- 1Department of Microbiology, Kitasato University School of Medicine, Sagamihara, Kanagawa, Japan
- 2Department of Immunology, Kitasato University School of Medicine, Sagamihara, Kanagawa, Japan
Attempts to immortalize effector T cells by fusion of normal T cells with T cell lymphomas have been extensively made after the successful establishment of the B cell hybridoma that made it possible to infinitely produce monoclonal antibodies (1, 2). However, T cell hybridomas, which were mostly produced by using the T cell lymphoma line, BW5147, have contributed to T cell biology with limited success, because immortalizing effector T cells by cell fusion dose not necessarily result in immortalizing the bona fide effector functions of parental T cells. Meanwhile, T cells have been becoming recognized as consisting of more diverse populations than previously anticipated (3–6). Furthermore, a new class of lymphocytes termed innate lymphoid cells has been identified; they lack rearranged antigen receptors and are involved not only in immunity but also in inflammation and in tissue remodeling and repair (6–9). Thus, in order to gain a better understanding of the functions of various lymphocytes and their roles in tissue microenvironment, it is desirable to produce functional hybridomas that preserve the effector functions of the parental lymphocytes faithfully. We briefly review phenotypic changes in T cell hybrids from the point of view of somatic cell genetics and describe the concepts and principles behind devising an improved method for immortalizing various lymphocyte functions.
Genetics of Somatic Cell Fusion
Somatic cell hybridization has been used over several decades to study the genetic basis of cellular differentiation (10, 11). Those studies revealed that intertypic cell hybrids of equal ploidity fail to express the tissue-specific differentiated traits of either parent, a phenomenon termed extinction (12), that activation of previously silent genes occurs in hybrids with biased gene dosages, and that re-expression of previously extinguished phenotypes can occur in the hybrid segregants that have eliminated chromosomes of one parental cell. The differentiated state of somatic cells is therefore thought to be regulated via trans-acting regulatory factors and maintained by continuous regulatory circuits (13).
It has recently been shown that fusion of pluripotent embryonic stem cells (ESCs) with differentiated somatic cells yields pluripotent hybrid cells, indicating that ESCs have the ability to reset or reprogram the epigenomes of somatic cell nuclei toward pluripotency, albeit at low frequencies, but not to completely identical levels with original ESCs (14–18). Accordingly, extinction and/or activation of tissue-specific genes in somatic cell hybrids may be elucidated by partial reprograming of somatic cell epigenomes (19). In fact, in heterokaryons formed by fusion between somatic cells and non-dividing myotubes, trans-acting factors have been shown to partially reprogram one parental cell epigenome toward another parental cell epigenome in a bi-directional manner depending on biased gene dosages (20–22). However, because previously extinguished phenotypes in growing hybrids can be re-expressed in hybrid segregants (10, 23), the epigenetic state of each differentiated cell genome is considered to be stably maintained in proliferating hybrids. In continuously proliferating hybrids, therefore, reprograming of one parental epigenome toward another parental epigenome appears to be less likely to occur than in non-dividing heterokaryons. The mechanisms of epigenetic inheritance during the cell cycle (24) and the cell cycle stage of hybrid parental cells (25, 26) may inevitably affect reprograming of somatic nuclei.
In contrast to intertypic cell hybrids, hybrids between the same cell types (intratypic hybrids) continue to express the tissue-specific products of both parental cells (10, 27). When intratypic fusions are conducted between cells whose maturation stages are different, resultant hybrids usually express a variety of phenotypes (10). This phenotypic diversity, as suggested by the epigenetic study on CD4+ T cell differentiation (28), may reflect intricate epigenetic changes that occur during the maturation processes in the same cell lineage. The T cell hybridomas that were hitherto produced by fusion of immature lymphomas, like BW5147, with differentiated T cells are considered to be such intratypic hybrids.
However, T cell hybridomas were in most cases isolated by selecting proliferating cells based on a particular trait from effector T cell phenotypes, e.g., antigenic responsiveness, production of a lymphokine or cytolytic activity, without thorough cytogenetic analyses. Thus, the properties expressed in some of the hybridomas so far reported might represent the traits of hybrids with unequal ploidity or the traits of the segregants that have eliminated or duplicated chromosomes of one parental cell.
