- 1 Finnish Forest Research Institute, Southern Finland Regional Unit, Vantaa, Finland
- 2 Department of Forest Sciences, University of Helsinki, Helsinki, Finland
- 3 Department of Environmental Science, University of Eastern Finland, Kuopio, Finland
- 4 Department of Bioresources/Microbiology, Austrian Institute of Technology, Seibersdorf, Austria
- 5 CSIRO Marine and Atmospheric Research, Hobart, TAS, Australia
- 6 School of Forest Sciences, University of Eastern Finland, Joensuu, Finland
It is known that Sphagnum associated methanotrophy (SAM) changes in relation to the peatland water table (WT) level. After drought, rising WT is able to reactivate SAM. We aimed to reveal whether this reactivation is due to activation of indigenous methane (CH4) oxidizing bacteria (MOB) already present in the mosses or to MOB present in water. This was tested through two approaches: in a transplantation experiment, Sphagna lacking SAM activity were transplanted into flark water next to Sphagna oxidizing CH4. Already after 3 days, most of the transplants showed CH4 oxidation activity. Microarray showed that the MOB community compositions of the transplants and the original active mosses had become more similar within 28 days thus indicating MOB movement through water between mosses. Methylocystis-related type II MOB dominated the community. In a following experiment, SAM inactive mosses were bathed overnight in non-sterile and sterile-filtered SAM active site flark water. Only mosses bathed with non-sterile flark water became SAM active, which was also shown by the pmoA copy number increase of over 60 times. Thus, it was evident that MOB present in the water can colonize Sphagnum mosses. This colonization could act as a resilience mechanism for peatland CH4 dynamics by allowing the re-emergence of CH4 oxidation activity in Sphagnum.
Introduction
Peatlands store over one third of global terrestrial carbon (Gorham, 1991). Although these ecosystems are carbon dioxide (CO2) sinks they are also a major source of methane (CH4), formed as the final product of anaerobic degradation of organic matter. Most carbon in these systems is derived from Sphagnum mosses (Clymo and Hayward, 1982), the dominant plant in bog-type northern peatlands. Mosses sequester atmospheric CO2 directly through photosynthesis. Methanotrophic bacteria (MOB) living inside the moss hyaline cells and on leaf surfaces (Raghoebarsing et al., 2005; Kip et al., 2010) also play an important role in carbon binding. These bacteria provide CO2 for the plant via CH4 oxidation, a mechanism that is especially important in submerged conditions where CO2 diffusion is slow (Kip et al., 2010). This phenomenon is of local and global importance as it has been detected in all 23 Sphagnum species of a peatland area (Larmola et al. (2010)) and in geographically distant peatlands (Kip et al., 2010) and may be partly responsible for the lower CH4 emissions of Sphagnum bogs in relation to other peatland types (Nykänen et al., 1998).
About 10–15 (Raghoebarsing et al., 2005) or 10–30% (Larmola et al., 2010) of Sphagnum biomass carbon is from CH4 oxidation by MOB. Thus, it seems clear that mosses benefit from their partners and the relationship has been discussed to be symbiotic (Raghoebarsing et al., 2005). Still, there is evidence that the bacteria involved are only loosely connected to Sphagnum (Basiliko et al., 2004; Larmola et al., 2010). The study by Larmola et al. (2010) showed that peatland water table (WT) level is the main factor influencing MOB activity in mosses. Sphagnum associated methanotrophy (SAM) became de-/reactivated upon natural WT fluctuation. However, Larmola et al. (2010) did not provide evidence whether reactivated CH4 oxidation was caused by reactivation of the original MOB community, invasion of new MOB from the surrounding water or by both mechanisms. The ability of MOB to colonize Sphagnum from surrounding water would make ecosystem CH4 dynamics less vulnerable to extended periods of drought than a tight symbiosis between MOB and Sphagnum or relaying on the reactivation of original community. To test the importance of colonization we examined the question more thoroughly. First, we conducted a similar transplantation trial as in Larmola et al. (2010) where inactive mosses were planted next to active ones. Colonization process was followed by measurement of CH4 oxidation potentials and community analysis by a microarray that profiles diversity within the pmoA gene coding for particulate methane mono-oxygenase (pMMO), a key enzyme in CH4 oxidation (Bodrossy et al., 2003). By using this method covering a wider range of MOB diversity than Larmola et al. (2010) we aimed to reveal more detailed changes in community compositions. We hypothesized that colonization of MOB through the water phase is a substantial reason for methanotrophic reactivation. Since we presume that all mosses are colonized through the same pathway this should be reflected in MOB of the neighboring mosses influencing the microbial community of the transplanted moss. Second, we tested the hypothesis in the laboratory by treating inactive Sphagnum mosses with water from a wet depression (flark) harboring methanotrophic active Sphagnum mosses. As a control, parallel samples were treated with the same water after MOB removal through filtration. CH4 oxidation potentials were measured and MOB communities analyzed by pmoA-based quantitative PCR (qPCR) and denaturing gradient gel electrophoresis (DGGE) analysis followed by sequencing.
Experimental Procedures
Transplantation Experiment
Sphagnum transplantation
The experiment was conducted at the Lakkasuo mire (61°48′N, 24°19′E; 150 m. a.s.l), a boreal raised bog complex in southern Finland. On 7 July 2008, patches (8 cm in diameter) of inactive Sphagnum rubellum from site O were transplanted to six different flark sites (A–F) showing high Sphagnum associated methanotrophic (SAM; CH4 oxidation) activity (Figure 1). To control the effect of transplantation, S. rubellum was replanted in the original site and the native Sphagnum species (Table A1 in Appendix) gathered from each of six different flark sites were also returned to their original places. Thus, all samples were of transplanted Sphagnum. Moss samples were gathered at the beginning of the experiment (0 day), after 3 days, and 28 days. After gathering, mosses were rinsed with deionized water and dried overnight at +4°C. Only upper 10 cm of the moss plants were included in the following analysis. Ecological variables of the transplantation sites are listed in Table A1 in Appendix.
