- Stomatological Hospital, School of Stomatology, Southern Medical University, Guangzhou, China
Oral squamous cell carcinoma (OSCC) remains a formidable malignancy with persistently poor clinical outcomes. Recent research has underscored the pivotal role of the innate immune system, particularly tumor-associated macrophages (TAMs), a key component of the myeloid lineage, in orchestrating the tumor microenvironment (TME) and shaping disease progression. As professional phagocytes of the innate immune system, macrophages not only mediate pathogen recognition and inflammatory responses but also undergo functional polarization in response to local cues. In OSCC, dysbiosis of the oral microbiota, marked by the overrepresentation of species such as Fusobacterium nucleatum and Porphyromonas gingivalis—acts as a chronic inflammatory trigger that promotes epithelial-mesenchymal transition (EMT), immune evasion, and tumor growth. These pathogenic bacteria actively engage innate immune signaling pathways such as TLRs and CSF-1R, skewing macrophages toward an immunosuppressive M2 phenotype. M2-like TAMs then contribute to tumor progression by secreting anti-inflammatory cytokines (IL-10, TGF-β), promoting angiogenesis, and expressing immune checkpoint ligands such as PD-L1. This review summarizes current knowledge on the bidirectional crosstalk between dysbiotic microbiota and innate immune macrophages in OSCC, highlighting key receptor-mediated pathways and their implications for immune suppression, metastasis, and therapy resistance. Targeting microbiota modulation or innate immune reprogramming represents a promising strategy for restoring anti-tumor immunity and enhancing therapeutic efficacy in OSCC.
1 Introduction
Oral squamous cell carcinoma (OSCC) accounts for 90% of oral malignancies and remains the predominant head and neck cancer subtype (1–3). Despite surgical resection being the standard of care, therapeutic options are limited, and the global 5-year survival rate remains below 50% (4). Thus, further research into the etiopathogenesis and mechanisms driving OSCC is essential. Since the WHO classification of Helicobacter pylori as a carcinogen, interest in bacteria–cancer associations has intensified, revealing diverse molecular mechanisms (5). The oral cavity harbors over 700 bacterial species, making it one of the most complex microbial ecosystems in the human body (6, 7). Although the link between oral microbiota and OSCC is debated, prevailing hypotheses posit that pathogenic bacteria directly promote oncogenesis or that oral dysbiosis accelerates tumor progression (8).
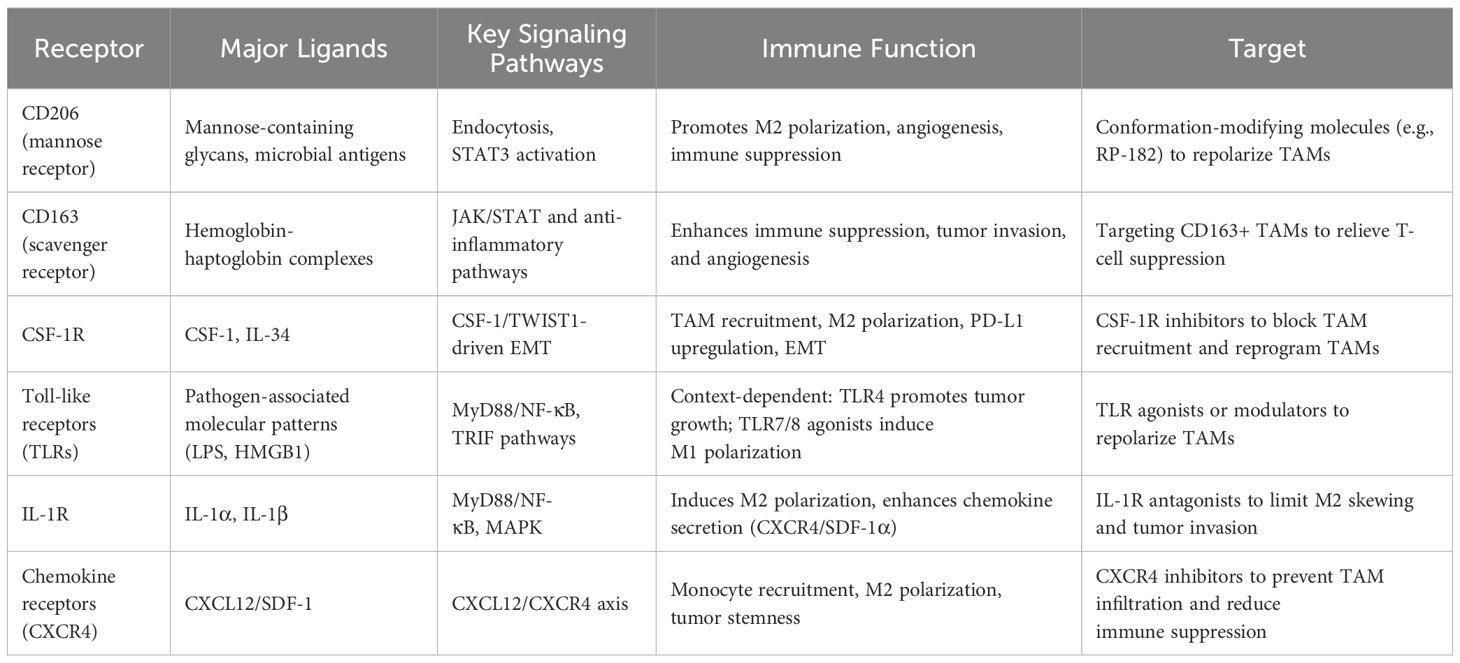
Beyond microbial influences, immune regulation is pivotal. Macrophages—key effectors of innate immunity—mediate pathogen clearance, modulate inflammation, and orchestrate tissue repair (9). Tumor-associated macrophages (TAMs), a dominant immune population within the OSCC stroma, critically shape the tumor microenvironment (TME) (10). TAMs display functional plasticity: the pro-inflammatory, anti-tumor M1 phenotype contrasts with the TME-induced M2 phenotype, which fosters invasion, metastasis, and immunosuppression (11, 12). In OSCC, TAMs express receptors such as CD206, CD163, and Toll-like receptors, as well as cytokine and chemokine receptors including IL-1R, CSF-1R, and CCR family members, which serve as biomarkers for TAM quantification and tumor dissemination (13, 14). This review summarizes current evidence on OSCC-associated oral microbiota shifts, TAM–tumor cell interactions mediated by surface receptors, and the emerging role of microbiota in modulating macrophage function to drive OSCC pathogenesis.
2 Microorganisms in OSCC
In OSCC, microbial populations vary significantly across tumor stages and oral sites. YANG et al. (8) analyzed microbial composition in adjacent normal tissues, tumor tissues, and saliva of OSCC patients, finding Streptococcus and Peptostreptococcus species most abundant in tumor tissues, while Neisseria species were prevalent in saliva. Similarly, SARKAR et al. (15) observed that Prevotella and Fusobacterium species were more abundant in tumor tissues compared to normal tissues. Li et al. (16) reported significant microbial differences between healthy individuals and OSCC patients, with Porphyromonas and Peptostreptococcus species enriched in tumor samples. Notably, Porphyromonas abundance correlated with elevated C-reactive protein levels in OSCC patients. Additionally, oral microbiota correlates with clinical tumor staging: Treponema species are abundant in early-stage tumors, while Moraxella species dominate advanced-stage tumor tissues (8). Periodontal disease is a recognized OSCC risk factor (17), linking oral microbiota-induced inflammation to cancer development. ZHANG et al. (18) confirmed increased Fusobacterium nucleatum abundance in OSCC, with F. nucleatum showing progressive enrichment from normal tissues to cancerous tissues (8, 19). Porphyromonas gingivalis is also associated with OSCC (20, 21), and higher abundance of Fusobacterium, Peptostreptococcus, and Prevotella species is found in tumor tissues compared to gingival squamous cell carcinoma and periodontal disease (22). Beyond these well-characterized taxa, other microorganisms, including Treponema denticola and Campylobacter rectus, have also been detected at increased abundance in the oral cavity of OSCC patients and are thought to contribute to chronic inflammation and immune modulation, potentially influencing tumor initiation and progression.
