- 1State Key Laboratory of Mariculture Biobreeding and Sustainable Goods, Yellow Sea Fisheries Research Institute, Chinese Academy of Fishery Sciences, Qingdao, China
- 2Laboratory for Marine Fisheries Science and Food Production Processes, Laoshan Laboratory, Qingdao, China
- 3College of Fisheries and Life Science, Shanghai Ocean University, Shanghai, China
- 4School of Marine Science and Fisheries, Jiangsu Ocean University, Lianyungang, China
- 5Seawater Research Department, Putian Institute of Aquaculture Science of Fujian Province, Putian, China
Temperature is an important environmental factor affecting metabolism, growth and life activities of marine bivalves. To reveal the molecular and metabolic responses to heat stress, comparative transcriptomics and metabolomics were performed in the commercially important bivalve, Manila clam Ruditapes philippinarum. Comparative transcriptomics revealed a total of 3980 differential genes (DEGs) mainly involved in energy metabolism, protein processing, glycerolphospholipid metabolism, inoxidizability and anti-apoptosis. Comparative metabolomic analysis identified 293 differential metabolites (DEMs) in the heat-stressed clams, including Acetyl-CoA, Beta-D-Glucose, phosphatidylcholine and free amino acids. For the heat-stress clams, they showed the significantly increased enzyme activities of AKP and ACP, as well as antioxidant-related enzyme activities (SOD and CAT) and oxidative damage substance (MDA). The changes in gene expression of glycolysis-related genes (GLUT1, GLK, HK1 and GPI6) and metabolites (Beta-D-Glucose and Acetyl-CoA) reflect the adjusted energy metabolism in the heat-stressed clams. The protein processing in endoplasmic reticulum may play the important roles in the stability of cell membrane structure during heat adaptation. The increased contents of glycerophospholipid (PC) may be essential to maintain the stability and permeability of cell membrane in the heat-stressed clams. Molecular chaperones (HSP70 and HSP90) may participate in the degradation of misfolded proteins to maintain cell homeostasis during the heat stress. To defense the heat stress, clams also initiate the protection and defense mechanisms by activating antioxidant system, innate immune system and anti-apoptotic system. These findings will shed new lights on molecular and metabolic adaptation to heat stress in the intertidal mollusks in a warming climate at the global scale.
1 Introduction
Temperature is one of the major environmental stressors in marine organism. As the frequency of global extreme weather increases, the temperature in the sea area increases year by year (Bindoff et al., 2019). The molecular and physiological responses of aquatic animals to heat stress in their living environment has drawn more and more attention (Pörtner and Farrell, 2008; Somero, 2010). As previously reported, the metabolic activities, growth development and physiological conditions of aquatic animals have been remarkably affected by the heat stress, especially for the intertidal mollusks (Freytes-Ortiz and Stallings, 2018; Jiang et al., 2019; Dong et al., 2022a). Relative to the swimming ability of fish and crustaceans, most of intertidal mollusks have poor locomotion activity to escape from their local habitat (Trueman, 1983). For mollusks, the appropriate temperature can promote their growth and development, but when temperature exceeds a certain limit, it will negatively affect the normal metabolism and even cause their death (Velasco and Barros, 2008).
The potential effects of heat stress on survival, behavior, physiological and metabolic changes have been widely evidenced in a variety of mollusk species, contributing to the current status of species distribution patterns (Dong et al., 2022a). For mollusks, when temperature exceeded their optimal range, they have to decrease oxygen consumption rate and ammonia removal rate, resulting in the disordered physiological function (Ansell and Sivadas, 1973; Aldridge et al., 1995; Bougrier et al., 1995). As indicated, high temperature can significantly affect the oxygen consumption, ammonia-N excretion, enzyme activities, as well as valve closing behavior of scallop Mizuhopecten yessoensis (Hao et al., 2014). The aerobic metabolism of abalone Haliotis iris was remarkably limited under the heat stress, mainly due to the reduced binding ability of hemocyanin to oxygen (Wells et al., 1998). In order to cope with heat stress, the clam Chamelea gallina increased their metabolic intensity, leading to aggravated oxidative damage (Monari et al., 2007). The network analysis of oyster transcriptome revealed a cascade of gene responses during recovery after heat shock, including senescence-associated protein, inhibitor of apoptosis protein, cAMP response element-binding protein, and heat shock protein 70 (Zhang et al., 2012b). Despite this, the capacity of mollusks to adapt to heat stress from molecular to metabolic functions remains largely unknown in most species. Therefore, it is essential to conduct the integrative studies on different levels to illustrate the adaptive strategy in response to heat stress in mollusks.
Transcriptomics is a discipline that reveals complex biological pathways and molecular mechanisms of character regulatory networks through comprehensive research on gene expression and regulatory laws (Manalo et al., 2005). Transcriptomics has been widely used in molecular research of mollusks, and has been reported in oyster Crassostrea gigas (Meng et al., 2013), clam Ruditapes philippinarum (Nie et al., 2020), clam Panopea globose (Juárez et al., 2018), and scallop Patinopecten yessoensis (Jiang et al., 2018). Owing to recent technical advances, metabolomics has been utilized extensively for the identification of single metabolites, representing the apogee of the omics trilogy (Patti et al., 2012). However, metabolomics has been rarely applied to in the field of mollusks studies. Association analysis of different omics helps reveal more complex biological regulation mechanisms (Gracey and Connor, 2016; Xu et al., 2016). Therefore, the combination of multiple omics such as transcriptomics and metabolomics may help to further clarify the molecular and metabolic mechanisms of bivalve mollusks in response to heat stress.
The Manila clam R. philippinarum has the advantages of fast growth and high nutritional value, and is an important economic mollusk cultivated in coastal areas (Zhao and Zhang, 2016; Nie et al., 2017; Yan et al., 2019). Due to the rising of global temperatures, summer temperatures can frequently threaten the survival of this species (Zhang and Yan, 2006; Dong et al., 2022a). Although gene expression responses to thermal stress has been reported, the critical information about thermal effects at different levels is still lacking in clams (Nie et al., 2017; Ding et al., 2018). In this study, comparative transcriptome and metabolomics were combined to reveal the molecular and metabolic mechanisms of clams in response to heat stress, and further understand the thermal sensitivities of clams and their capacities to adapt to temperature changes. The findings of this study are of great significance to the study of molecular mechanism of aquatic organisms and the practice of aquaculture.
2 Materials and methods
2.1 Animal collection and acclimation
In this study, clams were collected from aquaculture ponds in Qingdao, and healthy individuals were selected for subsequent experiments. The clams (shell length 31.83 ± 1.94mm) were raised in a seawater bath for seven days before the start of the experiment. After feeding, the clams were randomly divided into two groups, including a control group (20°C) and a heat stress group (30°C). The heat stress experiment was set up according the previous studies, which showed the decreased survival and depressed physiological condition of heat-stressed clams at 30°C, compared to the healthy condition at 20°C (Han et al., 2008; Munari et al., 2011; Zhang et al., 2023). After 24 h of stress, whole-tissue samples from nine clams in each group were used for metabolite extraction in the metabolome (nine biological repetitions), and three samples of the same amount in two group were mixed for RNA extraction in the transcriptome (three biological repetitions). The whole-tissue sampling was performed to investigate transcripts and metabolite changes of clams in response to heat stress at the whole-organism level. According to the results of transcriptomics, we selected six enzyme activities and nine genes for subsequent experiments. The gill tissues of clam under heat stress (three biological repetitions) for 24 h, 48 h, 72 h, 96 h and 120 h were taken for enzyme activity determination, reference gene screening and gene expression determination. Ethical review and approval was not required for the study on clams in accordance with the local legislation and institutional requirements.
2.2 RNA extraction and transcriptome sequencing
The total RNA from the entire tissue sample was extracted using Trizol reagent. The concentration and purity of total RNA were detected by NanoPhotometer spectrophotometer and agarose gel electrophoresis. The qualified total RNA was used in subsequent experiments. In short, RNA was randomly disrupted at elevated temperature and the fragmented RNA was used as a reverse transcription template to produce a second cDNA strand, which was sequenced using the Illumina HiSeqTM 4000 platform. After filtering the original data, the remaining clean reads were further mapped to the reference genome. Differential expressed genes (DEGs) between the two groups were screened using DESeq2 software. We selected DEGs according to the following criteria: FDR < 0.05, |log2FC| > 1 (FDR: false discovery rate; FC: fold change). All the DEGs were then mapped to the GO term (FDR < 0.05) to characterize its primary biological function. Enrichment of the KEGG pathway was performed to identify the major biochemical and signal transduction pathways in DEGs.
2.3 Metabolome sample treatment and LC-MS/MS detection
Metabolites were extracted from 50 mg treated clam samples. Metabolites were extracted conventionally using a mixture of 1 ml of acetonitrile and methanol according to our previous study (Sun et al., 2021). After milling, low-temperature ultrasonic treatment and centrifugation, the sample was transferred to the sample bottle for subsequent analysis. LC-MS/MS analysis was performed using an UHPLC (1290, Agilent Technologies). For the positive mode, mobile phase A consists of 0.1% aqueous formic acid, and mobile phase B consists of acetonitrile. For the negative mode, mobile phase A was composed of 5mM ammonium acetate in water and mobile phase B was set as acetonitrile. Isolate the compound at 0.5 mL/min using the gradient method. The XCMS software was used for the correction of retention time correction, peak recognition, peak extraction, peak integration and peak alignment. The OPLS-DA (Orthogonal projection to latent structures-discriminant analysis) model was used in the following analysis. The differential metabolites (DEMs) were screened according to the combination of P value of t-test and VIP (variable importance in the projection) of the OPLS-DA model. The screening criteria were set as P value < 0.05 and VIP ≥ 1. The KEGG analysis was then used for the pathway enrichment of DEMs.
2.4 The integrated analysis of transcriptomic and metabolomic data
The transcriptomic and metabolomic datasets were combined for the integrated analysis. For transcriptomics and metabolomics association analyses, we used KEGG pathway analysis to determine the association between the transcript and the metabolite. Based on previous studies, we also analyzed the association between the transcript and the metabolite using different methods, including Two-way Orthogonal PLS O2PLS (Bylesjö et al., 2007), the joint loading plots (Bouhaddani et al., 2016) and pearson correlation coefficients (Copley et al., 2017).
2.5 Enzyme activity analysis
The enzyme activities including catalase (CAT), malondialdehyde (MDA), Na+K+-ATPase (NAK), alkaline phosphatase (AKP), acid phosphatase (ACP) and superoxide dismutase (SOD) were determined in gill of clams. All these enzyme activities were detected using the corresponding kits (Aidisheng, Jiangsu, China).
2.6 Selection of reference gene
In order to obtain accurate gene expression results, the optimal internal reference gene for clam under heat stress was screened out. The total RNA in the gills of clam at five sampling time points in the two groups was extracted by TRIzol. The purity and quality of RNA were confirmed by NanoPhotometer spectrophotometer (Implen, CA, USA). The cDNA was synthesized using the HiScript III RT SuperMix for qPCR (+gDNA wiper) (Vazyme, China) according to the instructions of the manufacture. In this study, the most appropriate internal reference gene was selected from seven common internal reference genes, including β-actin (ACT), elongation factor 1-α (EF1A), tubulin (TUB), ribosomal protein L31 (RPL31), ribosomal protein S23 (RPS23), histone (HIS) and growth factor receptor-binding protein 2 (GFRP2). The primers were designed by Primer3 online (https://bioinfo.ut.ee/primer3-0.4.0/) and synthesized by Sangon Biotech (Shanghai) (Supplementary Table S1).
The qRT-PCR reactions were amplified in CFX Connect Real-Time PCR Detection System. The reaction solution included 10 μL 2×ChamQ STBR Color qPCR master mix (Vazyme, China), 0.4 μL each for forward and reverse primer, 2 μL cDNA template, 7.2 μL ddH2O. The reaction procedure of qRT-PCR was 95°C for 30 s, 40 cycles at 95°C for 10 s and 60°C for 30 s. The reverse transcribed cDNA was diluted into five gradients to obtain a standard curve and calculate the amplification efficiency in qRT-PCR instrument. GeNorm (Vandesompele et al., 2002), NormFinder (Andersen et al., 2004) and BestKeeper (Pfaffl et al., 2004) were used to statistically analyze the stability of gene expression for each candidate reference gene. The best candidate reference gene ranking was obtained by RefFinder (http://www.leonxie.com/reference gene.php).
2.7 Gene expression level determination
In order to verify the expression patterns of important genes, nine genes were selected for qRT-PCR analysis. The selected genes include energy metabolism genes, including succinate dehydrogenase (SDH), phosphoenolpyruvate carboxykinase (PEPCK) and ADP-dependent glucokinase (ADPGLK). Four genes involved in protein processing, including nucleotide exchange factor SIL1 (SIL1), heat shock protein 70 (HSP70), GTP-binding protein SAR1B (SAR1B) and heat shock protein 90 (HSP90). Furthermore, there are also antioxidation gene cytochrome P450 (CYP450) and immune gene nuclear factor kappa-B (NF-κB). The screened optimal reference gene was used as an internal control, and relative gene expression levels were analyzed by the comparative 2-ΔCT method. The related information on the selected genes and relevant primers was listed in Supplementary Table S2.
2.8 Data analysis
Significant differences in enzyme content and gene expression between control group and heat stress group were assessed by t-tests. Results are presented as mean ± standard error of the mean.
3 Results
3.1 Transcriptome analysis of clam under heat stress
A total of 3980 DEGs including 1876 up-regulated genes and 2104 down-regulated genes were identified between the control and heat stress groups. In order to directly observe the expression pattern of DEGs between the two groups, a volcanic map with FDR and log2FC values was drawn (Figure 1A). The genes with the greatest difference in expression between the heat stress group and control group were located at both ends of the horizontal axis, including lymphocyte antigen 6D, c-type lectin, serine/threonine-protein kinase ULK4, receptor-type tyrosine-protein phosphatase mu and serine protease inhibitor Cvsi. The heat map shows the DEGs expression levels between the control group (C1, C2 and C3) and the heat stress group (T1, T2 and T3) (Figure 1B). Gene expression levels in different samples were shown as light blue (low expression level) to dark red (high expression level). Three samples in the control group were divided into one category, and three samples in the heat stress group were divided into another category.