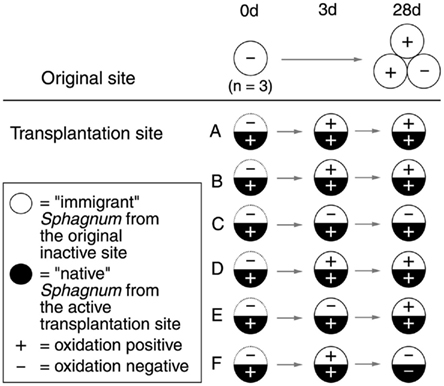
Figure 1. Diagram of the transplantation experiment. CH4 oxidation potentials at the time of sampling are included (±; values over 0.005 μmol g dry mass−1 h−1 were considered positive). Dominant Sphagnum species on transplantation sites were S. papillosum (sites A–C), S. cuspidatum (sites D and E), and S. majus (site F). The original site O and the transplanted immigrant mosses consisted of S. rubellum.
Methane oxidation potential
Methane oxidation potentials were measured as described in Larmola et al. (2010). Briefly, 30 g of moss was incubated in a 600 mL flask with an initial CH4 concentration of 10000 ppm in the dark at +15°C and the oxidation was monitored after 24 and 48 h by gas chromatography. Results are presented in micromole CH4 per gram dry weight per hour (μmol g dw−1 h−1).
Analysis of methanotrophic community composition by pmoA-microarray
Community composition of MOB in Sphagnum samples was investigated using a microarray (Bodrossy et al., 2003) designed to detect diversity within the pmoA gene. DNA was isolated as in Siljanen et al. (2011). A fragment of the pmoA gene was amplified using a semi-nested PCR approach with primer pairs A189f/T7–A682r and A189f/T7–mb661r as in Siljanen et al. (2011) with the exception that after the first PCR-step, products not detectable on the gel were diluted 1:10 before being used as templates in the second PCR. Concentration of PCR products was quantified using a Qubit fluorometer (Invitrogen, Carlsbad, CA, USA). In vitro transcription and hybridization was performed as in Stralis-Pavese et al. (2004) and the applied probe set was similar to that applied by Abell et al. (2009). One of three parallel original inactive moss samples from the 0 day time point could be successfully analyzed (see Larmola et al. (2010) that SAM inactive Sphagnum mosses host MOB DNA). Probing pmoA diversity cannot detect Methylocella or the recently discovered Methyloferula (Vorobev et al., 2011) methanotrophs as the pMMO enzyme is not present in these bacteria. It should be noted that another newly discovered methanotroph group, Verrucomicrobia, is also not detectable by the probes we used.
Statistical analysis of microarray data
The quantitative nature of the microarray data was converted to a binary matrix (presence = 1, absence = 0) to reveal the community changes caused by different groups of MOB colonizing mosses after transplantation, and also to prevent false interpretation originating from non-quantitative nested PCR approach. The data were then analyzed using principal component analysis (PCA) carried out with CANOCO Version 4.52 (ter Braak and Smilauer, 2002). Sites A–F (Figure 1) were analyzed separately (n = 1) and as parallel samples (n = 6). Universal MOB probes (positive controls) and probes not hybridizing to any of the samples were excluded from the analyses (threshold for positive samples ≥3 after normalization of the data to the scale of 0–100).
Bathing Experiment
Flark water bathing
Sphagnum mosses were gathered from Sallie’s Fen in NH, USA (43°12.5′N, 71°03.5′W 110 m. a.s.l.). Triplicate (n = 3) fresh samples of approximately 30 mL (volume based on the volume of water replaced by the mosses) of inactive S. magellanicum were subjected to the following treatments: (I) no treatment; (II) overnight (11 h) incubation in SAM active S. majus 200 mL flark; (III) overnight incubation in 200 mL 0.45 μm filtered SAM active S. majus flark water; (IV) overnight incubation in 200 mL SAM active S. majus flark water followed by rinsing with deionized water. In the final treatment, S. majus gathered from the active (flark) site was included as a positive control (n = 3) in the analyses described below. SAM active flark water was collected directly from a wet depression next to S. majus vegetation and did not contain any macroscopic plant material. Each overnight incubation was conducted in the dark at +20°C. Following treatment, all mosses were dried overnight at +4°C. Only upper 10 cm of the moss plants were included in the following analysis.
Methane oxidation potential and statistical analysis
Methane oxidation potentials were measured as above in the transplantation experiment but on a Shimadzu 14A gas chromatograph equipped with a flame ionization detector (Shimadzu Corp., Kyoto, Japan). The results are presented in micromole CH4 per gram dry weight per hour. The difference between sample treatments was tested using Kruskal–Wallis non-parametric analysis of variance followed by Nemenyi test for pairwise comparisons (p < 0.05; Zar, 1999).
Analysis of community composition by DGGE and sequencing
Diversity of MOB in Sphagnum samples from the bathing experiment was explored by pmoA-based PCR–DGGE analysis and sequencing as previously described (Tuomivirta et al., 2009; Larmola et al., 2010) using the primer pair A189f/GC-621r designed to target methanotrophs abundant in boreal peatlands (Tuomivirta et al., 2009). DNA was isolated as above in the transplantation experiment. Determined pmoA gene sequences were submitted to Genbank under accession numbers HQ651182 and HQ651183.
Quantification of pmoA genes and statistical analysis
Quantitative PCR was carried out as previously described (Tuomivirta et al., 2009) using the same primer pair as in the DGGE analysis (A189f/GC-621r). Results are expressed in pmoA copy number per gram dry weight. To test the difference between sample treatments, values were ln transformed to normalize the data followed by ANOVA and Tukey’s HSD test (p < 0.05).
