- 1The Sri Lankan Blue Whale Project and Oceanswell, Colombo, Sri Lanka
- 2Menzies Institute for Medical Research, University of Tasmania, Hobart, TAS, Australia
- 3Trace and Environmental DNA Laboratory, Molecular and Life Sciences, Curtin University, Perth, WA, Australia
- 4Environomics Future Science Platform, CSIRO National Collections and Marine Infrastructure, Crawley, WA, Australia
Blue whales are little studied, face significant anthropogenic threats and within the Northern Indian Ocean, have a restricted range, making them an archetype for conservation needs of megafauna around the world. We studied feeding behavior of blue whales using dietary DNA metabarcoding of fecal samples. While globally blue whale populations feed predominantly on Euphausiidae, 87% of prey DNA amplicons extracted from fecal samples from this population were sergestid shrimp, demonstrating that blue whales can locate and feed on dense swarms of other types of prey when they occur. Within the Indian Ocean sergestids are present within the top 300 m, which correlates with the deep scattering layer observed by hydroacoustics. Studies suggest that this requirement to dive deeper in search of prey likely explains the prevalence of fluke up diving within this population of blue whales relative to other parts of the globe. Furthermore, this study revealed the presence of acanthocephalan endoparasites within the stomach and intestines of the Northern Indian Ocean blue whales. This represents the first record of Acanthocephala in blue whales in the Northern Indian Ocean and highlights the need for further studies on both the ecto- and endoparasitic flora and monitoring of health of these cetaceans for their management and conservation.
Introduction
Baleen whales feed on small animals that can be filtered from the ocean using their baleen plates, which act like nets to separate their food from seawater (Pivorunas, 1979). They regularly feed on euphausiids, copepods, mysids, small fish, and amphipods. Blue whales (Balaenoptera musculus) are the archetypal euphausiid-feeding great whales. They are thought to fulfill their energetic needs by feeding on dense schools of their preferred prey, euphausiids, worldwide (Yochem and Leatherwood, 1985; Schoenherr, 1991; Croll et al., 1998; Gill, 2002).
Large marine predators forage on prey that are spatially and temporally patchy (Haury et al., 1978; Denman and Powell, 1984). To support their energy requirements, it is assumed that animals target areas that are resource abundant (Fauchald and Tveraa, 2003). Euphausiids have extremely patchy distributions in comparison to other animals of similar size and in some areas they form aggregations where density is as high as 1,500 individuals per m3 (Higginbottom and Hosie, 1989), making them an ideal target for lunge filter-feeding of blue whales. Blue whale movements within and between feeding areas are thought to be driven by the availability of dense patches of such prey (Fiedler et al., 1998). Most baleen whale species undertake long-range seasonal migrations between productive high latitude feeding grounds and unproductive, low latitude breeding grounds (Brodie, 1975; Corkeron and Connor, 1999). However, because blue whales need to seek regions of exceptionally high productivity (Croll et al., 2005), there is evidence that they feed at high, mid and low latitudes (Reilly and Thayer, 1990; Silva et al., 2013).
Northern Indian Ocean (NIO) blue whales (Balenoptera musculus indica) however, usually remain in low latitude waters throughout the year (Alling et al., 1991; de Vos et al., 2012). A number of their areas of aggregation have been identified, including waters near Somalia (Small and Small, 1991), the Seychelles (Mikhalev, 2000), the Maldives (Ballance et al., 2001; Anderson, 2005), and Sri Lanka (de Vos et al., 2014a). Sighting, stranding and acoustic data (Alling et al., 1991; Ilangakoon, 2006; Branch et al., 2007; Afsal et al., 2008; de Vos et al., 2012) indicate that this population occupies Sri Lankan waters almost year round. Off southern Sri Lanka, groups of NIO blue whales have been observed engaging in breeding activity (specifically “ménage à trois” behaviors as described by Sears et al., 2009). More significantly, they have been observed feeding (de Vos et al., 2014a) and defecating during the Northeast monsoon (December to March) (de Vos et al., 2012) indicating that this is an important foraging area.
Many wild animals are elusive or inhabit hostile environments making it nearly impossible to observe them as they undertake critical activities such as feeding, breeding, or escaping predators. The diet of whales while an important aspect of their ecology, is a particularly difficult one to study because of the challenges in linking observed surface behaviors to subsurface foraging activities and ultimately to the food that is consumed. Furthermore, whales often have vast ranges, are difficult to observe closely at sea, and are available for observation over limited spatial and temporal scales (Samuels and Tyack, 1999; Au et al., 2000).
In this study of blue whale diet, we used DNA metabarcoding of fecal samples. Feces have previously been extensively used to study diet composition in blue whales and a variety of other species across the world (Jarman et al., 2002; Jarman and Wilson, 2004; Meekan et al., 2009; Pompanon et al., 2012). More recently fecal samples have also been used for the assessment of parasites in baleen whale species (Hunt et al., 2013). Protozoan parasites have been detected in both North Atlantic right whales (Eubalaena glacialis) and bowhead whale feces (Balaena mysticetus) (Hughes-Hanks et al., 2005). Therefore the main purpose of this work was to ascertain if the Northern Indian Ocean blue whale also preferentially preyed on euphausiids, as visual observations of diving behavior suggest they may be targeting deeper water species around Sri Lanka (de Vos et al., 2013). Secondarily, these samples were used to identify other species such as parasites that exist within the gut flora of the blue whales within the Northern Indian Ocean.
Materials and Methods
Sampling Protocol
A total of 34 blue whale fecal samples were collected from 16 different encounters during the months of January, February and March 2013 off the southern coast of Sri Lanka. The samples were collected opportunistically from the sea surface using a strainer with mesh size 0.5 mm when a blue whale was at least 50 m away to minimize disturbance caused by the research boat. Each sample was placed into a sterile sample collection bottle sufficient to fill half the bottle. The sample was then preserved in 80% ethanol, labeled and placed in an ice box until we reached shore when the sample was placed in a freezer that was maintained at −20°C. The sampled material was a coarse paste of animal particles (see Supplementary Material). It was not possible to sample the water where the whale was about to defecate with the same sampling procedure because a coarse mesh sieve cannot be used to sample water and it is not possible to predict when a whale will defecate. Differences among the samples in the DNA sequences produced were used as a proxy for negative water controls.
