- Laboratory of Microbiology and Infectious Diseases, Institute of Scientific and Industrial Research, Osaka University, Ibaraki, Osaka, Japan
Salmonella enterica is an important enteric pathogen, and its various serovars cause both systemic and intestinal diseases in humans and domestic animals. The emergence of multidrug-resistant strains of Salmonella, leading to increased morbidity and mortality, has further complicated its management. Hfq is an RNA chaperon that mediates the binding of small RNAs to mRNA and assists in post-transcriptional gene regulation in bacteria. Although Hfq is related to important phenotypes including virulence in Salmonella, its role in the drug resistance of this organism is unknown. The aim of this study was to investigate the role of Hfq in intrinsic drug resistance of S. enterica serovar Typhimurium. hfq Mutant was susceptible to acriflavine. Although there is a relationship between the production of the AcrB multidrug efflux pump and Hfq in Escherichia coli, the deletion of the drug efflux acrB did not impair the effect of hfq deletion on Salmonella susceptibility. In contrast, the deletion of another drug efflux gene, smvA, impaired the effect of hfq deletion on acriflavine susceptibility. These results indicate that Hfq regulates the intrinsic drug resistance, and it may influence drug susceptibility by regulating SmvA in Salmonella.
Introduction
Salmonella causes a variety of diseases in humans, ranging from gastroenteritis to bacteremia and typhoid fever (Scherer and Miller, 2001). In the 1990s, the prevalence of multidrug-resistant Salmonella enterica increased dramatically in the United Kingdom, the United States, and Canada (Hosek et al., 1997; Threlfall et al., 1997; Glynn et al., 1998; Ng et al., 1999). Many other countries have also documented outbreaks associated with drug-resistant Salmonella in poultry, beef, and pork (Davies et al., 1996; Cody et al., 1999; Grein et al., 1999; Molbak et al., 1999; Villar et al., 1999). Emerging resistance in Salmonella has been observed in both humans and animals, and thus, this is a potentially serious public health problem (Cloeckaert and Chaslus-Dancla, 2001; Piddock, 2002). Drug resistance in bacteria is often associated with multidrug efflux pumps that decrease cellular drug accumulation (Nikaido, 1996; Zgurskaya and Nikaido, 2000).
The phenomenon of multidrug resistance is associated with the ability of pumps to expel from cells multiple drugs with different modes of action. Multidrug resistance is a serious problem in treatment of human ailments caused by pathogenic bacteria, fungi, parasites, and cancer. Functional studies identified multidrug efflux pumps classified in five families: ATP-binding cassette (ABC), the major facilitator (MFS), resistance-nodulation-cell division (RND), small multidrug resistance, and multidrug and toxic compound extrusion families (Brown et al., 1999; Putman et al., 2000; Paulsen et al., 2001). The sequencing of bacterial genomes enables us to trace putative drug resistance genes (Paulsen et al., 1998, 2000). There are many putative and proven drug efflux pumps in the Salmonella genome. We previously demonstrated that S. enterica serovar Typhimurium has nine functional drug efflux pumps (Nishino et al., 2006). In addition to these pumps, it has been reported that SmvA is an important efflux pump for acriflavine and related compounds (Villagra et al., 2008). Because many of these multidrug efflux pumps have overlapping substrate spectra, it is intriguing that bacteria, with their economically organized genomes, harbor such large sets of multidrug efflux genes. The key to understanding how bacteria utilize these multiple pumps lies in the regulation of pump expression. Currently, available data indicate that multidrug efflux pumps are often expressed under precise and elaborate transcriptional control (Ahmed et al., 1994; Lomovskaya et al., 1995; Brooun et al., 1999; Grkovic et al., 2002).