Results and Discussion
In the transplantation experiment SAM inactive Sphagnum rubellum mosses were planted on six different SAM active sites. Most of the originally inactive mosses showed detectable CH4 oxidation potentials (>0.005 μmol CH4 g−1 h−1) after 3 days and 28 days (Figure 1). Comparing transplantation sites individually indicated that initially different MOB communities became more similar with time (Figure A1 in Appendix). Averaging over the entire data set, the pmoA-microarray showed that the MOB community of the originally inactive mosses started to resemble that of the native mosses in the active site after 3 days (Figure 2). After 28 days, MOB communities from the majority of inactive Sphagnum mosses transplanted to active sites (immigrants in Figure 1) were more similar to those of native active mosses than to those in the inactive site at 0 day. Thus, this field experiment indicated that MOB could be transferred between mosses through the water phase. In addition, the original inactive Sphagnum (site O) became active after transplantation in its original site and had a different MOB community than before transplantation, demonstrating possible new MOB movement through the water phase (Figures 2 and 3 and Figure A1 in Appendix). However, although SAM activity was induced in most of the samples together with invasion of new MOB this was not always the case and SAM activity could also be induced without major changes in community composition (Site E, Figure A1 in Appendix). Thus there are methanotrophs that move through water but we cannot state that for all community members. Some members of the methanotroph community seem to be permanently associated with the mosses regardless of whether conditions favor CH4 oxidation or not. In our transplantation experiment this factor hindered us from seeing the invading members of the community and partly explains why large changes in community composition were not always seen when CH4 oxidation was reactivated. As Methylocystis-related type II MOB were present in practically all samples in the transplantation experiment (Figure 3 and Figure A2 in Appendix), as shown by the probes Mcy413, Mcy522, Mcy270, Mcy459, Msi232, Peat264, II509, and II630, their movement between mosses could not be clearly detected. Still, probes targeting Methylocystis strain M (McyM309), Methylocystis subgroup B (McyB304), and the genus Methylosinus (Mst214, Mst314, MsS475, Msi294, NMsiT.271) gave more variable presence patterns among the type II methanotrophs, suggesting water-mediated movement. Compared to α-proteobacterial type II MOB, the other main methanotrophic lineage, the γ-proteobacterial type I, was present more rarely and with more variability. Prior to transplantation, no type I methanotrophs were detected in inactive mosses. After 28 day, they were present in four of six “immigrant” samples transplanted to active sites. Type Ia subgroup was found only in mosses transplanted in the active flarks and the native mosses of these sites, suggesting that this group moved from the native mosses to the transplanted “immigrants.” Type I MOB could not be clearly linked to the emergence of CH4 oxidation activity, although they were present in most of the active samples and absent from most of the inactive ones. Consequently, based on the transplantation experiment alone, we cannot exclusively state that invasion by nearby MOB is an imperative route in the reactivation of SAM activity.
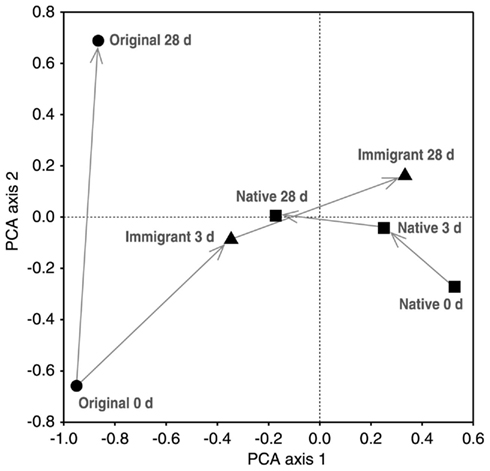
Figure 2. Principal component analysis (PCA) of binary microarray data. Sample means from different sampling times in days (d) are showed [original 0 days = original inactive moss (n = 1), original 28 days = original moss transplanted back to the original site (n = 3), immigrant = original moss transplanted to the active flark sites (n = 6), native = native mosses of the active flark sites (n = 6)]. The first and second PCA axes explain 25 and 11% of the compositional variation, respectively.
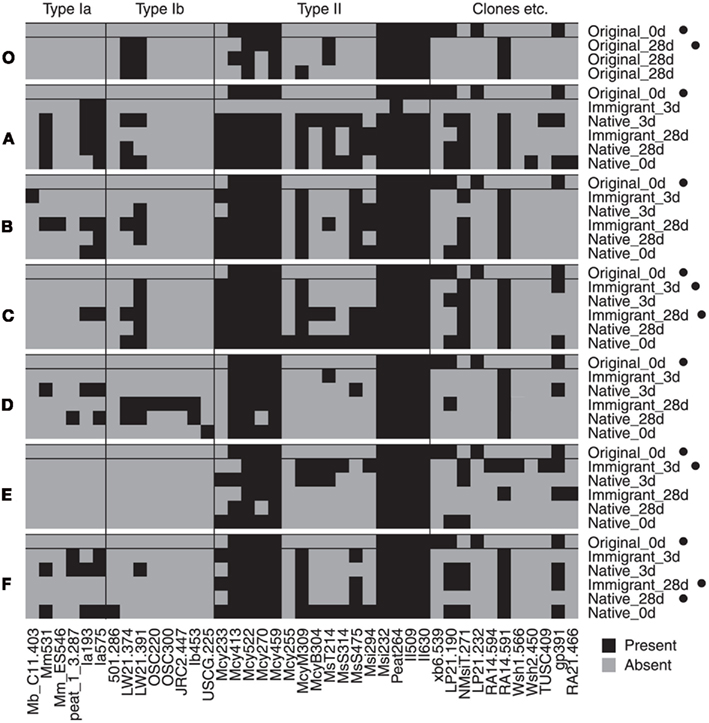
Figure 3. Binary map of microarray results. Each site (A–F and O) is showed separately together with the original 0 day sample data for comparison. Microarray analysis was successful for only one of three parallel inactive original_0 day samples. Samples that were inactive in CH4 oxidation at the time of sampling are marked with a black dot (•). Only probes with a positive hybridization signal (≥3 after normalization of the data to the scale of 0–100) on any of the samples are shown (universal MOB probes were not included). Binary results were converted from the semiquantitative data presented as a heatmap in the Figure A2 in Appendix.
