- 1Department of International Health, Johns Hopkins Bloomberg School of Public Health, Baltimore, MD, USA
- 2International Centre for Diarrhoeal Disease Research, Dhaka, Bangladesh
The presence of Vibrio cholerae in the environment is key to understanding the epidemiology of cholera. The gold standard for laboratory confirmation of V. cholerae from water is a culture method, but this requires laboratory infrastructure. A rapid diagnostic test that is simple, inexpensive, and can be deployed widely would be useful for confirming V. cholerae in samples of environmental water. Here, we evaluated a dipstick test to detect V. cholerae O1 and O139 from environmental water samples in spiked samples and under field conditions. When environmental water samples were incubated in alkaline peptone water for 24 h at room temperature, samples spiked with <10 CFU could be detected using the dipstick test. When compared to culture, the test was 89% sensitive and 100% specific with environmental samples.
Introduction
Cholera is transmitted through water contaminated with Vibrio cholerae. Methods for identifying culturable V. cholerae from water and sewage include culture and PCR techniques (Barrett et al., 1980; Fields et al., 1992; Alam et al., 2010), but efforts for detecting V. cholerae in environmental samples have previously been limited by laboratory testing capacity in areas where cholera most often occurs and the difficulty of achieving reliable results in these areas.
Recently a dipstick rapid test, based on detection of LPS antigen was introduced for detecting V. cholerae directly from fecal samples (Crystal VC kit, Span Diagnostics Ltd., Udhna, Surat, India). When watery fecal samples were tested directly, using culture method as gold standard the test was found to be sensitive (>90%) and moderately specific (~70%; Ingram et al., 1996; Rabbani et al., 2001; Bhuiyan et al., 2003; Nato et al., 2003; Wang et al., 2006; Harris et al., 2009; Mukherjee et al., 2010; Rosewell et al., 2012; Sinha et al., 2012; Boncy et al., 2013). The test can also be carried out with rectal swab specimens if these samples are first incubated in alkaline peptone water (APW) for 4 to 6 h, a vibrio-selective enrichment step, prior to testing with the dipstick (Bhuiyan et al., 2003). We felt that this dipstick test might be adapted for use in detecting V. cholerae O1 and O139 from environmental water samples in areas at risk for cholera if the specimen was incubated in APW first in order to amplify the concentration of vibrios in the specimen.
Materials and Methods
First, we carried out dipstick experiments with water samples from the Chesapeake Bay that were spiked with 10-fold serial dilutions of V. cholerae serotype O1 or O139. We placed 1 mL of the spiked water samples into 9 mL of APW, followed by incubation at temperatures of 22, 30, and 37°C (Table 1). The concentration of bacteria added to spike the water was determined by plating serial dilutions of the initial inoculum. We tested the APW after 2, 4, 6 and 24 h with the dipstick using methods as described by the manufacturer for fecal samples.
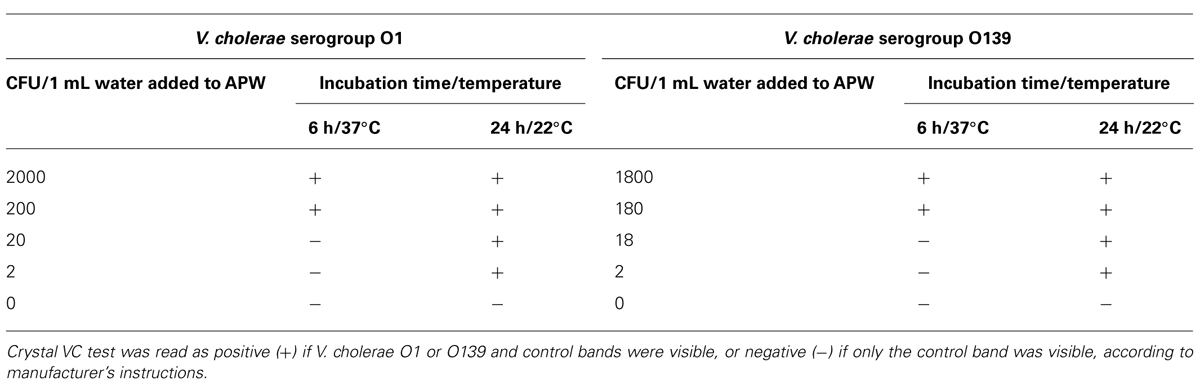
TABLE 1. Dipstick detection of Vibrio cholerae-containing water samples incubated in alkaline peptone water (APW).