The Hfq protein is a conserved RNA chaperone protein first characterized as a host factor (HF-1) for phage Qβ RNA replication (Franze de Fernandez et al., 1968) and subsequently shown to be widely distributed in the bacterial kingdom with multiple homologs in the annotated genomic database (Brennan and Link, 2007). As a bacterial homolog of the eukaryotic and archaeal Sm/LSm proteins, Hfq is known largely for its global post-transcriptional regulation by binding AU-rich sequences of target mRNA and facilitating pairing between sRNAs and mRNAs (Moller et al., 2002; Zhang et al., 2002; Valentin-Hansen et al., 2004; Waters and Storz, 2009). Most Hfq homologs are known to function as homohexamers with two independent RNA-binding motifs (Brennan and Link, 2007), and hfq mutants exhibit pleiotropic phenotypes (Tsui et al., 1994). In recent years, Hfq has been established as an important virulence factor in bacterial pathogens (Hansen and Kaper, 2009). Deletion of hfq has long been known to impair the expression of σS (Brown and Elliott, 1996), a general stress sigma factor essential for Salmonella virulence in mice (Fang et al., 1992). hfq mutation was also revealed to attenuate the ability of Salmonella to invade epithelial cells, secrete virulence factors, infect mice, and survive inside cultured macrophages (Sittka et al., 2007). Transcriptomic analysis revealed that Hfq controls the expression of Salmonella genes in several horizontally acquired pathogenicity islands (SPI-1, -2, -4, -5), two sigma factor regulons, and the flagellar gene cascade (Sittka et al., 2008). However, the role Hfq in the drug resistance of Salmonella is unknown.
In this study, we demonstrate that Hfq affects drug susceptibilities in Salmonella. In addition, we reveal that SmvA and not the AcrB drug efflux system contributes to the Hfq-mediated drug resistance of Salmonella, whereas it has been reported that AcrB contributes to the Hfq-mediated drug resistance of Escherichia coli. Our data suggest that Hfq plays an important role in controlling drug susceptibility against acriflavine and that the SmvA efflux pump is involved in this susceptibility in Salmonella.
Materials and Methods
Bacterial Strains, Plasmids, and Growth Conditions
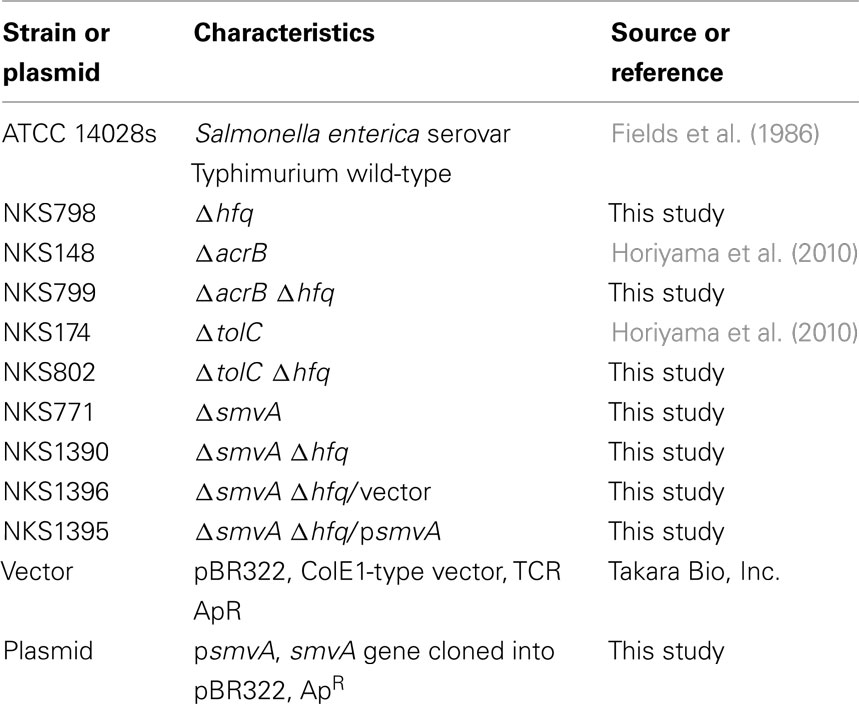
The bacterial strains and plasmids used in this study are listed in Table 1. The strains of S. enterica serovar Typhimurium used in this study were derived from the wild-type strain ATCC 14028s (Fields et al., 1986). Bacterial strains were grown at 37°C in Lysogeny Broth (LB). Ampicillin was added to the growth medium at a final concentration of 100 μg/ml for plasmid maintenance.