To examine the hypothesis “the colonization of MOB through the water phase is a substantial reason for methanotrophic reactivation” further, a more simplified experiment was conducted in laboratory conditions. In this bathing experiment, SAM inactive S. magellanicum mosses exposed to unfiltered, SAM active flark water began CH4 oxidation within 11 h, as indicated by CH4 oxidation potential measurements and community analyses including pmoA-based qPCR, DGGE fingerprinting and sequencing. Maximum CH4 oxidation potentials and pmoA copy numbers were measured for mosses treated with unfiltered water (averages for rinsed mosses 0.33 μmol CH4 g−1 h−1, 1.9 × 107 pmoA copies g dw−1, and for unrinsed 0.29 μmol CH4 g−1 h−1, 2.6 × 107 pmoA copies g dw−1) and positive control S. majus mosses (average 0.63 μmol CH4 g−1 h−1, 20 × 107 pmoA copies g dw−1) from the active flark site (Figure 4). Respective values for mosses treated with filtered water (no CH4 oxidation detected, 0.03 × 107 pmoA copies g dw−1) and the negative control (<0.005 μmol CH4 g−1 h−1, 0.06 × 107 pmoA copies g dw−1) were clearly lower. The pmoA copy number between filtered and non-filtered flark water treated S. magellanicum increased in average by a factor of 63. Also, DGGE revealed the transfer of two Methylocystis-related methanotrophs through unfiltered water (Figure 5). Filtered water did not induce CH4 oxidation activity. This experiment clearly demonstrated the water-mediated dispersal of MOB, but it also showed that compared to invasion by new MOB, reactivation of the original MOB was not a major mechanism in the reactivation of CH4 oxidation process in the studied mosses. In the DGGE gel the two Methylocystis bands are faintly present already in the unbathed negative control. Bathing of the mosses with sterile-filtered water caused these bands to fade away as shown also by the qPCR. On the contrary, treatment with unfiltered water caused emergence of high numbers of MOB and also high SAM activity. Thus the reactivation of the CH4 oxidation activity must have been brought up by MOB invading the moss through the water phase or the growth in MOB numbers should have been seen also in the moss bathed with filtered water. Moreover, known Methylocystis strains have doubling times of several hours when growing on CH4 in laboratory conditions (Wise et al., 1999; Dedysh et al., 2007; Baani and Liesack, 2008) suggesting that it would be highly unlikely for these MOB to increase their numbers over 60 times higher in 11 h, as seen in the bathing experiment. Therefore we accept the posed hypothesis with a minor modification: MOB colonization through the water phase occurs and it obviously supports the reactivation of CH4 oxidation in Sphagnum mosses, but our experiments cannot rule out the possibility of reactivation of original community members. This could be investigated in a prolonged bathing experiment in combination with diurnal light rhythm, but was beyond the scope of this investigation.
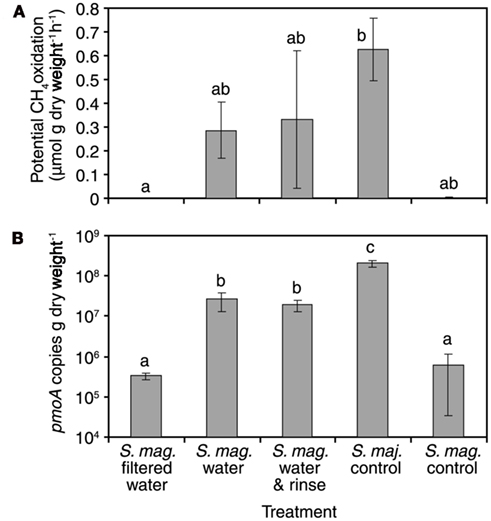
Figure 4. CH4 oxidation potentials (A) and pmoA copy numbers (B) of differently treated moss samples (S. mag. = Sphagnum magellanicum, S. maj. = Sphagnum majus). Standard error bars are shown (n = 3). Same letter superscripts denote non-significant differences among treatments in the potential oxidation rates. Note the logarithmic scale on the y-axis (B).
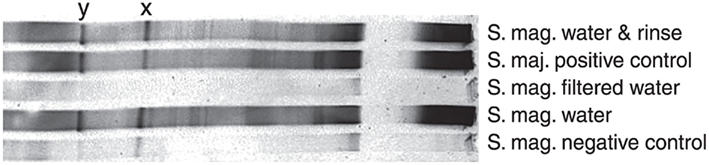

Figure 5. Denaturing gradient gel electrophoresis gel showing differently treated samples. Two Methylocystis bands are marked by x and y (S. mag. = Sphagnum magellanicum, S. maj. = Sphagnum majus). Closest database match to sequence x (HQ651182) was methanotrophic sequence from an acidic fen (FR726177) with 95% similarity. Sequence y (HQ651183) was identical to a Methylocystis sequence previously found on Finnish peatlands (GQ468279, GQ279344, FJ930091) and Sphagnum mosses (GQ121280).
