- 1Center for Virology and Vaccine Research, Beth Israel Deaconess Medical Center, Harvard Medical School, Boston, MA, United States
- 2Ragon Institute of Massachusetts General Hospital, MIT, and Harvard, Cambridge, MA, United States
Despite efficient suppression of plasma viremia in people living with HIV (PLWH) on cART, evidence of HIV-induced immunosuppression remains, and normally benign and opportunistic pathogens become major sources of co-morbidities, including virus-induced cancers. In fact, cancer remains a primary cause of death even in virally suppressed PLWH. Natural killer (NK) cells provide rapid early responses to HIV infection, contribute substantially to disease modulation and vaccine protection, and are also major therapeutic targets for cancer immunotherapy. However, much like other lymphocyte populations, recent burgeoning evidence suggests that in chronic conditions like HIV, NK cells can become functionally exhausted with impaired cytotoxic function, altered cytokine production and impaired antibody-dependent cell-mediated cytotoxicity. Recent work suggests functional anergy is likely due to low-level ongoing virus replication, increased inflammatory cytokines, or increased presence of MHClow target cells. Indeed, HIV-induced loss of NK cell-mediated control of lytic EBV infection has been specifically shown to cause lymphoma and also increases replication of CMV. In this review, we will discuss current understanding of NK cell modulation of HIV disease, reciprocal exhaustion of NK cells, and how this may impact increased cancer incidences and prospects for NK cell-targeted immunotherapies. Finally, we will review the most recent evidence supporting adaptive functions of NK cells and highlight the potential of adaptive NK cells for cancer immunotherapy.
NK Cells Have Therapeutic Potential to Enhance Control of Both HIV and HIV-Related Cancers
While efficient suppression of plasma viremia by combination antiretroviral therapy (cART) has substantially decreased mortality of people living with HIV (PLWH), burgeoning evidence suggests a higher occurrence of a vast range of comorbidities linked to long-term treatment and aging among PLWH, including cancers. The incidence of AIDS-defining cancers such as Kaposi Sarcoma, Non-Hodgkin lymphoma, and cervical cancer, has substantially decreased with access to cART. However, cART-treated PLWH still have a higher susceptibility to non-AIDS defining cancers (NADCs) compared to the general population, and NADCs currently represent a major cause of mortality among PLWH (1, 2). In particular, lymphomas, including Burkitt and classical Hodgkin lymphomas, have been reported at a significantly higher frequency in PLWH, yet many other cancers associated with infections (i.e., anus, oropharynx, liver) and some cancers associated with cigarette smoking (i.e., lung, kidney) were also found to be elevated among PLWH (3). Several mechanisms have been proposed to explain the predisposition of cART-treated PLWH to NADCs (4). Nevertheless, as cART treatment only partially prevents HIV-induced chronic inflammation and immune senescence, it is very likely that immune dysregulation in PLWH is an important determinant of NADCs and explains why most cancers predominantly found in PLWH are related to viral infections (4, 5).
NK Cell Subpopulations
Natural killer (NK) cells are large granular leukocytes that play a central role in the control of viral infections and neoplasms. Human NK cells are defined as CD3negCD56pos lymphocytes (6) and can be subdivided into functionally distinct subpopulations based on expression levels of CD56 and CD16 (7). CD56brightCD16neg NK cells have a high proliferation potential and the ability to secrete a large amount of cytokines, notably IFN-γ in response to IL-12, with limited cytotoxic functions (8), while CD56dimCD16pos NK cells display strong cytolytic activity as well as a significant capacity to secrete cytokines upon triggering of activating receptors (6, 9). In addition, a subset of CD56negCD16pos NK cells appears to expand in chronic viral infections including HIV and might represent an exhausted/anergic subset of NK cells (10–12).
Our understanding of human NK cells has essentially been acquired while studying peripheral blood NK cells, yet it is now clear that subsets other than CD56bright and CD56dim NK cell subpopulations can be found in peripheral tissues. Tissue-resident NK cells differ from circulating NK cells and are found not only in secondary lymphoid organs but also in many peripheral tissues including the uterus, lung, and liver where they represent up to 50% of lymphocytes (13–15). Findings from recent studies have allowed reliable identification of tissue-resident NK cells based on their expression of CD69, CD49a, or CD103, three markers functionally involved in the retention of lymphocytes in tissues. Besides the uterus, lung and liver, NK cells have been characterized in many additional tissues such as the intestinal mucosa, skin, and kidneys. However, in a majority of older studies it is not clear if those NK cells represent tissue-resident NK cells, NK cells circulating between tissues and blood, or innate lymphoid cells (ILCs). Indeed, ILCs can express markers associated with NK cells such as CD56, NKp46, or NKp44, and it was only lately appreciated that a deeper analysis of expressed transcription factors and produced cytokines is required to discriminate NK cells and ILCs. Until recently, NK cells were even considered as part of ILC group 1 due to the common innate lymphoid progenitors. However, NK cells are now distinguished from other ILCs because of their unique development and cytotoxic functions (16). In summary, tissue-resident NK cells likely play a crucial role in select tissues or organs involved in cancer and HIV disease, yet due to the scarcity of data on the contribution of tissue-resident NK cells in HIV infection or cancer development, herein we will focus primarily on circulating NK cells.
NK Cell Function
NK cells can efficiently discriminate between transformed or virally-infected cells and normal cells without the need for prior sensitization, and have the capacity to kill abnormal cells before adaptive immunity develops, thereby containing viral replication or tumor development. NK cells can clear cellular targets by a number of different mechanisms, including (i) exocytosis of cytotoxic granules containing perforin and granzyme that results in cell lysis, (ii) signaling through Fas ligand or TRAIL death receptors which induces apoptosis, (iii) release of cytokines with potent anti-viral and anti-tumor activities, and (iv) antibody-dependent cellular cytotoxicity (ADCC), triggered through binding of the FcγRIIIA receptor (CD16) on NK cells by the constant (Fc) domain of IgG antibodies. NK cells also play major roles in tuning and controlling adaptive immune responses (17).
NK Cell Receptors
Unlike other lymphocytes, NK cells lack antigen-specific receptors but lyse target cells following the integration of inhibitory and activating signals. These signals are generated by an arsenal of germline encoded cell surface molecules, with effector functions taking place when activating signals overcome inhibitory ones (18). The major NK cell receptors, which allow NK cells to discriminate between “self” and a variety of pathological cell states belong to three main categories: (i) natural cytotoxicity receptors (NCRs) such as NKp46, NKp30, and NKp44, which can bind to several viral or tumor-associated molecules (19, 20), (ii) NKG2A/C/E-CD94 heterodimers and NKG2D homodimers, which are c-type lectins binding to the non-classical Human Leukocyte Antigen E (HLA-E) molecule and stress-induced ligands, respectively, and (iii) the killer-cell immunoglobulin-like receptors (KIRs), which primarily recognize HLA class Ia (HLA-Ia) and Ib (HLA-Ib) molecules and related surface molecules (21).
The classical HLA-Ia group includes the highly polymorphic and ubiquitously expressed HLA-A, -B and -C antigens. Non-classical HLA-Ib antigens comprise HLA-E, -F, and -G molecules which are expressed in a tissue-specific manner, display low genetic diversity, and limited peptide repertoire (22). While the biological function and clinical relevance of most HLA-Ia and -Ib antigens have been investigated in detail, HLA-F was only recently recognized for its important immune-regulatory functions in cancer (23–26) and potentially in HIV infection (27). Besides their role in mediating recognition and elimination of unhealthy cells, a direct interaction between inhibitory KIRs and their HLA class I ligands during NK cell development is necessary for NK cells to acquire self-tolerance and functionality through an education process termed “licensing.” Besides NK cell licensing, which involves engagement of self-HLA class Ia molecules by their inhibitory ligand, non-classical HLA class I as well as non-HLA class I molecules also contribute to NK cell education (28).
While NKp30, NKp46, NKG2D, and NKG2C are expressed at relatively comparable levels on circulating CD56dim and CD56bright NK cells, other major NK cell receptors are differentially expressed on distinct subsets of NK cells (11). Peripheral blood CD56bright NK cells have been proposed to represent a mixture of immature NK cells that are direct precursors of CD56dim NK cells (29, 30), and mature NK cells, including CD56dim NK cells that have upregulated CD56 and lost CD16 upon activation in peripheral tissues (31). Immature CD56bright NK cells lack expression of KIRs, which are sequentially acquired during the differentiation process into mature CD56dim NK cells, a process that occurs in parallel with a progressive decrease in NKG2A expression and acquisition of the marker of terminal differentiation CD57 (32). NKp44 is usually not expressed on peripheral blood NK cells and up-regulated upon IL-2- or IL-15-mediated NK cell activation (33).
Other groups of receptors have received attention because their expression on NK cells is modulated in HIV and/or cancer and impacts NK cell function. These include Signaling Lymphocyte Activation Molecule (SLAM)-related receptors such as 2B4 (34–37) that displays co-stimulatory functions on NK cells and binds to CD48, or sialic acid-binding immunoglobulin-type lectins (Siglec), which are HLA class I-independent inhibitory receptors that recognize sialic acid-containing carbohydrates (38, 39). T cell immunoglobulin and mucin-domain containing-3 (Tim-3), which can binds to galectin-9, carcinoembryonic antigen cell adhesion molecule 1 (Ceacam1), high-mobility group box 1 (HMGB1) or phosphatidylserine (PtdSer), is another immunoregulatory molecule highly expressed on NK cells with relevance for NK cell function in both HIV and cancer (40–47). NK cells also express members of the immunoglobulin (Ig) superfamily such as the activating receptor DNAM-I (48–51), which has been shown to recognize CD112 (PVR) and CD155 (Nectin-2), two ligands expressed on tumor cells.
NK Cell Control of Cancers and HIV Infection
NK cells were originally defined as immune cells capable of lysing tumor cell lines. Since then, their capacity to kill primary cancer cells in vitro as well as their ability to prevent growth and metastasis of certain tumors in vivo, principally hematological cancers, has been clearly established (52–54). In particular, protection against development of cancer has been associated with higher NK cell cytotoxicity (55) and increasing evidence has highlighted the implication of NK cells in defense against leukemia. Importantly, in the context of hematopoietic stem cell transplantation (HSCT), it has been demonstrated that allogeneic NK cells from the donor can prevent relapse of myeloid leukemia via graft-vs.-leukemia effect (56, 57). However, thus far clinical trials aimed at harnessing NK cell anti-tumor activity have shown marginal therapeutic efficacy (58–61), with beneficial effects reported mainly against hematologic malignancies (62). Development of therapeutic strategies to enhance NK cell activity against tumor cells in vivo has therefore become a major field of investigation.
Besides NK cell anti-metastatic properties, numerous studies have emphasized the early and pivotal role of NK cells in the control of HIV infection. Notably, particular KIR genes expressed in conjunction with their HLA ligands are associated with significantly slower HIV disease progression and lower viral set-point (63, 64), elite control of HIV (65), and protection against disease acquisition (66, 67). In particular, activating KIR3DS1 has been associated with delayed HIV disease progression in individuals with specific HLA-B alleles since a first study by Martin et al. (63), yet a ligand for KIR3DS1 was only recently described, underscoring the relevance of HLA-F in regulating immunity to HIV (27). Indeed, HLA-F open conformers (OCs), which constitute heavy chains not bound to β2-microglobulin, can be recognized by several KIRs but have the highest affinity for KIR3DS1 (27, 68). HLA-F OCs trigger polyfunctional responses by KIR3DS1pos NK cells, which efficiently suppress HIV replication in vitro. HLA-F is expressed on activated CD4pos T cells and may act as a marker of cellular stress in specific conditions including viral infections and cellular transformation.
Control of HIV infection has also been associated with NK cells displaying potent cytotoxic function and IFN-γ expression after stimulation (69) as well as with polyfunctional CD8αpos NK cells (70). Moreover, it has been demonstrated that NK cells expand in the peripheral blood during early acute HIV infection, can inhibit HIV replication in vitro, and can mediate in vivo immune pressure in infected individuals, resulting in viral escape (71–77). Finally, indirect NK cell-mediated ADCC has been linked to vaccine-induced protective immunity against HIV infection (78), elite control of HIV (79–81) and slower HIV disease progression (82, 83). Therefore, in cART-treated PLWH, therapeutic interventions targeting NK cells might result in improved control of HIV and other viral infections as well as in decreased incidence of cancers.
Aberrant Expression of Key NK Cell Receptors May Contribute to Decreased Control of Pre-cancerous Cells in PLWH
NK cell-mediated immunosurveillance is decreased in PLWH, mostly as a long-term consequence of chronic HIV infection. While administration of suppressive cART partly restores NK cell properties, NK cells undergo many HIV-associated functional and phenotypic alterations, which are likely to severely impair NK cell-mediated control of viruses as well as of pre-cancerous cells.
