- 1Department of Hematology, Iuliu Hatieganu University of Medicine and Pharmacy, Cluj-Napoca, Romania
- 2Research Center for Functional Genomics, Biomedicine and Translational Medicine, Iuliu Hatieganu University of Medicine and Pharmacy, Cluj-Napoca, Romania
- 3“Octavian Fodor” Institute of Gastroenterology and Hepatology, Cluj-Napoca, Romania
- 4The Oncology Institute Prof. Dr. Ion Chiricuta, Cluj-Napoca, Romania
- 5Department of Gastroenterology, Iuliu Hatieganu University of Medicine and Pharmacy, Cluj-Napoca, Romania
Macrophages are known to have an impact in cytokine signaling in the myriad of organs in which they reside and are classically known to be either pro-inflammatory (M1), anti-inflammatory (M2). Different classes of signaling molecules influence these states, of which, microRNAs represent key modulators. These are short RNA species approximately 21 to 23 nucleotides long that generally act by binding to the 3′ untranslated region of mRNAs, regulating their translation, and, thus, the quantity of protein they encode. From these species, microRNA-155 was observed to be of great importance for M1 polarization. Because of it’s major implication in M1 polarization microRNA-155 was shown to be implicated in different inflammatory diseases. To name a few, microRNA-155 was shown to be modified in patients with asthma and to correlate with asthma symptoms in mouse model; it has been shown to modulate the activity of foam cells and influence the dimensions of the atherosclerotic plaque and it has also been shown to be of crucial influence in transducing the signal of LPS in septic shock. Because of this, the current review aims to offer an overview of the role of microRNA-155 in M1 polarization, the implication that this poses for the pathophysiology of inflammatory diseases and the potential therapeutic possibilities that this knowledge might bring. Currently, microRNA-155 has been used in clinical trials as a marker of inflammation, but the question remains if it’s inhibition will be useful in inflammatory diseases, as other products might have a better cost/benefit ratio.
Introduction
Macrophages have been known for their implication in the immune system, either through their phagocytic and/or antigen presenting abilities. Nonetheless, another important property of this type of cells is represented by their role in immune signaling. At the basis of their inflammatory functions stays the concept of macrophage polarization, which, in a simplified manner, is considered to be classic/proinflammatory (M1) or alternative/anti-inflammatory (M2) (1). The transition between different states of polarization is regulated by several classes of molecules, of which microRNAs present high importance. These are short RNA species of approximately 21 to 23 nucleotides that bind to different RNA species modifying their quantity or changing the rate at which they are translated. Other mechanisms through which microRNAs can act as regulatory molecules are represented by DNA binding at the promoter regions, influencing transcription or direct protein binding (2).
Because of the high number of microRNAs and targets that these species have, we decided to further focus on the implication of microRNA-155 in inflammatory diseases.
Macrophage Polarization
The continuum of macrophage polarization starts with the M0 polarized macrophage. One protein with key importance in macrophage polarization is represented by PU.1, which opens the chromatin conformation of macrophage-specific genes and further allow transcription factors to act in a cell-specific manner (3–6).
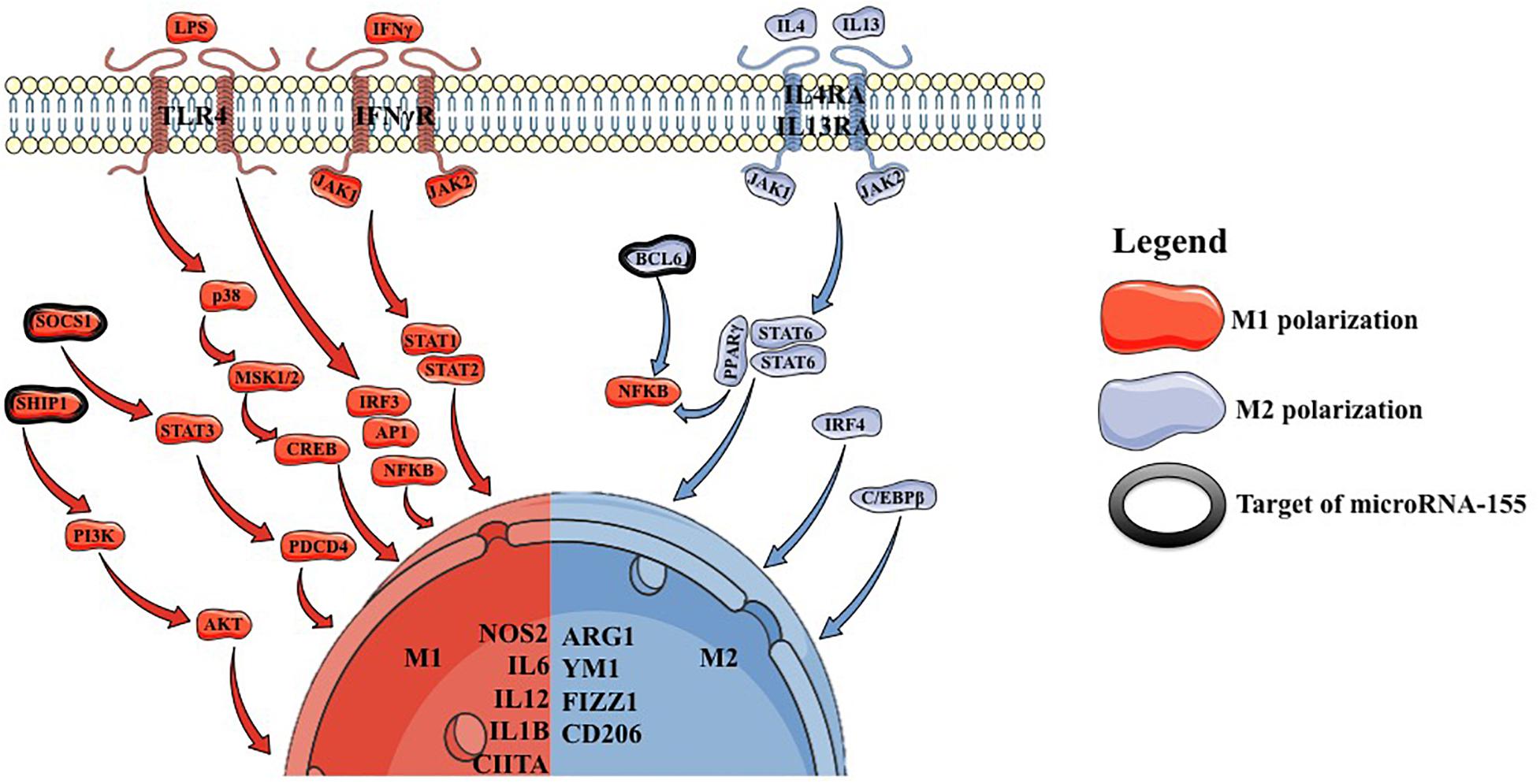
Conventionally, it was considered that macrophages can polarize either to an M1 or an M2 phenotype (Figure 1). Nowadays it is known that macrophages take a dynamic state between these two phenotypes.
