- Department of Infectious Diseases, Leiden University Medical Center, Leiden, Netherlands
In view of the fact that only a small part of the Mtb expressome has been explored for identification of antigens capable of activating human T-cell responses, which is critically required for the design of better TB vaccination strategies, more emphasis should be placed on innovative ways to discover new Mtb antigens and explore their function at the several stages of infection. Better protective antigens for TB-vaccines are urgently needed, also in view of the disappointing results of the MVA85 vaccine, which failed to induce additional protection in BCG-vaccinated infants (1). Moreover, immune responses to relevant antigens may be useful to identify TB-specific biomarker signatures. Here, we describe the potency of novel tools and strategies to reveal such Mtb antigens. Using proteins specific for different Mtb infection phases, many new antigens of the latency-associated Mtb DosR-regulon as well as resuscitation promoting factor proteins, associated with resuscitating TB, were discovered that were recognized by CD4+ and CD8+ T-cells. Furthermore, by employing MHC binding algorithms and bioinformatics combined with high-throughput human T-cell screens and tetramers, HLA-class Ia restricted polyfunctional CD8+ T-cells were identified in TB patients. Comparable methods, led to the identification of HLA-E-restricted Mtb epitopes recognized by CD8+ T-cells. A genome-wide unbiased antigen discovery approach was applied to analyze the in vivo Mtb gene expression profiles in the lungs of mice, resulting in the identification of IVE-TB antigens, which are expressed during infection in the lung, the main target organ of Mtb. IVE-TB antigens induce strong T-cell responses in long-term latently Mtb infected individuals, and represent an interesting new group of TB antigens for vaccination. In summary, new tools have helped expand our view on the Mtb antigenome involved in human cellular immunity and provided new candidates for TB vaccination.
Introduction
M. tuberculosis Antigen Discovery: Traditional Approaches
Antigen discovery efforts have been a core component of mycobacterial research for over several decades, and have been markedly facilitated since the availability of the Mycobacterium tuberculosis (Mtb) genome sequence (2). A lot of different antigen discovery approaches have been described, among which are biochemical-, genetic-, expression library-, and peptide T-cell epitope-based approaches. Using animal models, antigens with protective potential against Mtb infection have been discovered, some of which have moved into human clinical trials (phase 1/2a) (3). Nevertheless, T-cell epitopes have been identified in only 7% of all predicted 4000 open reading frames (ORFs) of Mtb, and the top 30 most frequently studied protein antigens contain 65% of the known epitopes (4). This leaves the Mtb “antigenome” incompletely identified, which may be especially relevant in relation to Mtb’s phase-dependent variation in gene expression (see below) in response to varying environmental factors (5, 6).
Selection of vaccine candidates has been based largely on empirical observations in the mouse, guinea pig, and non-human primate Mtb infection models and only a few antigens (mostly those secreted during active replication of Mtb) have been exploited as human vaccine candidates (7). Since Mtb alters its gene expression profile significantly during intracellular stress inside host macrophages, its antigen repertoire which is expressed and exposed to the immune system varies considerably during different stages of infection under the pressure of various human host defense mechanisms. A more profound understanding of the actual Mtb antigenome expressed during different phases of Mtb infection, particularly in the lung, the main target organ of Mtb and the identification of the major T-cell epitopes involved, is key to the design of better TB-vaccines and TB correlates of immunity.
Almost all TB-vaccine antigen discovery approaches have implicitly relied on the assumption that the Mtb antigens studied are expressed and presented by infected cells, where they are supposedly recognized by T-cells that execute an appropriate effector response. The latter either assist phagocytes in controlling or eliminating live bacteria through various intracellular pathways (phagosomal maturation and phagolysosomal fusion; oxidative/nitrate intermediates; oxygen/nutrient deprivation; the activity of defensins and other anti-microbial peptides and enzymes; autophagy; apoptosis) (8), or – alternatively direct killing of infected cells. Despite the significant advances made recently, relatively little is known about the Mtb antigen repertoire, which is truly expressed by the tubercle bacillus during its infection cycle in human cells. Although abundantly expressed proteins of Mtb are often the primary targets of research, less prominently expressed antigens may have equally good or even superior vaccine potential. Better insight into the antigen repertoire available for immune recognition on infected cells, its dynamic changes as well as the quantitative relationship between the various antigens expressed, should provide new directions for antigen discovery and vaccine testing, with the potential to complement or change current strategies used in TB-vaccine-design.
Classical Mtb Antigen Discovery Methods and the Risk of Tunnel Vision
Indirect discovery approaches have mostly been the basis for currently available evidence supporting the recognition of Mtb antigens, including those on infected cells. Whereas the protective potential of Mtb antigens is typically demonstrated using effective vaccine platforms in animal models, the selection of the antigens to be tested in such platforms is often biased and limited by the antigen discovery procedures used. For example, as outlined above, many studies have concentrated on antigens that are highly expressed by bacteria under in vitro culture conditions in liquid growth media. However, it is improbable that the same bacterial transcriptomic or proteomic profiles expressed under optimal laboratory growth conditions are the same as those expressed during in vivo host infection.
A further important bias is that most Mtb antigens recognized by human cells have been identified using IFN-γ assays as read outs. As mentioned above, a considerable number of antigens eliciting CD4+ IFN-γ Th1-cell responses has been identified (9), but this represents almost certainly only a fraction of the potential Mtb “antigenome.” The number of IFN-γ-releasing antigen-specific T-cells and the amount of total IFN-γ released have remained widely used surrogate markers for the pro-inflammatory immune response against Mtb. Thus, the antigens activating other immune cells, including CD4+ T-cells that produce other cytokines than IFN-γ such as Th2 cells, Tregs, Th17/22, cells and cytolytic cells, as well as non-classically restricted (MHC-Ib, see below) human T-cell subsets, remain largely incomplete or even unknown. However, the lack of (application of) well-developed methods to identify Mtb-responsive T-cells other than classical Th1 or proliferative responses have contributed to this bias in Mtb antigens identified. Thus, it is vital to develop better and more diverse assays that can be applied to detect Mtb-induced responses across all relevant human T-cell compartments.