Engagement of the well-described NCRs, NKG2D, and CD16 receptors represent major pathways to promote potent NK cell activation and cytotoxic responses. In both chronic HIV infection and cancer, NK cell recognition of abnormal cells through those activating receptors is defective, mainly as a result of chronic exposure to the respective ligands, which results in persistent down-modulation of NCRs, NKG2D, and CD16 on NK cells. In this section, we will review known effects that malignancies and HIV infection have on the expression of key NK cell receptors (Figure 1, left panel). It is important to note that a simplified definition of NK cells as CD3negCD56pos lymphocytes or different gating strategies to identify the major NK cell subsets represent a caveat of some older studies, precluding any definite conclusions on phenotypic alterations specifically affecting individual NK cell subsets.
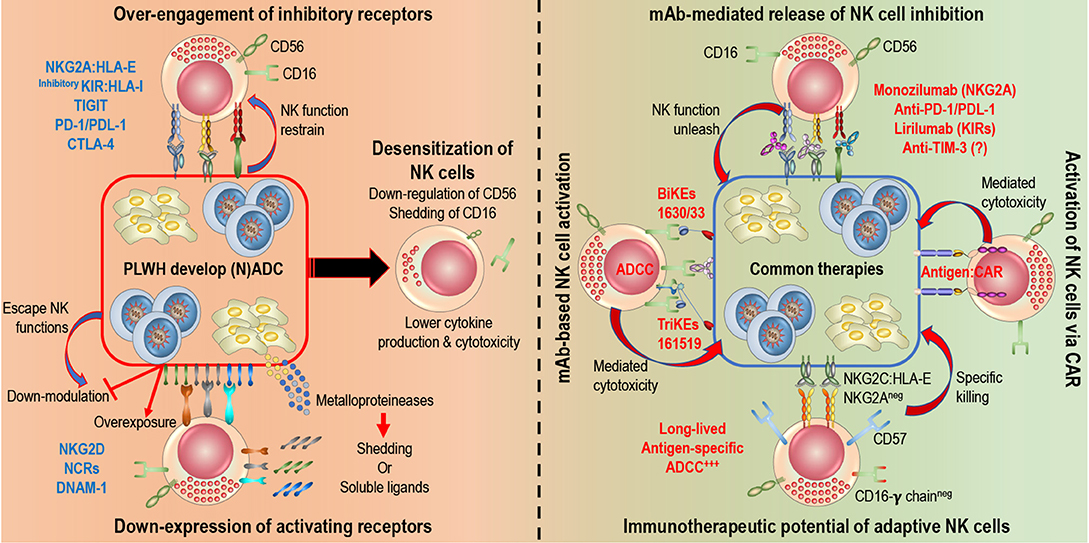
Figure 1. Rescuing and harnessing NK cell potency in PLWH developing cancers. Left: HIV-infected and cancer cells share common NK cell escape mechanisms. 1. Over-engagement of inhibitory receptors (i.e., NKG2A, inhibitory KIRs, PD-1…) blocks killing abilities of NK cells. 2. Down-modulation (blocking expression of ligand or shedding of ligand) or over-exposure (constant expression of ligands or release of soluble ligands) induce down-expression of activating receptors (NKG2D, NCRs, DNAM-1…) on NK cells. Right: Novel immunotherapies are being develop to harness NK cell potency and target HIV-infected and cancer cells. 1. Monoclonal antibodies (mAb) release engagement of inhibitory receptors, unleash NK cell cytotoxicity and engage Fc receptors (CD16) to induce ADCC. Several clinical trials are in progress. 2. Engineered proteins, Bi-specific or Tri-specific Killer engagers (BiKEs or TriKEs) and Chimeric Antigen Receptors (CARs), act as a link between NK cells and target cells to induce cytotoxicity. BiKEs or TriKEs induce ADCC by engaging CD16 receptors and bind to antigen on target cells. 3. Adaptive features of NK cells, defined by a higher expression of CD57, NKG2C, and/or absence of FcR-γ, could be harnessed to elicit specific killing of target cells.
NCRs represent a particularly important family of activating receptors in NK cell-mediated elimination of tumor cells, with a few tumor-associated ligands described for those molecules thus far (19). Accordingly, strategies to escape immune recognition by NCRs have been reported in both HIV infection and cancer, and have been associated with dysfunctional NK cells expressing lower levels of NCRs than NK cells from control subjects in many studies. Upon HIV infection, a population of dysfunctional CD56negCD16pos NK cells expands at the expense of the CD56dimCD16pos NK cell subset and is progressively eliminated with cART treatment. In PLWH, decreased NK cell expression of NKp30 and NKp46 receptors has been reported, and appears to be a characteristic of CD56negCD16pos NK cells, reducing their cytokine production and cytotoxicity, notably against tumor target cells, as well as their ability to interact with other immune cells (84–86). Similarly, decreased NK cell cytotoxicity in patients with acute or chronic myeloid leukemia (AML or CML) correlates with lower levels of NKp30 and NKp46 expression on NK cells compared to healthy individuals (87–89). NCRs downregulation on NK cells is induced by cell-to-cell contact with AML blasts and linked to poor survival in AML patients (87), whereas high levels of NKp30 and NKp46 expression on NK cells at AML diagnostic are predictive of better outcomes (90, 91). In AML, high expression of the immunosuppressive glycoprotein CD200 on tumor cells has been shown to directly impair NK cell anti-tumor responses and is associated with downregulated expression of NKp44 and NKp46 receptors on NK cells (92).
As overexposure to their ligands promotes decreased NCRs expression on NK cells, it is not surprising that shedding of NCR ligands is a hallmark of tumor escape, underscoring further the importance of this family of receptors in anti-metastatic NK cell functions. The A disintegrin and metalloproteinase ADAM-10 and ADAM-17 can cleave B7–H6, a ligand for NKp30, from the surface of tumors, likely leading to reduced NKp30 expression on NK cells surrounding the tumor (93, 94). Shedding of NKp30 ligands has also been described in chronic lymphocytic leukemia (CLL), in which exosomal expression of BAG6 mediates NK cell activation, whereas soluble BAG6 suppresses NK cell cytotoxicity (95). Galectin-3 is another molecule released by tumor cells that can serve as ligand for NKp30 and prevent NK cell activation (96). As another immune escape mechanism, catabolites specifically generated in tumor microenvironments, such as L-kynurenine, can also directly down-modulate NKp46 expression on NK cells (97). Whether HIV infection-associated NCR ligands are shed from the surface of infected cells remains to be fully determined, but likely contributes to impaired NCRpos NK cell function in HIV infection.
Altogether, these data suggest that fully restoring and even enhancing NCR-mediated signaling in NK cells might be crucial to efficiently control pre-cancerous cells in PLWH. Of note, B7–H6 is the only NCR ligand expressed on tumors that has been characterized so far. Identification of NCR ligands specifically expressed in cancer or HIV infection would represent a milestone in the development of therapeutic interventions aimed at maintaining NCR-mediated NK cell function in PLWH. Finally, given the crucial role played by NCRs in regulating NK cell function in both blood and tissues, therapeutic interventions to enhance tumor surveillance by NK cells and targeting NCR signaling are currently being explored (20).
NKG2D is one of the most important NK cell activating receptor in terms of recognition and elimination of abnormal cells expressing stress-induced ligands. Similarly to NCRs, tumors and HIV evolved immune escape mechanisms to specifically circumvent NKG2D-mediated recognition by NK cells. In HIV infection, reduced NKG2D expression on NK cells and dampened NK cell function have been linked to elevated levels of the soluble form of its major histocompatibility complex I-related chains A (MICA) ligand in patient sera (98). MICA is likely released by HIV-infected CD4pos T cells based on their increased expression levels of matrix metalloproteinases MMP-2 and -7, a family of enzymes previously described for their role in proteolytic shedding of NKG2D ligands in human tumors (99, 100). UL16 binding proteins (ULBP) also serve as ligands for NKG2D and their expression is induced on HIV-infected cells (34), yet levels of ULBP-1 and -2 is down-modulated by the HIV accessory protein Nef, thereby dampening NKG2D-mediated NK cell cytotoxicity (101).
Tumor progression has been associated with lower levels of NKG2D (as well as NKp30 and NKp46) expression on NK cells from patients with cervical cancer (102), and defective NK cell function owing to NKG2D downregulation has been linked to high-risk myelodysplastic syndrome (MDS) (103). Shedding of NKG2D ligands also plays a central role in tumor escape. In AML patients, chronic exposure to MICA/B decreases expression of NKG2D on NK cells (104) and the concentration of NKG2D soluble ligands in the peripheral blood correlates with reduced NK cell cytotoxicity in AML and CML (105). MICA is released in multiple myeloma (106, 107), and MICA/B as well as ULBP-6 are shed from leukemic cells (108). NKG2D ligand shedding is also involved in Hodgkin lymphoma, in which lymph node stromal cells express proteases that shed MICA and ULBP-3 from the surface of the lymphoma cells (109). Thus, NKG2D and its well-described ligands represent additional promising therapeutic target to enhance immunosurveillance by NK cells in PLWH. Accordingly, it has been recently demonstrated that antitumor responses by NK cells can be efficiently promoted by antibodies against MICA by blocking MICA/B shedding and coating MICA-expressing tumor cells, rendering them susceptible to ADCC (110).
Function of additional NK cell receptors is modulated by HIV infection and play an important role in NK cell responses to cancerous cells, including the activating receptor DNAM-1 that is expressed on the majority of peripheral blood NK cells (111–115). The CD155 ligand for DNAM-I has been shown to be present on HIV-infected T cells and, although discrepant results were obtained based on the cell culture model used, some studies found CD155 to be counter-regulated by the HIV proteins Nef and Vpu, thereby preventing NK cell activation (50, 51, 116). Many tumors also express ligands for DNAM-1, triggering NK cell cytokine production and cytotoxicity (117, 118). Tumor escape from DNAM-1 has been described and associated with DNAM-1 downregulation on NK cells isolated from patients with cancer (119–124).
Siglec receptors, and particularly Siglec-7 and -9, have also gained a lot of attention in the past decade for their involvement in immune evasion of tumor and virus-infected cells. Siglec-7 and Siglec-9 are constitutively expressed on all peripheral blood NK cells and on a mature subset of cytotoxic CD56dim NK cells, respectively (125). Reduced Siglec-7 expression marks a subset of dysfunctional NK cells that appears in early stages of HIV infection, prior to downmodulation of CD56, in subjects with elevated HIV replication, and also characterizes the dysfunctional CD56neg NK cell subset in chronic HIV infection (126, 127). Interestingly, an association between downregulation of Siglec-7 and dysfunction of NK cells has also been described in HIV-2 infection (128).
Siglec-7 and -9 ligands are widely expressed on distinct tumor cells and shield them from Siglec-7pos and Siglec-9pos NK cells (125). Siglec-10, another member of the Siglec family expressed by NK cells, is associated with decreased survival and impaired NK cell function in hepatocellular carcinoma (129). Therefore, targeting Siglec molecules on NK cells, or their ligands on malignant cells, might prove an attractive immunotherapeutic strategy to augment NK cell antitumor immunity (130). Supporting this hypothesis, a Siglec-7neg NK-92 cell line exhibited high cytotoxicity against leukemia cells in vitro (131).
Overall, interactions between ligands and major activating receptors on NK cells are impaired in both HIV and cancer, with some common underlying mechanisms such as cleavage of membrane-bound receptor molecules by zinc-dependent endopeptidases such as MMPs and ADAMs. Therefore, drugs that prevent shedding of NK cell-activating ligands or receptors may enhance protection against development of cancer in PLWH. Several inhibitors of the metalloproteinase ADAM17, for instance, have already entered clinical trials and are being tested in combination with other therapeutics against cancer (132). Metalloproteinase inhibitors would also prevent CD16 shedding form the surface of NK cells, a mechanism that naturally occurs following CD16 ligation (133, 134) yet is dysregulated in HIV infection and cancer, thereby decreasing ADCC activity and cytotoxicity against HIV-infected or tumor cells. Moreover, increased levels of inhibitory receptors such as inhibitory KIRs or TIGIT on NK cells further contribute to decreased NK cell functions in PLWH (135, 136). Finally, unresolved inflammation is a hallmark of chronic HIV infection and is widely accepted to elicit malignant transformation of cells and carcinogenesis (137, 138). Several inflammatory mediators, such as TNF-α, IL-6, tumor-derived transforming growth factor β (TGF-β), and IL-10 have been shown to play a role in carcinogenesis. For instance, TGF-β is a cytokine endowed with immune-suppressing and anti-inflammatory properties that plays a key role in promoting NK cell dysfunction and is found elevated in both the tumor microenvironement and plasma of PLWH. In addition, TGF-β has been shown to elicit production of vascular endothelial growth factor by NK cells, thereby promoting tumor growth along with other cytokines chronically found elevated in PLWH (137, 139).