Properties of T Cell Hybridomas Produced by Cell Fusion between BW5147 Lymphoma and Normal T Lymphocytes
The BW5147/helper T cell hybridomas that expressed the T cell receptor (TCR) of both parental origins produced IL-2 upon engagement of the TCR with its cognate ligands (29, 30). On the other hand, BW5147 cells produced IL-2 by themselves when stimulated with calcium ionophore and phorbol ester mimicking the activation signals delivered when TCRs are engaged with cognate antigen (31). Furthermore, even cytolytic BW5147 hybridomas produced by fusion with cytotoxic T cells (CTLs) secreted IL-2 (32, 33) although CTLs usually had no ability to produce IL-2. Thus, the ability of hybridomas to produce IL-2 is most likely attributed to a property of the parental BW5147 lymphoma. Nevertheless, because of this ability, BW5147/T cell hybridomas – especially through the use of TCR-deficient BW5147 variants (34–36) – have made significant contributions to dissecting the structure and specificity of the TCR and also to analyzing antigen presentation by multiple types of antigen-presenting cells (37–41).
BW5147/T cell hybridomas have also contributed to the characterization of lymphokines, particularly the T cell replacing factor or eosinophil differentiation factor, now called IL-5 (42–45). However, because BW5147 lymphoma itself expresses a variety of lymphokines, including suppressor factors (31, 46, 47), it is necessary to be cautious when interpreting the results obtained with these lymphokine-producing hybridomas.
Fusion of BW5147 with CTLs usually yielded non-cytolytic hybridomas in which CD8α gene expression was suppressed by a trans-acting regulator derived from BW5147. Despite this, BW5147/T cell hybridomas with cytolytic activity were produced (32, 48), and their cytolytic activity was subsequently ascribed to the Fas/FasL cytolytic pathway (49). Because the Fas/FasL cytolytic activity may be weekly expressed by the BW5147 lymphoma itself (50), BW5147/T cell hybridomas might be permissive for the expression of the Fas/FasL cytolytic pathway, but not the perforin-mediated cytolytic pathway of parental CTLs (51).
Cytolytic BW5147/T cell hybrids, which grow in an IL-2-dependent manner were also produced (52), but when autonomously growing hybridomas were derived from them, they lost not only their dependence on IL-2, but also all other attributes of the CTL phenotypes (53) due to epigenetic mechanisms, not chromosomal segregation (54). In proliferating hybrids, all the gene expression programs of one parental cell genome that governs proliferation of the hybrid may conspire to determine the phenotype of the hybrid for unknown mechanisms. The above report is also reminiscent of the phenomenon termed “phenotypic exclusion” shown in melanoma/hepatoma hybrids (23).
In summary, BW5147/T cell hybridomas have considerably helped analyze certain aspects of T cell biology. However, since BW5147 genome simultaneously imposes, albeit not completely, its own programing on the hybridomas, BW5147 dose not faithfully immortalize effector functions of T cells.
Production of Growth-Arrested Hybrids through Fusion of Lymphomas and Terminally Differentiated T Cells
The quiescent state of non-dividing differentiated cells is actively controlled by regulatory mechanisms (55, 56). When terminally differentiated cells are fused with tumor cells in the same cell lineage, the regulatory mechanisms may operate dominantly over tumor transformation mechanisms (57). This notion has been substantiated by a study showing that fusion of lymphomas such as EL4, S1A, and YACUT, with antigen-responsive T cell line G4 yielded hybrid cells that ceased to proliferate (58–60). These growth-arrested hybrids proliferate by antigenic stimulation and thus have a phenotype similar to that of the G4 parental cells, which divide several times by antigenic stimulation and then enter into the quiescent phase of the cell cycle.
Upon antigenic stimulation, T lymphocytes proliferate, differentiate, and mature into effector cells. This is followed by either cessation of proliferation to become memory cells or initiation of apoptotic cell death to avoid over-immune-responses, which are harmful for the body. Thus, we can infer from the above experiments that when lymphomas are fused with normal activated lymphocytes, resultant hybrids will cease to proliferate or undergo cell death, though not necessarily always. This scenario explains why attempts to produce hybridomas between normal T cells and a variety of lymphomas were in many cases unsuccessful (47).
