- 1Department of Internal Medicine I, BMT, Medical University of Vienna, Vienna, Austria
- 2Department of Radiation Oncology, Klinikum rechts der Isar, Technische Universität München, Munich, Germany
- 3Department of Hematology, Hemostasis, Oncology and Stem Cell Transplantation, Transplantation Biology, Hannover Medical School, Hannover, Germany
- 4Hematological Sciences, Institute of Cellular Medicine, Newcastle University, Newcastle upon Tyne, UK
- 5Division of Hematology, Medical University of Graz, Graz, Austria
- 6Department of Internal Medicine III, University Hospital of Regensburg, Regensburg, Germany
Allogeneic hematopoietic stem cell transplantation (allo-HSCT) is the main curative therapy for hematological malignancy such as leukemias, lymphomas, or multiple myelomas and some other hematological disorders. In this therapy, cure of hematological diseases relies on graft-versus-malignancy effects by allogenic immune cells. However, severe posttransplant treatment-associated complications such as acute graft-versus-host disease (aGvHD) and chronic graft-versus-host disease (cGvHD) limit this approach. Most research into GvHD has concentrated on the aGvHD, while the more complex and multifaceted chronic form has been largely poorly investigated. cGvHD is a multi-organ autoimmune disorder and is the major cause of non-relapse morbidity and mortality following allo-HSCT, occurring in about 50% of patients, or 13,000–15,000 patients per year worldwide. Therefore, there is a high medical need for an early prediction of these therapy-associated toxicities. Biomarkers have gained importance over the last decade in diagnosis, in prognosis, and in prediction of pending diseases or side effects. Biomarkers can be cells, factors isolated from target tissues, or soluble factors that can be detected in body fluids. In this review, we aim to summarize some of the recent developments of biomarkers in the field of allo-HSCT. We will focus on cell-based biomarkers (B-cell subsets) for cGvHD and soluble factors including microRNA (miRNA), which are excreted into serum/plasma and urine. We also discuss the potential role of cytosolic and extracellular 70 kDa heat shock proteins (HSP70) as potential biomarkers for aGvHD and their role in preclinical models. Proteomic biomarkers in the blood have been used as predictors of treatment responses in patients with aGvHD for many years. More recently, miRNAs have been found to serve as a biomarker to diagnose aGvHD in the plasma. Another development relates to urine-based biomarkers that are usually detected by capillary electrophoresis and mass spectrometry. These biomarkers have the potential to predict the development of severe aGvHD (grades III–IV), overall mortality, and the pending development of cGvHD in patients posttransplant.
Complications of Hematopoietic Stem Cell Transplantation (HSCT)
Acute and chronic graft-versus-host disease (a/cGvHD) are serious and frequent complications after allogeneic hematopoietic stem cell transplantation (allo-HSCT) that negatively impact on survival and quality of life of patients (1, 2). GvHD develops in approximately 40–60% of recipients after allo-HSCT. Donor-derived T-cells targeting alloantigens of the recipient play a key role in the induction of GvHD. Donor T-cells generally destroy host tissues by secreting pro-inflammatory cytokines such as interleukin-1 (IL-1), tumor necrosis factor alpha (TNF-α), and interferon gamma (IFN-γ) or by direct cytolytic activities of immune effector cells (1, 2). So far, GvHD is diagnosed based on unique diagnostic clinical signs and symptoms, as recommended by the National Institutes of Health (NIH) consensus development conferences (3, 4). In 2014, the NIH Consensus Conference updated requirements for the integration of assessment of potential biomarkers in prospective clinical studies of GvHD (4). Biomarkers that could be obtained by minimal invasive methods would be beneficial to predict GvHD and thus would increase safety and quality of life of patients. Biomarkers should fulfill certain requirements such as the confirmation of an aGvHD or cGvHD, objectively measure disease activity, allow a distinction of organ damage, should provide prognostic risk assessment, and predict responses to therapy (5). Detailed documentations on collection of specimens including excellent clinical data on patient characteristics and HSCT course, and standardized data analyses are crucial for further data processing (5). cGvHD is a multi-organ autoimmune disorder and is the major cause of non-relapse morbidity and mortality following allo-HSCT, occurring in about 50% of patients, or 13,000–15,000 patients per year worldwide that causes a plethora of comorbidities including cardiovascular, gastrointestinal (GI), liver, pulmonary, endocrine (diabetes, hyperlipidemia, thyroid and adrenal insufficiency, hypogonadism), neuropsychiatric (e.g., depression, chronic neurologic diseases), bone and joint (osteoarthritis, osteoporosis) disorders, infections (bacterial, viral, and fungal), and other more specific comorbidities (solid malignancy, obesity, and infertility) (1–5). Herein, we discuss B-cell-based, stress protein-, and microRNA (miRNA)-based biomarkers as predictors for a/cGvHD.
Cellular Biomarkers
The Role of T/B-Cells As Cellular Biomarkers in cGvHD
For a long time it has been known that cross-reactive allo-T cells that were immunized by environmental antigens derived from the donor are key players in the induction of GvHD. Therefore, the determination of these cells and the measurement of their cytolytic activities against host tissues have been used to determine GvHD (6). The outcome of haploidentical HSCT can be improved by the depletion of T cells from the donor graft. However, this procedure is often accompanied by graft failure and an increased incidence of GvHD, which could be overcome by megadose HSCT, injecting of pathogen-specific T cells to rebuild immunity or engineered T cells to induce suicide in case of allo-reactivity (7). The sequential infusion of regulatory T cells (Tregs) (CD4+/CD25+ and CD4+/CD25−) after HSCT, the selective ex vivo (photo) depletion of certain autoreactive T cell clones, the preservation of γ/δ T cells in the stem cell graft, and the selection of the best stem cells provide other options to improve GVL effects while GvHD is not increased (7). All these procedures contribute to fewer infection and toxicity rates and leukemia-related death cases.
Recent research has demonstrated that apart from T cells, B-cells also play key roles in the pathogenesis of cGvHD. Therefore, the presence of auto- and alloantibodies, elevated plasma levels of B-cell activation factor (BAFF), a cytokine of the tumor necrosis family, and an accumulation of CD19+CD21low B-cells serve as biomarkers for GvHD. Apart from the depletion of T-cells by antibodies, the depletion of certain B-cell subpopulations might also provide a promising strategy to avoid GvHD (8–10). A delayed B-cell reconstitution with relative B-cell lymphopenia can result in downregulated B-cell counts in patients after HSCT (9–12). Low B-cell counts in the circulation may be explained in part by the insufficient production of B-cells in the bone marrow, as previously reported in patients with both, aGvHD and cGvHD (13). In contrast, a dysregulated B-cell homeostasis with persistent high BAFF levels can induce an upregulation of certain subpopulations of B-cells. In patients who do not develop cGvHD, elevated BAFF levels normalize after 6 months, whereas these remain highly elevated in patients developing cGvHD at later time points (11, 12). The observed high BAFF/B-cell ratio in patients with cGvHD suggests that during B-cell deficiency, autoreactive B-cell clones that would otherwise undergo negative selection could potentially survive due to an excess of BAFF, which in turn could possibly contribute to the pathophysiology of cGvHD (14–16). Furthermore, increased B-cell activation, aberrant B-cell signaling, and prolonged survival of activated B-cells have been found to be associated with cGvHD (17).
Perturbation of B-cell homeostasis can be associated with elevated or decreased numbers of different B-cell subpopulations during cGvHD (8, 11, 12, 16, 18, 19). Greinix and colleagues reported on elevated relative numbers of CD19+CD21low B-cells in patients with active cGvHD compared to those without cGvHD in a retrospective study on 70 patients (8). In addition, CD19+CD21low B-cell counts higher than 15% in patients with active cGvHD were found to be significantly associated with the presence of severe opportunistic infections (8). Furthermore, the memory B-cell compartment showed significantly lower relative and absolute numbers of both, non-class-switched CD19+CD27+IgD+ and class-switched CD19+CD27+IgD− memory B-cells. This observed perturbation of circulating B-cell subpopulations could be useful for assessing cGvHD activity and for identifying cGvHD patients at risk for severe infectious complications (8).
Kuzmina and colleagues investigated whether the number of CD19+CD21low B-cells could predict the outcome of extracorporeal photopheresis (ECP), which is used as one option for an immunomodulatory treatment of cGvHD (19). ECP non-responders had significantly higher (p = 0.02) relative numbers of CD19+CD21low B-cells (mean = 22%) in the peripheral blood prior to the start of ECP compared to patients achieving a complete response (CR) (mean = 8%) and partial response (mean = 16%) after 6 months of ECP therapy. These data suggest that CD19+CD21low B-cell counts could serve as a predictive cellular biomarker. Moreover, CR patients had significantly lower relative numbers of CD19+CD21low B-cells 6, 12, and 21 months after start of ECP compared to non-responders, confirming that CD19+CD21low B-cells could be potential cellular biomarkers for objective response assessment in cGvHD (19).
CD19+CD21low B-cells were further investigated in cGvHD patient cohorts in the context of impaired humoral immunity defined by increased or decreased serum immunoglobulin G (IgG) levels (11). cGvHD patients with hypogammaglobulinemia had significantly decreased absolute numbers of CD19+ B-cells with elevated percentages of CD19+CD21low B-cells and transitional CD19+CD21int-highCD38highIgMhigh B-cells compared to cGvHD cohorts with normogammaglobulinemia or hypergammaglobulinemia, respectively. Furthermore, cGvHD patients with hypogammaglobulinemia also had a significant reduction of non-class-switched CD19+CD27+IgD+ and class-switched CD19+CD27+IgD− memory B-cells compared to the other two cohorts. Of note, cGvHD patients with hypergammaglobulinemia presented with significantly higher BAFF/B-cell ratios and frequently had significantly more autoantibodies present compared to the hypogammaglobulinemia cohort (11). Taken together, these data suggested that B-cell subpopulations could indicate different pathogenic mechanisms involved in cGvHD and might allow a distinction between immunodeficiency and autoimmunity (11, 16).
Investigation of B-cell subpopulations in the context of specific organ involvement by cGvHD revealed significantly decreased absolute and relative numbers of CD19+ B-cells in patients with newly diagnosed lung involvement seen as bronchiolitis obliterans syndrome (BOS) (4, 16). The prognosis of BOS is poor, and therefore, the identification of patients at an early disease stage when they may have an improved response to immunosuppressive therapy is a major clinical challenge. Kuzmina and colleagues observed that patients with newly diagnosed BOS had significantly increased relative numbers of CD19+CD21low B-cells (25.5 versus 6.6%, p < 0.0001) and BAFF/CD19+ ratio (0.18 versus 0.02 ng/103 CD19+ B-cells, p = 0.007) compared with patients without cGvHD (16). Asymptomatic patients with a drop in pulmonary function tests and NIH-defined changes in high-resolution CT scan could be distinguished from patients without cGvHD due to significantly higher CD19+CD21low B-cells and ratio of BAFF/CD19+ B-cells. Interestingly, relative numbers of CD19+CD21low B-cells were significantly elevated both at onset of BOS as well as in patients with long-lasting BOS not responding to immunosuppressive treatment confirming a role of CD19+CD21low B-cells as cellular biomarkers for objective diagnosis of lung involvement as well as continued cGvHD activity during the course of therapy.