M1 polarization can be induced in vitro by stimulating M0 macrophages with IFNγ (interferon γ) or with lipopolysaccharide (LPS), which mimics in vivo conditions of M0 to M1 transition. IFNγ acts on the IFNγR, which further recruits JAK1/2 leading to the formation STAT1/STAT2 heterodimers, which act as a transcription factor for NOS2, MHC2 and IL12 (7, 8). LPS acts through TLR4 (toll like receptor 4), stimulating IFNα/β autocrine signaling and leading to the activation of STAT1 and STAT2, with the formation of STAT1/STAT2 heterodimers, which act as a transcription factor for NOS2, CIITA and IL12, leading to a similar phenotype as generated through IFNγ direct stimulation. Additionally, TLR4 can also lead to the activation of NFKB and mitogen associated protein kinase (MAPK) pathways with similar effects (9). These stimulatory factors lead to the deposition of M1-specific transcription factors, followed by the upregulation of IL12 and downregulation of IL10 (10, 11).
M2 polarization is induced by a combination of IL4 and IL13 stimulation of M0 macrophages (12). This leads to STAT6 homodimer formation followed by the upregulation of MRC1, CD206, FIZZ1 and YM1 (13, 14). Additionally, STAT6 represents a cofactor for PPARγ leading to an inhibition of NFKB pathway (15–18). C/EBP was shown to increase the transcription of ARG1, IL10 and MRC1 (19–21). Moreover, mice lacking C/EBP, were shown to have a reduced number of M2 macrophages, but with no influence on M1 macrophages (22).
As M1 and M2 macrophages have opposite effects it has also been shown that STAT1 has opposite effects compared to STAT6 regarding the transcription profile they induce (23). Interestingly, JMJD3 was shown to play roles in both M1 and M2 polarization. In the case of M1 polarization, JMJD3 was shown to respond to LPS and induce IL12 and CCL5 (24). In the case of M2 polarization, it was shown that the macrophages of mice lacking JMJD3 are unable to undergo M2 polarization (14).
Although the general classification tends to picture these two macrophage polarization phenotypes as steady states, in vivo these represent a continuum and can transit from one to another. In this direction, one study has shown that after a period of LPS stimulation, the M1 response genes require a more intense stimulus for the same activation, while the response to IL10 stimulation is kept (25). Another mechanism through which the proinflammatory processes are controlled is represented by a multistep process activated by the NFKB pathway, which leads to the upregulation of microRNA-155 and an initial upregulation loop through microRNA-155 mediated inhibition of SHIP1, with subsequent activation of the PI3K/AKT pathway. In parallel the NFKB pathway upregulates microRNA-146a, which inhibits IRAK1 and TRAF6, leading to the resolution of the inflammatory process (26).
MicroRNA-155 Biogenesis and Mechanism of Action
MicroRNA-155 is located on chromosome 21 and is encoded by the gene MIRHA155. Its biogenesis is similar to the classical microRNA maturation. Firstly, it is transcribed in the form of pri-microRNA, which undergoes 3′ guanine capping and poly adenylation, after which it is processed in pre-microRNA by DROSHA. The pre-microRNA form is exported from the nucleus through XPO5 using a GTP-dependent process. In the cytoplasm, the pre-microRNA is processed by DICER, which cleaves the hairpin, resulting in the formation of a double-stranded RNA, which is loaded in AGO2. This, in turn, interacts with other proteins forming the RISC complex. In this complex the passenger strand is cleaved and only the guide strand remains loaded (2, 27, 28). Regarding microRNA-155, it is generally considered that the 5p form is the guide strand, thus being the one kept in the RISC complex after processing. Nonetheless, there have been studies showing that the 3p form can also act as a guide (29).
The Regulation of MicroRNA-155 Expression and Its Effect on Target Genes
MicroRNA-155 regulation is highly linked to its role as an immune modulator. This is being shown by its rapid increase in macrophages in infection or other inflammatory processes. These processes are characterized by the stimulation of DAMP, PAMP, IL1A, IL1B, TNFα, and IFNγ, while being suppressed by antiinflammatory molecules, e.g., IL10, TGFβ, and glucocorticoids (30–33). Therapeutically, it has been shown that glucocorticoids can indirectly inhibit microRNA-155 expression by inhibiting NFKB pathway (34, 35). MicroRNA-155 implication in inflammatory processes can also be inferred from the regulatory sites the MIRHA155 gene presents, these being influenced by NFKB pathway, TGFβ pathway through SMAD4, and through IL10 and IFNγ stimulation (33, 36).