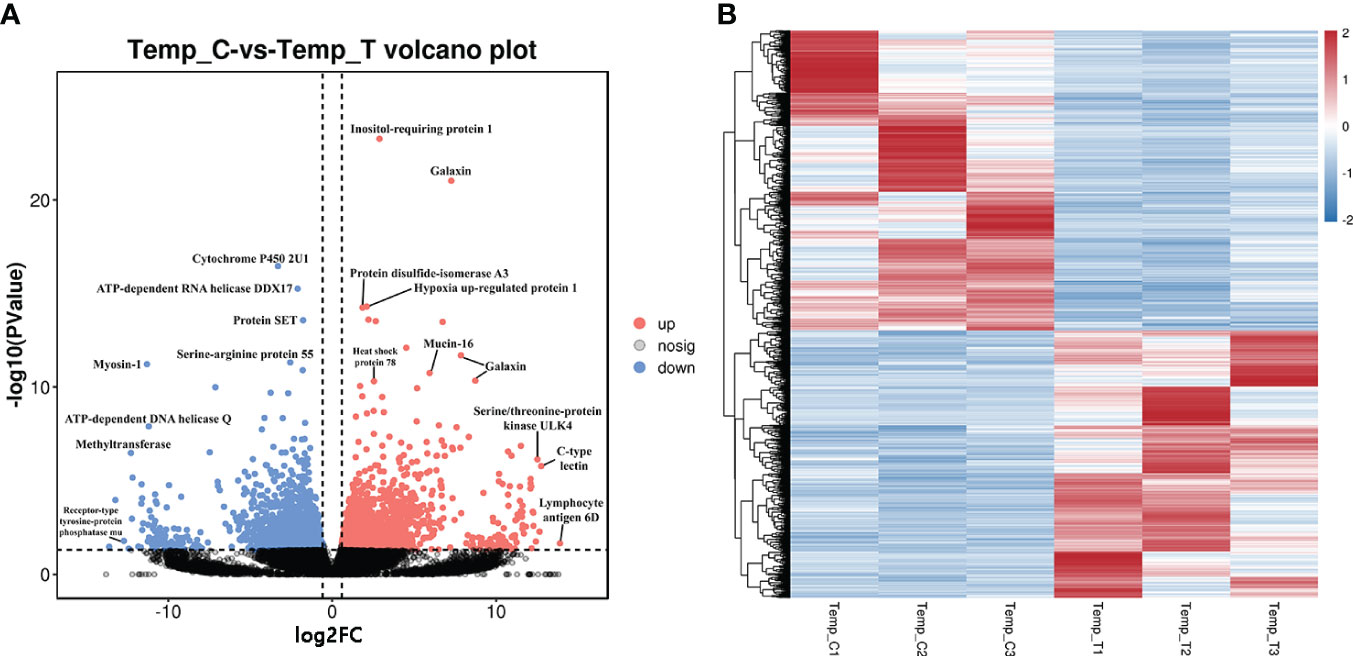
Figure 1 The volcanic plot and heat map of DEGs for the clam R. philippinarum in response to heat stress. (A) The volcanic plot shows the distribution of FDR and log2FC values of the DEGs, Red dots, significantly up-regulated genes; Blue dots, significantly down-regulated genes; Black dots, no significant difference. (B) The heat map of DEGs of the clam, the expression level of genes in different samples is represented by different colors. The scale at the top right denoted normalized expression levels.
GO annotation and KEGG enrichment analysis were carried out to further explore the biological processes and metabolic pathways for these DEGs. For the biological process, most genes were enriched in three GO term processes: cellular process, metabolic process and single-organism process. For the cellular component, most DEGs were concentrated in cell, cell part and membrane. For the molecular functions, most of DEGs were significantly enriched in the subcategory of binding and catalytic activity (Figure 2). The KEGG analysis enriched several important metabolic pathways, such as “protein processing in endoplasmic reticulum”, “olfactory transduction”, “inflammatory mediator regulation of TRP channels”, “glucagon signaling pathway”, “cGMP - PKG signaling pathway” and “spliceosome” (Figure 3).
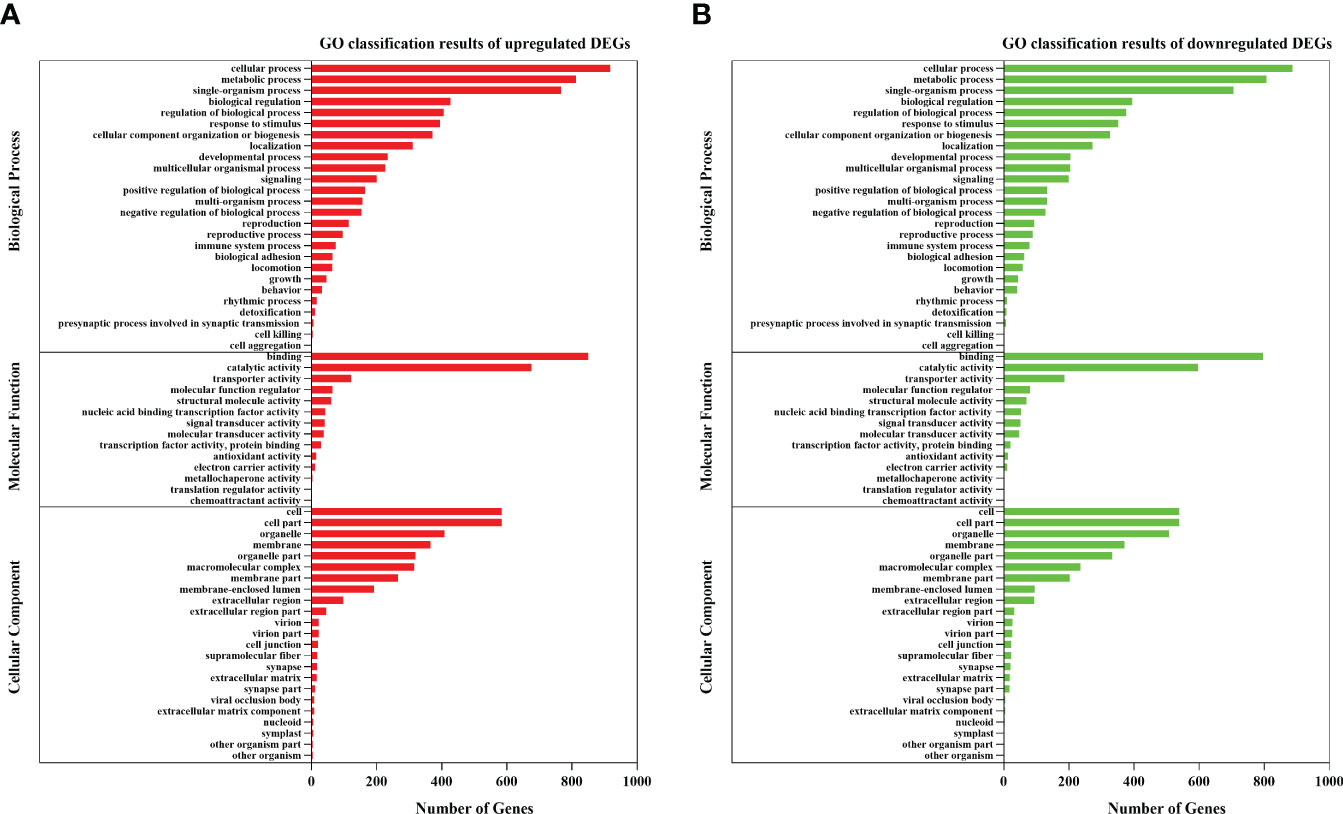
Figure 2 GO classification results for the DEGs of the clam R. philippinarum in response to heat stress. (A) GO classification results of upregulated DEGs. (B) GO classification results of downregulated DEGs.
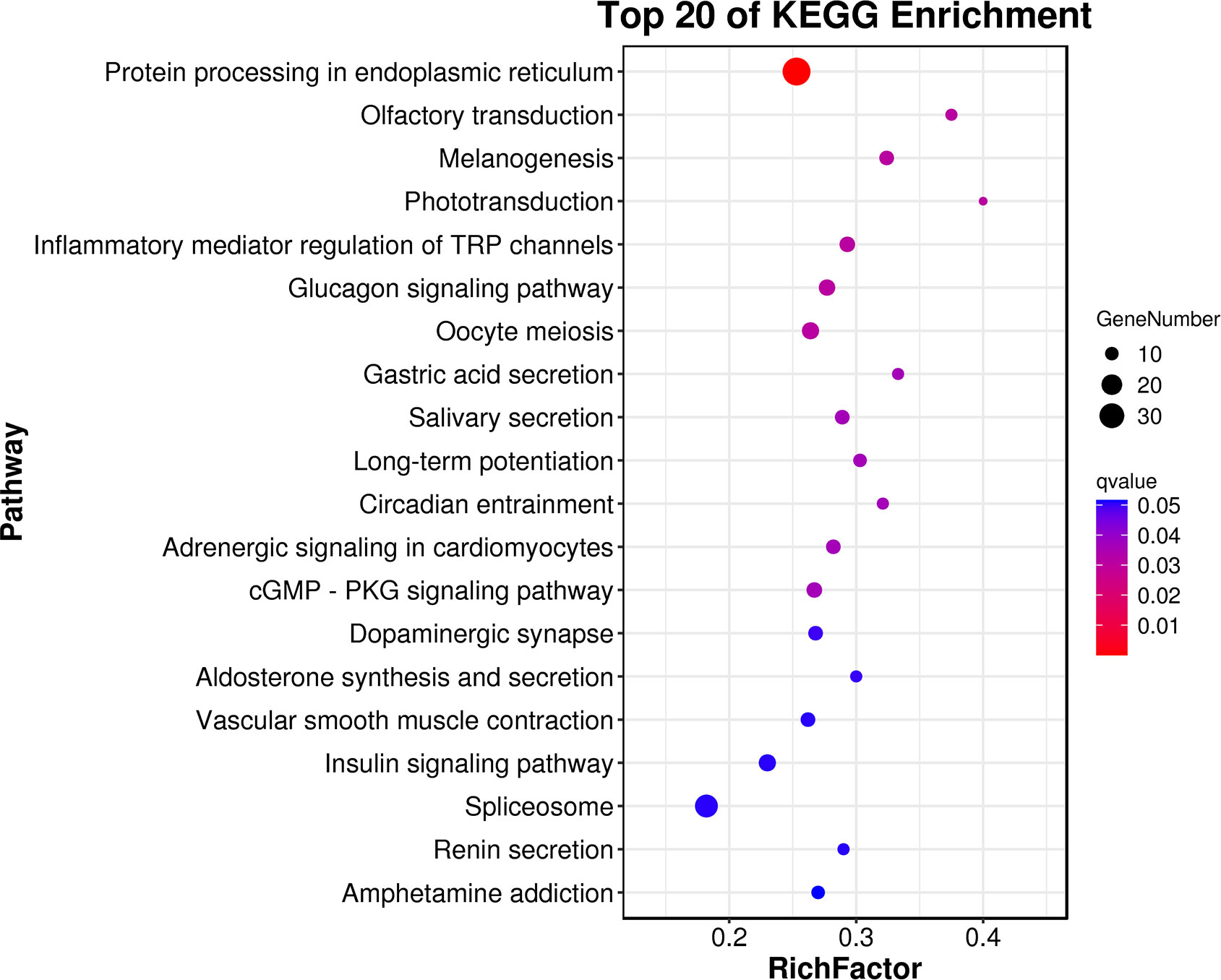
Figure 3 The top20 KEGG enrichment results for the DEGs of the clam R. philippinarum in response to heat stress.