In a large prospective study with 227 patients, Greinix and colleagues performed serial analyses starting on day 100 after HSCT to investigate CD19+CD21low B-cells as diagnostic and predictive cellular biomarkers in patients with newly diagnosed cGvHD (18). Higher relative numbers of CD19+CD21low B-cells, analyzed on day 100 after HSCT and compared to time-matched controls without cGvHD, were significantly associated with later development of cGvHD independently of clinical parameters (23.5 versus 15.2%, p = 0.004). Furthermore, significantly higher percentages of CD19+CD21low B-cells were also associated with first diagnosis of cGvHD (18.3 versus 9.9%, p = 0.001) (18). Although their exact biological functions still need to be elucidated, it is known that CD19+CD21low B-cells express inflammatory tissue-homing receptors, such as CXCR3 reflecting an increased capacity to home to inflammatory sites (20). Suryani and colleagues reported the production of significantly higher amounts of anti-nuclear autoantibodies by CD19+CD21low B-cells compared to other B-cell subpopulations such as naïve CD19+CD10−CD21highCD27− and memory CD19+CD27+IgD− B-cells, respectively (21). However, recent findings suggest that CD19+CD21low B-cells share the phenotype of anergic B-cells and do not proliferate in response to normal B-cell stimulation factors (22, 23).
Recently, regulatory B-cells (Bregs) have been shown to be involved in the pathogenesis of cGvHD (22, 24). Khoder and colleagues reported an enrichment of interleukin-10 (IL-10)-producing Bregs within memory CD19+IgM+CD27+ and transitional CD19+CD24hiCD38hi B-cells in healthy individuals (25). In patients with cGvHD, Breg cells were found to be decreased and thus less likely to produce IL-10 compared to healthy donors and patients without cGvHD (25). Another study reported a decrease of CD24hiCD27+ B-cells and IL-10-producing CD24hiCD27+ B-cells in patients with active cGvHD (26). Moreover, de Masson and colleagues observed increased CD24−CD38hi plasmablast frequencies, but decreased IL-10-producing plasmablasts in patients with active cGvHD compared to ones without cGvHD (22). Since CD24hiCD27+ B-cells and plasmablasts are among the most elevated cellular subsets within the Breg cell compartment, these observations could further support a possible role of Breg cells in the pathogenesis of cGvHD (22). Whether these novel cellular subpopulations could serve as biomarkers of active cGvHD, however, has to await further studies on well-defined patient cohorts including time-matched controls without cGvHD.
CD4+CD25+Foxp3+ Tregs have an indispensable role in the maintenance of tolerance after allo-HSCT. Poor reconstitution of Tregs after HSCT correlated with subsequent development of cGvHD (27–29). Furthermore, patients with long-lasting cGvHD are known to have a deficiency of Tregs in the circulation (26, 27). These findings led to therapeutic interventions aiming at enhancing Treg cell numbers by administration of low-dose interleukin-2 and thus, suppressing clinical manifestations of cGvHD (26). Based on these results, Tregs can be considered to be diagnostic and predictive cellular biomarkers of cGvHD.
Regarding other T-cell subpopulations, an increase in CCR7−CD45RA+CD8 T-cells that are effector memory T-cells and a decrease of CCR7+CD45RA+ naïve T-cells as well as CCR7+CD45RA− central memory T-cells has been reported in patients with cGvHD (30). The authors speculated whether their findings were the consequence of prolonged alloantigen exposure or interleukin-15 (IL-15) stimulation since patients with cGvHD reportedly have elevated IL-15 levels and effector memory T-cells can be generated by IL-15 stimulation. Yamashita and colleagues also observed a significantly higher percentage of CCR7−CD62LlowCD4+ effector memory T-cells in patients with cGvHD compared to ones without cGvHD or healthy donors (31). Furthermore, a preponderance of effector memory CD4+ T-cells relative to CCR7+CD45RA− central memory T-cells was observed in severe cGvHD by these investigators. In view of the small patient numbers in these single-center studies, confirmation and validation of these findings is required prior to using these T-cell subpopulations as diagnostic cellular biomarkers of cGvHD.
Greinix and colleagues recently reported that CD4+CD45RA+ naïve T-cells and CD4+CD45RA+CD31+ T-cells were significantly increased in patients with newly diagnosed cGvHD compared to a time-matched patient cohort without cGvHD (18). Furthermore, this T-cell subset when measured prospectively on day +100 after HSCT was also significantly elevated in patients subsequently developing cGvHD compared to patients never experiencing cGvHD. Besides these T-cell subpopulations, CD3+CD56+ NKT cells were significantly increased on day +100 after HSCT in patients subsequently developing cGvHD and thus, could also serve as a predictive cellular biomarker.
Soluble Biomarkers
70-kDa Heat Shock Protein Family (HSP70) As a Biomarker for Tumors and GvHD
Members of the HSP70 family are known to play an important role in transport, folding, and unfolding of proteins and also in the induction of immune responses (32–38). The major stress-inducible Hsp70 has been found to be upregulated in many different diseases including inflammation, autoimmunity, and tumors (32, 39). Hsp70 is also found on the cell surface of many tumor types including leukemic cells via a tumor-specific lipid anchorage (40). Furthermore, it was shown that membrane Hsp70 positive, viable tumor cells are able to secrete Hsp70 into the circulation in lipid vesicles, termed exosomes (33, 41). Therefore, exosomal Hsp70 serves as a tumor-specific biomarker for viable tumor mass (42). Functionally, extracellular HSPs, either alone or in combination with antigenic peptides, play important roles in the induction of inflammatory immune responses (34, 35, 43, 44). Apart from lipid-bound Hsp70, Hsp70 can be released by necrotic and inflamed tissues as a free molecule. This means that the major stress-inducible Hsp70 is secreted into the circulation either by living tumor cells in lipid vesicles, presumably tumor exosomes (41), or as a free molecule by dying necrotic and/or inflamed tissues. In cGvHD, inflammatory responses can occur in skin (77%), lung (50%), mouth (63%), liver (58%), eye (54%), joints (32%), GI (20%), and genital tract (16%). Therefore, free Hsp70, antibodies directed against Hsp70, mRNA levels might provide potential novel molecular biomarkers to diagnose and predict the onset of inflammation in cGvHD. Apart from members of the HSP70 family other stress proteins are discussed as potential markers for GvHD.
In a rat skin explant model, Novota et al could demonstrate that the two major stress-inducible genes Hsp70-1 and Hsp70-2 were upregulated. Therefore, elevated mRNA levels were associated with the grade of graft-versus-host reactions (GvHR) (45). Moreover, in the study by Kim et al., it was shown that a polymorphism in Hsp70-hom plays an important role in the prognosis of patients who received a sibling HLA-matched allo-HSCT (46). Thus, Hsp70-hom gene polymorphism might also serve as a prognostic marker for GvHD (47). A subsequent study by Bogunia-Kubik et al showed that patients who were homozygous for the A allele of the Hsp70-hom +2,663 SNP presented more frequently with grade II and IV toxic lesions and aGvHD compared to patients with different genotypes (48).
Other studies clearly demonstrated an involvement of the Hsp70 protein levels in the pathogenesis of GvHD. In a rat GvHD model that was induced by the injection of DA parental lymphoid cells into irradiated (LEW × DA)F1 rats, it was shown that the expression levels of Hsp70 were significantly increased in spleen and lymph nodes 7 and 14 days after induction of the disease (49). At later time points of the disease, the Hsp70 levels dropped to levels that were comparable to that of untreated control animals.
Different results exist with respect to HSP70-specific antibodies in the circulation in a rat model and human patients. In a rat model, elevated levels of antibodies (IgM, IgG2a, and IgG2b) directed against HSP70 have been found to be associated with the onset of symptoms that were associated with the development of GvHD (50), whereas in pediatric patients, this association was not detected after allo-HSCT (51). In subsequent human studies however, elevated anti-HSP70 and anti-HSP90 antibody levels were found to be associated with the development of a/cGvHD (52).
An association of the severity of GvHD disease and the expression of Hsp70 might be explained by the immunomodulatory activity of Hsp70. To date, several studies reported an involvement of Hsp70 in the production of pro-inflammatory cytokines such as TNF-α, IL-1, IL-6, and IL-12 and the secretion of nitric oxide (NO) and C-C chemokines by dendritic cells, monocytes, and macrophages (53, 54). Moreover, Hsp70 can activate intracellular signaling cascades that influence immunoregulatory functions of immune cells through binding of either free or lipid-bound Hsp70 to specific cell surface receptors such as Toll-like receptors 2 and 4 (TLR2/4), scavenger receptor CD36, low-density lipoprotein receptor-related protein CD91, C-type lectin receptor LOX-1, scavenger receptor SR-A, and CD40 (35). Through the induction of pro-inflammatory cytokines, Hsp70 contributes to the pathogenesis of autoimmune and various chronic inflammatory diseases (55–58). Thus, Hsp70 can serve as a damage-associated molecular pattern that activates the host’s adaptive and innate immune system by initiating alloimmunity (59). Another mechanism of the immunoregulatory activity of Hsp70 is based on its peptide-binding capacity. Exogenous Hsp70 can chaperone and cross-present antigens and cargo them to the antigen-presenting cells and thus elicit adaptive immune responses (44, 60). Therefore, it is not too surprising that Hsp70 is frequently upregulated in allografts, which in turn results in the progression of disease and allograft rejection (61–63).
Presumably, the modulation of Hsp70 levels might provide a therapeutic option for improving the outcome of GvHD. Previously, Oh et al showed, in a skin graft model, in which Hsp70 (Hsp70.1 gene) knock-out (KO) mice were used either as a donor or recipient, the importance of Hsp70 in acute allograft rejection (64). Allogeneic cells derived from Hsp70 KO mice were shown to induce lower rejection rates in recipients than those of Hsp70 wild-type animals. Therefore, the application of reagents that are able to silence Hsp70 expression may provide a promising strategy to reduce the risk for GvHD. The reduction of HSP70 levels by the administration of 15-deoxyspergualin (DSG), an immunosuppressive agent that binds to a constitutively expressed member of the 70-kDa heat shock protein family, has been shown to significantly reduce GvHD-associated mortality (65). DSG treatment reduced HSP70 levels in spleen and lymph nodes, inhibited the anti-HSP70 antibody production, and reduced the serum levels of IL-2, IFN-γ, TNF-α, and IL-10 (64).