Construction of Gene Deletion Mutants
The ΔacrB (NKS148) and ΔtolC (NKS174) mutants were constructed as described previously (Horiyama et al., 2010). To construct Δhfq and ΔsmvA mutants, gene disruption was performed as described by Datsenko and Wanner (2000). The following oligonucleotide primers were used for the construction of the mutants: hfq-P1 (GAAAGGTTCAAAGTACAAATAAGCATATAAGGAAAAGAGAGTGTAGGCTGGAGCTGCTTC); hfq-P2 (ATTATCCGACGCCCCCGACATGGATAAACAGCGCGTGAACCATATGAATATCCTCCTTAG); smvA-P1 (CTGGACAAGCGTCCAAATTTGAGTTTTTGAAGGGAGAGTTGTGTAGGCTGGAGCTGCTTC); and smvA-P2 (CCAGCTAGCGCATTAAGCGCTTATCTCACCAGGCGTTATGCATATGAATATCCTCCTTAG). The chloramphenicol resistance gene cat, flanked by Flp recognition sites, was amplified by PCR using the primers listed above. The resulting PCR products were used to transform the recipient ATCC 14028s strain that harbors the plasmid pKD46, which expresses Red recombinase. The chromosomal structure of the mutated loci was verified by PCR. cat was eliminated using the plasmid pCP20, as described previously (Datsenko and Wanner, 2000). To construct the ΔacrBΔhfq, ΔtolCΔhfq, and ΔsmvAΔhfq double mutants, the deletions were transferred to strains by P22 transduction as described by Davis et al. (1980).
Plasmid Construction
smvA was amplified from ATCC 14028s genomic DNA by using the primers GCGCATGCCATTCGTTCAACTTACCGAGG and GCGTCGACGGAAATGGACTCCCCCTGCC, which introduced SphI and SalI sites (underlined in the primer sequences above). The fragment was cleaved with SphI and SalI and then cloned into the corresponding sites of pBR322, resulting in psmvA (Table 1).
Determination of the Minimum Inhibitory Concentration of Toxic Compounds
The antibacterial activities of various agents were determined on LB agar (1% tryptone, 0.5% yeast extract, 0.5% NaCl) plates containing nalidixic acid, acriflavine, rhodamine 6G, benzalkonium, oxacillin, cefamandole, sodium dodecyl sulfate, or norfloxacin (Sigma, St. Louis, MO, USA) at various concentrations as indicated in Table 2. Agar plates were made by the twofold agar dilution technique, as described previously (Horiyama et al., 2010). To determine minimum inhibitory concentrations (MICs), bacteria were grown in LB broth at 37°C overnight, diluted into the same medium, and then tested at a final inoculum size of 1 × 105 cfu/μl using a multipoint inoculator (Sakuma Seisakusyo, Tokyo, Japan) after incubation at 37°C for 20 h. The MIC was the lowest concentration of a compound that inhibited cell growth.
