAGATCATAAGGT-3′
CD36: fwd 5′-GGAGGCATTCTCATGCCAGTCGGAGAC-3′
rev 5′-CAAAACTGTCTGTACACAGTGGTGCCTG-3′
Slamf1: fwd 5′-GGGAGCTATCCAGATCACCTG-3′
rev 5′-CGTTCTCCTCCACGCTCAC-3′
Cd23: fwd 5′-GACACTGCAATTCAGAATGTCTCTCATG-3′
rev 5′-GCTTCTGTTCAGCTTGGAGTTCTTGCAAG-3′
Nflil3: fwd 5′-CTCACGGACCAGGGAGCAGAACCACG-3′
rev 5′-CAGGTCTTAAGGACTTCAGCCTCTCATCC-3′
Obf1: fwd 5′-CGGTGTTGACCTATGCTTCTCCACC-3′
rev 5′-GAGGGGCGCCTGGTGCTCGGGACCC-3′
Vdr: fwd 5′-CGCTATGACCTGTGAAGGCTGCAAGGG-3′
rev 5′-GCCAATGTCCACGCAGCGTTTGAGCC-3′
Xbp1: fwd 5′-AGCAGCAAGTGGTGGATTTG-3′
rev 5′-CCAAGCGTGTTCTTAACTCCT-3′
Hmbs (for normalization): fwd 5′-GACCTGGTTGTTCACTCCCTGAAG-3′
rev 5′-GACAACAGCATCACAAGGGTTTTC-3′
Chromatin immunoprecipitation PCR and qPCR were performed on triplicate samples in all cases, and the data presented as means ± SD. Statistical significance was determined using the unpaired Student’s t-test.
Immunization
Mice stably reconstituted with Oct2+/+ or Oct2−/−fetal liver were immunized i.p with 2 × 109 sheep red blood cells (Applied Biological Product Management, Australia) in 100 μl of PBS and sacrificed after 9 days.
Author Contributions
Lynn Corcoran performed many of the experiments and wrote the manuscript; Dianne Emslie and Tobias Kratina together performed all of the qPCR and protein analyses; Wei Shi performed all bioinformatics analysis for the RNAseq experiments; Susanne Hirsch performed the Slamf1 studies; Nadine Taubenheim contributed to the Xbp1 studies; Stephane Chevrier led most of the flow cytometric analyses.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Drs. Laurie Glimcher for conditional Xbp1 mutant mice and Patrick Matthias for Obf1 mice, Gordon Smythe for bioinformatics assistance and Stephen Nutt for critical comments on the manuscript. Jennifer Vasiliadis and Louise Inglis provided expert animal care. This work was made possible through Victorian State Government Operational Infrastructure Support and Australian Government NHMRC IRIIS and research grants from the NHMRC (#637306 and #575500).
References
1. Staudt LM, Clerc RG, Singh H, Lebowitz JH, Sharp PA, Baltimore D. Cloning of a lymphoid-specific cDNA encoding a protein binding the regulatory octamer DNA motif. Science (1988) 241:577–80. doi: 10.1126/science.3399892
2. Herr W, Sturm RA, Clerc RG, Corcoran LM, Baltimore D, Sharp PA, et al. The POU domain: a large conserved region in the mammalian pit-1, oct-1, oct-2, and Caenorhabditis elegans unc-86 gene products. Genes Dev (1988) 2:1513–6. doi:10.1101/gad.2.12a.1513
3. Parslow TG, Blair DL, Murphy WJ, Granner DK. Structure of the 5’ ends of immunoglobulin genes: a novel conserved sequence. Proc Natl Acad Sci U S A (1984) 81:2650–4. doi:10.1073/pnas.81.9.2650
4. Landolfi NF, Capra JD, Tucker PW. Interaction of cell-type-specific nuclear proteins with immunoglobulin VH promoter region sequences. Nature (1986) 323:548–51. doi:10.1038/323548a0
5. Strubin M, Newell JW, Matthias P. OBF-1, a novel B cell-specific coactivator that stimulates immunoglobulin promoter activity through association with octamer-binding proteins. Cell (1995) 80:497–506. doi:10.1016/0092-8674(95)90500-6
6. Luo Y, Roeder RG. Cloning, functional characterization, and mechanism of action of the B-cell-specific transcriptional coactivator Oca-B. Mol Cell Biol (1995) 15:4115–24.