In conclusion, monitoring levels of HSP70 protein and/or anti-HSP70 antibody levels in the serum after HSCT might serve as a diagnostic tool to predict the onset of GvHD. In addition, genetic or pharmacological modulation of the HSP70 expression may have a therapeutic potential in the treatment of GvHD.
Protein Markers in Body Fluids for Diagnosis of aGvHD
Serum, plasma (66–68), and saliva have been analyzed for prediction or diagnosis of aGvHD and cGvHD (69–71). A surface-enhanced laser desorption/ionization (SELDI) mass spectrometer was used to analyze plasma/serum in patients with and without GvHD post-HSCT by the group of Barrett in 2006 at the National Heart, Lung, and Blood Institute, National Institute of Health (NIH). They used SELDI to identify proteins/peptides differentially excreted into plasma (69–71). Another group from the NIH analyzed salvia samples using SELDI, no further data have been reported for either study. Therefore, plasma proteomics may be a more promising approach. The group of Ferrara and colleagues have intensively studied the serum/plasma proteomic approach using enzyme-linked immunosorbent assays to detect the presence of several proteins in the serum/plasma of patients post allo-HSCT (66–68). Among other markers, they have described plasma markers for GI aGvHD (68), regenerating islet-derived protein 3a (Reg-3a) and suppression of tumorigenicity (ST2) as diagnostic markers for aGvHD and survival after HSCT. Reg-3a was tested in samples from 1,014 HSCT patients from three transplantation centers and Reg3a concentrations were threefold higher in patients at onset of GI GvHD than in all other patients and correlated with lower GI GvHD. Reg3a concentrations at time of GvHD onset predicted response to therapy at 4 weeks, 1-year NRM, and 1-year survival (p ≤ 0.001). In a multivariate analysis, advanced clinical stage, severe histologic damage, and high Reg3α concentrations at GvHD diagnosis independently predicted 1-year NRM. The combination of Reg3a with clinical stage and histologic grade of GvHD can be used to improve risk stratification of patients.
The described biomarkers have been tested in two different centers (67, 68). To fulfill the criteria of a reliable biomarker, the biomarkers have to be defined in a test set, confirmed in a first validation set and then validated in a multicenter validation trial (5), as biomarkers need to be robust under different clinical conditions and prophylaxis strategies.
The group of Levine and Ferrara has recently achieved this goal for the so-called Ann Arbor Score of aGvHD, which relies on three biomarkers (Reg3 alpha, ST2, and sTNFR1). If this score is high at the time of onset of GvHD, it strongly predicts day 28 treatment response and 6-month NRM irrespective of center-specific strategies (72).
An international Mount Sinai Acute GvHD International Consortium (MAGIC consortium) has been recently founded, which prospectively monitors clinical data and samples from more than 1,000 pts receiving allogeneic SCT/year and now allows to develop biomarker score-based treatment stratification at the time of onset of GvHD (73).
None of the serum/plasma markers described to date can predict aGvHD development, but they can help to diagnose aGvHD, especially GI aGvHD and to define prognosis at a very early time point where clinical presentation fails to allow exact prediction. Earlier biomarker scores (e.g., at day 7) after HSCT are currently developed and hopefully will allow stratification of preemptive treatment in the future. Taken together, the proteomic monitoring of patients holds promise for early diagnosis as well as risk stratification of patients.
Serum/plasma biomarker development for cGvHD is yet not at the state of prospective multicenter monitoring. Chemokine (C-X-C motif) ligand 9 (CXCL9), suppression of tumorigenicity 2 (ST2), osteopontin, and soluble BAFF levels are candidates among others (74–77). In particular, recent work by Yu et al has described a biomarker panel for cGvHD using quantitative proteomic profiling by high-resolution tandem mass spectrometry. Pooled plasma from patients with and without cGvHD at matched time points posttransplant were tested in a discovery set and two independent validation cohorts.
The matrix metalloproteinase 3 (MMP3) in the plasma correlates with BOS, a serious complication after allo-HSCT. An area under the receiver-operating characteristic (ROC) curve for MMP3 indicated a value of 0.77 (78).
Another biomarker that may be linked to cGvHD or to the maintenance of remission is the presence of H-Y specific antibodies that relate to a minor antigen mismatch. These antibodies appear approximately 4–12 months after HSCT predominantly in male patients who were transplanted with female donor grafts. In the presence of alloantibodies, the cumulative incidence of cGvHD reached 89% at 5 years after HSCT compared with only 31% in the absence of H-Y antibodies. However, the cumulative incidence of relapse reached 48% at 5 years in the absence of H-Y antibodies compared with 0% in the presence of H-Y alloantibodies. Therefore, the authors concluded that antibody responses to H-Y antigens were associated with maintenance of disease remission in gender-mismatched HSCT. In a follow-up research project, the same group demonstrated that H-Y antigen-binding B-cells developed in male recipients of female hematopoietic cells that were associated with cGvHD (79). Of note, B-cells specific for the dominant H-Y epitope, DEAD box protein (DBY-2) appeared in significantly higher frequency in the circulation 6 months after HSCT in individuals who developed cGvHD later and thus, may predict cGvHD (79, 80). However, this single center study requires validation in larger prospective clinical studies to allow firm conclusions.
Biofluid miRNAs As Biomarkers for Graft-versus-Host Disease
Within the last decade, circulating short single-stranded miRNA has been identified in human plasma, serum, and also in urine (81, 82). Interestingly, these miRNAs were found to be resistant to RNase, boiling, changes in pH, extended storage, and freeze-thaw cycles (83). Functionally, miRNA molecules are considered as one of the major groups of translational regulators with the ability to regulate differentiation of blood cells and immune functions (84, 85). The stability of miRNA in the circulation can be attributed to three major RNase protection mechanisms: miRNAs can be bound to protective proteins, such as nucleophosmin 1 (NPM1) and/or Agonaute 2 (Ago2) (86–88), miRNA can form complexes with lipid or lipoproteins including high-density lipoprotein and low-density lipoprotein, and miRNA can be encapsulated in extracellular lipid vesicles, such as exosomes (89, 90). The complex of miRNA and these proteins and/or vesicles allows the selective export of miRNAs into the circulation and the protection of them within the extracellular environment (86). Circulating miRNAs have been shown to act as robust biomarkers for a various diseases, including autoimmune conditions, such as rheumatoid arthritis and systemic lupus erythematosus and tumors (91). A number of studies have also assessed the association between circulating miRNAs and the development of GvHD, with promising results (92–96).
Circulating miRNAs are robust and can be detected in biofluids by minimal invasive methods using relatively simple and accurate technologies. Furthermore, circulating miRNAs may offer advantages over protein-based biomarkers as they are lower in complexity, conserved among clinically relevant species, expressed specifically in different tissues or biological stages and easily measured using common laboratory techniques (97).
Although miRNA studies in relation to GvHD are still in their infancy, miRNA-155 was the first miRNA to be associated with aGvHD (98). miRNA-155 expression is upregulated in T cells of mice with severe aGvHD after allo-HSCT and a reduction of miRNA-155 results in a decreased severity of aGvHD and a prolonged survival of mice (99). miRNA-155 is encoded within the B cell integration cluster and is important for the regulation of acute inflammation and innate immunity (98). Expression of miRNA-155 can be activated by inflammatory mediators, such as IFN-α/γ and TNF-α (100), and Ceppi et al proposed that it functions as part of the negative feedback loop controlling the secretion of inflammatory cytokines by LPS-induced DC activation (101). Thus, miRNA-155 is pivotal in fine-tuning of the immune response (Figure 1). Physiologically, miRNA-155 is upregulated during T cell differentiation, plays a role in CD4+ and CD8+ T cell-mediated immunity and promotes the development of T cells, including Th17 and Treg subsets. Mice with a germ-line depletion of miRNA-155 have a normal lymphocyte development but defective T and B cell immunity (102, 103). Within Tregs, miRNA-155 targets forkhead box P3 (FoxP3), which regulates in vivo Treg survival (104). miRNA-155 also directly targets the IL-2 signaling protein suppressor of cytokine signaling 1 (SOCS1), whereby miRNA-155 deficiency in Tregs results in increased SOCS1 expression (96). This, in turn, leads to impaired activation of signal transducer and activator of transcription factor 5 (STAT5) phosphorylation and IL-2 receptor signaling (105). miRNA-155 is also a key regulator of CD8+ T cell responses via SOCS1, where a deficiency of miRNA-155 results in defective STAT5-mediated cytokine signaling (106). In CD4+ T cells, miRNA-155 targets phosphatidylinositol 3,4,5-triphosphate 5-phosphatase 1 (SHIP1) for downregulation, which normally functions to suppress Th1 responses and T cells by modulating IFN-γ production (102, 107).
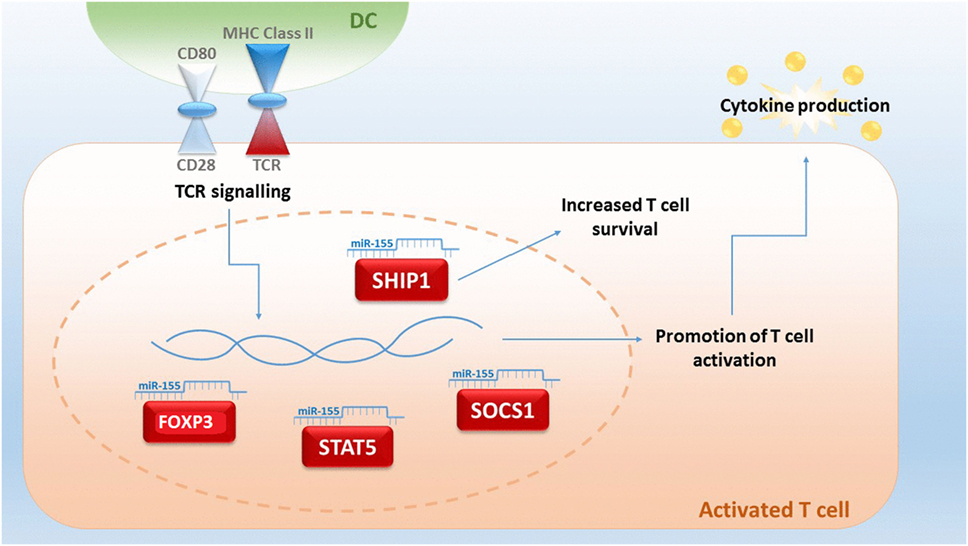
Figure 1. An upregulated expression of microRNA (miRNA)-155 in T-cells is indicative for lethal acute graft-versus-host disease (aGvHD) in mice. miR-155 plays a central role in fine-tuning the immune response, and expression can be triggered by T cell receptor (TCR) signaling. miRNA-155 regulates FOXP3, which is important for the survival of T regulatory cells and influences the targets signal transducer and activator of transcription factor 5 (STAT5) and suppressor of cytokine signaling 1 (SOCS1). Targeting of 3,4,5-triphosphate 5-phosphatase 1 (SHIP1) by miRNA-155 results in an increased T-cell survival by an upregulated production of IFN-γ (93, 104–108).