Results and Discussion
Hfq Affects the Intrinsic Drug Susceptibility of Salmonella
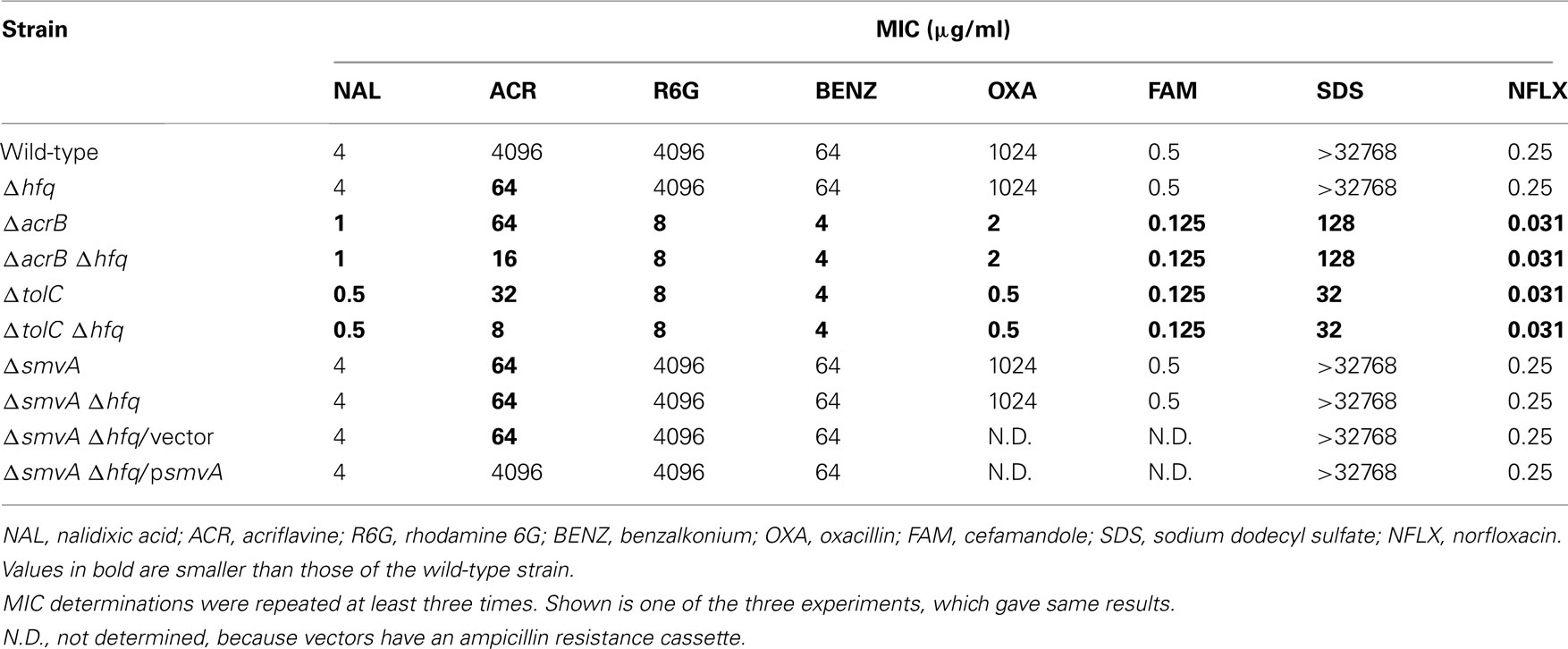
To investigate the role of Hfq in drug susceptibilities, hfq was deleted from S. enterica serovar Typhimurium strain ATCC 14028s, as described in the Section “Materials and Methods.” Δhfq mutant was more sensitive to acriflavine (64-fold) than the wild-type strain (Table 2). The MICs of nalidixic acid, rhodamine 6G, benzalkonium, oxacillin, cefamandole, sodium dodecyl sulfate, and norfloxacin were the same for Δhfq mutant as those for the wild-type strain. These data indicate that Hfq affects the intrinsic acriflavine resistance of Salmonella.
AcrB is not Involved in Hfq-Mediated Drug Susceptibility of Salmonella
In E. coli, it has been demonstrated that the AcrB multidrug efflux pump is involved in Hfq-mediated multidrug resistance (Yamada et al., 2010). To investigate whether AcrB in Salmonella is also involved in Hfq-mediated drug susceptibility of this organism, we measured MICs of several toxic compounds against ΔacrB mutant (Table 2). The ΔacrB mutant was sensitive to nalidixic acid (fourfold), acriflavine (64-fold), rhodamine 6G (512-fold), benzalkonium (16-fold), oxacillin (512-fold), cefamandole (fourfold), sodium dodecyl sulfate (>256-fold), and norfloxacin (eightfold). Although ΔacrB mutant was as sensitive to acriflavine as Δhfq mutant, the drug susceptibility pattern for other compounds was very different among these mutants. ΔacrBΔhfq double mutant was more sensitive to acriflavine (fourfold) than ΔacrB mutant, indicating that the deletion of acrB did not impair the effect of hfq deletion on Salmonella susceptibility. Based on these data, it was suggested that factors other than AcrB may be involved in the Hfq-mediated drug susceptibility of Salmonella because the drug susceptibility pattern of ΔacrB was very different from that of Δhfq, and the deletion of hfq from ΔacrB mutant made this strain more sensitive to acriflavine.