Next, we applied the dipstick testing method to environmental water samples from Bangladesh. The samples included specimens from both urban and rural sites that were being collected as part of cholera epidemiology studies at the International Centre for Diarrheal Disease Research, Bangladesh (icddr,b) during March 2011–June 2012. Use of these samples allowed for a direct comparison of the dipstick testing method to bacterial culture methods used at icddr,b.
A total of 550 environmental water samples (200 mL each) were filtered through individual 0.22 micron polycarbonate membrane filters. The filters were then placed in 10 mL of phosphate buffered saline (PBS). After resuspension, 2 mL of the PBS solution was added to 18 mL of APW, followed by incubation for 24 h at room temperature (~22°C). The APW broth was then tested using the dipstick. The same APW was streaked onto taurocholate tellurite gelatin agar (TTGA) and thiosulfate-citrate-bile salts-sucrose (TCBS) plates, and colonies suspected as being V. cholerae from either plate were identified as V. cholerae O1 or O139 using standard methods. The results from the dipstick and cultures were recorded without knowledge of results from the other test.
Results
All spiked specimens were negative at 2 and 4 h, and only the APW spiked with greater than ~200 CFU were positive after 6 h of incubation at 37°C. However, after 24 h, all specimens were positive, including those spiked with <10 CFU and incubated at 30°C or at room temperature (22°C). The APW inoculated with unspiked water was consistently negative at all time points and temperatures (Table 1).
A positive dipstick reading occurred when the concentration of V. cholerae exceeded ~107 per mL; thus, the small number of bacteria in the water that were added to the APW had to grow to this density to be detectable.
Among the 550 environmental samples tested in Bangladesh, 55 were positive for V. cholerae serotype O1 by the bacterial culture method, and 48 of these were also positive with the dipstick test (89% sensitivity). Of the 495 samples that were negative by culture, none were positive with the dipstick test (100% specificity). None of the samples were positive for V. cholerae O139 by dipstick or culture.
Discussion
This dipstick method following incubation in APW provides a simple procedure for detecting V. cholerae O1 and O139 from environmental water. Although the bacterial culture method did detect more positive specimens than the dipstick test, the dipstick test was still very sensitive. It should be noted that the culture methods used at the icddr,b included streaking onto both TCBS and TTGA agars. Incubation on these two media will generally be more sensitive than methods which use only one type of agar, as is carried out in most laboratories (Alam et al., 2010). Thus, the sensitivity of the dipstick method may be as sensitive as using culture with TCBS alone.
While our preliminary study used samples spiked with O139 as well as O1 serotypes, there were no O139 strains detected with the environmental samples from Bangladesh. It seems plausible that this method can be used for the detection of serotype O139 from environmental water, but this has not yet been validated in the field. Recently, there have been no clinical cases of cholera nor environmental isolations related to V. cholerae O139 in Bangladesh. The finding that there were no positive dipstick tests for O139 was reassuring.
The high specificity of the dipstick test (100%) in this evaluation differs from the results observed in clinical studies of fecal samples in which many dip stick positive tests could not be confirmed by culture suggesting a high rate of false positives (Rabbani et al., 2001; Harris et al., 2009; Rosewell et al., 2012). We believe that this difference is because we carried out the test on water samples after incubation in APW rather than directly testing as done in fecal specimens. The reason that some stool samples yield false positive results is not known, but by using an APW selective enrichment step, cross reacting substances are diluted and vibrio antigens are amplified. Our results confirm that APW enrichment of samples helps to improve the specificity of the test, as previously demonstrated (Wang et al., 2006). For water samples, incubation in APW allows growth of vibrio species to concentrations detectable with the dipstick.
This study used the Crystal VC dipstick to detect V cholerae as this has been the test most widely used in developing country settings. Other rapid tests for V. cholerae are also available (Qadri et al., 1995) and might also be useful since this method primarily relies on the ability of vibrios to grow and for the LPS signal to be amplified in the APW broth prior to testing.