Based on our results that MOB colonize the mosses from water, the relationship between MOB and Sphagnum seems to be a loose, mutually beneficial association rather than a tight symbiosis. This result is in line with a recent finding by Bragina et al. (in press) who showed by pyrosequencing that some bacteria are passed from the Sphagnum sporophyte to the gametophyte but no known methanotrophs were among them. Representatives of the genus Methylocystis, however, were detected in the gametophyte. Still, even though methanotrophs may not be obligately dependent on the mosses, they most likely prefer the plant cells over life in the water phase. This is supported by our results from the bathing experiment. Despite having slightly lowered the amount of pmoA detected, mosses rinsed with sterile water had almost the same potential CH4 oxidation activity as unrinsed ones, indicating that loosely attached methanotrophs play only a minor role in the process. In addition, the rapid (<11 h) increase in pmoA copy number suggests that methanotrophs present in the water phase quickly colonize Sphagnum. Compared to free-living bacteria, those associated with plants may gain an advantage from a stable CH4 gradient and supply of oxygen from photosynthesis, but it has yet to be demonstrated. In the bathing experiment methanotrophs moved to the mosses even in the dark when oxygen was not formed in photosynthesis, indicating that at the very least oxygen is not the only advantage bacteria gain from the mosses. In another study no CH4 oxidation activity was detected in peat water surrounding Sphagnum mosses (Kip et al., 2010), also indicating that, although present, MOB are not actively oxidizing CH4 in the water phase. Since the only MOB, Methylocystis, detected in the bathing experiment, is non-motile (Dedysh, 2009), it remains open how these MOB cells end up on the moss surface and inside the hyaline cells. On the other hand another peatland inhabiting type II MOB genus, Methylosinus (Dedysh et al., 2003; Chen et al., 2008), does contain motile species (Bowman et al., 1993) and has been isolated from Sphagnum mosses (Kip et al., 2011a).
Similar to previous studies (Tuomivirta et al., 2009; Larmola et al., 2010; Yrjälä et al., 2011) of Finnish peatlands, DGGE of our bathing experiment samples from Sallie’s Fen, located in NH, USA, detected only Methylocystis-like MOB. These were also the dominant methanotrophs in our transplantation experiment, run on the Finnish Lakkasuo raised bog complex, when the pmoA-microarray was used. Dominance of type II MOB in our samples is in line with previous studies. Especially the high prevalence of Methylocystis in Sphagnum samples is not surprising as it is commonly found in northern peatlands (McDonald et al., 1996; Morris et al., 2002; Jaatinen et al., 2005; Dedysh et al., 2006; Chen et al., 2008). Although Kip et al. (2010) found, in contrast to our results, a high diversity of type I methanotrophs, Methylocystis was still the dominant species in their globally gathered Sphagnum samples (Kip et al., 2010) and also very abundant in mosses from a Dutch peat bog (Kip et al., 2011b).
We have demonstrated that water serves as an essential route for methanotroph dispersal and is thus an imperative part of Sphagnum–methanotroph association. This is likely to act as a backup mechanism for peatland CH4 dynamics. Drainage of peatlands can alter the methanotroph community composition (Jaatinen et al., 2005; Yrjälä et al., 2011) and reduce Sphagnum coverage (Yrjälä et al., 2011), consequently compromising this mutualistic association. A case study (Yrjälä et al., 2011) found that the particular Methylocystis sp., which was found now also in the mosses of Sallie’s Fen (northeastern USA) of our bathing experiment and in the mosses of Lakkasuo (Larmola et al. (2010)), was lost when the WT dropped by 14 cm, which is similar to the predicted drawdown for northern peatlands in the global warming scenario by 3°C (Roulet et al., 1992). Restoration of drained peatlands aims to reactivate ecosystem function and restart methanogenesis (Tuittila et al., 2000). Any peatland restoration program should also aim to re-establish the conditions for the mutualistic association between methanotrophs and Sphagnum. Our study indicates that this could be done via transplantations of Sphagnum from donor sites with undisturbed CH4 dynamics. In natural environments Sphagnum associated methanotrophic communities may reduce the methane flux by as much as 80% (Kip et al., 2010). It is not yet known whether this phenomenon can reach that scale also in compromised ecosystems.
Conclusion
Here we showed, by two complementing experiments, that invasion of new MOB through water occurs and that it can be an important mechanism in the reactivation of CH4 oxidation in Sphagnum mosses. Based on this result, the relationship between Sphagnum and methanotrophs is a loose, mutually beneficial association, although some methanotrophs may have an even tighter connection to the mosses.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank T. Ronkainen and L. Maanavilja for field and laboratory assistance, R. Varner, J. Bubier, A. Saari, and P. J. Martikainen for facilities, P. Crill, S. Whitlow, J. Digg, and N. Blake for access to Sallie’s Fen, M. Hardman for checking the language, and S. Elomaa for the graphics. This work was mainly funded by Maj and Tor Nessling Foundation and the Academy of Finland (Project 121535). Additional funding was received from Maa- ja vesitekniikan tuki Foundation, Kone Foundation, and the Academy of Finland (Projects 118493 and 218101).
References
Abell, G. C. J., Stralis-Pavese, N., Sessitsch, A., and Bodrossy, L. (2009). Grazing affects methanotroph activity and diversity in an alpine meadow soil. Environ. Microbiol. Rep. 1, 457–465.
Baani, M., and Liesack, W. (2008). Two isozymes of particulate methane monooxygenase with different methane oxidation kinetics are found in Methylocystis sp. strain SC2. Proc. Natl. Acad. Sci. U.S.A. 105, 10203–10208.
Basiliko, N., Knowles, R., and Moore, T. R. (2004). Roles of moss species and habitat in methane consumption potential in a northern peatland. Wetlands 24, 178–185.
Bodrossy, L., Stralis-Pavese, N., Murrell, J. C., Radajewski, S., Weilharter, A., and Sessitsch, A. (2003). Development and validation of a diagnostic microbial microarray for methanotrophs. Environ. Microbiol. 5, 566–582.