In oral cancer, microorganisms induce inflammation by producing cytokines and chemokines, promoting tumor cell proliferation and epithelial-mesenchymal transition (EMT), which enhances migration and invasion abilities (20, 23). Chronic infections with F. nucleatum and P. gingivalis increase the severity of tongue tumors, with experimental tumors in mice being approximately 2.5 times larger than controls (23, 24). F. nucleatum promotes OSCC progression and is associated with poor prognosis, enhancing tumor cell proliferation, migration, and immune regulation in the TME (23, 25, 26). It regulates EMT through the lncRNA MIR4435-2HG/miR-296-5p/Akt2/SNAI1 signaling pathway (27), and activates STAT3 in CRC to upregulate EMT genes (28). P. gingivalis activates ERK1/2-Ets1, p38/HSP27, and PAR2/NF-κB pathways, promoting matrix metalloproteinase (MMP) expression and tumor invasiveness (29). Both F. nucleatum and P. gingivalis activate the TLR/MyD88-triggered integrin/FAK pathway, enhancing tumor invasiveness (30). Microorganisms play a critical role in exacerbating oral diseases and potentially in the development and progression of oral cancer (31).
3 Macrophages in OSCC
3.1 Role of macrophages in tumors
TME comprises diverse non-malignant components, including fibroblasts and immune cells, that collectively shape tumor biology (32, 33). TAMs, through the secretion of cytokines, chemokines, and growth factors, exert profound influence on tumor initiation, progression, and metastasis (34). Exhibiting notable phenotypic plasticity, macrophages polarize along a continuum from classically activated M1 to alternatively activated M2 states (35–37). M2 macrophages are subclassified into M2a (IL-4/IL-13–induced), M2b (immune complex/LPS–induced), M2c (glucocorticoid/IL-10–induced), and M2d (IL-6/adenosine–induced), each performing specialized functions in OSCC (38, 39). M2a macrophages promote wound healing and extracellular matrix (ECM) deposition, facilitating tissue remodeling (40, 41). M2b cells generate both pro- and anti-inflammatory cytokines, establishing an immunoregulatory milieu (42, 43). M2c macrophages mediate potent immunosuppression via IL-10 and TGF-β secretion, supporting tumor survival (44). Importantly, M2d macrophages—also referred to as tumor-associated macrophages with an “angiogenic” phenotype—are particularly relevant to OSCC because they release high levels of vascular endothelial growth factor (VEGF), platelet-derived growth factor (PDGF), and matrix metalloproteinases (MMP−9 and MMP−2) (45–47). By contrast, M1 macrophages, though less well classified and capable of subtype interconversion, dominate early tumor stages, releasing TNF-α, CXCL9, CXCL10, iNOS, and ROS to induce inflammation and eliminate tumor cells (48–51). M2 phenotypes predominate in advanced disease, sustaining chronic inflammation, ECM remodeling, migration, invasion, and neovascularization via PDGF, TGF-β, and related mediators (34, 52). Among these, M2a, M2c, and M2d subtypes are particularly enriched in TAM populations, functioning as principal drivers of OSCC progression (53, 54).
3.2 TAM surface receptors in OSCC progression
3.2.1 Human mannose receptor CD206 and CD163
Human mannose receptor CD206 is an endocytic transmembrane protein containing multiple carbohydrate-binding domains. Its ligands can include bacterial products, tumor metabolites, or other synthetic proteins (55). Structural modulation of CD206 using the synthetic ligand RP-182 suppresses tumor growth, prolongs survival, and reprograms M2 macrophages toward an M1 phenotype through endocytosis, phagolysosome formation, and autophagy (56). In OSCC, plasminogen activator inhibitor-1 (PAI-1) and IL-8 inhibit monocyte differentiation into CD206+ TAMs (57). CD206+ TAMs secrete higher epidermal growth factor (EGF) levels than CD163+ or CD204+ TAMs, enhancing OSCC proliferation and invasion via EGF receptor signaling, an effect abrogated by EGFR blockade (58). Through STAT3 activation, CD206+ M2 TAMs sustain a pro-tumor milieu, releasing VEGF, TGF-β, EGF, urokinase-type plasminogen activator (uPA), and multiple matrix metalloproteinases (MMPs), thereby promoting tumor growth, immune suppression, angiogenesis, metastasis, and chemoresistance (59). CD163, widely applied as an immunohistochemical TAM marker, is associated with tumor infiltration and invasion, with expression levels correlating with tumor stage, nodal involvement, and metastasis (60). Functionally, CD163+ TAMs foster tumor immune evasion; their depletion in melanoma models increases cytotoxic T cell infiltration and inflammatory monocyte recruitment, markedly enhancing tumor regression (61). In OSCC, co-expression of CD163 and CD204 in M2 TAMs coincides with IL-10 secretion and PD-L1 expression, potentially suppressing T cell function and facilitating invasion and metastasis (62). Moreover, PFKFB3 expression positively correlates with CD163 levels in OSCC, implicating this glycolytic regulator in TAM-mediated angiogenesis (63). Targeting CD163 may thus attenuate tumor-induced immunosuppression.
3.2.2 The role of colony-stimulating Factor 1 receptor
Colony-stimulating factor receptors (CSFR) are essential for the differentiation of myeloid stem cells into monocytes and macrophages. Chemoresistant colorectal cancer cells release CSF, which binds to CSFR, recruiting TAMs and upregulating PD-L1, leading to chemotherapy resistance and poor prognosis (64, 65). Pharmacological inhibition of CSF1R has been shown to reprogram TAMs from an M2-like immunosuppressive phenotype toward an M1-like pro-inflammatory state, thereby restoring antitumor immunity (66). A large-scale OSCC study found that CSF-1 levels correlated with TAM infiltration, and CSF-1 signaling blockade with BL2945 not only inhibited OSCC growth but also significantly reduced CSF-1 expression and TAM infiltration (67). These findings suggest that CSF-1, abundantly produced by OSCC cells and surrounding stromal components, binds chemotactically to CSF1R on TAMs, thereby fueling tumor cell proliferation. Moreover, in oral cancer, CSF-1 co-expresses with the transcriptional activator TWIST1 of epithelial-mesenchymal transition (EMT), accompanied by TAM infiltration, suggesting that CSF-1 induces TAM chemotaxis through CSFR activation, enhancing OSCC’s epithelial-mesenchymal transition and invasion (68). Therefore, targeting the CSF-1 receptor on TAMs represents an important strategy for combating tumor growth (Table 1).
3.2.3 The role of toll-like receptors
Toll-like receptors (TLRs), including TLR2, TLR4, TLR7, TLR8, and TLR9, are critical pattern recognition receptors on TAMs, with diverse ligands such as tumor proteins, acute-phase proteins, drugs, and bacterial metabolites. TLR4 signaling promotes tumor growth, while TLR7, TLR8, and TLR9 signaling may exert anti-tumor effects (69). Ferroptosis, a regulated cell death driven by lipid peroxidation, serves as a tumor-suppressive mechanism (70). Tumor cells undergoing ferroptosis release HMGB1, which binds to TLR4 on myeloid cells, enhancing differentiation into mature cells (71). Activation of TLR4/NF-κB signaling through carbon nanotubes induces M2-polarized TAMs to shift to M1, inhibiting metastasis in mouse models (72). Alpha 1-acid glycoprotein (AGP) activates TLR4 through CD14, modulating PD-L1 expression and IL-6 production, thus enhancing tumor immune suppression (73). Ginseng-derived nanoparticles (GDNPs) induce M2 to M1 polarization via TLR4/MyD88, promoting apoptosis in head and neck squamous cell carcinoma (74). In colorectal cancer, cathepsin K secretion disrupts the gut microbiome and activates TLR4 to promote M2 macrophages, accelerating progression (75). OSCC may follow similar mechanisms. Overexpression of ALDH3A1 in Cal27 cells inhibits IL-6 and suppresses TLR4 activation in TAMs, reducing inflammation (76). Oral administration of Dendrobium officinale polysaccharide (DOP) promotes M1 polarization via TLR2 on TAMs, inhibiting tumor growth (77). TLR7 and TLR8 agonists reverse TAMs from M2 to M1, reducing radiation resistance and tumor growth (78, 79). Immune-modulating agents delivered via polymer micelles targeting TLR7 reduce immune suppression in the TME (80).