It has been shown that microRNA-155 is upregulated by the stimulation of TLR4 and action of IFNγ, known drivers of M1 polarization. MicroRNA-155 role in promoting inflammation can also be observed as it inhibits INPP5D, an inhibitor of PI3K/AKT pathway, which is needed to relay the signal from the TLR4 signaling. Another important target of microRNA-155 affects is represented by SOCS1, which inhibits type 1 cytokine receptor/STAT pathways. Moreover, it has been shown that microRNA-155 inhibits BCL6, an inhibitor of NFKB pathway, with important implications in the signaling of foam cells (37–39).
Primary macrophages derived from mice lacking microRNA-155 and the RAW264.7 cell line with an induced deficit in microRNA-155 have been shown to present a resistance to LPS stimulation, associated with an increase in INPP5D expression (32, 37, 40). Not only does microRNA-155 stimulate M1 polarization, but it has also been shown to inhibit M2 polarization through the inhibition of IL13 and IL4 pathway components, like IL13RA and C/EBP (41–45).
MicroRNA-155-5P Vs MicroRNA-155-3P, Is There Only One Guide?
Generally, it is considered that microRNA-155-5p represents the guide strand as it targets a larger array of transcripts and it is considered to be the form with a higher thermodynamic stability. Nonetheless, there have been studies showing that microRNA-155-3p can also play biologically important roles (46, 47). Although the evidence is not as vast as in the case of microRNA-155-5p, there have been studies showing that microRNA-155-3p also has a role in promoting inflammation. For example, in the case of plasmacytoid dendritic cells, the initial response to TLR7 stimulation has been shown to be represented by microRNA-155-3p upregulation and subsequent upregulation of IFNα/β autocrine signaling. These processes add a parallel to LPS stimulation of macrophages with potential similar signaling effects (48, 49).
MicroRNA-155 in Inflammatory Diseases
MicroRNA-155 is implicated in the M1 polarization in a variety of inflammatory diseases with the following representing some examples in this direction (Figure 2).
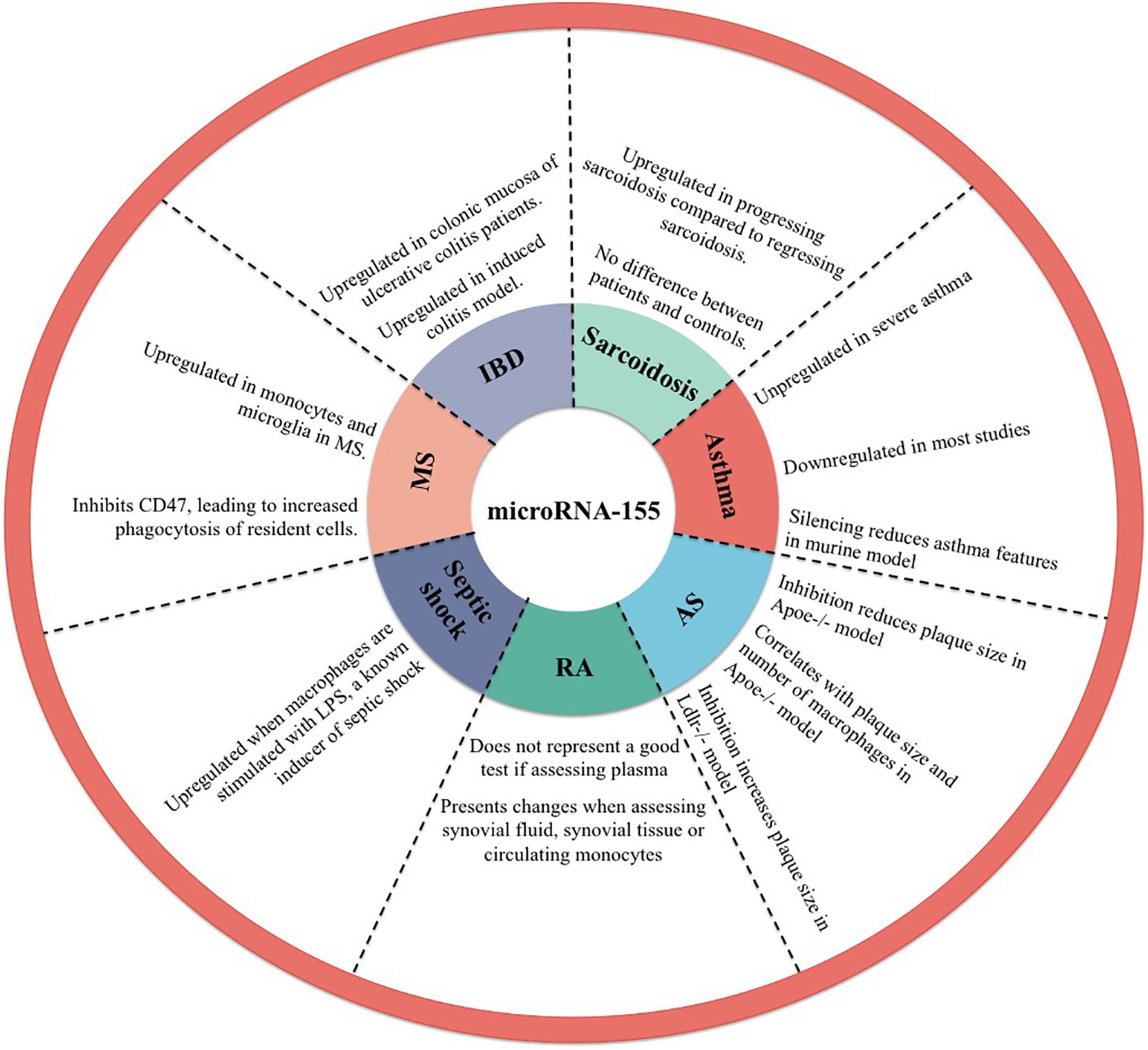
Figure 2. Implication of microRNA-155 in inflammatory diseases. AS, atherosclerosis; RA, rheumatoid arthritis; MS, multiple sclerosis; IBD, inflammatory bowel disease.