Amplification of DNA Metabarcode Regions and HTS
DNA was purified from 34 blue whale fecal samples with the QIAamp DNA stool kit following the manufacturer's instructions (Qiagen, Hilden, Germany). Only 16 of these samples had PCR amplifiable DNA, which was used for the subsequent analysis. A small fragment (~120–160 bp) of the nuclear small subunit rRNA gene of eukaryotes was amplified by PCR from the fecal DNA as previously described (Jarman et al., 2013). Amplifications were performed in triplicate with each 10 uL reaction contained 1 x Phusion PCR mastermix (NEB), 10 x BSA (NEB), 1 x EvaGreen dye (Biotium) and 1 uL of template DNA. Primers included were 1 uM of each primer 18Sf-msq (5′-TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCACCGCCCGTCGCTACTACCG-3′), 18Sr-msq (5′-GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGGTTCACCTACGGAAACCTTGTTACG-3′) and 10 uM of TetrapodBlockc3 (5′-CCTTGTTACGACTTTTACTTCCTCTAGATAG-c_3-3′) where c_3 refers to a C3 “spacer” modification (McInnes et al., 2016). Thermal cycling conditions were 98°C, 120 s; followed by 35 cycles of 98°C for 5 s, 67°C for 20 s, 72°C for 20 s. The first-round PCRs were performed in triplicate and a template-free control reaction was run. Reactions were run on an LC480 real time quantitative thermal cycler. Amplification of the template was monitored by EvaGreen fluorescence. Samples that produced three positive amplifications with a “crossing threshold” (Ct) of less than 30 were pooled and used for second round PCR. These low Ct-values indicate exponential amplification of reasonably concentrated DNA template.
A second round of PCR was used to add a unique combination of 10 bp index sequences to the pool of PCR products amplified from each sample (McInnes et al., 2016). Reaction conditions were as for the first round PCR, but without the EvaGreen dye or BSA and the template was 2 uL of a 1:10 dilution of the three first round PCR products. Thermal cycling conditions were 98°C, 120 s; followed by 35 cycles of 98°C for 5 s, 55°C for 20 s, 72°C for 20 s. No combinations of forward and reverse indexes are re-used in the laboratory as a measure to control contamination. A negative DNA purification control and two no template controls where water was added instead of DNA were also PCR amplified with unique index combinations. Fragments were sequenced on an Illumina MiSeq Sequencer (Ilumina) using a 300 cycle v2 Nano kit and 150 bp paired-end reads.
Data Processing and Sequence Identification
The reads were assigned to a sample based on the combination of their forward and reverse indices using the Illumina MiSeq Reporter software. Subsequent sequence processing steps were performed with Usearch7 (Edgar, 2010) and custom Python scripts to apply the process to the sequences from each of the 16 samples. The processing started with quality filtering, where reads less than 100 bp long or having a Q < 10 base were removed. Paired reads were then merged with staggered overlaps allowed and the truncqual parameter set to 2. Sequences that occurred only once were removed and clusters created with the clusterfast algorithm with an identity threshold of 0.9 set for cluster formation as used previously for this fragment (Jarman et al., 2013). Centroids for each cluster were identified to group level by BLASTn searches (Altschul et al., 1990) of the complete nucleotide database of GenBank. Centroid sequences were assigned to taxa that they matched most closely as in Jarman et al. (2013). No anonymous MOTUs were identified. Singleton sequences were removed during processing, which is likely to have been effective for removing chimaeric PCR products. There is not a validated, high-quality database available for chimera detection for the 3′ end of the SSU rDNA, but the lack of orphan sequences indicates that chimera formation was not significant. The DNA sequence data produced in this study was archived at www.datadryad.org and its DOI is doi: 10.5061/dryad.qt352sg.
Results
The DNA metabarcoding methodology that we employed has the advantages of being present in most eukaryotes and being PCR-amplifiable from degraded DNA. This provides a reasonable basis for metazoan diversity analysis. Our results indicate that overall, 87% of DNA detected in the blue whale fecal samples collected off Sri Lanka comprised Dendrobranchiata, with Euphausiacea representing the second most ingested prey group (8%). The results also indicate that blue whales ingested small proportions of other species groups ranging from Amphipoda to Cephalopoda. It is also notable that 94% of the blue whale feces sampled had evidence of Dendrobranchiata, specifically Sergestid shrimp DNA, followed by 56% having Euphausiacea. Thirty seven percent also indicated consumption of Amphipoda, but generally in lower quantities (Figure 1). The short DNA metabarcode that we used only allowed identification to groups of thes taxonomic levels (Jarman et al., 2013). No metazoan sequences were identified in the template-free negative control reactions.
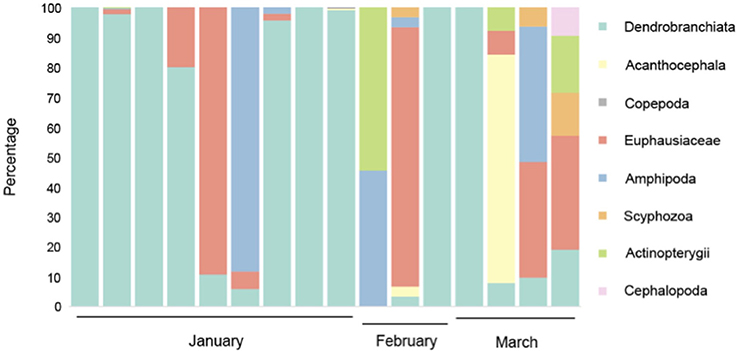
Figure 1. Prey species detected in Northern Indian Ocean blue whale fecal samples. Each column represents a single sample. Percentage of amplicons belonging to each prey group extracted from samples is indicated. The lack of a consistent, diverse background of amplicons suggests that the primary source of the amplicons amplified from the fecal samples is dietary rather than derived from seawater.