Scope of This Review
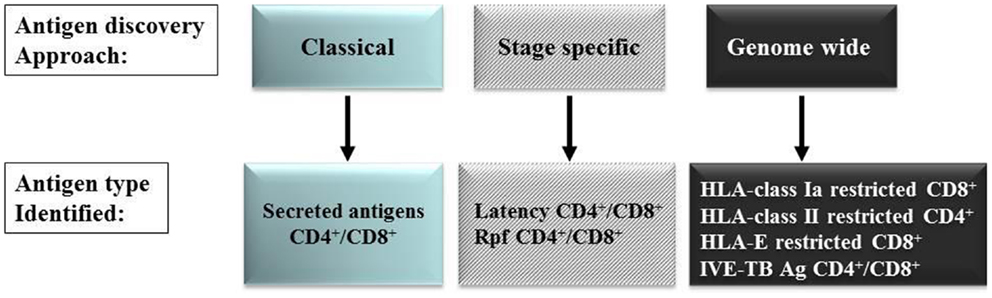
Since our knowledge of the human Mtb antigenome is far from complete as most currently known Mtb antigens were identified using IFN-γ (Th1) production as read-out, and because there is limited knowledge about the vaccine potential (protective efficacy) of most antigens, we have followed several alternative strategies to discover new Mtb antigens. Below, we will review the five approaches we have pursued in recent years (Figure 1). It is likely that many unexplored antigens for classical or non-classical T-cells exist in the Mtb antigenome that may possess vaccine potential, but appropriate tools and technologies are required to reveal these. Improved and rational selection is needed to identify candidate antigens with vaccine potential, based on comprehensive knowledge of their patterns of expression, broad immunogenicity, suitability for processing, HLA-binding and induction of protective immunity in relevant model systems.
New Mtb Antigen Discovery Approaches
Infection Stage-Specific Mtb Gene Expression Analyses
Mtb DosR-regulon encoded proteins as latency antigens
As mentioned above, we have hypothesized that the design of improved vaccination strategies requires a better understanding of the Mtb proteins that are expressed during the different phases of the Mtb intracellular life cycle, and their recognition by human T-cell subsets.
During in vivo Mtb infection, environmental and host immune factors induce bacterial dormancy, in the course of which Mtb enters a state of non-or slowly replicating persistence, decreases its metabolic activity and alters its gene expression pattern (5, 10). This adjustment by the bacterium is thought to enhance its resistance to environmental and host immune stress. Under in vitro conditions that are considered to imitate part of the environment encountered by tubercle bacilli in vivo in the immunocompetent host (11) such as hypoxia, low dose nitric oxide, CO exposure or in IFN-γ-activated macrophages, Mtb induces the expression of the 48 gene encoding DosR (Rv3133c) regulon (12). In view of this infection phase-specific upregulation, we first decided to study T-cell responses to these 48 DosR-regulon encoded antigens (so-called TB latency antigens) in various ethnically and geographically distinct cohorts in Europe (Italy, Germany, The Netherlands) and Africa (The Gambia, Ethiopia, Uganda, and South Africa) (13–19). In all cohorts we observed T-cell recognition of Mtb DosR-regulon encoded latency antigens, particularly in TST+ individuals [designated latent TB (LTBI)] compared to (ex-) TB patients. Thus, T-cell proliferation and IFN-γ production to Mtb DosR-regulon encoded latency antigens are characteristically seen in LTBI in diverse genetic and geographic populations.
An unexpected but important finding was that Mtb DosR-regulon encoded antigens were very poorly if at all recognized in previously BCG-vaccinated individuals (20). Additionally, BCG-vaccinated HLA-DR3 and HLA-A2 transgenic mice (21) failed to respond to these antigens. The explanation for this lack of induction of immune responses was not due to BCG’s ability to express the DosR-regulon because transcriptional profiling of 14 different BCG strains, cultured under hypoxia, and nitric oxide exposure in vitro, showed that genes of the DosR-regulon (except Rv3133) were expressed to similar extents as observed in Mtb. Moreover, comparison of the amino acid sequences of M. bovis BCG and Mtb showed at least 97% homology with 85% of the genes being completely identical (20). The explanation for this phenomenon thus remains unknown, although it may relate to an inability of BCG to enter a state of latency following intradermal immunization. In agreement with the findings in human LTBI, the Mtb DosR-regulon encoded latency antigens Rv1733c, Rv2031c, and Rv2626c, were more prominently recognized in chronic infected mice than in acute phase infection, whereas the secreted protein Ag85B (Rv3804c) showed the opposite phenotype (18), again highlighting the preferential recognition of these antigens in latent or long term as opposed to acute, early phase infection.
An important group of individuals, the study of which shed more light into this matter was composed of individuals who had been Mtb infected for over 30 years ago, as witnessed by their positive Mantoux skin tests, yet never developed disease despite not having received any therapeutic treatment. Detailed immune profiling experiments with cells from these long-term infected individuals demonstrated strong Mtb DosR-regulon encoded antigen-specific T-cell responses. Various epitopes were identified that induced mono- and polyfunctional CD4+ and CD8+ T-cell responses, in particular IFN-γ+TNF-α+ CD8+ effector memory- or effector T-cells (22).
Considering the inability of BCG to induce immune responses to latency antigens, TB vaccination strategies started to consider incorporating DosR-regulon encoded antigens as a strategy to complement and improve the current BCG vaccine. This was demonstrated in TB mouse models using rBCG_ureC:hly expressing defined latency-associated antigens and test this construct for long-term protection against an isolate of the Mtb Beijing/W lineage (23). Expression of Rv2659c, Rv3407, and Rv1733c by rBCG_ureC:hly showed long-term protection superior to BCG in both lung and spleen compared to rBCG not expressing latency antigens at day 200 post-infection after intradermal vaccination of mice. Even a latency antigen hsp16-derived peptide linked to PAM2Cys adjuvant induced more efficient protection than BCG (24).