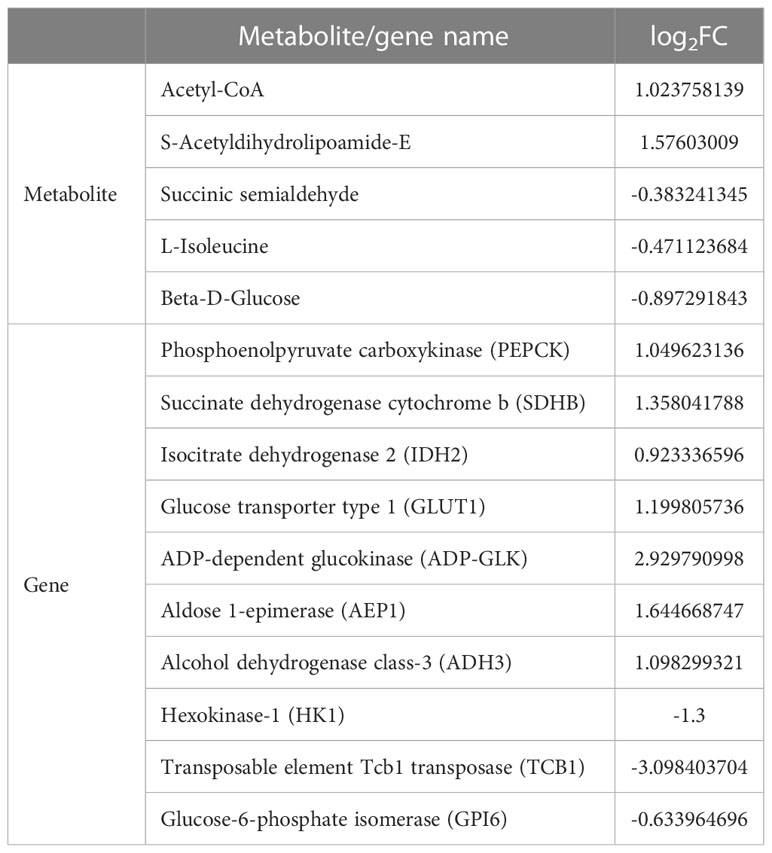
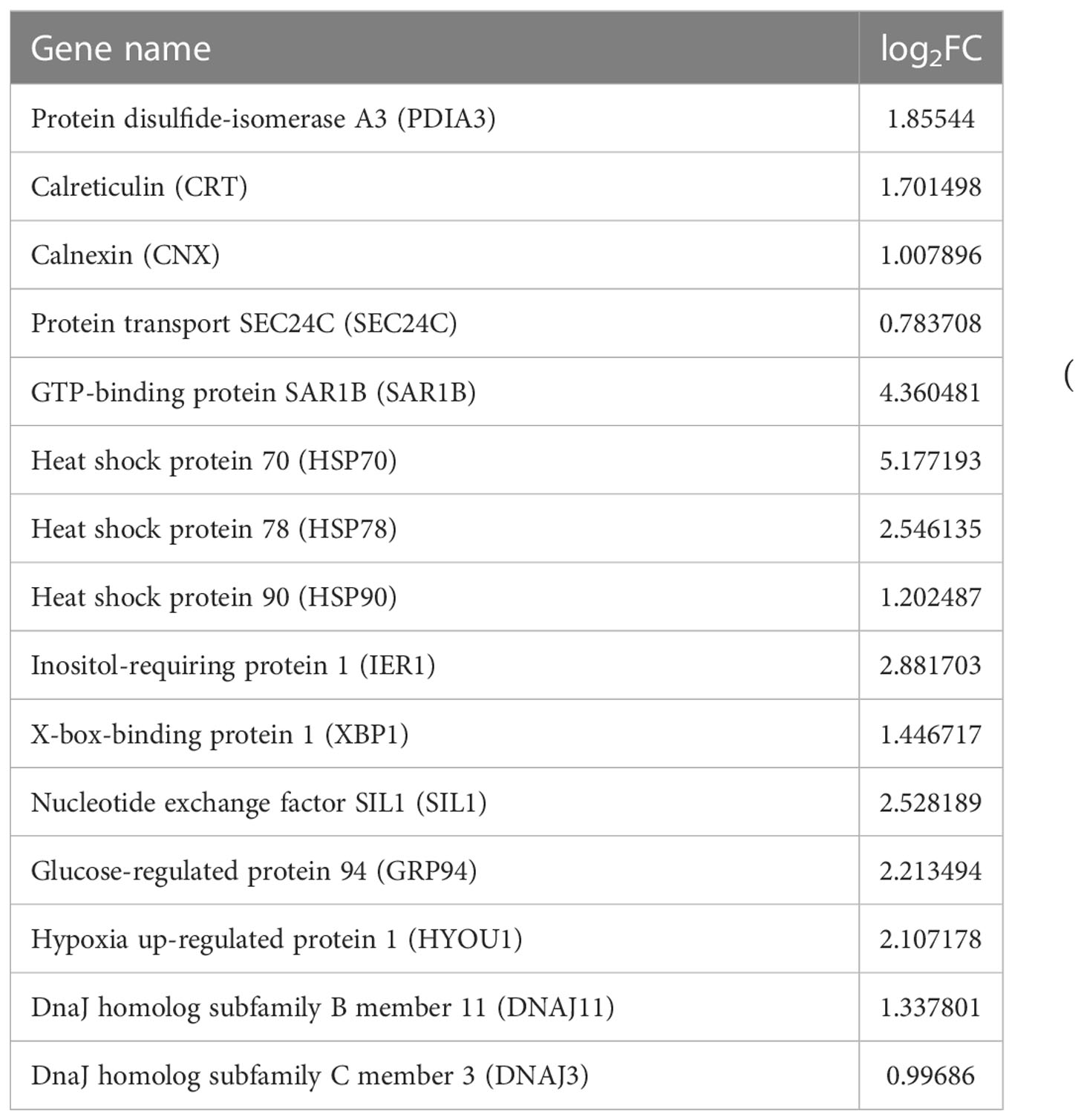
According the comparative transcriptomics, many DEGs were involved in energy metabolism, including phosphoenolpyruvate carboxykinase (PEPCK), succinate dehydrogenase cytochrome b (SDHB), glucose transporter type 1 (GLUT1), ADP-dependent glucokinase (ADP-GLK), glucose-6-phosphate isomerase (GPI6), and etc. (Table 1). A lot of DEGs related to protein processing were also detected in the heat-stressed clams, such as calreticulin (CRT), protein transport SEC24C (SEC24C), heat shock protein 70 (HSP70B2), X-box-binding protein 1 (XBP1), and DnaJ homolog subfamily C member 3 (DNAJ3; Table 2). As summarized in Table 3, multiple DEGs were found to be involved in substance metabolism (e.g. glycerophospholipid), including phosphoethanolamine N-methyltransferase 1 (PEMT1), Stearoyl-CoA Desaturase (SCD) and fatty acid synthase (FAS). Additionally, we detected some DEGs involved in inoxidizability, immunology and anti-apoptotic, such as cytochrome P450, glutathione S-transferase 1 (SGST), C-type lectin (CLEC), and E3 ubiquitin-protein ligase XIAP (Table 4).
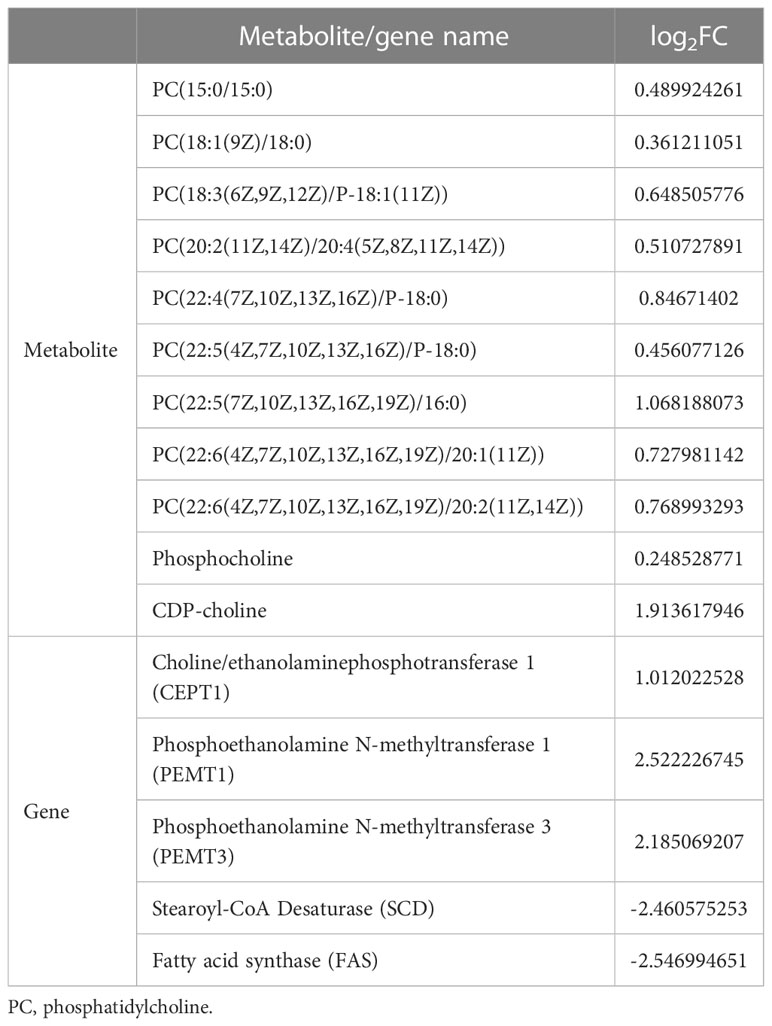
Table 3 The substance metabolism-related DEMs and DEGs list of clam R. philippinarum under heat stress.
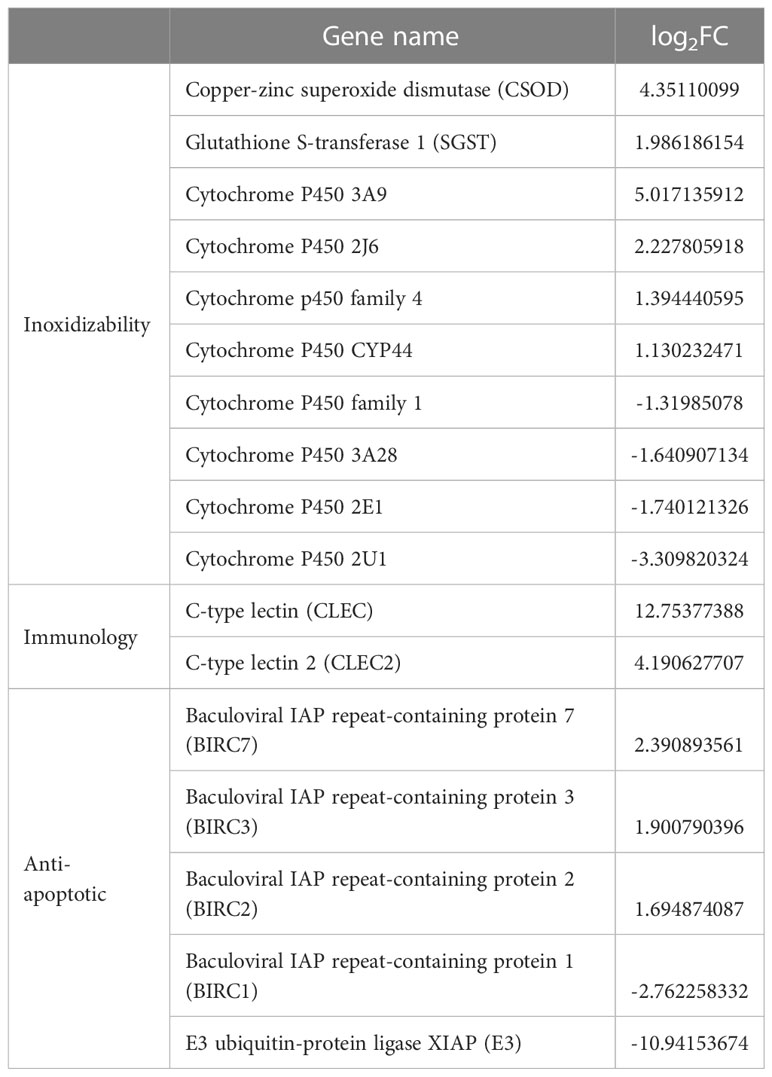
Table 4 Inoxidizability, immunology and anti-apoptotic-related DEGs list of clam R. philippinarum under heat stress.
3.2 Metabonomic analysis of clam under heat stress
In the metabolome analysis, positive ion (POS) and negative ion (NEG) modes were used to collect LC-MS information. First, basal peak chromatograms (BPC) of all samples were visually examined to provide an overall analysis of the metabolites in the control and heat stress groups. Based on OPLS-DA model analysis, Figures 4A, B show significant separation of each sample cluster between the two groups. For positive modes, values for categorical parameters were R2X = 0.235, R2Y = 0.868, Q2Y = 0.108, and for negative modes, R2X = 0.473, R2Y = 0.762, Q2Y = 0.003.
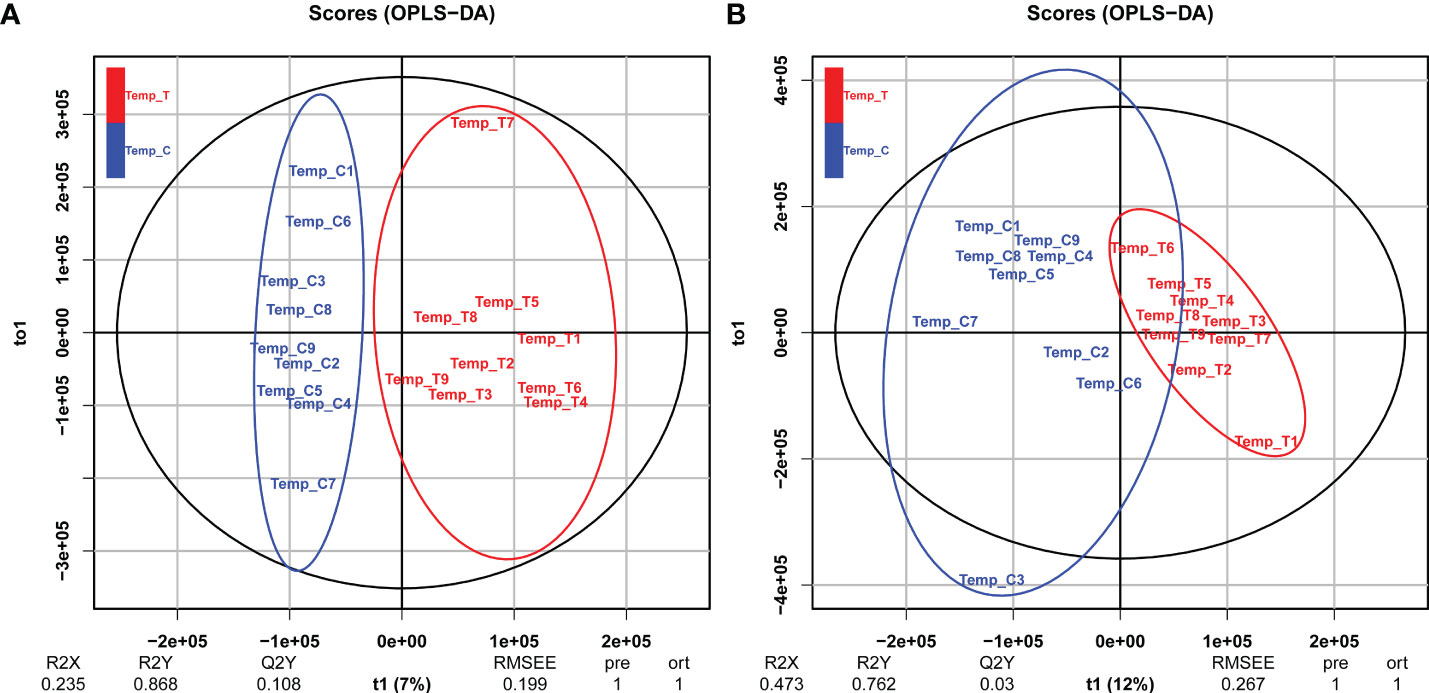
Figure 4 OPLS-DA score plots for the clam R. philippinarum in response to heat stress. (A) positive mode; (B) negative mode.
There are 293 kinds of DEMs detected in positive and negative ion modes, including 213 in the positive mode and 80 in the negative mode. For the positive mode, 176 up-regulated and 37 down-regulated metabolites were found in the heat-stressed group compared with that in the control group. For the negative mode, 37 up-regulated and 43 down-regulated metabolites were detected in the heat-stressed group compared with that in the control group. As a result, the top 20 enrichment pathways of KEGG enrichment analysis of DEMs mainly related to “fatty acid metabolism”, “glycerophospholipid metabolism”, “linoleic acid metabolism”, “valine, leucine and isoleucine biosynthesis”, “fatty acid biosynthesis” and “HIF-1 signaling pathway” (Figure 5). The energy metabolism-related DEMs related to heat stress was summarized in Table 1, including succinic semialdehyde, L-Isoleucine and Beta-D-Glucose and Acetyl-CoA. In addition, there were many DEMs involved in substance metabolism, including phosphatidylcholine (PC), phosphocholine and CDP-choline (Table 3).
