- Department of Biology, South Texas Center for Emerging Infectious Diseases, University of Texas San Antonio, San Antonio, TX, USA
Francisella tularensis is a facultative intracellular pathogen that causes the disease tularemia. F. tularensis subsp. tularensis causes the most severe disease in humans and has been classified as a Category A select agent and potential bioweapon. There is currently no vaccine approved for human use, making genetic manipulation of this organism critical to unraveling the genetic basis of pathogenesis and developing countermeasures against tularemia. The development of genetic techniques applicable to F. tularensis have lagged behind those routinely used for other bacteria, primarily due to lack of research and the restricted nature of the biocontainment required for studying this pathogen. However, in recent years, genetic techniques, such as transposon mutagenesis and targeted gene disruption, have been developed, that have had a dramatic impact on our understanding of the genetic basis of F. tularensis virulence. In this review, we describe some of the methods developed for genetic manipulation of F. tularensis.
Introduction
Francisella tularensis is the causative agent of the human disease tularemia. Various subspecies of this organism have been described, with F. tularensis subsp. tularensis (also referred to as “Type A”) being the most virulent for humans. F. tularensis subsp. holarctica (also referred to as “Type B”) also causes serious disease in humans, and both subsp. tularensis and subsp. holarctica have been classified as category A select agents that require biosafety level 3 (BSL-3) containment for routine laboratory culture (McLendon et al., 2006). A closely related bacterium, F. novicida (also referred to as F. tularensis subsp. novicida, although it is not officially classified as a F. tularensis subsp.), is considered avirulent for healthy humans and can be maintained under BSL-2 conditions. A live attenuated vaccine strain (LVS) derived from F. tularensis subsp. holarctica is also avirulent for humans and can also be maintained under BSL-2 conditions (Eigelsbach and Downs, 1961). F. novicida and LVS cause lethal diseases in mice, and share similar pathogenic mechanism(s) as the select agent designated forms of F. tularensis, such as intramacrophage replication (Figure 1). Because F. novicida and LVS can be grown and maintained without the requirement for restrictive BSL-3 containment, these bacteria have been invaluable for the development of genetic manipulation techniques and the dissection of virulence mechanisms of F. tularensis.
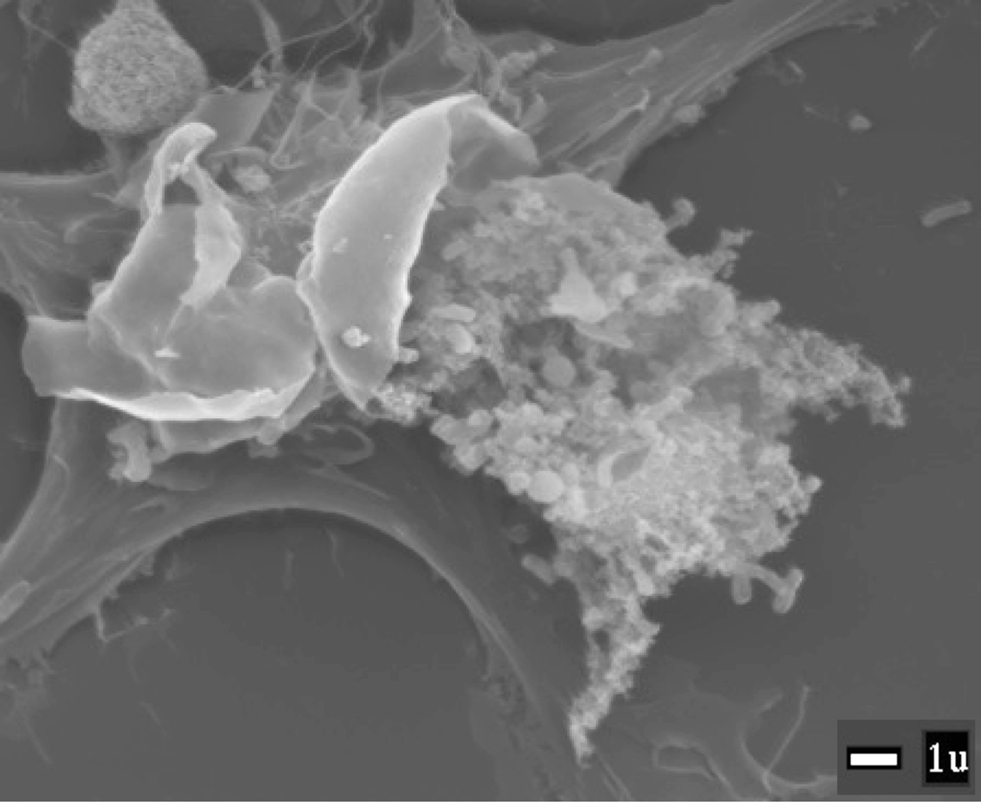
Figure 1. Francisella novicida being released from a lysed macrophage. SEM image provided by M. Neal Guntzel and Annette R. Rogriguez.
Initial techniques for genetic manipulation were developed in F. novicida, for the reasons given above, and the fact that F. novicida has proven to be the most amenable of all Francisella spp. to genetic manipulation, for reasons that are not obvious. F. novicida is closely related to F. tularensis, as has become clear through whole genome sequencing (Rohmer et al., 2007; Champion et al., 2009), yet F. novicida can be transformed with linear DNA fragments which integrate into the chromosome through homologous recombination (Lauriano et al., 2003), a property that does not seem to apply to F. tularensis subsp. Additionally, use of antibiotic resistance markers is restricted in select agent strains of F. tularensis (discussed below), but not F. novicida. Taken together, this has allowed for a wider variety of techniques to be attempted in F. novicida than in F. tularensis.
Despite these various obstacles, a number of recent advances have allowed for increasingly easier genetic manipulation in select agent forms of F. tularensis. Both random and directed mutagenesis can now be performed readily in F. tularensis subsp. tularensis, and several plasmids are available to facilitate complementation and expression studies. These tools are facilitating the illumination of the genetic basis of F. tularensis pathogenesis, and several of these will be discussed below.
Antibiotic Resistance
Antibiotic resistance markers are critical for the development of genetic techniques. Genetic techniques utilizing various antibiotic resistance genes as selectable markers have been exploited in F. novicida (Anthony et al., 1994; Gallagher et al., 2007; Liu et al., 2007; Rodriguez et al., 2008) but the use of antibiotic resistance in F. tularensis subsp. (with the exception of LVS) is restricted in the US. Resistance to any antibiotic that may be used therapeutically to treat tularemia (Urich and Petersen, 2008; Hofinger et al., 2009) may not be introduced into select agent forms of F. tularensis, which should preclude the introduction of resistance to chloramphenicol, ciprofloxacin, levofloxacin, doxycycline, gentamicin, streptomycin, and tetracycline (Enderlin et al., 1994; Maurin et al., 2000; Antibiotic Selection Guide, 2004). This has restricted the use of antibiotic resistance markers to kanamycin, erythromycin, spectinomycin, rifampin, and hygromycin resistance in these organisms (LoVullo et al., 2006; Qin and Mann, 2006; Buchan et al., 2008; Kalivoda et al., 2010; Klose, unpublished data). These restrictions do not necessarily apply to scientists outside the US, and thus examples can be found in the published literature of other antibiotic resistance markers being introduced into, e.g., F. tularensis subsp. tularensis (Norqvist et al., 1996; Pomerantsev et al., 2001; Forslund et al., 2006; Thomas et al., 2007).