7. Gstaiger M, Georgiev O, Van Leeuwen H, Van Der Vliet P, Schaffner W. The B cell coactivator Bob1 shows DNA sequence-dependent complex formation with Oct-1/Oct-2 factors, leading to differential promoter activation. EMBO J (1996) 15:2781–90.
8. Tomilin A, Remenyi A, Lins K, Bak H, Leidel S, Vriend G, et al. Synergism with the coactivator OBF-1 (OCA-B, BOB-1) is mediated by a specific POU dimer configuration. Cell (2000) 103:853–64. doi:10.1016/S0092-8674(00)00189-6
9. Tantin D. Oct transcription factors in development and stem cells: insights and mechanisms. Development (2013) 140:2857–66. doi:10.1242/dev.095927
10. Lins K, Remenyi A, Tomilin A, Massa S, Wilmanns M, Matthias P, et al. OBF1 enhances transcriptional potential of Oct1. EMBO J (2003) 22:2188–98. doi:10.1093/emboj/cdg199
11. Karnowski A, Chevrier S, Belz GT, Mount A, Emslie D, D’Costa K, et al. B and T cells collaborate in antiviral responses via IL-6, IL-21, and transcriptional activator and coactivator, Oct2 and OBF-1. J Exp Med (2012) 209:2049–64. doi:10.1084/jem.20111504
12. Stoykova AS, Sterrer S, Erselius JR, Hatzopoulos AK, Gruss P. Mini-Oct and Oct-2c: two novel, functionally diverse murine Oct-2 gene products are differentially expressed in the CNS. Neuron (1992) 8:541–58. doi:10.1016/0896-6273(92)90282-I
13. Dent CL, Lillycrop KA, Estridge JK, Thomas NS, Latchman DS. The B-cell and neuronal forms of the octamer-binding protein Oct-2 differ in DNA-binding specificity and functional activity. Mol Cell Biol (1991) 11:3925–30.
14. Ninkina NN, Buchman VL, Akopian AN, Lawson SN, Yamamoto M, Campbell E, et al. Nerve growth factor-regulated properties of sensory neurones in Oct-2 null mutant mice. Brain Res Mol Brain Res (1995) 33:233–44. doi:10.1016/0169-328X(95)00128-F
15. Zhou L, Nazarian AA, Xu J, Tantin D, Corcoran LM, Smale ST. An inducible enhancer required for Il12b promoter activity in an insulated chromatin environment. Mol Cell Biol (2007) 27:2698–712. doi:10.1128/MCB.00788-06
16. Cockerill PN, Klinken SP. Octamer-binding proteins in diverse hemopoietic cells. Mol Cell Biol (1990) 10:1293–6.
17. Kamps MP, Corcoran L, Lebowitz JH, Baltimore D. The promoter of the human interleukin-2 gene contains two octamer-binding sites and is partially activated by the expression of Oct-2. Mol Cell Biol (1990) 10:5464–72.
18. Kang SM, Tsang W, Doll S, Scherle P, Ko HS, Tran AC, et al. Induction of the POU domain transcription factor Oct-2 during T-cell activation by cognate antigen. Mol Cell Biol (1992) 12:3149–54.
19. Corcoran LM, Karvelas M, Nossal GJ, Ye ZS, Jacks T, Baltimore D. Oct-2, although not required for early B-cell development, is critical for later B-cell maturation and for postnatal survival. Genes Dev (1993) 7:570–82. doi:10.1101/gad.7.4.570
20. Hatzopoulos AK, Stoykova AS, Erselius JR, Goulding M, Neuman T, Gruss P. Structure and expression of the mouse Oct2a and Oct2b, two differentially spliced products of the same gene. Development (1990) 109:349–62.
21. Annweiler A, Zwilling S, Wirth T. Functional differences between the Oct2 transactivation domains determine the transactivation potential of individual Oct2 isoforms. Nucleic Acids Res (1994) 22:4250–8. doi:10.1093/nar/22.20.4250
22. Corcoran LM, Koentgen F, Dietrich W, Veale M, Humbert PO. All known in vivo functions of the Oct-2 transcription factor require the C-terminal protein domain. J Immunol (2004) 172:2962–9.