Another type of latency antigens is represented by the so-called “starvation” antigens, which are expressed under conditions of nutrient deprivation. A hybrid protein consisting of two secreted, acute phase-specific antigens, Ag85B and ESAT-6, fused to the nutrient stress-induced antigen Rv2660c, was constructed. In pre-exposure mouse models this hybrid protein formulated in CAF01, promoted hybrid-specific T-cells, in particular polyfunctional CD4 + T-cells and more efficient containment of late-stage infection than the Ag85B-ESAT-6 vaccine and BCG (25). Moreover, in latent TB mouse models, post-exposure immunization with this hybrid controlled TB reactivation and significantly lowered the number of bacteria in the lung compared to adjuvant control mice.
Immunization of cynomolgus macaques with the multistage Ag85B-ESAT-6-Rv2660c protein in IC31 adjuvant as a boost to vaccination with BCG delayed and reduced clinical disease after challenge with M. tuberculosis and also prevented reactivation of latent infection (26). This boosting regimen resulted in efficient control of M. tuberculosis infection and reduced rates of clinical disease. Importantly, it improved survival of the NHP compared to BCG alone and the vaccinated monkeys did not reactivate latent infection after treatment with anti-TNF antibody.
Considering routine BCG vaccination practice in most TB endemic countries, improved, rationally designed Mtb subunit vaccines could be employed by simultaneous vaccination with BCG, or booster vaccination on top of BCG, or by constructing improved recombinant BCG strains expressing such antigens.
Resuscitation promoting factors as antigens
Although the factors that trigger bacterial reactivation and resumption of intracellular growth remain largely known, Mtb resuscitation promoting factors (Rpfs), which are secreted proteins with high homology to the hormone-like protein secreted by Micrococcus luteus (M. luteus) are believed to play an important role in this respect (27, 28). The Mtb genome encodes five such rpf genes [Rv0867c (rpfA), Rv1009 (rpfB), Rv1884c (rpfC), Rv2389c (rpfD), and Rv2450c (rpfE)] that are able to stimulate the growth of dormant mycobacteria and expression of the five Rpf proteins is observed in vitro in actively replicating Mtb, in BCG as well as in Mtb-infected human tissue (29–31). Addition of Rpf proteins to sputa from TB patients improves the sensitivity of culture based detection of live Mtb in certain conditions (32, 33). There is differential rpf expression in cultures grown under hypoxia, nutrient starvation, acidic conditions, stationary, non-cultivable, and resuscitation phase-like conditions, suggesting that the role of Mtb Rpfs may be infection stage dependent but likely is not identical for all Rpfs (34).
In view of the role of Mtb Rpf proteins in the resuscitation of mycobacteria, immunity against these proteins may reflect the ability to detect the presence of actively replicating Mtb organisms at an early stage. Thus, immunity against Rpf proteins may play a role in host control bacterial reactivation. In line with the immunogenicity found in mice for Rv0867c, Rv1009, Rv2389c, and Rv2450c (35), we identified the first human Mtb Rpf-specific T-cell responses against Mtb Rpfs (19), showing IFN-γ production in TST+ individuals in response to Rv1009, Rv1884c, and Rv2450c and to a lesser extent Rv0867c, whereas hardly any IFN-γ was detected in individuals without a positive Mantoux. More detailed analyses of the immune responses to Mtb Rpf proteins in Mtb-exposed individuals, including long-term LTBI non-progressors showed frequent and significant T-cell responses against both Rv0867c and Rv2389c and identified novel Mtb Rpf epitopes, including a single highly dominant peptide epitope in Rv2389c. Of note, Mtb Rpf-specific polyfunctional memory CD4+ and particularly CD8+ T-cell memory responses were observed in response to Rv0867c and Rv2389c Rpf proteins. The polyfunctional phenotype of these cells was both single and double cytokine producing CD4+ and CD8+ T-cells, supporting the concept that CD8+ T-cells may be important in long-term control of Mtb infection. Based on these collective studies, we envisage that a combination of multiple phase-specific antigens may significantly improve the protective potential of new TB-vaccines (25, 36). Thus, subunit vaccines based on a combination of latency and Rpf antigens could be designed that induce responses able to eliminate Mtb bacilli in their dormant state, as well as inhibit Mtb bacilli that are trying to recommence active replication.
Unbiased Mtb “Genome Wide” Antigen Discovery Approaches
HLA-class Ia presented human CD8+ T-cell epitopes
Despite the fact that CD4+ T-cells play a vital role in immunity against Mtb, it is becoming evident that also CD8+ T-cells contribute to host defense against Mtb by virtue of their ability to produce pro-inflammatory cytokines (9, 37, 38), lyse infected host cells (39), and kill mycobacteria (40). Despite the fact that the antigens and epitopes activating human Mtb-specific CD8+ T-cell responses have been less well defined than those for CD4+ T-cells, some groups have been able to isolate such CD8+ T-cells from humans (41–44). Unexpectedly, it was reported that ex vivo frequencies of CD8+ T-cells recognizing epitopes from six different Mtb proteins in patients with active TB were lower as evaluated by specific tetramers, but normalized following therapy to frequencies comparable to subjects with LTBI. Additionally, CD8+ T-cells with an IL-2+/IFN-γ+ phenotype were particularly reduced or found absent in active TB patients (45). Nevertheless, it has remained challenging to identify the role and function of CD8+ T-cells in TB, urging for new strategies and appropriate tools to decipher their specificities. Using classic approaches it would not be feasible to screen the approximately 1 million possible 9-mers in the Mtb proteome for CD8+ T-cell responses.
A recently described approach addressed this issue exploiting an integrated computational and proteomic approach to screen 10% of the Mtb proteome for antigens that are recognized by CD8+ T-cells: using a synthetic Mtb peptide library consisting of 15-mers (11 aa overlap) with high probability of containing CD8+ T-cell epitopes, IFN-γ release by Mtb-specific, HLA-class I-restricted CD8+ T-cell clones was measured by ELISPOT assay (46). This study identified the EsxJ family, PE9, and PE_PGRS42 as three novel CD8 antigens and validated the use of peptide library-based approaches a new tool for identification of Mtb epitopes recognized by CD8+ T-cells.