Francisella tularensis/F. novicida strains are all highly resistant to ampicillin and other penicillin derivatives, and thus these antibiotics are not used therapeutically to treat tularemia. Although two β-lactamases are encoded in the genome, only one of these, BlaB, contributes in a significant manner to the ampicillin resistance of Francisella strains (Lauriano et al., 2003; Bina et al., 2006; LoVullo et al., 2006). This has allowed for the use of blaB as a selectable marker in a blaB− F. tularensis strain, thus expanding of the use of antibiotic resistance markers in select agent strains to include ampicillin (LoVullo et al., 2006).
Transformation
The first step to genetic manipulation is getting exogenous DNA into the bacterial cell. Host restriction/modification systems inhibit the acquisition of foreign DNA into the cell, and both F. novicida and F. tularensis subsp. tularensis have restriction enzymes that inhibit the acquisition of non-self DNA (Gallagher et al., 2008; Klose, unpublished data); interestingly, LVS has been reported to lack significant restriction against foreign DNA. Restriction of foreign DNA in F. novicida was eliminated by disruption of four different restriction enzymes (Gallagher et al., 2008), and this strain (MFN245) is ∼10,000-fold enhanced for transformation with foreign DNA. Because the modification systems are intact, MFN245 can be used to transform with, e.g., plasmids, which can then be easily transformed in any F. novicida strain, due to the F. novicida-specific modifications. F. tularensis subsp. tularensis restricts DNA from F. novicida, indicating it has different restriction/modification systems (Klose, unpublished data), and thus transformation of MFN245 cannot be used to enhance subsequent transformation into F. tularensis subsp. tularensis.
Francisella novicida can be transformed with chromosomal DNA and linear fragments, indicating it likely has a natural competence system. The same is not true for the F. tularensis subsp., suggesting that natural competence was lost when F. novicida and F. tularensis diverged. There are several methods to transform F. novicida/F. tularensis. One of the most unique techniques for transformation is referred to as cryotransformation (Norqvist et al., 1996; Lauriano et al., 2003). Cryotransformation is a method that is easy to perform, and it has been shown to be effective in F. novicida and all F. tularensis subsp. The cryotransformation protocol is: (a) F. tularensis/F. novicida cells are washed and resuspended into 0.2 M RbCl2; (b) DNA and cryotransformation buffer (10% Glycerol, 10 mM HEPES, pH 6.5, 0.1 M Calcium chloride, and 10 mM RbCl2) are added to the cells and left on ice for 15 min; (c) the cells are then quick frozen in dry ice/ethanol bath for 5 min; (d) the cells are then thawed until at room temperature, and culture media is added and the mix is incubated for 2–3 h; (e) finally the cells are spread onto agar media with appropriate antibiotics (Liu et al., 2007; Rodriguez et al., 2008; Zogaj et al., 2008; Barker et al., 2009).
Electroporation is also commonly used for transformation into F. novicida/F. tularensis. There are reports indicating that electroporation is even more efficient than cryotransformation (LoVullo et al., 2006; Le Pihive et al., 2009). The typical protocol for electroporation of F. novicida/F. tularensis is (a) exponentially growing Francisella cells are centrifuged to concentrate them, (b) cells are washed three times and finally resuspended in 0.5 M sucrose, (c) DNA is mixed with cells for 10 min at RT, (d) cells are pulsed in electroporator (2.5 kV, 600 Ω, 25 μF), (e) culture media is added and the mix is incubated for 2–3 h, (f) finally the cells are spread onto agar media with appropriate antibiotics.
Chemical transformation is not commonly used in Francisella species, but was one of the first methods used to introduce DNA into F. novicida (Tyeryar and Lawton, 1969, 1970; Anthony et al., 1991). On the other hand, the direct transfer of DNA from donor bacteria into F. novicida/F. tularensis via conjugation is becoming a common technique, and has been used to introduce DNA into the various Francisella spp. (Golovliov et al., 2003a, Forslund et al., 2006; Mohapatra et al., 2007; Thomas et al., 2007). Transformation frequencies of approximately 1% in F. tularensis have been observed under optimal conditions when using E. coli donors to conjugate DNA (Golovliov et al., 2003a, Mohapatra et al., 2007). The conjugation procedure is simple: (a) incubate the donor E. coli and recipient F. novicida/F. tularensis cells together at 25°C for 18 h on non-selective agar medium (by cross-streaking or place both on filter), (b) transfer the mating mixture to selective medium and grown at 37°C to isolate desired recombinants. The selective medium typically contains antibiotic to select for the conjugated plasmid, and must also provide counter selection against the donor strain, e.g., polymyxin B (50–100 μg/ml) can be used to prevent the growth of E. coli without affecting the growth of F. tularensis (Frank and Zahrt, 2007).
Plasmids
Plasmids are important for the development of various reporter constructs and in trans complementation systems. Commonly used plasmids in other bacteria, such as those with pUC, p15a, and pSC101 origins, do not replicate in F. novicida/F. tularensis; this property has been exploited in the development of F. tularensis suicide plasmids, as described below. A plasmid, pFNL10, was isolated from F. novicida strain F6186 that can also replicate in F. tularensis (but is unable to replicate in, e.g., E. coli). Norqvist et al. modified this plasmid and added a p15a origin to allow replication in E. coli, resulting in the first F. tularensis shuttle vector, pKK202 (Norqvist et al., 1996; Frank and Zahrt, 2007). Many of the pKK202 derivatives contain tetracycline and/or chloramphenicol resistance markers, which are not allowed to be introduced into select agent forms of F. tularensis in the US. In our laboratory (e.g., pKEK1140; see below) and others’, these markers have been replaced with kanamycin resistance, which allows these pKK202 derivatives to be used in F. tularensis subsp. tularensis (LoVullo et al., 2006; Buchan et al., 2008).
It is difficult to transform pKK202 derivatives into several F. tularensis subsp., leading Maier et al. to create a new shuttle vector by first selecting for pFNL10 derivatives that would replicate better in LVS, which resulted in the deletion of a portion of the pFNL10 plasmid present in pKK202 encoding an apparent toxin–antitoxin system (ORF4–ORF5). This “modified” Francisella ori is the basis for the pFNLTP series of plasmids, which also contain the pUC ori for replication in E. coli, as well as kanamycin resistance for use in select agent forms of F. tularensis. The pFNLTP shuttle vectors have high transformation efficiency in LVS and contain multiple cloning sites (Maier et al., 2004). Additionally, these researchers identified a temperature-sensitive allele in the replicon that has become an extremely useful tool for removal of pFNL10-based plasmids from Francisella strains.