Based on the samples collected during this study it is also evident that feeding varied through the study period which ranged from January to March 2013. While diet in January comprised predominantly of Dendrobranchiata, the samples from March comprised a wider range of prey species with no species being particularly predominant (Figure 1).
Discussion
The discovery that blue whales in warm tropical waters off Sri Lanka feed predominantly on Dendrobranchiata rather than Euphausiids, their typical primary prey in other areas, demonstrates that blue whales are able to locate and feed on dense swarms of other types of prey when they occur. This parallels observations by Fiedler et al. (1998) that blue whales in the Channel Islands of California were taking advantage of the seasonal swarming behavior of Thyasanoessa spinifera, at shallower depths and higher density than their more typical prey Euphausia pacifica, during the relatively short periods when seasonal swarming of T. spinifera and blue whales movements into that area overlap (Fiedler et al., 1998). Previously, Alling et al. (1991) proposed that Northern Indian Ocean blue whales fed on mysids, however no systematic examination of prey samples is available to verify this statement. Further, the molecular assay that we applied in this study can detect Mysida in mixed samples (Jarman et al., 2013) but none were detected here. We have presented our DNA metabarcoding results as proportions of reads in each sample because the whale fecal samples are inevitably contaminated with small amounts of trace environmental DNA in seawater. However, the PCR primer binding sites for the amplicon we sequenced are highly conserved (Jarman et al., 2013), which provides a good basis for the results being semi-quantitative (Willerslev et al., 2014; Evans et al., 2016). If environmental DNA contamination was contributing significantly to the overall result, there would be a more consistent influence on the DNA signal found during each time period. The large differences in read composition among samples suggest that the environmental DNA contribution to the DNA pool is small compared to that from the fresh fecal material.
Large baleen whales require considerable concentrations of food to make their energetically demanding lunge feeding strategy viable (Brodie et al., 1978; Macaulay et al., 1995). The range of species documented in the diet of blue whales during this study is unsurprising given that their feeding behavior involves lunging into prey patches and filtering prey (Pivorunas, 1979; Goldbogen et al., 2011). This feeding adaptation makes it impossible to feed selectively on small prey and the results support a non-selective feeding strategy. The presence of Cephalapoda in one sample reflects this non-selectivity in feeding (Clarke, 1996).
Sergestidae, the predominant prey of NIO blue whales as suggested by this study, primarily ingest euphausiids and copepods, while the rest of their diet comprises chaetognaths, ostracods, and radiolarians with some evidence of phytoplankton, protists and cnidarian nematocysts (Tavares and Martin, 2010). Given that sergestids, like other small zooplankton species are movement limited and restricted to diel vertical migrations (Steele, 1978), they are likely densest in areas where they can encounter their favored prey items. This further explains the small quantities of chaetognaths, copepods and euphausiids in the diet of the blue whales in this study (Figure 1). The variation in prey availability during the months of January, February and March indicate that seasonal variation occurs within the water column and is likely driven by changes in circulation resulting from Sri Lanka's monsoonal climate (de Vos et al., 2014b).
Sergestidae predominantly comprise species that are deep benthic dwellers or members of the meso- and bathypelagic fauna (Pérez-Farfante and Kensley, 1997). They are primarily mesopelagic during the day with much of the assemblage exhibiting diel vertical migration into the epipelagic zone at night (Donaldson, 1975; Walters, 1976). Sergestids are an abundant component of oceanic micronekton and because of their zooplanktivorous habits are an important link between zooplankton and higher trophic levels in pelagic ecosystems (Yousuf, 2006). The Indian Ocean has upwards of 38 species of Sergestidae (Palomares and Pauly, 2016). Research in the Arabian and Northern Arabian seas showed an increase in sergestids in net trawls northwards between 13 and 17° and in day time trawls individuals of the species Sergestes seminudus were collected at depths between 0 and 300 m (Karuppasamy et al., 2010). Other species of the Genus Sergia have been recorded in day time trawls at varying depths up to 2,000 m (Vereshchaka, 2009). The deep scattering layer that has been previously recorded at approximately 300 m within the study area may therefore comprise this prey species (see Supplementary Material).
Feeding on these epibenthic animals necessitates deeper foraging dives. Within the NIO, blue whales have been frequently documented “fluking up,” or lifting their tail flukes before a deep dive (de Vos et al., 2013). These whales have been observed accelerating in to the terminal surfacing as they present the high arch and then the fluke (de Vos et al., 2013), enabling them to flip vertically and sink to depth likely because the high descent angles enable faster acceleration when the body is vertically oriented, due to lowered pressure drag as the projected area is significantly decreased compared to when the body is broadside to vertically acting buoyant forces (Goldbogen et al., 2006).
Blue whale dive depths are known to closely track the depth distributions of euphausiids, as indicated by a correlation in vertical space off the coast of California (Croll et al., 1998). Research from Monterey Bay indicates that blue whales concentrated their foraging activity on deeper layers of euphausiids located between 150 and 200 m on the canyon edge (Croll et al., 2005) with mean maximum foraging dive depth of two tagged whales reaching 155 and 172 m (Croll et al., 2005) compared to non-foraging dives that reached only 67 m on average (Croll et al., 2001). Research off the Channel Islands, California indicated that blue whales foraged at similar dive depths (Croll et al., 1998; Fiedler et al., 1998; Calambokidis et al., 2001). More recently, Goldbogen et al. (2011) recorded average dive depths and durations of 201 m and 9.8 min respectively over 200 blue whale foraging dives off the eastern North Pacific. In Sri Lanka, blue whale dive duration averaged 10.7 min (de Vos et al., 2013) which is longer than the deepest recorded dive for this species of 293 m that lasted 9.8 min within the Southern California Bight (Calambokidis et al., 2007). Based on this available data, we hypothesize that NIO blue whales are deep divers with the potential to reach depths of approximately 300 m. This coincides with the depths at which some species of Sergestidae have been recorded within the Indian Ocean (Karuppasamy et al., 2010). Further, this coincides with the depth of the deep scattering layer documented within this area during pilot hydroacoustic surveys conducted in March 2012 (see Supplementary Material).