23. Qin XF, Reichlin A, Luo Y, Roeder RG, Nussenzweig MC. Oca-B integrates B cell antigen receptor-, CD40L- and IL 4-mediated signals for the germinal center pathway of B cell development. EMBO J (1998) 17:5066–75. doi:10.1093/emboj/17.17.5066
24. Zwilling S, Dieckmann A, Pfisterer P, Angel P, Wirth T. Inducible expression and phosphorylation of coactivator BOB.1/OBF.1 in T cells. Science (1997) 277:221–5. doi:10.1126/science.277.5323.221
25. Yu X, Wang L, Luo Y, Roeder RG. Identification and characterization of a novel Oca-B isoform. Implications for a role in B cell signaling pathways. Immunity (2001) 14:157–67. doi:10.1016/S1074-7613(09)00090-9
26. Siegel R, Kim U, Patke A, Yu X, Ren X, Tarakhovsky A, et al. Nontranscriptional regulation of Syk by the coactivator Oca-B is required at multiple stages of B cell development. Cell (2006) 125:761–74. doi:10.1016/j.cell.2006.03.036
27. Humbert PO, Corcoran LM. Oct-2 gene disruption eliminates the peritoneal B-1 lymphocyte lineage and attenuates B-2 cell maturation and function. J Immunol (1997) 159:5273–84.
28. Samardzic T, Gerlach J, Muller K, Marinkovic D, Hess J, Nitschke L, et al. CD22 regulates early B cell development in BOB.1/OBF.1-deficient mice. Eur J Immunol (2002) 32:2481–9. doi:10.1002/1521-4141(200209)32:9<2481::AID-IMMU2481>3.0.CO;2-C
29. Emslie D, D’Costa K, Hasbold J, Metcalf D, Takatsu K, Hodgkin PO, et al. Oct2 enhances antibody-secreting cell differentiation through regulation of IL-5 receptor alpha chain expression on activated B cells. J Exp Med (2008) 205:409–21. doi:10.1084/jem.20072049
30. Hess J, Nielsen PJ, Fischer KD, Bujard H, Wirth T. The B lymphocyte-specific coactivator BOB.1/OBF.1 is required at multiple stages of B-cell development. Mol Cell Biol (2001) 21:1531–9. doi:10.1128/MCB.21.5.1531-1539.2001
31. Jankovic M, Nussenzweig MC. OcaB regulates transitional B cell selection. Int Immunol (2003) 15:1099–104. doi:10.1093/intimm/dxg109
32. Chevrier S, Kratina T, Emslie D, Karnowski A, Corcoran LM. Germinal center-independent, IgM-mediated autoimmunity in sanroque mice lacking OBF1. Immunol Cell Biol (2013) 92(1):12–9. doi:10.1038/icb.2013.71
33. Samardzic T, Marinkovic D, Nielsen PJ, Nitschke L, Wirth T. BOB.1/OBF.1 deficiency affects marginal-zone B-cell compartment. Mol Cell Biol (2002) 22:8320–31. doi:10.1128/MCB.22.23.8320-8331.2002
34. Kim U, Qin XF, Gong S, Stevens S, Luo Y, Nussenzweig M, et al. The B-cell-specific transcription coactivator OCA-B/OBF-1/Bob-1 is essential for normal production of immunoglobulin isotypes. Nature (1996) 383:542–7. doi:10.1038/383542a0
35. Nielsen PJ, Georgiev O, Lorenz B, Schaffner W. B lymphocytes are impaired in mice lacking the transcriptional co-activator Bob1/Oca-B/OBF1. Eur J Immunol (1996) 26:3214–8. doi:10.1002/eji.1830261255
36. Schubart DB, Rolink A, Kosco-Vilbois MH, Botteri F, Matthias P. B-cell-specific coactivator OBF-1/OCA-B/BOB1 required for immune response and germinal centre formation. Nature (1996) 383:538–42. doi:10.1038/383538a0
37. Fehr T, Lopez-Macias C, Odermatt B, Torres RM, Schubart DB, O’Keefe TL, et al. Correlation of anti-viral B cell responses and splenic morphology with expression of B cell-specific molecules. Int Immunol (2000) 12:1275–84. doi:10.1093/intimm/12.9.1275
38. Corcoran LM, Karvelas M. Oct-2 is required early in T cell-independent B cell activation for G1 progression and for proliferation. Immunity (1994) 1:635–45. doi:10.1016/1074-7613(94)90035-3
39. Schubart K, Massa S, Schubart D, Corcoran LM, Rolink AG, Matthias P. B cell development and immunoglobulin gene transcription in the absence of OCT-2 and OBF-1. Nat Immunol (2001) 2:69–74. doi:10.1038/83190
40. Greenwalt DE, Lipsky RH, Ockenhouse CF, Ikeda H, Tandon NN, Jamieson GA. Membrane glycoprotein CD36: a review of its roles in adherence, signal transduction, and transfusion medicine. Blood (1992) 80:1105–15.
