- 1State Key Laboratory of Agricultural Microbiology, College of Life Sciences and Technology, Huazhong Agricultural University, Wuhan, P. R. of China
- 2Department of Plant and Environmental Sciences, University of Copenhagen, Frederiksberg, Denmark
In order to construct a more universal model for understanding the genetic requirements for bacterial AsIII oxidation, an in silico examination of the available sequences in the GenBank was assessed and revealed 21 conserved 5–71 kb arsenic islands within phylogenetically diverse bacterial genomes. The arsenic islands included the AsIII oxidase structural genes aioBA, ars operons (e.g., arsRCB) which code for arsenic resistance, and pho, pst, and phn genes known to be part of the classical phosphate stress response and that encode functions associated with regulating and acquiring organic and inorganic phosphorus. The regulatory genes aioXSR were also an island component, but only in Proteobacteria and orientated differently depending on whether they were in α-Proteobacteria or β-/γ-Proteobacteria. Curiously though, while these regulatory genes have been shown to be essential to AsIII oxidation in the Proteobacteria, they are absent in most other organisms examined, inferring different regulatory mechanism(s) yet to be discovered. Phylogenetic analysis of the aio, ars, pst, and phn genes revealed evidence of both vertical inheritance and horizontal gene transfer (HGT). It is therefore likely the arsenic islands did not evolve as a whole unit but formed independently by acquisition of functionally related genes and operons in respective strains. Considering gene synteny and structural analogies between arsenate and phosphate, we presumed that these genes function together in helping these microbes to be able to use even low concentrations of phosphorus needed for vital functions under high concentrations of arsenic, and defined these sequences as the arsenic islands.
Introduction
Microbial arsenite (AsIII) oxidation converts the more toxic AsIII to the less toxic AsV which is known to be catalyzed by a molybdenum-containing enzyme. The AsIII oxidase enzyme is encoded by aioBA (previously referred to as aoxAB) (Silver and Phung, 2005). Putative AioA were found to be specific for AsIII-oxidizing bacteria, therefore the usefulness of the aioBA as a functional marker indicating the ability to oxidize AsIII of a strain was proposed (Inskeep et al., 2007; Quéméneur et al., 2008; Hamamura et al., 2009). However, aioA is not a suitable marker for microbial diversity studies because its phylogeny does not always strictly correlate with that of the 16S rRNA genes, due to horizontal gene transfer (HGT) (Heinrich-Salmeron et al., 2011). In addition, in some AsIII-oxidizing strains, an aioA sequence was not detected either by PCR nor genome sequencing (Richey et al., 2009). It is now known that a new type of AsIII oxidase gene arxA, which was only distantly related to aioA, could be identified in a number of strains (Zargar et al., 2010, 2012).
Currently, the genetics underlying AsIII oxidation and its regulation are perhaps best understood in Agrobacterium tumefaciens 5A. The aioBA genes are part of the aioX-aioS-aioR-aioB-aioA-cytc2-chlE operon that has been shown to be regulated by a two-component regulatory pair comprised of the sensor kinase AioS and its cognate response regulator AioR in conjunction with the periplasmic AsIII-binding protein AioX (Kashyap et al., 2006; Koechler et al., 2010; Liu et al., 2012). In addition, the σ54 factor (RpoN) has been shown by two different groups to play a role in aioBA regulation (Koechler et al., 2010; Kang et al., 2012). A −24/−12 box for RpoN binding has been detected upstream of aioB and shown to be important for aioBA expression by 5′ RACE (rapid-amplification of cDNA ends) (Sardiwal et al., 2010) and precision deletion experiments (Koechler et al., 2010). Furthermore, AioR also contains a conserved domain for response regulators that could regulate σ54-type promoters (Sardiwal et al., 2010). RpoN is viewed to form a close complex with RNA polymerase, which requires energy provided by regulators for transcriptional initiation. The σ54-dependent regulators such as AioR may bind to upstream activation sequences (UAS) of σ54-type promoters for energy conservation (Shingler, 2011).
Regarding sequences in the vicinity of the aio operon, Silver and Phung (2005) first proposed the concept of an “arsenic island” based on a 71-kb DNA region of the Alcaligenes faecalis genome which contains over 20 functionally related genes such as those encoding AsIII oxidase AioBA, ArsAB for AsIII efflux, and a variety of oxyanion ABC transporters. Muller et al. (2007) reported the gene sequences in the vicinity of aioBA in Herminiimonas arsenicoxydans ULPAs1 and several other strains. Later, Arsène-Ploetze et al. (2010) proposed that aioBA was located in a genomic island which may have been acquired by HGT in Thiomonas sp. 3As. However, since only a few aio operons were known until recently, the definition and distribution of such arsenic islands was unclear and speculative. Due to the development and usage of high-throughput sequencing, more aio operons could be identified in microbial genomes (Hao et al., 2012; Huang et al., 2012; Li et al., 2012; Lin et al., 2012; Phung et al., 2012). From visual inspection, gene patterns associated with the aioBA genes became apparent and warranted a more detailed characterization.
One such pattern is the frequent physical association of genes involved with As and phosphorus (P) metabolism (Moreno-Sanchez et al., 2012). As and P are both members of Group 15 on the periodic table, resulting in their being structural analogs, such that AsV and phosphate may be co-metabolized, with the best examples involving AsV substituting for phosphate as substrate for phosphate transporters or interfering with ATP metabolism (Moreno-Sanchez et al., 2012). The Pho regulation is induced by P starvation and has been reported to control about 30 genes in 9 transcripts, including phnCDEFGHIJKLMNOP genes for phosphonate assimilation, phoE for outer membrane phosphoporin, phoA for alkaline phosphatase, pstSCAB genes for specific phosphate transport and ugpABCD genes for glycerol-3-phosphate transporter (Hsieh and Wanner, 2010). Recently, Kang et al. (2012) demonstrated that in A. tumefaciens strain 5A the close genomic association of the aioXSRBA genes with genes coding for functions involved with acquiring P under P-stress conditions is not simply coincidental. Surprisingly, induction of aioBA is repressed under high phosphate conditions and involves regulatory components of the phosphate stress response. Either or both PhoB response regulators, PhoB1 and PhoB2, are required for normal transcriptional kinetics of aioBA and aioSR. In addition, genes usually regulated by environmental phosphate levels, pstS1 and phoU, were found to be regulated by ArsR in an AsIII-dependent manner (Kang et al., 2012).