Unique Properties of the Hybrids between YACUT Lymphoma and Differentiated Lymphocytes
YACUT lymphoma derives from a Moloney leukemia virus (MoLV)-induced lymphoma originating from a T cell in an immature stage of development (61, 62). In growth-arrested YACUT/G4 hybrids, the G4 cell genome introduced into the YACUT lymphoma suppressed the immature phenotypes as well as the transformed phenotype of YACUT and imposed its own programing of terminally differentiated traits on the hybrids (62). Unlike growth-arrested EL4/G4 and S1A/G4 hybrids, prolonged growth of growth-arrested YACUT/G4 hybrids resulted in the appearance of continuously proliferating cells with the differentiated phenotype due to an increase in the number of the tumor-derived chromosome 15 carrying the MoLV-inserted pvt-1 gene (62). Furthermore, no segregants expressing immature phenotypes appeared after prolonged propagation of the re-transformed hybrids. From these observations, we can hypothesize that the epigenome of immature lymphoma nuclei differentiates into the mature epigenetic state of the G4 cell nuclei in the growth-arrested hybrids. This concept of association of cell growth-arrest with the induced differentiation of the immature lymphoma in lymphoma/lymphocyte hybrids (62) is parallel to the notion that tumor/normal cell hybrids in which tumor malignancy is suppressed are executing the differentiation program of a normal parental cell under the microenvironment in vivo (63, 64). This hypothesis predicts that fusion of the YACUT lymphoma with T lymphocytes followed by re-transformation will generate functional T cell hybridomas.
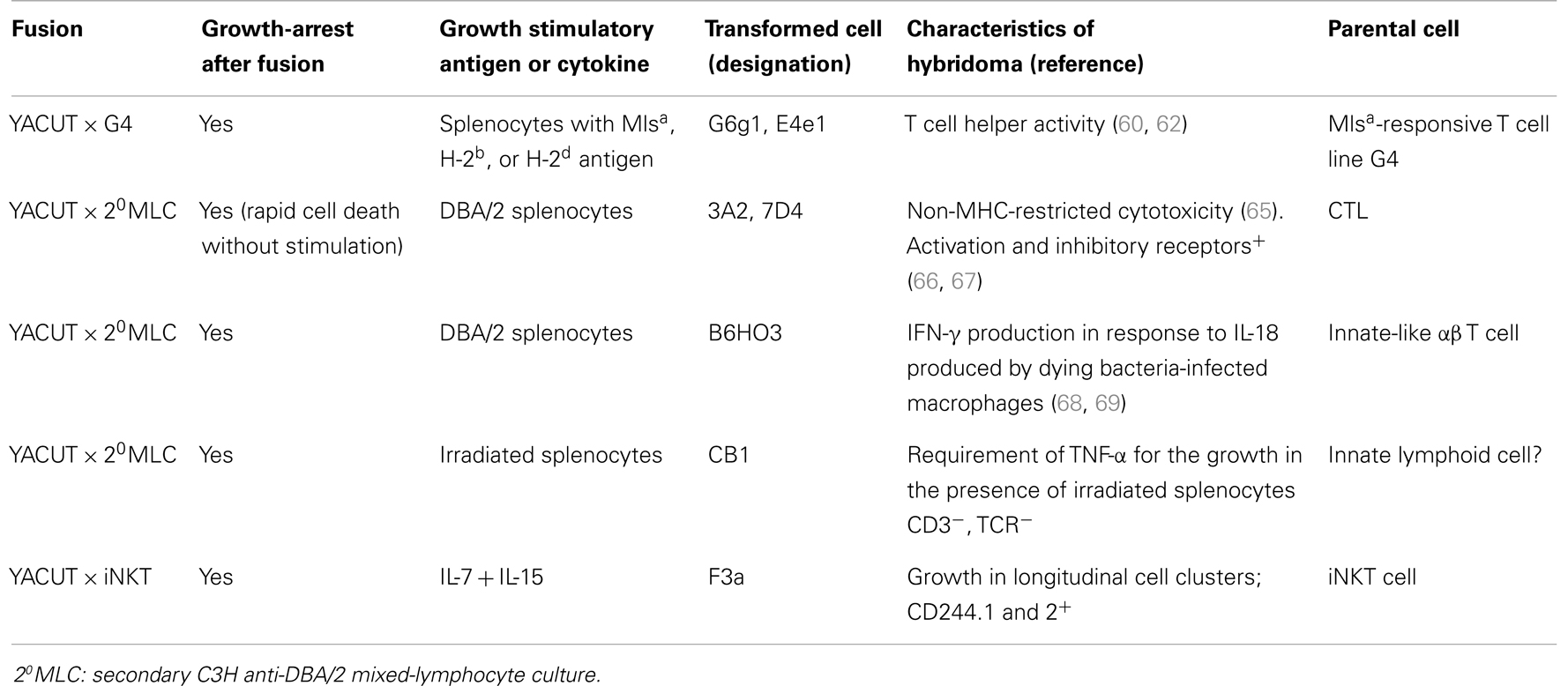
On the basis of this prediction, we have produced several functional hybridomas as summarized in Table 1. From rapidly dying growth-arrested hybrids produced by fusion of YACUT with a secondary mixed-lymphocyte culture cell, cytolytic hybridomas expressing the natural killer (NK) cell activation and inhibitory receptors were produced (65–67). From long-lived growth-arrested hybrids of the same fusion, we obtained another hybridoma termed B6HO3, which formed cell conjugates with bacteria-infected macrophages and produced IFN-γ in response to IL-18 secreted from dying macrophages caused by bacterial infection (68, 69). The parental αβ T cell of this hybridoma was a cell that belongs to minor innate-like αβ T cells. This B6HO3 hybridoma has helped reveal that subsets of innate lymphocytes respond to macrophage cell death caused by bacterial infection with the production of innate IFN-γ, which plays an important defensive role at an early stage of bacterial infections (70, 71).