Some limitations to this dipstick method should be mentioned. V. cholerae can exist as viable but non-culturable (VBNC) bacteria (Alam et al., 2007). The dipstick test relies on the ability of the bacteria to grow in the APW medium; thus, it is not able to detect VBNC vibrios. Secondly, the Crystal VC dipstick test detects the LPS of V. cholerae but does not differentiate toxigenic from non-toxigenic strains. Non-toxigenic strains of serogroups O1 and O139 may also occur in environmental water but do not carry the same epidemic implications (Faruque et al., 2004). Thus, it would seem that some positive samples should be validated with follow up cultures to determine if they are toxigenic and to more fully characterize the strains. However, it should be noted that standard culture methods also do not determine if the vibrio strains isolated produce toxin. Depending on the epidemiological circumstances, the finding of V. cholerae O1 and O139 in water samples should stimulate a public health response and it is imperative that the finding of a positive dip stick test is rapidly confirmed.
Another limitation of this method, as it was carried out in Bangladesh, included filtration through a 0.22 micron filter. For some cholera endemic areas, this method of filtration may not be possible. Thus, we are evaluating a less complex filtration method through cotton gauze, similar to the method used by Spira (Spira and Ahmed, 1981). Though filtration through gauze is likely to be somewhat less sensitive, it may still be useful in monitoring water samples. In fact the sensitivity of any assay also depends on the volume of water assayed, and no single test will be 100% sensitive.
We conclude that this dipstick method for testing environmental water samples is sensitive and specific for V. cholerae O1 and should be useful for water monitoring in cholera-endemic settings and in areas at risk for cholera. The method is technically simple and does not require use of an incubator. It is not “rapid” since an overnight incubation step is necessary but is “rapid” compared to traditional culture method.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
Support for the research was provided by National Institutes of Health grant no. R01AI039129-13 and by a grant from the Bill and Melinda Gates Foundation to Johns Hopkins University for the DOVE project (Delivering Oral Vaccine Effectively). In addition, the International Centre for Diarrhoeal Disease Research, Bangladesh acknowledges its donors for providing unrestricted support including the, governments of Bangladesh, Canada, the United Kingdom, Sweden, and Australia. Span Diagnostics, Ltd, Udhna, Surat, India donated the test kits.
References
Alam, M., Hasan, N. A., Sultana, M., Nair, G. B., Sadique, A., Faruque, A. S., et al. (2010). Diagnostic limitations to accurate diagnosis of cholera. J. Clin. Microbiol. 48, 3918–3922. doi: 10.1128/JCM.00616-10
Alam, M., Sultana, M., Nair, G. B., Siddique, A. K., Hasan, N. A., Sack, R. B., et al. (2007). Viable but nonculturable Vibrio cholerae O1 in biofilms in the aquatic environment and their role in cholera transmission. Proc. Natl. Acad. Sci. U.S.A. 104, 17801–17806. doi: 10.1073/pnas.0705599104
Barrett, T. J., Blake, P. A., Morris, G. K., Puhr, N. D., Bradford, H. B., and Wells, J. G. (1980). Use of Moore swabs for isolating Vibrio cholerae from sewage. J. Clin. Microbiol. 11, 11385–11388.
Bhuiyan, N. A., Qadri, F., Faruque, A. S., Malek, M. A., Salam, M. A., Nato, F., et al. (2003). Use of dipsticks for rapid diagnosis of cholera caused by Vibrio cholerae O1 and O139 from rectal swabs. J. Clin. Microbiol. 41, 3939–3941. doi: 10.1128/JCM.41.8.3939-3941.2003
Boncy, J., Rossignol, E., Dahourou, G., Hast, M., Buteau, J., Stanislas, M., et al. (2013). Performance and utility of a rapid diagnostic test for cholera: notes from Haiti. Diagn. Microbiol. Infect. Dis. 76, 521–523. doi: 10.1016/j.diagmicrobio.2013.03.010
Faruque, S. M., Chowdhury, N., Kamruzzaman, M., Dziejman, M., Rahman, M. H., Sack, D. A., et al. (2004). Genetic diversity and virulence potential of environmental Vibrio cholerae population in a cholera-endemic area. Proc. Natl. Acad. Sci. U.S.A. 101, 2123–2128. doi: 10.1073/pnas.0308485100
Fields, P. I., Popovic, T., Wachsmuth, K., and Olsvik, O. (1992). Use of polymerase chain reaction for detection of toxigenic Vibrio cholerae O1 strains from the Latin American cholera epidemic. J. Clin. Microbiol. 30, 2118–2121.