In comparison, spinner dolphins caught in the tuna driftnet fishery in the Sulu Sea dove to 200–300 m to feed on sergestids (Dolar et al., 2003). In Hawaii, spinner dolphins are known to closely track the mesopelagic boundary community as they migrate vertically through the water column diurnally. This community reaches a depth of 400–700 m during the day and is spread between 0 and 400 m at night (Benoit-Bird and Au, 2003).
Baleen whales have an estimated trophic level of 3.2–3.4, with blue whales ranked at the lowest level within that group (Pauly et al., 1998) because of the assumption that they feed on a limited variety of prey like euphausiids that themselves feed on primary producers (Trites, 2001). In the southern ocean, blue whales like other baleen whale species feed on the abundantly available Euphausia superba (Wiedenmann et al., 2011), while within the Bonney upwelling off South Australia, the pygmy blue whale (B. m. brevicauda) feeds predominantly on the indigenous coastal euphausiid, Nyctiphanes australis (Gill, 2002). In Sri Lanka, this species has adapted to feed on the locally abundant Sergestidae, which along with other Dendrobranchiata are generally restricted to warm waters between 40°N and 40°S (Tavares and Martin, 2010). Dendrobranchiata are voracious omnivores that feed on euphausiids and mysids (Tavares and Martin, 2010), changing the trophic position of blue whales in the food chain of the NIO.
The finding that NIO blue whales prey on Sergestidae, a predator of euphausiids, introduces an additional trophic level to the blue whale food web in Sri Lankan waters. This may result in increased bioaccumulation of toxins, the potential for trophic cascades and reduced energy transfer to higher trophic levels. While studies on female blue whales and their offspring in the Gulf of St. Lawrence indicated that bioaccumulation occurs through transplacental and lactational transfer (Metcalfe et al., 2004), a comprehensive review on the impacts of organochlorine contaminants showed no conclusive evidence for any impact on blue whales (O'Shea and Brownell, 1994). Despite the addition of a trophic level in Sri Lankan waters, blue whales remain near the bottom of the food chain and therefore biomagnification would not be expected to have a strong effect on these individuals (O'Shea and Brownell, 1994). The addition of a third trophic level in this system is theoretically predicted to lead to a trophic cascade whereby a decrease in the density of the second trophic level results in an increase in biomass at the first trophic level. However, trophic cascades are rarer in complicated food webs and the addition of an extra species on the density of those at lower trophic levels may be less pronounced (Strong, 1992). Reduced energy transfer could potentially contribute to the shorter length of these blue whales (Mikhalev, 2000), however based on measurements made during Soviet whaling operations, blubber thickness in NIO blue whales during the months of November-December (coinciding with our sample collection period) were comparable to those for blue whales within Antarctic feeding regions (Mikhalev, 1996).
The large size of blue whales, much like that of other low trophic level feeders such as whale sharks confers numerous advantages including increased swimming efficiency, reduced risk of predation, reduced metabolic rate, increased energy storage capacity, and increased foraging range (Kozlowski, 2000), enabling long distance travel between food patches of worthwhile richness (Martin, 2007). However, unlike higher trophic level feeders, the reliance of these wide-ranging, low trophic level feeders on zooplankton (Stevens, 2007; Couturier et al., 2012) strongly links their sightings to environmental variables, as the distribution and abundance of their prey respond rapidly to changes in the environment (Boucher et al., 1987; Richardson, 2008; de Vos et al., 2014a). Further, as this population has adapted to feed on locally available prey the need for long-range movement is reduced, possibly explaining their limited range (Zemsky and Sazhinov, 1994). We hypothesize therefore that while most other baleen whale species feed almost exclusively in high productivity areas of the high latitudes after which they rarely if ever feed during their migration until they return to their feeding grounds (Mackintosh, 1942, 1965) using a “gorger” strategy, blue whales in the NIO have adapted a “grazer” strategy where they feed continuously throughout the year on smaller patches of shrimp over a restricted range in tropical waters that are unusually productive (de Vos et al., 2014b).
A further finding of significance was the detection of acanthocephalans in three of the 16 fecal samples. In particular, the sample from March was dominated by this obligate parasite and likely represented a significant infestation in the individual concerned. This represents the first record of acanthocephalan parasitic worms from the gut flora of blue whales in the Northern Indian Ocean (Measures, 1993). Only two species of two genera have been found to reproduce in the intestines and stomach of marine mammals. These species are similar to those found in aquatic birds (Raga et al., 2009). Bolbosoma spp. are the main genus of acanthocephalan found in the small intestine of large baleen whales (Baylis, 1932). They are not thought to be host specific, however, acanthocephalans with aquatic life cycles use arthropod intermediate hosts such as amphipods, ostracods, or other crustaceans (Measures, 1992). The life cycle of marine acanthocephalans is heteroxenous, involving one or two intermediate hosts, being represented by crustaceans and fishes. It has been suggested that large whales become infected either by the oral uptake of Bolbosoma-infected euphausiid species, or by feeding on a second intermediate host (fishes) (Hermosilla et al., 2015).
Parasitic diseases may have profound influences on individual, population, and ecosystem health. Acanthocephala infections have been shown to cause changes in host phenotype, impacting host behavior and immunity (Gunalan et al., 2013). High parasitic burdens have been associated with intestinal wall performation and mucosal ulceration (Morejohn et al., 1975; Hermosilla et al., 2016). A detailed post mortem of a male Bryde's whale off the east coast of the Malaysian peninsula suggested that Acanthocephala may have resulted in lung, liver, kidney, and heart complications in the individual (Gunalan et al., 2013). Further, Acanthocephala of the species Bolbosoma balanae caused multifocal transmutal abscesses and ultimately contributed to the death of a juvenile gray whale, Eschrichtius robustus, stranded along the northern California coast (Dailey et al., 2000).