Asthma
Classically, asthma is characterized by a chronic imbalance between type 2 and type 1 immune responses and chronic hyper-reactivity of the airways. In this condition a myriad of microRNAs have been identified with implication in either M1 or M2 polarization (50, 51). Human airway smooth muscle cells from asthmatic patients were shown to present elevated levels of microRNA-155 compared to controls (52). Conversely, some studies have shown that microRNA-155 presents lower levels in breath condensates, plasma, nasal mucosa, epithelial cells and sputum of asthmatic patients (50, 52–55). Others have shown that severe asthma patients present elevated plasma microRNA-155 when compared to mild-moderate asthma patients and to healthy controls. Interestingly, in this study there was no difference observed between the microRNA-155 plasma levels of mild-moderate asthma patients compared to healthy controls (56). Moreover, it has been shown that microRNA-155 silencing in an ovalbumin asthma mouse model reduces asthma features, this, in turn, showing a potential therapeutic path for the future (57).
Atherosclerosis
Atherosclerosis is characterized by the formation of atheromatous plaques on arterial walls with subsequent reduction in blood flow, hypertension and an increased risk of embolus formation. One key cell with importance in the pathophysiology of atherosclerotic plaques is represented by the macrophage, which accumulate lipids and forms foam cells with a proinflammatory influence and leading to the evolution of the atherosclerotic plaque (58). Studies on APOE–/– mouse models with partial carotid ligation showed a decreased plaque formation and lower number of infiltrating macrophages when the microRNA-155 levels were inhibited (37). These results were reproduced by others showing that downregulating microRNA-155 in APOE–/– mouse model lowers the dimensions of the atherosclerotic plaque (59). The effect of microRNA-155 on atherosclerotic plaque formation has been explained through the modulation of the SOCS1-STAT3-PDCD4 pathway (60, 61). Considering these results, it could be though that microRNA-155 silencing might have a role in the future as therapy for decreasing the size of atherosclerotic plaques or to slow their evolution. In contrast to the above mentioned results, LDLR–/– mice models showed that transplantation with microRNA-155 deficient macrophages leads to an increase in the size of atherosclerotic plaque and leads to a decrease in IL10 levels secreted by peritoneal macrophages (62).
Rheumatoid Arthritis
Rheumatoid arthritis (RA) is characterized by progressive joint inflammation leading to joint damage and subsequent disability. One important hint that macrophage present an important role in this condition is represented by an upregulation in IL6 that these patients present, as this represents an important cytokine secreted by M1 macrophages (63). In RA patients it has been observed that microRNA-155 is upregulated in synovial tissue and synovial fibroblasts (64). Murata et al. did not observe any difference between the peripheral blood concentration of microRNA-155 of rheumatoid arthritis patients and healthy controls (65). Nonetheless, others have shown that microRNA-155 is upregulated in peripheral blood mononuclear cells (PBMCs) from patients with RA (66, 67). These results show that it is more likely that microRNA-155 concentration in the synovial fluid or in the cells involved in the process of joint inflammation is an indicator of disease, rather than the plasma levels of microRNA-155.
Septic Shock
Septic shock represents the effect of LPS stimulation of the TLR4 receptor on macrophages with a subsequent discharge of proinflammatory molecules leading to severe hypotension and organ dysfunction. Because it is known that LPS stimulation of macrophages lead to M1 polarization, it is highly probable that microRNA-155 also plays a role in this septic shock (68). In vitro, microRNA-155 was observed to be upregulated in THP1 cells when stimulated with LPS (69). In a BALB/c mouse model it has been observed that LPS stimulation increases microRNA-155 expression in the liver. Moreover, this increase in microRNA-155 was observed to be inhibited by pretreatment with dexamethasone, known as a therapeutic approach in septic shock and an indirect inhibitor of microRNA-155 (70).
Multiple Sclerosis
Multiple sclerosis is characterized by the progressive demyelination of axons with subsequent reduction in the functions deserved by them. One important mechanism through which demyelination occurs is through myelin phagocytosis by the local microglia (71). In multiple sclerosis patients it has been shown that microRNA-155 is upregulated in CD14+ monocytes and in microglia of multiple sclerosis patients compared to healthy controls (72). In a multiple sclerosis mouse model it has been observed that the focal lesions contain microRNA-155 upregulation. This, in turn, has been shown to inhibit the expression of CD47 in brain resident cells, leading to an increase in myelin phagocytosis by macrophages (73).
Inflammatory Bowel Disease
Inflammatory bowel diseases are characterized by an inflammatory state of the mucosa and submucosa in the case of ulcerative colitis and of all the layers of the bowel in the case of Crohn’s disease with subsequent functional impairment of these organs. Currently, there is not a precisely known pathogenesis, but inflammation was shown to play an important role and immunosuppressive and antiinflammatory therapies were shown to have an effect on these diseases (74). In a murine model of induced colitis it was shown that microRNA-155 is upregulated (75). Moreover, microRNA-155 was observed to be upregulated in the colonic mucosa of patients with ulcerative colitis (76).
Sarcoidosis
Sarcoidosis is a disease characterized by a systemic granulomatous response of unknown origin. An important microscopic clue to the pathophysiology of the disease is represented by the presence of giant cells, formed by the fusion of multiple macrophages, at the center of the granulomas. This, in turn, shows the possible inflammatory state caused by these cells (77). MicroRNA-155 was observed to be upregulated in progressing sarcoidosis compared to regressing sarcoidosis, but no difference was observed when comparing sarcoidosis patients with healthy controls (78).