Altogether, these observations show that the ability of NK cells to eliminate tumor cells is impaired by the tumor microenvironment and further constrained in HIV infection, and that NK cell dysfunction in cART-treated PLWH may significantly contribute to their enhanced susceptibility to develop malignancies.
Recent Advances in Development of NK Cell-Based Strategies for the Treatment of Cancer
NK cell-based immunotherapies rely on enhancement of endogenous NK cell activities in the tumor microenvironment or on adoptive transfer of NK cells with improved function. Strategies so far have included blockade of inhibitory NK cell receptors or immunosuppressive processes in the tumor microenvironment as well as enhancement of NK cell activation via cytokine stimulation or chimeric receptor expression (140). In this section, we will focus on strategies that could be of particular benefit in PLWH for elimination of HIV as well as HIV-associated cancers (Figure 1, right panel).
mAb-Mediated Release of NK Cell Inhibition
Autologous NK cells are oftentimes suppressed by self HLA class-I molecules expressed on tumor cells that bind to inhibitory CD94/NKG2A or KIR. This can be circumvented by adoptive therapy of allogeneic NK cells with a KIR-HLA class I mismatch. Alternatively, release of inhibitory signals using mAbs that target HLA class I-binding NK cell inhibitory receptors represent another strategy to enhance NK cell antitumor functions. This approach might be particularly beneficial in PLWH, as HIV infection results in downmodulation of the major activating NK cell receptors.
NKG2A is a c-type lectin that has been shown to mediate NK cell suppression in both HIV infection and cancer. In particular, elevated expression of HLA-A has been linked to poor control of HIV (141). This deleterious effect is mediated by NKG2Apos NK cells that are functionally suppressed by increased levels of HLA-E; whose expression is directly regulated by the availability of HLA class I-derived peptides. Whether increased HLA-A also correlates with poor outcome in cancer remains to be determined. Overexpression of HLA-E by tumor cells has long been proposed as a mechanism of escape from the action of NK cells (142). For instance, enhanced expression of HLA-E in hepatocarcinomas is driven by IL-10 released in the tumor micro-environment and is associated with enhanced NKG2A expression, a profile that correlates with NK cell exhaustion/anergy, as measured by low IFN-γ intracellular production upon stimulation with IL-12, and with a poorer prognosis (143). Failure to achieve remission in AML patients has been linked to impaired function of NK cells that upregulated NKG2A (144), and expression of HLA-E in multiple myeloma cells decreases NK cell cytotoxicity (145). Accordingly, efficacy of a specific IgG4 mAb that targets NKG2A (Monalizumab) is currently being assessed in various tumor settings along with other mAbs (61). Promising results were obtained in phase II trials in combination with the anti-EGFR antibody Cetuximab in head and neck cancers (146). Interestingly, Monalizumab targets both T cell and NK cell responses, promoting effector T cell responses in combination with anti-PDL1 and enhancing NK cell effector functions, including ADCC. Whether therapeutic blockade of HLA-E:NKG2A interaction, potentially in combination with PD-1 signaling blockade, could significantly improve control of HIV remains to be evaluated. NKG2A also significantly contributes to NK cell education in the early stages of NK cell ontogenesis. Accordingly, administration of Monalizumab has been suggested to promote NK cell alloreactivity against malignant cells when administered early after haplo-HSCT, thereby circumventing the need for a KIR-mismatched donor (147).
Inhibitory KIRs represent another interesting target for immunotherapies. For instance, Lirilumab, an IgG4 mAb that targets KIR2DL1/2/3 and KIR2DS1/2 has been evaluated in several clinical trials in combination with different mAbs in AML (phase II NCT02399917), MDS (phase II NCT02599649), lymphoma (phase II NCT01592370), and CLL (phase I NCT02481297). However, long-term use of inhibitory KIR blocking agents might lead to desensitization of NK cells (60). Finally, the recent discovery of HLA-F OCs' ability to bind KIRs, and particularly KIR3DS1 that has a widespread influence in human diseases including HIV, has made HLA-F a target of significant interest for therapies to enhance anti-tumor function of NK cells that might be particularly relevant for PLWH with malignancies.
Numerous antibody-based immune checkpoint inhibitors currently under investigation target the interaction of PD-1 or CTLA-4 and their cognate ligands on tumor cells, in order to boost the power of tumor-specific CD8pos T cells. In particular, clinical studies assessing the blockade of PD-1 or its ligand PD-L1 reported potent therapeutic efficacy against several cancers such as melanoma and non-small cell lung cancer. Selective PD-1 expression on CD56dimCD57pos mature NK cells in some but not all healthy individuals has been reported (148) and associated with functional defects (149). However, overall expression and functional relevance of those markers on NK cells in health and disease is still unclear, and recent studies suggest that blockade of CTLA-4 and PD-1 might enhance NK cell anti-tumor activity mostly via indirect mechanisms (150). Interestingly, PD-1 also mediates T-cell exhaustion in chronic HIV infection, and dual immune checkpoint blockade targeting PD-1 and IL-10 significantly enhances NK cell function through reversal of adaptive immune exhaustion in PLWH (151). Therefore, immunotherapeutic interventions targeting PD-1 may augment NK cell responses against both HIV and tumors in PLWH.
Another immune checkpoint inhibitor currently tested in clinic is a mAb targeting Tim-3, a receptor associated with exhaustion in T cells. Tim-3 has been proposed to mark mature NK cells, with chronic Tim-3 upregulation being associated with NK cell dysfunction, yet the precise impact of TIM-3 expression on NK cell function require further investigations (152). While Tim-3 has been shown to be upregulated on NK cells in various tumors, studies dissecting the effects of Tim-3 blockade on NK cell function in cancer settings have yielded mixed results (150).
Finally, even though it represents a promising approach for the treatment of cancer, administration of mAbs targeting regulatory immune checkpoint molecules has been associated with toxicities known as immune-related adverse events (irAEs) (153). irAEs are mainly caused by the release of inhibitory mechanisms that normally constrain the immune response, leading to various local and systemic autoimmune responses. Clinical benefit of immune checkpoint therapy is also restricted to a subset of patients. Several mechanisms of resistance to immune checkpoint inhibition have been described (154, 155). Notably, cancer therapy based on administration of mAbs promotes the induction of antibodies against such humanized mAbs and it is not clear yet whether such antibodies do or do not play a role by neutralizing the effects of the therapy. However, the potential of such antibodies to induce hypersensitivity reactions need to be considered.
mAb-Based NK Cell Activation
Antibody therapy that targets activating NK cell receptors is another strategy that has shown efficacy in certain malignancies. Elotuzumab, an antibody that targets SLAMF7, directly activates NK cells and can simultaneously induce ADCC by coating multiple myeloma cells, which express SLAMF7. The ability of a therapeutic mAb to induce ADCC results in potent NK cell activation and led to the design of bi-specific and tri-specific killer cell engagers, BiKEs and TriKEs, respectively. These single-chain variable fragment recombinant reagents can bind the tumor cells and NK cells via CD16 to induce direct killing via ADCC. This technique has been used in clinical trials where Hodgkin target cells expressing CD30 were linked to CD16 expressed on NK cells (156). The anti-CD16XCD33 BiKE activation can override the inhibitory signals mediated by ligation of inhibitory NK cell receptors and their HLA class I ligands expressed on AML (157) and MDS (158) targets. However, BiKEs do not promote in vivo proliferation and survival of NK cells. To overcome this issue, TriKEs were manufactured to engage the IL-15 receptor and are evaluated in different tumor pathologies (159–161). Use of therapeutic mAbs with potent ADCC activity may lead to substantial benefit in PLWH who present high frequencies of NK cells with enhanced antibody-dependent activation, as described in the last section.
Activation of NK Cells via CAR
A new tool for immunotherapy is chimeric antigen receptor (CAR)-engineered NK cells. CAR are artificial receptors composed of an extracellular antibody-derived tumor antigen binding domain as well as transmembrane and intracellular domains for activating signal transduction (162). Thus far, CAR T cells have been developed and successfully employed in the treatment of hematological malignancies. However, use of CAR-T cells has been limited as therapy for solid tumors and triggered numerous severe side effects in clinical trials that can be overcome with the use of CAR-NK cells. These include graft-vs.-host disease, cytokine release syndrome, and off-target toxicities. Moreover, CAR-NK cells can also eliminate tumor cells in a CAR-independent manner through recognition of ligands expressed on tumor cells by a range of activating receptors such as NKp30, NKG2D, DNAM-I, providing another advantage to use CAR-NK cells over CAR-T cells for cancer immunotherapies (163, 164). However, safety and efficacy of CAR-NK cells in humans need to be fully evaluated as only few clinical trials have been using CAR-NK cells up to now. One issue pertaining to CAR-NK cells is their limited in vivo persistence. To circumvent this restriction, a phase II trial is currently assessing the persistence and anti-tumor activity of IL-15- and caspase-9 suicide gene-transduced CD28-CAR-NK cells in B cell lymphoma (NCT03056339). Alternatively, CAR expression in adaptive NK cell subsets discussed in the next section may overcome expansion and persistence issues while simultaneously boosting anti-tumor activity. Finally, implementation of CAR-based strategies optimized for NK cells is warranted. For instance, induced pluripotent stem cell (iPSC)-derived NK cells transduced with novel CAR constructs that include NK cell-specific signaling domains instead of CD3ζ signaling-based domains are being evaluated and may significantly enhance their potency (165). Importantly, while CAR-T cell-based clinical trials have failed to provide clinical benefit and HIV viral suppression in PLWH, advanced CAR strategies that are developed specifically for NK cells in the cancer field can benefit PLWH as they could be applied to efficiently redirect NK cell functions toward HIV-infected cells (166).
Immunotherapeutic Potential of Adaptive NK Cells
While NK cells are classically viewed as non-specific effector cells of the innate immune system, a vast amount of independent studies has demonstrated that subsets of murine, non-human primate and human NK cells are capable of adaptive immune functions, including antigen-dependent expansion and long-lived immunological memory (167, 168). Adaptive NK cell-based immunotherapies may circumvent many of the limitations inherent to the various strategies tested thus far to harness anti-tumor functions of conventional NK cells.
The best characterized adaptive NK cell subset in humans is the one driven by HCMV infection, originally identified as a population of NK cells expressing high levels of the activating CD94/NKG2C receptor and the marker of terminal differentiation CD57, which expand upon HCMV infection or reactivation and can persist for years at high frequency in HCMV-seropositive individuals (169–173). Corroborating the adaptive features of this NK cell subset, it was recently shown that expansion and differentiation of this CD94/NKG2Cpos NK cell subset is driven by the HCMV UL40 peptide presented by HLA-E, the ligand for NKG2C (174).
The CD94/NKG2Cpos NK cell population largely overlaps with an FcεRIγ adaptor protein-deficient memory NK cell subset with enhanced antibody-dependent functions (FcγRΔg NK cells) that has more recently also been characterized in HCMV-seropositive subjects (175–184) and rhCMV-positive macaques (185). Adaptive characteristics of FcγRΔg NK cells include a distinctive epigenetic signature close to that of memory CD8pos T cells, endowing these adaptive NK cells with specialized functions such as enhanced responses to CD16 cross-linking, potent IFN-γ production to selective stimuli and reduced activation by innate cytokines.
Interestingly, adaptive CD94/NKG2Cpos NK cells proliferate not only in response to CMV reactivation or infection in patients receiving hematopoietic transplantation (169, 172, 186–188), but also upon de novo infection with different viruses including HIV and upon HCMV reactivation in PLWH (171, 189, 190). Several reports strongly suggest that HCMV-associated adaptive NK cells improve control of HIV infection. Higher frequencies of CD94/NKG2Cpos NK cells during primary HIV infection are linked to lower viral set points, are predictive of higher CD4pos T cell counts and of an overall better outcome in treated PLWH (191, 192). In contrast, individuals with NKG2C gene deletions are more susceptible to HIV infection and once infected may have accelerated disease progression (193). Finally, in HCMV-seropositive PLWH, CD94/NKG2Cpos NK cells exhibiting adaptive signatures of FcγRΔg NK cells present conserved effector functions (190). The beneficial effect of adaptive CD94/NKG2Cpos NK cells has also been demonstrated in cancer settings. HCMV reactivation has been linked to longer relapse-free survival in patients with hematological malignancies receiving allogeneic hematopoietic cell transplantation (194). More specifically, expansion of adaptive NKG2CposCD57pos NK cells upon HCMV reactivation after HCT is associated with reduced leukemia relapse (195, 196). Of note, specific phenotypic signatures have been associated with this NK cell adaptive subset and include lack of NKG2A expression. As a result, these cells are intrinsically insensitive to tumor-mediated suppression through HLA-E. Therefore, HCMV-associated adaptive NK cells represent an attractive subset of NK cells that could be exploited instead of conventional NK cells to limit cancer incidence in PLWH, particularly in combination with tumor-targeting therapeutic antibodies that efficiently promote NK cell-mediated ADCC.