The primary aim of this study was to characterize the physical arrangement and functional relatedness of aio, pho, pst, and ars genes among the available arsenic islands. In addition, we also examined the phylogenetic relationships of these genes so as to assess mode of inheritance and used this information to begin assimilating a broader picture of how AsIII oxidation is regulated in different organisms and speculate the functional relationships of some genes that appear to have been repeatedly co-inherited in nature.
Results
Detection of Genes in the Vicinity of aio Operon Revealed Putative Arsenic Islands
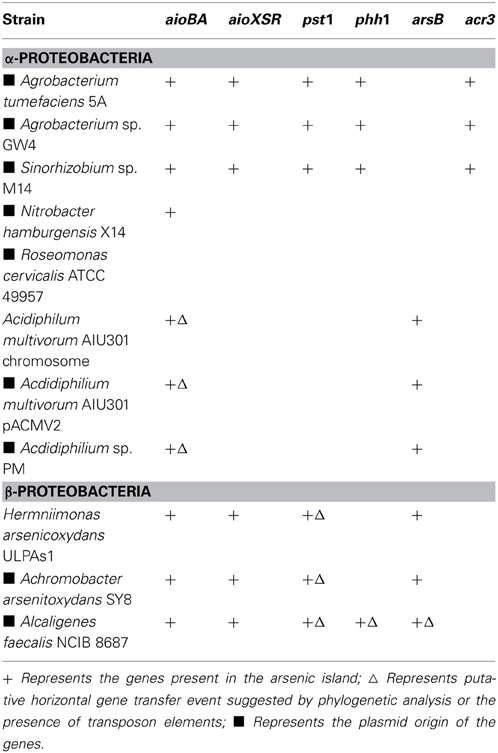
A total of 57 full-length aioBA operons encoding arsenite oxidase were detected in 55 strains using a BLAST search in GenBank. Among them, genes in the vicinity of the aioBA operons showed significant synteny among 21 genome sequences (ranging 5–71 kb). These genes that were all responsible for AsIII oxidation (21 aio operons, aioBA, or aioXSRBACD), arsenic resistance (23 ars operons, e.g., arsR, arsC, arsB, and acr3), phosphate transport (10 pst1 operons, e.g., pstS, pstC, pstA, and pstB) and phosphonate transport (6 phn1 operons, e.g., phnC, phnD, phnE, and phnE) were frequently detected (Figure 1). Considering the major function for AsIII's oxidation, we refer to these 21 sequences as arsenic oxidase gene islands (Figure 1). In some closely related bacterial strains, gene arrangements of the islands showed excellent synteny as similar gene and operon arrangements were found in A. tumefaciens 5A, Agrobacterium sp. GW4 and Sinorhizobium sp. M14. In addition, the same arrangement was also shared by Acidiphilium multivorum AIU301 and Acidiphilium sp. PM. However, other distantly related bacteria containing an arsenic island did not show a similar arrangement (Figure 1). Later on, Agrobacterium albertimagni strain AOL15 (Trimble et al., 2012a) and Achromobacter pichaudii strain HLE (Trimble et al., 2012b), were sequenced, but showed similar arsenic gene islands to A. arsenitoxydans SY8 (Li et al., 2012) and A. tumefaciens 5A, respectively (Hao et al., 2012) (data not shown).
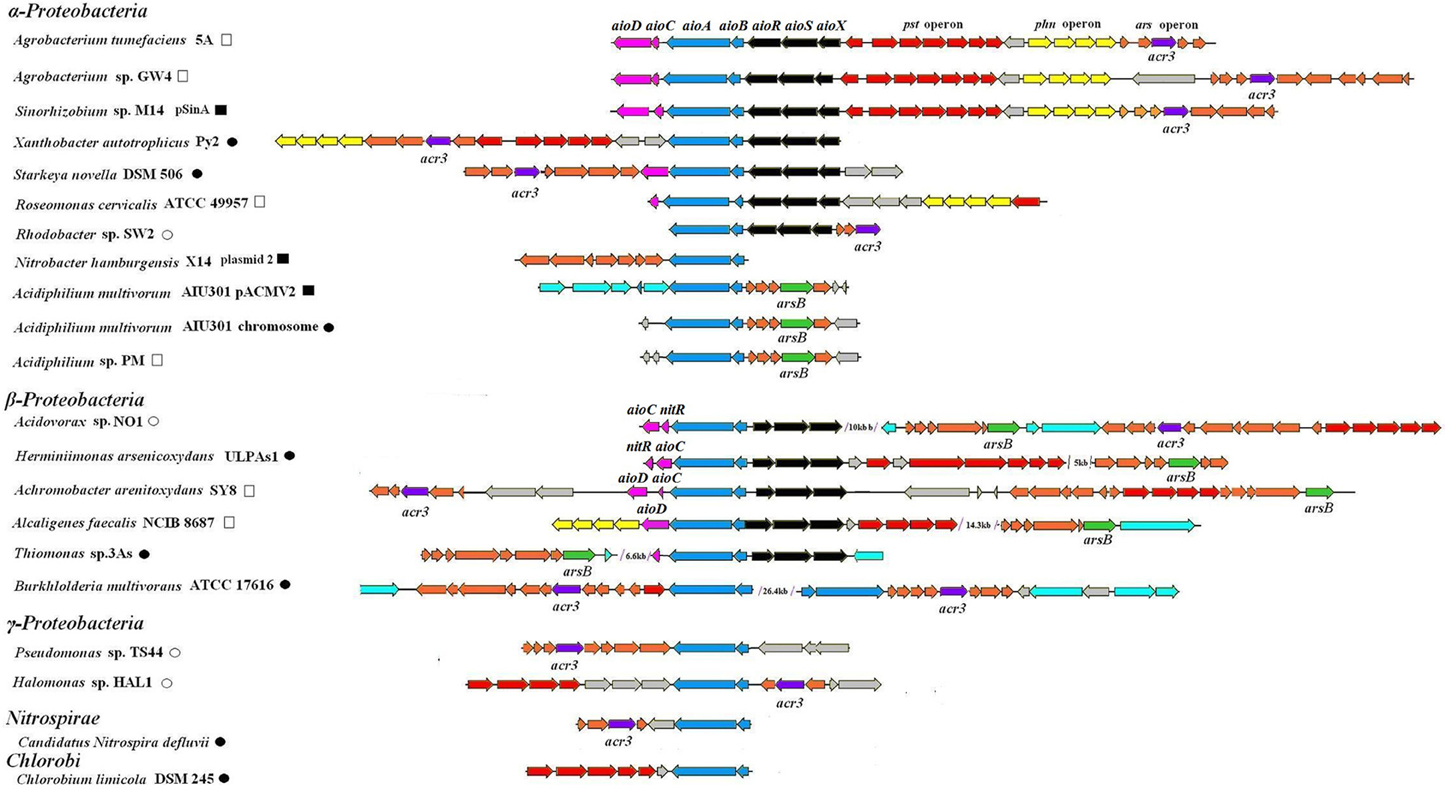
Figure 1. Gene arrangements of the 21 arsenic islands. Arrows with different colors represent the following genes: blue for aioBA, black for aioXSR, pink for aioCD or nitR (encoding cytochrome c and molybdenum biosynthesis protein or nitroreductase, respectively), red for pst operon, yellow for phn operon, orange for ars operon in which green for arsB, purple for acr3, light blue for mobile element. ■ and □ represent the reported and predicted plasmid-originated sequences, respectively. • and ° represent the reported and predicted chromosome-originated sequences, respectively. GenBank accession numbers are as follows: Acidovorax sp. NO1 (AGTS01000000), Herminiimonas arsenicoxydans ULPAs1 (CU207211), Alcaligenes faecalis NCIB 8687 (AY297781), Achromobacter arsenitoxydans SY8 (AGUF01000000), Sinorhizobium sp. M14 (GU990088), Agrobacterium sp. GW4 (JQ423942), Roseomonas cervicalis ATCC 49957 (NZ_ADVL01000677), Xanthobacter autotrophicus Py2 (NC_009720), Rhodobacter sp. SW2 (NZ_ACYY01000001), Starkeya novella DSM 506 (NC_014217), Nitrobacter hamburgensis X14 (NC_007960), Halomonas sp. HAL1 (EU651834), Pseudomonas sp. TS44 (EU311944), Candidatus nitrospira defiuvii (NC_014355), Chlorobium limicola DSM 245 (CP001097), Burkholderia multivorans ATCC17616 (NC_010087), Acidiphilium multivorum AIU301 (NC_015186 and NC_015187).
By scanning the genomes, we found that aio operons were only present as a single copy and often located within the arsenic islands. To better interpret the results of this analysis, it is important to point out that in addition to the pst or phn operons on the arsenic islands (here referred to as pst1 and phn1), almost all strains possessed another pst or phn operon located distantly (here referred to as pst2 or phn2). The phylogenies of the representative amino acid sequences (AioA, PstS, and PhnC) for aio, pst and phn operons were compared to their 16S rDNAs in order to determine whether possible HGT had taken place. Since ars operons have frequently been shown to associate with HGT events (Tuffin et al., 2005; Cai et al., 2009a), only the representative ars genes (acr3 or arsB) on the arsenic islands were analyzed in this study (see following sections for detailed results), although there are orthologs in many other phyla.