In a seminal study in 2012, Ranganathan et al showed that miRNA-155 is upregulated in T-cells from mice developing GvHD following HSCT (98). Moreover, the use of miRNA-155 inhibitors decreased disease severity and prolonging survival (98). In addition, the group found increased miRNA-155 expression in small and large bowel biopsies compared to normal bowel tissue in patients with GI aGvHD (98). However, these studies focused on miRNA expression within tissue biopsies, and more recently, several groups have sought to exploit the biomarker potential of circulating miRNAs in the context of GvHD. Although the presence of miRNAs has been established in a number of biological fluids (103), studies to date have concentrated on serum and plasma fractions of the blood.
A comprehensive report published by Xiao et al (92) in 2013 used a high-throughput qRT-PCR-based array to profile the expression of 345 miRNAs in the plasma of patients with aGvHD compared to those with no GvHD. They employed a discovery and training cohort to identify a final signature of four miRNAs (miRNA-423, miRNA-199-3p, miRNA-93*, and miRNA-377). These four miRNAs were able to predict aGvHD 6 weeks post-HSCT, prior to the onset of symptoms. The model was significant in ROC analysis (AUC = 0.76, p < 0.001) and was also associated with disease severity and poor overall survival (92). The miRNAs were shown to be specific for aGvHD, as they were not detected in the plasma of lung transplant or non-transplant sepsis patients. This study highlighted the potential of biofluid miRNAs as independent markers for prediction, prognosis, and diagnosis of GvHD.
Although miRNA-155 has been previously associated with the development of GvHD, it was not included in the final model proposed by Xiao et al. While expression was significantly upregulated in the plasma of aGvHD patients, levels were the lowest among the miRNA candidates identified (92). However, a recent study by Xie et al focusing on serum showed that miRNA-155 was significantly upregulated in aGvHD patients (p = 0.003) and also correlated with disease severity (p < 0.001) (93). Furthermore, expression was elevated in cGvHD patients compared to those who did not develop the disease (p = 0.005); however, levels were not sufficient to distinguish between aGvHD and cGvHD (p = 0.96) (93). Although this study showed promise for the inclusion of miRNA-155 as a GvHD biofluid biomarker, the study was restricted to one cohort of Han Chinese population (n = 64). Thus, the results need to be validated in independent prospective cohorts.
A small study by Sang et al confirmed elevated miRNA-155 plasma levels in patients with aGvHD, and additionally reported upregulation of miRNA-92b, while miRNA-150 and miRNA-181 were significantly downregulated (95). Interestingly, levels of the miRNAs were also altered prior to disease onset, highlighting their biomarker potential for predicting incidence (95). However, the difference in miRNA-181 was most pronounced, with no variation observed in control patients compared to reduced levels of expression in 19/22 (86%) patients prior to disease development (95). The group also demonstrated miRNA-181 to act as an effective predictor of aGvHD in a murine allo-HSCT model (95); however, larger cohorts are required in order to validate these findings.
More recently, Wang et al performed a study focusing on miRNA-586 expression in the plasma of HSCT patients (96). miRNA-586 was significantly upregulated in patients who developed aGvHD compared to no aGvHD as early as 7 days post-HSCT; however, expression was influenced by infection that reduced the significance of the association (96). As infections are common in early post-HSCT and can make distinguishing a diagnosis of aGvHD more challenging, it is important to validate the findings in larger studies as well as to elucidate the biological role of miRNA-586 in infections. Thus, although these results shown promise, miRNA-586 may play a greater role as a clinical biomarker to differentiate aGvHD from infectious complications.
In a separate study, Gimondi et al focused on lymphoma patients receiving MUD allo-transplants to profile the plasma using qRT-PCR for global miRNA expression (94). Assessing samples collected 28 days post-HSCT, 113 miRNAs were detected in all samples and of these, 27 could collectively discriminate between aGvHD versus no aGvHD and miRNA-194 and miRNA-518f were significantly upregulated in the patients who later developed the disease (94). The authors did not detect differential expression of miRNAs identified by Xiao et al (92), and there was no validation cohort included in the investigation. Thus, although results showed potential for the identification of aGvHD biomarkers, additional confirmatory studies are required.
Although circulating miRNAs as biomarkers for GvHD show great promise, these studies are still in their infancy and few overlapping targets between reports have been identified. A summary of presently known miRNAs for a/cGvHD have been presented in Table 1. Much work is needed to validate the findings in independent cohorts that reflect the heterogeneity in conditioning and prophylaxis regimens employed by different clinical centers. Moreover, collaboration between groups will allow for the standardization of protocols and technologies, which may greatly influence the reproducibility of findings and is a likely explanation for the lack of concordance in results to date. For blood studies, the choice of serum or plasma needs to be considered, with slightly more biomarker studies currently focusing on serum over plasma, while few whole blood or PBMC studies have been performed (108). Indeed, it has been proposed that profiles from isolated PBMC may yield information on the immune status of the disease, while the serum and plasma levels reflect the disease-dependent secretion and expression of miRNAs (108). The effect of hemolysis and platelet contamination as well as the choice of anti-coagulant should also be considered (109–111). Selection of the miRNA detection platform and normalization as well as the RNA isolation methodology employed has also been shown to affect results. Indeed, analytical variables have a huge potential to bias results, and this is particularly dependent on the normalization methods used (112). Some level of correction may be achieved by spiking in synthetic miRNAs (113); however, this approach can correct for technical variation such as the efficiency of RNA isolation and reverse transcription, but it will not account for intrinsic biological variation (114). Thus, the choice of normalization controls and/or global normalization becomes integral to the data analysis strategy. With regard to the miRNA detection platform, although qRT-PCR is a commonly used approach, variations on the fluorescent molecules including TaqMan and Sybr-Green, as well as the development of newer techniques including NanoString and Next-generation sequencing, have increased heterogeneity. This is important, as variation in results have been observed depending on the analysis platform, partly attributed to the difficulty in distinguishing small molecules at low abundance with high sequence homology (115).
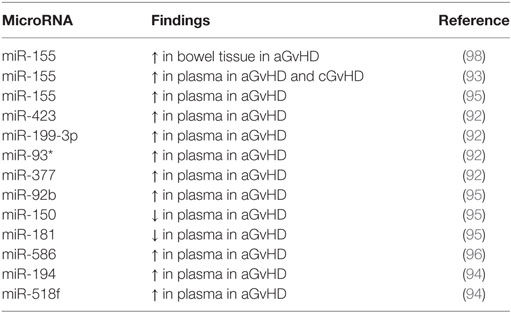
Table 1. Role of different microRNAs (miRNAs) in acute and chronic graft-versus-host disease (a/cGvHD).
Despite these considerations, it is expected that over the next few years, as the number of circulating miRNA biomarker studies increases, specific miRNA patterns for GvHD will be proposed and validated in the clinic. Although studies to date have focused on fractions of the blood, the potential for discovery of miRNA signatures in other fluids, including urine, is attractive. Indeed, initial data promise that miRNA signatures may be identified to predict the incidence of GvHD prior to onset, the severity of disease, to distinguish GvHD from other complications, and even to differentiate between aGvHD and cGvHD, particularly in relation to late onset aGvHD. Given the heterogeneity of transplantation protocols employed throughout different clinics, miRNA models appropriate to different transplant protocols may also be possible. Further investigation of validated miRNA signatures will allow the impact of miRNA dysregulation on the pathogenesis of GvHD to be studied, and conclusively, prospective investigations assessing the outcome of treatments selected by miRNA status will confirm their prognostic strength. Ultimately, the aim will be to diagnose GvHD and outcome before clinical symptoms manifest, allowing earlier introduction of therapy, tailored treatments, and reduced mortality and morbidity outcomes.
Urine Proteomics Based on Capillary Electrophoresis and Mass Spectrometry (CE-MS)
Proteomics for Prediction of aGvHD after HSCT
Proteomics of tissues or body fluids has gained significant importance over the last 10 years.
For example, urinary proteome-based classifiers, developed for aGvHD prediction, were first described in 2004 (116) and adapted in 2007 (117) and additional prospective evaluation was provided in 2014 (118). The urinary proteome profiling for aGvHD is done on prospectively collected urine samples from patients undergoing HSCT. The urine is collected prior to HSCT and conditioning, weekly after HSCT until day +35 and afterward biweekly until day +100. Urine can be frozen at −20°C until preparation of the sample for analyses using CE-MS (119–122). An extension to include patients with cGvHD was carried out in 2008; thus, samples were collected bimonthly after day +100 or at diagnosis of cGvHD (119, 123).
Acute and chronic GvHD are currently diagnosed on clinical parameters defined in NIH Consensus (2, 124, 125) and German Consensus Conferences (126–128). Prediction of aGvHD using the investigator-independent, unbiased proteomic classifier aGvHD_MS17 was done on more than 500 patients in at least 4 centers in Germany and UK and resulted in stable prediction of aGvHD about 7–10 days prior to clinical diagnosis or biopsy positivity (118, 119, 123, 129). The classifier aGvHD_MS17 consists of 17 peptides, 10 of which have been sequenced to date (117–119). The identified peptides are various fragments of collagen (indicating disturbances in metabolism of collagen and/or early organ damage) and fragments of fibronectin, beta-2-microglobulin (β2M), and CD99, an activation marker of T-cells. Multivariate regression analysis showed that aGvHD_MS17 positivity was the highest predictive parameter for aGvHD development (p < 0.0001, Figure 2).
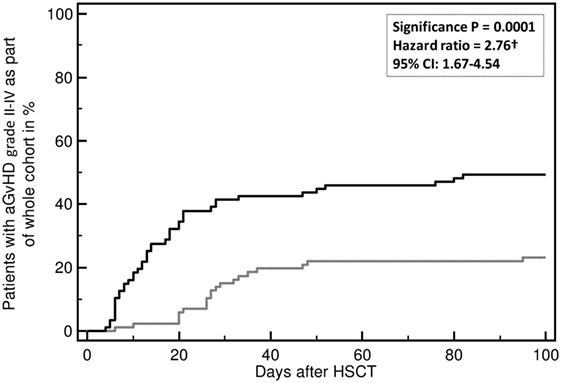
Figure 2. Incidence of developing an acute graft-versus-host disease (aGvHD) grades II–IV. Incidence of aGvHD grades II–IV is predicted by aGvHD_MS17 positivity (p = 0.0001) when compared to patients without aGvHD_MS17 pattern positivity. Shown is the percentage (%) of patients with aGvHD II–IV with (black line) or without aGvHD_MS17 positivity over time [days after allogeneic hematopoietic stem cell transplantation (allo-HSCT)] (gray line).