An alternative method of combined bioinformatics- and functional immunological screening strategies, called “reverse antigen discovery,” was applied by our group to identify HLA-class Ia restricted, CD8+ T-cell antigens. Through peptide-binding prediction algorithms, potential peptide epitopes of Mtb antigens were identified restricted by three major HLA-class Ia supertypes (HLA-A2, -A3, and -B7) which together cover more than 80% of the population from different ethnic groups (47). Over 400 synthetic Mtb peptides with predicted binding affinities for HLA-A*0201, HLA-A*0301, and HLA-B*0702 (representing the above supertypes) were tested for HLA-binding and induction of proliferation of CD8+ T-cells. This study led to the identification of >60 new Mtb epitopes. Further validation of the most interesting epitopes was executed using HLA-class I-tetramers and assessment of peptide-induced intracellular cytokine staining to measure multifunctional CD8+ T-cell responses in cured TB patients and healthy control individuals. In depth analysis of 18 prominently recognized HLA-A*0201-binding, Mtb peptides using CD8+ T-cells of cured TB patients, showed IFN-γ, IL-2, and TNF-α, mono-, dual-, and triple-positive CD8+ T-cells. Interestingly, in Mtb-non-infected individuals these polyfunctional CD8+ T-cells were absent, which argues for their priming during in vivo Mtb infection. Thus, this study is consistent with the notion that there is a much broader repertoire of CD8+ T-cells, which can be identified with specific bioinformatic approaches combined with functional immune assays.
An additional alternative strategy, not yet applied in the identification of new Mtb antigens, could be the use of HLA-conditional ligands for high-throughput tetramer generation (48). This technique allows production of HLA ligands that form stable complexes with HLA molecules but can be cleaved upon UV irradiation. The resulting empty, peptide-receptive HLA molecules can be loaded under native conditions with selected epitopes in a high-throughput and HLA-epitope tetramers can be generated and subsequently used for T-cell detection.
Summarized innovative detection methods for CD8+ T-cells can and will allow identification of protective- and pathogenic immunity, which can be used to monitor vaccine- and treatment efficacy.
HLA-class II presented+ human T-cell epitopes
A similar genome-wide approach was recently followed by Sette et al. to identify CD4+ T-cell epitopes: the Mtb genome was mined for potential peptide epitopes presented by HLA-class II molecules to CD4+ T-cells (49). The approach relied on predictions of HLA-binding capacity for a panel of DR, DP, and DQ alleles representative of those most commonly expressed in the general population, coupled with high-throughput ELISPOT assays. They found that secreted antigens as well as proteins involved in the active secretion process were dominant targets of the CD4+ T-cell response in the latently infected individuals tested in San Diego who successfully contain Mtb infection.
Responses were highly focused on three broadly immunodominant antigenic islands, all related to bacterial secretion systems and composed by several distinct ORFs. These data suggest that vaccination with one or few defined antigens will fail to replicate the response associated with natural immunity. Importantly, they also found that the CD4+ T-cells responding were largely restricted to the CXCR3+CCR6+ memory subset. The identification of this immunodominant population of memory T-cells characterized suggests that the response is shaped uniquely by Mtb-associated factors.
In analogy to the above results for CD8+ T-cells, these results again underline the power of an unbiased, genome-wide, Mtb antigen discovery approaches.
HLA-class Ib presented CD8+ human T-cell epitopes
The identification of Mtb antigens for human CD8+ T-cells focus has been focused almost entirely on classical HLA-class Ia and to a lesser extent, on CD1 a, b, c restricted CD8+ T-cell responses (50). In this respect, the polymorphism of the classical HLA-class Ia molecules (HLA-A, -B, -C) which include more than 500, 850, or 270 unique alleles, respectively, is an important factor to be considered for vaccine development due to the considerable variations in peptides that can bind to each HLA-class Ia molecule. In contrast, the HLA-class Ib genes HLA-E, -F, and -G exhibit restricted polymorphism with only 3, 4, and 10 alleles, respectively (51). This might argue for distinct roles for MHC-class Ia and class Ib molecules in host defense to infectious diseases. Non-classical HLA-class I molecules can present antigens of both self and foreign (microbial) origin to CD8+ T-cells (52–54). The fact that their allelic variation is limited provides an opportunity for vaccine-design. Since the two most important variants of HLA-E, HLA-ER (E*0101), and HLA-EG (E*0103), occur in equal frequencies amongst different populations (55) and can present antigens derived from pathogens including Mtb, Mtb antigen(s) recognized by HLA-E restricted CD8+ T-cell clones represent interesting targets for vaccine development. Moreover, HLA-E is not downregulated by HIV in contrast to HLA-A and -B offering possibilities for vaccination of HIV-infected individuals as well (56). However, the nature of the epitopes recognized by HLA-E restricted CD8+ T-cells remained unknown until recently (57).
Although HLA-E molecules can clearly interact with CD94 molecular complexes expressed predominantly by NK cells, HLA-E is also known to trigger microbial-specific cytotoxic CD8+ T-cells (58, 59). In mice, CD8+ T-cells restricted by the murine HLA-E equivalent, Qa-1 were found to have the ability to induce immune-suppression (60), revealing yet another function of non-classically restricted CD8+ T-cells.
To identify HLA-E-restricted Mtb antigens which could be exploited for TB vaccination, we applied bioinformatics, HLA-E peptide-binding assays, and immunological screening, in analogy to the approach used for identification of classical MHC-Ia restricted CD8 T-cell epitopes (61). With this method we identified 69 Mtb peptides, derived from a large variety of Mtb antigens, which were presented by HLA-E molecules to human CD8+ T-cells. We could demonstrate that CD8+ T-cells from both mycobacterium responsive adults and BCG-vaccinated infants, but not negative controls, proliferated in response to the identified Mtb peptides. These CD8+ T-cells displayed cytotoxic activity against target-cells expressing HLA-E loaded with specific peptides, in the absence of any class Ia molecules, and were able to lyse M. bovis BCG infected human macrophages demonstrating that HLA-E restricted antigens are naturally processed during infection. Besides, several HLA-E-restricted, Mtb-specific CD8+ T-cells were able to suppress proliferation of bystander CD4+ T-cells. This suppression depended on cell–cell contact and was mediated, at least in part, by membrane bound TGFβ1. These data underscore that human CD8+ T-cells are highly multifunctional, and can combine diverse functions such as suppressive and cytotoxic functions. If BCG vaccination would be able to prime HLA-E restricted T-cell responses, HLA-E peptide-based vaccines might be able to boost such BCG-primed responses. The dual character of the response induced in the context of HLA-E may lead to induction of protection as well as balanced regulation of inflammation, which, might be exploited to limit inflammatory pathology in TB.