The Arsenic Islands are Localized on Chromosomes or on Plasmids
Of the 21 arsenic islands analyzed in this study, 3 have been shown to be localized on a plasmid (GenBank GU990088, CP000321 and AP012037) and 8 on a chromosome (GenBank CP000781, CP003126, AP012037, NC_009138, FP475956, NC_010087, FP929003, and CP001097) (Figure 1). To determine where all of the 21 arsenic islands are located, we performed a bioinformatics analysis to predict localization on a plasmid using the cBar program (Zhou and Xu, 2010). According to our analysis, nine arsenic islands were predicted to be located on a plasmid and 12 on a chromosome (Figure 1). The previously determined localization of 3 arsenic islands on a plasmid and eight on a chromosome were all correctly predicted, demonstrating good reliability of plasmid prediction using cBar. The plasmid-borne arsenic islands were prevalent in α-Proteobacteria (7/11). Notably, strains Agrobacterium sp. GW4, A. tumefaciens 5A, and Sinorhizobium sp. M14 shared similar “arsenic island” arrangements, all predicted to be located on a plasmid. Again, these predictions were in agreement with the known localization of the arsenic gene island on plasmid pSinA (GenBank GU990088) of strain Sinorhizobium sp. M14 (Figure 1). Furthermore, the three arsenic islands from Acidiphilium multivorum AIU301 and Acidiphilium sp. PM had a similar arrangement of their genes and were all localized on a plasmid (Figure 1), two of the arsenic islands were predicted by cBar whereas one occurs on pACMV2 (AP012037). It appears likely that these strains may have acquired their respective arsenic islands by HGT.
Wide Spread Distribution and Genomic Stability of Aioba
The phylogenetic tree of AioA was generally in accordance with the 16S rDNA phylogeny which branched into Proteobacteria, Chlorobi, Deinococcus-Thermus, Chloroflexi, and Archaea (Figure 2), and the AioBA phylogeny was similar to that of AioA (data not shown). Most of the sequences encoding AioA were found in Proteobacteria, and mainly be divided into two groups. Group I is made up of sequences of α-Proteobacteria and Group II is comprised of β- and γ-Proteobacteria. This distribution was consistent with a previous analysis in which partial AioA sequences obtained by degenerate primers was distributed along a similar pattern (Heinrich-Salmeron et al., 2011). However, eight AioA-like proteins from marine α- or γ-Proteobacteria clustered together into a separate branch and exhibited a unique arrangement. The genes encoding this subfamily of AioA were all located downstream of two genes encoding the cytochrome c peroxidase MauG and were arranged in the gene order of mauG-mauG-aioBA. This phylogenetically distinct clade of these AioA-like proteins may have evolved in their marine environment due to unique conditions. Compared to the 16S rDNA phylogenetic tree, there were three conflicts with the AioA phylogenetic tree, which suggests the occurrence of HGT. For example, the two AioAs in Acidiphilium multivorum AIU301 (one located from the chromosome and one from a plasmid) and the AioA in Acidiphilium sp. PM fell into a clade together with Chlorobi and Deinococcus-Thermus, respectively (Figure 2). The Chlorobi or Deinococcus-Thermus strains appears to had transferred the aioA into A. multivorum AIU301 and Acidiphilium sp. PM since they all have been isolated from a similar acidic environment (San Martin-Uriz et al., 2011). In addition, a HGT might also be more likely since the two AioAs in A. multivorum AIU301 and Acidiphilium sp. PM were also predicted by cBar to be located on plasmids (Figure 1). Chlorflexus aggregans DSM 9485, Chloroflexus sp. J-10-fl and Y-400-fl are closely related based on 16S rDNA analysis, while the AioA of C. aggregans DSM 9485 clustered with the AioAs from Deinococcus-Thermus (Figure 2). It appears that A. arsenitoxydans SY8 had transferred aioA into Ralstonia sp. 22 by HGT (Lieutaud et al., 2010), and here again we found that the aioA of A. arsenitoxydans SY8 is located on plasmid.