3.2.4 Interleukin-1 and chemokine receptors
The interleukin-1 receptor (IL-1R) is activated by its canonical ligands, IL-1α and IL-1β, whereas the interleukin-1 receptor antagonist competitively inhibits this signaling. Ligand binding to IL-1R predominantly triggers the MyD88–NF-κB and mitogen-activated protein kinase (MAPK) pathways. Accumulating evidence indicates that the IL-1R/MyD88 signaling axis regulates programmed death-1 (PD-1) expression, thereby sustaining the immunosuppressive activity of TAMs and facilitating tumor progression in melanoma and other malignancies (81, 82). OSCC cells release IL-1β to recruit monocytes, and IL-1R-mediated TAM conversion to the M2 phenotype imparts immune-suppressive capabilities (83). Following IL-1β-induced M2 polarization, TAMs continue to secrete IL-1β. OSCC and TAMs induce the expression of CXCR4 or stromal cell-derived factor-1 alpha (SDF-1α) through IL-1β/IL-1R signaling and hypoxia-induced activation of ERK signaling, thus promoting the expression of MMP-9 and MMP-13, which facilitate OSCC migration and invasion (84). Therefore, IL-1R modulation in TAMs could be a promising target for cancer therapy. Chemokine receptors are part of the seven-transmembrane G-protein-coupled receptor family, with many receptors present on TAMs. Tumor-associated fibroblasts (CAFs) in the microenvironment attract monocytes via CXCL12/CXCR4 signaling, which induces the polarization of TAMs to the M2 phenotype. These polarized TAMs contribute to the formation of tumor stem cells in OSCC, promoting proliferation, reducing apoptosis, and enhancing migration (85). This receptor family, with its large array of ligands, presents considerable research potential and could become a therapeutic target in TAM-based OSCC treatment.
3.2.5 The role of other receptors
TAMs express numerous surface receptors, representing an area of active research for receptor-based therapies. Tumor-associated endothelial cells aberrantly express and secrete HSPA12B, which can be partially engulfed by macrophages through the oxidized low-density lipoprotein receptor 1 (OLR1), triggering PI3K/Akt/mTOR signaling and increasing the expression of M2 markers (86, 87). Immunohistochemical staining has revealed that TAMs in endogenous OSCC express more patched-1 (PTCH) receptors than in exogenous OSCC, with sonic hedgehog (SHH) ligands inducing tumor invasion through autocrine signaling and regulating OSCC stroma-parenchyma interactions via paracrine signaling (88). In addition, TAMs obtain CD73 from exosomes secreted by head and neck squamous carcinoma cells and express it on their cell membrane. This exosome-mediated transfer enhances immune evasion and tumor invasiveness through activation of CD73–NF-κB p65 signaling, and promotes the secretion of immunosuppressive mediators such as PD-1, PD-L1, and pro-inflammatory cytokines (89). Furthermore, microbial-derived tryptophan metabolites have been shown to activate the aryl hydrocarbon receptor on TAMs, thereby amplifying their immunosuppressive functions and further attenuating antitumor immunity (90).
4 Microbial and macrophage crosstalk in the development of OSCC
4.1 Crosstalk between oral microbiota and macrophages
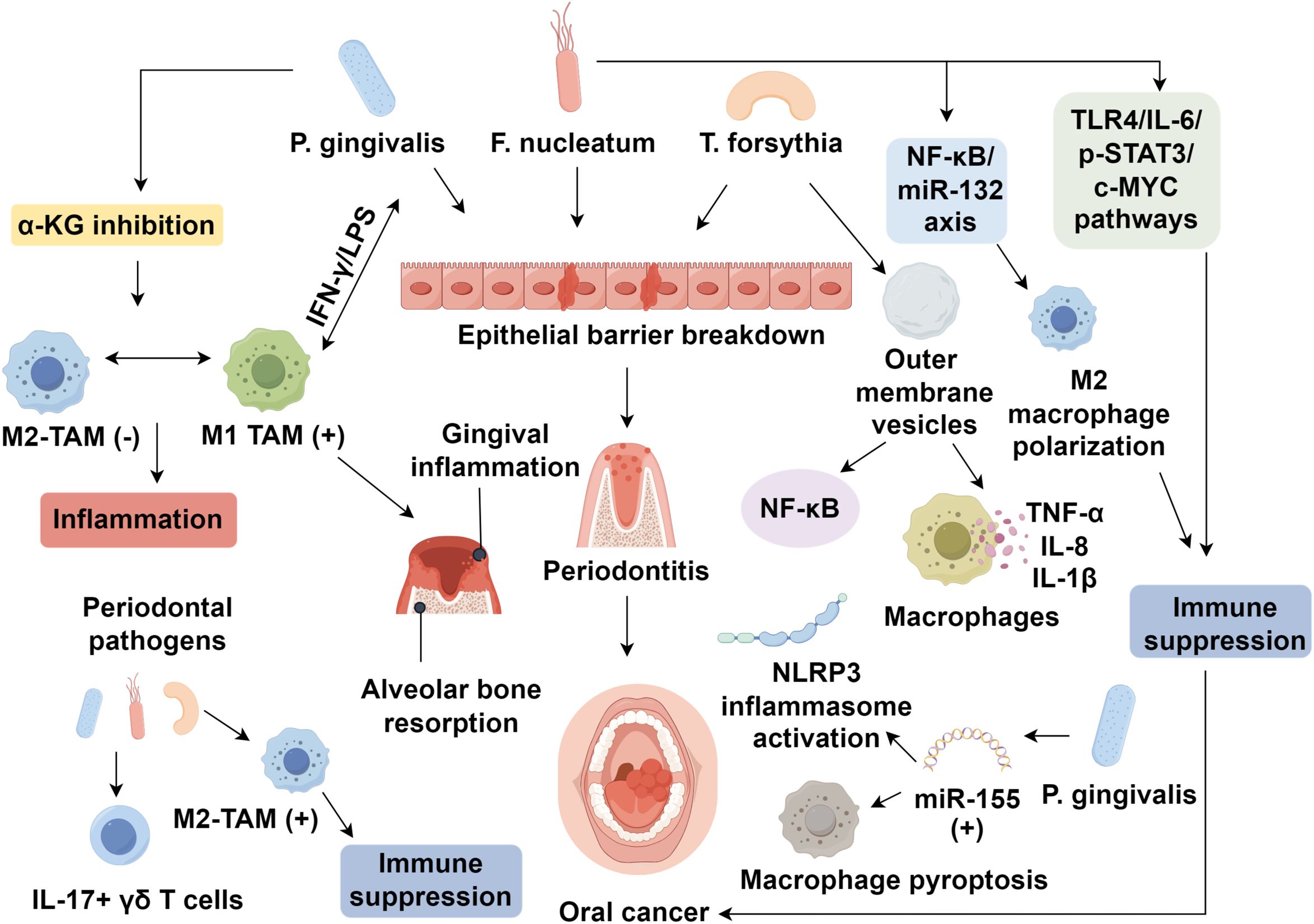
The balance between oral microbiota, immune cells, and the epithelial barrier is crucial for maintaining oral microbiome stability. Disruption of this balance, caused by pathogenic microbes, can lead to inflammatory responses, mucosal barrier breakdown, and oral diseases, such as periodontitis, eventually promoting oral cancer. Periodontitis, the sixth most prevalent global disease (91), is influenced by bacterial infections and immune dysregulation. Key bacteria, including P. gingivalis, F. nucleatum, and T. forsythia, have been extensively studied in periodontitis models (92), with macrophages playing a significant role in disease progression. Macrophages polarize into M1 macrophages in response to periodontal pathogens, contributing to the inflammatory environment (93). P. gingivalis inhibits α-ketoglutarate (α-KG) expression, suppressing M2 macrophage generation and inducing M1 macrophage polarization to maintain inflammation (94). It drives M1 macrophage infiltration into deep periodontal tissues, promoting gingival inflammation and alveolar bone resorption (94, 95). Additionally, macrophages interacting with oral commensal bacteria, such as through IFN-γ/LPS stimulation, can enhance pathogen survival and disease progression (96). In the tumor microenvironment, oral microbiota promotes M2 polarization of macrophages. Periodontal pathogens activate IL-17+ γδ T cells and increase M2-type TAM infiltration in OSCC (97). P. gingivalis has also been shown to protect OSCC cells from macrophage phagocytosis and induce M2 macrophage polarization (98). Furthermore, T. forsythia produces outer membrane vesicles that activate NF-κB and upregulate TNF-α, IL-8, and IL-1β in macrophages (99), while P. gingivalis upregulates miR-155, promoting NLRP3 inflammasome activation and macrophage pyroptosis (100). Other less frequently discussed species, such as Treponema denticola and Campylobacter rectus, have also been reported to modulate immune responses in periodontal disease and may contribute to macrophage activation and polarization in the OSCC microenvironment, though their specific mechanisms remain less well characterized.