TolC is not Involved in the Hfq-Mediated Drug Susceptibility of Salmonella
TolC is a major outer membrane channel, and a variety of inner membrane and accessory protein interact with TolC to expel structurally diverse molecules. We previously identified that seven drug efflux systems, AcrAB, AcrD, AcrEF, MdsAB, MdtABC, EmrAB, and MacAB, in Salmonella that require TolC to function (Horiyama et al., 2010). To investigate whether TolC-dependent type drug efflux systems are involved in Hfq-mediated drug susceptibility, we measured MICs of compounds against ΔtolC mutant (Table 2). ΔtolC mutant was sensitive to nalidixic acid (eightfold), acriflavine (128-fold), rhodamine 6G (512-fold), benzalkonium (16-fold), oxacillin (2048-fold), cefamandole (fourfold), sodium dodecyl sulfate (>1024-fold), and norfloxacin (eightfold). The susceptibilities of ΔtolC mutant to oxacillin and sodium dodecyl sulfate were higher than those of the ΔacrB mutant probably because TolC-dependent type efflux systems other than AcrB are involved in the efflux of these compounds. The deletion of hfq from ΔtolC mutant made this strain more sensitive to acriflavine (fourfold), meaning that the TolC-dependent type drug efflux systems are not involved in the Hfq-mediated drug susceptibility of Salmonella.
Involvement of SmvA Efflux Pump in the Hfq-Mediated Acriflavine Susceptibility
Among the tested compounds, Δhfq mutant was specifically susceptible to acriflavine (Table 2) as mentioned above. Because it has been reported that SmvA is an important efflux pump for acriflavine (Villagra et al., 2008), we hypothesized that SmvA may be involved in the Hfq-mediated acriflavine susceptibility of Salmonella. Similarly, as Δhfq mutant, ΔsmvA mutant was more sensitive to acriflavine (64-fold) than the wild-type strain (Table 2). This phenotype is in good agreement with a previous report (Villagra et al., 2008). MIC of acriflavine against ΔsmvAΔhfq double mutant was similar to that against ΔsmvA mutant, indicating that deletion of smvA impaired the effect of hfq deletion on acriflavine susceptibility. Moreover, psmvA, which expressed smvA, conferred acriflavine resistance to ΔsmvAΔhfq double mutant. MIC of acriflavine against ΔsmvAΔhfq/psmvA strain is similar to that against the wild-type strain (Table 2). Taken together, these results indicated that Hfq regulates the intrinsic acriflavine resistance of Salmonella and SmvA plays an important role in this resistance because the drug susceptibility pattern of ΔsmvA was same as that of Δhfq, and the deletion of hfq from ΔsmvA mutant did not change the acriflavine susceptibility of this strain.