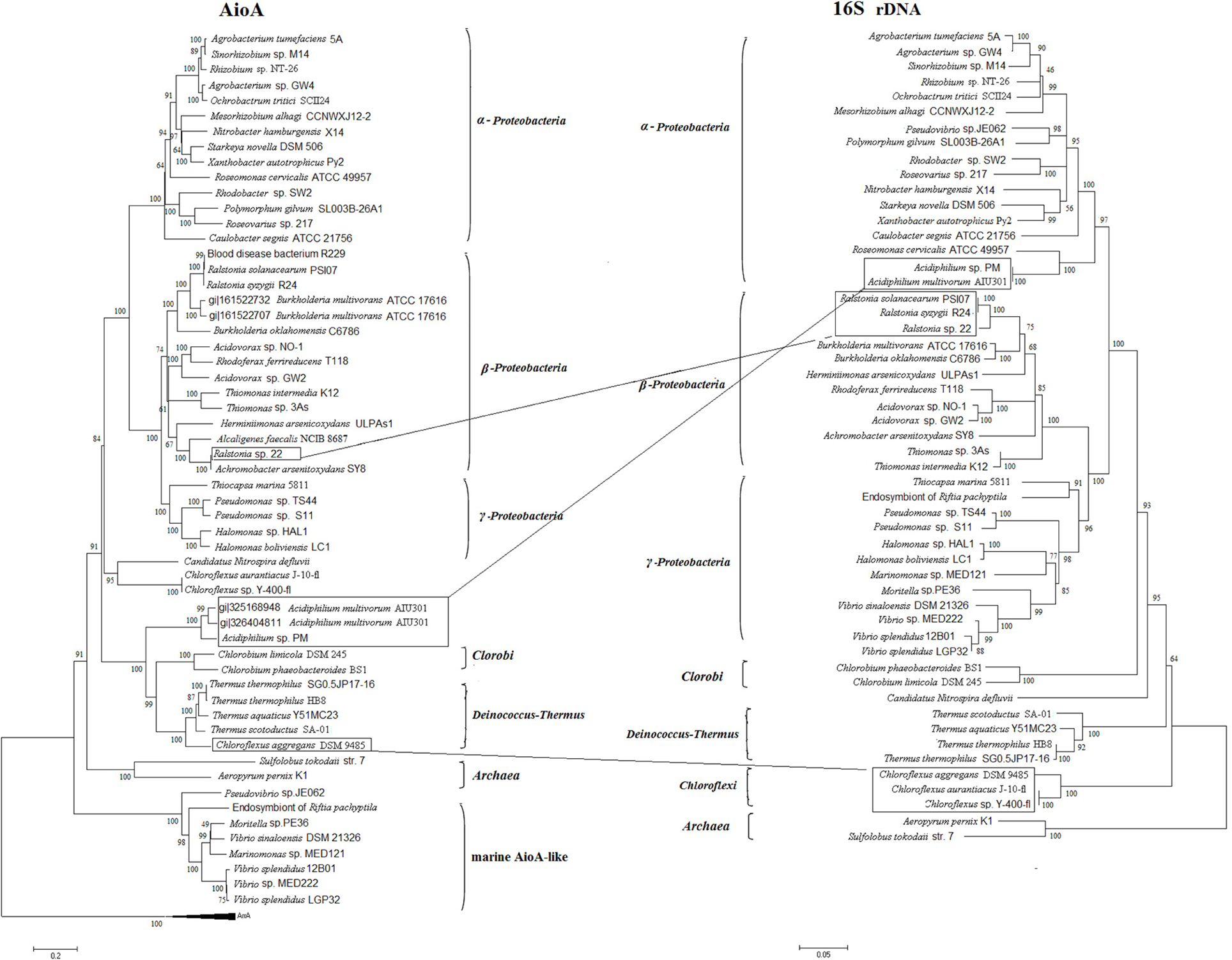
Figure 2. The neighbor-joining (NJ) phylogenetic trees of AioA sequences and 16S rDNA sequences. Putative horizontal gene transfer events (labeled with frames) have been compared based on the inconsistency of AioA amino acid tree (on the left) and the 16S rDNA tree (on the right).
Analysis of the Putative Regulators for aioBA Operons Suggested Different Mechanisms of aio Regulation
The genes encoding regulators AioXSR located upstream of aioBA were only identified in 12 strains of Proteobacteria among the 21 analyzed strains encoding aioBA as part of their arsenic island. All of these 12 strains belonged to either α-Proteobacteria or β-Proteobacteria (Figure 1). The transcriptional orientation of aioXSR genes differed between α- and β-Proteobacteria, which is in agreement with the AioA phylogeny (Figure 2). The aioXSR genes from α-Proteobacteria displayed the same transcriptional orientation as aioBA, while those from β-Proteobacteria displayed the opposite orientation (Figure 1). The other nine sequences without aioXSR genes were distributed in different taxonomic groups such as Proteobacteria, Chlorobi and Nitrospriae. In some of these identified species such as Pseudomonas sp. TS44 and Halomonas sp. HAL1, AsIII oxidation could be verified (Cai et al., 2009b; Lin et al., 2012). The mode and mechanism regulating expression of aioBA in these strains is unknown but might involve distantly located regulators. It is interesting that there are nitRs (encoding nitroreductases) after the aioC instead of aioD in Acidovorax sp. NO1 (AGTS01000000), Herminiimonas arsenicoxydans ULPAs1 (Figure 1). Recently, a disruption of the nitR in strain NO1 resulted in the delay of AsIIII oxidation indicating that the nitR may participate to the electron transfer in the strain (data not shown).
Phylogenetical Analyses of Arsenite Efflux Proteins ArsB or ACR3 Encoded in ars Operons of the Arsenic Islands.