NK cell memory has been described against multiple viral, bacterial, and tumor antigens, and can also be induced by brief exposure to specific cytokines. Indeed, NK cells can differentiate into cytokine-induced memory-like (CIML) NK cells that display enhanced effector functions after a short pre-activation with a combination of IL-12, IL-15, and IL-18 followed by a prolonged rest period (197). Re-stimulation of CIML NK cells using leukemia target cells, cytokines or FcγRIIIa ligation is associated with increased responsiveness that can be retained for several weeks following their initial pre-activation (197–202). CD56bright and CD56dim NK cells both have the potential to differentiate into CIML NK cells (197). Potent effector functions of CIML NK cells have been linked to expression of the high-affinity IL-2 receptor αβγ (IL-2Rαβγ), demethylation of the conserved upstream non-coding enhancer region of the IFN-γ gene, recruitment of anergic unlicensed NK cells, enhanced antibody-mediated functions and release from KIR-mediated inhibition (198, 200, 201, 203). Therefore, superior functionality of CIML NK cells is not affected by prior licensing through HLA class-I molecules. Compared to control NK cells, CIML NK cells have been shown to express higher levels of CD56, CD94, NKG2A, NKG2D, NKp46, CD25, NKp30, NKp44, CD62L, CD27, TRAIL, perforin and granzyme B, and lower levels of CD16, whereas NKG2C expression was found similar between control and CIML NK cells (197, 199).
The long-lived properties of CIML NK cells have tremendous potential to be exploited for cancer immunotherapy, and preliminary results from a first-in-human phase 1 clinical trial have shown that NK cells pre-activated with IL-12, IL-15, and IL-18 can expand in vivo and exert robust responses against leukemia targets, leading to remission in a subset of AML patients (199). A better understanding of the mechanisms behind CIML NK cell responses may lead to novel strategies to further enhance their antitumor function. For instance, recent studies suggested that targeting the interaction between SEMA7A, a potent immunomodulator expressed by cytokine-activated NK cells, and integrin-β1 might provide a novel immunotherapeutic approach to potentiate antitumor activity of CIML NK cells (204).
Strikingly, burgeoning evidence also supports the existence of true antigen-specific memory NK cells in humans (174, 177, 205), including a recent report of human HIV-specific memory NK cells (168). While further studies are warranted to fully characterize human antigen-specific NK cells and define the mechanisms underlying NK cell memory formation and maintenance, it is possible that adaptive NK cells that can specifically recognize tumor-associated antigens and efficiently eliminate cancerous cells develop in cancer patients. Vaccines including components to boost tumor-specific NK cells or infusion of expanded tumor-specific NK cells represent attractive avenues for the development of novel therapeutic interventions.
Overall, the immunotherapeutic potential of adaptive NK cells is expected to exceed that of conventional NK cells as they may overcome some of the major limitations faced in NK cell-based cancer therapies that have been evaluated so far in pre-clinical or clinical studies. For instance, adaptive NK cells can be expanded ex vivo, are long-lived and persist in vivo, are less sensitive to regulatory T cells-mediated suppression (206) or myeloid-derived suppressor cell inhibition (207) and can achieve significantly enhanced antibody-dependent functions (194) or antigen-specific cytotoxicity (168). Importantly, HCMV-dependent adaptive NK cells are increased 7-fold (181) and confer protection in PLWH (191, 192). Therefore, exploitation of adaptive NK cells may represent an attractive strategy to efficiently prevent or treat malignancies in PLWH.
Author Contributions
OL and SJ contributed to writing of specific sections. RR and SJ edited the final version of the manuscript.
Funding
This work was supported by National Institutes of Health (NIH) grants R01 AI116363 (SJ), R21 AI137835 (SJ), and R01 AI120828 (RR). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1. Rodger AJ, Lodwick R, Schechter M, Deeks S, Amin J, Gilson R, et al. Mortality in well controlled HIV in the continuous antiretroviral therapy arms of the SMART and ESPRIT trials compared with the general population. AIDS. (2013) 6:973–9. doi: 10.1097/QAD.0b013e32835cae9c
2. Zucchetto A, Virdone S, Taborelli M, Grande E, Camoni L, Pappagallo M, et al. Non-AIDS-defining cancer mortality: emerging patterns in the late HAART Era. J Acquir Immune Defic Syndr. (2016) 2:190–6. doi: 10.1097/QAI.0000000000001033
3. Shiels MS, Cole SR, Kirk GD, Poole C. A meta-analysis of the incidence of non-AIDS cancers in HIV-infected individuals. J Acquir Immune Defic Syndr. (2009) 5:611–22. doi: 10.1097/QAI.0b013e3181b327ca
4. Borges AH, Dubrow R, Silverberg MJ. Factors contributing to risk for cancer among HIV-infected individuals, and evidence that earlier combination antiretroviral therapy will alter this risk. Curr Opin HIV AIDS. (2014) 1:34–40. doi: 10.1097/COH.0000000000000025
5. Nasi M, De Biasi S, Gibellini L, Bianchini E, Pecorini S, Bacca V, et al. Ageing and inflammation in patients with HIV infection. Clin Exp Immunol. (2017) 1:44–52. doi: 10.1111/cei.12814
6. Cooper MA, Fehniger TA, Caligiuri MA. The biology of human natural killer-cell subsets. Trends Immunol. (2001) 11:633–40. doi: 10.1016/S1471-4906(01)02060-9
7. Lanier LL, Le AM, Civin CI, Loken MR, Phillips JH. The relationship of CD16 (Leu-11) and Leu-19 (NKH-1) antigen expression on human peripheral blood NK cells and cytotoxic T lymphocytes. J Immunol. (1986) 12:4480–6.
8. Cooper MA, Fehniger TA, Turner SC, Chen KS, Ghaheri BA, Ghayur T, et al. Human natural killer cells: a unique innate immunoregulatory role for the CD56(bright) subset. Blood. (2001) 10:3146–51. doi: 10.1182/blood.V97.10.3146
9. Caligiuri MA, Zmuidzinas A, Manley TJ, Levine H, Smith KA, Ritz J. Functional consequences of interleukin 2 receptor expression on resting human lymphocytes. Identification of a novel natural killer cell subset with high affinity receptors. J Exp Med. (1990) 5:1509–26. doi: 10.1084/jem.171.5.1509
10. Alter G, Teigen N, Davis BT, Addo MM, Suscovich TJ, Waring MT, et al. Sequential deregulation of NK cell subset distribution and function starting in acute HIV-1 infection. Blood. (2005) 10:3366–9. doi: 10.1182/blood-2005-03-1100
11. Bjorkstrom NK, Ljunggren HG, Sandberg JK. CD56 negative NK cells: origin, function, and role in chronic viral disease. Trends Immunol. (2010) 11:401–6. doi: 10.1016/j.it.2010.08.003
12. Mavilio D, Benjamin J, Daucher M, Lombardo G, Kottilil S, Planta MA, et al. Natural killer cells in HIV-1 infection: dichotomous effects of viremia on inhibitory and activating receptors and their functional correlates. Proc Natl Acad Sci USA. (2003) 25:15011–6. doi: 10.1073/pnas.2336091100
13. Bjorkstrom NK, Ljunggren HG, Michaelsson J. Emerging insights into natural killer cells in human peripheral tissues. Nat Rev Immunol. (2016) 5:310–20. doi: 10.1038/nri.2016.34
14. Panda SK, Colonna M. Innate lymphoid cells in mucosal immunity. Front Immunol. (2019) 10:861. doi: 10.3389/fimmu.2019.00861
15. Sojka DK, Yang L, Yokoyama WM. Uterine natural killer cells. Front Immunol. (2019) 10:960. doi: 10.3389/fimmu.2019.00960
16. Vivier E, Artis D, Colonna M, Diefenbach A, Di Santo JP, Eberl G, et al. Innate lymphoid cells: 10 years on. Cell. (2018) 5:1054–66. doi: 10.1016/j.cell.2018.07.017
17. Crome SQ, Lang PA, Lang KS, Ohashi PS. Natural killer cells regulate diverse T cell responses. Trends Immunol. (2013) 7:342–9. doi: 10.1016/j.it.2013.03.002
18. Lanier LL. NK cell recognition. Annu Rev Immunol. (2005) 23:225–74. doi: 10.1146/annurev.immunol.23.021704.115526
19. Pazina T, Shemesh A, Brusilovsky M, Porgador A, Campbell KS. Regulation of the functions of natural cytotoxicity receptors by interactions with diverse ligands and alterations in splice variant expression. Front Immunol. (2017) 8:369. doi: 10.3389/fimmu.2017.00369
20. Barrow AD, Martin CJ, Colonna M. The natural cytotoxicity receptors in health and disease. Front Immunol. (2019) 10:909. doi: 10.3389/fimmu.2019.00909
21. Pegram HJ, Andrews DM, Smyth MJ, Darcy PK, Kershaw MH. Activating and inhibitory receptors of natural killer cells. Immunol Cell Biol. (2011) 2:216–24. doi: 10.1038/icb.2010.78
22. Lin A, Yan WH. The emerging roles of human leukocyte antigen-F in immune modulation and viral infection. Front Immunol. (2019) 10:964. doi: 10.3389/fimmu.2019.00964
23. Harada A, Ishigami S, Kijima Y, Nakajo A, Arigami T, Kurahara H, et al. Clinical implication of human leukocyte antigen (HLA)-F expression in breast cancer. Pathol Int. (2015) 11:569–74. doi: 10.1111/pin.12343
24. Feng E, Liang T, Wang X, Du J, Tang K, Wang F, et al. Correlation of alteration of HLA-F expression and clinical characterization in 593 brain glioma samples. J Neuroinflamm. (2019) 1:33. doi: 10.1186/s12974-019-1418-3
25. Wu B, Yang H, Ying S, Lu H, Wang W, Lv J, et al. High HLA-F expression is a poor prognosis factor in patients with nasopharyngeal carcinoma. Anal Cell Pathol. (2018) 2018:7691704. doi: 10.1155/2018/7691704
26. Zhang X, Lin A, Zhang JG, Bao WG, Xu DP, Ruan YY, et al. Alteration of HLA-F and HLA I antigen expression in the tumor is associated with survival in patients with esophageal squamous cell carcinoma. Int J Cancer. (2013) 1:82–9. doi: 10.1002/ijc.27621
27. Garcia-Beltran WF, Holzemer A, Martrus G, Chung AW, Pacheco Y, Simoneau CR, et al. Open conformers of HLA-F are high-affinity ligands of the activating NK-cell receptor KIR3DS1. Nat Immunol. (2016) 9:1067–74. doi: 10.1038/ni.3513
28. He Y, Tian Z. NK cell education via nonclassical MHC and non-MHC ligands. Cell Mol Immunol. (2017) 4:321–30. doi: 10.1038/cmi.2016.26
29. Freud AG, Yu J, Caligiuri MA. Human natural killer cell development in secondary lymphoid tissues. Semin Immunol. (2014) 2:132–7. doi: 10.1016/j.smim.2014.02.008
30. Luetke-Eversloh M, Killig M, Romagnani C. Signatures of human NK cell development and terminal differentiation. Front Immunol. (2013) 4:499. doi: 10.3389/fimmu.2013.00499
31. Robertson MJ. Role of chemokines in the biology of natural killer cells. J Leukoc Biol. (2002) 2:173–83. doi: 10.1189/jlb.71.2.173
32. Bjorkstrom NK, Riese P, Heuts F, Andersson S, Fauriat C, Ivarsson MA, et al. Expression patterns of NKG2A, KIR, and CD57 define a process of CD56dim NK-cell differentiation uncoupled from NK-cell education. Blood. (2010) 19:3853–64. doi: 10.1182/blood-2010-04-281675