Another hybridoma termed CB1 was also produced; its membrane phenotype was CD3−, TCR−, B220−, CD19−, CD244+, and FcRII/III+, suggesting that the parental cell was a NK-like cell or a cell belonging to innate lymphoid cells; CB1 was of interest, because the proliferation of the cells was suppressed in the presence of irradiated spleen cells and TNF-α was intimately involved in this growth. Further, by fusing YACUT with invariant natural killer T (iNKT) cells and culturing the resultant growth-arrested hybrids in the presence of IL-7 and IL-15, we obtained hybridoma F3a, which was unique in that it proliferated in longitudinal cell clusters. The functions of CB1 and F3a and their parental lymphocyte functions remain to be determined by further studies, but insofar as our prediction is valid, previously unknown functions of lymphocytes will be unveiled through these hybridomas.
Concluding Remarks
Fusion of lymphomas with normal lymphocytes yields mostly growth-arrested hybrids in which epigenomes of immature lymphomas are thought to be induced to differentiate toward the mature epigenetic state of normal lymphocyte nuclei. Thus, if we devise some methods of making the growth-arrested hybrids re-proliferate indefinitely, it will make it possible to immortalize effector functions of normal lymphocytes. In this respect, the growth-arrested YACUT/normal T cell hybrids are of importance. After prolonged growth by repeated stimulation, they resume indefinite proliferation presumably because of chromosomal instability of the YACUT genome, thereby becoming functional hybridomas that exhibit the phenotypes of their parental lymphocytes. This YACUT hybridization system and further research based on our hypothesis will shed light on hitherto unknown functions of lymphocytes and contribute to a better understanding of lymphocyte biology.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1. Köhler G, Milstein C. Continuous cultures of fused cells secreting antibody of predefined specificity. Nature (1975) 256:495–7. doi:10.1038/256495a0
3. Bendelac A, Bonneville M, Kearney JF. Autoreactivity by design: innate B and T lymphocytes. Nat Rev Immunol (2001) 1:177–86. doi:10.1038/35105052
4. Zhu J, Paul WE. Heterogeneity and plasticity of T helper cells. Cell Res (2010) 20:4–12. doi:10.1038/cr.2009.138
5. Ramiscal RR, Vinuesa CG. T-cell subsets in germinal center. Immunol Rev (2013) 252:146–55. doi:10.1111/imr.12031
6. Rankin L, Groom J, Mielke LA, Seillet C, Belz GT. Diversity, function, and transcriptional regulation of gut innate lymphocytes. Front Immunol (2013) 4:22. doi:10.3389/fimmu.2013.00022
7. Spits H, Cupedo T. Innate lymphoid cells: emerging insights in development, lineage relationships, and function. Annu Rev Immunol (2012) 30:647–75. doi:10.1146/annurev-immunol-020711-075053
8. Russell SE, Walsh PT. Sterile inflammation – do innate lymphoid cell subsets play a role? Front Immunol (2012) 3:246. doi:10.3389/fimmu.2012.00246
9. Spits H, Artis D, Colonna M, Diefenbach A, Di Santo JP, Eberi G, et al. Innate lymphoid cells – a proposal for uniform nomenclature. Nat Rev Immunol (2013) 13:145–9. doi:10.1038/nri3365
10. Gourdeau H, Fournier RE. Genetic analysis of mammalian cell differentiation. Annu Rev Cell Biol (1990) 6:69–94. doi:10.1146/annurev.cellbio.6.1.69