Bowman, J. P., Sly, L. I., Nichols, P. D., and Hayward, A. C. (1993). Revised taxonomy of the methanotrophs: description of Methylobacter gen. nov., emendation of Methylococcus, validation of Methylosinus and Methylocystis species, and a proposal that the family Methylococcaceae includes only the group I methanotrophs. Int. J. Syst. Bacteriol. 43, 735–753.
Bragina, A., Berg, C., Cardinale, M., Shcherbakov, A., Chebotar, V., and Berg, G. (in press). Sphagnum mosses harbour highly specific bacterial diversity during their whole lifecycle. ISME J. doi: 10.1038/ismej.2011.151
Chen, Y., Dumont, M. G., McNamara, N. P., Chamberlain, P. M., Bodrossy, L., Stralis-Pavese, N., and Murrell, J. C. (2008). Diversity of the active methanotrophic community in acidic peatlands as assessed by mRNA and SIP-PLFA analyses. Environ. Microbiol. 10, 446–459.
Clymo, R. S., and Hayward, P. M. (1982). “The ecology of Sphagnum,” in Bryophyte Ecology, ed. A. J. E. Smith (London: Chapman and Hall), 229–289.
Dedysh, S. N. (2009). Exploring methanotroph diversity in acidic northern wetlands: molecular and cultivation-based studies. Microbiology 78, 655–669.
Dedysh, S. N., Belova, S. E., Bodelier, P. L. E., Smirnova, K. V., Khmelenina, V. N., Chidthaisong, A., Trotsenko, Y. A., Liesack, W., and Dunfield, P. F. (2007). Methylocystis heyeri sp. nov., a novel type II methanotrophic bacterium possessing ‘signature’ fatty acids of type I methanotrophs. Int. J. Syst. Evol. Microbiol. 57, 472–479.
Dedysh, S. N., Dunfield, P. F., Derakshani, M., Stubner, S., Heyer, J., and Liesack, W. (2003). Differential detection of type II methanotrophic bacteria in acidic peatlands using newly developed 16S rRNA-targeted fluorescent oligonucleotide probes. FEMS Microbiol. Ecol. 43, 299–308.
Dedysh, S. N., Pankratov, T. A., Belova, S. E., Kulichevskaya, I. S., and Liesack, W. (2006). Phylogenetic analysis and in situ identification of bacteria community composition in an acidic Sphagnum peat bog. Appl. Environ. Microbiol. 72, 2110–2117.
Gorham, E. (1991). Northern peatlands: role in the carbon cycle and probable responses to climatic warming. Ecol. Appl. 1, 182–195.
Jaatinen, K., Tuittila, E. S., Laine, J., Yrjälä, K., and Fritze, H. (2005). Methane-oxidizing bacteria in a Finnish raised mire complex: effects of site fertility and drainage. Microb. Ecol. 50, 429–439.
Kip, N., Ouyang, W., van Winden, J., Raghoebarsing, A., van Niftrik, L., Pol, A., Pan, Y., Bodrossy, L., van Donselaar, E. G., Reichart, G.-J., Jetten, M. S. M., Sinninghe Damsté, J. S., and Op den Camp, H. J. M. (2011a). Detection, isolation and characterization of acidophilic methanotrophs from Sphagnum mosses. Appl. Environ. Microbiol. 77, 5643–5654.
Kip, N., Dutilh, B. E., Pan, Y., Bodrossy, L., Neveling, K., Kwint, M. P., Jetten, M. S. M., and Op den Camp, H. J. M. (2011b). Ultra-deep pyrosequencing of pmoA amplicons confirms the prevalence of Methylomonas and Methylocystis in Sphagnum mosses from a Dutch peat bog. Environ. Microbiol. Rep. 3, 667–673.
Kip, N., van Winden, J., Pan, Y., Bodrossy, L., Reichart, G.-J., Smolders, A. J. P., Jetten, M. S. M., Sinninghe Damsté, J. S., and Op den Camp, H. J. M. (2010). Global prevalence of methane oxidation by symbiotic bacteria in peat-moss ecosystems. Nat. Geosci. 3, 617–621.
Larmola, T., Tuittila, E.-S., Tiirola, M., Nykänen, H., Martikainen, P. J., Yrjälä, K., Tuomivirta, T., and Fritze, H. (2010). The role of Sphagnum mosses in the methane cycling of a boreal mire. Ecology 91, 2356–2365.
McDonald, I. R., Hall, G. H., Pickup, R. W., and Murrell, C. J. (1996). Methane oxidation potential and preliminary analysis of methanotrophs in blanket bog peat using molecular ecology techniques. FEMS Microbiol. Ecol. 21, 197–211.
Morris, S. A., Radajewski, S., Willison, T. W., and Murrell, J. C. (2002). Identification of the functionally active methanotroph population in a peat soil microcosm by stable-isotope probing. Appl. Environ. Microbiol. 68, 1446–1453.
Nykänen, H., Alm, J., Silvola, J., Tolonen, K., and Martikainen, P. (1998). Methane fluxes on boreal peatlands of different fertility and the effect of long-term experimental lowering of the water table on flux rates. Global Biogeochem. Cycles 12, 53–69.
Raghoebarsing, A. A., Smolders, A. J. P., Schmid, M. C., Rijpstra, W. I. C., Wolters-Arts, M., Derksen, J., Jetten, M. S. M., Schouten, S., Sinninghe Damsté, J. S., Lamers, L. P. M., Roelofs, J. G. M., Op den Camp, H. J. M., and Strous, M. (2005). Methanotrophic symbionts provide carbon for photosynthesis in peat bogs. Nature 436, 1153–1156.