33. Arnon TI, Markel G, Mandelboim O. Tumor and viral recognition by natural killer cells receptors. Semin Cancer Biol. (2006) 5:348–58. doi: 10.1016/j.semcancer.2006.07.005
34. Ward J, Bonaparte M, Sacks J, Guterman J, Fogli M, Mavilio D, et al. HIV modulates the expression of ligands important in triggering natural killer cell cytotoxic responses on infected primary T-cell blasts. Blood. (2007) 4:1207–14. doi: 10.1182/blood-2006-06-028175
35. Mercier-Bataille D, Sanchez C, Baier C, Le Treut T, Mounier N, Mokhtari S, et al. Expression of activating receptors on natural killer cells from AIDS-related lymphoma patients. AIDS Res Ther. (2014) 11:38. doi: 10.1186/1742-6405-11-38
36. Ostrowski SR, Ullum H, Pedersen BK, Gerstoft J, Katzenstein TL. 2B4 expression on natural killer cells increases in HIV-1 infected patients followed prospectively during highly active antiretroviral therapy. Clin Exp Immunol. (2005) 3:526–33. doi: 10.1111/j.1365-2249.2005.02869.x
37. Claus M, Meinke S, Bhat R, Watzl C. Regulation of NK cell activity by 2B4, NTB-A and CRACC. Front Biosci. (2008) 13:956–65. doi: 10.2741/2735
38. Crocker PR, Paulson JC, Varki A. Siglecs and their roles in the immune system. Nat Rev Immunol. (2007) 4:255–66. doi: 10.1038/nri2056
39. Falco M, Biassoni R, Bottino C, Vitale M, Sivori S, Augugliaro R, et al. Identification and molecular cloning of p75/AIRM1, a novel member of the sialoadhesin family that functions as an inhibitory receptor in human natural killer cells. J Exp Med. (1999) 6:793–802. doi: 10.1084/jem.190.6.793
40. Taborda NA, Hernandez JC, Lajoie J, Juno JA, Kimani J, Rugeles MT, et al. Short communication: low expression of activation and inhibitory molecules on NK cells and CD4(+) T cells is associated with viral control. AIDS Res Hum Retroviruses. (2015) 6:636–40. doi: 10.1089/aid.2014.0325
41. Xu L, Huang Y, Tan L, Yu W, Chen D, Lu C, et al. Increased Tim-3 expression in peripheral NK cells predicts a poorer prognosis and Tim-3 blockade improves NK cell-mediated cytotoxicity in human lung adenocarcinoma. Int Immunopharmacol. (2015) 2:635–41. doi: 10.1016/j.intimp.2015.09.017
42. Han G, Chen G, Shen B, Li Y. Tim-3: an activation marker and activation limiter of innate immune cells. Front Immunol. (2013) 4:449. doi: 10.3389/fimmu.2013.00449
43. Jost S, Moreno-Nieves UY, Garcia-Beltran WF, Rands K, Reardon J, Toth I, et al. Dysregulated Tim-3 expression on natural killer cells is associated with increased Galectin-9 levels in HIV-1 infection. Retrovirology. (2013) 10:74. doi: 10.1186/1742-4690-10-74
44. So EC, Khaladj-Ghom A, Ji Y, Amin J, Song Y, Burch E, et al. NK cell expression of Tim-3: First impressions matter. Immunobiology. (2019) 224:362–70. doi: 10.1016/j.imbio.2019.03.001
45. Yin M, Di G, Bian M. Dysfunction of natural killer cells mediated by PD-1 and Tim-3 pathway in anaplastic thyroid cancer. Int Immunopharmacol. (2018) 64:333–39. doi: 10.1016/j.intimp.2018.09.016
46. Finney CA, Ayi K, Wasmuth JD, Sheth PM, Kaul R, Loutfy M, et al. HIV infection deregulates Tim-3 expression on innate cells: combination antiretroviral therapy results in partial restoration. J Acquir Immune Defic Syndr. (2013) 2:161–7. doi: 10.1097/QAI.0b013e318285cf13
47. de Kivit S, Lempsink LJ, Plants J, Martinson J, Keshavarzian A, Landay AL. Modulation of TIM-3 expression on NK and T cell subsets in HIV immunological non-responders. Clin Immunol. (2015) 1:28–35. doi: 10.1016/j.clim.2014.10.009
48. Lakshmikanth T, Burke S, Ali TH, Kimpfler S, Ursini F, Ruggeri L, et al. NCRs and DNAM-1 mediate NK cell recognition and lysis of human and mouse melanoma cell lines in vitro and in vivo. J Clin Invest. (2009) 5:1251–63. doi: 10.1172/JCI36022
49. Han B, Mao FY, Zhao YL, Lv YP, Teng YS, Duan M, et al. Altered NKp30, NKp46, NKG2D, and DNAM-1 expression on circulating NK cells is associated with tumor progression in human gastric cancer. J Immunol Res. (2018) 2018:6248590. doi: 10.1155/2018/6248590
50. Matusali G, Potesta M, Santoni A, Cerboni C, Doria M. The human immunodeficiency virus type 1 Nef and Vpu proteins downregulate the natural killer cell-activating ligand PVR. J Virol. (2012) 8:4496–504. doi: 10.1128/JVI.05788-11
51. Davis ZB, Sowrirajan B, Cogswell A, Ward JP, Planelles V, Barker E. CD155 on HIV-infected cells is not modulated by HIV-1 Vpu and Nef but synergizes with NKG2D ligands to trigger NK cell lysis of autologous primary HIV-infected cells. AIDS Res Hum Retroviruses. (2017) 2:93–100. doi: 10.1089/aid.2015.0375
52. Lopez-Soto A, Gonzalez S, Smyth MJ, Galluzzi L. Control of metastasis by NK cells. Cancer Cell. (2017) 2:135–54. doi: 10.1016/j.ccell.2017.06.009
53. Morvan MG, Lanier LL. NK cells and cancer: you can teach innate cells new tricks. Nat Rev Cancer. (2016) 1:7–19. doi: 10.1038/nrc.2015.5
54. Guillerey C, Huntington ND, Smyth MJ. Targeting natural killer cells in cancer immunotherapy. Nat Immunol. (2016) 9:1025–36. doi: 10.1038/ni.3518
55. Imai K, Matsuyama S, Miyake S, Suga K, Nakachi K. Natural cytotoxic activity of peripheral-blood lymphocytes and cancer incidence: an 11-year follow-up study of a general population. Lancet. (2000) 9244:1795–9. doi: 10.1016/S0140-6736(00)03231-1
56. Ruggeri L, Capanni M, Urbani E, Perruccio K, Shlomchik WD, Tosti A, et al. Effectiveness of donor natural killer cell alloreactivity in mismatched hematopoietic transplants. Science. (2002) 5562:2097–100. doi: 10.1126/science.1068440
57. Ruggeri L, Mancusi A, Burchielli E, Capanni M, Carotti A, Aloisi T, et al. NK cell alloreactivity and allogeneic hematopoietic stem cell transplantation. Blood Cells Mol Dis. (2008) 1:84–90. doi: 10.1016/j.bcmd.2007.06.029
58. Fang F, Xiao W, Tian Z. NK cell-based immunotherapy for cancer. Semin Immunol. (2017) 31:37–54. doi: 10.1016/j.smim.2017.07.009
59. Malmberg KJ, Carlsten M, Bjorklund A, Sohlberg E, Bryceson YT, Ljunggren HG. Natural killer cell-mediated immunosurveillance of human cancer. Semin Immunol. (2017) 31:20–29. doi: 10.1016/j.smim.2017.08.002
60. Chiossone L, Dumas PY, Vienne M, Vivier E. Natural killer cells and other innate lymphoid cells in cancer. Nat Rev Immunol. (2018) 11:671–88. doi: 10.1038/s41577-018-0061-z
61. Chiossone L, Vienne M, Kerdiles YM, Vivier E. Natural killer cell immunotherapies against cancer: checkpoint inhibitors and more. Semin Immunol. (2017) 31:55–63. doi: 10.1016/j.smim.2017.08.003
62. Cooley S, Parham P, Miller JS. Strategies to activate NK cells to prevent relapse and induce remission following hematopoietic stem cell transplantation. Blood. (2018) 10:1053–62. doi: 10.1182/blood-2017-08-752170
63. Martin MP, Gao X, Lee JH, Nelson GW, Detels R, Goedert JJ, et al. Epistatic interaction between KIR3DS1 and HLA-B delays the progression to AIDS. Nat Genet. (2002) 4:429–34. doi: 10.1038/ng934
64. Martin MP, Qi Y, Gao X, Yamada E, Martin JN, Pereyra F, et al. Innate partnership of HLA-B and KIR3DL1 subtypes against HIV-1. Nat Genet. (2007) 6:733–40. doi: 10.1038/ng2035
65. Malnati MS, Ugolotti E, Monti MC, Battista D, Vanni I, Bordo D, et al. Activating killer immunoglobulin receptors and HLA-C: a successful combination providing HIV-1 control. Sci Rep. (2017) 7:42470. doi: 10.1038/srep42470
66. Boulet S, Kleyman M, Kim JY, Kamya P, Sharafi S, Simic N, et al. A combined genotype of KIR3DL1 high expressing alleles and HLA-B*57 is associated with a reduced risk of HIV infection. AIDS. (2008) 12:1487–91. doi: 10.1097/QAD.0b013e3282ffde7e
67. Boulet S, Sharafi S, Simic N, Bruneau J, Routy JP, Tsoukas CM, et al. Increased proportion of KIR3DS1 homozygotes in HIV-exposed uninfected individuals. AIDS. (2008) 5:595–9. doi: 10.1097/QAD.0b013e3282f56b23
68. Dulberger CL, McMurtrey CP, Holzemer A, Neu KE, Liu V, Steinbach AM, et al. Human leukocyte antigen F presents peptides and regulates immunity through interactions with NK cell receptors. Immunity. (2017) 6:1018–29.e7. doi: 10.1016/j.immuni.2017.06.002
69. Vieillard V, Fausther-Bovendo H, Samri A, Debre P. Specific phenotypic and functional features of natural killer cells from HIV-infected long-term nonprogressors and HIV controllers. J Acquir Immune Defic Syndr. (2010) 5:564–73. doi: 10.1097/QAI.0b013e3181d0c5b4
70. Ahmad F, Hong HS, Jackel M, Jablonka A, Lu IN, Bhatnagar N, et al. High frequencies of polyfunctional CD8+ NK cells in chronic HIV-1 infection are associated with slower disease progression. J Virol. (2014) 21:12397–408. doi: 10.1128/JVI.01420-14
71. Alter G, Heckerman D, Schneidewind A, Fadda L, Kadie CM, Carlson JM, et al. HIV-1 adaptation to NK-cell-mediated immune pressure. Nature. (2011) 7358:96–100. doi: 10.1038/nature10237
72. Alter G, Martin MP, Teigen N, Carr WH, Suscovich TJ, Schneidewind A, et al. Differential natural killer cell-mediated inhibition of HIV-1 replication based on distinct KIR/HLA subtypes. J Exp Med. (2007) 12:3027–36. doi: 10.1084/jem.20070695
73. Alter G, Rihn S, Walter K, Nolting A, Martin M, Rosenberg ES, et al. HLA class I subtype-dependent expansion of KIR3DS1+ and KIR3DL1+ NK cells during acute human immunodeficiency virus type 1 infection. J Virol. (2009) 13:6798–805. doi: 10.1128/JVI.00256-09
74. Boulet S, Song R, Kamya P, Bruneau J, Shoukry NH, Tsoukas CM, et al. HIV protective KIR3DL1 and HLA-B genotypes influence NK cell function following stimulation with HLA-devoid cells. J Immunol. (2010) 4:2057–64. doi: 10.4049/jimmunol.0902621
75. O'Connell KA, Han Y, Williams TM, Siliciano RF, Blankson JN. Role of natural killer cells in a cohort of elite suppressors: low frequency of the protective KIR3DS1 allele and limited inhibition of human immunodeficiency virus type 1 replication in vitro. J Virol. (2009) 10:5028–34. doi: 10.1128/JVI.02551-08
76. Parsons MS, Boulet S, Song R, Bruneau J, Shoukry NH, Routy JP, et al. Mind the gap: lack of association between KIR3DL1*004/HLA-Bw4-induced natural killer cell function and protection from HIV infection. J Infect Dis. (2010) S356–60. doi: 10.1086/655966
77. Strbo N, de Armas L, Liu H, Kolber MA, Lichtenheld M, Pahwa S. IL-21 augments natural killer effector functions in chronically HIV-infected individuals. AIDS. (2008) 13:1551–60. doi: 10.1097/QAD.0b013e3283089367
78. Haynes BF, Gilbert PB, McElrath MJ, Zolla-Pazner S, Tomaras GD, Alam SM, et al. Immune-correlates analysis of an HIV-1 vaccine efficacy trial. N Engl J Med. (2012) 366:1275–86. doi: 10.1056/NEJMoa1113425
79. Madhavi V, Wines BD, Amin J, Emery S, Lopez E, Kelleher A, et al. HIV-1 Env- and Vpu-specific antibody-dependent cellular cytotoxicity responses associated with elite control of HIV. J Virol. (2017) 18:e00700-17. doi: 10.1128/JVI.00700-17