In vivo expressed Mtb (IVE-TB) antigen-derived human T-cell epitopes
Since the lung represents the principal organ where TB disease is manifested, vaccine-induced immune responses need to target Mtb during pulmonary infection. In this respect, it is important to note the significant impact of the alteration of Mtb’s gene expression profile during intracellular stress inside host alveolar macrophages, on the Mtb antigen repertoire that is presented to the immune system. Thus, we hypothesized that a more profound understanding of the real time Mtb “antigenome” expressed during infection in the lung and its recognition by the human immune system, including non-classical T- and B-cells, is vital to develop better vaccination strategies. To identify novel antigens with vaccine potential, we used an unbiased genome-wide approach based on comprehensive gene expression data from Mtb during infection of four genetically related but distinct mouse strains (62). These strains represent key features of human TB with a spectrum of TB susceptibility, including the development of necrotic lesions and granuloma formation, which are regulated by the super-susceptibility to tuberculosis 1 (sst1) locus as well as modifier background genes (62). We investigated the in vivo expression of 2170 Mtb genes, most of which represent the first gene of each predicted Mtb operon, during infection in the lungs. To select candidate antigens, stringent selection approaches were then applied to identify a list of in vivo expressed Mtb (designated IVE-TB) genes. The resulting 16 most consistently expressed Mtb genes were produced as recombinant proteins and their immunogenicity was analyzed in PBMC of TST+ healthy, TB affected individuals, TB patients as well as long-term LTBI. Seven of the identified IVE-TB antigens were strongly immunogenic in TST+, ESAT-6/CFP10-responsive individuals, but not in E/C negative TST+ individuals and healthy mycobacterial naïve individuals, indicating that these antigens are presented during natural Mtb infection. Importantly, IVE-TB antigen-specific responses could be detected in long-term LTBI, who had been exposed to Mtb many years ago yet never developed TB symptoms despite not having had preventive treatment. The most pronounced T-cell subsets recognizing IVE-TB antigens were identified as IFN-γ+/TNF-α+ CD8+ T-cells and TNF-α+/IL-2+ CD154+CD4+ T-cells of which the former were major contributors to IFN-γ production. Since IFN-γ+/TNF-α+ CD8+ T-cells were also the most prominent subset in the response to Rpf and DosR proteins, this suggests that the development of specific differential T-cell subsets may be unrelated to the nature of the specific protein antigen involved.
Thus, the analysis of in vivo expression patterns during pulmonary Mtb infection to identify IVE-TB antigens, combined with detailed immune profiling in humans, led to the identification of IVE-TB as a new class of TB antigens, with the potential for TB vaccination. Our most recent results indeed reveal their protective efficacy in various animal models (63). Importantly, IVE-TB antigen discovery strategies can be applied also to other infectious diseases caused by complex pathogens and represent a novel approach for antigen discovery in general.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We gratefully acknowledge the support of the following organizations: the European Commission (FP7 projects NEWTBVAC, IDEA, ADITEC, EURIPRED, and VACTRAIN); the Bill and Melinda Gates Foundation Grand Challenges in Global Health (grants GC6#74); EDCTP through a project entitled AE-TBC under Grant Agreement No. IP_09_32040, the Netherlands Leprosy Relief Foundation (NLR) together with the Turing Foundation (ILEP#: 701.02.49), the Heiser Program for Research in Leprosy in The New York Community Trust (P13-000392), the Order of Malta-Grants-for-Leprosy-Research (MaLTALEP), and the Q. M. Gastmann Wichers Foundation. We thank all members of the laboratory for their great support. We would also like to express our gratitude to all other colleagues which have contributed greatly to parts of this research, in particular: May Young Lin, Francesco Dieli, Nadia Caccamo, Ole Lund, Sheila Tan, Marta Romano, Kris Huygen, Gregory Dolganov, Gary Schoolnik and Fredrik Oftung, and members of the BMGF supported Biomarkers for TB consortium.
References
1. Tameris MD, Hatherill M, Landry BS, Scriba TJ, Snowden MA, Lockhart S, et al. Safety and efficacy of MVA85A, a new tuberculosis vaccine, in infants previously vaccinated with BCG: a randomised, placebo-controlled phase 2b trial. Lancet (2013) 381(9871):1021–8. doi: 10.1016/S0140-6736(13)60177-4
2. Cole ST, Brosch R, Parkhill J, Garnier T, Churcher C, Harris D, et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature (1998) 393:537–44. doi:10.1038/31159
3. Ottenhoff TH, Doherty TM, van Dissel JT, Bang P, Lingnau K, Kromann I, et al. First in humans: a new molecularly defined vaccine shows excellent safety and strong induction of long-lived Mycobacterium tuberculosis-specific Th1-cell like responses. Hum Vaccin (2010) 6:1007–15. doi:10.4161/hv.6.12.13143
4. Blythe MJ, Zhang Q, Vaughan K, de Castro R Jr, Salimi N, Bui HH, et al. An analysis of the epitope knowledge related to mycobacteria. Immunome Res (2007) 3:10. doi:10.1186/1745-7580-3-10
5. Betts JC, Lukey PT, Robb LC, McAdam RA, Duncan K. Evaluation of a nutrient starvation model of Mycobacterium tuberculosis persistence by gene and protein expression profiling. Mol Microbiol (2002) 43:717–31. doi:10.1046/j.1365-2958.2002.02779.x
6. Voskuil MI, Schnappinger D, Visconti KC, Harrell MI, Dolganov GM, Sherman DR, et al. Inhibition of respiration by nitric oxide induces a Mycobacterium tuberculosis dormancy program. J Exp Med (2003) 198:705–13. doi:10.1084/jem.20030205
7. Ottenhoff TH, Kaufmann SH. Vaccines against tuberculosis: where are we and where do we need to go? PLoS Pathog (2012) 8:e1002607. doi:10.1371/journal.ppat.1002607
8. Ottenhoff TH. New pathways of protective and pathological host defense to mycobacteria. Trends Microbiol (2012) 20:419–28. doi:10.1016/j.tim.2012.06.002
9. Ottenhoff TH, Lewinsohn DA, Lewinsohn DM. Human CD4 and CD8 T cell responses to Mycobacterium tuberculosis: antigen specificity, function, implications and applications. In: Kaufmann SH, Britton WJ, editors. Handbook of Tuberculosis. Weinheim: Wiley-VCH Verlag GmbH & Co. KGaA (2008). p. 119–156.