LoVullo et al. (2006, 2009) have constructed a series of Francisella shuttle vectors (pMP series) with varying portions of pFNL10. These studies established that shuttle vectors containing the ori, repA, and ORF2 regions of pFNL10, along with kanamycin resistance expressed from the Francisella groEL promoter, can be transformed at high efficiency into F. tularensis strains. Moreover, the addition of either the ORF4–ORF5 toxin–antitoxin system, or ORF3 (which increases plasmid copy number within the cell) from pFNL10 to the shuttle vector allowed for stable maintenance of the plasmid in the absence of antibiotic selection. It should be noted that the pFNLTP and pMP vectors and the pKK202 derivatives arose from the same progenitor, pFNL10, and thus belong to the same incompatibility group.
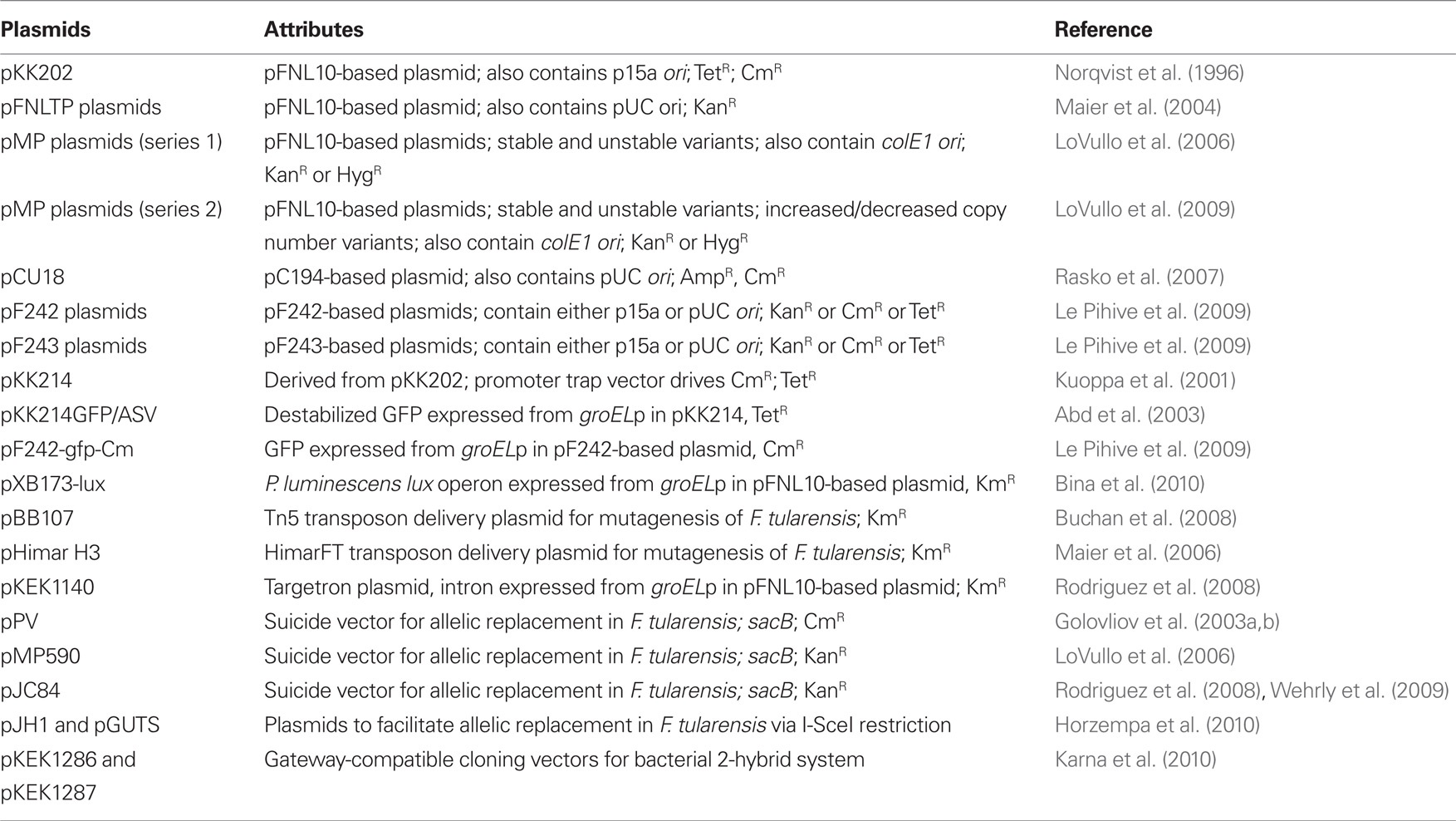
A Francisella shuttle vector based on the replicon of a different incompatibility group has been constructed, utilizing the origin from the Staphylococcus aureus pC194 plasmid, along with the pUC ori. This plasmid, pCU18, can be maintained in F. novicida/F. tularensis cells already carrying plasmids derived from pFNL10 (Rasko et al., 2007). This plasmid has not yet been adopted for widespread use in F. novicida/F. tularensis, and the presence of the chloramphenicol resistance marker prevents its usage in select agent forms of F. tularensis. However, additional shuttle vectors have recently been developed from F. philomiragia plasmids pF242 and pF243 (Le Pihive et al., 2009). pF242 is closely related to pC194, and can coexist in the same F. tularensis cells as pFNL10-based plasmids, and pF243 is closely related to pFNL10-based plasmids. Useful shuttle vectors utilizing these origins of replication have been developed for F. tularensis with a variety of antibiotic resistance markers (Table 1).
Gene Complementation
The pFNL10 derivative vectors discussed above have been used to achieve gene expression/complementation in trans. Researchers have typically either used native promoters for in trans expression, or frequently the F. tularensis groE promoter. The groE promoter was one of the first Francisella promoters identified for high-level expression within host cells (Ericsson et al., 1997), and this promoter has subsequently been utilized for expression of genes in trans, as well as antibiotic resistance markers (Forslund et al., 2006). Another promoter identified for high-level expression in F. novicida in vivo is that for FTN1451, which encodes a protein of unknown function (Gallagher et al., 2007); the FTN1451 promoter has been used to drive high-level expression of antibiotic resistance markers in various F. tularensis strains (Liu et al., 2007). Full complementation of chromosomal mutations utilizing these promoters to drive in trans expression off a pFNL10 derivative plasmid has been notoriously difficult in Francisella strains, especially in tissue culture. This incomplete complementation is likely due to a combination of increased copy number due to the relatively high-copy number plasmid, non-native expression due to the groE promoter being used, and possibly instability of plasmids lacking the toxin–antitoxin system.