80. Lambotte O, Ferrari G, Moog C, Yates NL, Liao HX, Parks RJ, et al. Heterogeneous neutralizing antibody and antibody-dependent cell cytotoxicity responses in HIV-1 elite controllers. AIDS. (2009) 8:897–906. doi: 10.1097/QAD.0b013e328329f97d
81. Ackerman ME, Mikhailova A, Brown EP, Dowell KG, Walker BD, Bailey-Kellogg C, et al. Polyfunctional HIV-specific antibody responses are associated with spontaneous HIV control. PLoS Pathog. (2016) 1:e1005315. doi: 10.1371/journal.ppat.1005315
82. Kulkarni A, Kurle S, Shete A, Ghate M, Godbole S, Madhavi V, et al. Indian long-term non-progressors show broad ADCC responses with preferential recognition of V3 region of envelope and a region from tat protein. Front Immunol. (2017) 8:5. doi: 10.3389/fimmu.2017.00005
83. Wren LH, Chung AW, Isitman G, Kelleher AD, Parsons MS, Amin J, et al. Specific antibody-dependent cellular cytotoxicity responses associated with slow progression of HIV infection. Immunology. (2013) 2:116–23. doi: 10.1111/imm.12016
84. Brunetta E, Hudspeth KL, Mavilio D. Pathologic natural killer cell subset redistribution in HIV-1 infection: new insights in pathophysiology and clinical outcomes. J Leukoc Biol. (2010) 6:1119–30. doi: 10.1189/jlb.0410225
85. Fogli M, Costa P, Murdaca G, Setti M, Mingari MC, Moretta L, et al. Significant NK cell activation associated with decreased cytolytic function in peripheral blood of HIV-1-infected patients. Eur J Immunol. (2004) 8:2313–21. doi: 10.1002/eji.200425251
86. De Maria A, Fogli M, Costa P, Murdaca G, Puppo F, Mavilio D, et al. The impaired NK cell cytolytic function in viremic HIV-1 infection is associated with a reduced surface expression of natural cytotoxicity receptors (NKp46, NKp30 and NKp44). Eur J Immunol. (2003) 9:2410–8. doi: 10.1002/eji.200324141
87. Fauriat C, Just-Landi S, Mallet F, Arnoulet C, Sainty D, Olive D, et al. Deficient expression of NCR in NK cells from acute myeloid leukemia: evolution during leukemia treatment and impact of leukemia cells in NCRdull phenotype induction. Blood. (2007) 1:323–30. doi: 10.1182/blood-2005-08-027979
88. Costello RT, Knoblauch B, Sanchez C, Mercier D, Le Treut T, Sebahoun G. Expression of natural killer cell activating receptors in patients with chronic lymphocytic leukaemia. Immunology. (2012) 2:151–7. doi: 10.1111/j.1365-2567.2011.03521.x
89. Costello RT, Sivori S, Marcenaro E, Lafage-Pochitaloff M, Mozziconacci MJ, Reviron D, et al. Defective expression and function of natural killer cell-triggering receptors in patients with acute myeloid leukemia. Blood. (2002) 10:3661–7. doi: 10.1182/blood.V99.10.3661
90. Chretien AS, Fauriat C, Orlanducci F, Rey J, Borg GB, Gautherot E, et al. NKp30 expression is a prognostic immune biomarker for stratification of patients with intermediate-risk acute myeloid leukemia. Oncotarget. (2017) 30:49548–63. doi: 10.18632/oncotarget.17747
91. Chretien AS, Devillier R, Fauriat C, Orlanducci F, Harbi S, Le Roy A, et al. NKp46 expression on NK cells as a prognostic and predictive biomarker for response to allo-SCT in patients with AML. Oncoimmunology. (2017) 12:e1307491. doi: 10.1080/2162402X.2017.1307491
92. Coles SJ, Wang EC, Man S, Hills RK, Burnett AK, Tonks A, et al. CD200 expression suppresses natural killer cell function and directly inhibits patient anti-tumor response in acute myeloid leukemia. Leukemia. (2011) 5:792–9. doi: 10.1038/leu.2011.1
93. Pesce S, Tabellini G, Cantoni C, Patrizi O, Coltrini D, Rampinelli F, et al. B7-H6-mediated downregulation of NKp30 in NK cells contributes to ovarian carcinoma immune escape. Oncoimmunology. (2015) 4:e1001224. doi: 10.1080/2162402X.2014.1001224
94. Schlecker E, Fiegler N, Arnold A, Altevogt P, Rose-John S, Moldenhauer G, et al. Metalloprotease-mediated tumor cell shedding of B7-H6, the ligand of the natural killer cell-activating receptor NKp30. Cancer Res. (2014) 13:3429–40. doi: 10.1158/0008-5472.CAN-13-3017
95. Reiners KS, Topolar D, Henke A, Simhadri VR, Kessler J, Sauer M, et al. Soluble ligands for NK cell receptors promote evasion of chronic lymphocytic leukemia cells from NK cell anti-tumor activity. Blood. (2013) 18:3658–65. doi: 10.1182/blood-2013-01-476606
96. Wang W, Guo H, Geng J, Zheng X, Wei H, Sun R, et al. Tumor-released Galectin-3, a soluble inhibitory ligand of human NKp30, plays an important role in tumor escape from NK cell attack. J Biol Chem. (2014) 48:33311–9. doi: 10.1074/jbc.M114.603464
97. Della Chiesa M, Carlomagno S, Frumento G, Balsamo M, Cantoni C, Conte R, et al. The tryptophan catabolite L-kynurenine inhibits the surface expression of NKp46- and NKG2D-activating receptors and regulates NK-cell function. Blood. (2006) 13:4118–25. doi: 10.1182/blood-2006-03-006700
98. Nolting A, Dugast AS, Rihn S, Luteijn R, Carrington MF, Kane K, et al. MHC class I chain-related protein A shedding in chronic HIV-1 infection is associated with profound NK cell dysfunction. Virology. (2010) 1:12–20. doi: 10.1016/j.virol.2010.05.014
99. Waldhauer I, Goehlsdorf D, Gieseke F, Weinschenk T, Wittenbrink M, Ludwig A, et al. Tumor-associated MICA is shed by ADAM proteases. Cancer Res. (2008) 15:6368–76. doi: 10.1158/0008-5472.CAN-07-6768
100. Salih HR, Rammensee HG, Steinle A. Cutting edge: down-regulation of MICA on human tumors by proteolytic shedding. J Immunol. (2002) 8:4098–102. doi: 10.4049/jimmunol.169.8.4098
101. Cerboni C, Neri F, Casartelli N, Zingoni A, Cosman D, Rossi P, et al. Human immunodeficiency virus 1 Nef protein downmodulates the ligands of the activating receptor NKG2D and inhibits natural killer cell-mediated cytotoxicity. J Gen Virol. (2007) 88(Pt 1):242–50. doi: 10.1099/vir.0.82125-0
102. Garcia-Iglesias T, Del Toro-Arreola A, Albarran-Somoza B, Del Toro-Arreola S, Sanchez-Hernandez PE, Ramirez-Duenas MG, et al. Low NKp30, NKp46 and NKG2D expression and reduced cytotoxic activity on NK cells in cervical cancer and precursor lesions. BMC Cancer. (2009) 9:186. doi: 10.1186/1471-2407-9-186
103. Epling-Burnette PK, Bai F, Painter JS, Rollison DE, Salih HR, Krusch M, et al. Reduced natural killer (NK) function associated with high-risk myelodysplastic syndrome (MDS) and reduced expression of activating NK receptors. Blood. (2007) 11:4816–24. doi: 10.1182/blood-2006-07-035519
104. Sanchez-Correa B, Morgado S, Gayoso I, Bergua JM, Casado JG, Arcos MJ, et al. Human NK cells in acute myeloid leukaemia patients: analysis of NK cell-activating receptors and their ligands. Cancer Immunol Immunother. (2011) 8:1195–205. doi: 10.1007/s00262-011-1050-2
105. Hilpert J, Grosse-Hovest L, Grunebach F, Buechele C, Nuebling T, Raum T, et al. Comprehensive analysis of NKG2D ligand expression and release in leukemia: implications for NKG2D-mediated NK cell responses. J Immunol. (2012) 3:1360–71. doi: 10.4049/jimmunol.1200796
106. Kaiser BK, Yim D, Chow IT, Gonzalez S, Dai Z, Mann HH, et al. Disulphide-isomerase-enabled shedding of tumour-associated NKG2D ligands. Nature. (2007) 7143:482–6. doi: 10.1038/nature05768
107. Jinushi M, Vanneman M, Munshi NC, Tai YT, Prabhala RH, Ritz J, et al. MHC class I chain-related protein A antibodies and shedding are associated with the progression of multiple myeloma. Proc Natl Acad Sci USA. (2008) 4:1285–90. doi: 10.1073/pnas.0711293105
108. Szczepanski MJ, Szajnik M, Welsh A, Whiteside TL, Boyiadzis M. Blast-derived microvesicles in sera from patients with acute myeloid leukemia suppress natural killer cell function via membrane-associated transforming growth factor-beta1. Haematologica. (2011) 9:1302–9. doi: 10.3324/haematol.2010.039743
109. Zocchi MR, Catellani S, Canevali P, Tavella S, Garuti A, Villaggio B, et al. High ERp5/ADAM10 expression in lymph node microenvironment and impaired NKG2D ligands recognition in Hodgkin lymphomas. Blood. (2012) 6:1479–89. doi: 10.1182/blood-2011-07-370841
110. Ferrari de Andrade L, Tay RE, Pan D, Luoma AM, Ito Y, Badrinath S, et al. Antibody-mediated inhibition of MICA and MICB shedding promotes NK cell-driven tumor immunity. Science. (2018) 6383:1537–42. doi: 10.1126/science.aao0505
111. Chan CJ, Andrews DM, McLaughlin NM, Yagita H, Gilfillan S, Colonna M, et al. DNAM-1/CD155 interactions promote cytokine and NK cell-mediated suppression of poorly immunogenic melanoma metastases. J Immunol. (2010) 2:902–11. doi: 10.4049/jimmunol.0903225
112. Guillerey C, Ferrari de Andrade L, Vuckovic S, Miles K, Ngiow SF, Yong MC, et al. Immunosurveillance and therapy of multiple myeloma are CD226 dependent. J Clin Invest. (2015) 7:2904. doi: 10.1172/JCI82646
113. Croxford JL, Tang ML, Pan MF, Huang CW, Kamran N, Phua CM, et al. ATM-dependent spontaneous regression of early Emu-myc-induced murine B-cell leukemia depends on natural killer and T cells. Blood. (2013) 13:2512–21. doi: 10.1182/blood-2012-08-449025
114. Pende D, Spaggiari GM, Marcenaro S, Martini S, Rivera P, Capobianco A, et al. Analysis of the receptor-ligand interactions in the natural killer-mediated lysis of freshly isolated myeloid or lymphoblastic leukemias: evidence for the involvement of the Poliovirus receptor (CD155) and Nectin-2 (CD112). Blood. (2005) 5:2066–73. doi: 10.1182/blood-2004-09-3548
115. Castriconi R, Dondero A, Corrias MV, Lanino E, Pende D, Moretta L, et al. Natural killer cell-mediated killing of freshly isolated neuroblastoma cells: critical role of DNAX accessory molecule-1-poliovirus receptor interaction. Cancer Res. (2004) 24:9180–4. doi: 10.1158/0008-5472.CAN-04-2682
116. Bolduan S, Reif T, Schindler M, Schubert U. HIV-1 Vpu mediated downregulation of CD155 requires alanine residues 10, 14 and 18 of the transmembrane domain. Virology. (2014) 464–5:375–84. doi: 10.1016/j.virol.2014.07.034
117. Martinet L, Ferrari De Andrade L, Guillerey C, Lee JS, Liu J, Souza-Fonseca-Guimaraes F, et al. DNAM-1 expression marks an alternative program of NK cell maturation. Cell Rep. (2015) 1:85–97. doi: 10.1016/j.celrep.2015.03.006
118. Tahara-Hanaoka S, Shibuya K, Onoda Y, Zhang H, Yamazaki S, Miyamoto A, et al. Functional characterization of DNAM-1 (CD226) interaction with its ligands PVR (CD155) and nectin-2 (PRR-2/CD112). Int Immunol. (2004) 4:533–8. doi: 10.1093/intimm/dxh059
119. Stannard KA, Lemoine S, Waterhouse NJ, Vari F, Chatenoud L, Gandhi MK, et al. Human peripheral blood DNAM-1(neg) NK cells are a terminally differentiated subset with limited effector functions. Blood Adv. (2019) 11:1681–94. doi: 10.1182/bloodadvances.2018030676
120. El-Sherbiny YM, Meade JL, Holmes TD, McGonagle D, Mackie SL, Morgan AW, et al. The requirement for DNAM-1, NKG2D, and NKp46 in the natural killer cell-mediated killing of myeloma cells. Cancer Res. (2007) 18:8444–9. doi: 10.1158/0008-5472.CAN-06-4230