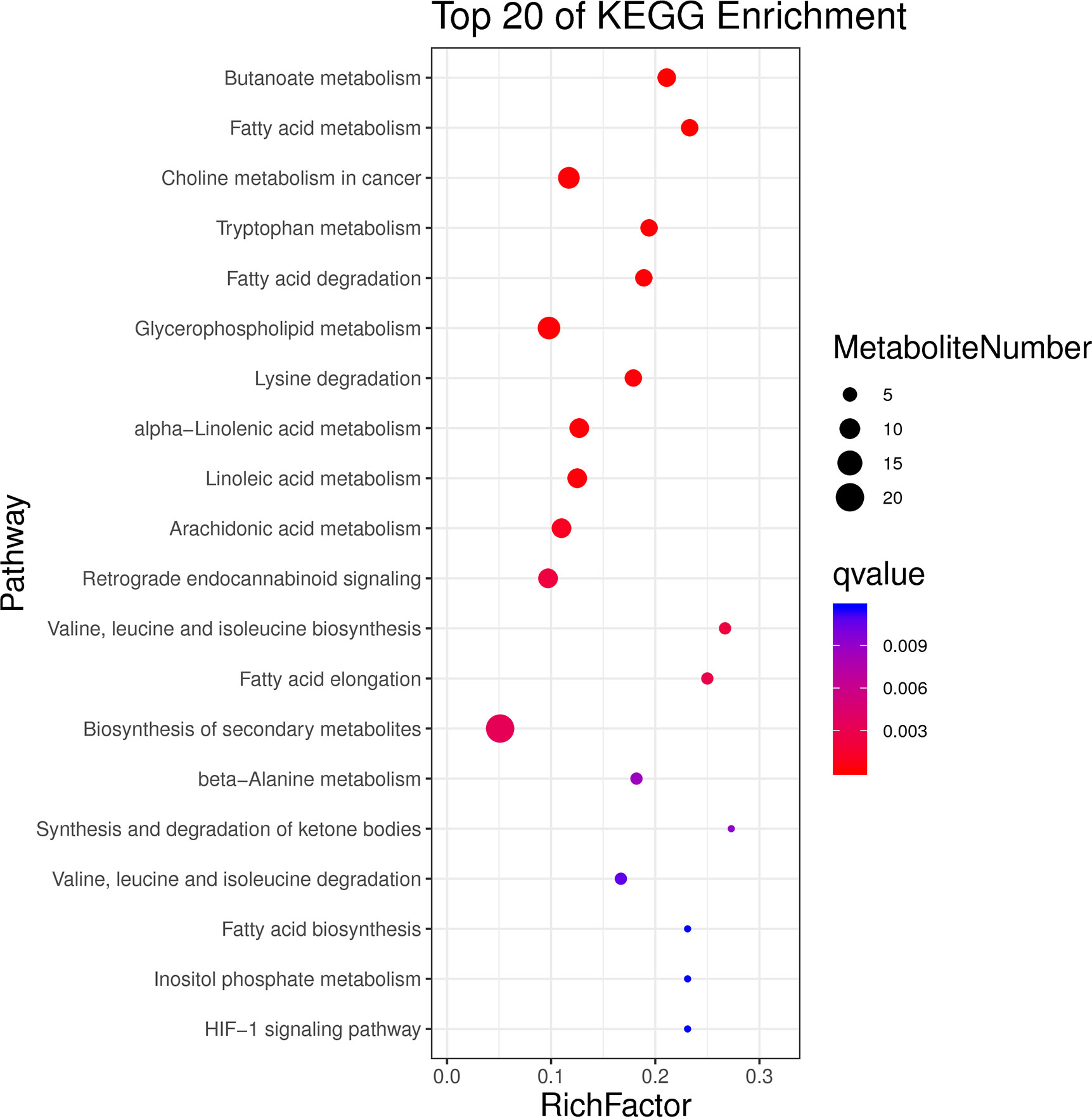
Figure 5 The top20 KEGG enrichment results for the DEMs of the clam R. philippinarum in response to heat stress.
3.3 Correlation analysis of transcriptome and metabolome of clam under heat stress
KEGG analysis showed that a total of 53 functional pathways might be involved in heat stress. The top 20 of enrichment pathway are mainly associated with “glucagon signaling pathway”, “alanine aspartate and glutamate metabolism”, “purine metabolism”, “arachidonic acid metabolism”, “glycolysis” and “pentose phosphate pathway”. The effect of association analysis was assessed using an O2PLS model based on previous studies (Bouhaddani et al., 2016). The O2PLS model calculations (control vs. heat stress) were calculated as R2Xcorr = 0.953, R2Ycorr = 0.926, which reflected that the obtained O2PLS model was a reliable model for correlation analysis of transcriptome and metabolome. The joint loading plot is shown in Figures 6A, B. The metabolite joint loading revealed many metabolites closely related to genes, such as phosphatidylcholine, Homo-L-arginine, Acetyl-CoA, L-Histidine and L-Isoleucine (Figure 6A). The transcript joint loading uncovered thousands of genes closely related to metabolites, such as small heat shock protein, heat shock protein 70, inhibitor of apoptosis 1, Cyclic GMP-AMP synthase, glutamine–tRNA ligase and cytochrome family genes (Figure 6B). In this study, pearson correlation coefficient of DEGs and DEMs was calculated. In order to show the correlation characteristics, the top 250 DEGs and DEMs correlations of correlation coefficients are selected to show in heat map (Supplementary Figure S1). Metabolite and gene data from the heat map are presented in Supplementary File S1.
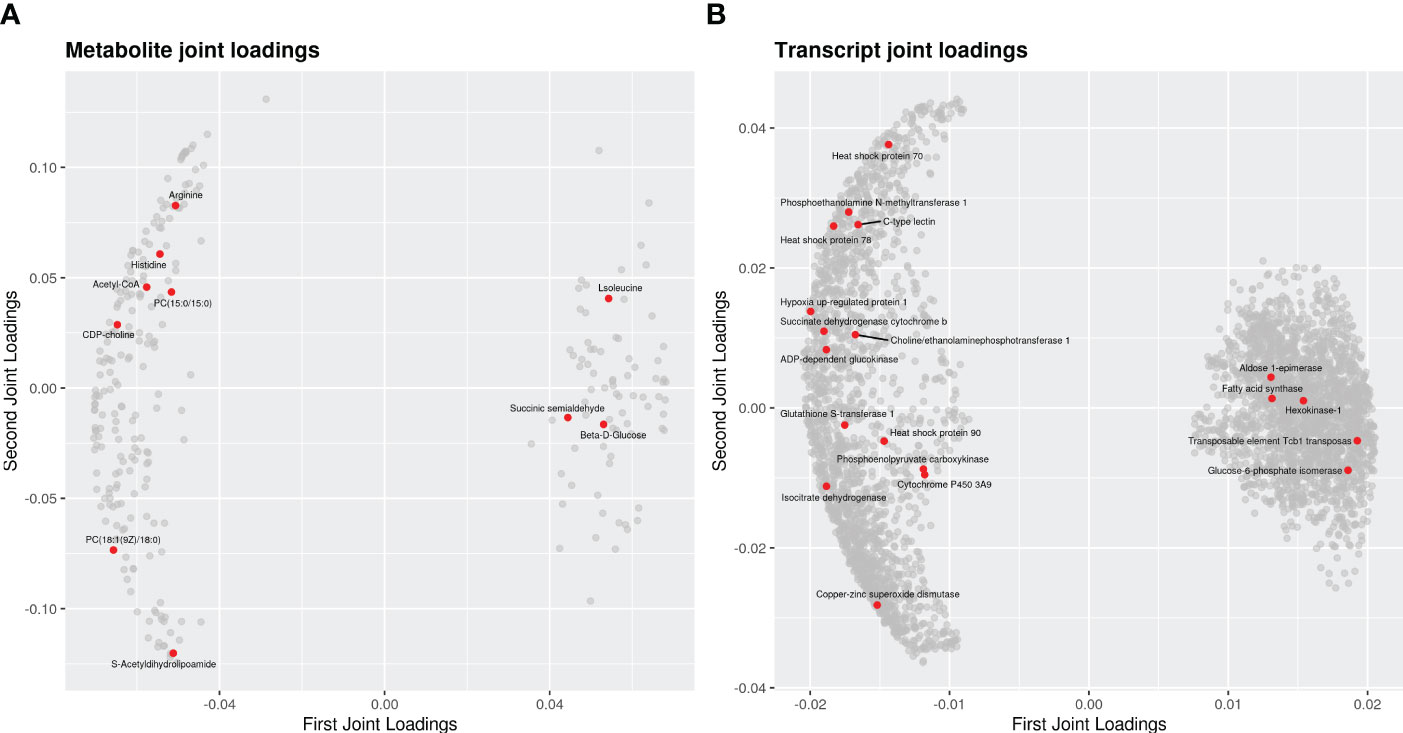
Figure 6 The loading plots for the differential genes and metabolites of the clams in response to heat stress. (A) metabolite joint loadings; (B) transcript joint loadings. Each dot represents a metabolite or a gene. The horizontal axis is the first-dimension coordinate of the joint part, and the ordinate axis of is the second-dimension coordinate of the joint part. The absolute values of an element in the coordinate represent the degree of association between this element and another omics.
3.4 Selection for the optimal internal reference gene under heat stress
After analysis, the amplification efficiency of the candidate reference gene is between 90.4% and 93.8% (Supplementary Table S1). After qRT-PCR analysis, a single peak in the melting curve was observed for each reference gene. Because GFRP2 could not detect Ct values in some samples, it was excluded from the ranking. The Ct values of the remaining candidate reference genes ranged from 21.58 to 35.49, with the greatest change in ACT and the smallest change in RPS23 (Supplementary Figure S2). Stability of candidate reference genes was ranked by geNorm: RPL31/HIS > TUB > RPS23 > EF1A > ACT; Normfinder: RPL31 > TUB > HIS > EF1A > RPS23 > ACT; Bestkeeper: RPS23 > HIS > TUB > RPL31 > EF1A > ACT. The RefFinder was used to consolidate these results from different programs, resulting in the final ranking as following: RPL31 > HIS > TUB > RPS23 > EF1A > ACT (Supplementary Table S3).
3.5 Gene expression pattern by qRT-PCR analysis
According to qRT-PCR analysis, it was found that the expression levels of all target genes tended to increase after heat stress (Figure 7). For instance, SDH expression was significantly increased at 48 h, 72 h and 120 h. In contrast, PEPCK and ADPGLK were significantly increased at 24 h and 48 h. Moreover, the significantly up-regulated expression of SAR1B was found at 24 h, 48 h, and 120 h, and SIL1 at 24 h. For HSP70, HSP90 and CYP450, the significantly increased expression was detected at 24 h, 48 h and 72 h. For the immune gene (NF-κB), it was found to be significantly up-regulated at all of the time points.
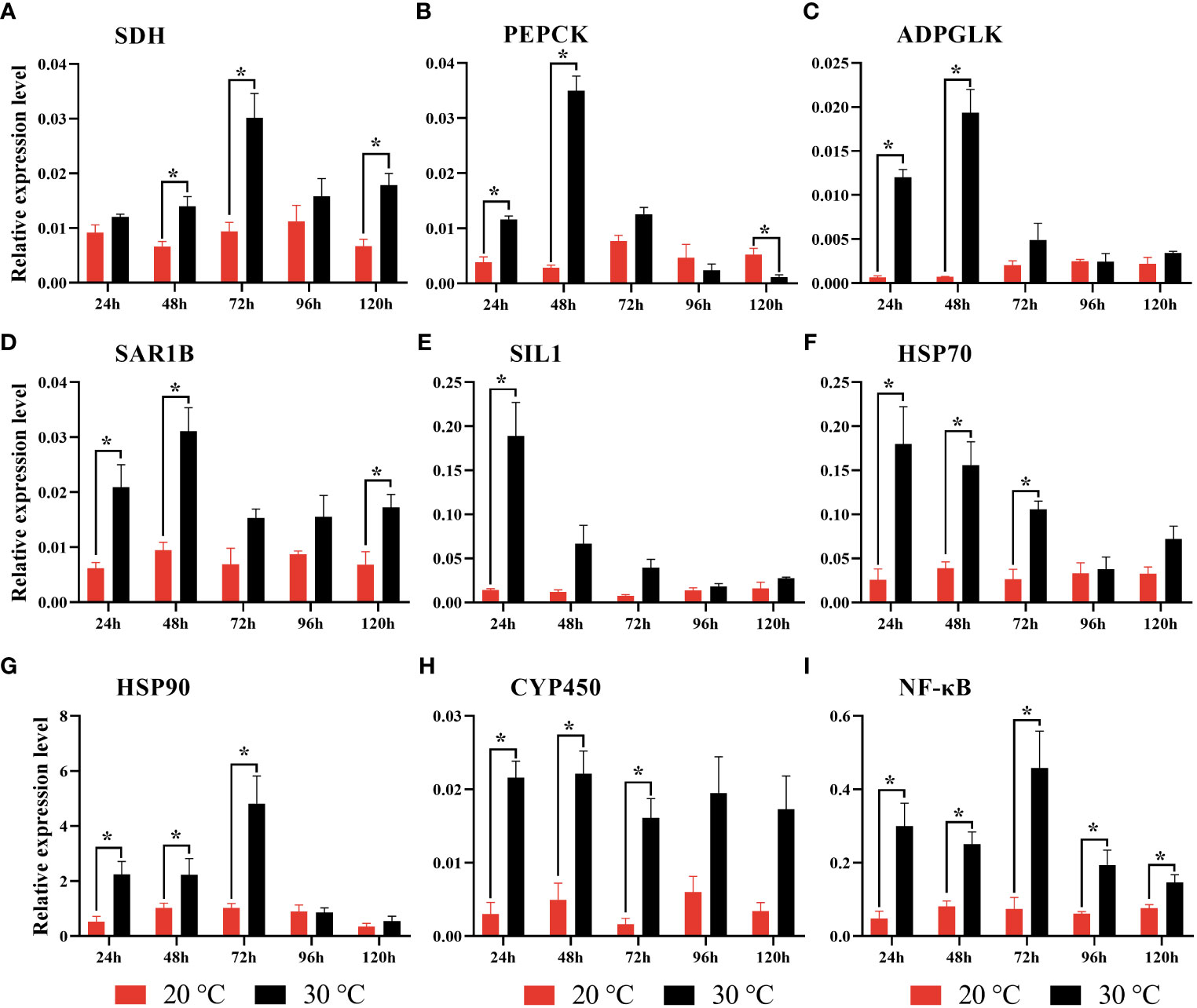
Figure 7 Gene expression revealed by qRT-PCR in gill tissue of clam R. philippinarum after heat stress. Values are mean ± standard error of the mean. Results are normalized against the RPL31 gene. Significance was considered at *p < 0.05. (A) Succinate dehydrogenase (SDH); (B) Phosphoenolpyruvate carboxykinase (PEPCK); (C) ADP-dependent glucokinase (ADPGLK); (D) GTP-binding protein SAR1B (SAR1B); (E) Nucleotide exchange factor SIL1 (SIL1); (F) Heat shock protein 70 (HSP70); (G) Heat shock protein 90 (HSP90); (H) Cytochrome P450 (CYP450); (I) Nuclear factor kappa-B (NF-κB).