4.2 Impact of microbial and macrophage regulation on OSCC development
The imbalance between the oral mucosa, microbiota, and immune system leads to oral diseases. In germ-free mice, intestinal mucosal atrophy and reduced macrophage numbers recover after microbiota colonization, with full recovery by five weeks (101). In dysbiotic mice, macrophages release IL-6, promoting colorectal cancer proliferation and EMT, with macrophage depletion reversing tumor-promoting effects (102). Co-culturing Faecalibacterium prausnitzii with macrophages induces morphological changes in intestinal epithelial cells, highlighting microbiota-immune-epithelial regulation (103). Dysbiosis weakens the mucosal barrier, facilitating bacterial translocation and chronic inflammation, which leads to DNA damage, cytokine/chemokine production, and tumor proliferation, migration, and apoptosis inhibition (104). Fusobacterium nucleatum (F. nucleatum) plays a crucial role in OSCC, promoting M2-type TAM aggregation and immune suppression by inhibiting macrophages and T cells (105). F. nucleatum disrupts epithelial tight junctions and induces immune suppression via indoleamine 2,3-dioxygenase (IDO) upregulation (106). It activates the NF-κB/miR-132 axis, promoting M2 macrophage polarization and CRC metastasis, as well as TLR4/IL-6/p-STAT3/c-MYC pathways to enhance immunosuppressive effects (107–110).
P. gingivalis also induces M2 polarization, aiding OSCC immune evasion (98). Beyond direct effects on macrophages, these dysbiotic microbes and M2-polarized TAMs also profoundly shape the wider immune landscape of the OSCC tumor microenvironment. TAM-derived IL-10, TGF-β, and chemokines drive the recruitment and expansion of regulatory T cells (Tregs), which further suppress effector T cell function. Similarly, TAM-secreted factors such as CCL2 and CSF-1 promote the accumulation of myeloid-derived suppressor cells (MDSCs), reinforcing an immunosuppressive milieu. In parallel, persistent inflammatory signaling and PD-L1 expression by TAMs induce functional exhaustion of CD8+ cytotoxic T lymphocytes, characterized by upregulation of inhibitory receptors (PD-1, TIM-3, TIGIT) and diminished cytotoxicity. These interconnected suppressive networks, orchestrated by microbial signals and macrophage-derived mediators, contribute to a deeply immune-evasive tumor niche that fosters OSCC progression and resistance to therapy. Microbial-infected macrophages play a pivotal role in tumor development, providing insights for early oral cancer diagnosis and treatment (Figure 1). Recent clinical investigations have shown that the presence of dysbiotic microbial communities, particularly Fusobacterium nucleatum and Porphyromonas gingivalis, correlates with higher tumor stage, lymph node metastasis, and poorer survival outcomes in OSCC patients (111, 112). Elevated infiltration of M2-polarized TAMs in biopsy specimens is consistently associated with immune evasion, advanced stage disease, and resistance to immune checkpoint inhibitors (113, 114). These findings underscore the potential for using the oral microbiota composition and TAM-related markers as prognostic biomarkers, which also provide a rationale for integrating microbiota modulation and TAM-targeting therapies into clinical strategies to improve immunotherapy efficacy and survival in OSCC patients.
5 Conclusion
The intricate interplay between dysbiotic oral microbiota and TAMs—innate immune sentinels—shapes a profoundly immunosuppressive tumor microenvironment in OSCC. Pathogenic bacteria such as Fusobacterium nucleatum and Porphyromonas gingivalis promote chronic inflammation, epithelial-mesenchymal transition, and M2 polarization of macrophages via pattern recognition receptors and cytokine signaling. These M2-like TAMs, in turn, sustain immune evasion, angiogenesis, and tumor progression.
Currently, clinical interventions that directly target the oral microbiota–immune axis in OSCC remain limited. Approaches such as probiotics or oral microbiome modulation are under early investigation for improving periodontal health and reducing chronic inflammation, but robust evidence in OSCC prevention or treatment is lacking. Similarly, antibiotics have been explored for altering microbial communities, yet their nonspecific effects and potential to disrupt beneficial bacteria limit their clinical utility. Fecal microbiota transplantation has shown promise in gastrointestinal cancers for modulating systemic immunity, but its application in head and neck cancers, including OSCC, remains largely unexplored and without clinical validation.
These gaps underscore the need for carefully designed trials to evaluate whether microbiome-focused interventions can complement immunotherapy or standard treatments in OSCC. Future therapeutic strategies should prioritize targeting this microbiome–innate immunity axis. Interventions that reprogram TAMs toward a pro-inflammatory M1 phenotype or modulate microbial composition may restore antitumor immunity. In-depth mechanistic studies and clinical validation of dual-targeted therapies, focusing on innate immune modulation and microbiota regulation, could pave the way for precision immunotherapy in OSCC.
Author contributions
XD: Writing – original draft. SH: Writing – original draft, Writing – review & editing.
Funding
The author(s) declare financial support was received for the research and/or publication of this article. This work was supported by the Science Research Cultivation Program of Stomatological Hospital, Southern Medical University (Grant No. PY2023010), which was provided to support the postdoc research of the first and senior author Dr. Xin Deng.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Generative AI statement
The authors declare that no Generative AI was used in the creation of this manuscript.