12. Boshart M, Nitsch D, Schütz G. Extinction of gene expression in somatic cell hybrids – a reflection of important regulatory mechanisms? Trends Genet (1993) 9:240–5. doi:10.1016/0168-9525(93)90088-Y
13. Blau HM. Differentiation requires continuous active control. Annu Rev Biochem (1992) 61:1213–30. doi:10.1146/annurev.bi.61.070192.010025
14. Ambrosi DJ, Tanasijevic B, Kaur A, Obergfell C, O’Neill RJ, Krueger W, et al. Genome-wide reprogramming in hybrids of somatic cells and embryonic stem cells. Stem Cells (2007) 25:1104–13. doi:10.1634/stemcells.2006-0532
15. Yamanaka S, Blau HM. Nuclear reprogramming to a pluripotent state by three approaches. Nature (2010) 465:704–12. doi:10.1038/nature09229
16. Serov OL, Matveeva NM, Khabarova AA. Reprogramming mediated by cell fusion technology. Int Rev Cell Mol Biol (2011) 291:155–90. doi:10.1016/B978-0-12-386035-4.00005-7
17. Soza-Ried J, Fisher AG. Reprogramming somatic cells towards pluripotency by cellular fusion. Curr Opin Genet Dev (2012) 22:459–65. doi:10.1016/j.gde.2012.07.005
18. Foshay KM, Looney TJ, Chari S, Mao FF, Lee JH, Zhang L, et al. Embryonic stem cells induce pluripotency in somatic cell fusion through biphasic reprogramming. Mol Cell (2012) 46:159–70. doi:10.1016/j.molcel.2012.02.013
19. Massa S, Junker S, Matthias P. Molecular mechanisms of extinction: old findings and new ideas. Int J Biochem Cell Biol (2000) 32:23–40. doi:10.1016/S1357-2725(99)00102-8
20. Terranova R, Pererira CF, Du Roure C, Merkenschlager M, Fisher AG. Acquisition and extinction of gene expression programs are separable events in heterokaryon reprogramming. J Cell Sci (2006) 119:2065–72. doi:10.1242/jcs.02945
21. Zhang F, Pomerantz JH, Sen G, Palermo AT, Blau HM. Active tissue-specific DNA demethylation conferred by somatic cell nuclei in stable heterokaryons. Proc Natl Acad Sc U S A (2007) 104:4395–400. doi:10.1073/pnas.0700181104
22. Palermo A, Doyonnas R, Bhutani N, Pomerantz J, Alkan O, Blau HM. Nuclear reprogramming in heterokaryons is rapid, extensive, and bidirectional. FASEB J (2009) 23:1431–40. doi:10.1096/fj.08-122903
23. Fougère C, Weiss MC. Phenotypic exclusion in mouse melanoma-rat hepatoma hybrid cells: pigment and albumin production are not reexpressed simultaneously. Cell (1978) 15:843–54. doi:10.1016/0092-8674(78)90269-6
24. Probst AV, Dunleavy E, Almouzni G. Epigenetic inheritance during the cell cycle. Nat Rev Mol Cell Biol (2009) 10:192–206. doi:10.1038/nrm2640
25. Wilmut I, Schnieke AE, McWhir J, Kind AJ, Campbell KHS. Viable offspring derived from fetal and adult mammalian cells. Nature (1997) 385:810–3. doi:10.1038/385810a0
26. Sullivan S, Pells S, Hooper M, Gallagher E, McWhir J. Nuclear reprogramming of somatic cells by embryonic stem cells is affected by cell cycle stage. Cloning Stem Cells (2006) 8:174–88. doi:10.1089/clo.2006.8.174
27. Havran WL, Fitch FW. Characterization of murine cytolytic-helper hybrid T cell clones. Nature (1987) 325:65–7. doi:10.1038/325065a0