The copy number effect of in trans complementation can be overcome by in cis complementation, by introducing the expression construct onto the Francisella chromosome. The first examples of complementation in Francisella involved in cis complementation, but this technique has not subsequently been utilized extensively, although there are notable examples of its use in F. novicida, as well as in F. tularensis subsp. holarctica (Mdluli et al., 1994; Baron and Nano, 1998; Forslund et al., 2006; de Bruin et al., 2007). A technique for in cis complementation in F. novicida has been developed that targets the FTN_1758 gene (de Bruin et al., 2007); because this gene is not found in F. tularensis strains this method is not applicable for use in these organisms. LoVullo et al. (2009) have recently developed two systems for in cis complementation in F. tularensis, using suicide vectors to target a Tn7 insertion site near the glmS gene, and to replace the blaB gene.
Random Mutagenesis
Transposon-mediated random mutagenesis is an invaluable tool for the generation of populations of mutant bacteria, which can then be screened for the phenotype of interest; the mutated gene can then be easily identified by the identification of the location of the transposon. The first transposon mutants were constructed in F. novicida, utilizing a Tn5-based transposon to mutagenize F. novicida DNA that was subsequently used to transform F. novicida, which resulted in the identification of genes important for intramacrophage growth and virulence (Baron and Nano, 1998). While this first study was successful, the technique was cumbersome and only applicable to F. novicida, and the transposon was discovered to be unstably integrated (Lauriano et al., 2003).
A major milestone was achieved with the creation of the first transposon mutants in LVS, utilizing the Tn5 derivative transposon EZ::TN Kan (Kawula et al., 2004). This technique uses a Tn5-based transposon complexed with transposase, which is introduced directly into the Francisella cells via transformation; once the complex enters the cell, the transposase mediates random insertion of the transposon into the chromosome. The transposon insertion frequency is high and allowed for the creation of a large pool of Tn mutants in LVS (Kawula et al., 2004). A further adaptation of this technique was used to generate “tagged” transposons for a signature-tag mutagenesis study of LVS (Kawula et al., 2004; Su et al., 2007). Derivatives of the Tn5-based EZ::TN transposon have since been used to make transposon mutant libraries in F. novicida and F. tularensis subsp. tularensis (Qin et al., 2004; Qin and Mann, 2006; Weiss et al., 2007).
The Manoil laboratory utilized a EZ::TN-based approach to create a comprehensive transposon mutant library in F. novicida. Through saturation mutagenesis and high-throughput sequencing, these researchers were able to identify 396 genes that lacked Tn insertions, indicating that these are essential genes. An important observation during this effort was that the antibiotic resistance gene present in the transposon requires a Francisella promoter to achieve truly random insertion in non-essential genes. The resulting comprehensive library contains at least two independent Tn insertions in each of the 1,490 non-essential F. novicida genes. This library is a valuable resource that has been subsequently used to identify genes involved in virulence (Gallagher et al., 2007).
Variants of the EZ::TN transposon have been created that contain additional elements, including FRT sites for subsequent excision of the antibiotic resistance gene via FLP-mediated recombination (Gallagher et al., 2007), R6Kγori, a conditional origin of replication that is used to rescue the transposon after insertion into the genome (Qin et al., 2004), and outward facing T7 promoters to facilitate identification of the insertion site (Tempel et al., 2006; Weiss et al., 2007).
A further variation on Tn5-based mutagenesis was the development of a plasmid delivery system for transposon mutagenesis. The Tn5-based transposon, rather than being directly transformed as a transposon–transposase complex, is instead delivered on the plasmid pBB107, which is a shuttle vector that also contains the transposase under inducible control, as well as a temperature-sensitive pFNL10-based ori (Maier et al., 2004; Buchan et al., 2008); following Tn insertion into the chromosome the plasmid can be removed with elevated temperature. Variants of this transposon contain additional useful attributes, including FLP recombination sites, the R6K ori, and transcriptional reporters (luxCDABE and lacZ; Buchan et al., 2008).
The Himar-based transposons are characterized to exhibit the most random insertion of all known transposons, due to the minimal 2 bp recognition sequence (AT) utilized by this transposon. Due to the AT-rich nature of the F. tularensis genome, Himar transposons would be predicted to be particularly useful, and a Himar1-based transposon has been developed for use in Francisella (HimarFT). HimarFT was used to create transposon mutants in LVS (Maier et al., 2006), which allowed for the identification of additional virulence genes (Maier et al., 2007).
Targeted Gene Disruption
Targeted gene disruption is one of the major goals when developing genetic techniques for any organism. F. novicida is readily manipulated to quickly inactivate target genes in techniques that rely on double homologous recombination, primarily due to its ability to be transformed with linear DNA. However, the techniques utilizing this capability in F. novicida have failed to work in F. tularensis, as mentioned above. Still, techniques are available to specifically inactivate genes in F. tularensis through a two-step process, where the first homologous recombination event is selected through positive selection, and the second homologous recombination event is counter selected. An alternative technique involving inactivation by retargeted Group II introns has also been developed that works efficiently in F. tularensis strains.
Genes in F. novicida can be inactivated by transformation with linear PCR products that consist of an antibiotic resistance gene flanked by at least 500 bp of flanking homology to the gene of interest (Lauriano et al., 2003; Liu et al., 2007). Increasing the amount of flanking homology in the linear fragment increases the efficiency of mutant generation, and expression of the antibiotic resistance gene by a Francisella promoter is important for successful mutagenesis. Although this technique results in an antibiotic cassette inserted in the target gene, our studies suggest that the cassette does not inhibit expression of downstream genes. This technique has allowed for rapid generation of F. novicida mutants for the study of the genetic basis of pathogenesis.
For targeted mutagenesis in F. tularensis, suicide vectors have been developed to mediate the two-step chromosomal gene replacement technique. The first targeted mutant constructed in a F. tularensis strain was an iglC deletion mutant in LVS (Golovliov et al., 2003b). These researchers utilized a plasmid, pPV, with a pUC-based ori (which fails to function in F. tularensis), an antibiotic resistance marker (for selection), and the counter selectable marker sacB (which causes toxicity in the presence of sucrose). The two-step process consists of (a) transforming the cells with the suicide plasmid containing the deletion and/or insertion in the targeted gene, (b) selection for transformants with the antibiotic resistance gene located on the suicide plasmid (this selects for the first homologous recombination event that results in the plasmid integrating into the chromosome at the target site), (c) counter selection by growth on sucrose (this selects for second homologous recombination event that replaces chromosomal gene with the construct on plasmid). This technique has been used to modify or inactivate a number of genes in LVS (Golovliov et al., 2003a; Bakshi et al., 2006; Gil et al., 2006; LoVullo et al., 2006; Pechous et al., 2006; Ramakrishnan et al., 2008; Sen et al., 2010) and F. tularensis subsp. tularensis (Twine et al., 2005; LoVullo et al., 2006; Lindgren et al., 2007; Thomas et al., 2007; Michell et al., 2010).