41. Konig H, Pfisterer P, Corcoran LM, Wirth T. Identification of CD36 as the first gene dependent on the B-cell differentiation factor Oct-2. Genes Dev (1995) 9:1598–607. doi:10.1101/gad.9.13.1598
42. Shore P, Dietrich W, Corcoran LM. Oct-2 regulates CD36 gene expression via a consensus octamer, which excludes the co-activator OBF-1. Nucleic Acids Res (2002) 30:1767–73. doi:10.1093/nar/30.8.1767
43. Pfisterer P, Hess J, Wirth T. Identification of target genes of the lymphoid-specific transcription factor Oct2. Immunobiology (1997) 198:217–26. doi:10.1016/S0171-2985(97)80042-3
44. Corcoran L, Vremec D, Febbraio M, Baldwin T, Handman E. Differential regulation of CD36 expression in antigen-presenting cells: Oct-2 dependence in B lymphocytes but not dendritic cells or macrophages. Int Immunol (2002) 14:1099–104. doi:10.1093/intimm/dxf075
45. Hasbold J, Corcoran LM, Tarlinton DM, Tangye SG, Hodgkin PD. Evidence from the generation of immunoglobulin G-secreting cells that stochastic mechanisms regulate lymphocyte differentiation. Nat Immunol (2004) 5:55–63. doi:10.1038/ni1016
46. Nurieva RI, Chung Y, Hwang D, Yang XO, Kang HS, Ma L, et al. Generation of T follicular helper cells is mediated by interleukin-21 but independent of T helper 1, 2, or 17 cell lineages. Immunity (2008) 29:138–49. doi:10.1016/j.immuni.2008.05.009
47. Kallies A, Hasbold J, Tarlinton DM, Dietrich W, Corcoran LM, Hodgkin PD, et al. Plasma cell ontogeny defined by quantitative changes in blimp-1 expression. J Exp Med (2004) 200:967–77. doi:10.1084/jem.20040973
48. Corcoran LM, Tawfilis S, Barlow LJ. Generation of B lymphoma cell lines from knockout mice by transformation in vivo with an Emu-myc transgene. J Immunol Methods (1999) 228:131–8. doi:10.1016/S0022-1759(99)00094-0
49. Mackay F, Schneider P, Rennert P, Browning J. BAFF AND APRIL: a tutorial on B cell survival. Annu Rev Immunol (2003) 21:231–64. doi:10.1146/annurev.immunol.21.120601.141152
50. Schwartzberg PL, Mueller KL, Qi H, Cannons JL. SLAM receptors and SAP influence lymphocyte interactions, development and function. Nat Rev Immunol (2009) 9:39–46. doi:10.1038/nri2456
51. Cannons JL, Qi H, Lu KT, Dutta M, Gomez-Rodriguez J, Cheng J, et al. Optimal germinal center responses require a multistage T cell:B cell adhesion process involving integrins, SLAM-associated protein, and CD84. Immunity (2010) 32:253–65. doi:10.1016/j.immuni.2010.01.010
52. Corcoran LM, Hasbold J, Dietrich W, Hawkins E, Kallies A, Nutt SL, et al. Differential requirement for OBF-1 during antibody-secreting cell differentiation. J Exp Med (2005) 201:1385–96. doi:10.1084/jem.20042325
53. Wurster AL, Tanaka T, Grusby MJ. The biology of Stat4 and Stat6. Oncogene (2000) 19:2577–84. doi:10.1038/sj.onc.1203485
54. Tirosh B, Iwakoshi NN, Glimcher LH, Ploegh HL. Rapid turnover of unspliced XBP-1 as a factor that modulates the unfolded protein response. J Biol Chem (2006) 281:5852–60. doi:10.1074/jbc.M509061200
55. Cancro MP. Peripheral B-cell maturation: the intersection of selection and homeostasis. Immunol Rev (2004) 197:89–101. doi:10.1111/j.0105-2896.2004.0099.x