3.6 Enzyme activity
According to enzyme activity analysis in clams, enzymes related to substance metabolism and energy metabolism were significantly affected by the heat stress. For instance, the enzyme activity of NAK was significantly decreased at 48 h and 72 h (Figure 8). AKP content increased significantly at 48 h and 72 h and decreased at 120 h, while ACP content increased significantly at 24 h and 48 h. Regarding to antioxidant defenses, we found that antioxidant enzymes of SOD and CAT were significantly increased at 48 h, 72 h and 96 h. The MDA content of oxidative damage substances increased significantly at 48 h and 72 h (Figure 8).
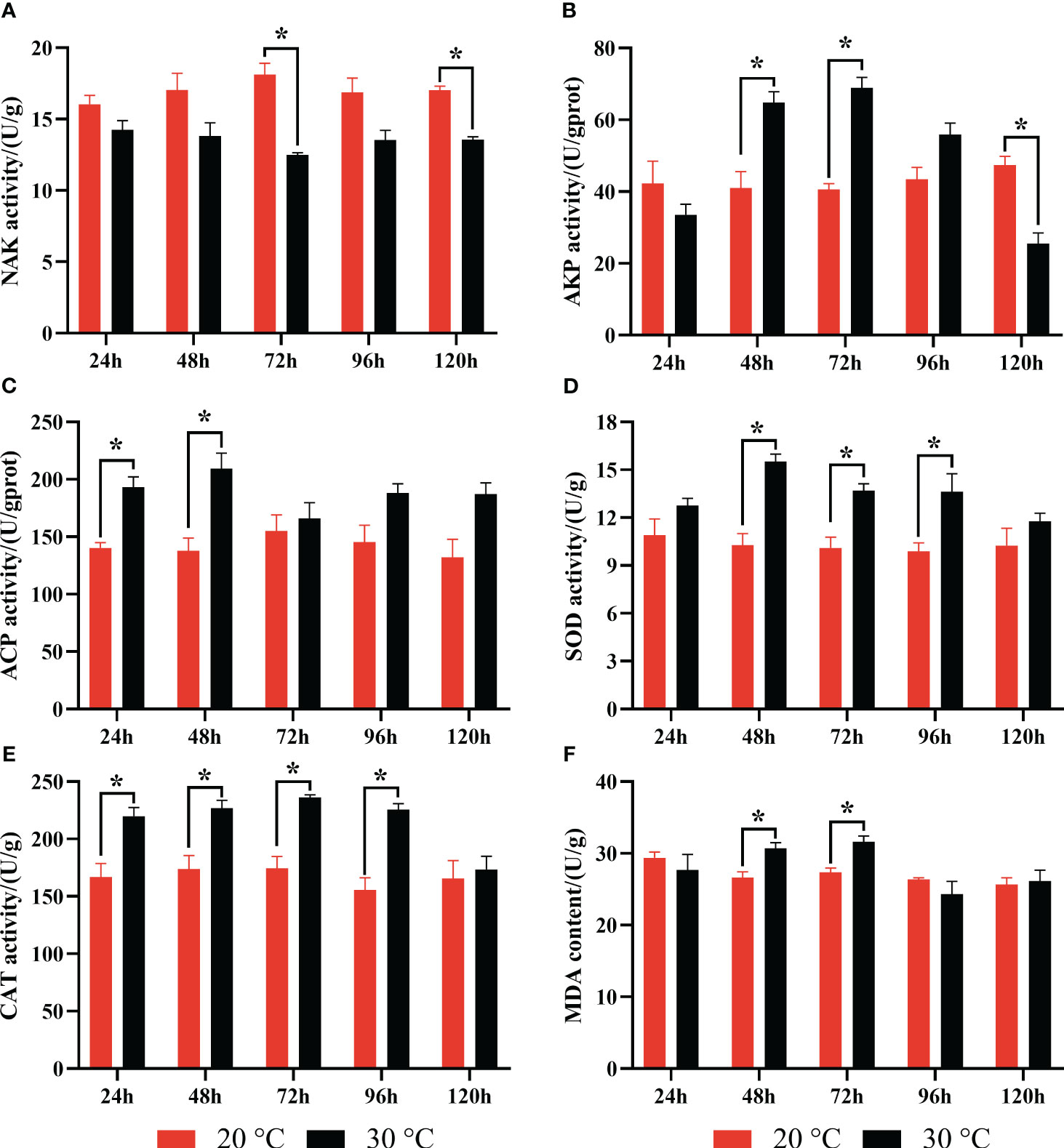
Figure 8 Effect of heat stress on enzyme activities in the gill of clam R. philippinarum. Values are mean ± standard error of the mean. Significance was considered at *p < 0.05. (A) Na+K+-ATPase (NAK); (B) Alkaline phosphatase (AKP); (C) Acid phosphatase (ACP); (D) Superoxide dismutase (SOD); (E) Catalase (CAT); (F) Malondialdehyde (MDA).
4 Discussion
4.1 Energy metabolism
Energy absorption and energy distribution of aquatic invertebrates will be affected by heat stress (Sokolova et al., 2012; Han et al., 2013). The energetic costs of mollusks under heat stress may be substantial and vary sharply with intensity of heat stress. For mollusks, these energy costs are likely to be instrumental in governing the suitability of different habitats and may play key roles in defining the distribution pattern over latitudinal and vertical temperature gradients (Dong et al., 2022a). Under heat and hypoxia stress, mollusks maintain the energy balance in a specific way, promoting carbohydrate metabolism by stimulating glycolysis (Sun et al., 2021; Hu et al., 2022a). As well-known, glycolysis is the first stage of carbohydrate metabolism, an anaerobic process producing two molecules of ATP from one molecule of glucose through enzymatic reactions (Lauer et al., 2012). For instance, air exposure could induced the significantly higher expression of glycolysis-related genes in oyster C. gigas, including hexokinase (HK), phosphofructokinase (PFK), glucose transporter 1 (GLUT1), phosphoenolpyruvate carboxykinase (PEPCK) and glyceraldehyde-3-phosphate dehydrogenase (GAPDH; Meng et al., 2018). After the heat and hypoxia stress of clam Mercenaria mercenaria, the glycolysis process was remarkably increased, showing the content of fructose and mannose significantly higher than that in the control group (Hu et al., 2022a). Consistently, our present study also revealed the significant enrichment of the glycolysis pathway in the heat-stressed clams. The metabolite of Beta-D-Glucose in the heat-stressed clams showed the significantly decreased pattern under heat stress, while Acetyl-CoA displayed increased (Table 1). Similarly, gene expression levels showed the significant increasing for the glycolysis-related genes, including GLUT1, PEPCK, ADP-dependent glucokinase (ADP-GLK), aldose 1-epimerase (AEP1) and alcohol dehydrogenase class-3 (ADH3). Moreover, we also detected the significantly increased expression levels of SDH, PEPCK and ADPGLK in the heat-stressed clams. In contrast, gene expression of hexokinase-1 (HK1), transposable element tcb1 transposase (TCB1) and glucose-6-phosphate isomerase (GPI6) was significantly decreased in the heat-stressed clams (Table 1). Therefore, the changes in glycolysis-related metabolites (Beta-D-Glucose and Acetyl-CoA) and gene expression suggests that glycolysis may serve as the main energy source in the heat-stressed clams. This may represent the survival strategy of the hard-stressed clams, which may help them to maintain the energy balance by promoting glycolysis during heat stress adaptation.
TCA cycle is the ultimate metabolic pathway, serving as the metabolic hub of carbohydrates, lipids and amino acids in aerobic organisms (Lu et al., 2020). In TCA cycle, pyruvic acid, a glycolysis product, will be gradually oxidized and decomposed into CO2 and H2O, generating NADH, FADH2 and ATP. In this study, the metabolites of Acetyl-CoA and S-Acetyldihydrolipoamide-E involved in TCA cycle were found to be significantly increased after heat stress, while succinic semialdehyde was significantly decreased in the heat-stress clams. Acetyl-CoA is served as the key metabolic node in TCA, allowing energy supply to match energy requirement in animal tissues (Shi and Tu, 2015). S-Acetyldihydrolipoamide-E is a key metabolite linking the TCA cycle, pyruvate metabolism, and gluconeogenesis (Ma et al., 2021). Succinic semialdehyde can be converted to succinic acid, which is a key intermediate product of the TCA cycle. In this study, our finding confirmed that genes involved in TCA cycle (succinate dehydrogenase cytochrome b, SDHB; isocitrate dehydrogenase 2, IDH2) were significantly up-regulated after heat stress (Table 1). SDH is a key enzyme linking oxidative phosphorylation and electron transfer, and its activity can generally be used as an indicator to evaluate the degree of TCA operation (Rutter et al., 2010). Furthermore, IDH is a key metabolic enzyme in TCA cycle, which catalyzes the conversion of isocitrate acid into α -ketoglutarate and plays an important role in cell energy metabolism and biosynthesis (Zheng et al., 2013). The present findings suggest that heat-stressed clams might strengthen their TCA flux to produce more energy to maintain energy balance for heat adaptation. This phenomenon is also supported by a previous study, which revealed the enhanced TCA cycle in scallop Chlamys farreri under heat stress (Dong et al., 2022b). Arginine and histidine are anaplerotic substrates, which are converted to α-ketoglutarate acid and then enter TCA cycle (Hara et al., 2021). In contrast, isoleucine enters TCA cycle by converting into succinyl-CoA, and then decomposes to produce ATP (Hara et al., 2021; Dong et al., 2022b). In this study, the increasing of arginine and histidine, accompanied by the reduction of isoleucine, suggests the important roles of free amino acids involved in TCA flux to produce more energy in the heat-stressed clams.
4.2 Protein processing
In this study, protein processing in endoplasmic reticulum (ER) pathway is the most significantly enriched pathway revealed by comparative transcriptomics analysis in clam R. philippinarum under the heat stress. In eukaryotic cells, ER plays an important role in the synthesis of a series of important substances such as protein, lipids and carbohydrate (Ellgaard and Frickel, 2003). The protein of ER is supervised by the quality control (QC) system, which ensures that the protein is correctly folded. The QC system includes several important genes, including calnexin (CNX), calreticulin (CRT) and glycoprotein specific thiol-disulfide oxidoreductase (ERP57; Ellgaard and Frickel, 2003; Gupta, 2012). In this research, comparative transcriptomics revealed the significantly up-regulated expression of CRT, protein disulfide-isomerase A3 (PDIA3) and CNX genes in the heat-stressed clams, probably involved in promoting the protein folding in ER (Table 2). Furthermore, secretory proteins of eukaryotic cells will be transported from the ER to Golgi apparatus through ER-Golgi intermediate compartment (Appenzeller-Herzog and Hauri, 2006). Proteins synthesized within the ER are transported to the Golgi via coat protein complex II (COPII)-coated vesicles, which comprise small GTPase Sar1, the inner-coat complex (SEC23/SEC24) and the outer-coat complex (SEC13/SEC31) (Saito et al., 2017). In this study, the significantly increased expression of protein transport SEC24C (SEC24C) and GTP-binding protein SAR1B (SAR1B) suggests that clams may adapt the heat stress by regulating protein transporting and processing in endoplasmic reticulum.
To cope with environmental stress, ER proteins will misfold and accumulate in the cells, affecting cell normal function, and even causing diseases (Ruggiano et al., 2014; Almanza et al., 2019). Endoplasmic reticulum-associated protein degradation (ERAD) is one of the most important pathways regulating ER pressure in eukaryotic cells. As indicated, multiprotein complexes in the ER are able to identify, remove, ubiquitinate and deliver misfolded proteins to the 26S proteasome for degradation in the cytosol (Needham et al., 2019). During the process, molecular chaperones of heat shock protein (HSP) family participate in ERAD process by combining with ubiquitin ligases (Goldberg, 2003). In this study, it is evidenced by the significantly increased expression levels of HSP70 and HSP90, probably aiming to promote the degradation of misfolded proteins and maintain cell homeostasis (Table 2). Moreover, molecular chaperones of GRP94 and HSP40 in ER are capable of recognizing misfolded protein and assisting in targeting malformed protein to ERAD (Haas, 1994; Plemper et al., 1997; Tyedmers et al., 2003; Eletto et al., 2010). In the present study, we also detect the up-regulation of molecular chaperones in clams after the heat stress, such as nucleotide exchange factor SIL1, glucose-regulated protein 94 (GRP94), dnaJ homolog subfamily C member 3 (DNAJ3), and DNAJ11 (Table 2). This suggests that molecular chaperones in the ER may participate in the degradation of misfolded proteins and maintain cell homeostasis during the heat stress in clam tissues.
Additionally, IRE1α can specifically cut the mRNA of X-box binding protein (XBP1), which regulates protein folding, transport and degradation at the transcriptional level, and thereby controls lipid synthesis and restores ER homeostasis by increasing ER volume (Tirasophon et al., 1998; Hetz et al., 2020). In this study, the up-regulated expression of inositol-requiring protein 1 (IER1) and XBP1 may be involved in maintaining the stability of ER by the degradation of misfolded proteins in the heat-stressed clams.
4.3 Substance metabolism
Glycerophospholipid (phosphatidylcholine, PC; phosphatidylethanolamine, PE), as the important component of cell membranes, is closely related to cell membrane permeability and ATPase activity (Overgaard et al., 2008; Nemova et al., 2017; Nguyen et al., 2020). Glycerophospholipid has been recently recognized as an important metabolite of cell membrane damage caused by high temperature stress (Calzada et al., 2016). For instance, the increasing of PC is an adaptive response to heat stress to maintain biofilm stability and function in animal tissues (Qu and Ajuwon, 2018). The increased PE plays an important role in the heat tolerance of heat-resistant intertidal zone snail Echinolittorina malaccana (Chen et al., 2021). Moreover, the hard clam M. mercenaria increased their glycerophospholipid content after heat stress to enhance cell membrane stability (Hu et al., 2022a). Similarly, the razor clam Sinonovacula constricta also showed the increased contents of PC and PE under heat stress (Zhang and Dong, 2021). In this study, the increasing of PC contents in the heat-stressed clams was consistently detected by comparative metabonomics (Table 3).