Future Perspectives
Numerous candidates for biomarkers are currently available, which include plasma and serum markers, cell-based markers such as B-cell subpopulations, and T follicular helper cells as regulators of B-cell immunity in determining cGvHD in patients (130), miRNAs, different stress proteins, or proteomic approaches using plasma, serum, or urine. Proteomic mass spectrometry analysis coupled with computational biology approaches will further lead to the identification of novel biomarkers for GvHD such as BOS in the lung (131). Recently, the diversity of the intestinal microbiome as analyzed by 16 s rRNA sequencing of stool microbiota (131–134) and the assessment of bacterial metabolites (135) very early after transplantation has identified patients at high risk of lethal complications. On the other hand, biomarkers can also be used to tailor immunosuppressive therapies in transplanted patients and thus might predict the severity of GvHD.
The main challenge remains to integrate all these candidates and to validate them as biomarkers that are valid in multiple centers independent of the center’s strategy of prophylaxis, and consortia on prospective testing of these markers together with clinical data collection as shown for the MAGIC consortium are urgently needed. The expectations have to be defined clearly, do we want to predict outcome in general or treatment response for aGvHD or cGvHD or do we want to make a diagnosis as precise as possible (e.g., for GI GvHD, for cGvHD) and the current standards and endpoints for each of these parameters have to be defined separately (e.g., d28 response for aGvHD, 2- or 3-month response for cGvHD, 6 months NRM for aGvHD, composite endpoints of GvHD-free and relapse-free survivals) (136).
New candidates have to be integrated in this process step by step, and bio-mathematical approaches are needed to define those markers that give true additive information. In addition, integration of older markers has not been performed (e.g., calprotectin in stool samples for GI GvHD (137), the poor man’s biomarker albumin for GI GvHD) (138, 139), and more importantly, testing biomarkers against clinical predictors or markers of outcome is needed. For aGvHD, factors indicating poor prognosis are the extent of organ involvement and number of involved organs. For cGvHD, the type of onset and simultaneous occurrence of thrombocytopenia should be addressed. It might well be that some of the markers behave as highly sensitive and specific markers due to the correlation with extent and type of involved organ and the specific pathophysiology, which might be reflected by the correlation with clinical parameters (140).
Author Contributions
MJ, MS, PM, and JO equally contributed to the manuscript. MJ, MS, PM, JO, and RC wrote the manuscript and prepared the figures. AD, HG, EH, EW, and GM outlined and proof-read the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors thank Katie Gray and Anett Lange for editorial support of the manuscript.
Funding
The work of the authors was supported by the European Union grant FP7-PEOPLE-2012-ITN-315963 (CELLEUROPE), by the DFG SFB-824/2, BMBF (01GU0823, 02NUK038A), and DKTK-ROG. This work was supported by the German Research Foundation (DFG) and the Technische Universität München (TUM) within the funding program Open Access Publishing.
References
1. Socie G, Stone JV, Wingard JR, Weisdorf D, Henslee-Downey PJ, Bredeson C, et al. Long-term survival and late deaths after allogeneic bone marrow transplantation. Late Effects Working Committee of the International Bone Marrow Transplant Registry. N Engl J Med (1999) 341:14–21. doi:10.1056/NEJM199907013410103
2. Lee SJ, Vogelsang G, Flowers ME. Chronic graft-versus-host disease. Biol Blood Marrow Transplant (2003) 9:215–33. doi:10.1053/bbmt.2003.50026
3. Filipovich AH, Weisdorf D, Pavletic S, Socie G, Wingard JR, Lee SJ, et al. National Institutes of Health consensus development project on criteria for clinical trials in chronic graft-versus-host disease: I. Diagnosis and staging working group report. Biol Blood Marrow Transplant (2005) 11:945–56. doi:10.1016/j.bbmt.2005.09.004
4. Jagasia MH, Greinix HT, Arora M, Williams KM, Wolff D, Cowen EW, et al. National Institutes of Health consensus development project on criteria for clinical trials in chronic graft-versus-host disease: I. The 2014 diagnosis and staging working Group report. Biol Blood Marrow Transplant (2015) 21(389–401):e1. doi:10.1016/j.bbmt.2014.12.001
5. Paczesny S, Hakim FT, Pidala J, Cooke KR, Lathrop J, Griffith LM, et al. National Institutes of Health consensus development project on criteria for clinical trials in chronic graft-versus-host disease: III. The 2014 Biomarker Working Group Report. Biol Blood Marrow Transplant (2015) 21:780–92. doi:10.1016/j.bbmt.2015.01.003
6. Ferrara JL, Levine JE, Reddy P, Holler E. Graft-versus-host disease. Lancet (2009) 373:1550–61. doi:10.1016/S0140-6736(09)60237-3
7. Fuchs E. Haploidentical transplantation for hematologic malignancies: where do we stand? Hematolopgy Am Soc Hematol Educ Program (2012) 23(15):230–6. doi:10.1182/asheducation-2012.1.230
8. Greinix HT, Pohlreich D, Kouba M, Kormoczi U, Lohmann I, Feldmann K, et al. Elevated numbers of immature/transitional CD21- B lymphocytes and deficiency of memory CD27+ B cells identify patients with active chronic graft-versus-host disease. Biol Blood Marrow Transplant (2008) 14:208–19. doi:10.1016/j.bbmt.2007.10.009
9. Sarantopoulos S, Ritz J. Aberrant B-cell homeostasis in chronic GVHD. Blood (2015) 125:1703–7. doi:10.1182/blood-2014-12-567834
10. Socie G. Chronic GVHD: B cells come of age. Blood (2011) 117:2086–7. doi:10.1182/blood-2010-12-322297
11. Kuzmina Z, Greinix HT, Weigl R, Kormoczi U, Rottal A, Frantal S, et al. Significant differences in B-cell subpopulations characterize patients with chronic graft-versus-host disease-associated dysgammaglobulinemia. Blood (2011) 117:2265–74. doi:10.1182/blood-2010-07-295766
12. Sarantopoulos S, Stevenson KE, Kim HT, Cutler CS, Bhuiya NS, Schowalter M, et al. Altered B-cell homeostasis and excess BAFF in human chronic graft-versus-host disease. Blood (2009) 113:3865–74. doi:10.1182/blood-2008-09-177840
13. Storek J, Witherspoon RP, Webb D, Storb R. Lack of B cell precursors in marrow transplant recipients with chronic graft-versus-host disease. Am J Hematol (1996) 52:82–9. doi:10.1002/(SICI)1096-8652(199606)52:2<82::AID-AJH3>3.0.CO;2-1
14. Rolink AG, Melchers F. BAFFled B cells survive and thrive: roles of BAFF in B-cell development. Curr Opin Immunol (2002) 14:266–75. doi:10.1016/S0952-7915(02)00332-1
15. Sarantopoulos S, Stevenson KE, Kim HT, Bhuiya NS, Cutler CS, Soiffer RJ, et al. High levels of B-cell activating factor in patients with active chronic graft-versus-host disease. Clin Cancer Res (2007) 13:6107–14. doi:10.1158/1078-0432.CCR-07-1290
16. Kuzmina Z, Krenn K, Petkov V, Kormoczi U, Weigl R, Rottal A, et al. CD19(+)CD21(low) B cells and patients at risk for NIH-defined chronic graft-versus-host disease with bronchiolitis obliterans syndrome. Blood (2013) 121:1886–95. doi:10.1182/blood-2012-06-435008
17. Allen JL, Fore MS, Wooten J, Roehrs PA, Bhuiya NS, Hoffert T, et al. B cells from patients with chronic GVHD are activated and primed for survival via BAFF-mediated pathways. Blood (2012) 120:2529–36. doi:10.1182/blood-2012-06-438911
18. Greinix HT, Kuzmina Z, Weigl R, Kormoczi U, Rottal A, Wolff D, et al. CD19+CD21low B cells and CD4+CD45RA+CD31+ T cells correlate with first diagnosis of chronic graft-versus-host disease. Biol Blood Marrow Transplant (2015) 21:250–8. doi:10.1016/j.bbmt.2014.11.010
19. Kuzmina Z, Greinix HT, Knobler R, Worel N, Kouba M, Weigl R, et al. Proportions of immature CD19+CD21- B lymphocytes predict the response to extracorporeal photopheresis in patients with chronic graft-versus-host disease. Blood (2009) 114:744–6. doi:10.1182/blood-2009-05-221028
20. Rakhmanov M, Keller B, Gutenberger S, Foerster C, Hoenig M, Driessen G, et al. Circulating CD21low B cells in common variable immunodeficiency resemble tissue homing, innate-like B cells. Proc Natl Acad Sci U S A (2009) 106:13451–6. doi:10.1073/pnas.0901984106
21. Suryani S, Fulcher DA, Santner-Nanan B, Nanan R, Wong M, Shaw PJ, et al. Differential expression of CD21 identifies developmentally and functionally distinct subsets of human transitional B cells. Blood (2010) 115:519–29. doi:10.1182/blood-2009-07-234799
22. de Masson A, Bouaziz JD, Le Buanec H, Robin M, O’Meara A, Parquet N, et al. CD24(hi)CD27(+) and plasmablast-like regulatory B cells in human chronic graft-versus-host disease. Blood (2015) 125:1830–9. doi:10.1182/blood-2014-09-599159
23. Cambier JC, Gauld SB, Merrell KT, Vilen BJ. B-cell anergy: from transgenic models to naturally occurring anergic B cells? Nat Rev Immunol (2007) 7:633–43. doi:10.1038/nri2133
24. de Masson A, Socie G, Bagot M, Bensussan A, Bouaziz JD. Deficient regulatory B cells in human chronic graft-versus-host disease. Oncoimmunology (2015) 4:e1016707. doi:10.1080/2162402X.2015.1016707
25. Khoder A, Sarvaria A, Alsuliman A, Chew C, Sekine T, Cooper N, et al. Regulatory B cells are enriched within the IgM memory and transitional subsets in healthy donors but are deficient in chronic GVHD. Blood (2014) 124:2034–45. doi:10.1182/blood-2014-04-571125
26. Koreth J, Matsuoka K, Kim HT, McDonough SM, Bindra B, Alyea EP III, et al. IL-2 and regulatory T cells in graft-versus-host disease. N Engl J Med (2011) 365(22):2055–66. doi:10.1056/NEJMoa1108188
27. Miura Y, Thoburn CJ, Bright EC, Phleps ML, Shin T, Matsui WH, et al. Association of Foxp3 regulatory gene expression with graft-versus-hist disease. Blood (2004) 104(7):2187–93. doi:10.1182/blood-2004-03-1040
28. Zorn E, Kim HT, Lee SJ, Floyd BH, Litsa D, Arumugarajah S, et al. Reduced frequency of FOXP3+CD4+CD25+ regulatory T cells in patients with chronic graft-versus-host disease. Blood (2005) 106(8):2903–11. doi:10.1182/blood-2005-03-1257
29. Matsuoka KI, Kim HT, McDonough S, Bascug G, Warshauer B, Koreth J, et al. Altered regulatory T cell homeostasis in patients with CD4+ lymphopenia following allogeneic hematopoietic stem cell transplantation. J Clin Invest (2010) 120(5):1479–93. doi:10.1172/JCI41072
30. D`Asaro M, Dieli F, Caccamo N, Mussa M, Porretto F, Salerno A. Increase of CCR7-CD45RA+CD8 T cells (T(EMRA)) in chronic graft-versus-host diesease. Leukemia (2006) 20(3):545–7. doi:10.1038/sj.leu.2404079
31. Yamashita K, Choi U, Woltz PC, Foster SF, Sneller MC, Hakim FT, et al. Severe chronic graft-versus-host disease is characterized by a preponderance of CD4(+) effector memory cells relative to central memory cells. Blood (2004) 103(10):3986–8. doi:10.1182/blood-2003-09-3286
32. Gehrmann M, Schmetzer H, Eissner G, Haferlach T, Hiddemann W, Multhoff G. Membrane-bound heat shock protein 70 (Hsp70) in acute myeloid leukemia: a tumor specific recognition structure for the cytolytic activity of autologous NK cells. Haematologica (2003) 88:474–6.