Any alternative text (alt text) provided alongside figures in this article has been generated by Frontiers with the support of artificial intelligence and reasonable efforts have been made to ensure accuracy, including review by the authors wherever possible. If you identify any issues, please contact us.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
References
1. Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. (2021) 71:209–49. doi: 10.3322/caac.21660
2. Sur S, Davray D, Basu S, Kheur S, Pal JK, Nagar S, et al. Novel insights on oral squamous cell carcinoma management using long non-coding RNAs. Oncol Res. (2024) 32:1589–612. doi: 10.32604/or.2024.052120
3. Shan F, Liang L, Feng C, Xu H, Wang Z, Liu W, et al. LAMC2 regulates proliferation, migration, and invasion mediated by the Pl3K/AKT/mTOR pathway in oral. Oncol Res. (2023) 31:481–93. doi: 10.32604/or.2023.029064
4. Sklenicka S, Gardiner S, Dierks EJ, Potter BE, and Bell RB. Survival analysis and risk factors for recurrence in oral squamous cell carcinoma: does surgical salvage affect outcome? J Oral Maxillofac Surg. (2010) 68:1270–5. doi: 10.1016/j.joms.2009.11.016
5. Khatoon J, Rai RP, and Prasad KN. Role of Helicobacter pylori in gastric cancer: Updates. World J Gastrointest Oncol. (2016) 8:147–58. doi: 10.4251/wjgo.v8.i2.147
6. Dewhirst FE, Chen T, Izard J, Paster BJ, Tanner AC, Yu WH, et al. The human oral microbiome. J Bacteriol. (2010) 192:5002–17. doi: 10.1128/JB.00542-10
7. Deo PN and Deshmukh R. Oral microbiome: Unveiling the fundamentals. J Oral Maxillofac Pathol. (2019) 23:122–8. doi: 10.4103/jomfp.JOMFP_304_18
8. Yang K, Wang Y, Zhang S, Zhang D, Hu L, Zhao T, et al. Oral microbiota analysis of tissue pairs and saliva samples from patients with oral squamous cell carcinoma - A pilot study. Front Microbiol. (2021) 12:719601. doi: 10.3389/fmicb.2021.719601
9. Kim SY and Nair MG. Macrophages in wound healing: activation and plasticity. Immunol Cell Biol. (2019) 97:258–67. doi: 10.1111/imcb.12236
10. Abou Khouzam R, Goutham HV, Zaarour RF, Chamseddine AN, Francis A, Buart S, et al. Integrating tumor hypoxic stress in novel and more adaptable strategies for cancer immunotherapy. Semin Cancer Biol. (2020) 65:140–54. doi: 10.1016/j.semcancer.2020.01.003
11. Kondoh N, Mizuno-Kamiya M, Umemura N, Takayama E, Kawaki H, Mitsudo K, et al. Immunomodulatory aspects in the progression and treatment of oral Malignancy. Jpn Dent Sci Rev. (2019) 55:113–20. doi: 10.1016/j.jdsr.2019.09.001
12. Xie H, Xi X, Lei T, Liu H, and Xia Z. CD8(+) T cell exhaustion in the tumor microenvironment of breast cancer. Front Immunol. (2024) 15:1507283. doi: 10.3389/fimmu.2024.1507283
13. Frafjord A, Skarshaug R, Hammarström C, Stankovic B, Dorg LT, Aamodt H, et al. Antibody combinations for optimized staining of macrophages in human lung tumours. Scand J Immunol. (2020) 92:e12889. doi: 10.1111/sji.12889
14. Yoshida C, Kadota K, Yamada K, Fujimoto S, Ibuki E, Ishikawa R, et al. Tumor-associated CD163(+) macrophage as a predictor of tumor spread through air spaces and with CD25(+) lymphocyte as a prognostic factor in resected stage I lung adenocarcinoma. Lung Cancer. (2022) 167:34–40. doi: 10.1016/j.lungcan.2022.03.016
15. Sarkar P, Malik S, Laha S, Das S, Bunk S, Ray JG, et al. Dysbiosis of oral microbiota during oral squamous cell carcinoma development. Front Oncol. (2021) 11:614448. doi: 10.3389/fonc.2021.614448
16. Li Z, Chen G, Wang P, Sun M, Zhao J, Li A, et al. Alterations of the oral microbiota profiles in chinese patient with oral cancer. Front Cell Infect Microbiol. (2021) 11:780067. doi: 10.3389/fcimb.2021.780067
17. Komlós G, Csurgay K, Horváth F, Pelyhe L, and Németh Z. Periodontitis as a risk for oral cancer: a case-control study. BMC Oral Health. (2021) 21:640. doi: 10.1186/s12903-021-01998-y
18. Zhang L, Liu Y, Zheng HJ, and Zhang CP. The oral microbiota may have influence on oral cancer. Front Cell Infect Microbiol. (2019) 9:476. doi: 10.3389/fcimb.2019.00476
19. Ganly I, Yang L, Giese RA, Hao Y, Nossa CW, Morris LGT, et al. Periodontal pathogens are a risk factor of oral cavity squamous cell carcinoma, independent of tobacco and alcohol and human papillomavirus. Int J Cancer. (2019) 145:775–84. doi: 10.1002/ijc.32152
20. Olsen I and Yilmaz Ö. Possible role of Porphyromonas gingivalis in orodigestive cancers. J Oral Microbiol. (2019) 11:1563410. doi: 10.1080/20002297.2018.1563410
21. Wen L, Mu W, Lu H, Wang X, Fang J, Jia Y, et al. Porphyromonas gingivalis promotes oral squamous cell carcinoma progression in an immune microenvironment. J Dent Res. (2020) 99:666–75. doi: 10.1177/0022034520909312
22. Li Y, Tan X, Zhao X, Xu Z, Dai W, Duan W, et al. Composition and function of oral microbiota between gingival squamous cell carcinoma and periodontitis. Oral Oncol. (2020) 107:104710. doi: 10.1016/j.oraloncology.2020.104710
23. Harrandah AM, Chukkapalli SS, Bhattacharyya I, Progulske-Fox A, and Chan EKL. Fusobacteria modulate oral carcinogenesis and promote cancer progression. J Oral Microbiol. (2020) 13:1849493. doi: 10.1080/20002297.2020.1849493
24. Binder Gallimidi A, Fischman S, Revach B, Bulvik R, Maliutina A, Rubinstein AM, et al. Periodontal pathogens Porphyromonas gingivalis and Fusobacterium nucleatum promote tumor progression in an oral-specific chemical carcinogenesis model. Oncotarget. (2015) 6:22613–23. doi: 10.18632/oncotarget.4209
25. Neuzillet C, Marchais M, Vacher S, Hilmi M, Schnitzler A, Meseure D, et al. Prognostic value of intratumoral Fusobacterium nucleatum and association with immune-related gene expression in oral squamous cell carcinoma patients. Sci Rep. (2021) 11:7870. doi: 10.1038/s41598-021-86816-9
26. Nosho K, Sukawa Y, Adachi Y, Ito M, Mitsuhashi K, Kurihara H, et al. Association of Fusobacterium nucleatum with immunity and molecular alterations in colorectal cancer. World J Gastroenterol. (2016) 22:557–66. doi: 10.3748/wjg.v22.i2.557
27. Zhang S, Li C, Liu J, Geng F, Shi X, Li Q, et al. Fusobacterium nucleatum promotes epithelial-mesenchymal transiton through regulation of the lncRNA MIR4435-2HG/miR-296-5p/Akt2/SNAI1 signaling pathway. FEBS J. (2020) 287:4032–47. doi: 10.1111/febs.15233
28. Shao W, Fujiwara N, Mouri Y, Kisoda S, Yoshida K, Yoshida K, et al. Conversion from epithelial to partial-EMT phenotype by Fusobacterium nucleatum infection promotes invasion of oral cancer cells. Sci Rep. (2021) 11:14943. doi: 10.1038/s41598-021-94384-1
29. Inaba H, Sugita H, Kuboniwa M, Iwai S, Hamada M, Noda T, et al. Porphyromonas gingivalis promotes invasion of oral squamous cell carcinoma through induction of proMMP9 and its activation. Cell Microbiol. (2014) 16:131–45. doi: 10.1111/cmi.12211
30. Kamarajan P, Ateia I, Shin JM, Fenno JC, Le C, Zhan L, et al. Periodontal pathogens promote cancer aggressivity via TLR/MyD88 triggered activation of Integrin/FAK signaling that is therapeutically reversible by a probiotic bacteriocin. PloS Pathog. (2020) 16:e1008881. doi: 10.1371/journal.ppat.1008881
31. Bouaoud J, Foy JP, Tortereau A, Michon L, Lavergne V, Gadot N, et al. Early changes in the immune microenvironment of oral potentially Malignant disorders reveal an unexpected association of M2 macrophages with oral cancer free survival. Oncoimmunology. (2021) 10:1944554. doi: 10.1080/2162402X.2021.1944554
32. Chanmee T, Ontong P, and Itano N. Hyaluronan: A modulator of the tumor microenvironment. Cancer Lett. (2016) 375:20–30. doi: 10.1016/j.canlet.2016.02.031
33. Yan J, Ye G, Jin Y, Miao M, Li Q, and Zhou H. Identification of novel prognostic circRNA biomarkers in circRNA-miRNA-mRNA regulatory network in gastric cancer and immune infiltration analysis. BMC Genomics. (2023) 24:323. doi: 10.1186/s12864-023-09421-2