This technique is a powerful tool to create chromosomal deletion mutations, which have the dual advantages of lack of polarity and lack of antibiotic resistance. Some important aspects of this technique need to be noted. The second recombination event counter selected by growth on sucrose results in the loss of the plasmid, but sucrose resistant colonies need to be screened to identify the substituted chromosomal locus, because the second recombination event can also regenerate the wildtype locus. Also, the amount of flanking homology surrounding the deletion and/or insertion is important, with 1000 bp required for efficient recombination. Finally, efficient transformation of the vector is critical, since the plasmid cannot replicate in F. tularensis and therefore must recombine into the chromosome immediately following transformation. This last issue can be alleviated by utilizing a vector that can be mobilized into F. tularensis via conjugation, which is frequently a more efficient process than transformation, and Celli and colleagues have developed a mobilizable suicide vector, pJC84, with these attributes (Wehrly et al., 2009).
Recently, a technique has been developed for Francisella targeted mutagenesis using a clever approach to generating a double-strand break in the chromosome that serves as a substrate for the bacterial DNA repair/recombination machinery. This method utilizes the I-SceI restriction enzyme, which has an unusually large 18 bp recognition sequence, such that the restriction site is not normally found in the chromosomal sequence. By first allowing a deletion construct in a suicide plasmid with a I-SceI site (pJH1) to integrate into the F. tularensis chromosome, and then transforming this merodiploid strain with a plasmid that expresses the I-SceI enzyme (pGUTS), these researchers were able to generate F. tularensis strains with unmarked deletions in the chromosome (Horzempa et al., 2010).
An alternate strategy for targeted gene inactivation in F. tularensis has been developed that utilizes group II intron insertion into the gene of interest. Group II introns form a ribonucleoprotein complex that targets the site for insertion by basepairing between the RNA and the chromosomal DNA. Thus Group II introns can be retargeted to insert at specific sites by altering the sequence of the RNA component, and this technique (referred to as Targetron) has been exploited to disrupt genes in Gram-positive and Gram-negative bacteria (Karberg et al., 2001; Frazier et al., 2003; Perutka et al., 2004; Chen et al., 2005; Yao et al., 2006; Rodriguez et al., 2008, 2009). Our laboratory has optimized this technique for use in F. tularensis (Rodriguez et al., 2008, 2009), which involves (a) identification of target sites in specific gene sequence via computer algorithm, (b) “retargeting” the Group II intron via PCR and subsequent cloning into Targetron plasmid (pKEK1140), (c) transformation of Targetron plasmid into F. tularensis, (d) screen colonies for insertion of intron into target gene, and (e) remove temperature-sensitive Targetron plasmid by growth at elevated temperature. The ribonucleoprotein complex functions independent of host recombination machinery, so the key to this technique functioning in any given organism is achieving expression of the ribonucleoprotein complex, which is accomplished by a Francisella promoter.
This technique has several advantages, including the continuous expression of the ribonucleoprotein complex within the cells, which leads to efficient target site insertion. Also, the intron lacks an antibiotic resistance marker, which is advantageous when working with select agent strains of F. tularensis. Finally, this technique allows for the inactivation of two identical genes simultaneously, which has allowed for simultaneous inactivation of both copies of the duplicated FPI genes in F. tularensis subsp. tularensis. The disadvantage of this technique is that it results in an insertion in, not a deletion of, the gene of interest, which may affect downstream gene expression.
Reporter Constructs
Plasmids and transposons can be engineered to serve as reporters of gene expression, and a number of these genetic constructs now exist to facilitate the study of F. tularensis pathogenesis. The first attempt to identify genes upregulated during F. tularensis growth inside cells utilized a promoter “trap” construct in which chromosomal DNA fragments were cloned in the plasmid pKK214 (Kuoppa et al., 2001), to identify promoters that drive expression of a chloramphenicol reporter gene, which led to survival in chloramphenicol-treated host cells. This initial study led to the identification of the groE promoter, which is still used to drive expression of various genes in F. tularensis constructs. Additional studies have also utilized this construct (Baron et al., 1999; Abd et al., 2003), but restrictions on antibiotic resistance prevent its usage in select agent forms of F. tularensis in the US. Promoter trap plasmids have also been developed for F. tularensis that drive expression of GFP (Rasko et al., 2007). Utilizing a promoter trap plasmid that measured expression of red fluorescent protein in the heterologous host E. coli, the glucose-repressible promoter for the LVS gene FTL_0580 was identified and characterized, which may provide a useful tool for regulated expression in F. tularensis in response to glucose concentration (Horzempa et al., 2008). Transposons have also been engineered to function as reporters of gene expression in F. tularensis; utilizing both lacZ and luciferase containing transposons, Buchan et al. (2008) were able to identify iron-regulated genes.
Additional useful reporter plasmids have been created for the visualization of F. tularensis. The plasmid pKK214GFP-ASV was created to provide high-level GFP expression in F. tularensis for use in fluorescent imaging experiments; the GFP in this plasmid has been destabilized by the addition of a C-terminal tag, which prevents build-up of GFP in the cell (Abd et al., 2003). Likewise, pF242- and pF243-derived plasmids were developed that express GFP (Le Pihive et al., 2009). Finally, Francisella plasmids expressing the Photorhabdus luminescens lux operon and GFP have been developed to facilitate in vivo imaging of F. tularensis infected live animals (Bina et al., 2010).
Gateway Cloning
Gateway® cloning (Invitrogen) is a method that utilizes bacteriophage lambda recombination proteins to circumvent the need for traditional restriction/ligation cloning. In this method, ORFs can be easily moved from “entry” plasmids to “destination” plasmids by addition of the lambda proteins (Aguiar et al., 2004; Gao et al., 2008; Matsuyama and Yoshida, 2009). A complete Gateway® entry clone set representing all the ORFs from the F. tularensis subsp. tularensis genome has been constructed and is available through the Pathogen Functional Genomics Resource Center (PFGRC; http://pfgrc.jcvi.org/). The only thing limiting broad use of this clone set is a lack of appropriate destination vectors modified for use in F. tularensis. Our laboratory has recently developed a Gateway-compatible bacterial 2-hybrid system that can be used to study the interactions between proteins, and demonstrated that it can be used to detect interactions between F. tularensis ORFs present in the clone set (IglA and IglB; Karna et al., 2010). Hopefully more Gateway-compatible Francisella plasmids will be created in the future, e.g., for routine complementation, to take advantage of this tremendous resource.
Perspective
Genetic techniques and tools for the study of F. tularensis have expanded dramatically in the past several years. With this new arsenal of methods, the near future should represent the “golden age” of the study of the genetic basis of F. tularensis pathogenesis. This will afford greater insight into why F. tularensis is able to cause disease and evade host immune mechanisms, and ultimately lead to novel ways to prevent and treat tularemia.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgment
Karl E. Klose is supported by NIH AI57986.
References
Abd, H., Johansson, T., Golovliov, I., Sandstrom, G., and Forsman, M. (2003). Survival and growth of Francisella tularensis in Acanthamoeba castellanii. Appl. Environ. Microbiol. 69, 600–606.