As recently reported, PC biosynthesis occurs mainly via the CDP-choline pathway in eukaryotes (Islam et al., 2022). Choline can be converted to phosphocholine through phosphorylation by the activity of cytosolic enzyme choline kinase (CK). Phosphocholine is catalytically converted to CDP-choline by a rate-limiting enzyme such as phosphocholine cytidylyltransferase (CT), which is then catalyzed by c choline/ethanolaminephosphotransferase (CEPT1) to produce PC (Henneberry et al., 2002). In our study, we consistently find the significantly increased metabolites of CDP-choline and phosphocholine in the heat-stressed clams, supporting the conclusion of PC biosynthesis by the CDP-choline pathway. Moreover, another possible pathway of PC synthesis occurs when PE converts to PC through three successive methylation reactions catalyzed by phosphatidylethanolamine N-methyltransferase (PEMT) (van der Veen et al., 2017). In this study, PC synthesis derived from PE is also evidenced by the increased gene expression of CEPT1 and PEMT in the heat-stress clams (Table 3). Taken together, the increasing of PC synthesis is essential to maintain the stability and permeability of cell membrane during the heat stress in clams. These findings provide the molecular and metabolite evidences for heat stress adaptation in clams, as well as other bivalves.
Stearoyl-CoA Desaturase (SCD) is an important enzyme in fat metabolism and plays a central regulatory role in fatty acid biosynthesis (Ntambi, 1995; Kim and Ntambi, 1999; Ntambi et al., 2004). As reported, SCD plays an important role in membrane composition and fluidity regulation (Castro et al., 2011). The changes in SCD gene transcripts has been considered an important adaptative mechanism of aquatic organisms in response to temperature fluctuations (Hsieh and Kuo, 2005; Hsieh et al., 2007). In addition, fatty acid synthase (FAS) can catalyze the synthesis of long-chain fatty acids, which form as the important membrane substrates for cellular energy metabolism (Wakil, 1989; Smith et al., 2003). In the present study, the decreased gene expression of SCD and FAS suggests that clams may adapt to the heat stress by the regulation of membrane composition and fluidity.
In this study, the activities of several enzymes (e.g. NAK, AKP and ACP) were significantly affected by the heat stress in clams (Figure 8). As reported, NAK is an integral membrane protein that couples the hydrolysis of ATP to ion transport across the membranes in animal cells (Jorgensen and Pedersen, 2001). In this study, NAK activity in clams was significantly decreased at 72 h and 120 h after the heat stress, suggesting the reduction of ion transport across cell membranes during the heat stress. In addition to NAK, two important phosphatases (AKP and ACP) play an important role in stress signal transduction and physiological adaptation to the environment stress, probably by the degradation of exogenous protein, carbohydrates and lipids (Liu et al., 2004). More recently, ACP and AKP have also been reported to be involved in heat stress responses in scallop P. yessoensis and mussel Mytilus coruscus (Hu et al., 2015; Jiang et al., 2019). In this study, we consistently reveal that the activities of ACP and AKP were significantly increased in the heat-stressed clams. These findings suggests that the increased activities of ACP and AKP may be essential for stress signal transduction, and thereby play an important role in physiological adaptation to the heat stress in bivalves.
4.4 Inoxidizability
For mollusks, the increasing of seawater temperature can accelerate metabolic rates, resulting in the accumulation of reactive oxygen species (ROS) and activation of anti-oxidative enzymes (De Zoysa et al., 2009). However, excess ROS (including superoxide anion, peroxide and hydroxyl radical) may affect the normal cellular function and even cause the damages of DNA and protein function (Song et al., 2006; Kong et al., 2022). To control ROS production and accumulation, aquatic organisms usually activate antioxidant enzymes, such as superoxide dismutase (SOD), catalase (CAT), glutathione-S-transferases (GST) and cytochrome P450 (CYP) (Rosic et al., 2010; Zhang et al., 2021). Different from those antioxidant enzymes, malonyldialdehyde (MDA) is usually served as a signature product of antioxidant damage (Wang et al., 2011). In mollusks, these antioxidant enzymes are usually activated by environmental stress, especially for the heat stress. For instance, gene expression of GST and CYP in clam Sinonovacula constricta was activated after the heat shock (Kong et al., 2022). Similarly, the increased GST expression was also observed in bivalve Laternula elliptica and scallop P. yessoensis under heat stress (Truebano et al., 2010; Jiang et al., 2018). Additionally, the mRNA levels of SOD and GST were found to be substantially increased in the mussels Mytilus galloprovincialis after the exposure of acute heat stress (Banni et al., 2014). In this study, we also confirm the activation of antioxidant system in the heat-stressed clams by the elevated gene expression of SOD, GST and CYP, and the increased enzyme activities of SOD, CAT and MDA (Table 4). These findings reflect the important roles of antioxidant enzymes in physiological adaptation to the heat stress in bivalves.
4.5 Immunology/anti-apoptotic
Under the infection of external pathogens, bivalve mollusks can improve the expression of immune-related genes, relying on innate immune system to defense against invaders (Manfredi et al., 2008; Huang et al., 2013; Thornton et al., 2017; Shen et al., 2018). As reported in the clam S. constricta, C-type lectins (CLEC) are served as pattern recognition receptors, playing the crucial roles in innate immune response to thermal stress (Kong et al., 2022). Similarly, CLEC expression was found to be significantly up-regulated after the bacteria infection in bay scallop Argopecten irradians and clam Sinonovacula constricta (Huang et al., 2013; Shen et al., 2018). In addition to CLEC, NF-κB is an important transcription factor for regulating inflammation and innate immune function (Ghosh et al., 1998; Dixit and Mak, 2002; Wang et al., 2006). For instance, the increased expression level of NF-κB reflects the immune responses in the clam R. philippinarum under hypoxia stress (Nie et al., 2020). In this study, we consistently detect the significantly increased expression of CLEC and NF-κB in the heat-stressed clams, indicating that the clams might activate the innate immune system in response to the heat stress.
Besides the innate immune system, bivalves may also respond to the external environmental pressure through an effective anti-apoptotic system (Li et al., 2017; Song et al., 2021; Hu et al., 2022b). Inhibitors of apoptosis (IAP) and E3 ubiquitin-protein ligases (E3) are critical regulators of programmed cell death and apoptosis (Broemer and Meier, 2009; Song et al., 2021). For instance, IAPs and E3 played the important roles in survival of hard clam M. mercenaria and oyster C. gigas under air exposure and heat stress (Zhang et al., 2012a; Zhou et al., 2021; Hu et al., 2022b). In this study, the changes in expression levels of IAP and E3 genes suggest that clams may adapt to heat stress by regulating its anti-apoptotic system.
4.6 The predicted model for clams in response to heat stress
Based on the obtained data, we proposed a hypothetical pathway for clam adaptation to heat stress, as shown in Figure 9. During the heat stress, glycolysis may serve as the main energy source to maintain energy balance in the heat-stressed clams. Another important energy source may be provided by the TCA cycle linking with amino acid metabolism, which can facilitate energy production via the TCA cycle for heat adaptation. As illustrated, the changes in gene expression of glycolysis-related genes (GLUT1, GLK, HK1, GPI6) and metabolites (Beta-D-Glucose and Acetyl-CoA) reflect the adjusted energy metabolism in the heat-stressed clams. The protein processing in endoplasmic reticulum plays the important roles in the stability of cell membrane structure during heat adaptation. Molecular chaperones (HSP70 and HSP90) participate in the degradation of misfolded proteins to maintain cell homeostasis during the heat stress. The increased contents of glycerophospholipid are essential to maintain the stability and permeability of cell membrane in the heat-stressed clams. To defense the heat stress, clams also initiate the protection mechanisms by activating antioxidant system, innate immune system and anti-apoptotic system.
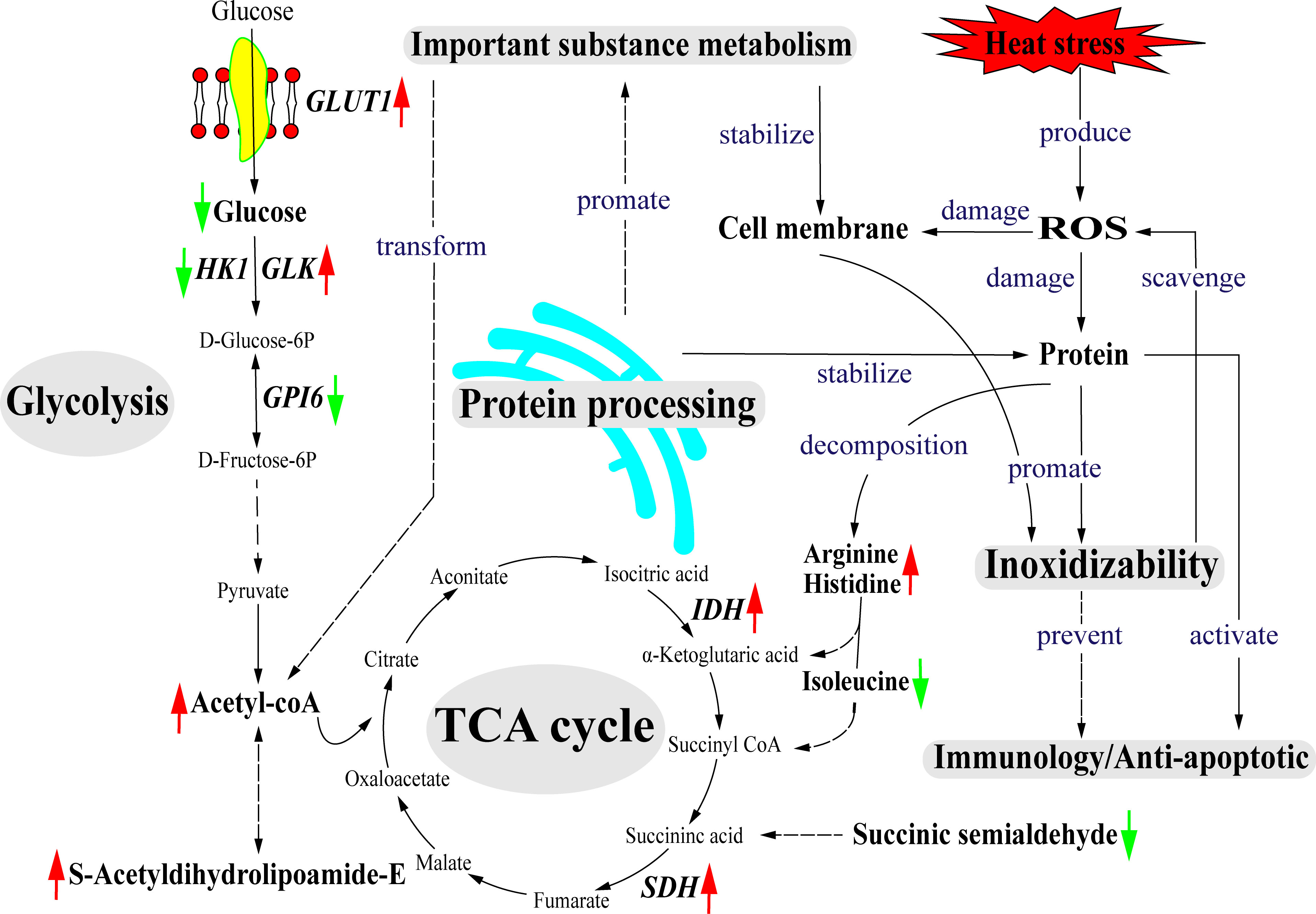
Figure 9 The predicted model for clams in response to heat stress. Solid line represents having a direct association and das lines represent having a potential association. The red arrows represent up-regulated and the green arrows represent down-regulated. GLUT1, glucose transporter type 1; HK1, hexokinase-1; ADP-GLK, ADP-dependent glucokinase; GPI6, glucose-6-phosphate isomerase; IDH, isocitrate dehydrogenase; SDH, succinate dehydrogenase cytochrome.
For aquatic invertebrates, moderate stress may cause metabolic and ATP turnover acceleration, allowing to compensate for additional energy cost for increased activity and cellular maintenance (Sokolova et al., 2012). However, extreme stress may give rise to the slow-down of activity and suppression of metabolic rate for time-limited survival. In this study, a large number of genes and metabolites, such as GLUT1, HSP, Beta-D-Glucose and Acetyl-CoA, were enriched in the energy metabolism pathway of clam under heat stress, indicating that clam may produce more energy to maintain the survival under heat stress.
To sum up, glycolysis, TCA cycle and amino acid metabolism may serve as the main energy sources to maintain energy balance in clams during heat stress. Protein processing in endoplasmic reticulum and glycerophospholipid may play an important role in heat stress adaptation, especially for maintaining the stability of cell membrane. Furthermore, the protection mechanism during heat stress will be maintained by antioxidant system, accompanied by innate immune system and anti-apoptotic system. Therefore, the time-limited survival of clams during heat stress may result from the adjusted energy metabolism, cell membrane stability and activated protection mechanisms. These novel findings may also contribute to global distribution pattern of clams over temperature gradients from an ecological perspective.
5 Conclusion
The transcriptome results indicated that DEGs were mainly involved energy metabolism, protein processing, glycerolphospholipid metabolism, inoxidizability and anti-apoptosis. Metabolome data revealed that DEMs are mainly involved in energy supply and glycerophospholipid metabolism. According the present findings, we proposed the graphical model of molecular and metabolic mechanism for heat stress adaptation in clams. As illustrated, the protein processing, substance metabolism, inoxidizability and immunology/anti-apoptosis play the important roles in the physiological adaptation of clams in response to heat stress. The present finding will shed lights on the adaptive strategies to heat stress in clams, as well as other mollusks, in a warming climate at the global scale.
Data availability statement
The datasets presented in this study can be found in online repositories. The BioProject for the clam transcriptomes has been registered as PRJNA933718, and the transcriptome sequencing data has been deposited under the accession numbers from SRR23410264 to SRR23410269. This information has been provided in the Supplementary Material.
Ethics statement
Ethical review and approval was not required for the study on clams in accordance with the local legislation and institutional requirements.
Author contributions
XS, HJ, and LZ contributed to conception and design of the study. HJ and YG wrote the first draft of the manuscript. LZ, KT, ZL and BW wrote sections of the manuscript. All authors contributed to the article and approved the submitted version.