28. Vahedi G, Poholek AC, Hand TW, Laurence A, Kanno Y, O’Shea JJ, et al. Helper T-cell identity and evolution of differential transcriptomes and epigenomes. Immunol Rev (2013) 252:24–40. doi:10.1111/imr.12037
29. Goldsby RA, Osborne BA, Simpson E, Herzenberg LA. Hybrid cell lines with T-cell characteristics. Nature (1977) 267:707–8. doi:10.1038/267707a0
30. Kappler JW, Skidmore B, White J, Marrack P. Antigen-inducible, H-2-restricted, interleukin-2-producing T cell hybridomas. Lack of independent antigen and H-2 recognition. J Exp Med (1981) 153:1198–214. doi:10.1084/jem.153.5.1198
31. Hagiwara H, Yokota T, Luh J, Lee F, Arai K, Arai N, et al. The AKR thymoma BW5147 is able to produce lymphokines when stimulated with calcium ionophore and phorbol ester. J Immunol (1988) 140:1561–5.
32. Kaufmann Y, Berke G. Monoclonal cytotoxic T lymphocyte hybridomas capable of specific killing activity, antigenic responsiveness, and inducible interleukin secretion. J Immunol (1983) 131:50–6.
33. Gu JJ, Gottlieb PD. Inducible functions in hybrids of a Lyt-2+ BW5147 transfectant and the 2C CTL line. Immunogenetics (1992) 36:283–93. doi:10.1007/BF00215656
34. White J, Blackman M, Bill J, Kappler J, Marrack P, Gold DP, et al. Two better cell lines for making hybridomas expressing specific T cell receptors. J Immunol (1989) 142:1822–5.
35. Burgert H-G, White J, Weltzien H-U, Marrack P, Kappler JW. Reactivity of Vβ17a+ CD8+ T cell hybrids. Analysis using a new CD8+ T cell fusion partner. J Exp Med (1989) 170:1887–904. doi:10.1084/jem.170.6.1887
36. Sanderson S, Shastri N. LacZ inducible, antigen/MHC-specific T cell hybrids. Int Immunol (1994) 6:369–76. doi:10.1093/intimm/6.3.369
37. Hedrick SM, Matis LA, Hecht TT, Samelson LE, Longo DL, Heber-Katz E, et al. The fine specificity of antigen and Ia determinant recognition by T cell hybridoma clones specific for pigeon cytochrome c. Cell (1982) 30:141–52. doi:10.1016/0092-8674(82)90020-4
38. Kappler J, Kubo R, Haskins K, White J, Marrack P. The mouse T cell receptor: comparison of MHC-restricted receptors on two cell hybridomas. Cell (1983) 34:727–37. doi:10.1016/0092-8674(83)90529-9
39. Bartley MB, Canaday DH. T cell hybridomas to study MHC-II restricted B-cell receptor-mediated antigen presentation by human B cells. J Immunol Methods (2011) 370:35–42. doi:10.1016/j.jim.2011.05.005
40. Gumperz JE, Roy C, Makowska A, Lum D, Sugita M, Podrebarac T, et al. Murine CD1d-restricted T cell recognition of cellular lipids. Immunity (2000) 12:211–21. doi:10.1016/S1074-7613(00)80174-0
41. Singh N, Pacholczyk R, Iwashima M, Ignatowicz L. Generation of T cell hybridomas from naturally occurring FoxP3+ regulatory T cells. Methods Mol Biol (2011) 707:39–44. doi:10.1007/978-1-61737-979-6_3
42. Takatsu K, Tanaka K, Tominaga A, Kumahara Y, Hamaoka T. Antigen-induced T cell-replacing factor (TRF). III. Establishment of T cell hybrid clone continuously producing TRF and functional analysis of released TRF. J Immunol (1980) 125:2646–53.