10. Wayne LG, Hayes LG. An in vitro model for sequential study of shiftdown of Mycobacterium tuberculosis through two stages of nonreplicating persistence. Infect Immun (1996) 64:2062–9.
11. Shi L, Jung YJ, Tyagi S, Gennaro ML, North RJ. Expression of Th1-mediated immunity in mouse lungs induces a Mycobacterium tuberculosis transcription pattern characteristic of nonreplicating persistence. Proc Natl Acad Sci U S A (2003) 100:241–6. doi:10.1073/pnas.0136863100
12. Schnappinger D, Ehrt S, Voskuil MI, Liu Y, Mangan JA, Monahan IM, et al. Transcriptional adaptation of Mycobacterium tuberculosis within macrophages: insights into the phagosomal environment. J Exp Med (2003) 198:693–704. doi:10.1084/jem.20030846
13. Black GF, Thiel BA, Ota MO, Parida SK, Adegbola R, Boom WH, et al. Immunogenicity of novel DosR regulon-encoded candidate antigens of Mycobacterium tuberculosis in three high-burden populations in Africa. Clin Vaccine Immunol (2009) 16:1203–12. doi:10.1128/CVI.00111-09
14. Goletti D, Butera O, Vanini V, Lauria FN, Lange C, Franken KL, et al. Response to Rv2628 latency antigen associates with cured tuberculosis and remote infection. Eur Respir J (2010) 36:135–42. doi:10.1183/09031936.00140009
15. Leyten EM, Lin MY, Franken KL, Friggen AH, Prins C, van Meijgaarden KE, et al. Human T-cell responses to 25 novel antigens encoded by genes of the dormancy regulon of Mycobacterium tuberculosis. Microbes Infect (2006) 8:2052–60. doi:10.1016/j.micinf.2006.03.018
16. Lin MY, Ottenhoff TH. Host-pathogen interactions in latent Mycobacterium tuberculosis infection: identification of new targets for tuberculosis intervention. Endocr Metab Immune Disord Drug Targets (2008) 8:15–29. doi:10.2174/187153008783928398
17. Lin MY, Ottenhoff TH. Not to wake a sleeping giant: new insights into host-pathogen interactions identify new targets for vaccination against latent Mycobacterium tuberculosis infection. Biol Chem (2008) 389:497–511. doi:10.1515/BC.2008.057
18. Roupie V, Romano M, Zhang L, Korf H, Lin MY, Franken KL, et al. Immunogenicity of eight dormancy regulon-encoded proteins of Mycobacterium tuberculosis in DNA-vaccinated and tuberculosis-infected mice. Infect Immun (2007) 75:941–9. doi:10.1128/IAI.01137-06
19. Schuck SD, Mueller H, Kunitz F, Neher A, Hoffmann H, Franken KL, et al. Identification of T-cell antigens specific for latent Mycobacterium tuberculosis infection. PLoS One (2009) 4:e5590. doi:10.1371/journal.pone.0005590
20. Lin MY, Geluk A, Smith SG, Stewart AL, Friggen AH, Franken KL, et al. Lack of immune responses to Mycobacterium tuberculosis DosR regulon proteins following Mycobacterium bovis BCG vaccination. Infect Immun (2007) 75:3523–30. doi:10.1128/IAI.01999-06
21. Geluk A, Lin MY, van Meijgaarden KE, Leyten EM, Franken KL, Ottenhoff TH, et al. T-cell recognition of the HspX protein of Mycobacterium tuberculosis correlates with latent M. tuberculosis infection but not with M. bovis BCG vaccination. Infect Immun (2007) 75:2914–21. doi:10.1128/IAI.01990-06
22. Commandeur S, Lin MY, van Meijgaarden KE, Friggen AH, Franken KL, Drijfhout JW, et al. Double- and monofunctional CD4 and CD8 T-cell responses to Mycobacterium tuberculosis DosR antigens and peptides in long-term latently infected individuals. Eur J Immunol (2011) 41:2925–36. doi:10.1002/eji.201141602
23. Reece ST, Nasser-Eddine A, Dietrich J, Stein M, Zedler U, Schommer-Leitner S, et al. Improved long-term protection against Mycobacterium tuberculosis Beijing/W in mice after intra-dermal inoculation of recombinant BCG expressing latency associated antigens. Vaccine (2011) 29:8740–4. doi:10.1016/j.vaccine.2011.07.144
24. Gowthaman U, Singh V, Zeng W, Jain S, Siddiqui KF, Chodisetti SB, et al. Promiscuous peptide of 16 kDa antigen linked to Pam2Cys protects against Mycobacterium tuberculosis by evoking enduring memory T-cell response. J Infect Dis (2011) 204:1328–38. doi:10.1093/infdis/jir548
25. Aagaard C, Hoang T, Dietrich J, Cardona PJ, Izzo A, Dolganov G, et al. A multistage tuberculosis vaccine that confers efficient protection before and after exposure. Nat Med (2011) 17:189–94. doi:10.1038/nm.2285
26. Lin PL, Dietrich J, Tan E, Abalos RM, Burgos J, Bigbee C, et al. The multistage vaccine H56 boosts the effects of BCG to protect cynomolgus macaques against active tuberculosis and reactivation of latent Mycobacterium tuberculosis infection. J Clin Invest (2012) 122:303–14. doi:10.1172/JCI46252
27. Biketov S, Mukamolova GV, Potapov V, Gilenkov E, Vostroknutova G, Kell DB, et al. Culturability of Mycobacterium tuberculosis cells isolated from murine macrophages: a bacterial growth factor promotes recovery. FEMS Immunol Med Microbiol (2000) 29:233–40. doi:10.1111/j.1574-695X.2000.tb01528.x
28. Mukamolova GV, Kaprelyants AS, Young DI, Young M, Kell DB. A bacterial cytokine. Proc Natl Acad Sci U S A (1998) 95:8916–21. doi:10.1073/pnas.95.15.8916