DNA metabarcoding analyses revealed the first incidence of blue whales feeding on prey other than euphausiids, which highlights an interesting local adaptation of this species within the NIO. Further, the analysis also revealed the presence of parasitic infections within this population. This adaptation and high specialization within a restricted range coupled with the presence of a range of anthropogenic and natural threats such as ship-strike (de Vos et al., 2016) and parasitic infections, highlights the urgent need for the implementation of conservation measures to protect this population, which is considered a subspecies of the blue whale complex, B. m. indica (SMM Committee on Taxonomy, 2014).
Author Contributions
AdV: Conceived the project, conducted the fieldwork and wrote the paper; SJ: Designed the DNA metabarcoding approach, analyzed the data, and wrote the paper; CF, JM, and JD: Conducted the DNA metabarcoding analyses and reviewed the manuscript. All authors approved final version to be published.
Funding
The fieldwork and DNA extractions were conducted with funds from the Ocean Park Conservation Foundation, Hong Kong, a convocation travel grant from the University of Western Australia and a Duke University Global Fellow's mini grant. DNA metabarcoding analysis was funded by Australian Antarctic Science grant 4014.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
All research reported in this manuscript was conducted under a Department of Wildlife Conservation, Sri Lanka permit (number WL/3/2/1/18). The authors wish to thank Rachel Sullivan-Lord and Salva Yatamar for support in the field; Deishini Herat for extracting prey DNA from fecal samples; Priyan Fernando for assistance with plotting the graphs included in this paper and Alexander Vereshchaka for advice on zooplankton species identification.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmars.2018.00104/full#supplementary-material
References
Afsal, V. V., Yousuf, K. S. S. M., Anoop, B., Anoop, A. K., Kannan, P., Rajagopalan, M., et al. (2008). A note on cetacean distribution in the Indian EEZ and contiguous seas during 2003-07. J. Cetac. Res. Manage. 10, 209–215.
Alling, A. K., Dorsey, E. M., and Gordon, J. C. D. (1991). “Blue whales (Balaenoptera musculus) off the northeast coast of Sri Lanka: distribution, feeding and individual identification,” in Cetaceans and Cetacean Research in the Indian Ocean Sanctuary: Marine Mammal Technical Report 3, eds S. Leatherwood and G. P. Donovan (Nairobi: United Nations Environment Programme Oceans and Coastal Areas Programme Activity Centre), 247–258.
Altschul, S. F., Gish, W., Miller, W., Myers, E. W., and Lipman, D. J. (1990). Basic local alignment search tool. J. Mol. Biol. 215, 403–410. doi: 10.1016/S0022-2836(05)80360-2
Anderson, R. C. (2005). Observations of cetaceans in the Maldives, 1990-2002. J. Cetac. Res. Manage. 7, 119–135.
Au, W. W. L., Fay, R. R., and Popper, A. N. (2000). Hearing by Whales and Dolphins. New York, NY: Springer Science and Business Media.
Ballance, L. T., Anderson, R. C., Pitman, R. L., Stafford, K. M., Shaan, A., Waheed, Z., et al. (2001). Cetacean sightings around the Republic of the Maldives, April 1998. J. Cetac. Res. Manage. 3, 213–218.
Benoit-Bird, K. J., and Au, W. W. L. (2003). Prey dynamics affect foraging by a pelagic predator (Stenella longirostris) over a range of spatial and temporal scales. Behav. Ecol. Sociobiol. 53, 364–373.
Boucher, J., Ibanez, F., and Prieur, L. (1987). Diel and seasonal variations in the spatial distribution of zooplankton populations in relation to the physical structure of the Ligurian Sea front. J. Mar. Res. 45, 133–173. doi: 10.1357/002224087788400891
Branch, T. A., Stafford, K. M., Palacios, D. M., Allison, C., Bannister, J. L., Burton, C. L. K., et al. (2007). Past and present distribution, densities and movements of blue whales Balaenoptera musculus in the Southern Hemisphere and northern Indian Ocean. Mamm. Rev. 37, 116–175. doi: 10.1111/j.1365-2907.2007.00106.x
Brodie, P. F. (1975). Cetacean energetics, an overview of intraspecific size variation. Ecology 56, 152–161. doi: 10.2307/1935307
Brodie, P. F., Sameoto, D. D., and Shelkdon, R. W. (1978). Population densities of euphausiids off Nova Scotia as indicated by net samples, whale stomach contents and sonar. Limnol. Oceanogr. 23, 1264–1267. doi: 10.4319/lo.1978.23.6.1264
Calambokidis, J., Schorr, G., Steiger, G. H., Francis, J., Bakhitiari, M., Marshall, G., et al. (2007). Insights into the underwater diving, feeding and calling behaviour of blue whales from a suction-cup-attached video-imaging tag (CRITTERCAM). Mar. Technol. Soc. J. 41, 19–29. doi: 10.4031/002533207787441980
Calambokidis, J., Steiger, G. H., Straley, J. M., Herman, L. M., Cerchio, S., Salden, D. R., et al. (2001). Movements and population structure of humpback whales in the North Pacific. Mar. Mamm. Sci. 17, 769–794. doi: 10.1111/j.1748-7692.2001.tb01298.x
Clarke, M. R. (1996). The role of cephalopods in the world's oceans - Cephalopods as prey. III. Cetaceans. Philos. Trans. R. Soc. London Ser. B 351, 1053–1065.
Corkeron, P. J., and Connor, R. C. (1999). Why do baleen whales migrate? Mar. Mamm. Sci. 15, 1228–1245.