43. Puré E, Isakson PC, Takatsu K, Hamaoka T, Swain SL, Dutton RW, et al. Induction of B cell differentiation by T cell factors. I. Stimulation of IgM secretion by products of a T cell hybridoma and a T cell line. J Immunol (1981) 127:1953–8.
44. Warren DJ, Sanderson CJ. Production of a T-cell hybrid producing a lymphokine stimulating eosinophil differentiation. Immunology (1985) 54:615–23.
45. Kinashi T, Harada N, Severinson E, Tanabe T, Sideras P, Konishi M, et al. Cloning of complementary DNA encoding T-cell replacing factor and identity with B-cell growth factor II. Nature (1986) 324:70–3. doi:10.1038/324070a0
47. Beezley BB, Ruddle NH. A critical analysis of the T cell hybrid technique. J Immunol Methods (1982) 52:269–81. doi:10.1016/0022-1759(82)90001-1
48. Kaufmann Y, Berke G, Eshhar Z. Cytotoxic T lymphocyte hybridomas that mediate specific tumor-cell lysis in vitro. Proc Natl Acad Sci U S A (1981) 78:2502–6. doi:10.1073/pnas.78.4.2502
49. Berke G. The Fas-based mechanism of lymphocytotoxicity. Hum Immunol (1997) 54:1–7. doi:10.1016/S0198-8859(97)00009-8
50. Kaneda R, Iwabuchi K, Onoé K. Dissociation of Fas-mediated cytotoxicity and FasL expression in a cytotoxic CD4+ T-cell clone: comparative analysis of Fas-mediated cytotoxicity between a T-hybridoma and a T-cell clone. Immunol Lett (1997) 55:53–60. doi:10.1016/S0165-2478(96)02683-1
51. Lowin B, Hahne M, Mattmann C, Tschopp J. Cytolytic T-cell cytotoxicity is mediated through perforin and Fas lytic pathways. Nature (1994) 370:650–2. doi:10.1038/370650a0
52. Nabholz M, Cianfriglia M, Acuto O, Conzelmann A, Haas W, von Boehmer H, et al. Cytolytically active murine T-cell hybrids. Nature (1980) 287:437–40. doi:10.1038/287437a0
53. Nguyen TT, Nabholz M. Glucocorticoid resistance is a dominant trait in hybrids between cytolytic T-lymphocyte lines and AKR thymomas. Somat Cell Mol Genet (1985) 11:593–604. doi:10.1007/BF01534724
54. Somoza C, Fernandez-Ruiz E, Jotterand-Bellomo M, Sanz E, Nabholz M, Silva A. Loss of interleukin 2 dependence in cloned interleukin 2-dependent rat T lymphocyte x BW5147 hybridomas is not associated with segregation of a specific pair of rat chromosomes. Eur J Immunol (1989) 19:1177–81. doi:10.1002/eji.1830190704
55. Blomen VA, Boonstra J. Cell fate determination during G1 phase progression. Cell Mol Life Sci (2007) 64:3084–104. doi:10.1007/s00018-007-7271-z
56. Yusuf I, Fruman DA. Regulation of quiescence in lymphocytes. Trends Immunol (2003) 24:380–6. doi:10.1016/S1471-4906(03)00141-8
57. Szpirer J, Szpirer C, Wanson J-C. Control of serum protein production in hepatocyte hybridomas: immortalization and expression of normal hepatocyte genes. Proc Natl Acad Sci U S A (1980) 77:6616–20. doi:10.1073/pnas.77.11.6616
58. Kubota K, Katoh H. Cessation of autonomous proliferation of mouse lymphoma EL4 by fusion with a T cell line. Int J Cancer (1990) 45:540–5. doi:10.1002/ijc.2910450328
59. Kubota K, Katoh H. Cessation of autonomous proliferation of mouse S1A by fusion with Mls-antigen-dependent T cell line. Proc Jap Cancer Ass (1990) 9:163.