Clinical Trials
At the time of this review, 26 clinical trials were found when searching on ClinicalTrials.gov for the term miR155. The search term microRNA-155 yielded fewer results, which were overlapping with the previous search term. Of those, 20 clinical trials include patients with inflammatory diseases and mention using microRNA-155 as a marker for assessing the activity in these diseases. Of those, one trial assessed microRNA-155 in asthma (NCT02719145); none assessed microRNA-155 in patients with atherosclerosis, but there were three trials assessing it for cardiovascular diseases (NCT02605512, NCT02997462, and NCT04277390); two trials assessed microRNA-155 in RA, one as a response to tocilizumab, an anti IL6 antibody (NCT03149796) and one as a response to tofacitinib, a JAK inhibitor (NCT03815578); one trial assessed microRNA-155 dynamics in days 1, 2, 5 and 7 after septic shock onset (NCT02464371); one trial plans to assess the levels of microRNA-155 in the serum of patients with multiple sclerosis (NCT04300543) and no trials plan to evaluate the levels of microRNA-155 in patients with inflammatory bowel disease or sarcoidosis.
Conclusion
MicroRNA-155 represents both a target of proinflammatory signals and an initiator of inflammation with an important impact in M1 polarization. Moreover, it has been repeatedly shown that microRNA-155 upregulation represents an important signal in various inflammatory diseases, with some clinical trials harboring this association as a biomarker for inflammation. As there are no clinical trials using microRNA-155 inhibition as an intervention in inflammatory diseases it can be asked if microRNA-155 will ever be used as more than a biomarker. MicroRNA-155 might have some interventional clinical trials in the future for inflammatory disease, but with the reserve that it might not have a suitable cost/benefit ratio or that other already existing compounds might have a more important effect in modulating inflammation.
Author Contributions
SP, AJ, and BP wrote the manuscript. CT and DM coordinated the team members and made appropriate changes to the final version of the manuscript. All authors agreed on the final version of the manuscript.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1. Lawrence T, Natoli G. Transcriptional regulation of macrophage polarization: enabling diversity with identity. Nat Rev Immunol. (2011) 11:750–61. doi: 10.1038/nri3088
2. O’Brien J, Hayder H, Zayed Y, Peng C. Overview of MicroRNA biogenesis, mechanisms of actions, and circulation. Front Endocrinol. (2018) 9:402. doi: 10.3389/fendo.2018.00402
3. Olson MC, Scott EW, Hack AA, Su GH, Tenen DG, Singh H, et al. PU. 1 is not essential for early myeloid gene expression but is required for terminal myeloid differentiation. Immunity. (1995) 3:703–14. doi: 10.1016/1074-7613(95)90060-8
4. Nerlov C, Graf T. PU.1 induces myeloid lineage commitment in multipotent hematopoietic progenitors. Genes Dev. (1998) 12:2403–12. doi: 10.1101/gad.12.15.2403
5. DeKoter RP. Regulation of B Lymphocyte and macrophage development by graded expression of PU.1. Science. (2000) 288:1439–41. doi: 10.1126/science.288.5470.1439
6. Rosenbauer F, Tenen DG. Transcription factors in myeloid development: balancing differentiation with transformation. Nat Rev Immunol. (2007) 7:105–17. doi: 10.1038/nri2024
7. Darnell J, Kerr I, Stark G. Jak-STAT pathways and transcriptional activation in response to IFNs and other extracellular signaling proteins. Science. (1994) 264:1415–21. doi: 10.1126/science.8197455
8. Gordon S, Taylor PR. Monocyte and macrophage heterogeneity. Nat Rev Immunol. (2005) 5:953–64. doi: 10.1038/nri1733
9. Park C, Li S, Cha E, Schindler C. Immune response in Stat2 knockout mice. Immunity. (2000) 13:795–804. doi: 10.1016/s1074-7613(00)00077-7
10. Krausgruber T, Blazek K, Smallie T, Alzabin S, Lockstone H, Sahgal N, et al. IRF5 promotes inflammatory macrophage polarization and TH1-TH17 responses. Nat Immunol. (2011) 12:231–8. doi: 10.1038/ni.1990
11. Takaoka A, Yanai H, Kondo S, Duncan G, Negishi H, Mizutani T, et al. Integral role of IRF-5 in the gene induction programme activated by Toll-like receptors. Nature. (2005) 434:243–9. doi: 10.1038/nature03308
12. Mosser DM, Edwards JP. Exploring the full spectrum of macrophage activation. Nat Rev Immunol. (2008) 8:958–69. doi: 10.1038/nri2448
13. Takeda K, Tanaka T, Shi W, Matsumoto M, Minami M, Kashiwamura S, et al. Essential role of Stat6 in IL-4 signalling. Nature. (1996) 380:627–30. doi: 10.1038/380627a0
14. Satoh T, Takeuchi O, Vandenbon A, Yasuda K, Tanaka Y, Kumagai Y, et al. The Jmjd3-Irf4 axis regulates M2 macrophage polarization and host responses against helminth infection. Nat Immunol. (2010) 11:936–44. doi: 10.1038/ni.1920
15. Szanto A, Balint BL, Nagy ZS, Barta E, Dezso B, Pap A, et al. STAT6 transcription factor Is a facilitator of the nuclear receptor PPARγ-regulated gene expression in macrophages and dendritic cells. Immunity. (2010) 33:699–712. doi: 10.1016/j.immuni.2010.11.009
16. Ricote M, Li AC, Willson TM, Kelly CJ, Glass CK. The peroxisome proliferator-activated receptor-γ is a negative regulator of macrophage activation. Nature. (1998) 391:79–82. doi: 10.1038/34178
17. Pascual G, Fong AL, Ogawa S, Gamliel A, Li AC, Perissi V, et al. SUMOylation-dependent pathway mediates transrepression of inflammatory response genes by PPAR-γ. Nature. (2005) 437:759–63. doi: 10.1038/nature03988
18. Huang N, Zhang Y, Chen M, Jin H, Nie J, Luo Y, et al. Resveratrol delays 6-hydroxydopamine-induced apoptosis by activating the PI3K/Akt signaling pathway. Exp Gerontol. (2019) 124:110653. doi: 10.1016/j.exger.2019.110653
19. El Kasmi KC, Qualls JE, Pesce JT, Smith AM, Thompson RW, Henao-Tamayo M, et al. Toll-like receptor–induced arginase 1 in macrophages thwarts effective immunity against intracellular pathogens. Nat Immunol. (2008) 9:1399–406. doi: 10.1038/ni.1671
20. Ruffell D, Mourkioti F, Gambardella A, Kirstetter P, Lopez RG, Rosenthal N, et al. A CREB-C/EBP cascade induces M2 macrophage-specific gene expression and promotes muscle injury repair. Proc Natl Acad Sci USA. (2009) 106:17475–80. doi: 10.1073/pnas.0908641106
21. Friedman AD. Transcriptional control of granulocyte and monocyte development. Oncogene. (2007) 26:6816–28. doi: 10.1038/sj.onc.1210764
22. Gordon S, Martinez FO. Alternative activation of macrophages: mechanism and functions. Immunity. (2010) 32:593–604. doi: 10.1016/j.immuni.2010.05.007
23. Ohmori Y, Hamilton TA. IL-4-induced STAT6 suppresses IFN-gamma-stimulated STAT1-dependent transcription in mouse macrophages. J Immunol. (1997) 159:5474–82.