Couturier, L. I. E., Marshall, A. D., Jaine, F. R. A., Kashiwagi, T., Pierce, S. J., Townsend, K. A., et al. (2012). Biology, ecology and conservation of the Mobulidae. J. Fish. Biol. 80, 1075–1119. doi: 10.1111/j.1095-8649.2012.03264.x
Croll, D. A., Clark, C. W., Calambokidis, J., Ellison, W. T., and Tershy, B. R. (2001). Effect of anthropogenic low-frequency noise on the foraging ecology of Balaenoptera whales. Anim. Conserv. 4, 13–27. doi: 10.1017/S1367943001001020
Croll, D. A., Marinovic, B., Benson, S., Chavez, F. P., Black, N., Ternullo, R., et al. (2005). From wind to whales: trophic links in a coastal upwelling system. Mar. Ecol. Prog. Ser. 289, 117–130. doi: 10.3354/meps289117
Croll, D. A., Tershy, B. R., Hewitt, R. P., Demer, D. A., Fiedler, P. C., Smith, S. E., et al. (1998). An integrated approach to the foraging ecology of marine birds and mammals. Deep Sea Res. Part II Top. Stud. Oceanogr. 45, 1353–1371. doi: 10.1016/S0967-0645(98)00031-9
Dailey, M. D., Gulland, F. M. D., Lowenstine, L. J., Silvagni, P., and Howard, D. (2000). Prey, parasites and pathology associated with the mortality of a juvenile gray whale (Eschrichtius robustus) stranded along the northern California coast. Dis. Aquat. Org. 42, 111–117. doi: 10.3354/dao042111
Denman, K. L., and Powell, T. M. (1984). Effects of physical processes on planktonic ecosystems in the coastal ocean. Oceanogr. Mar. Biol. 22, 125–168.
de Vos, A., Brownell, R. L. Jr., Tershy, B. R., and Croll, D. A. (2016). Anthropogenic threats and conservation needs of blue whales, Balaneoptera musculus indica, around Sri Lanka. J. Mar. Biol. 2016, 12. doi: 10.1155/2016/8420846
de Vos, A., Christiansen, F., Harcourt, R. G., and Pattiaratchi, C. B. (2013). Surfacing characteristics and diving behaviour of blue whales in Sri Lankan waters. J. Exp. Mar. Biol. Ecol. 449, 149–153. doi: 10.1016/j.jembe.2013.09.010
de Vos, A., Clark, R., Johnson, C., Johnson, G., Kerr, I., Payne, R., et al. (2012). Sightings and acoustic detections of cetaceans in the offshore waters of Sri Lanka: March - June 2003. J. Cetac. Res. Manage. 12, 185–193.
de Vos, A., Pattiaratchi, C. B., and Wijeratne, E. M. S. (2014b). Surface circulation and upwelling patterns around Sri Lanka. Biogeosciences 11, 5909–5930. doi: 10.5194/bg-11-5909-2014
de Vos, A., Pattiaratchi, C., and Harcourt, R. (2014a). Inter-annual variability in blue whale distribution off Southern Sri Lanka between 2011 and 2012. J. Mar. Sci. Eng. 2, 534–550. doi: 10.3390/jmse2030534
Dolar, M. L. L., Walker, W. A., Kooyman, G. L., and Perrin, W. F. (2003). Comparative feeding ecology of spinner dolphins (Stenella longirostris) and Fraser's dolphins (Lagenodelphis hosei) in the Sulu Sea. Mar. Mamm. Sci. 19, 1–19. doi: 10.1111/j.1748-7692.2003.tb01089.x
Donaldson, H. A. (1975). Vertical distribution and feeding of sergestid shrimps (Decapoda: Natantia) collected near Bermuda. Mar. Biol. 31, 37–50. doi: 10.1007/BF00390646
Edgar, R. C. (2010). Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461. doi: 10.1093/bioinformatics/btq461
Evans, N. T., Olds, B. P., Renshaw, M. A., Turner, C. R., Li, Y., Jerde, C. L., et al. (2016). Quantification of mesocosm fish and amphibian species diversity via environmental DNA metabarcoding. Mol. Ecol. Resour. 16, 29–41. doi: 10.1111/1755-0998.12433
Fauchald, P., and Tveraa, T. (2003). Using first-passage time in the analysis of area-restricted search and habitat selection. Ecology 84, 282–288. doi: 10.1890/0012-9658(2003)084[0282:UFPTIT]2.0.CO;2
Fiedler, P. C., Reilly, S. B., Hewitt, S. B., Demer, D., Philbrick, V. A., Smith, S., et al. (1998). Blue whale habitat and prey in the California Channel Islands. Deep Sea Res. Part II Top. Stud. Oceanogr. 45, 1781–1801. doi: 10.1016/S0967-0645(98)80017-9
Gill, P. C. (2002). A blue whale (Balaenoptera musculus) feeding ground in a southern Australian upwelling zone. J. Cetac. Res. Manage. 4, 179–184.
Goldbogen, J. A., Calambokidis, J., Oleson, E., Potvin, J., Pyenson, N. D., Schorr, G., et al. (2011). Mechanics, hydrodynamics and energetics of blue whale lunge feeding: efficiency dependence on krill density. J. Exp. Biol. 214, 131–146. doi: 10.1242/jeb.048157
Goldbogen, J. A., Calambokidis, J., Shadwick, R. E., Oleson, E. M., McDonald, M. A., and Hildebrand, J. A. (2006). Kinematics of foraging dives and lunge-feeding in fin whales. J. Exp. Biol. 209, 1231–1244. doi: 10.1242/jeb.02135
Gunalan, S., Kamaliah, G., Chadrawathani, P., Fatiah, M. A., Wan, S., Azizul, M. O., et al. (2013). Post mortem findings in a stranded Bryde's whale on the east coast of peninsular Malaysia. Malays. J. Vet. Res. 4, 37–44.
Haury, L. H., McGowan, J. A., and Wiebe, P. H. (1978). “Patterns and processes in the time-space scales of plankton distributions,” in Spatial Pattern in Plankton Communities, ed J. H. Steele (New York, NY: Plenum Press), 277–328.