Aguiar, J. C., LaBaer, J., Blair, P. L., Shamailova, V. Y., Koundinya, M., Russell, J. A., Huang, F., Mar, W., Anthony, R. M., Witney, A., Caruana, S. R., Brizuela, L., Sacci, J. B. Jr., Hoffman, S. L., and Carucci, D. J. (2004). High-throughput generation of P. falciparum functional molecules by recombinational cloning. Genome Res. 14, 2076–2082.
Anthony, L. S., Cowley, S. C., Mdluli, K. E., and Nano, F. E. (1994). Isolation of a Francisella tularensis mutant that is sensitive to serum and oxidative killing and is avirulent in mice: correlation with the loss of MinD homologue expression. FEMS Microbiol. Lett. 124, 157–165.
Anthony, L. S., Gu, M. Z., Cowley, S. C., Leung, W. W., and Nano, F. E. (1991). Transformation and allelic replacement in Francisella spp. J. Gen. Microbiol. 137, 2697–2703.
Antibiotic Selection Guide. (2004). Johns Hopkins Division of Infectious Diseases. Available at: http://hopkins-abxguide.org/
Bakshi, C. S., Malik, M., Regan, K., Melendez, J. A., Metzger, D. W., Pavlov, V. M., and Sellati, T. J. (2006). Superoxide dismutase B gene (sodB)-deficient mutants of Francisella tularensis demonstrate hypersensitivity to oxidative stress and attenuated virulence. J. Bacteriol. 188, 6443–6448.
Barker, J. R., Chong, A., Wehrly, T. D., Yu, J. J., Rodriguez, S. A., Liu, J., Celli, J., Arulanandam, B. P., and Klose, K. E. (2009). The Francisella tularensis pathogenicity island encodes a secretion system that is required for phagosome escape and virulence. Mol. Microbiol. 74, 1459–1470.
Baron, G. S., and Nano, F. E. (1998). MglA and MglB are required for the intramacrophage growth of Francisella novicida. Mol. Microbiol. 29, 247–259.
Baron, G. S., Reilly, T. J., and Nano, F. E. (1999). The respiratory burst-inhibiting acid phosphatase AcpA is not essential for the intramacrophage growth or virulence of Francisella novicida. FEMS Microbiol. Lett. 176, 85–90.
Bina, X. R., Miller, M. A., and Bina, J. E. (2010). Construction of a bioluminescence reporter plasmid for Francisella tularensis. Plasmid 64, 156–161.
Bina, X. R., Wang, C., Miller, M. A., and Bina, J. E. (2006). The Bla2 beta-lactamase from the live-vaccine strain of Francisella tularensis encodes a functional protein that is only active against penicillin-class beta-lactam antibiotics. Arch. Microbiol. 186, 219–228.
Buchan, B. W., McLendon, M. K., and Jones, B. D. (2008). Identification of differentially regulated Francisella tularensis genes by use of a newly developed Tn5-based transposon delivery system. Appl. Environ. Microbiol. 74, 2637–2645.
Champion, M. D., Zeng, Q., Nix, E. B., Nano, F. E., Keim, P., Kodira, C. D., Borowsky, M., Young, S., Koehrsen, M., Engels, R., Pearson, M., Howarth, C., Larson, L., White, J., Alvarado, L., Forsman, M., Bearden, S. W., Sjostedt, A., Titball, R., Michell, S. L., Birren, B., and Galagan, J. (2009). Comparative genomic characterization of Francisella tularensis strains belonging to low and high virulence subspecies. PLoS Pathog. 5, e1000459. doi: 10.1371/journal.ppat.1000459
Chen, Y., McClane, B. A., Fisher, D. J., Rood, J. I., and Gupta, P. (2005). Construction of an alpha toxin gene knockout mutant of Clostridium perfringens type A by use of a mobile group II intron. Appl. Environ. Microbiol. 71, 7542–7547.
de Bruin, O. M., Ludu, J. S., and Nano, F. E. (2007). The Francisella pathogenicity island protein IglA localizes to the bacterial cytoplasm and is needed for intracellular growth. BMC Microbiol. 7, 1. doi: 10.1186/1471-2180-7-1
Eigelsbach, H. T., and Downs, C. M. (1961). Prophylactic effectiveness of live and killed tularemia vaccines. I. Production of vaccine and evaluation in the white mouse and guinea pig. J. Immunol. 87, 415–425.
Enderlin, G., Morales, L., Jacobs, R. F., and Cross, J. T. (1994). Streptomycin and alternative agents for the treatment of tularemia: review of the literature. Clin. Infect. Dis. 19, 42–47.
Ericsson, M., Golovliov, I., Sandstrom, G., Tarnvik, A., and Sjostedt, A. (1997). Characterization of the nucleotide sequence of the groE operon encoding heat shock proteins chaperone-60 and -10 of Francisella tularensis and determination of the T-cell response to the proteins in individuals vaccinated with F. tularensis. Infect. Immun. 65, 1824–1829.
Forslund, A. L., Kuoppa, K., Svensson, K., Salomonsson, E., Johansson, A., Bystrom, M., Oyston, P. C., Michell, S. L., Titball, R. W., Noppa, L., Frithz-Lindsten, E., Forsman, M., and Forsberg, A. (2006). Direct repeat-mediated deletion of a type IV pilin gene results in major virulence attenuation of Francisella tularensis. Mol. Microbiol. 59, 1818–1830.
Frank, D. W., and Zahrt, T. C. (2007). Genetics and genetic manipulation in Francisella tularensis. Ann. N. Y. Acad. Sci. 1105, 67–97.
Frazier, C. L., San Filippo, J., Lambowitz, A. M., and Mills, D. A. (2003). Genetic manipulation of Lactococcus lactis by using targeted group II introns: generation of stable insertions without selection. Appl. Environ. Microbiol. 69, 1121–1128.
Gallagher, L. A., McKevitt, M., Ramage, E. R., and Manoil, C. (2008). Genetic dissection of the Francisella novicida restriction barrier. J. Bacteriol. 190, 7830–7837.
Gallagher, L. A., Ramage, E., Jacobs, M. A., Kaul, R., Brittnacher, M., and Manoil, C. (2007). A comprehensive transposon mutant library of Francisella novicida, a bioweapon surrogate. Proc. Natl. Acad. Sci. U.S.A. 104, 1009–1014.
Gao, H., Pattison, D., Yan, T., Klingeman, D. M., Wang, X., Petrosino, J., Hemphill, L., Wan, X., Leaphart, A. B., Weinstock, G. M., Palzkill, T., and Zhou, J. (2008). Generation and validation of a Shewanella oneidensis MR-1 clone set for protein expression and phage display. PLoS ONE 3, e2983. doi: 10.1371/journal.pone.0002983
Gil, H., Platz, G. J., Forestal, C. A., Monfett, M., Bakshi, C. S., Sellati, T. J., Furie, M. B., Benach, J. L., and Thanassi, D. G. (2006). Deletion of TolC orthologs in Francisella tularensis identifies roles in multidrug resistance and virulence. Proc. Natl. Acad. Sci. U.S.A. 103, 12897–12902.