24. De Santa F, Narang V, Yap ZH, Tusi BK, Burgold T, Austenaa L, et al. Jmjd3 contributes to the control of gene expression in LPS-activated macrophages. EMBO J. (2009) 28:3341–52. doi: 10.1038/emboj.2009.271
25. Foster SL, Hargreaves DC, Medzhitov R. Gene-specific control of inflammation by TLR-induced chromatin modifications. Nature. (2007) 447:972–8. doi: 10.1038/nature05836
26. Mann M, Mehta A, Zhao JL, Lee K, Marinov GK, Garcia-Flores Y, et al. An NF-κB-microRNA regulatory network tunes macrophage inflammatory responses. Nat Commun. (2017) 8:851. doi: 10.1038/s41467-017-00972-z
27. Gebert LFR, MacRae IJ. Regulation of microRNA function in animals. Nat Rev Mol Cell Biol. (2019) 20:21–37. doi: 10.1038/s41580-018-0045-7
28. Treiber T, Treiber N, Meister G. Regulation of microRNA biogenesis and its crosstalk with other cellular pathways. Nat Rev Mol Cell Biol. (2019) 20:5–20. doi: 10.1038/s41580-018-0059-1
29. Alivernini S, Gremese E, McSharry C, Tolusso B, Ferraccioli G, McInnes IB, et al. MicroRNA-155—at the critical interface of innate and adaptive immunity in arthritis. Front Immunol. (2018) 8:1932. doi: 10.3389/fimmu.2017.01932
30. Quinn SR, Mangan NE, Caffrey BE, Gantier MP, Williams BRG, Hertzog PJ, et al. The role of Ets2 transcription factor in the induction of MicroRNA-155 (miR-155) by lipopolysaccharide and its targeting by interleukin-10. J Biol Chem. (2014) 289:4316–25. doi: 10.1074/jbc.M113.522730
31. O’Connell RM, Taganov KD, Boldin MP, Cheng G, Baltimore D. MicroRNA-155 is induced during the macrophage inflammatory response. Proc Natl Acad Sci USA. (2007) 104:1604–9. doi: 10.1073/pnas.0610731104
32. Kurowska-Stolarska M, Hasoo MK, Welsh DJ, Stewart L, McIntyre D, Morton BE, et al. The role of microRNA-155/liver X receptor pathway in experimental and idiopathic pulmonary fibrosis. J Allergy Clin Immunol. (2017) 139:1946–56. doi: 10.1016/j.jaci.2016.09.021
33. Bruning U, Cerone L, Neufeld Z, Fitzpatrick SF, Cheong A, Scholz CC, et al. MicroRNA-155 promotes resolution of hypoxia-inducible factor 1 Activity during prolonged hypoxia. Mol Cell Biol. (2011) 31:4087–96. doi: 10.1128/MCB.01276-10
34. Chinenov Y, Coppo M, Gupte R, Sacta MA, Rogatsky I. Glucocorticoid receptor coordinates transcription factor-dominated regulatory network in macrophages. BMC Genomics. (2014) 15:656. doi: 10.1186/1471-2164-15-656
35. Zheng Y, Xiong S, Jiang P, Liu R, Liu X, Qian J, et al. Glucocorticoids inhibit lipopolysaccharide-mediated inflammatory response by downregulating microRNA-155: a novel anti-inflammation mechanism. Free Radic Biol Med. (2012) 52:1307–17. doi: 10.1016/j.freeradbiomed.2012.01.031
36. Elton TS, Selemon H, Elton SM, Parinandi NL. Regulation of the MIR155 host gene in physiological and pathological processes. Gene. (2013) 532:1–12. doi: 10.1016/j.gene.2012.12.009
37. Nazari-Jahantigh M, Wei Y, Noels H, Akhtar S, Zhou Z, Koenen RR, et al. MicroRNA-155 promotes atherosclerosis by repressing Bcl6 in macrophages. J Clin Invest. (2012) 122:4190–202. doi: 10.1172/JCI61716
38. O’Connell RM, Chaudhuri AA, Rao DS, Baltimore D. Inositol phosphatase SHIP1 is a primary target of miR-155. PNAS. (2009) 106:7113–8. doi: 10.1073/pnas.0902636106
39. An H, Xu H, Zhang M, Zhou J, Feng T, Qian C, et al. Src homology 2 domain-containing inositol-5-phosphatase 1 (SHIP1) negatively regulates TLR4-mediated LPS response primarily through a phosphatase activity- and PI-3K-independent mechanism. Blood. (2005) 105:4685–92. doi: 10.1182/blood-2005-01-0191
40. Jing W, Zhang X, Sun W, Hou X, Yao Z, Zhu Y. CRISPR. /CAS9-Mediated Genome Editing of miRNA-155 Inhibits Proinflammatory Cytokine Production by RAW264.7 Cells. BioMed Res Int. (2015) 2015:1–7. doi: 10.1155/2015/326042
41. He M, Xu Z, Ding T, Kuang D-M, Zheng L. MicroRNA-155 regulates inflammatory cytokine production in tumor-associated macrophages via targeting C/EBPβ. Cell Mol Immunol. (2009) 6:343–52. doi: 10.1038/cmi.2009.45
42. Kim H. The transcription factor MafB promotes anti-inflammatory M2 polarization and cholesterol efflux in macrophages. Sci Rep. (2017) 7:7591. doi: 10.1038/s41598-017-07381-8
43. Martinez-Nunez RT, Louafi F, Sanchez-Elsner T. The interleukin 13 (IL-13) pathway in human macrophages is modulated by MicroRNA-155 via direct targeting of interleukin 13 receptor α1 (IL13Rα1). J Biol Chem. (2011) 286:1786–94. doi: 10.1074/jbc.M110.169367