A total of 23 ars operons were detected in the arsenic islands and their inheritance models were analyzed by phylogenetic analyses of ArsB or ACR3, and comparing their phylogeny with those obtained using 16S rDNA. Among the 23 ars operons, eight ArsB and 13 ACR3 were detected. Some ars operons without the arsB or acr3 genes such as those from N. hamburgensis X14 and A. arsenitoxydans SY8 could not be analyzed in this context.
Phylogenetic analysis suggested most ArsB arsenite efflux proteins were congruent with 16S rDNAs (Supplementary materials, Figure S1). However, a notable exception included the ArsBs from Acidovorax sp. NO1, Thiomonas sp. 3A and A. faecalis NCIB8687, which clustered together. All of these ArsB proteins were encoded as part of a transposon, again indicating HGT events by transposon insertion accounted for acquisition of these ars operons in the respective arsenic islands.
The ACR3s separated into two clades in previous studies (Achour et al., 2007) and we also found that ACR3s on the arsenic islands that could be divided into ACR3 (1) and ACR3 (2). In the respective ACR3 clades, their phylogenies were both in accordance with 16S rDNA phylogeny, therefore, suggesting genomic stability (Figure S2).
Phylogenetic Analyses of the Phosphorus Related pst And phn Operons
The pst1 or phn1 genes are localized within arsenic islands, while pst2 and phn2 genes are localized distantly on the respective chromosomes. Phylogenetic analysis indicated that all of the Pst2 branched in accordance with the 16S rDNAs (Figure S3). Therefore, pst2 operons appear to follow vertical inheritance. However, Pst1 did not strictly branch as the phylogenetic tree based on the 16S rDNA sequences (Figure 3). The Pst1 of Alcaligenes faecalis NCIB 8687 (β-Proteobacteria) clustered together with the Pst1 of α-Proteobacteria strains Agrobacterium tumefaciens 5A, Agrobacterium sp. GW4, Sinorhizobium sp. M14 and Xanthobacter autotrophicus Py2. The Pst1 of A. arsenitoxydans SY8 and H. arsenicoxydans ULPAs1 (β-Proteobacteria) were more related to those from γ-Proteobacteria. These results suggest that HGT may have occurred in transmission of the pst1 operon.
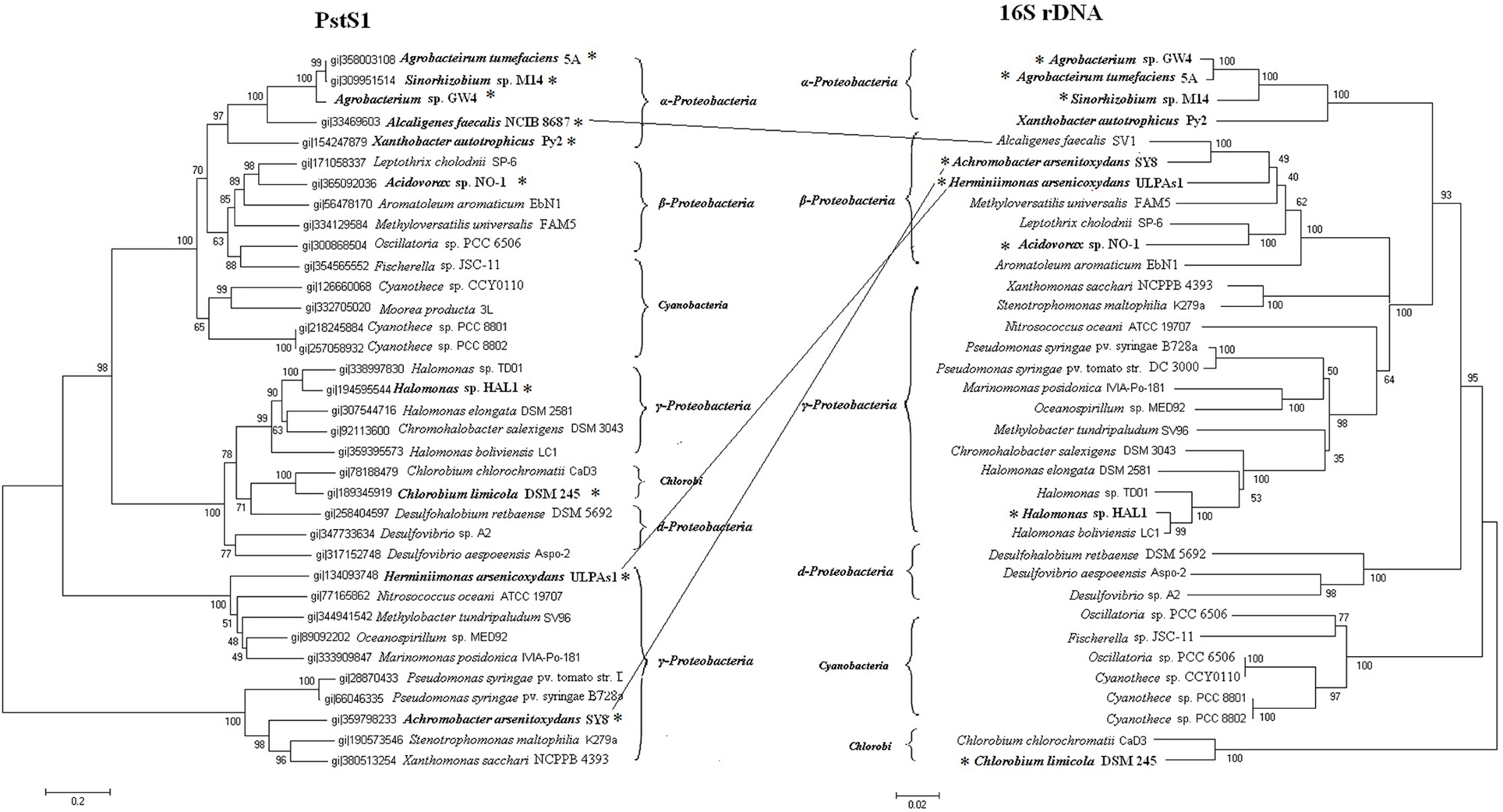
Figure 3. Phylogenetic trees based on PstS1 sequences and 16S rDNA sequences. Bold and *symbol represent proteins from the strains of the arsenic islands while the others are not. Putative horizontal gene transfer events (connected lines) have been compared based on the inconsistency of the amino acid sequence tree (on the left) and the 16S rDNA tree (on the right).