Funding
This study was supported by research grants from the Science and Technology Innovation Program of the Laoshan Laboratory (LSKJ202203803), National Key R&D Program of China (2018YFD0900702) and National Natural Science Foundation of China (32273107). This work was also supported by Central Public-interest Scientific Institution Basal Research Fund, YSFRI, CAFS (20603022022001), Shinan District Science and Technology Plan Project (2022-2-026-ZH) and the project of Putian Science and Technology Department (2021NJJ002) and the Guiding Funds of Central Government for Supporting the Development of the Local Science and Technology (2022L3043).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmars.2023.1204598/full#supplementary-material
References
Aldridge D. W., Payne B. S., Miller A. C. (1995). Oxygen consumption, nitrogenous excretion, and filtration rates of Dreissena polymorpha at acclimation temperatures between 20 and 32°C. Can. J. Fish. Aquat. Sci. 52, 1761–1767. doi: 10.1139/f95-768
Almanza A., Carlesso A., Chintha C., Creedican S., Doultsinos D., Leuzzi B., et al. (2019). Endoplasmic reticulum stress signalling - from basic mechanisms to clinical applications. FEBS J. 286, 241–278. doi: 10.1111/febs.14608
Andersen C. L., Jensen J. L., Orntoft T. F. (2004). Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 64, 5245–5250. doi: 10.1158/0008-5472.CAN-04-0496
Ansell A. D., Sivadas P. (1973). Some effects of temperature and starvation on the bivalve Donax vittatus (da Costa) in experimental laboratory populations. J. Exp. Mar. Biol. Ecol. 13, 229–262. doi: 10.1016/0022-0981(73)90069-5
Appenzeller-Herzog C., Hauri H. P. (2006). The ER-golgi intermediate compartment (ERGIC): in search of its identity and function. J. Cell. Sci. 119, 2173–2183. doi: 10.1242/jcs.03019
Banni M., Hajer A., Sforzini S., Oliveri C., Boussetta H., Viarengo A. (2014). Transcriptional expression levels and biochemical markers of oxidative stress in Mytilus galloprovincialis exposed to nickel and heat stress. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 160, 23–29. doi: 10.1016/j.cbpc.2013.11.005
Bindoff N. L., Cheung W. W., Kairo J. G., Arístegui J., Guinder V. A., Hallberg R., et al. (2019). Changing ocean, marine ecosystems, and dependent communities,” in IPCC special report on the ocean and cryosphere in a changing climate. Eds. Pörtner H.-O., Roberts D. C., Masson-Delmotte V., Zhai P., Tignor M., Poloczanska E., et al (Cambridge, UK and New York, NY, USA: Cambridge University Press), 447–587.
Bougrier S., Geairon P., Deslous-Paoli J. M., Bacher C., Jonquières G. (1995). Allometric relationships and effects of temperature on clearance and oxygen consumption rates of Crassostrea gigas (Thunberg). Aquaculture 134, 143–154. doi: 10.1016/0044-8486(95)00036-2
Bouhaddani S. E., Houwing-Duistermaat J., Salo P., Perola M., Jongbloed G., Uh H. W. (2016). Evaluation of O2PLS in omics data integration. BMC Bioinform. 17, 117–132. doi: 10.1186/s12859-015-0854-z
Broemer M., Meier P. (2009). Ubiquitin-mediated regulation of apoptosis. Trends Cell Biol. 19, 130–140. doi: 10.1016/j.tcb.2009.01.004
Bylesjö M., Eriksson D., Kusano M., Moritz T., Trygg J. (2007). Data integration in plant biology: the O2PLS method for combined modeling of transcript and metabolite data. Plant J. 52, 1181–1191. doi: 10.1111/j.1365-313X.2007.03293.x
Calzada E., Onguka O., Claypool S. M. (2016). Phosphatidylethanolamine metabolism in health and disease. Int. Rev. Cell Mol. Biol. 321, 29–88. doi: 10.1016/bs.ircmb.2015.10.001
Castro L. F. C., Wilson J. M., Gonçalves O., Galante-Oliveira S., Rocha E., Cunha I. (2011). The evolutionary history of the stearoyl-CoA desaturase gene family in vertebrates. BMC Evol. Biol. 11, 1–14. doi: 10.1186/1471-2148-11-132
Chen Y., Wang J., Liao M., Li X., Dong Y. (2021). Temperature adaptations of the thermophilic snail Echinolittorina malaccana: insights from metabolomic analysis. J. Exp. Biol. 224, jeb238659. doi: 10.1242/jeb.238659
Copley T. R., Aliferis K. A., Kliebenstein D. J., Jabaji S. H. (2017). An integrated RNAseq-1H NMR metabolomics approach to understand soybean primary metabolism regulation in response to rhizoctonia foliar blight disease. BMC Plant Biol. 17, 1–18. doi: 10.1186/s12870-017-1020-8
De Zoysa M., Whang I., Lee Y., Lee S., Lee J. S., Lee J. (2009). Transcriptional analysis of antioxidant and immune defense genes in disk abalone (Haliotis discus discus) during thermal, low-salinity and hypoxic stress. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 154, 387–395. doi: 10.1016/j.cbpb.2009.08.002
Ding J., Li J., Yang D., Yang F., Nie H., Huo Z., et al. (2018). Molecular characteristics of a novel HSP60 gene and its differential expression in Manila clams (Ruditapes philippinarum) under thermal and hypotonic stress. Cell Stress Chaperones 23, 179–187. doi: 10.1007/s12192-017-0796-7
Dixit V., Mak T. W. (2002). NF-κB signaling: many roads lead to Madrid. Cell 111, 615–619. doi: 10.1016/s0092-8674(02)01166-2
Dong Y., Liao M., Han G., Somero G. (2022a). An integrated, multi-level analysis of thermal effects on intertidal molluscs for understanding species distribution patterns. Biol. Rev. 97, 554–581. doi: 10.1111/brv.12811
Dong X., Yang Z., Liu Z., Wang X., Yu H., Peng C., et al. (2022b). Metabonomic analysis provides new insights into the response of zhikong scallop (Chlamys farreri) to heat stress by improving energy metabolism and antioxidant capacity. Antioxidants 11, 1084. doi: 10.3390/antiox11061084
Eletto D., Dersh D., Argon Y. (2010). GRP94 in ER quality control and stress responses. Semin. Cell Dev. Biol. 21, 479–485. doi: 10.1016/j.semcdb.2010.03.004
Ellgaard L., Frickel E. M. (2003). Calnexin, calreticulin, and ERp57. Cell Biochem. Biophys. 39, 223–247. doi: 10.1385/CBB:39:3:223
Freytes-Ortiz I. M., Stallings C. D. (2018). Elevated temperatures suppress inducible defenses and alter shell shape of intertidal mussel. Mar. Biol. 165, 1–9. doi: 10.1007/s00227-018-3371-6
Ghosh S., May M. J., Kopp E. B. (1998). NF-kappa b and REL proteins: evolutionarily conserved mediators of immune responses. Annu. Rev. Immunol. 16, 225. doi: 10.1146/annurev.immunol.16.1.225
Goldberg A. L. (2003). Protein degradation and protection against misfolded or damaged proteins. Nature 426, 895–899. doi: 10.1038/nature02263
Gracey A. Y., Connor K. (2016). Transcriptional and metabolomic characterization of spontaneous metabolic cycles in Mytilus californianus under subtidal conditions. Mar. Genomics 30, 35–41. doi: 10.1016/j.margen.2016.07.004
Gupta G. S. (2012). “Lectins in quality control: calnexin and calreticulin,” in Animal lectins: form, function and clinical applications (Vienna: Springer), 29–56. doi: 10.1007/978-3-7091-1065-2
Haas I. G. (1994). BiP (GRP78), an essential hsp70 resident protein in the endoplasmic reticulum. Experientia 50, 1012–1020. doi: 10.1007/BF01923455
Han K. N., Lee S. W., Wang S. Y. (2008). The effect of temperature on the energy budget of the Manila clam, Ruditapes philippinarum. Aquac. Int. 16, 143–152. doi: 10.1007/s10499-007-9133-y
Han G., Zhang S., Marshall D. J., Ke C., Dong Y. (2013). Metabolic energy sensors (AMPK and SIRT1), protein carbonylation, and cardiac failure as biomarkers of thermal stress in an intertidal limpet: linking energetic allocation with environmental temperature during aerial emersion. J. Exp. Biol. 216, 3273–3282. doi: 10.1242/jeb.084269
Hao Z. L., Tang X. J., Ding J., Ben Y., Chang Y. Q. (2014). Effect of high temperature on survival, oxygen consumption, behavior, ammonia-n excretion, and related immune indicators of the Japanese scallop Mizuhopecten yessoensis. Aquac. Int. 22, 1863–1876. doi: 10.1007/s10499-014-9788-0
Hara Y., Kume S., Kataoka Y., Watanabe N. (2021). Changes in TCA cycle and TCA cycle-related metabolites in plasma upon citric acid administration in rats. Heliyon 7, e08501. doi: 10.1016/j.heliyon.2021.e08501
Henneberry A. L., Wright M. M., McMaster C. R. (2002). The major sites of cellular phospholipid synthesis and molecular determinants of fatty acid and lipid head group specificity. Mol. Biol. Cell 13, 3148–3161. doi: 10.1091/mbc.01-11-0540
Hetz C., Zhang K., Kaufman R. J. (2020). Mechanisms, regulation and functions of the unfolded protein response. Nat. Rev. Mol. Cell Biol. 21, 421–438. doi: 10.1038/s41580-020-0250-z
Hsieh S. L., Hu C. Y., Hsu Y. T., Hsieh T. J. (2007). Influence of dietary lipids on the fatty acid composition and stearoyl-CoA desaturase expression in hybrid tilapia (Oreochromis niloticus× o. aureus) under cold shock. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 147, 438–444. doi: 10.1016/j.cbpb.2007.02.010
Hsieh S. L., Kuo C. M. (2005). Stearoyl–CoA desaturase expression and fatty acid composition in milkfish (Chanos chanos) and grass carp (Ctenopharyngodon idella) during cold acclimation. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 141, 95–101. doi: 10.1016/j.cbpc.2005.02.001
Hu Z., Feng J., Song H., Zhou C., Yang M. J., Shi P., et al. (2022a). Metabolic response of Mercenaria mercenaria under heat and hypoxia stress by widely targeted metabolomic approach. Sci. Total Environ. 809, 151172. doi: 10.1016/j.scitotenv.2021.151172
Hu Z., Feng J., Song H., Zhou C., Yu Z. L., Yang M. J., et al. (2022b). Mechanisms of heat and hypoxia defense in hard clam: insights from transcriptome analysis. Aquaculture 549, 737792. doi: 10.1016/j.aquaculture.2021.737792
Hu M., Li L., Sui Y., Li J., Wang Y., Lu W., et al. (2015). Effect of pH and temperature on antioxidant responses of the thick shell mussel Mytilus coruscus. Fish Shellfish Immunol. 46, 573–583. doi: 10.1016/j.fsi.2015.07.025
Huang M., Song X., Zhao J., Mu C., Wang L., Zhang H., et al. (2013). A c-type lectin (AiCTL-3) from bay scallop Argopecten irradians with mannose/galactose binding ability to bind various bacteria. Gene 531, 31–38. doi: 10.1016/j.gene.2013.08.042
Islam R., Melvin S. D., Kit Yu R. M., O'Connor W. A., Anh Tran T. K., Andrew-Priestley M., et al. (2022). Estrogenic mixtures induce alterations in lipidomic profiles in the gonads of female oysters. Chemosphere 291, 132997. doi: 10.1016/j.chemosphere.2021.132997
Jiang W., Du M., Fang J., Gao Y., Mao Y., Chen Q., et al. (2019). Response of yesso scallop Patinopecten yessoensis to acute temperature challenge: physiological and biochemical parameters. J. Oceanol. Limnol. 37, 321–329. doi: 10.1007/s00343-019-7245-7
Jiang W., Lin F., Fang J., Gao Y., Du M., Fang J., et al. (2018). Transcriptome analysis of the yesso scallop, Patinopecten yessoensis gills in response to water temperature fluctuations. Fish Shellfish Immunol. 80, 133–140. doi: 10.1016/j.fsi.2018.05.038
Jorgensen P. L., Pedersen P. A. (2001). Structure-function relationships of na+, k+, ATP, or Mg2+ binding and energy transduction in Na,K-ATPase. Biochim. Biophys. Acta Biomembr. 1505, 57–74. doi: 10.1016/s0005-2728(00)00277-2
Juárez O. E., Lafarga-De la Cruz F., Leyva-Valencia I., López-Landavery E., García-Esquivel Z., Díaz F., et al. (2018). Transcriptomic and metabolic response to chronic and acute thermal exposure of juvenile geoduck clams Panopea globosa. Mar. Genomics 42, 1–13. doi: 10.1016/j.margen.2018.09.003
Kim Y. C., Ntambi J. M. (1999). Regulation of stearoyl-CoA desaturase genes: role in cellular metabolism and preadipocyte differentiation. Biochem. Biophys. Res. Commun. 266, 1–4. doi: 10.1006/bbrc.1999.1704
Kong X., Lv L., Ren J., Liu Y., Lin Z., Dong Y. (2022). Comparative transcriptome analyses unravel the response to acute thermal stress in the razor clam, Sinonovacula constricta. Aquac. Rep. 23, 101079. doi: 10.1016/j.aqrep.2022.101079
Lauer M. M., de Oliveira C. B., Yano N. L. I., Bianchini A. (2012). Copper effects on key metabolic enzymes and mitochondrial membrane potential in gills of the estuarine crab Neohelice granulata at different salinities. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 156, 140–147. doi: 10.1016/j.cbpc.2012.08.001
Li Y., Huang J., Liu Z., Zhou Y., Xia B., Wang Y., et al. (2017). Transcriptome analysis provides insights into hepatic responses to moderate heat stress in the rainbow trout (Oncorhynchus mykiss). Gene 619, 1–9. doi: 10.1016/j.gene.2017.03.041
Liu S., Jiang X., Hu X., Gong J., Hwang H., Mai K. (2004). Effects of temperature on non-specific immune parameters in two scallop species: Argopecten irradians (Lamarck 1819) and Chlamys farreri (Jones & Preston 1904). Aquac. Res. 35, 678–682. doi: 10.1111/j.1365-2109.2004.01065.x
Lu Z., Wang S., Ji C., Li F., Cong M., Shan X., et al. (2020). iTRAQ-based proteomic analysis on the mitochondrial responses in gill tissues of juvenile olive flounder Paralichthys olivaceus exposed to cadmium. Environ. pollut. 257, 113591. doi: 10.1016/j.envpol.2019.113591