29. Davies AP, Dhillon AP, Young M, Henderson B, McHugh TD, Gillespie SH. Resuscitation-promoting factors are expressed in Mycobacterium tuberculosis-infected human tissue. Tuberculosis (Edinb) (2008) 88:462–8. doi:10.1016/j.tube.2008.01.007
30. Mukamolova GV, Turapov OA, Young DI, Kaprelyants AS, Kell DB, Young M. A family of autocrine growth factors in Mycobacterium tuberculosis. Mol Microbiol (2002) 46:623–35. doi:10.1046/j.1365-2958.2002.03184.x
31. Rachman H, Strong M, Ulrichs T, Grode L, Schuchhardt J, Mollenkopf H, et al. Unique transcriptome signature of Mycobacterium tuberculosis in pulmonary tuberculosis. Infect Immun (2006) 74:1233–42. doi:10.1128/IAI.74.2.1233-1242.2006
32. Mukamolova GV, Turapov O, Malkin J, Woltmann G, Barer MR. Resuscitation-promoting factors reveal an occult population of tubercle bacilli in sputum. Am J Respir Crit Care Med (2010) 181:174–80. doi:10.1164/rccm.200905-0661OC
33. Huang W, Qi Y, Diao Y, Yang F, Zha X, Ren C, et al. Use of resuscitation-promoting factors proteins improves the sensitivity of culture-based tuberculosis testing in special samples. Am J Respir Crit Care Med (2014) 189(5):612–4. doi:10.1164/rccm.201310-1899LE
34. Gupta RK, Srivastava BS, Srivastava R. Comparative expression analysis of rpf-like genes of Mycobacterium tuberculosis H37Rv under different physiological stress and growth conditions. Microbiology (2010) 156:2714–22. doi:10.1099/mic.0.037622-0
35. Yeremeev VV, Kondratieva TK, Rubakova EI, Petrovskaya SN, Kazarian KA, Telkov MV, et al. Proteins of the Rpf family: immune cell reactivity and vaccination efficacy against tuberculosis in mice. Infect Immun (2003) 71:4789–94. doi:10.1128/IAI.71.8.4789-4794.2003
36. Geluk A, van den Eeden SJ, van Meijgaarden KE, Dijkman K, Franken KL, Ottenhoff TH. A multistage-polyepitope vaccine protects against Mycobacterium tuberculosis infection in HLA-DR3 transgenic mice. Vaccine (2012) 30(52):7513–21. doi:10.1016/j.vaccine.2012.10.045
37. Cho S, Mehra V, Thoma-Uszynski S, Stenger S, Serbina N, Mazzaccaro RJ, et al. Antimicrobial activity of MHC class I-restricted CD8+ T cells in human tuberculosis. Proc Natl Acad Sci U S A (2000) 97:12210–15. doi:10.1073/pnas.210391497
38. Flynn JL, Chan J. Tuberculosis: latency and reactivation. Infect Immun (2001) 69:4195–201. doi:10.1128/IAI.69.7.4195-4201.2001
39. Lalvani A, Brookes R, Wilkinson RJ, Malin AS, Pathan AA, Andersen P, et al. Human cytolytic and interferon gamma-secreting CD8+ T lymphocytes specific for Mycobacterium tuberculosis. Proc Natl Acad Sci U S A (1998) 95:270–5. doi:10.1073/pnas.95.1.270
40. Stenger S, Mazzaccaro RJ, Uyemura K, Cho S, Barnes PF, Rosat JP, et al. Differential effects of cytolytic T cell subsets on intracellular infection. Science (1997) 276:1684–7. doi:10.1126/science.276.5319.1684
41. Ab BK, Kiessling R, Van Embden JD, Thole JE, Kumararatne DS, Pisa P, et al. Induction of antigen-specific CD4+ HLA-DR-restricted cytotoxic T lymphocytes as well as nonspecific nonrestricted killer cells by the recombinant mycobacterial 65-kDa heat-shock protein. Eur J Immunol (1990) 20:369–77. doi:10.1002/eji.1830200221
42. Lewinsohn DA, Winata E, Swarbrick GM, Tanner KE, Cook MS, Null MD, et al. Immunodominant tuberculosis CD8 antigens preferentially restricted by HLA-B. PLoS Pathog (2007) 3:1240–9. doi:10.1371/journal.ppat.0030127
43. Smith SM, Brookes R, Klein MR, Malin AS, Lukey PT, King AS, et al. Human CD8+ CTL specific for the mycobacterial major secreted antigen 85A. J Immunol (2000) 165:7088–95. doi:10.4049/jimmunol.165.12.7088
44. Smith SM, Klein MR, Malin AS, Sillah J, McAdam KP, Dockrell HM. Decreased IFN-gamma and increased IL-4 production by human CD8(+) T cells in response to Mycobacterium tuberculosis in tuberculosis patients. Tuberculosis (Edinb) (2002) 82:7–13. doi:10.1054/tube.2001.0317
45. Caccamo N, Guggino G, Meraviglia S, Gelsomino G, Di CP, Titone L, et al. Analysis of Mycobacterium tuberculosis-specific CD8 T-cells in patients with active tuberculosis and in individuals with latent infection. PLoS One (2009) 4:e5528. doi:10.1371/journal.pone.0005528
46. Lewinsohn DM, Swarbrick GM, Cansler ME, Null MD, Rajaraman V, Frieder MM, et al. Human CD8 T Cell antigens/epitopes identified by a proteomic peptide library. PLoS One (2013) 8:e67016. doi:10.1371/journal.pone.0067016
47. Tang ST, van Meijgaarden KE, Caccamo N, Guggino G, Klein MR, van Weeren P, et al. Genome-based in silico identification of new Mycobacterium tuberculosis antigens activating polyfunctional CD8+ T cells in human tuberculosis. J Immunol (2011) 186:1068–80. doi:10.4049/jimmunol.1002212
48. Rodenko B, Toebes M, Hadrup SR, van Esch WJ, Molenaar AM, Schumacher TN, et al. Generation of peptide-MHC class I complexes through UV-mediated ligand exchange. Nat Protoc (2006) 1:1120–32. doi:10.1038/nprot.2006.121