The phylogenies of Phn2 were in accordance with those calculated for the 16S rDNAs (Figure S4). However, Phn1 showed some conflicts (Figure 4). A Phn1 (A. faecalis NCIB 8687) from β-Proteobacteria clustered with the α-Proteobacteria. The phn2 loci were usually arranged as phnCDEE' and located in the vicinity of other phosphonate utilizing genes, such as phnFGHIJKLMNOP (Jochimsen et al., 2011). The phn1 locus A. faecalis NCIB 8687 was arranged as phnDCEE', which had no other functional related genes in vicinity. Thus, the phn1 and phn2 may be functional different operons.
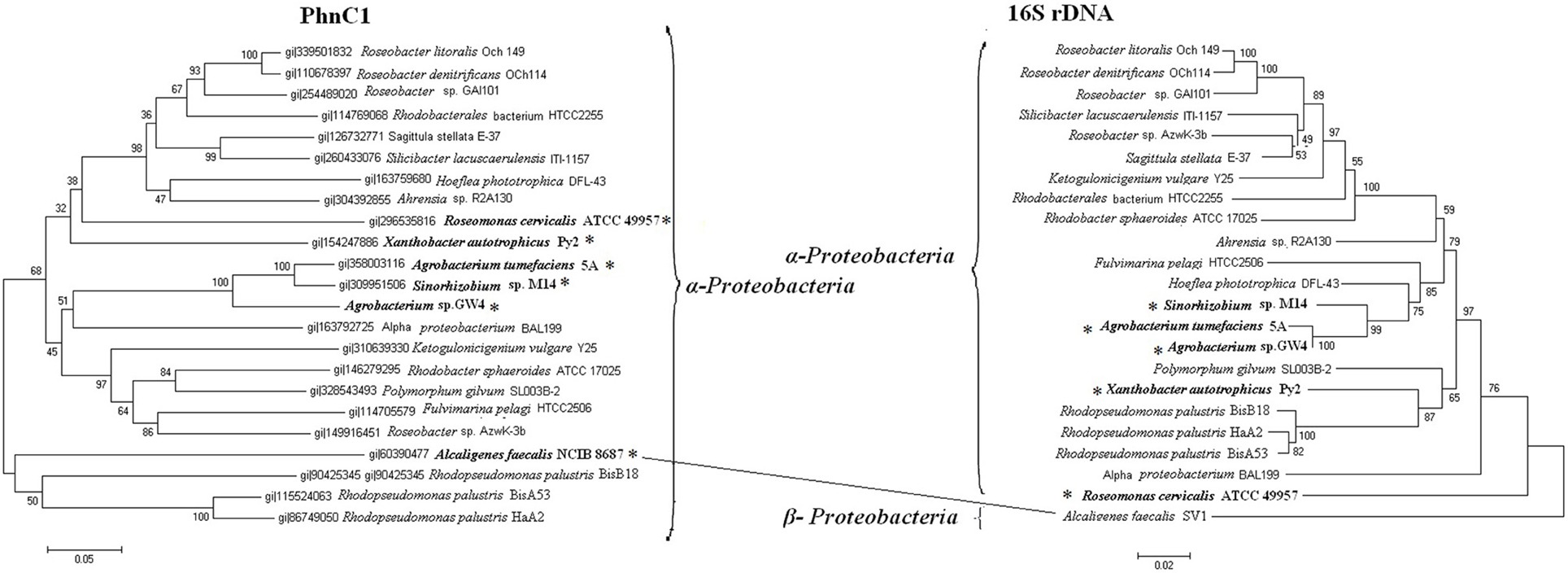
Figure 4. Phylogenetic trees of PhnC1 and 16S rDNA sequences. Bold and *symbol represent proteins from the strains of the arsenic islands while the others are not. Putative horizontal gene transfer events (connected lines) have been compared based on the inconsistency of the amino acid sequence tree (on the left) and the 16S rDNA tree (on the right).
Discussion
This study provides a comprehensive analysis of most of the available full-length AioBA sequences. Large scale scanning of the sequences in the vicinity of aioBA operons revealed the frequent occurrence of genes related to arsenic and phosphorous metabolism, such as the regulatory aioXSR operon and pst, phn, and ars operons (Silver and Phung, 2005). Considering gene synteny and structural analogies between arsenate and phosphate, we presumed that these genes function together in helping these microbes to be able to use even low concentrations of phosphorus needed for vital functions under high concentrations of arsenic, and defined these sequences as the arsenic islands. The aioBA operons function to convert AsIII to the less toxic AsV but frequently also use this as a chemolithotrophic energy source. In contrast, ars operons are responsible for arsenic efflux after arsenate reduction and have a purely protective role (Silver and Phung, 2005). We found that some strains contain pst1 or phn1 operons encoding putative phosphate and phosphonate uptake transport systems in the vicinity of the aio operons in addition to the distantly located pst2 or phn2 operons, which raises the question about the functional role of pst1 and phn1 operons. Previous results indicated that arsenate can increase the Vmax of Pst2 for phosphate uptake (Moreno-Sanchez et al., 2012). AsIII may induce phosphate starvation as a competitive inhibitor of phosphate uptake, and cells may need to express more of these transporters or possibly more specific transporters for phosphate uptake. Similarly, we conjecture that the AsV generated by the AioBA may lead to phosphate starvation and the pst1 and phn1 may encode additional more specific uptake systems for P assimilation. This proposition is in accordance with the transcriptional profile of H. arsenicoxydans ULPAs1, in which pst1 operon was induced under conditions of As exposure (Cleiss-Arnold et al., 2010). Recently, it was reported that a pst1-like protein discriminated P from AsV 500–850-fold in phosphate-limited condition (Elias et al., 2012). In addition, one could envision PstS1 transporting AsV into the cells and deposited into acidicalcisomes or as part of polyphosphate granules. This had been suggested by Moreno-Sanchez et al. (2012).
In this study, we analyzed the localization of AsIII oxidation genes and found that aioBA of the α-Proteobacteria was prevalently localized on plasmids. As many arsenic islands were localized on plasmids, we predict that plasmids played a role in the widespread distribution of aioBA. Most of the aioBA sequences analyzed here could be retrieved from Proteobacteria and these sequences could be assigned to two groups, α-Proteobacteria and β-/γ-Proteobacteria, consistent with a previous analysis (Hamamura et al., 2009). The AioAs generally showed similar phylogeny as their 16S rDNA sequences (Figure 2) indicating an ancient origin of the enzyme (Cai et al., 2009b; Zhou and Xu, 2010). However, several strains showed putative HGT events with AioAs (Figure 2; Table 1) suggesting HGT also play a role during inheritance process (Arsène-Ploetze et al., 2010; Heinrich-Salmeron et al., 2011).