33. Guzhova IV, Shevtsov MA, Abkin SV, Pankratova KM, Margulis BA. Intracellular and extracellular Hsp70 chaperone as a target for cancer therapy. Int J Hyperthermia (2013) 29:399–408. doi:10.3109/02656736.2013.807439
34. Chen T, Cao X. Stress for maintaining memory: HSP70 as a mobile messenger for innate and adaptive immunity. Eur J Immunol (2010) 40:1541–4. doi:10.1002/eji.201040616
35. Tsan MF, Gao B. Heat shock proteins and immune system. J Leukoc Biol (2009) 85:905–10. doi:10.1189/jlb.0109005
36. Finka A, Sharma SK, Goloubinoff P. Multi-layered molecular mechanisms of polypeptide holding, unfolding and disaggregation by HSP70/HSP110 chaperones. Front Mol Biosci (2015) 2:29. doi:10.3389/fmolb.2015.00029
37. Clerico EM, Tilitsky JM, Meng W, Gierasch LM. How hsp70 molecular machines interact with their substrates to mediate diverse physiological functions. J Mol Biol (2015) 427:1575–88. doi:10.1016/j.jmb.2015.02.004
38. Duncan EJ, Cheetham ME, Chapple JP, van der Spuy J. The role of HSP70 and its co-chaperones in protein misfolding, aggregation and disease. Subcell Biochem (2015) 78:243–73. doi:10.1007/978-3-319-11731-7_12
39. Gross C, Holler E, Stangl S, Dickinson A, Pockley AG, Asea AA, et al. An Hsp70 peptide initiates NK cell killing of leukemic blasts after stem cell transplantation. Leuk Res (2008) 32:527–34. doi:10.1016/j.leukres.2007.03.027
40. Gehrmann M, Liebisch G, Schmitz G, Anderson R, Steinem C, De MA, et al. Tumor-specific Hsp70 plasma membrane localization is enabled by the glycosphingolipid Gb3. PLoS One (2008) 3:e1925. doi:10.1371/journal.pone.0001925
41. Gastpar R, Gehrmann M, Bausero MA, Asea A, Gross C, Schroeder JA, et al. Heat shock protein 70 surface-positive tumor exosomes stimulate migratory and cytolytic activity of natural killer cells. Cancer Res (2005) 65:5238–47. doi:10.1158/0008-5472.CAN-04-3804
42. Breuninger S, Erl J, Knape C, Gunther S, Regel I, Rodel F, et al. Quantitative analysis of liposomal heat shock protein 70 (Hsp70) in the blood of tumor patients using an novel lipHsp70 ELISA. Clin Cell Immunol (2014) 5:2–10. doi:10.4172/2155-9899.1000264
43. Asea A. Stress proteins and initiation of immune response: chaperokine activity of Hsp72. Exerc Immunol Rev (2005) 11:34–45.
44. Shevtsov M, Multhoff G. Heat shock protein-peptide and HSP-based immunotherapies for the treatment of Cancer. Front Immunol (2016) 7:171. doi:10.3389/fimmu.2016.00171
45. Novota P, Sviland L, Zinocker S, Stocki P, Balavarca Y, Bickeboller H, et al. Correlation of Hsp70-1 and Hsp70-2 gene expression with the degree of graft-versus-host reaction in a rat skin explant model. Transplantation (2008) 85:1809–16. doi:10.1097/TP.0b013e31817753f7
46. Kim I, Kim JH, Rhee JY, Kim JW, Cho HJ, Cho EY, et al. Patient HSP70-hom TG haplotype is associated with decreased transplant-related mortality and improved survival after sibling HLA-matched hematopoietic stem cell transplantation. Clin Transplant (2010) 24:459–66. doi:10.1111/j.1399-0012.2009.01094.x
47. Bogunia-Kubik K, Wysoczanska B, Lange A. Non-HLA gene polymorphisms and the outcome of allogeneic hematopoietic stem cell transplantation. Curr Stem Cell Res Ther (2006) 1:239–53. doi:10.2174/157488806776956850
48. Bogunia-Kubik K, Lange A. HSP70-hom gene polymorphism in allogeneic hematopoietic stem-cell transplant recipients correlates with the development of acute graft-versus-host disease. Transplantation (2005) 79:815–20. doi:10.1097/01.TP.0000153157.97736.2C
49. Goral J, Mathews HL, Clancy J Jr. Expression of 70-kDa heat-shock protein during acute graft-versus-host disease. Clin Immunol Immunopathol (1998) 86:252–8. doi:10.1006/clin.1997.4473
50. Goral J, Mathews HL, Clancy J Jr. Antibodies specific for the 70-kDa heat-shock protein parallel the development of acute graft-versus-host disease in (DA x LEW)F1 rats. Clin Immunol Immunopathol (1995) 75:147–53. doi:10.1006/clin.1995.1064
51. Zlacka D, Sedlacek P, Prucha M, Hromadnikova I. Antibodies to 60, 65 and 70 kDa heat shock proteins in pediatric allogeneic stem cell transplant recipients. Pediatr Transplant (2006) 10:794–804. doi:10.1111/j.1399-3046.2006.00566.x
52. Goral J, Shenoy S, Mohanakumar T, Clancy J Jr. Antibodies to 70 kD and 90 kD heat shock proteins are associated with graft-versus-host disease in peripheral blood stem cell transplant recipients. Clin Exp Immunol (2002) 127:553–9. doi:10.1046/j.1365-2249.2002.01770.x
53. Panjwani NN, Popova L, Srivastava PK. Heat shock proteins gp96 and hsp70 activate the release of nitric oxide by APCs. J Immunol (2002) 168:2997–3003. doi:10.4049/jimmunol.168.6.2997
54. Wang Y, Kelly CG, Singh M, McGowan EG, Carrara AS, Bergmeier LA, et al. Stimulation of Th1-polarizing cytokines, C-C chemokines, maturation of dendritic cells, and adjuvant function by the peptide binding fragment of heat shock protein 70. J Immunol (2002) 169:2422–9. doi:10.4049/jimmunol.169.5.2422
55. Mansilla MJ, Costa C, Eixarch H, Tepavcevic V, Castillo M, Martin R, et al. Hsp70 regulates immune response in experimental autoimmune encephalomyelitis. PLoS One (2014) 9:e105737. doi:10.1371/journal.pone.0105737
56. Mosenson JA, Eby JM, Hernandez C, Le Poole IC. A central role for inducible heat-shock protein 70 in autoimmune vitiligo. Exp Dermatol (2013) 22:566–9. doi:10.1111/exd.12183
57. Alam MU, Harken JA, Knorn AM, Elford AR, Wigmore K, Ohashi PS, et al. Transgenic expression of Hsc70 in pancreatic islets enhances autoimmune diabetes in response to beta cell damage. J Immunol (2009) 183:5728–37. doi:10.4049/jimmunol.0901288
58. Pockley AG. Heat Shock Proteins in Health and Disease: Therapeutic Targets or Therapeutic Agents? Expert Reviews in Molecular Medicine. Cambridge, UK: Cambridge University Press (2001). p. 1–21.