In this study, we investigated the role of Hfq in the drug susceptibility of S. enterica serovar Typhimurium ATCC 14028s and found that Hfq plays a role in its intrinsic acriflavine resistance and that SmvA efflux pump is involved in this resistance. Interestingly, Δhfq mutant of Salmonella was specifically sensitive to acriflavine among the tested compounds. This phenotype is very different from Δhfq mutant of E. coli W3104 or MC4100 (Yamada et al., 2010). In case of E. coli, Δhfq mutant was susceptible to various compounds including acriflavine, benzalkonium, cefamandole, chloramphenicol, crystal violet, nalidixic acid, novobiocin, oxacillin, and rhodamine 6G because Hfq positively regulates the production of the AcrB drug efflux pump (Yamada et al., 2010). However, AcrB was considered not to be involved in the Hfq-mediated intrinsic acriflavine resistance of Salmonella. These observations suggest the differential regulation of genes by Hfq between E. coli and Salmonella. Indeed, transcriptomic analysis revealed that Hfq controls the Salmonella gene expression in several horizontally acquired pathogenicity islands (SPI-1, -2, -4, -5) that are not present in E. coli (Sittka et al., 2008). Unlike the AcrAB drug efflux system, which is widely distributed throughout all Enterobacteriaceae, homologs of SmvA are not found in E. coli and Shigella spp. Villagra et al. (2008) suggested that acriflavine is a substrate for both AcrB and SmvA efflux pumps, but SmvA pump plays the major role in the efflux of acriflavine in Salmonella. This may explain why SmvA and not AcrB drug efflux system contributes to the Hfq-mediated drug resistance of Salmonella.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Eiji Nikaido for technical help in constructing the plasmids. This study was supported by Grants-in-Aid for Young Scientists from the Japan Society for the Promotion of Science (to Mitsuko Hayashi-Nishino), the Uehara Memorial Foundation (to Kunihiko Nishino) and the Funding Program for Next Generation World-Leading Researchers from the Cabinet Office, Government of Japan (to Kunihiko Nishino).
References
Ahmed, M., Borsch, C. M., Taylor, S. S., Vazquez-Laslop, N., and Neyfakh, A. A. (1994). A protein that activates expression of a multidrug efflux transporter upon binding the transporter substrates. J. Biol. Chem. 269, 28506–28513.
Brennan, R. G., and Link, T. M. (2007). Hfq structure, function and ligand binding. Curr. Opin. Microbiol. 10, 125–133.
Brooun, A., Tomashek, J. J., and Lewis, K. (1999). Purification and ligand binding of EmrR, a regulator of a multidrug transporter. J. Bacteriol. 181, 5131–5133.
Brown, L., and Elliott, T. (1996). Efficient translation of the RpoS sigma factor in Salmonella typhimurium requires host factor I, an RNA-binding protein encoded by the hfq gene. J. Bacteriol. 178, 3763–3770.
Brown, M. H., Paulsen, I. T., and Skurray, R. A. (1999). The multidrug efflux protein NorM is a prototype of a new family of transporters. Mol. Microbiol. 31, 394–395.
Cloeckaert, A., and Chaslus-Dancla, E. (2001). Mechanisms of quinolone resistance in Salmonella. Vet. Res. 32, 291–300.
Cody, S. H., Abbott, S. L., Marfin, A. A., Schulz, B., Wagner, P., Robbins, K., Mohle-Boetani, J. C., and Vugia, D. J. (1999). Two outbreaks of multidrug-resistant Salmonella serotype typhimurium DT104 infections linked to raw-milk cheese in Northern California. JAMA 281, 1805–1810.
Datsenko, K. A., and Wanner, B. L. (2000). One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. U.S.A. 97, 6640–6645.
Davies, A., O’Neill, P., Towers, L., and Cooke, M. (1996). An outbreak of Salmonella typhimurium DT104 food poisoning associated with eating beef. Commun. Dis. Rep. CDR Rev. 6, R159–R162.
Davis, R. W., Bolstein, D., and Roth, J. R. (1980). Advanced Bacterial Genetics. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press.
Fang, F. C., Libby, S. J., Buchmeier, N. A., Loewen, P. C., Switala, J., Harwood, J., and Guiney, D. G. (1992). The alternative sigma factor katF (rpoS) regulates Salmonella virulence. Proc. Natl. Acad. Sci. U.S.A. 89, 11978–11982.
Fields, P. I., Swanson, R. V., Haidaris, C. G., and Heffron, F. (1986). Mutants of Salmonella typhimurium that cannot survive within the macrophage are avirulent. Proc. Natl. Acad. Sci. U.S.A. 83, 5189–5193.
Franze de Fernandez, M. T., Eoyang, L., and August, J. T. (1968). Factor fraction required for the synthesis of bacteriophage Qbeta-RNA. Nature 219, 588–590.