Roulet, N., Moore, T., Bubier, J., and Lafleur, P. (1992). Northern fens: methane flux and climatic change. Tellus B Chem. Phys. Meteorol. 44, 100–105.
Siljanen, H. M. P., Saari, A., Krause, S., Lensu, A., Abell, G. C., Bodrossy, L., Bodelier, P. L. E., and Martikainen, P. J. (2011). Hydrology is reflected in the functioning and community composition of methanotrophs in the littoral wetland of a boreal lake. FEMS Microbiol. Ecol. 75, 430–445.
Stralis-Pavese, N., Sessitsch, A., Weilharter, A., Reichenauer, T., Riesing, J., Csontos, J., Murrell, J. C., and Bodrossy, L. (2004). Optimization of diagnostic microarray for application in analysing landfill methanotroph communities under different plant covers. Environ. Microbiol. 6, 347–363.
ter Braak, C. J. F., and Smilauer, P. (2002). CANOCO Reference Manual and CanoDraw for Windows User’s Guide: Software for Canonical Community Ordination. Version 4.5. Ithaca, NY: Microcomputer Power.
Tuittila, E.-S., Komulainen, V.-M., Vasander, H., Nykänen, H., Martikainen, P. J., and Laine, J. (2000). Methane dynamics of a restored cut-away peatland. Glob. Chang. Biol. 6, 569–581.
Tuomivirta, T. T., Yrjälä, K., and Fritze, H. (2009). Quantitative PCR of pmoA using a novel reverse primer correlates with potential methane oxidation in finnish fen. Res. Microbiol. 160, 751–756.
Vorobev, A. V., Baani, M., Doronina, N. V., Brady, A. L., Liesack, W., Dunfield, P. F., and Dedysh, S. N. (2011). Methyloferula stellata gen. nov., sp. nov., an acidophilic, obligately methanotrophic bacterium possessing only a soluble methane monooxygenase. Int. J. Syst. Evol. Microbiol. 61, 2456–2463.
Wise, M. G., McArthur, J. V., and Shimkets, L. J. (1999). Methanotroph diversity in landfill soil: isolation of novel type I and type II methanotrophs whose presence was suggested by culture-independent 16S ribosomal DNA analysis. Appl. Environ. Microbiol. 65, 4887–4897.
Yrjälä, K., Tuomivirta, T., Juottonen, H., Putkinen, A., Lappi, K., Tuittila, E.-S., Penttilä, T., Minkkinen, K., Laine, J., Peltoniemi, K., and Fritze, H. (2011). CH4 production and oxidation processes in a boreal fen ecosystem after long-term water table drawdown. Glob. Chang. Biol. 17, 1311–1320.
Appendix
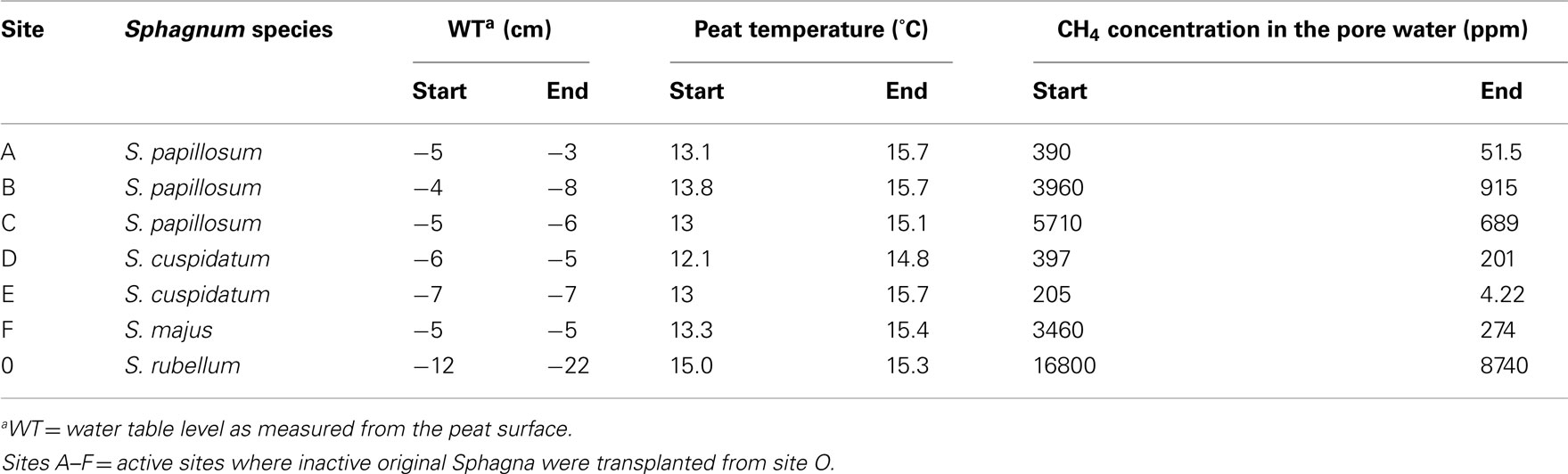
Table A1. Ecological variables of Lakkasuo sites in the beginning (0 day) and the end (28 days) of the transplantation experiment.