Golovliov, I., Baranov, V., Krocova, Z., Kovarova, H., and Sjostedt, A. (2003a). An attenuated strain of the facultative intracellular bacterium Francisella tularensis can escape the phagosome of monocytic cells. Infect. Immun. 71, 5940–5950.
Golovliov, I., Sjostedt, A., Mokrievich, A., and Pavlov, V. (2003b). A method for allelic replacement in Francisella tularensis. FEMS Microbiol. Lett. 222, 273–280.
Hofinger, D. M., Cardona, L., Mertz, G. J., and Davis, L. E. (2009). Tularemic meningitis in the United States. Arch. Neurol. 66, 523–527.
Horzempa, J., Carlson, P. E. Jr., O’Dee, D. M., Shanks, R. M., and Nau, G. J. (2008). Global transcriptional response to mammalian temperature provides new insight into Francisella tularensis pathogenesis. BMC Microbiol. 8, 172. doi: 10.1186/1471-2180-8-172
Horzempa, J., Shanks, R. M., Brown, M. J., Russo, B. C., O’Dee, D. M., and Nau, G. J. (2010). Utilization of an unstable plasmid and the I-SceI endonuclease to generate routine markerless deletion mutants in Francisella tularensis. J. Microbiol. Methods 80, 106–108.
Kalivoda, E. J., Horzempa, J., Stella, N. A., Sadaf, A., Kowalski, R. P., Nau, G. J., and Shanks, R. M. (2010). New vector tools with a hygromycin resistance marker for use with opportunistic pathogens. Mol. Biotechnol. doi: 10.1007/s12033-010-9342-x. [Epub ahead of print].
Karberg, M., Guo, H., Zhong, J., Coon, R., Perutka, J., and Lambowitz, A. M. (2001). Group II introns as controllable gene targeting vectors for genetic manipulation of bacteria. Nat. Biotechnol. 19, 1162–1167.
Karna, S. L., Zogaj, X., Barker, J., Seshu, J., Dove, S., and Klose, K. (2010). A bacterial two-hybrid system that utilizes Gateway cloning for rapid screening of protein-protein interactions. BioTechniques 49, 831–833.
Kawula, T. H., Hall, J. D., Fuller, J. R., and Craven, R. R. (2004). Use of transposon–transposase complexes to create stable insertion mutant strains of Francisella tularensis LVS. Appl. Environ. Microbiol. 70, 6901–6904.
Kuoppa, K., Forsberg, A., and Norqvist, A. (2001). Construction of a reporter plasmid for screening in vivo promoter activity in Francisella tularensis. FEMS Microbiol. Lett. 205, 77–81.
Lauriano, C. M., Barker, J. R., Nano, F. E., Arulanandam, B. P., and Klose, K. E. (2003). Allelic exchange in Francisella tularensis using PCR products. FEMS Microbiol. Lett. 229, 195–202.
Le Pihive, E., Blaha, D., Chenavas, S., Thibault, F., Vidal, D., and Valade, E. (2009). Description of two new plasmids isolated from Francisella philomiragia strains and construction of shuttle vectors for the study of Francisella tularensis. Plasmid 62, 147–157.
Lindgren, H., Shen, H., Zingmark, C., Golovliov, I., Conlan, W., and Sjostedt, A. (2007). The resistance of Francisella strains against reactive nitrogen and oxygen species with special reference to the role of KatG. Infect. Immun. 75, 1303–1309.
Liu, J., Zogaj, X., Barker, J. R., and Klose, K. E. (2007). Construction of targeted insertion mutations in Francisella tularensis subsp. novicida. Biotechniques 43, 487–492.
LoVullo, E. D., Molins-Schneekloth, C. R., Schweizer, H. P., and Pavelka, M. S., Jr. (2009). Single-copy chromosomal integration systems for Francisella tularensis. Microbiology 155, 1152–1163.
LoVullo, E. D., Sherrill, L. A., Perez, L. L., and Pavelka, M. S., Jr. (2006). Genetic tools for highly pathogenic Francisella tularensis subsp. tularensis. Microbiology 152, 3425–3435.
Maier, T. M., Casey, M. S., Becker, R. H., Dorsey, C. W., Glass, E. M., Maltsev, N., Zahrt, T. C., and Frank, D. W. (2007). Identification of Francisella tularensis Himar1-based transposon mutants defective for replication in macrophages. Infect. Immun. 75, 5376–5389.
Maier, T. M., Havig, A., Casey, M., Nano, F. E., Frank, D. W., and Zahrt, T. C. (2004). Construction and characterization of a highly efficient Francisella shuttle plasmid. Appl. Environ. Microbiol. 70, 7511–7519.
Maier, T. M., Pechous, R., Casey, M., Zahrt, T. C., and Frank, D. W. (2006). In vivo Himar1-based transposon mutagenesis of Francisella tularensis. Appl. Environ. Microbiol. 72, 1878–1885.
Matsuyama, A., and Yoshida, M. (2009). Systematic cloning of an ORFeome using the gateway system. Methods Mol. Biol. 577, 11–24.
Maurin, M., Mersali, N. F., and Raoult, D. (2000). Bactericidal activities of antibiotics against intracellular Francisella tularensis. Antimicrob. Agents Chemother. 44, 3428–3431.
McLendon, M. K., Apicella, M. A., and Allen, L. A. (2006). Francisella tularensis: taxonomy, genetics, and immunopathogenesis of a potential agent of biowarfare. Annu. Rev. Microbiol. 60, 167–185.
Mdluli, K. E., Anthony, L. S., Baron, G. S., McDonald, M. K., Myltseva, S. V., and Nano, F. E. (1994). Serum-sensitive mutation of Francisella novicida: association with an ABC transporter gene. Microbiology 140(Pt 12), 3309–3318.
Michell, S. L., Dean, R. E., Eyles, J. E., Hartley, M. G., Waters, E., Prior, J. L., Titball, R. W., and Oyston, P. C. (2010). Deletion of the Bacillus anthracis capB homologue in Francisella tularensis subspecies tularensis generates an attenuated strain that protects mice against virulent tularaemia. J. Med. Microbiol. 59, 1275–1284.
Mohapatra, N. P., Balagopal, A., Soni, S., Schlesinger, L. S., and Gunn, J. S. (2007). AcpA is a Francisella acid phosphatase that affects intramacrophage survival and virulence. Infect. Immun. 75, 390–396.
Norqvist, A., Kuoppa, K., and Sandstrom, G. (1996). Construction of a shuttle vector for use in Francisella tularensis. FEMS Immunol. Med. Microbiol. 13, 257–260.
Pechous, R., Celli, J., Penoske, R., Hayes, S. F., Frank, D. W., and Zahrt, T. C. (2006). Construction and characterization of an attenuated purine auxotroph in a Francisella tularensis live vaccine strain. Infect. Immun. 74, 4452–4461.