121. Peng YP, Xi CH, Zhu Y, Yin LD, Wei JS, Zhang JJ, et al. Altered expression of CD226 and CD96 on natural killer cells in patients with pancreatic cancer. Oncotarget. (2016) 41:66586–94. doi: 10.18632/oncotarget.11953
122. Mamessier E, Sylvain A, Thibult ML, Houvenaeghel G, Jacquemier J, Castellano R, et al. Human breast cancer cells enhance self tolerance by promoting evasion from NK cell antitumor immunity. J Clin Invest. (2011) 9:3609–22. doi: 10.1172/JCI45816
123. Carlsten M, Baumann BC, Simonsson M, Jadersten M, Forsblom AM, Hammarstedt C, et al. Reduced DNAM-1 expression on bone marrow NK cells associated with impaired killing of CD34+ blasts in myelodysplastic syndrome. Leukemia. (2010) 9:1607–16. doi: 10.1038/leu.2010.149
124. Sanchez-Correa B, Gayoso I, Bergua JM, Casado JG, Morgado S, Solana R, et al. Decreased expression of DNAM-1 on NK cells from acute myeloid leukemia patients. Immunol Cell Biol. (2012) 1:109–15. doi: 10.1038/icb.2011.15
125. Jandus C, Boligan KF, Chijioke O, Liu H, Dahlhaus M, Demoulins T, et al. Interactions between Siglec-7/9 receptors and ligands influence NK cell-dependent tumor immunosurveillance. J Clin Invest. (2014) 4:1810–20. doi: 10.1172/JCI65899
126. Brunetta E, Fogli M, Varchetta S, Bozzo L, Hudspeth KL, Marcenaro E, et al. The decreased expression of Siglec-7 represents an early marker of dysfunctional natural killer-cell subsets associated with high levels of HIV-1 viremia. Blood. (2009) 18:3822–30. doi: 10.1182/blood-2009-06-226332
127. Zulu MZ, Naidoo KK, Mncube Z, Jaggernath M, Goulder PJR, Ndung'u T, et al. Reduced expression of Siglec-7, NKG2A, and CD57 on terminally differentiated CD56(-)CD16(+) natural killer cell subset is associated with natural killer cell dysfunction in chronic HIV-1 clade C infection. AIDS Res Hum Retroviruses. (2017) 12:1205–13. doi: 10.1089/aid.2017.0095
128. Lucar O, Sadjo Diallo M, Bayard C, Samri A, Tarantino N, Debre P, et al. B7-H6-mediated downregulation of NKp30 in natural killer cells contributes to HIV-2 immune escape. AIDS. (2019) 1:23–32. doi: 10.1097/QAD.0000000000002061
129. Zhang P, Lu X, Tao K, Shi L, Li W, Wang G, et al. Siglec-10 is associated with survival and natural killer cell dysfunction in hepatocellular carcinoma. J Surg Res. (2015) 1:107–13. doi: 10.1016/j.jss.2014.09.035
130. Daly J, Carlsten M, O'Dwyer M. Sugar free: novel immunotherapeutic approaches targeting siglecs and sialic acids to enhance natural killer cell cytotoxicity against cancer. Front Immunol. (2019) 10:1047. doi: 10.3389/fimmu.2019.01047
131. Huang CH, Liao YJ, Fan TH, Chiou TJ, Lin YH, Twu YC. A developed NK-92MI cell line with Siglec-7(neg) phenotype exhibits high and sustainable cytotoxicity against leukemia cells. Int J Mol Sci. (2018) 19:E1073. doi: 10.3390/ijms19041073
132. Wu J, Mishra HK, Walcheck B. Role of ADAM17 as a regulatory checkpoint of CD16A in NK cells and as a potential target for cancer immunotherapy. J Leukoc Biol. (2019) 105:1297–303. doi: 10.1002/JLB.2MR1218-501R
133. Peruzzi G, Femnou L, Gil-Krzewska A, Borrego F, Weck J, Krzewski K, et al. Membrane-type 6 matrix metalloproteinase regulates the activation-induced downmodulation of CD16 in human primary NK cells. J Immunol. (2013) 4:1883–94. doi: 10.4049/jimmunol.1300313
134. Romee R, Foley B, Lenvik T, Wang Y, Zhang B, Ankarlo D, et al. NK cell CD16 surface expression and function is regulated by a disintegrin and metalloprotease-17 (ADAM17). Blood. (2013) 18:3599–608. doi: 10.1182/blood-2012-04-425397
135. Iannello A, Debbeche O, Samarani S, Ahmad A. Antiviral NK cell responses in HIV infection: II. viral strategies for evasion and lessons for immunotherapy and vaccination. J Leukoc Biol. (2008) 1:27–49. doi: 10.1189/jlb.0907649
136. Yin X, Liu T, Wang Z, Ma M, Lei J, Zhang Z, et al. Expression of the inhibitory receptor TIGIT is up-regulated specifically on NK cells with CD226 activating receptor from HIV-infected individuals. Front Immunol. (2018) 9:2341. doi: 10.3389/fimmu.2018.02341
137. Landskron G, De la Fuente M, Thuwajit P, Thuwajit C, Hermoso MA. Chronic inflammation and cytokines in the tumor microenvironment. J Immunol Res. (2014) 2014:149185. doi: 10.1155/2014/149185
138. Appay V, Sauce D. Immune activation and inflammation in HIV-1 infection: causes and consequences. J Pathol. (2008) 2:231–41. doi: 10.1002/path.2276
139. Bruno A, Focaccetti C, Pagani A, Imperatori AS, Spagnoletti M, Rotolo N, et al. The proangiogenic phenotype of natural killer cells in patients with non-small cell lung cancer. Neoplasia. (2013) 2:133–42. doi: 10.1593/neo.121758
140. Di Vito C, Mikulak J, Zaghi E, Pesce S, Marcenaro E, Mavilio D. NK cells to cure cancer. Semin Immunol. (2019). doi: 10.1016/j.smim.2019.03.004. [Epub ahead of print].
141. Ramsuran V, Naranbhai V, Horowitz A, Qi Y, Martin MP, Yuki Y, et al. Elevated HLA-A expression impairs HIV control through inhibition of NKG2A-expressing cells. Science. (2018) 6371:86–90. doi: 10.1126/science.aam8825
142. Lo Monaco E, Tremante E, Cerboni C, Melucci E, Sibilio L, Zingoni A, et al. Human leukocyte antigen E contributes to protect tumor cells from lysis by natural killer cells. Neoplasia. (2011) 9:822–30. doi: 10.1593/neo.101684
143. Sun C, Xu J, Huang Q, Huang M, Wen H, Zhang C, et al. High NKG2A expression contributes to NK cell exhaustion and predicts a poor prognosis of patients with liver cancer. Oncoimmunology. (2017) 1:e1264562. doi: 10.1080/2162402X.2016.1264562
144. Stringaris K, Sekine T, Khoder A, Alsuliman A, Razzaghi B, Sargeant R, et al. Leukemia-induced phenotypic and functional defects in natural killer cells predict failure to achieve remission in acute myeloid leukemia. Haematologica. (2014) 5:836–47. doi: 10.3324/haematol.2013.087536
145. Sarkar S, van Gelder M, Noort W, Xu Y, Rouschop KM, Groen R, et al. Optimal selection of natural killer cells to kill myeloma: the role of HLA-E and NKG2A. Cancer Immunol Immunother. (2015) 8:951–63. doi: 10.1007/s00262-015-1694-4
146. Andre P, Denis C, Soulas C, Bourbon-Caillet C, Lopez J, Arnoux T, et al. Anti-NKG2A mAb is a checkpoint inhibitor that promotes anti-tumor immunity by unleashing both T and NK cells. Cell. (2018) 7:1731–43 e13. doi: 10.1016/j.cell.2018.10.014
147. Zaghi E, Calvi M, Marcenaro E, Mavilio D, Di Vito C. Targeting NKG2A to elucidate natural killer cell ontogenesis and to develop novel immune-therapeutic strategies in cancer therapy. J Leukoc Biol. (2019) 6:1243–51. doi: 10.1002/JLB.MR0718-300R
148. Pesce S, Greppi M, Tabellini G, Rampinelli F, Parolini S, Olive D, et al. Identification of a subset of human natural killer cells expressing high levels of programmed death 1: a phenotypic and functional characterization. J Allergy Clin Immunol. (2017) 1:335–46 e3. doi: 10.1016/j.jaci.2016.04.025
149. Della Chiesa M, Pesce S, Muccio L, Carlomagno S, Sivori S, Moretta A, et al. Features of memory-like and PD-1(+) human NK cell subsets. Front Immunol. (2016) 7:351. doi: 10.3389/fimmu.2016.00351
150. Kim N, Lee HH, Lee HJ, Choi WS, Lee J, Kim HS. Natural killer cells as a promising therapeutic target for cancer immunotherapy. Arch Pharm Res. (2019) 42:591–606. doi: 10.1007/s12272-019-01143-y
151. Porichis F, Hart MG, Massa A, Everett HL, Morou A, Richard J, et al. Immune checkpoint blockade restores HIV-specific CD4 T cell help for NK cells. J Immunol. (2018) 3:971–81. doi: 10.4049/jimmunol.1701551
152. Carotta S. Targeting NK cells for anticancer immunotherapy: clinical and preclinical approaches. Front Immunol. (2016) 7:152. doi: 10.3389/fimmu.2016.00152
153. Marin-Acevedo JA, Dholaria B, Soyano AE, Knutson KL, Chumsri S, Lou Y. Next generation of immune checkpoint therapy in cancer: new developments and challenges. J Hematol Oncol. (2018) 1:39. doi: 10.1186/s13045-018-0582-8
154. Lim SY, Rizos H. Immune cell profiling in the age of immune checkpoint inhibitors: implications for biomarker discovery and understanding of resistance mechanisms. Mamm Genome. (2018) 11–12:866–78. doi: 10.1007/s00335-018-9757-4
155. Jenkins RW, Barbie DA, Flaherty KT. Mechanisms of resistance to immune checkpoint inhibitors. Br J Cancer. (2018) 1:9–16. doi: 10.1038/bjc.2017.434
156. Reusch U, Burkhardt C, Fucek I, Le Gall F, Le Gall M, Hoffmann K, et al. A novel tetravalent bispecific TandAb (CD30/CD16A) efficiently recruits NK cells for the lysis of CD30+ tumor cells. MAbs. (2014) 3:728–39. doi: 10.4161/mabs.28591
157. Wiernik A, Foley B, Zhang B, Verneris MR, Warlick E, Gleason MK, et al. Targeting natural killer cells to acute myeloid leukemia in vitro with a CD16 x 33 bispecific killer cell engager and ADAM17 inhibition. Clin Cancer Res. (2013) 14:3844–55. doi: 10.1158/1078-0432.CCR-13-0505
158. Gleason MK, Ross JA, Warlick ED, Lund TC, Verneris MR, Wiernik A, et al. CD16xCD33 bispecific killer cell engager (BiKE) activates NK cells against primary MDS and MDSC CD33+ targets. Blood. (2014) 19:3016–26. doi: 10.1182/blood-2013-10-533398
159. Felices M, Kodal B, Hinderlie P, Kaminski MF, Cooley S, Weisdorf DJ, et al. Novel CD19-targeted TriKE restores NK cell function and proliferative capacity in CLL. Blood Adv. (2019) 6:897–907. doi: 10.1182/bloodadvances.2018029371
160. Sarhan D, Brandt L, Felices M, Guldevall K, Lenvik T, Hinderlie P, et al. 161533 TriKE stimulates NK-cell function to overcome myeloid-derived suppressor cells in MDS. Blood Adv. (2018) 12:1459–69. doi: 10.1182/bloodadvances.2017012369
161. Vallera DA, Felices M, McElmurry R, McCullar V, Zhou X, Schmohl JU, et al. IL15 Trispecific Killer Engagers (TriKE) make natural killer cells specific to CD33+ targets while also inducing persistence, in vivo expansion, and enhanced function. Clin Cancer Res. (2016) 14:3440–50. doi: 10.1158/1078-0432.CCR-15-2710
162. Rezvani K, Rouce R, Liu E, Shpall E. Engineering natural killer cells for cancer immunotherapy. Mol Ther. (2017) 8:1769–81. doi: 10.1016/j.ymthe.2017.06.012
163. Hu Y, Tian ZG, Zhang C. Chimeric antigen receptor (CAR)-transduced natural killer cells in tumor immunotherapy. Acta Pharmacol Sin. (2018) 2:167–76. doi: 10.1038/aps.2017.125
164. Glienke W, Esser R, Priesner C, Suerth JD, Schambach A, Wels WS, et al. Advantages and applications of CAR-expressing natural killer cells. Front Pharmacol. (2015) 6:21. doi: 10.3389/fphar.2015.00021
165. Li Y, Hermanson DL, Moriarity BS, Kaufman DS. Human iPSC-derived natural killer cells engineered with chimeric antigen receptors enhance anti-tumor activity. Cell Stem Cell. (2018) 2:181–92.e5. doi: 10.1016/j.stem.2018.06.002