Unlike aioBAs, which are widely distributed among Proteobacteria, Chlorobi, Deinococcus-Thermus, Chloroflexi, and even Archaea, the three component regulator genes genes aioXSR were only found in Proteobacteria and displayed opposite transcriptional orientation between α- and β-Proteobacteria. It was possible that the aioXSR genes emerged in Proteobacteria after the introduction of aioBA. The regulation of these aioBA operons with no aioXSR genes is not clear, but they may be controlled by distantly located regulators, or quorum sensing, as proposed by Kashyap et al. (Kashyap et al., 2006). Thus, the regulatory genes aioXSR may have evolved independently from aioBA. In a few strains including A. tumefaciens 5A, AioSR regulation of aioBA was RpoN-dependent, and the -24/-12 region for RpoN (σ54 factor for RNA polymerase) binding was also detected (Kang et al., 2012). The arsenite oxidase regulator AioR belonged to the NtrC family indicating that aioBA may be under the regulation of RpoN-dependent σ54-type promoter. However, the molecular details of AioR interacting with the promoter, and of the RpoN-RNA polymerase complex initiating transcription are still not known. Here we identified two tandem repeats of palindrome-like sequences which are located 100–200 nt upstream of the aioB start codon (Figure 5). The palindrome-like sequences are probably the upstream activating sequences (UAS) of σ54-type promoters which function in binding of AioR. The two palindromes and the -24/-12 regions were detected in all of the 12 aioBA operons that contained the aioXSR three-component system, but absent in other aioBA operons without aioXSR. Thus, we have to propose that the aioBAs without the upstream sequences of aioXSR are regulated differently.
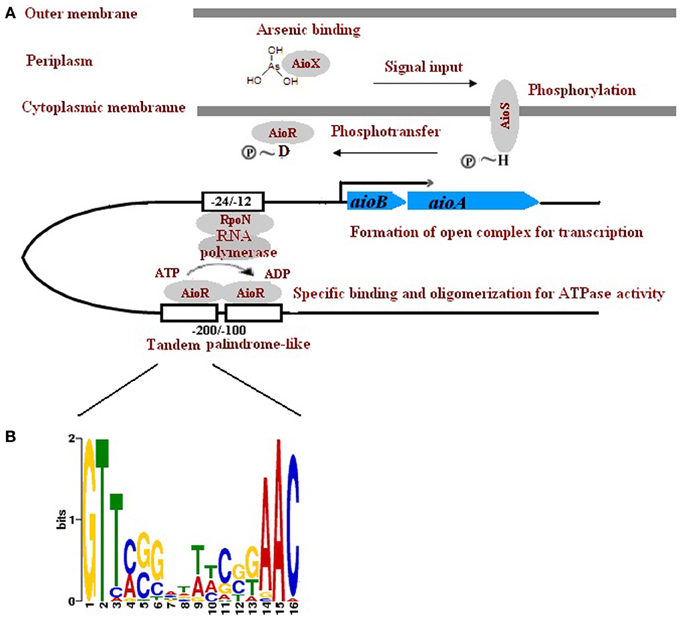
Figure 5. A proposed mechanism aioXSR mediated aioBA regulation. (A) AioX binds to arsenite and delivers the signal to the AioS/AioR two-component system. The phosphorylated AioR binds to the palindrome-like sequence, leading to oligomerization for ATPase activity. Energy conserved by AioR would open the RopN-RNA polymerase complex, and initiate the aioBA transcription. (B) Sequence logo of palindrome-like sequences located upstream of aioBA. Higher bit scores indicate more conservation at respective site.
The ars, pst, and phn operons were frequently detected on the arsenic gene islands but did not display a similar arrangement in various strains. Some plasticity was found even in the taxonomically closely related strains A. tumefaciens 5A, Agrobacterium sp. GW4 and Sinorhizobium sp. M14. These strains shared the same arrangement in aio, pst, and phn operons, but not in ars operons. The large scale synteny of aio, pst and phn operons in these three strains may be due to vertical inheritance, while ars operons were integrated independently into the arsenic islands.
The ars operons encoded either ArsB or ACR3 as the AsIII efflux pump but did not display the same arrangement of the remaining genes such as arsC or arsH. This does not indicate a common origin of the different ars operons on the respective arsenic islands. The phylogenetic relatedness of ArsB or ACR3 seems to be in accordance with the corresponded tree predicted by 16S rDNA comparison. This suggests that ArsB or ACR3 were both mostly vertically inherited from the gene pool of the respective taxonomic clade. Vertical inheritance and HGT may have contributed to the origin of pst1 and phn1 operons (Table 1). It is therefore likely the arsenic islands did not evolve as a whole unit but formed independently by acquisition of functionally related genes and operons in respective strains. The elucidation of the phylogeny and distribution of aio genes might provide further insight into the evolution of the aioBA operon, and lead to better understanding of the arsenic island.
Methods
Data Sources
The amino acids sequence of AioA from Agrobacterium tumefaciens 5A was used as the initial query for a BLASTP search at the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov). Partial AioA sequences obtained from degenerate primers were ignored, as there was usually no flanking sequence information for them. We selected the full-length AioA sequences with the following threshold: sequence identity >30%, coverage >80%, starting with methionine and harboring the conserved domain TIGR02693 specific for arsenite oxidase. The selected BLASTP hits were used as query sequences for additional BLASTP searches, until no more full-length AioA hits were found. The corresponding nucleotide sequences where aioA was located, as well as the gene annotation information were downloaded in GenBank format for further analysis.
Detection of Gene Synteny in the Arsenic Islands
The GenBank formatted sequences containing 57 aioA genes were loaded in the CLC sequence viewer program (http://www.clcbio.com). And the downstream and upstream sequences were scanned over 100 kb. Twenty-one sequences were found in vicinity of aioBA which were called arsenic islands, the others are single aioBA. The genes in the arsenic islands were exported as image files with the same genes represented by the same colors to detect synteny.