49. Lindestam Arlehamn CS, Gerasimova A, Mele F, Henderson R, Swann J, Greenbaum JA, et al. Memory T cells in latent Mycobacterium tuberculosis infection are directed against three antigenic islands and largely contained in a CXCR3+CCR6+ Th1 subset. PLoS Pathog (2013) 9:e1003130. doi:10.1371/journal.ppat.1003130
50. Kaufmann SH. How can immunology contribute to the control of tuberculosis? Nat Rev Immunol (2001) 1:20–30. doi:10.1038/35095558
51. Available from: http://hla.alleles.org/alleles/class1.html
52. Mazzarino P, Pietra G, Vacca P, Falco M, Colau D, Coulie P, et al. Identification of effector-memory CMV-specific T lymphocytes that kill CMV-infected target cells in an HLA-E-restricted fashion. Eur J Immunol (2005) 35:3240–7. doi:10.1002/eji.200535343
53. Pietra G, Romagnani C, Mazzarino P, Falco M, Millo E, Moretta A, et al. HLA-E-restricted recognition of cytomegalovirus-derived peptides by human CD8+ cytolytic T lymphocytes. Proc Natl Acad Sci U S A (2003) 100:10896–901. doi:10.1073/pnas.1834449100
54. Salerno-Goncalves R, Fernandez-Vina M, Lewinsohn DM, Sztein MB. Identification of a human HLA-E-restricted CD8+ T cell subset in volunteers immunized with Salmonella enterica serovar typhi strain Ty21a typhoid vaccine. J Immunol (2004) 173:5852–62. doi:10.4049/jimmunol.173.9.5852
55. Strong RK, Holmes MA, Li P, Braun L, Lee N, Geraghty DE. HLA-E allelic variants. Correlating differential expression, peptide affinities, crystal structures, and thermal stabilities. J Biol Chem (2003) 278:5082–90. doi:10.1074/jbc.M208268200
56. Cohen GB, Gandhi RT, Davis DM, Mandelboim O, Chen BK, Strominger JL, et al. The selective downregulation of class I major histocompatibility complex proteins by HIV-1 protects HIV-infected cells from NK cells. Immunity (1999) 10:661–71. doi:10.1016/S1074-7613(00)80065-5
57. Heinzel AS, Grotzke JE, Lines RA, Lewinsohn DA, McNabb AL, Streblow DN, et al. HLA-E-dependent presentation of Mtb-derived antigen to human CD8+ T cells. J Exp Med (2002) 196:1473–81. doi:10.1084/jem.20020609
58. Garcia P, Llano M, de Heredia AB, Willberg CB, Caparros E, Aparicio P, et al. Human T cell receptor-mediated recognition of HLA-E. Eur J Immunol (2002) 32:936–44. doi:10.1002/1521-4141(200204)32:4<936::AID-IMMU936>3.3.CO;2-D
59. Pietra G, Romagnani C, Falco M, Vitale M, Castriconi R, Pende D, et al. The analysis of the natural killer-like activity of human cytolytic T lymphocytes revealed HLA-E as a novel target for TCR alpha/beta-mediated recognition. Eur J Immunol (2001) 31:3687–93. doi:10.1002/1521-4141(200112)31:12<3687::AID-IMMU3687>3.0.CO;2-C
60. Sarantopoulos S, Lu L, Cantor H. Qa-1 restriction of CD8+ suppressor T cells. J Clin Invest (2004) 114:1218–21. doi:10.1172/JCI200423152
61. Joosten SA, van Meijgaarden KE, van Weeren PC, Kazi F, Geluk A, Savage ND, et al. Mycobacterium tuberculosis peptides presented by HLA-E molecules are targets for human CD8 T-cells with cytotoxic as well as regulatory activity. PLoS Pathog (2010) 6:e1000782. doi:10.1371/journal.ppat.1000782
62. Commandeur S, van Meijgaarden KE, Prins C, Pichugin AV, Dijkman K, van den Eeden SJ, et al. An unbiased genome-wide Mycobacterium tuberculosis gene expression approach to discover antigens targeted by human T cells expressed during pulmonary infection. J Immunol (2013) 190:1659–71. doi:10.4049/jimmunol.1201593
63. Commandeur S, van den Eeden SJF, Dijkman K, Clark SO, van Meijgaarden KE, Wilson L, et al. The in vivo expressed Mycobacterium tuberculosis (IVE-TB) antigen Rv2034 induces CD4+ T-cells that protect against pulmonary infection in HLA-DR transgenic mice and guinea pigs. Vaccine (2014) 32:3580–8. doi:10.1016/j.vaccine.2014.05.005
Keywords: cellular immunity, CD4, CD8, Mycobacterium tuberculosis, TB, T-cell epitopes, vaccines
Citation: Geluk A, van Meijgaarden KE, Joosten SA, Commandeur S and Ottenhoff THM (2014) Innovative strategies to identify M. tuberculosis antigens and epitopes using genome-wide analyses. Front. Immunol. 5:256. doi: 10.3389/fimmu.2014.00256
Received: 31 March 2014; Paper pending published: 02 May 2014;
Accepted: 16 May 2014; Published online: 25 June 2014.
Edited by:
Juraj Ivanyi, King’s College London, UKReviewed by:
Alexis DeHaven Dunkle, Lawrence Livermore National Laboratory, USARajko Reljic, St George’s University of London, UK
Copyright: © 2014 Geluk, van Meijgaarden, Joosten, Commandeur and Ottenhoff. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Annemieke Geluk, Department of Infectious Diseases, Leiden University Medical Center, PO Box 9600, Leiden 2300 RC, Netherlands e-mail:YS5nZWx1a0BsdW1jLm5s
 Annemieke Geluk
Annemieke Geluk Krista E. van Meijgaarden
Krista E. van Meijgaarden Simone A. Joosten
Simone A. Joosten Susanna Commandeur
Susanna Commandeur Tom H. M. Ottenhoff
Tom H. M. Ottenhoff