166. Zenere G, Olwenyi OA, Byrareddy SN, Braun SE. Optimizing intracellular signaling domains for CAR NK cells in HIV immunotherapy: a comprehensive review. Drug Discov Today. (2019) 4:983–91. doi: 10.1016/j.drudis.2019.02.002
167. Paust S, Blish CA, Reeves RK. Redefining memory: building the case for adaptive NK cells. J Virol. (2017) 91:e00169-17. doi: 10.1128/JVI.00169-17
168. Nikzad R, Angelo LS, Aviles-Padilla K, Le DT, Singh VK, Bimler L, et al. Human natural killer cells mediate adaptive immunity to viral antigens. Sci Immunol. (2019) 35:eaat8116. doi: 10.1126/sciimmunol.aat8116
169. Foley B, Cooley S, Verneris MR, Pitt M, Curtsinger J, Luo X, et al. Cytomegalovirus reactivation after allogeneic transplantation promotes a lasting increase in educated NKG2C+ natural killer cells with potent function. Blood. (2012) 11:2665–74. doi: 10.1182/blood-2011-10-386995
170. Guma M, Angulo A, Vilches C, Gomez-Lozano N, Malats N, Lopez-Botet M. Imprint of human cytomegalovirus infection on the NK cell receptor repertoire. Blood. (2004) 12:3664–71. doi: 10.1182/blood-2004-05-2058
171. Guma M, Budt M, Saez A, Brckalo T, Hengel H, Angulo A, et al. Expansion of CD94/NKG2C+ NK cells in response to human cytomegalovirus-infected fibroblasts. Blood. (2006) 9:3624–31. doi: 10.1182/blood-2005-09-3682
172. Lopez-Verges S, Milush JM, Schwartz BS, Pando MJ, Jarjoura J, York VA, et al. Expansion of a unique CD57(+)NKG2Chi natural killer cell subset during acute human cytomegalovirus infection. Proc Natl Acad Sci USA. (2011) 36:14725–32. doi: 10.1073/pnas.1110900108
173. Hwang I, Zhang T, Scott JM, Kim AR, Lee T, Kakarla T, et al. Identification of human NK cells that are deficient for signaling adaptor FcRgamma and specialized for antibody-dependent immune functions. Int Immunol. (2012) 12:793–802. doi: 10.1093/intimm/dxs080
174. Hammer Q, Ruckert T, Borst EM, Dunst J, Haubner A, Durek P, et al. Peptide-specific recognition of human cytomegalovirus strains controls adaptive natural killer cells. Nat Immunol. (2018) 5:453–63. doi: 10.1038/s41590-018-0082-6
175. Azzi T, Lunemann A, Murer A, Ueda S, Beziat V, Malmberg KJ, et al. Role for early-differentiated natural killer cells in infectious mononucleosis. Blood. (2014) 16:2533–43. doi: 10.1182/blood-2014-01-553024
176. Hatton O, Strauss-Albee DM, Zhao NQ, Haggadone MD, Pelpola JS, Krams SM, et al. NKG2A-expressing natural killer cells dominate the response to autologous lymphoblastoid cells infected with epstein-barr virus. Front Immunol. (2016) 7:607. doi: 10.3389/fimmu.2016.00607
177. Hendricks DW, Balfour HH Jr, Dunmire SK, Schmeling DO, Hogquist KA, Lanier LL. Cutting edge: NKG2C(hi)CD57+ NK cells respond specifically to acute infection with cytomegalovirus and not Epstein-Barr virus. J Immunol. (2014) 10:4492–6. doi: 10.4049/jimmunol.1303211
178. Jud A, Kotur M, Berger C, Gysin C, Nadal D, Lunemann A. Tonsillar CD56brightNKG2A+ NK cells restrict primary Epstein-Barr virus infection in B cells via IFN-gamma. Oncotarget. (2017) 4:6130–41. doi: 10.18632/oncotarget.14045
179. Lunemann A, Vanoaica LD, Azzi T, Nadal D, Munz C. A distinct subpopulation of human NK cells restricts B cell transformation by EBV. J Immunol. (2013) 10:4989–95. doi: 10.4049/jimmunol.1301046
180. Oh JS, Ali AK, Kim S, Corsi DJ, Cooper CL, Lee SH. NK cells lacking FcepsilonRIgamma are associated with reduced liver damage in chronic hepatitis C virus infection. Eur J Immunol. (2016) 4:1020–9. doi: 10.1002/eji.201546009
181. Zhou J, Amran FS, Kramski M, Angelovich TA, Elliott J, Hearps AC, et al. An NK cell population lacking FcRgamma is expanded in chronically infected HIV patients. J Immunol. (2015) 10:4688–97. doi: 10.4049/jimmunol.1402448
182. Lee J, Zhang T, Hwang I, Kim A, Nitschke L, Kim M, et al. Epigenetic modification and antibody-dependent expansion of memory-like NK cells in human cytomegalovirus-infected individuals. Immunity. (2015) 3:431–42. doi: 10.1016/j.immuni.2015.02.013
183. Schlums H, Cichocki F, Tesi B, Theorell J, Beziat V, Holmes TD, et al. Cytomegalovirus infection drives adaptive epigenetic diversification of NK cells with altered signaling and effector function. Immunity. (2015) 3:443–56. doi: 10.1016/j.immuni.2015.02.008
184. Zhang T, Scott JM, Hwang I, Kim S. Cutting edge: antibody-dependent memory-like NK cells distinguished by FcRgamma deficiency. J Immunol. (2013) 4:1402–6. doi: 10.4049/jimmunol.1203034
185. Shah SV, Manickam C, Ram DR, Kroll K, Itell H, Permar SR, et al. CMV primes functional alternative signaling in adaptive deltag NK cells but is subverted by lentivirus infection in Rhesus Macaques. Cell Rep. (2018) 10:2766–74 e3. doi: 10.1016/j.celrep.2018.11.020
186. Muccio L, Bertaina A, Falco M, Pende D, Meazza R, Lopez-Botet M, et al. Analysis of memory-like natural killer cells in human cytomegalovirus-infected children undergoing alphabeta+T and B cell-depleted hematopoietic stem cell transplantation for hematological malignancies. Haematologica. (2016) 3:371–81. doi: 10.3324/haematol.2015.134155
187. Foley B, Cooley S, Verneris MR, Curtsinger J, Luo X, Waller EK, et al. Human cytomegalovirus (CMV)-induced memory-like NKG2C(+) NK cells are transplantable and expand in vivo in response to recipient CMV antigen. J Immunol. (2012) 10:5082–8. doi: 10.4049/jimmunol.1201964
188. Della Chiesa M, Falco M, Podesta M, Locatelli F, Moretta L, Frassoni F, et al. Phenotypic and functional heterogeneity of human NK cells developing after umbilical cord blood transplantation: a role for human cytomegalovirus? Blood. (2012) 2:399–410. doi: 10.1182/blood-2011-08-372003
189. Malmberg KJ, Beziat V, Ljunggren HG. Spotlight on NKG2C and the human NK-cell response to CMV infection. Eur J Immunol. (2012) 12:3141–5. doi: 10.1002/eji.201243050
190. Peppa D, Pedroza-Pacheco I, Pellegrino P, Williams I, Maini MK, Borrow P. Adaptive reconfiguration of natural killer cells in HIV-1 infection. Front Immunol. (2018) 9:474. doi: 10.3389/fimmu.2018.00474
191. Ma M, Wang Z, Chen X, Tao A, He L, Fu S, et al. NKG2C(+)NKG2A(-) natural killer cells are associated with a lower viral set point and may predict disease progression in individuals with primary HIV infection. Front Immunol. (2017) 8:1176. doi: 10.3389/fimmu.2017.01176
192. Gondois-Rey F, Cheret A, Granjeaud S, Mallet F, Bidaut G, Lecuroux C, et al. NKG2C(+) memory-like NK cells contribute to the control of HIV viremia during primary infection: optiprim-ANRS 147. Clin Transl Immunology. (2017) 7:e150. doi: 10.1038/cti.2017.22
193. Thomas R, Low HZ, Kniesch K, Jacobs R, Schmidt RE, Witte T. NKG2C deletion is a risk factor of HIV infection. AIDS Res Hum Retroviruses. (2012) 8:844–51. doi: 10.1089/aid.2011.0253
194. Capuano C, Pighi C, Battella S, Santoni A, Palmieri G, Galandrini R. Memory NK cell features exploitable in anticancer immunotherapy. J Immunol Res. (2019) 2019:8795673. doi: 10.1155/2019/8795673
195. Cichocki F, Taras E, Chiuppesi F, Wagner JE, Blazar BR, Brunstein C, et al. Adaptive NK cell reconstitution is associated with better clinical outcomes. JCI Insight. (2019) 2:e125553. doi: 10.1172/jci.insight.125553
196. Cichocki F, Cooley S, Davis Z, DeFor TE, Schlums H, Zhang B, et al. CD56dimCD57+NKG2C+ NK cell expansion is associated with reduced leukemia relapse after reduced intensity HCT. Leukemia. (2016) 2:456–63. doi: 10.1038/leu.2015.260
197. Romee R, Schneider SE, Leong JW, Chase JM, Keppel CR, Sullivan RP, et al. Cytokine activation induces human memory-like NK cells. Blood. (2012) 24:4751–60. doi: 10.1182/blood-2012-04-419283
198. Leong JW, Chase JM, Romee R, Schneider SE, Sullivan RP, Cooper MA, et al. Preactivation with IL-12, IL-15, and IL-18 induces CD25 and a functional high-affinity IL-2 receptor on human cytokine-induced memory-like natural killer cells. Biol Blood Marrow Transplant. (2014) 4:463–73. doi: 10.1016/j.bbmt.2014.01.006
199. Romee R, Rosario M, Berrien-Elliott MM, Wagner JA, Jewell BA, Schappe T, et al. Cytokine-induced memory-like natural killer cells exhibit enhanced responses against myeloid leukemia. Sci Transl Med. (2016) 357:357ra123. doi: 10.1126/scitranslmed.aaf2341
200. Wagner JA, Berrien-Elliott MM, Rosario M, Leong JW, Jewell BA, Schappe T, et al. Cytokine-induced memory-like differentiation enhances unlicensed natural killer cell antileukemia and FcgammaRIIIa-triggered responses. Biol Blood Marrow Transplant. (2017) 3:398–404. doi: 10.1016/j.bbmt.2016.11.018
201. Ewen EM, Pahl JHW, Miller M, Watzl C, Cerwenka A. KIR downregulation by IL-12/15/18 unleashes human NK cells from KIR/HLA-I inhibition and enhances killing of tumor cells. Eur J Immunol. (2018) 2:355–65. doi: 10.1002/eji.201747128
202. Terren I, Mikelez I, Odriozola I, Gredilla A, Gonzalez J, Orrantia A, et al. Implication of interleukin-12/15/18 and ruxolitinib in the phenotype, proliferation, and polyfunctionality of human cytokine-preactivated natural killer cells. Front Immunol. (2018) 9:737. doi: 10.3389/fimmu.2018.00737
203. Luetke-Eversloh M, Hammer Q, Durek P, Nordstrom K, Gasparoni G, Pink M, et al. Human cytomegalovirus drives epigenetic imprinting of the IFNG locus in NKG2Chi natural killer cells. PLoS Pathog. (2014) 10:e1004441. doi: 10.1371/journal.ppat.1004441
204. Ghofrani J, Lucar O, Dugan H, Reeves RK, Jost S. Semaphorin 7A modulates cytokine-induced memory-like responses by human natural killer cells. Eur J Immunol. (2019) 49:1153–66. doi: 10.1002/eji.201847931
205. Suliman S, Geldenhuys H, Johnson JL, Hughes JE, Smit E, Murphy M, et al. Bacillus Calmette-Guerin (BCG) revaccination of adults with latent Mycobacterium tuberculosis infection induces long-lived BCG-reactive NK cell responses. J Immunol. (2016) 4:1100–10. doi: 10.4049/jimmunol.1501996
206. Sarhan D, Hippen KL, Lemire A, Hying S, Luo X, Lenvik T, et al. Adaptive NK cells resist regulatory T-cell suppression driven by IL37. Cancer Immunol Res. (2018) 7:766–75. doi: 10.1158/2326-6066.CIR-17-0498
Keywords: HIV, cancer, natural killer, innate immunity, immunotherapy
Citation: Lucar O, Reeves RK and Jost S (2019) A Natural Impact: NK Cells at the Intersection of Cancer and HIV Disease. Front. Immunol. 10:1850. doi: 10.3389/fimmu.2019.01850
Received: 27 May 2019; Accepted: 23 July 2019;
Published: 14 August 2019.
Edited by:
Mirko Paiardini, Emory University School of Medicine, United StatesReviewed by:
Aftab A. Ansari, Emory University School of Medicine, United StatesNicolas Huot, Institut Pasteur, France
Copyright © 2019 Lucar, Reeves and Jost. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Stephanie Jost, c2pvc3RAYmlkbWMuaGFydmFyZC5lZHU=
 Olivier Lucar
Olivier Lucar R. Keith Reeves
R. Keith Reeves Stephanie Jost
Stephanie Jost