34. Chen Y, Jin H, Song Y, Huang T, Cao J, Tang Q, et al. Targeting tumor-associated macrophages: A potential treatment for solid tumors. J Cell Physiol. (2021) 236:3445–65. doi: 10.1002/jcp.30139
35. Xu T, Yu S, Zhang J, and Wu S. Dysregulated tumor-associated macrophages in carcinogenesis, progression and targeted therapy of gynecological and breast cancers. J Hematol Oncol. (2021) 14:181. doi: 10.1186/s13045-021-01198-9
36. Zhai X, Zhang H, Xia Z, Liu M, Du G, Jiang Z, et al. Oxytocin alleviates liver fibrosis via hepatic macrophages. JHEP Rep. (2024) 6:101032. doi: 10.1016/j.jhepr.2024.101032
37. Murray PJ. Macrophage polarization. Annu Rev Physiol. (2017) 79:541–66. doi: 10.1146/annurev-physiol-022516-034339
38. Arora S, Dev K, Agarwal B, Das P, and Syed MA. Macrophages: Their role, activation and polarization in pulmonary diseases. Immunobiology. (2018) 223:383–96. doi: 10.1016/j.imbio.2017.11.001
39. Huang X, Li Y, Fu M, and Xin HB. Polarizing macrophages in vitro. Methods Mol Biol. (2018) 1784:119–26. doi: 10.1007/978-1-4939-7837-3_12
40. Ryma M, Tylek T, Liebscher J, Blum C, Fernandez R, Böhm C, et al. Translation of collagen ultrastructure to biomaterial fabrication for material-independent but highly efficient topographic immunomodulation. Adv Mater. (2021) 33:e2101228. doi: 10.1002/adma.202101228
41. Gindele JA, Mang S, Pairet N, Christ I, Gantner F, Schymeinsky J, et al. Opposing effects of in vitro differentiated macrophages sub-type on epithelial wound healing. PloS One. (2017) 12:e0184386. doi: 10.1371/journal.pone.0184386
42. Chen J, Wang W, Ni Q, Zhang L, and Guo X. Interleukin 6-regulated macrophage polarization controls atherosclerosis-associated vascular intimal hyperplasia. Front Immunol. (2022) 13:952164. doi: 10.3389/fimmu.2022.952164
43. Lai H, Yip HC, Gong Y, Chan KF, Leung KK, Chan MS, et al. MFGE8 in exosomes derived from mesenchymal stem cells prevents esophageal stricture after endoscopic submucosal dissection in pigs. J Nanobiotechnol. (2024) 22:143. doi: 10.1186/s12951-024-02429-0
44. Li C, Zhang H, Liu Y, Zhang T, and Gu F. Gpr109A in TAMs promoted hepatocellular carcinoma via increasing PKA/PPARγ/MerTK/IL-10/TGFβ induced M2c polarization. Sci Rep. (2025) 15:18820. doi: 10.1038/s41598-025-02447-4
45. Lai W, Xian C, Chen M, Luo D, Zheng J, Zhao S, et al. Single-cell and bulk transcriptomics reveals M2d macrophages as a potential therapeutic strategy for mucosal healing in ulcerative colitis. Int Immunopharmacol. (2023) 121:110509. doi: 10.1016/j.intimp.2023.110509
46. Sezginer O and Unver N. Dissection of pro-tumoral macrophage subtypes and immunosuppressive cells participating in M2 polarization. Inflammation Res. (2024) 73:1411–23. doi: 10.1007/s00011-024-01907-3
47. Chiu KC, Lee CH, Liu SY, Chou YT, Huang RY, Huang SM, et al. Polarization of tumor-associated macrophages and Gas6/Axl signaling in oral squamous cell carcinoma. Oral Oncol. (2015) 51:683–9. doi: 10.1016/j.oraloncology.2015.04.004
48. Swann JB, Vesely MD, Silva A, Sharkey J, Akira S, Schreiber RD, et al. Demonstration of inflammation-induced cancer and cancer immunoediting during primary tumorigenesis. Proc Natl Acad Sci U.S.A. (2008) 105:652–6. doi: 10.1073/pnas.0708594105
49. van Dalen FJ, van Stevendaal M, Fennemann FL, Verdoes M, and Ilina O. Molecular repolarisation of tumour-associated macrophages. Molecules. (2018) 24:9. doi: 10.3390/molecules24010009
50. Brenner C, Galluzzi L, Kepp O, and Kroemer G. Decoding cell death signals in liver inflammation. J Hepatol. (2013) 59:583–94. doi: 10.1016/j.jhep.2013.03.033
51. Jiang M, Wang Z, He X, Hu Y, Xie M, Jike Y, et al. A risk-scoring model based on evaluation of ferroptosis-related genes in osteosarcoma. J Oncol. (2022) 2022:4221756. doi: 10.1155/2022/4221756
52. Potas JR, Haque F, Maclean FL, and Nisbet DR. Interleukin-10 conjugated electrospun polycaprolactone (PCL) nanofibre scaffolds for promoting alternatively activated (M2) macrophages around the peripheral nerve in vivo. J Immunol Methods. (2015) 420:38–49. doi: 10.1016/j.jim.2015.03.013
53. Asai A, Tsuchimoto Y, Ohama H, Fukunishi S, Tsuda Y, Kobayashi M, et al. Host antitumor resistance improved by the macrophage polarization in a chimera model of patients with HCC. Oncoimmunology. (2017) 6:e1299301. doi: 10.1080/2162402X.2017.1299301
54. Wu H, Xu JB, He YL, Peng JJ, Zhang XH, Chen CQ, et al. Tumor-associated macrophages promote angiogenesis and lymphangiogenesis of gastric cancer. J Surg Oncol. (2012) 106:462–8. doi: 10.1002/jso.23110
55. Cummings RD. The mannose receptor ligands and the macrophage glycome. Curr Opin Struct Biol. (2022) 75:102394. doi: 10.1016/j.sbi.2022.102394
56. Jaynes JM, Sable R, Ronzetti M, Bautista W, Knotts Z, Abisoye-Ogunniyan A, et al. Mannose receptor (CD206) activation in tumor-associated macrophages enhances adaptive and innate antitumor immune responses. Sci Transl Med. (2020) 12:eaax6337. doi: 10.1126/scitranslmed.aax6337
57. Kai K, Moriyama M, Haque A, Hattori T, Chinju A, Hu C, et al. Oral squamous cell carcinoma contributes to differentiation of monocyte-derived tumor-associated macrophages via PAI-1 and IL-8 production. Int J Mol Sci. (2021) 22:9475. doi: 10.3390/ijms22179475
58. Haque A, Moriyama M, Kubota K, Ishiguro N, Sakamoto M, Chinju A, et al. CD206(+) tumor-associated macrophages promote proliferation and invasion in oral squamous cell carcinoma via EGF production. Sci Rep. (2019) 9:14611. doi: 10.1038/s41598-019-51149-1
59. Bruna F and Scodeller P. Pro-tumorigenic macrophage infiltration in oral squamous cell carcinoma and possible macrophage-aimed therapeutic interventions. Front Oncol. (2021) 11:675664. doi: 10.3389/fonc.2021.675664
60. Chaudhari N, Prakash N, Pradeep GL, Mahajan A, Lunawat S, and Salunkhe V. Evaluation of density of tumor-associated macrophages using CD163 in histological grades of oral squamous cell carcinoma, an immunohistochemical study. J Oral Maxillofac Pathol. (2020) 24:577. doi: 10.4103/jomfp.JOMFP_109_20
61. Etzerodt A, Tsalkitzi K, Maniecki M, Damsky W, Delfini M, Baudoin E, et al. Specific targeting of CD163(+) TAMs mobilizes inflammatory monocytes and promotes T cell-mediated tumor regression. J Exp Med. (2019) 216:2394–411. doi: 10.1084/jem.20182124
62. Suárez-Sánchez FJ, Lequerica-Fernández P, Suárez-Canto J, Rodrigo JP, Rodriguez-Santamarta T, Domínguez-Iglesias F, et al. Macrophages in oral carcinomas: relationship with cancer stem cell markers and PD-L1 expression. Cancers (Basel). (2020) 12:1764. doi: 10.3390/cancers12071764
63. Li JJ, Mao XH, Tian T, Wang WM, Su T, Jiang CH, et al. Role of PFKFB3 and CD163 in oral squamous cell carcinoma angiogenesis. Curr Med Sci. (2019) 39:410–4. doi: 10.1007/s11596-019-2051-1
64. Chen TW, Hung WZ, Chiang SF, Chen WT, Ke TW, Liang JA, et al. Dual inhibition of TGFβ signaling and CSF1/CSF1R reprograms tumor-infiltrating macrophages and improves response to chemotherapy via suppressing PD-L1. Cancer Lett. (2022) 543:215795. doi: 10.1016/j.canlet.2022.215795
65. Wang X, Zhang J, Hu B, and Qian F. High expression of CSF-1R predicts poor prognosis and CSF-1R(high) tumor-associated macrophages inhibit anti-tumor immunity in colon adenocarcinoma. Front Oncol. (2022) 12:850767. doi: 10.3389/fonc.2022.850767
66. Rodriguez-Perdigon M, Jimaja S, Haeni L, Bruns N, Rothen-Rutishauser B, and Rüegg C. Polymersomes-mediated delivery of CSF1R inhibitor to tumor associated macrophages promotes M2 to M1-like macrophage repolarization. Macromol Biosci. (2022) 22:e2200168. doi: 10.1002/mabi.202200168
67. Guo XY, Zhang JY, Shi XZ, Wang Q, Shen WL, Zhu WW, et al. Upregulation of CSF-1 is correlated with elevated TAM infiltration and poor prognosis in oral squamous cell carcinoma. Am J Transl Res. (2020) 12:6235–49.