Hermosilla, C., Silva, L. M. R., Kleinertz, S., Prieto, R., Silva, M. A., and Taubert, A. (2016). Endoparasite survey of free-swimming baleen whales (Balaenoptera musculus, B. physalus, B. borealis) and sperm whales (Physeter macrocephalus) using non/minimally invasive methods. Parasitol. Res. 115, 889–896. doi: 10.1007/s00436-015-4835-y
Hermosilla, C., Silva, L. M. R., Prieto, R., Kleinertz, S., Taubert, A., and Silva, M. A. (2015). Endo- and ectoparasites of large whales (Cetartiodactyla: Balaenopteridae, Physeteridae): overcoming difficulties in obtaining appropriate samples by non- and minimally-invasive methods. Int. J. Parasitol. 4, 414–420. doi: 10.1016/j.ijppaw.2015.11.002
Higginbottom, I. R., and Hosie, G. W. (1989). Biomass and population structure of a large aggregation of krill near Prydz Bay, Antarctica. Mar. Ecol. Prog. Ser. 58, 197–203. doi: 10.3354/meps058197
Hughes-Hanks, J., Rickard, L., Panuska, C., Saucier, J., O'hara, T., Dehn, L., et al. (2005). Prevalence of Cryptosporidium spp. and Giardia spp. in five marine mammal species. J. Parasitol. 91, 1225–1228. doi: 10.1645/GE-545R.1
Hunt, K. E., Moore, M. J., Rolland, R. M., Kellar, N. M., Hall, A. J., Kershaw, J., et al. (2013). Overcoming the challenges of studying conservation physiology in large whales: a review of available methods. Conserv. Physiol. 1:cot006. doi: 10.1093/conphys/cot006
Ilangakoon, A. D. (2006). Preliminary analysis of large whale strandings in Sri Lanka 1889-2004. Pak. J. Oceanogr. 2, 61–68.
Jarman, S. N., Gales, N. J., Tierney, M., Gill, P. C., and Elliott, N. G. (2002). A DNA-based method for identification of krill species and its application to analysing the diet of marine vertebrate predators. Mol. Ecol. 11, 2679–2690. doi: 10.1046/j.1365-294X.2002.01641.x
Jarman, S. N., McInnes, J. C., Faux, C., Polanowski, A. M., Marthick, J., Deagle, B. E., et al. (2013). Adélie penguin population diet monitoring by analysis of food DNA in scats. PLoS ONE 8:e82227. doi: 10.1371/journal.pone.0082227
Jarman, S. N., and Wilson, S. G. (2004). DNA-based species identification of krill consumed by whale sharks. J. Fish Biol. 65, 586–591. doi: 10.1111/j.0022-1112.2004.00466.x
Karuppasamy, P. K., Muraleedharan, K. R., Dineshkumar, P. K., and Nair, M. (2010). Distribution of mesopelagic micronekton in the Arabian Sea during the winter monsoon. Indian J. Mar. Sci. 39, 227–237.
Kozlowski, J. (2000). “Does body size optimisation alter the allometries for production and life history traits,” in Scaling in Biology, eds J. H. Brown and G. B. West (Oxford: Oxford University Press), 237–252.
Macaulay, M. C., Wishner, K. F., and Daly, K. L. (1995). Acoustic scattering from zooplankton and micronekton in relation to a whale feeding site near Georges Bank and Cap Cod. Cont. Shelf Res. 15, 509–537. doi: 10.1016/0278-4343(94)00058-U
Martin, R. A. (2007). A review of behavioural ecology of whale sharks (Rhincodon typus). Fish. Res. 84, 10–16. doi: 10.1016/j.fishres.2006.11.010
McInnes, J. C., Emmerson, L., Southwell, C., Faux, C., and Jarman, S. N. (2016). Simultaneous DNA-based diet analysis of breeding, non-breeding and chick Adélie penguins. R. Soc. Open Sci. 3:150443. doi: 10.1098/rsos.150443
Measures, L. N. (1992). Bolbosoma turbinella new record acanthocephala in a blue whale Balaenoptera musculus stranded in the St. Lawrence Estuary, Quebec, Canada. J. Helminthol. Soc. Washington 59, 206–211.
Measures, L. N. (1993). Anotated list of metazoan parasites reported from the blue whale, Balaenoptera musculus. J. Helminthol. Soc. Washington 60, 62–66.
Meekan, M. G., Jarman, S. N., McLean, C., and Schultz, M. B. (2009). DNA evidence of whale sharks (Rhincodon typus) feeding on red crab (Gecarcoidea natalis) larvae at Christmas Island, Australia. Mar. Freshw. Res. 60, 607–609. doi: 10.1071/MF08254
Metcalfe, C., Koenig, B., Metcalfe, T., Paterson, G., and Sears, R. (2004). Intra-and inter-species differences in persistent organic contaminants in the blubber of blue whales and humpback whales from the Gulf of St. Lawrence, Canada. Mar. Environ. Res. 57, 245–260. doi: 10.1016/j.marenvres.2003.08.003
Mikhalev, Y. A. (1996). “Pygmy blue whales of the Northern-Western Indian Ocean,” in Paper Presented to the Scientific Committee of the IWC SC/48/SH30 (Moscow).
Mikhalev, Y. A. (2000). “Whaling in the Arabian Sea by the whaling fleets Slava and Sovetskaya Ukraina,” in Soviet Whaling Data (1949–1979), eds A. V. Yablokov and V. A. Zemsky (Moscow: Center for Russian Environmental Policy Marine Mammal Council), 141–181.
Morejohn, G. V., Ames, J. A., and Lewis, D. B. (1975). Post mortem studies of sea otters, Enhydra lutris L., in California. California Department of Fish and Game, Marine Resources Technical Report.
O'Shea, T. J., and Brownell, R. L. Jr. (1994). Organochlorine and metal contaminants in baleen whales: a review and evaluation of conservation implications. Sci. Total Environ. 154, 179–200. doi: 10.1016/0048-9697(94)90087-6
Palomares, M. L. D., and Pauly, D. (2016). SeaLifeBase [Online]. Available online at: www.sealifebase.org
Pauly, D., Trites, A. W., Capuli, E., and Christensen, V. (1998). Diet composition and trophic levels of marine mammals. ICES J. Mar. Sci. 55, 467–481. doi: 10.1006/jmsc.1997.0280
Pérez-Farfante, I., and Kensley, B. F. (1997). Penaeoid and sergestoid shrimps and prawns of the world. Keys and diagnoses for the families and genera. Mémoires du Muséum National d'Histoire Naturelle 175, 1–233.