Ingram, A., Phelan, A., Dunlop, J., and Clements, J. B. (1996). Immediate early protein IE63 of herpes simplex virus type 1 binds RNA directly. J. Gen. Virol. 77(Pt 8), 1847–1851. doi: 10.1099/0022-1317-77-8-1847
Harris, J. R., Cavallaro, E. C., de Nobrega, A. A., Dos, S. B. J. C., Bopp, C., Parsons, M. B., et al. (2009). Field evaluation of crystal VC Rapid Dipstick test for cholera during a cholera outbreak in Guinea-Bissau. Trop. Med. Int. Health 14, 1117–1121. doi: 10.1111/j.1365-3156.2009.02335.x
Mukherjee, P., Ghosh, S., Ramamurthy, T., Bhattacharya, M. K., Nandy, R. K., Takeda, Y., et al. (2010). Evaluation of a rapid immunochromatographic dipstick kit for diagnosis of cholera emphasizes its outbreak utility. Jpn. J. Infect. Dis. 63, 234–238.
Nato, F., Boutonnier, A., Rajerison, M., Grosjean, P., Dartevelle, S., Guenole, A., et al. (2003). One-step immunochromatographic dipstick tests for rapid detection of Vibrio cholerae O1 and O139 in stool samples. Clin. Diagn. Lab. Immunol. 10, 476–478.
Qadri, F., Hasan, J. A., Hossain, J., Chowdhury, A., Begum, Y. A., Azim, T., et al. (1995). Evaluation of the monoclonal antibody-based kit Bengal SMART for rapid detection of Vibrio cholerae O139 synonym Bengal in stool samples. J. Clin. Microbiol. 33, 732–734.
Rabbani, G. H., Teka, T., Zaman, B., Majid, N., Khatun, M., and Fuchs, G. J. (2001). Clinical studies in persistent diarrhea: dietary management with green banana or pectin in Bangladeshi children. Gastroenterology 121, 554–560. doi: 10.1053/gast.2001.27178
Rosewell, A., Addy, B., Komnapi, L., Makanda, F., Ropa, B., Posanai, E., et al. (2012). Cholera risk factors, Papua New Guinea, 2010. BMC Infect. Dis. 12:287. doi: 10.1186/1471-2334-12-287
Sinha, A., Sengupta, S., Ghosh, S., Basu, S., Sur, D., Kanungo, S., et al. (2012). Evaluation of a rapid dipstick test for identifying cholera cases during the outbreak. Indian J. Med. Res. 135, 523–528.
Spira, W. M., and Ahmed, Q. S. (1981). Gauze filtration and enrichment procedures for recovery of Vibrio cholerae from contaminated waters. Appl. Environ. Microbiol. 42, 730–733.
Keywords: cholera, dipstick, Bangladesh, environmental water, Vibrio cholerae
Citation: Chakraborty S, Alam M, Scobie HM and Sack DA (2013) Adaptation of a simple dipstick test for detection of Vibrio cholerae O1 and O139 in environmental water. Front. Microbiol. 4:320. doi: 10.3389/fmicb.2013.00320
Received: 19 August 2013; Paper pending published: 20 September 2013;
Accepted: 10 October 2013; Published online: 29 October 2013.
Edited by:
Daniela Ceccarelli, University of Maryland, USAReviewed by:
Shawn R. Campagna, University of Tennessee, Knoxville, USAMarie-Laure Quilici, Institut Pasteur, France
Copyright © 2013 Chakraborty, Alam, Scobie and Sack. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: David A. Sack, Department of International Health, Johns Hopkins Bloomberg School of Public Health, 615 North Wolfe Street, E 5537, Baltimore, MD 21205, USA e-mail:ZHNhY2tAamhzcGguZWR1
†Present address: Heather M. Scobie, Centers for Disease Control and Prevention, 1600 Clifton Road, MS A-04, Atlanta 30333, Georgia
 Subhra Chakraborty
Subhra Chakraborty Munirul Alam
Munirul Alam Heather M. Scobie1†
Heather M. Scobie1† David A. Sack
David A. Sack