59. Williams JH, Ireland HE. Sensing danger – Hsp72 and HMGB1 as candidate signals. J Leukoc Biol (2008) 83:489–92. doi:10.1189/jlb.0607356
60. Calderwood SK, Theriault JR, Gong J. Message in a bottle: role of the 70-kDa heat shock protein family in anti-tumor immunity. Eur J Immunol (2005) 35:2518–27. doi:10.1002/eji.200535002
61. O’Neill S, Ingman TG, Wigmore SJ, Harrison EM, Bellamy CO. Differential expression of heat shock proteins in healthy and diseased human renal allografts. Ann Transplant (2013) 18:550–7. doi:10.12659/AOT.889599
63. Lee YS, Sakurai M, Imai Y, Horinouchi T, Goto T, Yamada A. Induction of heat shock proteins in a rat composite tissue allotransplantation model of acute rejection. Plast Reconstr Surg (2006) 118:29–40. doi:10.1097/01.prs.0000220482.94982.01
64. Oh KH, Kim JY, Kim D, Lee EM, Oh HY, Seo JS, et al. Targeted gene disruption of the heat shock protein 72 gene (hsp70.1) in the donor tissue is associated with a prolonged rejection-free survival in the murine skin allograft model. Transpl Immunol (2004) 13:273–81. doi:10.1016/j.trim.2004.08.003
65. Goral J, Mathews HL, Nadler SG, Clancy J. Reduced levels of Hsp70 result in a therapeutic effect of 15-deoxyspergualin on acute graft-versus-host disease in (DA x LEW)F1 rats. Immunobiology (2000) 202:254–66. doi:10.1016/S0171-2985(00)80032-7
66. Harris AC, Ferrara JL, Braun TM, Holler E, Teshima T, Levine JE, et al. Plasma biomarkers of lower gastrointestinal and liver acute GVHD. Blood (2012) 119:2960–3. doi:10.1182/blood-2011-10-387357
67. Levine JE, Logan BR, Wu J, Alousi AM, Bolanos-Meade J, Ferrara JL, et al. Acute graft-versus-host disease biomarkers measured during therapy can predict treatment outcomes: a Blood and Marrow Transplant Clinical Trials Network study. Blood (2012) 119:3854–60. doi:10.1182/blood-2012-01-403063
68. Ferrara JL, Harris AC, Greenson JK, Braun TM, Holler E, Teshima T, et al. Regenerating islet-derived 3-alpha is a biomarker of gastrointestinal graft-versus-host disease. Blood (2011) 118:6702–8. doi:10.1182/blood-2011-08-375006
69. Paczesny S, Krijanovski OI, Braun TM, Choi SW, Clouthier SG, Kuick R, et al. A biomarker panel for acute graft-versus-host disease. Blood (2009) 113:273–8. doi:10.1182/blood-2008-07-167098
70. Srinivasan R, Daniels J, Fusaro V, Lundqvist A, Killian JK, Geho D, et al. Accurate diagnosis of acute graft-versus-host disease using serum proteomic pattern analysis. Exp Hematol (2006) 34:796–801. doi:10.1016/j.exphem.2006.02.013
71. Imanguli MM, Atkinson JC, Harvey KE, Hoehn GT, Ryu OH, Wu T, et al. Changes in salivary proteome following allogeneic hematopoietic stem cell transplantation. Exp Hematol (2007) 35:184–92. doi:10.1016/j.exphem.2006.10.009
72. Levine JE, Braun TM, Harris AC, Holler E, Taylor A, Miller H, et al. A prognostic score for acute graft-versus-host disease based on biomarkers: a multicentre study. Lancet Haematol (2015) 2:e21–9. doi:10.1016/S2352-3026(14)00035-0
73. Harris AC, Young R, Devine S, Hogan WJ, Ayuk F, Bunworasate U, et al. International, multicenter standardization of acute graft-versus-host disease clinical data collection: a report from the Mount Sinai acute GVHD international consortium. Biol Blood Marrow Transplant (2016) 22:4–10. doi:10.1016/j.bbmt.2015.09.001
74. Kitko CL, Levine JE, Storer BE, Chai X, Fox DA, Braun TM, et al. Plasma CXCL9 elevations correlate with chronic GVHD diagnosis. Blood (2014) 123:786–93. doi:10.1182/blood-2013-08-520072
75. Yu J, Storer BE, Kushekhar K, Abu Zaid M, Zhang Q, Gafken PR, et al. Biomarker panel for chronic graft-versus-host disease. J Clin Oncol (2016) 34:2583–90. doi:10.1200/JCO.2015.65.9615
76. Ahmed SS, Wang XN, Norden J, Pearce K, El-Gezawy E, Atarod S, et al. Identification and validation of biomarkers associated with acute and chronic graft versus host disease. Bone Marrow Transplant (2016) 51:890. doi:10.1038/bmt.2016.125
77. Pidala J, Sarwal M, Roedder S, Lee SJ. Biologic markers of chronic GVHD. Bone Marrow Transplant (2014) 49:324–31. doi:10.1038/bmt.2013.97
78. Liu X, Yue Z, Yu J, Daguindau E, Kushekhar K, Zhang Q, et al. Proteomic characterization reveals that MMP-3 correlates with bronchiolitis obliterans syndrome following allogeneic hematopoietic cell and lung transplantation. Am J Transplant (2016) 16:2342–51. doi:10.1111/ajt.13750
79. Sahaf B, Yang Y, Arai S, Herzenberg LA, Herzenberg LA, Miklos DB. H-Y antigen-binding B cells develop in male recipients of female hematopoietic cells and associate with chronic graft vs host disease. Proc Natl Acad Sci U S A (2013) 110:3005–10. doi:10.1073/pnas.1222900110
80. Miklos DB, Kim HT, Miller KH, Guo L, Zorn E, Lee SJ, et al. Antibody responses to H-Y minor histocompatibility antigens correlate with chronic graft-versus-host disease and disease remission. Blood (2005) 105:2973–8. doi:10.1182/blood-2004-09-3660
81. Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K, et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res (2008) 18:997–1006. doi:10.1038/cr.2008.282
82. Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A (2008) 105:10513–8. doi:10.1073/pnas.0804549105
83. Serody J. GvHD and miR: good things in small packages. Blood (2015) 126(11):1265–7. doi:10.1182/blood-2015-07-657114
84. Ameres SL, Zamore PD. Diversifying microRNA sequence and function. Nat Rev Mol Cell Biol (2013) 14:475–88. doi:10.1038/nrm3611
85. Hunter MP, Ismail N, Zhang X, Aguda BD, Lee EJ, Yu L, et al. Detection of microRNA expression in human peripheral blood microvesicles. PLoS One (2008) 3:e3694. doi:10.1371/journal.pone.0003694
86. Etheridge A, Lee I, Hood L, Galas D, Wang K. Extracellular microRNA: a new source of biomarkers. Mutat Res (2011) 717:85–90. doi:10.1016/j.mrfmmm.2011.03.004
87. Vickers KC, Palmisano BT, Shoucri BM, Shamburek RD, Remaley AT. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat Cell Biol (2011) 13:423–33. doi:10.1038/ncb2210
88. Turchinovich A, Weiz L, Langheinz A, Burwinkel B. Characterization of extracellular circulating microRNA. Nucleic Acids Res (2011) 39:7223–33. doi:10.1093/nar/gkr254
89. Gallo A, Tandon M, Alevizos I, Illei GG. The majority of microRNAs detectable in serum and saliva is concentrated in exosomes. PLoS One (2012) 7:e30679. doi:10.1371/journal.pone.0030679
90. Taylor DD, Gercel-Taylor C. MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol Oncol (2008) 110:13–21. doi:10.1016/j.ygyno.2008.04.033
91. Pauley KM, Cha S, Chan EK. MicroRNA in autoimmunity and autoimmune diseases. J Autoimmun (2009) 32:189–94. doi:10.1016/j.jaut.2009.02.012
92. Xiao B, Wang Y, Li W, Baker M, Guo J, Corbet K, et al. Plasma microRNA signature as a noninvasive biomarker for acute graft-versus-host disease. Blood (2013) 122:3365–75. doi:10.1182/blood-2013-06-510586
93. Xie LN, Zhou F, Liu XM, Fang Y, Yu Z, Song NX, et al. Serum microRNA155 is increased in patients with acute graft-versus-host disease. Clin Transplant (2014) 28:314–23. doi:10.1111/ctr.12314
94. Gimondi S, Dugo M, Vendramin A, Bermema A, Biancon G, Cavane A, et al. Circulating miRNA panel for prediction of acute graft-versus-host disease in lymphoma patients undergoing matched unrelated hematopoietic stem cell transplantation. Exp Hematol (2016) 44(624–34):e1. doi:10.1016/j.exphem.2016.03.005
95. Sang W, Zhang C, Zhang D, Wang Y, Sun C, Niu M, et al. MicroRNA-181a, a potential diagnosis marker, alleviates acute graft versus host disease by regulating IFN-gamma production. Am J Hematol (2015) 90:998–1007. doi:10.1002/ajh.24136
96. Wang Y, Zhao X, Ye X, Luo H, Zhao T, Diao Y, et al. Plasma microRNA-586 is a new biomarker for acute graft-versus-host disease. Ann Hematol (2015) 94:1505–14. doi:10.1007/s00277-015-2414-z
97. Wang K, Yuan Y, Cho JH, McClarty S, Baxter D, Galas DJ. Comparing the microRNA spectrum between serum and plasma. PLoS One (2012) 7:e41561. doi:10.1371/journal.pone.0041561
98. Ranganathan P, Heaphy CE, Costinean S, Stauffer N, Na C, Hamadani M, et al. Regulation of acute graft-versus-host disease by microRNA-155. Blood (2012) 119:4786–97. doi:10.1182/blood-2011-10-387522
99. Rodriguez A, Vigorito E, Clare S, Warren MV, Couttet P, Soond DR, et al. Requirement of bic/microRNA-155 for normal immune function. Science (2007) 316:608–11. doi:10.1126/science.1139253
100. O’Connell RM, Taganov KD, Boldin MP, Cheng G, Baltimore D. MicroRNA-155 is induced during the macrophage inflammatory response. Proc Natl Acad Sci U S A (2007) 104:1604–9. doi:10.1073/pnas.0610731104
101. Ceppi M, Pereira PM, Dunand-Sauthier I, Barras E, Reith W, Santos MA, et al. MicroRNA-155 modulates the interleukin-1 signaling pathway in activated human monocyte-derived dendritic cells. Proc Natl Acad Sci U S A (2009) 106:2735–40. doi:10.1073/pnas.0811073106
102. Huffaker TB, Hu R, Runtsch MC, Bake E, Chen X, Zhao J, et al. Epistasis between microRNAs 155 and 146a during T cell-mediated antitumor immunity. Cell Rep (2012) 2:1697–709. doi:10.1016/j.celrep.2012.10.025
103. Weber JA, Baxter DH, Zhang S, Huang DY, Huang KH, Lee MJ, et al. The microRNA spectrum in 12 body fluids. Clin Chem (2010) 56:1733–41. doi:10.1373/clinchem.2010.147405
104. Kohlhaas S, Garden OA, Scudamore C, Turner M, Okkenhaug K, Vigorito E. Cutting edge: the Foxp3 target miR-155 contributes to the development of regulatory T cells. J Immunol (2009) 182:2578–82. doi:10.4049/jimmunol.0803162
105. Lu LF, Thai TH, Calado DP, Chaudhry A, Kubo M, Tanaka K, et al. Foxp3-dependent microRNA155 confers competitive fitness to regulatory T cells by targeting SOCS1 protein. Immunity (2009) 30:80–91. doi:10.1016/j.immuni.2008.11.010