44. Wahl SM, Allen JB, Costa GL, Wong HL, Dasch JR. Reversal of acute and chronic synovial inflammation by anti-transforming growth factor beta. J Exp Med. (1993) 177:225–30. doi: 10.1084/jem.177.1.225
45. Louafi F, Martinez-Nunez RT, Sanchez-Elsner T. MicroRNA-155 targets SMAD2 and modulates the response of macrophages to transforming growth factor-β. J Biol Chem. (2010) 285:41328–36. doi: 10.1074/jbc.M110.146852
46. Filipowicz W, Bhattacharyya SN, Sonenberg N. Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet. (2008) 9:102–14. doi: 10.1038/nrg2290
47. Bushati N, Cohen SM. microRNA Functions. Annu Rev Cell Dev Biol. (2007) 23:175–205. doi: 10.1146/annurev.cellbio.23.090506.123406
48. Tarassishin L, Loudig O, Bauman A, Shafit-Zagardo B, Suh H-S, Lee SC. Interferon regulatory factor 3 inhibits astrocyte inflammatory gene expression through suppression of the proinflammatory miR-155 and miR-155∗. Glia. (2011) 59:1911–22. doi: 10.1002/glia.21233
49. Zhou H, Huang X, Cui H, Luo X, Tang Y, Chen S, et al. miR-155 and its star-form partner miR-155∗ cooperatively regulate type I interferon production by human plasmacytoid dendritic cells. Blood. (2010) 116:5885–94. doi: 10.1182/blood-2010-04-280156
50. Feketea G, Bocsan CI, Popescu C, Gaman M, Stanciu LA, Zdrenghea MT. A. Review of macrophage MicroRNAs’. Role Hum Asthma Cells. (2019) 8:420. doi: 10.3390/cells8050420
51. Li H, Jiang T, Li M-Q, Zheng X-L, Zhao G-J. Transcriptional regulation of macrophages polarization by MicroRNAs. Front Immunol. (2018) 9:1175. doi: 10.3389/fimmu.2018.01175
52. Comer BS, Camoretti-Mercado B, Kogut PC, Halayko AJ, Solway J, Gerthoffer WT. Cyclooxygenase-2 and MicroRNA-155 expression are elevated in asthmatic airway smooth muscle cells. Am J Respir Cell Mol Biol. (2015) 52:438–47. doi: 10.1165/rcmb.2014-0129OC
53. Suojalehto H, Lindström I, Majuri M-L, Mitts C, Karjalainen J, Wolff H, et al. Altered MicroRNA expression of nasal mucosa in long-term asthma and allergic rhinitis. Int Arch Allergy Immunol. (2014) 163:168–78. doi: 10.1159/000358486
54. Pinkerton M, Chinchilli V, Banta E, Craig T, August A, Bascom R, et al. Differential expression of microRNAs in exhaled breath condensates of patients with asthma, patients with chronic obstructive pulmonary disease, and healthy adults. J Allergy Clin Immunol. (2013) 132:217–9.e2. doi: 10.1016/j.jaci.2013.03.006
55. Malmhäll C, Johansson K, Winkler C, Alawieh S, Ekerljung L, Rådinger M. Altered miR-155 expression in allergic asthmatic airways. Scand J Immunol. (2017) 85:300–7. doi: 10.1111/sji.12535
56. Qiu L, Zhang Y, Do DC, Ke X, Zhang S, Lambert K, et al. miR-155 modulates cockroach allergen– and oxidative stress–induced cyclooxygenase-2 in asthma. J Immunol. (2018) 201:916–29. doi: 10.4049/jimmunol.1701167
57. Chen H, Xu X, Cheng S, Xu Y, Xuefei Q, Cao Y, et al. Small interfering RNA directed against microRNA-155 delivered by a lentiviral vector attenuates asthmatic features in a mouse model of allergic asthma. Exp Ther Med. (2017) 14:4391–6. doi: 10.3892/etm.2017.5093
58. Libby P, Buring JE, Badimon L, Hansson GK, Deanfield J, Bittencourt MS, et al. Atherosclerosis. Nat Rev Dis Primers. (2019) 5:56. doi: 10.1038/s41572-019-0106-z
59. Du F, Yu F, Wang Y, Hui Y, Carnevale K, Fu M, et al. MicroRNA-155 deficiency results in decreased macrophage inflammation and attenuated atherogenesis in apolipoprotein E–deficient mice. Arterioscler Thromb Vasc Biol. (2014) 34:759–67. doi: 10.1161/ATVBAHA.113.302701
60. Yang Y, Yang L, Liang X, Zhu G. MicroRNA-155 promotes atherosclerosis inflammation via targeting SOCS1. Cell Physiol Biochem. (2015) 36:1371–81. doi: 10.1159/000430303
61. Ye J, Guo R, Shi Y, Qi F, Guo C, Yang L. miR-155 regulated inflammation response by the SOCS1-STAT3-PDCD4 axis in atherogenesis. Mediators Inflamm. (2016) 2016: 8060182. doi: 10.1155/2016/8060182
62. Donners MMPC, Wolfs IMJ, Stöger LJ, van der Vorst EPC, Pöttgens CCH, Heymans S, et al. Hematopoietic miR155 deficiency enhances atherosclerosis and decreases plaque stability in hyperlipidemic mice. PLoS One. (2012) 7:e35877. doi: 10.1371/journal.pone.0035877