Glynn, M. K., Bopp, C., Dewitt, W., Dabney, P., Mokhtar, M., and Angulo, F. J. (1998). Emergence of multidrug-resistant Salmonella enterica serotype typhimurium DT104 infections in the United States. N. Engl. J. Med. 338, 1333–1338.
Grein, T., O’Flanagan, D., McCarthy, T., and Bauer, D. (1999). An outbreak of multidrug-resistant Salmonella typhimurium food poisoning at a wedding reception. Ir. Med. J. 92, 238–241.
Grkovic, S., Brown, M. H., and Skurray, R. A. (2002). Regulation of bacterial drug export systems. Microbiol. Mol. Biol. Rev. 66, 671–701.
Hansen, A. M., and Kaper, J. B. (2009). Hfq affects the expression of the LEE pathogenicity island in enterohaemorrhagic Escherichia coli. Mol. Microbiol. 73, 446–465.
Horiyama, T., Yamaguchi, A., and Nishino, K. (2010). TolC dependency of multidrug efflux systems in Salmonella enterica serovar Typhimurium. J. Antimicrob. Chemother. 65, 1372–1376.
Hosek, G., Leschinsky, D. D., Irons, S., and Safranek, T. J. (1997). Multidrug resistant Salmonella serotype Typhimurium-United States, 1996. MMWR Morb. Mortal. Wkly. Rep. 46, 308–310.
Lomovskaya, O., Lewis, K., and Matin, A. (1995). EmrR is a negative regulator of the Escherichia coli multidrug resistance pump EmrAB. J. Bacteriol. 177, 2328–2334.
Molbak, K., Baggesen, D. L., Aarestrup, F. M., Ebbesen, J. M., Engberg, J., Frydendahl, K., Gerner-Smidt, P., Petersen, A. M., and Wegener, H. C. (1999). An outbreak of multidrug-resistant, quinolone-resistant Salmonella enterica serotype typhimurium DT104. N. Engl. J. Med. 341, 1420–1425.
Moller, T., Franch, T., Hojrup, P., Keene, D. R., Bachinger, H. P., Brennan, R. G., and Valentin-Hansen, P. (2002). Hfq: a bacterial Sm-like protein that mediates RNA-RNA interaction. Mol. Cell 9, 23–30.
Ng, L. K., Mulvey, M. R., Martin, I., Peters, G. A., and Johnson, W. (1999). Genetic characterization of antimicrobial resistance in Canadian isolates of Salmonella serovar Typhimurium DT104. Antimicrob. Agents Chemother. 43, 3018–3021.
Nishino, K., Latifi, T., and Groisman, E. A. (2006). Virulence and drug resistance roles of multidrug efflux systems of Salmonella enterica serovar Typhimurium. Mol. Microbiol. 59, 126–141.
Paulsen, I. T., Chen, J., Nelson, K. E., and Saier, M. H. Jr. (2001). Comparative genomics of microbial drug efflux systems. J. Mol. Microbiol. Biotechnol. 3, 145–150.
Paulsen, I. T., Nguyen, L., Sliwinski, M. K., Rabus, R., and Saier, M. H. Jr. (2000). Microbial genome analyses: comparative transport capabilities in eighteen prokaryotes. J. Mol. Biol. 301, 75–100.
Paulsen, I. T., Sliwinski, M. K., and Saier, M. H. Jr. (1998). Microbial genome analyses: global comparisons of transport capabilities based on phylogenies, bioenergetics and substrate specificities. J. Mol. Biol. 277, 573–592.
Piddock, L. J. (2002). Fluoroquinolone resistance in Salmonella serovars isolated from humans and food animals. FEMS Microbiol. Rev. 26, 3–16.
Putman, M., Van Veen, H. W., and Konings, W. N. (2000). Molecular properties of bacterial multidrug transporters. Microbiol. Mol. Biol. Rev. 64, 672–693.
Scherer, C. A., and Miller, S. I. (2001). “Molecular pathogenesis of Salmonella,” in Principles of Bacterial Pathogenesis, ed. E. A. Groisman (New York: Academic Press), 266–333.