106. Dudda JC, Salaun B, Ji Y, Palmer DC, Monnot GC, Merck E, et al. MicroRNA-155 is required for effector CD8+ T cell responses to virus infection and cancer. Immunity (2013) 38:742–53. doi:10.1016/j.immuni.2012.12.006
107. Tarasenko T, Kole HK, Chi AW, Mentink-Kane MM, Wynn TA, Bolland S. T cell-specific deletion of the inositol phosphatase SHIP reveals its role in regulating Th1/Th2 and cytotoxic responses. Proc Natl Acad Sci U S A (2007) 104:11382–7. doi:10.1073/pnas.0704853104
108. De Guire V, Robitaille R, Tetreault N, Guerin R, Menard C, Bambace N, et al. Circulating miRNAs as sensitive and specific biomarkers for the diagnosis and monitoring of human diseases: promises and challenges. Clin Biochem (2013) 46:846–60. doi:10.1016/j.clinbiochem.2013.03.015
109. Kim DJ, Linnstaedt S, Palma J, Park JC, Ntrivalas E, Kwak-Kim JY, et al. Plasma components affect accuracy of circulating cancer-related microRNA quantitation. J Mol Diagn (2012) 14:71–80. doi:10.1016/j.jmoldx.2011.09.002
110. McDonald JS, Milosevic D, Reddi HV, Grebe SK, Algeciras-Schimnich A. Analysis of circulating microRNA: preanalytical and analytical challenges. Clin Chem (2011) 57:833–40. doi:10.1373/clinchem.2010.157198
111. Pritchard CC, Kroh E, Wood B, Arroyo JD, Dougherty KJ, Miyaji MM, et al. Blood cell origin of circulating microRNAs: a cautionary note for cancer biomarker studies. Cancer Prev Res (Phila) (2012) 5:492–7. doi:10.1158/1940-6207.CAPR-11-0370
112. Leidner RS, Li L, Thompson CL. Dampening enthusiasm for circulating microRNA in breast cancer. PLoS One (2013) 8:e57841. doi:10.1371/journal.pone.0057841
113. Sapre N, Selth LA. Circulating MicroRNAs as biomarkers of prostate cancer: the state of play. Prostate Cancer (2013) 2013:539680. doi:10.1155/2013/539680
114. Huang X, Liang M, Dittmar R, Wang L. Extracellular microRNAs in urologic malignancies: chances and challenges. Int J Mol Sci (2013) 14:14785–99. doi:10.3390/ijms140714785
115. Mestdagh P, Hartmann N, Baeriswyl L, Andreasen D, Bernard N, Chen C, et al. Evaluation of quantitative miRNA expression platforms in the microRNA quality control (miRQC) study. Nat Methods (2014) 11:809–15. doi:10.1038/nmeth.3014
116. Kaiser T, Kamal H, Rank A, Kolb HJ, Holler E, Ganser A, et al. Proteomics applied to the clinical follow-up of patients after allogeneic hematopoietic stem cell transplantation. Blood (2004) 104:340–9. doi:10.1182/blood-2004-02-0518
117. Weissinger EM, Schiffer E, Hertenstein B, Ferrara JL, Holler E, Stadler M, et al. Proteomic patterns predict acute graft-versus-host disease after allogeneic hematopoietic stem cell transplantation. Blood (2007) 109:5511–9. doi:10.1182/blood-2007-01-069757
118. Weissinger EM, Metzger J, Dobbelstein C, Wolff D, Schleuning M, Kuzmina Z, et al. Proteomic peptide profiling for preemptive diagnosis of acute graft-versus-host disease after allogeneic stem cell transplantation. Leukemia (2014) 28:842–52. doi:10.1038/leu.2013.210
119. Weissinger EM, Mischak H, Kontsendorn J, Hahn A, Hahn N, Morgan M, et al. Proteome analysis in hematology using capillary electrophoresis coupled on-line to mass spectrometry. Mini Rev Med Chem (2009) 9:627–623. doi:10.2174/138955709788167583
120. Weissinger EM, Mischak H. Application of proteomics to posttransplantational follow-up. Methods Mol Med (2007) 134:217–28. doi:10.1007/978-1-59745-223-6_15
121. Weissinger EM, Hertenstein B, Mischak H, Ganser A. Online coupling of capillary electrophoresis with mass spectrometry for the identification of biomarkers for clinical diagnosis. Expert Rev Proteomics (2005) 2:639–47. doi:10.1586/14789450.2.5.639
122. Weissinger EM, Mischak H, Ganser A, Hertenstein B. Value of proteomics applied to the follow-up in stem cell transplantation. Ann Hematol (2006) 85:205–11. doi:10.1007/s00277-005-0057-1
123. Weissinger EM, Dickinson AM. Immunogenomics and proteomics in hematopoietic stem cell transplantation: predicting post-hematopoietic stem cell transplant complications. Cancer Treat Res (2009) 144:95–129. doi:10.1007/978-0-387-78580-6_5
124. Pavletic SZ, Martin P, Lee SJ, Mitchell S, Jacobsohn D, Cowen EW, et al. Measuring therapeutic response in chronic graft-versus-host disease: National Institutes of Health consensus development project on criteria for clinical trials in chronic graft-versus-host disease: IV. Response criteria working group report. Biol Blood Marrow Transplant (2006) 12:252–66. doi:10.1016/j.bbmt.2006.01.008
125. Jacobsohn DA, Kurland BF, Pidala J, Inamoto Y, Chai X, Palmer JM, et al. Correlation between NIH composite skin score, patient-reported skin score, and outcome: results from the Chronic GVHD Consortium. Blood (2012) 120:2545–52;quiz774. doi:10.1182/blood-2012-04-424135
126. Landfried K, Wolff D, Holler E. Pathophysiology and management of graft-versus-host disease in the era of reduced-intensity conditioning. Curr Opin Oncol (2009) 21(Suppl 1):S39–41. doi:10.1097/01.cco.0000357475.66035.d2
127. Wolff D, Steiner B, Hildebrandt G, Edinger M, Holler E. Pharmaceutical and cellular strategies in prophylaxis and treatment of graft-versus-host disease. Curr Pharm Des (2009) 15:1974–97. doi:10.2174/138161209788453158
128. Wolff D, Gerbitz A, Ayuk F, Kiani A, Hildebrandt GC, Vogelsang GB, et al. Consensus conference on clinical practice in chronic graft-versus-host disease (GVHD): first-line and topical treatment of chronic GVHD. Biol Blood Marrow Transplant (2010) 16:1611–28. doi:10.1016/j.bbmt.2010.06.015
129. Weissinger EM, Zurbig P, Ganser A. Proteomics studies after hematopoietic stem cell transplantation. Methods Mol Biol (2009) 506:437–52. doi:10.1007/978-1-59745-409-4_29
130. Forcade E, Kim HT, Cutler C, Wang K, Alho AC, Nikiforow S, et al. Circulating T follicular helper cells with increased function during chronic graft-versus-host disease. Blood (2016) 127:2489–97. doi:10.1182/blood-2015-12-688895
131. Jenq RR, Taur Y, Devlin SM, Ponce DM, Goldberg JD, Ahr KF, et al. Intestinal Blautia is associated with reduced death from graft-versus-host disease. Biol Blood Marrow Transplant (2015) 21:1373–83. doi:10.1016/j.bbmt.2015.04.016
132. Taur Y, Jenq RR, Ubeda C, van den Brink M, Pamer EG. Role of intestinal microbiota in transplantation outcomes. Best Pract Res Clin Haematol (2015) 28:155–61. doi:10.1016/j.beha.2015.10.013
133. Holler E, Butzhammer P, Schmid K, Hundsrucker C, Koestler J, Peter K, et al. Metagenomic analysis of the stool microbiome in patients receiving allogeneic stem cell transplantation: loss of diversity is associated with use of systemic antibiotics and more pronounced in gastrointestinal graft-versus-host disease. Biol Blood Marrow Transplant (2014) 20:640–5. doi:10.1016/j.bbmt.2014.01.030
134. Taur Y, Jenq RR, Perales MA, Littmann ER, Morjaria S, Ling L, et al. The effects of intestinal tract bacterial diversity on mortality following allogeneic hematopoietic stem cell transplantation. Blood (2014) 124:1174–82. doi:10.1182/blood-2014-02-554725
135. Weber D, Oefner PJ, Hiergeist A, Koestler J, Gessner A, Weber M, et al. Low urinary indoxyl sulfate levels early after transplantation reflect a disrupted microbiome and are associated with poor outcome. Blood (2015) 126:1723–8. doi:10.1182/blood-2015-04-638858
136. Holtan SG, DeFor TE, Lazaryan A, Bejanyan N, Arora M, Brunstein CG, et al. Composite end point of graft-versus-host disease-free, relapse-free survival after allogeneic hematopoietic cell transplantation. Blood (2015) 125:1333–8. doi:10.1182/blood-2014-10-609032
137. Rodriguez-Otero P, Porcher R, Peffault de Latour R, Contreras M, Bouhnik Y, Xhaard A, et al. Fecal calprotectin and alpha-1 antitrypsin predict severity and response to corticosteroids in gastrointestinal graft-versus-host disease. Blood (2012) 119:5909–17. doi:10.1182/blood-2011-12-397968
138. Ayuk F, Bussmann L, Zabelina T, Veit R, Alchalby H, Wolschke C, et al. Serum albumin level predicts survival of patients with gastrointestinal acute graft-versus-host disease after allogeneic stem cell transplantation. Ann Hematol (2014) 93:855–61. doi:10.1007/s00277-013-1957-0
139. Rezvani AR, Storer BE, Storb RF, Mielcarek M, Maloney DG, Sandmaier BM, et al. Decreased serum albumin as a biomarker for severe acute graft-versus-host disease after reduced-intensity allogeneic hematopoietic cell transplantation. Biol Blood Marrow Transplant (2011) 17:1594–601. doi:10.1016/j.bbmt.2011.07.021
Keywords: biomarkers, graft-versus-host disease, proteomics, genomics, cellular, heat shock protein, B-cell subsets
Citation: Juric MK, Shevtsov M, Mozes P, Ogonek J, Crossland RE, Dickinson AM, Greinix HT, Holler E, Weissinger EM and Multhoff G (2017) B-Cell-Based and Soluble Biomarkers in Body Liquids for Predicting Acute/Chronic Graft-versus-Host Disease after Allogeneic Hematopoietic Stem Cell Transplantation. Front. Immunol. 7:660. doi: 10.3389/fimmu.2016.00660
Received: 03 August 2016; Accepted: 16 December 2016;
Published: 16 January 2017
Edited by:
Antoine Toubert, Paris Diderot University, FranceReviewed by:
Laurent Garderet, Hôpital Saint-Antoine, FrancePhilippe Saas, Etablissement français du sang Bourgogne Franche-Comté, France
Eric Spierings, Utrecht University, Netherlands
Copyright: © 2017 Juric, Shevtsov, Mozes, Ogonek, Crossland, Dickinson, Greinix, Holler, Weissinger and Multhoff. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Gabriele Multhoff, Z2FicmllbGUubXVsdGhvZmZAdHVtLmRl
†These authors contributed equally to this work.
 Mateja Kralj Juric1†
Mateja Kralj Juric1† Maxim Shevtsov
Maxim Shevtsov Petra Mozes
Petra Mozes Rachel E. Crossland
Rachel E. Crossland Anne M. Dickinson
Anne M. Dickinson Hildegard T. Greinix
Hildegard T. Greinix Ernst Holler
Ernst Holler Gabriele Multhoff
Gabriele Multhoff