56. Bartholdy B, Du Roure C, Bordon A, Emslie D, Corcoran LM, Matthias P. The Ets factor Spi-B is a direct critical target of the coactivator OBF-1. Proc Natl Acad Sci U S A (2006) 103:11665–70. doi:10.1073/pnas.0509430103
57. Garrett-Sinha LA, Su GH, Rao S, Kabak S, Hao Z, Clark MR, et al. PU.1 and Spi-B are required for normal B cell receptor-mediated signal transduction. Immunity (1999) 10:399–408. doi:10.1016/S1074-7613(00)80040-0
58. Hu CJ, Rao S, Ramirez-Bergeron DL, Garrett-Sinha LA, Gerondakis S, Clark MR, et al. PU.1/Spi-B regulation of c-rel is essential for mature B cell survival. Immunity (2001) 15:545–55. doi:10.1016/S1074-7613(01)00219-9
59. Dienz O, Eaton SM, Bond JP, Neveu W, Moquin D, Noubade R, et al. The induction of antibody production by IL-6 is indirectly mediated by IL-21 produced by CD4+ T cells. J Exp Med (2009) 206:69–78. doi:10.1084/jem.20081571
60. Eto D, Lao C, Ditoro D, Barnett B, Escobar TC, Kageyama R, et al. IL-21 and IL-6 are critical for different aspects of B cell immunity and redundantly induce optimal follicular helper CD4 T cell (Tfh) differentiation. PLoS One (2011) 6:e17739. doi:10.1371/journal.pone.0017739
61. Haynes NM, Allen CD, Lesley R, Ansel KM, Killeen N, Cyster JG. Role of CXCR5 and CCR7 in follicular Th cell positioning and appearance of a programmed cell death gene-1high germinal center-associated subpopulation. J Immunol (2007) 179:5099–108.
62. Glatman Zaretsky A, Taylor JJ, King IL, Marshall FA, Mohrs M, Pearce EJ. T follicular helper cells differentiate from Th2 cells in response to helminth antigens. J Exp Med (2009) 206:991–9. doi:10.1084/jem.20090303
63. Deenick EK, Chan A, Ma CS, Gatto D, Schwartzberg PL, Brink R, et al. Follicular helper T cell differentiation requires continuous antigen presentation that is independent of unique B cell signaling. Immunity (2010) 33:241–53. doi:10.1016/j.immuni.2010.07.015
64. Baumjohann D, Okada T, Ansel KM. Cutting edge: distinct waves of BCL6 expression during T follicular helper cell development. J Immunol (2011) 187:2089–92. doi:10.4049/jimmunol.1101393
65. Baumjohann D, Preite S, Reboldi A, Ronchi F, Ansel KM, Lanzavecchia A, et al. Persistent antigen and germinal center B cells sustain T follicular helper cell responses and phenotype. Immunity (2013) 38:596–605. doi:10.1016/j.immuni.2012.11.020
66. Goenka R, Barnett LG, Silver JS, O’Neill PJ, Hunter CA, Cancro MP, et al. Cutting edge: dendritic cell-restricted antigen presentation initiates the follicular helper T cell program but cannot complete ultimate effector differentiation. J Immunol (2011) 187:1091–5. doi:10.4049/jimmunol.1100853
67. Paul WE, Ohara J. B-cell stimulatory factor-1/interleukin 4. Annu Rev Immunol (1987) 5:429–59. doi:10.1146/annurev.immunol.5.1.429
68. Casellas R, Jankovic M, Meyer G, Gazumyan A, Luo Y, Roeder R, et al. OcaB is required for normal transcription and V(D)J recombination of a subset of immunoglobulin kappa genes. Cell (2002) 110:575–85. doi:10.1016/S0092-8674(02)00911-X