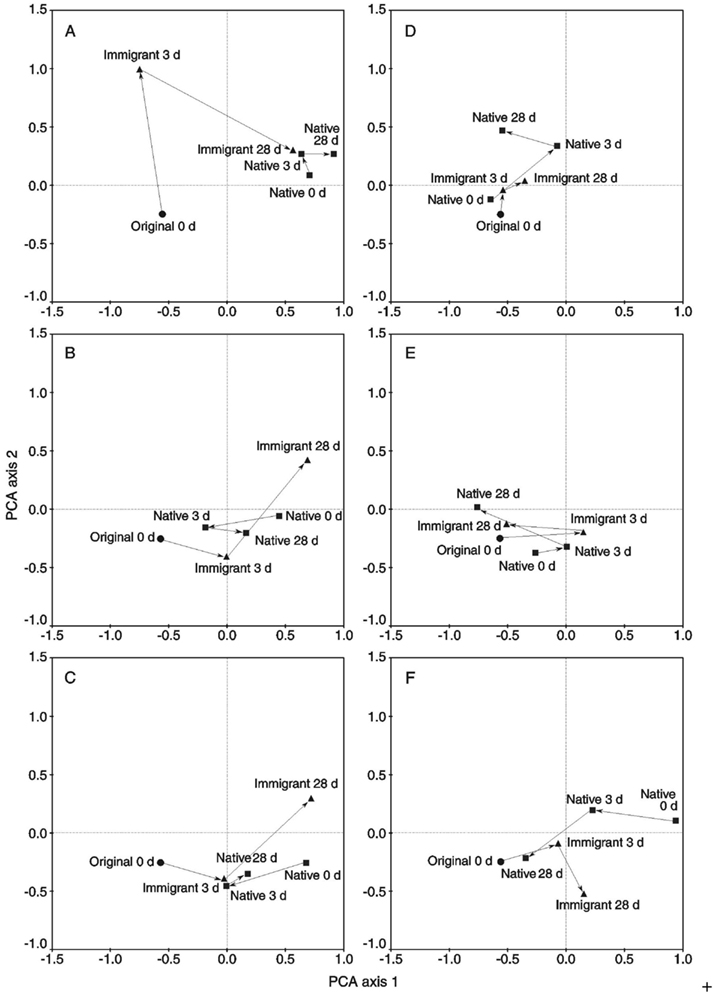
Figure A1. The principal component analysis (PCA) results of the binary microarray data divided into transplantation sites (A–F) (original 0 day = original inactive moss, original 28 days = original moss transplanted back to the original site, immigrant = original moss transplanted to the active flark site, native = native moss of the active flark site). The first and second PCA axes explain 25 and 11% of the compositional variation, respectively.
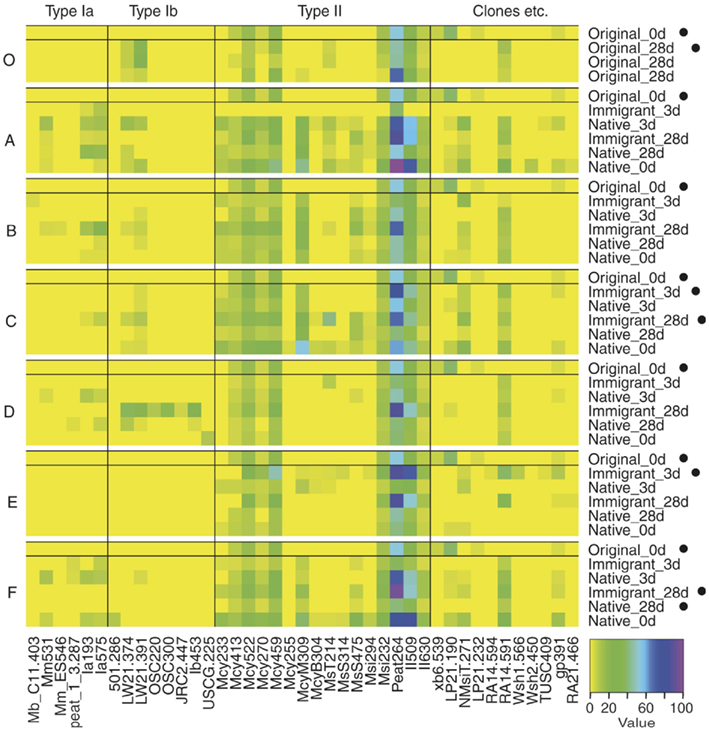
Figure A2. Heatmap of semiquantitative microarray results. Each site (A–F and O) is showed separately together with the original 0 day sample data for comparison. Microarray analysis was successful for only one of three parallel inactive original_0 day samples. Samples that were inactive in CH4 oxidation at the time of sampling are marked with a black dot (•). A value of 100 (see color code bar) indicates maximum signal for an individual probe and zero represents no hybridization. Only probes with a positive hybridization signal on any of the samples are shown.
Keywords: plant–microbe interaction, peatland, pmoA, microarray, qPCR, ecosystem resilience
Citation: Putkinen A, Larmola T, Tuomivirta T, Siljanen HMP, Bodrossy L, Tuittila E-S and Fritze H (2012) Water dispersal of methanotrophic bacteria maintains functional methane oxidation in Sphagnum mosses. Front. Microbio. 3:15. doi: 10.3389/fmicb.2012.00015
Received: 20 December 2011; Accepted: 09 January 2012;
Published online: 23 January 2012.
Edited by:
Svetlana N. Dedysh, Russian Academy of Sciences, RussiaReviewed by:
Svetlana N. Dedysh, Russian Academy of Sciences, RussiaYin Chen, University of Warwick, UK
Copyright: © 2012 Putkinen, Larmola, Tuomivirta, Siljanen, Bodrossy, Tuittila and Fritze. This is an open-access article distributed under the terms of the Creative Commons Attribution Non Commercial License, which permits non-commercial use, distribution, and reproduction in other forums, provided the original authors and source are credited.
*Correspondence: Anuliina Putkinen, Finnish Forest Research Institute, Southern Finland Regional Unit, P.O. Box 18 (Jokiniemenkuja 1), 01301 Vantaa, Finland. e-mail:YW51bGlpbmEucHV0a2luZW5AbWV0bGEuZmk=
†Present address: Tuula Larmola, Mount Holyoke College, South Hadley, MA 01075, USA.


 Tero Tuomivirta1
Tero Tuomivirta1