Sittka, A., Lucchini, S., Papenfort, K., Sharma, C. M., Rolle, K., Binnewies, T. T., Hinton, J. C., and Vogel, J. (2008). Deep sequencing analysis of small noncoding RNA and mRNA targets of the global post-transcriptional regulator, Hfq. PLoS Genet. 4, e1000163. doi:10.1371/journal.pgen.1000163
Sittka, A., Pfeiffer, V., Tedin, K., and Vogel, J. (2007). The RNA chaperone Hfq is essential for the virulence of Salmonella typhimurium. Mol. Microbiol. 63, 193–217.
Threlfall, E. J., Ward, L. R., Skinner, J. A., and Rowe, B. (1997). Increase in multiple antibiotic resistance in nontyphoidal salmonellas from humans in England and Wales: a comparison of data for 1994 and 1996. Microb. Drug Resist. 3, 263–266.
Tsui, H. C., Leung, H. C., and Winkler, M. E. (1994). Characterization of broadly pleiotropic phenotypes caused by an hfq insertion mutation in Escherichia coli K-12. Mol. Microbiol. 13, 35–49.
Valentin-Hansen, P., Eriksen, M., and Udesen, C. (2004). The bacterial Sm-like protein Hfq: a key player in RNA transactions. Mol. Microbiol. 51, 1525–1533.
Villagra, N. A., Hidalgo, A. A., Santiviago, C. A., Saavedra, C. P., and Mora, G. C. (2008). SmvA, and not AcrB, is the major efflux pump for acriflavine and related compounds in Salmonella enterica serovar Typhimurium. J. Antimicrob. Chemother. 62, 1273–1276.
Villar, R. G., Macek, M. D., Simons, S., Hayes, P. S., Goldoft, M. J., Lewis, J. H., Rowan, L. L., Hursh, D., Patnode, M., and Mead, P. S. (1999). Investigation of multidrug-resistant Salmonella serotype typhimurium DT104 infections linked to raw-milk cheese in Washington State. JAMA 281, 1811–1816.
Yamada, J., Yamasaki, S., Hirakawa, H., Hayashi-Nishino, M., Yamaguchi, A., and Nishino, K. (2010). Impact of the RNA chaperone Hfq on multidrug resistance in Escherichia coli. J. Antimicrob. Chemother. 65, 853–858.
Zgurskaya, H. I., and Nikaido, H. (2000). Multidrug resistance mechanisms: drug efflux across two membranes. Mol. Microbiol. 37, 219–225.
Keywords: drug efflux system, drug resistance, Hfq, Salmonella, small RNA
Citation: Hayashi-Nishino M, Fukushima A and Nishino K (2012) Impact of Hfq on the intrinsic drug resistance of Salmonella enterica serovar Typhimurium. Front. Microbio. 3:205. doi: 10.3389/fmicb.2012.00205
Received: 16 April 2012; Accepted: 18 May 2012
Published online: 04 June 2012.
Edited by:
Axel Cloeckaert, Institut National de la Recherche Agronomique, FranceReviewed by:
Fiona Walsh, Agroscope Changins Wädenswil, SwitzerlandCharles Knapp, University of Strathclyde, UK
Copyright: © 2012 Hayashi-Nishino, Fukushima and Nishino. This is an openaccess article distributed under the terms of the Creative Commons Attribution Non Commercial License, which permits non-commercial use, distribution, and reproduction in other forums, provided the original authors and source are credited.
*Correspondence: Kunihiko Nishino, Laboratory of Microbiology and Infectious Diseases, Institute of Scientific and Industrial Research, Osaka University, 8-1 Mihogaoka, Ibaraki, Osaka 567-0047, Japan. e-mail:bmlzaGlub0BzYW5rZW4ub3Nha2EtdS5hYy5qcA==
†Mitsuko Hayashi-Nishino and Aiko Fukushima have contributed equally to this work.
 Mitsuko Hayashi-Nishino†
Mitsuko Hayashi-Nishino†