69. Reimold AM, Iwakoshi NN, Manis J, Vallabhajosyula P, Szomolanyi-Tsuda E, Gravallese EM, et al. Plasma cell differentiation requires the transcription factor XBP-1. Nature (2001) 412:300–7. doi:10.1038/35085509
70. Iwakoshi NN, Lee AH, Vallabhajosyula P, Otipoby KL, Rajewsky K, Glimcher LH. Plasma cell differentiation and the unfolded protein response intersect at the transcription factor XBP-1. Nat Immunol (2003) 4:321–9. doi:10.1038/ni907
71. Shaffer AL, Shapiro-Shelef M, Iwakoshi NN, Lee AH, Qian SB, Zhao H, et al. XBP1, downstream of Blimp-1, expands the secretory apparatus and other organelles, and increases protein synthesis in plasma cell differentiation. Immunity (2004) 21:81–93. doi:10.1016/j.immuni.2004.06.010
72. Taubenheim N, Tarlinton DM, Crawford S, Corcoran LM, Hodgkin PD, Nutt SL. High rate of antibody secretion is not integral to plasma cell differentiation as revealed by XBP-1 deficiency. J Immunol (2012) 189:3328–38. doi:10.4049/jimmunol.1201042
73. Shen Y, Hendershot LM. Identification of ERdj3 and OBF-1/BOB-1/OCA-B as direct targets of XBP-1 during plasma cell differentiation. J Immunol (2007) 179:2969–78.
74. Nutt SL, Taubenheim N, Hasbold J, Corcoran LM, Hodgkin PD. The genetic network controlling plasma cell differentiation. Semin Immunol (2011) 23:341–9. doi:10.1016/j.smim.2011.08.010
75. Boyd AW, Goding JW, Schrader JW. The regulation of growth and differentiation of a murine B cell lymphoma. I. Lipopolysaccharide-induced differentiation. J Immunol (1981) 126:2461–5.
76. Krebs DL, Yang Y, Dang M, Haussmann J, Gold MR. Rapid and efficient retrovirus-mediated gene transfer into B cell lines. Methods Cell Sci (1999) 21:57–68. doi:10.1023/A:1009843325770
77. Hetz C, Lee AH, Gonzalez-Romero D, Thielen P, Castilla J, Soto C, et al. Unfolded protein response transcription factor XBP-1 does not influence prion replication or pathogenesis. Proc Natl Acad Sci U S A (2008) 105:757–62. doi:10.1073/pnas.0711094105
Keywords: Oct2, Obf1, Syk, Slamf1, B:T collaboration, cytokines
Citation: Corcoran L, Emslie D, Kratina T, Shi W, Hirsch S, Taubenheim N and Chevrier S (2014) Oct2 and Obf1 as facilitators of B:T cell collaboration during a humoral immune response. Front. Immunol. 5:108. doi: 10.3389/fimmu.2014.00108
Received: 24 December 2013; Accepted: 03 March 2014;
Published online: 20 March 2014.
Edited by:
Ananda L. Roy, Tufts University School of Medicine, USAReviewed by:
Mikael Sigvardsson, Linköping University, SwedenBruce David Mazer, Montreal Children’s Hospital, Canada
Copyright: © 2014 Corcoran, Emslie, Kratina, Shi, Hirsch, Taubenheim and Chevrier. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Lynn Corcoran, Walter and Eliza Hall Institute of Medical Research, 1G Royal Parade, Melbourne, VIC 3052, Australia e-mail:Y29yY29yYW5Ad2VoaS5lZHUuYXU=
†Present address: Stephane Chevrier, Institute of Molecular Life Sciences, University of Zurich, Zurich, Switzerland
 Lynn Corcoran
Lynn Corcoran Dianne Emslie1,2
Dianne Emslie1,2 Wei Shi
Wei Shi Susanne Hirsch
Susanne Hirsch Nadine Taubenheim
Nadine Taubenheim Stephane Chevrier
Stephane Chevrier