68. da Silva SD, Marchi FA, Su J, Yang L, Valverde L, Hier J, et al. Co-overexpression of TWIST1-CSF1 is a common event in metastatic oral cancer and drives biologically aggressive phenotype. Cancers (Basel). (2021) 13:153. doi: 10.3390/cancers13010153
69. Ge Z and Ding S. The crosstalk between tumor-associated macrophages (TAMs) and tumor cells and the corresponding targeted therapy. Front Oncol. (2020) 10:590941. doi: 10.3389/fonc.2020.590941
70. Zhou Q, Meng Y, Li D, Yao L, Le J, Liu Y, et al. Ferroptosis in cancer: from molecular mechanisms to therapeutic strategies. Signal Transduct Target Ther. (2024) 9:55. doi: 10.1038/s41392-024-01769-5
71. Zhao YY, Lian JX, Lan Z, Zou KL, Wang WM, and Yu GT. Ferroptosis promotes anti-tumor immune response by inducing immunogenic exposure in HNSCC. Oral Dis. (2023) 29:933–41. doi: 10.1111/odi.14077
72. Wu L, Tang H, Zheng H, Liu X, Liu Y, Tao J, et al. Multiwalled carbon nanotubes prevent tumor metastasis through switching M2-polarized macrophages to M1 via TLR4 activation. J BioMed Nanotechnol. (2019) 15:138–50. doi: 10.1166/jbn.2019.2661
73. Matsusaka K, Fujiwara Y, Pan C, Esumi S, Saito Y, Bi J, et al. α(1)-acid glycoprotein enhances the immunosuppressive and protumor functions of tumor-associated macrophages. Cancer Res. (2021) 81:4545–59. doi: 10.1158/0008-5472.CAN-20-3471
74. Cao M, Yan H, Han X, Weng L, Wei Q, Sun X, et al. Ginseng-derived nanoparticles alter macrophage polarization to inhibit melanoma growth. J Immunother Cancer. (2019) 7:326. doi: 10.1186/s40425-019-0817-4
75. Li R, Zhou R, Wang H, Li W, Pan M, Yao X, et al. Gut microbiota-stimulated cathepsin K secretion mediates TLR4-dependent M2 macrophage polarization and promotes tumor metastasis in colorectal cancer. Cell Death Differ. (2019) 26:2447–63. doi: 10.1038/s41418-019-0312-y
76. Wang B, He Y, Wang B, Li J, and Qin L. ALDH3A1 overexpression in OSCC inhibits inflammation via phospho-Ser727 at STAT3 in tumor-associated macrophages. Oral Dis. (2023) 29:1513–24. doi: 10.1111/odi.14161
77. Wang HY, Ge JC, Zhang FY, Zha XQ, Liu J, Li QM, et al. Dendrobium officinale polysaccharide promotes M1 polarization of TAMs to inhibit tumor growth by targeting TLR2. Carbohydr Polym. (2022) 292:119683. doi: 10.1016/j.carbpol.2022.119683
78. Bolli E, Scherger M, Arnouk SM, Pombo Antunes AR, Straßburger D, Urschbach M, et al. : targeted repolarization of tumor-associated macrophages via imidazoquinoline-linked nanobodies. Adv Sci (Weinh). (2021) 8:2004574. doi: 10.1002/advs.202004574
79. Zhang Y, Feng Z, Liu J, Li H, Su Q, Zhang J, et al. Polarization of tumor-associated macrophages by TLR7/8 conjugated radiosensitive peptide hydrogel for overcoming tumor radioresistance. Bioact Mater. (2022) 16:359–71. doi: 10.1016/j.bioactmat.2021.12.033
80. Wei X, Liu L, Li X, Wang Y, Guo X, Zhao J, et al. Selectively targeting tumor-associated macrophages and tumor cells with polymeric micelles for enhanced cancer chemo-immunotherapy. J Control Release. (2019) 313:42–53. doi: 10.1016/j.jconrel.2019.09.021
81. Tartey S, Neale G, Vogel P, Malireddi RKS, and Kanneganti TD. A myD88/IL1R axis regulates PD-1 expression on tumor-associated macrophages and sustains their immunosuppressive function in melanoma. Cancer Res. (2021) 81:2358–72. doi: 10.1158/0008-5472.CAN-20-3510
82. Zhang W, Borcherding N, and Kolb R. IL-1 signaling in tumor microenvironment. Adv Exp Med Biol. (2020) 1240:1–23. doi: 10.1007/978-3-030-38315-2_1
83. Niklander SE, Murdoch C, and Hunter KD. IL-1/IL-1R signaling in head and neck cancer. Front Oral Health. (2021) 2:722676. doi: 10.3389/froh.2021.722676
84. Petruzzi MN, Cherubini K, Salum FG, and de Figueiredo MA. Role of tumour-associated macrophages in oral squamous cells carcinoma progression: an update on current knowledge. Diagn Pathol. (2017) 12:32. doi: 10.1186/s13000-017-0623-6
85. Li X, Bu W, Meng L, Liu X, Wang S, Jiang L, et al. CXCL12/CXCR4 pathway orchestrates CSC-like properties by CAF recruited tumor associated macrophage in OSCC. Exp Cell Res. (2019) 378:131–8. doi: 10.1016/j.yexcr.2019.03.013
86. Zhou J, Zhang A, and Fan L. HSPA12B secreted by tumor-associated endothelial cells might induce M2 polarization of macrophages via activating PI3K/akt/mTOR signaling. Onco Targets Ther. (2020) 13:9103–11. doi: 10.2147/OTT.S254985
87. Veerasamy V, Veeran V, and Nagini S. Dysregulated PI3K/AKT signaling in oral squamous cell carcinoma: The tumor microenvironment and epigenetic modifiers as key drivers. Oncol Res. (2025) 33:1835–60. doi: 10.32604/or.2025.064010
88. Takabatake K, Shimo T, Murakami J, Anqi C, Kawai H, Yoshida S, et al. The role of sonic hedgehog signaling in the tumor microenvironment of oral squamous cell carcinoma. Int J Mol Sci. (2019) 20:5779. doi: 10.3390/ijms20225779
89. Lu T, Zhang Z, Zhang J, Pan X, Zhu X, Wang X, et al. CD73 in small extracellular vesicles derived from HNSCC defines tumour-associated immunosuppression mediated by macrophages in the microenvironment. J Extracell Vesicles. (2022) 11:e12218. doi: 10.1002/jev2.12218
90. Hezaveh K, Shinde RS, Klötgen A, Halaby MJ, Lamorte S, Ciudad MT, et al. Tryptophan-derived microbial metabolites activate the aryl hydrocarbon receptor in tumor-associated macrophages to suppress anti-tumor immunity. Immunity. (2022) 55:324–340.e328. doi: 10.1016/j.immuni.2022.01.006
91. Slots J. Periodontitis: facts, fallacies and the future. Periodontol 2000. (2017) 75:7–23. doi: 10.1111/prd.12221
92. Söder B, Källmén H, Yucel-Lindberg T, and Meurman JH. Periodontal microorganisms and diagnosis of Malignancy: A cross-sectional study. Tumour Biol. (2021) 43:1–9. doi: 10.3233/TUB-200066