Ma R., Xie Q., Wang J., Huang L., Guo X., Fan Y. (2021). Combination of urine and faeces metabolomics to reveal the intervention mechanism of Polygala tenuifolia compatibility with Magnolia officinalis on gastrointestinal motility disorders. J. Pharm. Pharmacol. 73, 247–262. doi: 10.1093/jpp/rgaa022
Manalo D. J., Rowan A., Lavoie T., Natarajan L., Kelly B. D., Ye S. Q., et al. (2005). Transcriptional regulation of vascular endothelial cell responses to hypoxia by HIF-1. Blood 105, 659–669. doi: 10.1182/blood-2004-07-2958
Manfredi A. A., Rovere-Querini P., Brunelli S., Clementi E. (2008). Cell death: tipping the balance of autoimmunity and tissue repair. Curr. Pharm. Des. 14, 269–277. doi: 10.2174/138161208783413275
Meng J., Wang T., Li L., Zhang G. (2018). Inducible variation in anaerobic energy metabolism reflects hypoxia tolerance across the intertidal and subtidal distribution of the pacific oyster (Crassostrea gigas). Mar. Environ. Res. 138, 135–143. doi: 10.1016/j.marenvres.2018.04.002
Meng J., Zhu Q., Zhang L., Li C., Li L., She Z., et al. (2013). Genome and transcriptome analyses provide insight into the euryhaline adaptation mechanism of Crassostrea gigas. PloS One 8, e58563. doi: 10.1371/journal.pone.0058563
Monari M., Matozzo V., Foschi J., Cattani O., Serrazanetti G. P., Marin M. G. (2007). Effects of high temperatures on functional responses of haemocytes in the clam Chamelea gallina. Fish Shellfish Immunol. 22, 98–114. doi: 10.1016/j.fsi.2006.03.016
Munari M., Matozzo V., Marin M. (2011). Combined effects of temperature and salinity on functional responses of haemocytes and survival in air of the clam Ruditapes philippinarum. Fish Shellfish Immunol. 30, 1024–1030. doi: 10.1016/j.fsi.2011.01.025
Needham P. G., Guerriero C. J., Brodsky J. L. (2019). Chaperoning endoplasmic reticulum–associated degradation (ERAD) and protein conformational diseases. Cold Spring Harb. Perspect. Biol. 11, a033928. doi: 10.1101/cshperspect.a033928
Nemova N. N., Kaivarainen E. I., Fokina N. N. (2017). Activity of Na+/K+-ATPase and the content of phospholipids in the blue mussel Mytilus edulis l. during environmental temperature changes. Appl. Biochem. Micro. 53, 699–702. doi: 10.1134/S0003683817060114
Nguyen T. P. L., Nguyen V. T. A., Do T. T. T., Nguyen Quang T., Pham Q. L., Le T. T. (2020). Fatty acid composition, phospholipid molecules, and bioactivities of lipids of the mud crab Scylla paramamosain. J. Chem-Ny. 2020, 1-9. doi: 10.1155/2020/8651453
Nie H., Liu L., Huo Z., Chen P., Ding J., Yang F., et al. (2017). The HSP70 gene expression responses to thermal and salinity stress in wild and cultivated Manila clam Ruditapes philippinarum. Aquaculture 470, 149–156. doi: 10.1016/j.aquaculture.2016.12.016
Nie H., Wang H., Jiang K., Yan X. (2020). Transcriptome analysis reveals differential immune related genes expression in Ruditapes philippinarum under hypoxia stress: potential HIF and NF-κB crosstalk in immune responses in clam. BMC Genom. 21, 1–16. doi: 10.1186/s12864-020-6734-6
Ntambi J. M. (1995). The regulation of stearoyl-CoA desaturase (SCD). Prog. Lipid Res. 34, 139–150. doi: 10.1016/0163-7827(94)00010-j
Ntambi J. M., Miyazaki M., Dobrzyn A. (2004). Regulation of stearoyl-CoA desaturase expression. Lipids 39, 1061–1065. doi: 10.1007/s11745-004-1331-2
Overgaard J., Tomcala A., Sørensen J. G., Holmstrup M., Krogh P. H., Simek P., et al. (2008). Effects of acclimation temperature on thermal tolerance and membrane phospholipid composition in the fruit fly Drosophila melanogaster. J. Insect Physiol. 54, 619–629. doi: 10.1016/j.jinsphys.2007.12.011
Patti G., Yanes O., Siuzdak G. (2012). Metabolomics: the apogee of the omics trilogy. Nat. Rev. Mol. Cell Biol. 13, 263–269. doi: 10.1038/nrm3314
Pfaffl M. W., Tichopad A., Prgomet C., Neuvians T. P. (2004). Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper–excel-based tool using pair-wise correlations. Biotechnol. Lett. 26, 509–515. doi: 10.1023/b:bile.0000019559.84305.47
Plemper R. K., Böhmler S., Bordallo J., Sommer T., Wolf D. H. (1997). Mutant analysis links the translocon and BiP to retrograde protein transport for ER degradation. Nature 388, 891–895. doi: 10.1038/42276
Pörtner H. O., Farrell A. P. (2008). Physiology and climate change. Science 322, 690–692. doi: 10.1126/science.1163156
Qu H., Ajuwon K. M. (2018). Metabolomics of heat stress response in pig adipose tissue reveals alteration of phospholipid and fatty acid composition during heat stress. J. Anim. Sci. 96, 3184–3195. doi: 10.1093/jas/sky127
Rosic N. N., Pernice M., Dunn S., Dove S., Hoegh-Guldberg O. (2010). Differential regulation by heat stress of novel cytochrome P450 genes from the dinoflagellate symbionts of reef-building corals. Appl. Environ. Microbiol. 76, 2823–2829. doi: 10.1128/AEM.02984-09
Ruggiano A., Foresti O., Carvalho P. (2014). ER-associated degradation: protein quality control and beyond. J. Cell Biol. 204, 869–879. doi: 10.1083/jcb.201312042
Rutter J., Winge D. R., Schiffman J. D. (2010). Succinate dehydrogenase–assembly, regulation and role in human disease. Mitochondrion 10, 393–401. doi: 10.1016/j.mito.2010.03.001
Saito K., Maeda M., Katada T. (2017). Regulation of the Sar1 GTPase cycle is necessary for large cargo secretion from the endoplasmic reticulum. Front. Cell Dev. Biol. 5. doi: 10.3389/fcell.2017.00075
Shen Y., Shao Y., Cui Y., Zhao X., Zhang W., Li C. (2018). Novel c-type lectin from razor clam Sinonovacula constricta agglutinates bacteria and erythrocytes in a Ca2+-dependent manner. Dev. Comp. Immunol. 86, 9–16. doi: 10.1016/j.dci.2018.04.016
Shi L., Tu B. P. (2015). Acetyl-CoA and the regulation of metabolism: mechanisms and consequences. Curr. Opin. Cell Biol. 33C, 125–131. doi: 10.1016/j.ceb.2015.02.003
Smith S., Witkowski A., Joshi A. K. (2003). Structural and functional organization of the animal fatty acid synthase. Prog. Lipid Res. 42, 289–317. doi: 10.1016/s0163-7827(02)00067-x
Sokolova I. M., Frederich M., Bagwe R., Lannig G., Sukhotin A. A. (2012). Energy homeostasis as an integrative tool for assessing limits of environmental stress tolerance in aquatic invertebrates. Mar. Environ. Res. 79, 1–15. doi: 10.1016/j.marenvres.2012.04.003
Somero G. N. (2010). The physiology of climate change: how potentials for acclimatization and genetic adaptation will determine 'winners' and 'losers'. J. Exp. Biol. 213, 912–920. doi: 10.1242/jeb.037473
Song H., Guo X., Sun L., Wang Q., Han F., Wang H., et al. (2021). The hard clam genome reveals massive expansion and diversification of inhibitors of apoptosis in bivalvia. BMC Biol. 19, 1–20. doi: 10.1186/s12915-020-00943-9
Song Y. G., Liu B., Wang L. F., Li M. H., Liu Y. (2006). Damage to the oxygen-evolving complex by superoxide anion, hydrogen peroxide, and hydroxyl radical in photoinhibition of photosystem II. Photosyn. Res. 90, 67–78. doi: 10.1007/s11120-006-9111-7
Sun X., Tu K., Li L., Wu B., Wu L., Liu Z., et al. (2021). Integrated transcriptome and metabolome analysis reveals molecular responses of the clams to acute hypoxia. Mar. Environ. Res. 168, 105317. doi: 10.1016/j.marenvres.2021.105317
Thornton C., Leaw B., Mallard C., Nair S., Jinnai M., Hagberg H. (2017). Cell death in the developing brain after hypoxia-ischemia. Front. Cell. Neurosci. 11. doi: 10.3389/fncel.2017.00248
Tirasophon W., Welihinda A. A., Kaufman R. J. (1998). A stress response pathway from the endoplasmic reticulum to the nucleus requires a novel bifunctional protein kinase/endoribonuclease (Ire1p) in mammalian cells. Genes Dev. 12, 1812–1824. doi: 10.1101/gad.12.12.1812
Truebano M., Burns G., Thorne M. A., Hillyard G., Peck L. S., Skibinski D. O., et al. (2010). Transcriptional response to heat stress in the Antarctic bivalve Laternula elliptica. J. Exp. Mar. Biol. Ecol. 391, 65–72. doi: 10.1016/j.jembe.2010.06.011
Trueman E. R. (1983). “Locomotion in molluscs,” in The Mollusca. Academic Press, 155–198. doi: 10.1016/B978-0-12-813627-0.00008-9
Tyedmers J., Lerner M., Wiedmann M., Volkmer J., Zimmermann R. (2003). Polypeptide-binding proteins mediate completion of co-translational protein translocation into the mammalian endoplasmic reticulum. EMBO Rep. 4, 505–510. doi: 10.1038/sj.embor.embor826
van der Veen J. N., Kennelly J. P., Wan S., Vance J. E., Vance D. E., Jacobs R. L., et al. (2017). The critical role of phosphatidylcholine and phosphatidylethanolamine metabolism in health and disease. Biochim. Biophys. Acta Biomembr. 1859, 1558–1572. doi: 10.1016/j.bbamem.2017.04.006
Vandesompele J., De Preter K., Pattyn F., Poppe B., Van Roy N., De Paepe A., et al. (2002). Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, RESEARCH0034. doi: 10.1186/gb-2002-3-7-research0034
Velasco L. A., Barros J. (2008). Experimental larval culture of the Caribbean scallops argopecten nucleus and Nodipecten nodosus. Aquac. Res. 39, 603–618. doi: 10.1111/j.1365-2109.2008.01917.x
Wakil S. J. (1989). Fatty acid synthase, a proficient multifunctional enzyme. Biochemistry 28, 4523–4530. doi: 10.1021/bi00437a001
Wang X. W., Tan N. S., Ho B., Ding J. L. (2006). Evidence for the ancient origin of the NF-κB/IκB cascade: its archaic role in pathogen infection and immunity. Proc. Natl. Acad. Sci. U.S.A. 103, 4204–4209. doi: 10.1073/pnas.0507044103
Wang X., Yang H., Liu G., Wang Q. (2011). Enzyme responses and lipid peroxidation in gills and hepatopancreas of clam Mactra vereformis, following cadmium exposure. Chin. J. Oceanol. Limnol. 29, 981–989. doi: 10.1007/s00343-011-0088-5
Wells R. M., Baldwin J., Speed S. R., Weber R. E. (1998). Haemocyanin function in the new Zealand abalones Haliotis iris and h. australis: relationships between oxygen-binding properties, muscle metabolism and habitat. Mar. Freshw. Res. 49, 143–149. doi: 10.1071/MF97206
Xu G., Du F., Li Y., Nie Z., Xu P. (2016). Integrated application of transcriptomics and metabolomics yields insights into population-asynchronous ovary development in Coilia nasus. Sci. Rep. 6, 31835. doi: 10.1038/srep31835
Yan X., Nie H., Huo Z., Ding J., Li Z., Yan L., et al. (2019). Clam genome sequence clarifies the molecular basis of its benthic adaptation and extraordinary shell color diversity. iScience 19, 1225–1237. doi: 10.1016/j.isci.2019.08.049
Zhang W., Dong Y. (2021). Membrane lipid metabolism, heat shock response and energy costs mediate the interaction between acclimatization and heat-hardening response in the razor clam Sinonovacula constricta. J. Exp. Biol. 224, jeb243031. doi: 10.1242/jeb.243031
Zhang G., Fang X., Guo X., Li L., Luo R., Xu F., et al. (2012a). The oyster genome reveals stress adaptation and complexity of shell formation. Nature 490, 49–54. doi: 10.1038/nature11413
Zhang L., Hou R., Su H., Hu X., Wang S., Bao Z. (2012b). Network analysis of oyster transcriptome revealed a cascade of cellular responses during recovery after heat shock. PloS One 7, e35484. doi: 10.1371/journal.pone.0035484
Zhang Y., Nie H., Yan X. (2023). Metabolomic analysis provides new insights into the heat-hardening response of Manila clam (Ruditapes philippinarum) to high temperature stress. Sci. Total Environ. 857, 159430. doi: 10.1016/j.scitotenv.2022.159430
Zhang X., Wang X., Yan B. (2021). Single and combined effects of phenanthrene and polystyrene microplastics on oxidative stress of the clam (Mactra veneriformis). Sci. Total Environ. 771, 144728. doi: 10.1016/j.scitotenv.2020.144728
Zhang G., Yan X. (2006). A new three-phase culture method for Manila clam, Ruditapes philippinarum, farming in northern China. Aquaculture 258, 452–461. doi: 10.1016/j.aquaculture.2006.04.046
Zhao H., Zhang S. (2016). Identification of jiaozhou bay clams (Ruditapes philippinarum) by multi-element fingerprinting technique. Food Anal. Method 9, 2691–2626. doi: 10.1007/s12161-016-0461-2
Zheng B., Yao Y., Liu Z., Deng L., Anglin J. L., Jiang H., et al. (2013). Crystallographic investigation and selective inhibition of mutant isocitrate dehydrogenase. ACS Med. Chem. Lett. 4, 542–546. doi: 10.1021/ml400036z
Keywords: heat stress, Ruditapes philippinarum, transcriptomic, metabolomic, enzymatic activity
Citation: Jing H, Zhou L, Gao Y, Liu Z, Wu B, Sun X and Tu K (2023) Transcriptomics and metabolomics reveal the molecular and metabolic adaptation to heat stress in Manila clam Ruditapes philippinarum. Front. Mar. Sci. 10:1204598. doi: 10.3389/fmars.2023.1204598
Received: 30 May 2023; Accepted: 03 July 2023;
Published: 21 July 2023.
Edited by:
Yingxiang Li, University of Michigan, United StatesReviewed by:
Qing Wang, Chinese Academy of Sciences (CAS), ChinaLei Wei, Ludong University, China
Xin Wang, Harvard Medical School, United States
Copyright © 2023 Jing, Zhou, Gao, Liu, Wu, Sun and Tu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xiujun Sun, eGpzdW5AeXNmcmkuYWMuY24=; Kang Tu, dHU0NXR1QG91dGxvb2suY29t
 Hao Jing1,2,3
Hao Jing1,2,3 Liqing Zhou
Liqing Zhou Biao Wu
Biao Wu Xiujun Sun
Xiujun Sun Kang Tu
Kang Tu