Perutka, J., Wang, W., Goerlitz, D., and Lambowitz, A. M. (2004). Use of computer-designed group II introns to disrupt Escherichia coli DExH/D box protein and DNA helicase genes. J. Mol. Biol. 336, 421–439.
Pomerantsev, A. P., Golovliov, I. R., Ohara, Y., Mokrievich, A. N., Obuchi, M., Norqvist, A., Kuoppa, K., and Pavlov, V. M. (2001). Genetic organization of the Francisella plasmid pFNL10. Plasmid 46, 210–222.
Qin, A., and Mann, B. J. (2006). Identification of transposon insertion mutants of Francisella tularensis tularensis strain Schu S4 deficient in intracellular replication in the hepatic cell line HepG2. BMC Microbiol. 6, 69. doi: 10.1186/1471-2180-6-69
Qin, A., Tucker, A. M., Hines, A., and Wood, D. O. (2004). Transposon mutagenesis of the obligate intracellular pathogen Rickettsia prowazekii. Appl. Environ. Microbiol. 70, 2816–2822.
Ramakrishnan, G., Meeker, A., and Dragulev, B. (2008). fslE is necessary for siderophore-mediated iron acquisition in Francisella tularensis Schu S4. J. Bacteriol. 190, 5353–5361.
Rasko, D. A., Esteban, C. D., and Sperandio, V. (2007). Development of novel plasmid vectors and a promoter trap system in Francisella tularensis compatible with the pFLN10 based plasmids. Plasmid 58, 159–166.
Rodriguez, S. A., Davis, G., and Klose, K. E. (2009). Targeted gene disruption in Francisella tularensis by group II introns. Methods 49, 270–274.
Rodriguez, S. A., Yu, J. J., Davis, G., Arulanandam, B. P., and Klose, K. E. (2008). Targeted inactivation of Francisella tularensis genes by group II introns. Appl. Environ. Microbiol. 74, 2619–2626.
Rohmer, L., Fong, C., Abmayr, S., Wasnick, M., Larson Freeman, T. J., Radey, M., Guina, T., Svensson, K., Hayden, H. S., Jacobs, M., Gallagher, L. A., Manoil, C., Ernst, R. K., Drees, B., Buckley, D., Haugen, E., Bovee, D., Zhou, Y., Chang, J., Levy, R., Lim, R., Gillett, W., Guenthener, D., Kang, A., Shaffer, S. A., Taylor, G., Chen, J., Gallis, B., D’Argenio, D. A., Forsman, M., Olson, M. V., Goodlett, D. R., Kaul, R., Miller, S. I., and Brittnacher, M. J. (2007). Comparison of Francisella tularensis genomes reveals evolutionary events associated with the emergence of human pathogenic strains. Genome Biol. 8, R102.
Sen, B., Meeker, A., and Ramakrishnan, G. (2010). The fslE homolog, FTL_0439 (fupA/B), mediates siderophore-dependent iron uptake in Francisella tularensis LVS. Infect. Immun. 78, 4276–4285.
Su, J., Yang, J., Zhao, D., Kawula, T. H., Banas, J. A., and Zhang, J. R. (2007). Genome-wide identification of Francisella tularensis virulence determinants. Infect. Immun. 75, 3089–3101.
Tempel, R., Lai, X. H., Crosa, L., Kozlowicz, B., and Heffron, F. (2006). Attenuated Francisella novicida transposon mutants protect mice against wild-type challenge. Infect. Immun. 74, 5095–5105.
Thomas, R. M., Titball, R. W., Oyston, P. C., Griffin, K., Waters, E., Hitchen, P. G., Michell, S. L., Grice, I. D., Wilson, J. C., and Prior, J. L. (2007). The immunologically distinct O antigens from Francisella tularensis subspecies tularensis and Francisella novicida are both virulence determinants and protective antigens. Infect. Immun. 75, 371–378.
Twine, S., Bystrom, M., Chen, W., Forsman, M., Golovliov, I., Johansson, A., Kelly, J., Lindgren, H., Svensson, K., Zingmark, C., Conlan, W., and Sjostedt, A. (2005). A mutant of Francisella tularensis strain SCHU S4 lacking the ability to express a 58-kilodalton protein is attenuated for virulence and is an effective live vaccine. Infect. Immun. 73, 8345–8352.
Tyeryar, F. J. Jr., and Lawton, W. D. (1969). Transformation of Pasteurella novicida. J. Bacteriol. 100, 1112–1113.
Tyeryar, F. J., and Lawton, W. D. (1970). Factors affecting transformation of Pasteurella novicida. J. Bacteriol. 104, 1312–1317.
Urich, S. K., and Petersen, J. M. (2008). In vitro susceptibility of isolates of Francisella tularensis types A and B from North America. Antimicrob. Agents Chemother. 52, 2276–2278.
Wehrly, T. D., Chong, A., Virtaneva, K., Sturdevant, D. E., Child, R., Edwards, J. A., Brouwer, D., Nair, V., Fischer, E. R., Wicke, L., Curda, A. J., Kupko, J. J. 3rd, Martens, C., Crane, D. D., Bosio, C. M., Porcella, S. F., and Celli, J. (2009). Intracellular biology and virulence determinants of Francisella tularensis revealed by transcriptional profiling inside macrophages. Cell. Microbiol. 11, 1128–1150.
Weiss, D. S., Brotcke, A., Henry, T., Margolis, J. J., Chan, K., and Monack, D. M. (2007). In vivo negative selection screen identifies genes required for Francisella virulence. Proc. Natl. Acad. Sci. U.S.A. 104, 6037–6042.
Yao, J., Zhong, J., Fang, Y., Geisinger, E., Novick, R. P., and Lambowitz, A. M. (2006). Use of targetrons to disrupt essential and nonessential genes in Staphylococcus aureus reveals temperature sensitivity of Ll.LtrB group II intron splicing. RNA 12, 1271–1281.
Keywords: tularemia, transposon, allelic exchange, plasmid, targetron, select agent
Citation: Zogaj X and Klose KE (2011) Genetic manipulation of Francisella tularensis. Front. Microbio. 1:142. doi: 10.3389/fmicb.2010.00142
Received: 11 November 2010;
Accepted: 14 December 2010;
Published online: 05 January 2011.
Edited by:
Anders Sjostedt, Umeå University, SwedenReviewed by:
Martin Pavelka, University of Rochester Medical Center, USAChandra Shekhar Bakshi, Albany Medical College, USA
Copyright: © 2011 Zogaj and Klose. This is an open-access article subject to an exclusive license agreement between the authors and the Frontiers Research Foundation, which permits unrestricted use, distribution, and reproduction in any medium, provided the original authors and source are credited.
*Correspondence: Karl E. Klose, South Texas Center for Emerging Infectious Diseases, University of Texas San Antonio, One UTSA Circle, San Antonio, TX 78249, USA. e-mail:a2FybC5rbG9zZUB1dHNhLmVkdQ==
 Xhavit Zogaj
Xhavit Zogaj