Pompanon, F., Deagle, B. E., Symondson, W. O. C., Brown, D. S., Jarman, S. N., and Taberlet, P. (2012). Who is eating what: diet assessment using next generation sequencing. Mol. Ecol. 21, 1931–1950. doi: 10.1111/j.1365-294X.2011.05403.x
Raga, J. A., Fernández, M., Balbuena, J. A., and Aznar, F. J. (2009). “Parasites,” in Encyclopaedia of Marine Mammals, eds W. F. Perrin, B. Wursig, and J. G. M. Thewissen (Amsterdam: Elsevier, Academic Press), 821–830.
Reilly, S. B., and Thayer, V. G. (1990). Blue whale (Balaenoptera musculus) distribution in the eastern tropical Pacific. Mar. Mammal Sci. 6, 265–277. doi: 10.1111/j.1748-7692.1990.tb00357.x
Richardson, A. J. (2008). In hot water: zooplankton and climate change. ICES J. Mar. Sci. 65, 279–295. doi: 10.1093/icesjms/fsn028
Samuels, A., and Tyack, P. L. (1999). “Flukeprints,” in Cetacean Societies: Field Studies of Dolphins and Whales, eds J. Mann, R. C. Connor, P. L. Tyack, and H. Whitehead (Chicago, IL: The University of Chicago Press), 9–14.
Schoenherr, J. R. (1991). Blue whales feeding on high concentrations of euphausiids around Monterey submarine canyon. Can. J. Zool. 69, 583–594. doi: 10.1139/z91-088
Sears, R., Doniol-Valcroze, T., Berchock, C., Palsbøll, P. J., Bendlin, A., and Ramp, C. (2009). “Sex structure and male-male competition in blue whale trios from the North Atlantic or simply a “ménage à trois,”” in Eighteenth Biennal Conference on the Biology of Marine Mammals (Quebec City).
Silva, M. A., Prieto, R., Jonsen, I., Baumgartner, M. F., and Santos, R. S. (2013). North Atlantic blue and fin whales suspend their spring migration to forage in middle latitudes: building up energy reserves for the journey? PLoS ONE 8:e76507. doi: 10.1371/journal.pone.0076507
Small, J. A., and Small, G. (1991). “Cetacean observations from the Somali democratic republic, September 1985 through May 1987,” in Cetaceans and Cetacean Research in the Indian Ocean Sanctuary: Marine Mammal Technical Report 3, eds S. Leatherwood and G. P. Donovan (Nairobi: United Nations Environment Programme Oceans and Coastal Areas Programme Activity Centre), 179–210.
SMM Committee on Taxonomy (2014). List of Marine Mammal Species and Subspecies. Available online at: www.marinemammalscience.org (Accessed July 24, 2017).
Stevens, J. D. (2007). Whale shark (Rhincodon typus) biology and ecology: a review of the primary literature. Fish. Res. 84, 4–9. doi: 10.1016/j.fishres.2006.11.008
Strong, D. R. (1992). Are trophic cascades all wet? Differentiation and donor-control in speciose ecosystems. Ecology 73, 747–754. doi: 10.2307/1940154
Tavares, C., and Martin, J. W. (2010). “Suborder Dendrobranchiata Bate, 1888,” in Treatise on Zoology - Anatomy, Taxonomy, Biology: The Crustacea, eds F. R. Schram and J. C. Von Vaupel Klein (Leiden: Koninkliijke Brill NV), 99–164.
Trites, A. W. (2001). “Marine mammal trophic levels and interactions,” in Encyclopedia of Ocean Sciences, ed J. H. Steele (Oxford: Academic Press), 1628–1633.
Vereshchaka, A. L. (2009). Revision of the Genus Sergia (Decapoda: Dendrobranchiata: Sergestidae): Taxonomy and distribution. Galathea Report 22, Moscow: Shirshov Institute of Oceanology; Russian Academy of Sciences. 7–140.
Walters, J. F. (1976). Ecology of Hawaiian sergestid shrimps (Penaeidea). Antarctic Seas XXII 58, 49–91.
Wiedenmann, J., Cresswell, K. A., Goldbogen, J., Potvin, J., and Mangel, M. (2011). Exploring the effects of reductions in krill biomass in the Southern Ocean on blue whales using a state-dependent foraging model. Ecol. Model. 222, 3366–3379. doi: 10.1016/j.ecolmodel.2011.07.013
Willerslev, E., Davison, J., Moora, M., Zobel, M., Coissac, E., Edwards, M. E., et al. (2014). Fifty thousand years of Arctic vegetation and megafaunal diet. Nature 506, 47–51. doi: 10.1038/nature12921
Yochem, P. K., and Leatherwood, S. (1985). “Blue whale - Balaenoptera musculus (Linnaeus, 1758),” in Handbook of Marine Mammals, Vol 3, The Sirenians and Baleen Whales, eds S. Ridgway and R. Harrison (London; Orlando, FL: Academic Press), 1–31.
Keywords: DNA metabarcoding, Sri Lanka, Northern Indian Ocean, feces, Sergestidae, Euphausiidae, acanthocephala, krill
Citation: de Vos A, Faux CE, Marthick J, Dickinson J and Jarman SN (2018) New Determination of Prey and Parasite Species for Northern Indian Ocean Blue Whales. Front. Mar. Sci. 5:104. doi: 10.3389/fmars.2018.00104
Received: 31 May 2017; Accepted: 14 March 2018;
Published: 06 April 2018.
Edited by:
Ellen Hines, San Francisco State University, United StatesReviewed by:
Mark Peter Simmonds, University of Bristol, United KingdomBrittany Hanser, National Marine Fisheries Service (NOAA), United States
Frank Cipriano, Retired, San Francisco, CA, United States
Copyright © 2018 de Vos, Faux, Marthick, Dickinson and Jarman. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Asha de Vos, YXNoYWRldm9zQGdtYWlsLmNvbQ==
 Asha de Vos
Asha de Vos Cassandra E. Faux2
Cassandra E. Faux2 Joanne Dickinson
Joanne Dickinson Simon N. Jarman
Simon N. Jarman