63. Aletaha D, Smolen JS. Diagnosis and management of rheumatoid arthritis: a review. JAMA. (2018) 320:1360–72. doi: 10.1001/jama.2018.13103
64. Stanczyk J, Pedrioli DML, Brentano F, Sanchez-Pernaute O, Kolling C, Gay RE, et al. Altered expression of MicroRNA in synovial fibroblasts and synovial tissue in rheumatoid arthritis. Arthritis Rheum. (2008) 58:1001–9. doi: 10.1002/art.23386
65. Murata K, Yoshitomi H, Tanida S, Ishikawa M, Nishitani K, Ito H, et al. Plasma and synovial fluid microRNAs as potential biomarkers of rheumatoid arthritis and osteoarthritis. Arthritis Res Ther. (2010) 12:R86. doi: 10.1186/ar3013
66. Elmesmari A, Fraser AR, Wood C, Gilchrist D, Vaughan D, Stewart L, et al. MicroRNA-155 regulates monocyte chemokine and chemokine receptor expression in Rheumatoid Arthritis. Rheumatology. (2016) 55:2056–65. doi: 10.1093/rheumatology/kew272
67. Pauley KM, Satoh M, Chan AL, Bubb MR, Reeves WH, Chan EK. Upregulated miR-146a expression in peripheral blood mononuclear cells from rheumatoid arthritis patients. Arthritis Res Ther. (2008) 10:R101. doi: 10.1186/ar2493
68. Wentowski C, Mewada N, Nielsen ND. Sepsis in 2018: a review. Anaesth Intensive Care Med. (2019) 20:6–13. doi: 10.1016/j.mpaic.2018.11.009
69. Taganov KD, Boldin MP, Chang K-J, Baltimore D. NF- B-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc Natl Acad Sci. (2006) 103:12481–6. doi: 10.1073/pnas.0605298103
70. Tili E, Michaille J-J, Cimino A, Costinean S, Dumitru CD, Adair B, et al. Modulation of miR-155 and miR-125b Levels following Lipopolysaccharide/TNF-α stimulation and their possible roles in regulating the response to endotoxin shock. J Immunol. (2007) 179:5082–9. doi: 10.4049/jimmunol.179.8.5082
71. Dobson R, Giovannoni G. Multiple sclerosis – a review. Eur J Neurol. (2019) 26:27–40. doi: 10.1111/ene.13819
72. Moore CS, Rao VTS, Durafourt BA, Bedell BJ, Ludwin SK, Bar-Or A, et al. miR-155 as a multiple sclerosis-relevant regulator of myeloid cell polarization: miR-155 and Cell Polarization. Ann Neurol. (2013) 74:709–20. doi: 10.1002/ana.23967
73. Junker A, Krumbholz M, Eisele S, Mohan H, Augstein F, Bittner R, et al. MicroRNA profiling of multiple sclerosis lesions identifies modulators of the regulatory protein CD47. Brain. (2009) 132:3342–52. doi: 10.1093/brain/awp300
74. Digby-Bell JL, Atreya R, Monteleone G, Powell N. Interrogating host immunity to predict treatment response in inflammatory bowel disease. Nat Rev Gastroenterol Hepatol. (2020) 17:9–20. doi: 10.1038/s41575-019-0228-5
75. Lu Z-J, Wu J-J, Jiang W-L, Xiao J-H, Tao K-Z, Ma L, et al. MicroRNA-155 promotes the pathogenesis of experimental colitis by repressing SHIP-1 expression. World J Gastroenterol. (2017) 23:976–85. doi: 10.3748/wjg.v23.i6.976
76. Takagi T, Naito Y, Mizushima K, Hirata I, Yagi N, Tomatsuri N, et al. Increased expression of microRNA in the inflamed colonic mucosa of patients with active ulcerative colitis: MicroRNA in ulcerative colitis. J Gastroenterol Hepatol. (2010) 25:S129–33. doi: 10.1111/j.1440-1746.2009.06216.x
77. Bargagli E, Prasse A. Sarcoidosis: a review for the internist. Intern Emerg Med. (2018) 13:325–31. doi: 10.1007/s11739-017-1778-6
78. Dyskova T, Fillerova R, Novosad T, Kudelka M, Zurkova M, Gajdos P, et al. Correlation network analysis reveals relationships between MicroRNAs, transcription factor T-bet, and deregulated cytokine/chemokine-receptor network in pulmonary sarcoidosis. Mediators Inflamm. (2015) 2015:1–16. doi: 10.1155/2015/121378
Keywords: macrophage, polarization, microRNA-155, inflammation, M1
Citation: Pasca S, Jurj A, Petrushev B, Tomuleasa C and Matei D (2020) MicroRNA-155 Implication in M1 Polarization and the Impact in Inflammatory Diseases. Front. Immunol. 11:625. doi: 10.3389/fimmu.2020.00625
Received: 29 January 2020; Accepted: 19 March 2020;
Published: 15 April 2020.
Edited by:
Luminita Aurelia Stanciu, Imperial College London, United KingdomReviewed by:
Zhanjun Guy, Fourth Hospital of Hebei Medical University, ChinaSahana Holla, National Cancer Institute (NCI), United States
Copyright © 2020 Pasca, Jurj, Petrushev, Tomuleasa and Matei. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Sergiu Pasca, cGFzY2Euc2VyZ2l1MTIzQGdtYWlsLmNvbQ==
 Sergiu Pasca
Sergiu Pasca Ancuta Jurj2
Ancuta Jurj2 Daniela Matei
Daniela Matei