60. Kubota K, Imreh S, Katoh H, Babonits M, Wiener F. Correlation of myc expression with the growth-arrested and transformed phenotypes in hybrids between a T lymphoma and an antigen-responsive T-cell line. Int J Cancer (1992) 51:927–34. doi:10.1002/ijc.2910510616
61. Klein G, Klein E. Antigenic properties of lymphoma induced by the Moloney agent. J Natl Cancer Inst (1964) 32:547–68.
62. Kubota K, Nakazato K. The role of duplication of tumor-derived chromosome 15 carrying the rearranged pvt-1 gene in the transformed phenotype of YACUT T-cell lymphoma x G4 T-cell line somatic cell hybrids in dictating the terminal differentiation program of the parental G4 cell. J Cell Sci (1994) 107:2761–8.
63. Peehl DM, Stanbridge EJ. The role of differentiation in the suppression of tumorigenicity in human cell hybrids. Int J Cancer (1982) 30:113–20. doi:10.1002/ijc.2910300119
64. Harris H. The role of differentiation in the suppression of malignancy. J Cell Sci (1990) 97:5–10.
65. Kubota K, Nakazato K, Tamauchi H, Sasahara T, Katoh H. Generation of novel killer hybridomas derived from proliferation-suppressed somatic cell hybrids between YACUT T cell lymphoma and normal lymphocytes activated in secondary mixed lymphocyte cultures. J Immunol Methods (1996) 192:137–47. doi:10.1016/0022-1759(96)00030-0
66. Kubota K. A killer cell protective antigen expressed by MHC-unrestricted killer hybridomas. Cell Immunol (1997) 181:50–8. doi:10.1006/cimm.1997.1195
67. Kubota K, Katoh H, Muguruma K, Koyama K. Characterization of a surface membrane molecule expressed by natural killer cells in most inbred mouse strains: monoclonal antibody C9.1 identifies an allelic form of the 2B4 antigen. Immunology (1999) 96:491–7. doi:10.1046/j.1365-2567.1999.00709.x
68. Kubota K. A novel functional T cell hybridoma recognizes macrophage cell death induced by bacteria: a possible role for innate lymphocytes in bacterial infection. J Immunol (2006) 176:7576–88. doi:10.4049/jimmunol.176.12.7576
69. Kubota K, Kadoya Y. IL-18 provided in dying bacterial-infected macrophages induces IFN-γ production in functional T-cell hybridoma B6HO3 through cell conjugates. Innate Immun (2014) 20:133–44. doi:10.1177/1753425913485474
70. Kubota K. Innate IFN-γ production by subsets of natural killer cells, natural killer T cells and γδ T cells in response to dying bacterial-infected macrophages. Scand J Immunol (2010) 71:199–209. doi:10.1111/j.1365-3083.2009.02366.x
Keywords: BW5147, cell growth-arrest, induction of differentiation, lymphocyte differentiation, reprograming, YACUT
Citation: Kubota K and Iwabuchi K (2014) Phenotypic changes in growth-arrested T cell hybrids: a possible avenue to produce functional T cell hybridoma. Front. Immunol. 5:229. doi: 10.3389/fimmu.2014.00229
Received: 24 March 2014; Accepted: 05 May 2014;
Published online: 19 May 2014.
Edited by:
Nick Gascoigne, National University of Singapore, SingaporeReviewed by:
Willi Born, National Jewish Health, USAWendy L. Havran, The Scripps Research Institute, USA
Copyright: © 2014 Kubota and Iwabuchi. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence:a29pY2hpLXRuQGpjb20uaG9tZS5uZS5qcA==
 Koichi Kubota
Koichi Kubota Kazuya Iwabuchi
Kazuya Iwabuchi