93. Almubarak A, Tanagala KKK, Papapanou PN, Lalla E, and Momen-Heravi F. Disruption of monocyte and macrophage homeostasis in periodontitis. Front Immunol. (2020) 11:330. doi: 10.3389/fimmu.2020.00330
94. Yu S, Ding L, Liang D, and Luo L. Porphyromonas gingivalis inhibits M2 activation of macrophages by suppressing α-ketoglutarate production in mice. Mol Oral Microbiol. (2018) 33:388–95. doi: 10.1111/omi.12241
95. Yuan G, Fu C, Yang ST, Yuh DY, Hajishengallis G, and Yang S. RGS12 drives macrophage activation and osteoclastogenesis in periodontitis. J Dent Res. (2022) 101:448–57. doi: 10.1177/00220345211045303
96. Croft AJ, Metcalfe S, Honma K, and Kay JG. Macrophage polarization alters postphagocytosis survivability of the commensal streptococcus gordonii. Infect Immun. (2018) 86:e00858–17. doi: 10.1128/IAI.00858-17
97. Wei W, Li J, Shen X, Lyu J, Yan C, Tang B, et al. Oral microbiota from periodontitis promote oral squamous cell carcinoma development via γδ T cell activation. mSystems. (2022) 7:e0046922. doi: 10.1128/msystems.00469-22
98. Liu S, Zhou X, Peng X, Li M, Ren B, Cheng G, et al. Porphyromonas gingivalis promotes immunoevasion of oral cancer by protecting cancer from macrophage attack. J Immunol. (2020) 205:282–9. doi: 10.4049/jimmunol.1901138
99. Cecil JD, O’Brien-Simpson NM, Lenzo JC, Holden JA, Singleton W, Perez-Gonzalez A, et al. Outer membrane vesicles prime and activate macrophage inflammasomes and cytokine secretion In Vitro and In Vivo. Front Immunol. (2017) 8:1017. doi: 10.3389/fimmu.2017.01017
100. Li C, Yin W, Yu N, Zhang D, Zhao H, Liu J, et al. miR-155 promotes macrophage pyroptosis induced by Porphyromonas gingivalis through regulating the NLRP3 inflammasome. Oral Dis. (2019) 25:2030–9. doi: 10.1111/odi.13198
101. Schmidt F, Dahlke K, Batra A, Keye J, Wu H, Friedrich M, et al. Microbial colonization in adulthood shapes the intestinal macrophage compartment. J Crohns Colitis. (2019) 13:1173–85. doi: 10.1093/ecco-jcc/jjz036
102. Wan G, Xie M, Yu H, and Chen H. Intestinal dysbacteriosis activates tumor-associated macrophages to promote epithelial-mesenchymal transition of colorectal cancer. Innate Immun. (2018) 24:480–9. doi: 10.1177/1753425918801496
103. Belogortseva N, Krezalek M, Guyton K, Labno C, Poroyko V, Zaborina O, et al. Media from macrophages co-incubated with Enterococcus faecalis induces epithelial cell monolayer reassembly and altered cell morphology. PloS One. (2017) 12:e0182825. doi: 10.1371/journal.pone.0182825
104. Karin M, Lawrence T, and Nizet V. Innate immunity gone awry: linking microbial infections to chronic inflammation and cancer. Cell. (2006) 124:823–35. doi: 10.1016/j.cell.2006.02.016
105. Kostic AD, Chun E, Robertson L, Glickman JN, Gallini CA, Michaud M, et al. Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe. (2013) 14:207–15. doi: 10.1016/j.chom.2013.07.007
106. Wu J, Li Q, and Fu X. Fusobacterium nucleatum contributes to the carcinogenesis of colorectal cancer by inducing inflammation and suppressing host immunity. Transl Oncol. (2019) 12:846–51. doi: 10.1016/j.tranon.2019.03.003
107. Chen T, Li Q, Wu J, Wu Y, Peng W, Li H, et al. Fusobacterium nucleatum promotes M2 polarization of macrophages in the microenvironment of colorectal tumours via a TLR4-dependent mechanism. Cancer Immunol Immunother. (2018) 67:1635–46. doi: 10.1007/s00262-018-2233-x
108. Xu C, Fan L, Lin Y, Shen W, Qi Y, Zhang Y, et al. Fusobacterium nucleatum promotes colorectal cancer metastasis through miR-1322/CCL20 axis and M2 polarization. Gut Microbes. (2021) 13:1980347. doi: 10.1080/19490976.2021.1980347
109. Mola S, Pandolfo C, Sica A, and Porta C. The macrophages-microbiota interplay in colorectal cancer (CRC)-related inflammation: prognostic and therapeutic significance. Int J Mol Sci. (2020) 21:6866. doi: 10.3390/ijms21186866
110. Gong X, Chi H, Xia Z, Yang G, and Tian G. Advances in HPV-associated tumor management: Therapeutic strategies and emerging insights. J Med Virol. (2023) 95:e28950. doi: 10.1002/jmv.28950
111. Wang B, Deng J, Donati V, Merali N, Frampton AE, Giovannetti E, et al. The Roles and Interactions of Porphyromonas gingivalis and Fusobacterium nucleatum in Oral and Gastrointestinal Carcinogenesis: A Narrative Review. Pathogens. (2024) 13:93. doi: 10.3390/pathogens13010093
112. McIlvanna E, Linden GJ, Craig SG, Lundy FT, and James JA. Fusobacterium nucleatum and oral cancer: a critical review. BMC Cancer. (2021) 21:1212. doi: 10.1186/s12885-021-08903-4
113. Ceci C, Atzori MG, Lacal PM, and Graziani G. Targeting tumor-associated macrophages to increase the efficacy of immune checkpoint inhibitors: A glimpse into novel therapeutic approaches for metastatic melanoma. Cancers (Basel). (2020) 12:3401. doi: 10.3390/cancers12113401
114. Kouketsu A, Sato I, Oikawa M, Shimizu Y, Saito H, Tashiro K, et al. Regulatory T cells and M2-polarized tumour-associated macrophages are associated with the oncogenesis and progression of oral squamous cell carcinoma. Int J Oral Maxillofac Surg. (2019) 48:1279–88. doi: 10.1016/j.ijom.2019.04.004
Keywords: oral squamous cell carcinoma, tumor-associated macrophages, microbiota, inflammation, chemokine receptors, immune suppression
Citation: Deng X and Huang S (2025) Microbiome-macrophage crosstalk in the tumor microenvironment: implications for oral squamous cell carcinoma progression and therapy. Front. Immunol. 16:1651837. doi: 10.3389/fimmu.2025.1651837
Received: 22 June 2025; Accepted: 11 August 2025;
Published: 29 August 2025.
Edited by:
Jin Bin, Shandong University, ChinaReviewed by:
Prabhakar Mujagond, Southern Medical University, ChinaCopyright © 2025 Deng and Huang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Shaohong Huang, Z2Q5MjBAMjFjbi5jb20=
 Xin Deng
Xin Deng Shaohong Huang*
Shaohong Huang*