Phylogenetic Analysis of Nucleotide or Amino Acid Sequences
All of the gene sequences were searched in the GenBank using the aioA sequence and a neighbor-joining (NJ) phylogenetic tree was constructed using ClustalX analysis (Thompson et al., 1997) and MEGA 4.0 software (Tamura et al., 2007). The parameters are as follows: phylogeny test and options (Bootstrap, 1000 replicates), Gaps/Missing Data (Pairwise Deletion), Substitution Model (Poisson correction for amino acids, Kimura 2P for nucleotides). Later on, other phylogenetic comparisons were made using the some methods for 16S rDNA, ArsB, Acr3, pstS, phnC, and the sequences were extracted from the corresponding genomes or other related genomes when necessary.
Predicting of the Chromosome and Plasmid Location of aioA Genes
The information on chromosome or plasmid location for all the 21 arsenic gene islands, if existing, was identified from the strain notes in GenBank. However, many of the 21 arsenic islands had no information on chromosome or plasmid location because they were from draft genomes. We predicted the chromosome and plasmid location of all the 21 “arsenic islands” by the cBar program (Zhou and Xu, 2010). The cBar program was developed for classifying metagenomes into chromosomal and plasmid sequences based on their different nucleotide pentamer frequencies.
Detection of Conserved Sequence Motifs
The upstream 300 bp sequences of all the 57 aioBA genes were selected. The conserved motifs were detected by The MEME Suite motif-based sequence analysis tools (http://meme.sdsc.edu/meme/intro.html). The sequence logo which graphically represents the sequence conservation was also automatically generated by MEME on-line program.
Authors' Contributions
Hang Li carried out data collection, participated in the bioinformatic analyses and wrote the draft of the manuscript. Mingshun Li participated in bioinformatic analyses and helped to draft the manuscript. Yinyan Huang participated in sequence alignment study. Christopher Rensing participated in the design of the study and drafted the manuscript. Gejiao Wang coordinated the study, participated in its design and wrote the draft of the manuscript. All authors read and approved the final manuscript. We thank Dr. Timothy McDermott for discussion of the study design, editing, and comments on the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by the National Natural Science Foundation of China 31010103903 and 31170106.
Supplementary Material
The Supplementary Material for this article can be found online at: http://www.frontiersin.org/journal/10.3389/fmicb.2013.00347/abstract
Supplementary Figure S1 | Phylogenetical trees of ArsB and 16S rDNA sequences. Bold and *symbol represent proteins from the strains of the arsenic islands while the others are not. Phylogenetic relationship have been compared based on the amino acid sequence tree (on the left) and the 16S rDNA tree (on the right).
Supplementary Figure S2 |Phylogenetical trees of ACR3 and 16S rDNA sequences. Bold and *symbol represent proteins from the strains of the arsenic islands while the others are not. Phylogenetic relationship have been compared based on the amino acid sequence tree (on the left) and a 16S rDNA tree (on the right).
Supplementary Figure S3 |Phylogenetic trees based on PstS2 and 16S rDNA sequences. Bold and *symbol represent proteins from the strains of the arsenic islands while the others are not. Phylogenetic relationship have been compared based on the inconsistency of the amino acid sequence tree (on the left) and the 16S rDNA tree (on the right).
Supplementary Figure S4 |Phylogenetic trees of PhnC2 and 16S rDNA sequences. Bold and *symbol represent proteins from the strains of the arsenic islands while the others are not. Phylogenetic relationship have been compared based on the amino acid sequence tree (on the left) and the16S rDNA tree (on the right).
References
Achour, A. R., Bauda, P., and Billard, P. (2007). Diversity of arsenite transporter genes from arsenic-resistant soil bacteria. Res. Microbiol. 158, 128–137. doi: 10.1016/j.resmic.2006.11.006
Arsène-Ploetze, F., Koechler, S., Marchal, M., Coppée, J. Y., Chandler, M., and Bonnefoy, V. (2010). Structure, function, and evolution of the Thiomonas spp. genome. PLoS Genet. 6:e1000859. doi: 10.1371/journal.pgen.1000859
Cai, L., Liu, G., Rensing, C., and Wang, G. (2009a). Genes involved in arsenic transformation and resistance associated with different levels of arsenic-contaminated soils. BMC Microbial. 9:4. doi: 10.1186/1471-2180-9-4
Cai, L., Rensing, C., Li, X., and Wang, G. (2009b). Novel gene clusters involved in arsenite oxidation and resistance in two arsenite oxidizers: Achromobacter sp. SY8 and Pseudomonas sp. TS44. Appl. Microbiol. Biotechnol. 83, 715–725. doi: 10.1007/s00253-009-1929-4
Cleiss-Arnold, J., Koechler, S., Proux, C., Fardeau, M. L., Dillies, M. A., and Coppee, J. Y. (2010). Temporal transcriptomic response during arsenic stress in Herminiimonas arsenicoxydans. BMC Genomics 11:709. doi: 10.1186/1471-2164-11-709
Elias, M., Wellner, A., Goldin-Azulay, K., Chabriere, E., Vorholt, J. A., and Erb, T. J. (2012). The molecular basis of phosphate discrimination in arsenate-rich environments. Nature 491, 134–137. doi: 10.1038/nature11517
Hamamura, N., Macur, R. E., Korf, S., Ackerman, G., Taylor, W. P., and Kozubal, M. (2009). Linking microbial oxidation of arsenic with detection and phylogenetic analysis of arsenite oxidase genes in diverse geothermal environments. Environ. Microbiol. 11, 421–431. doi: 10.1111/j.1462-2920.2008.01781.x
Hao, X., Lin, Y., Johnstone, L., Liu, G., Wang, G., Wei, G., et al. (2012). Genome sequence of the arsenite-oxidizing strain Agrobacterium tumefaciens 5A. J. Bcateriol. 194, 903. doi: 10.1128/JB.06585-11
Heinrich-Salmeron, A., Cordi, A., Brochier-Armanet, C., Halter, D., Pagnout, C., and Abbaszadeh-fard, E. (2011). Unsuspected diversity of arsenite-oxidizing bacteria as revealed by widespread distribution of the aoxB gene in prokaryotes. Appl. Environ. Microbiol. 77, 4685–4692. doi: 10.1128/AEM.02884-10
Hsieh, Y. J., and Wanner, B. L. (2010). Global regulation by the seven-component Pi signaling system. Curr. Opin. Microbiol. 13, 198–203. doi: 10.1016/j.mib.2010.01.014
Huang, Y., Li, H., Rensing, C., Zhao, K., Johnstone, L., and Wang, G. (2012). Genome sequence of the facultative anaerobic arsenite-oxidizing and nitrate-reducing bacterium Acidovorax sp. strain NO1. J. Bacteriaol. 194, 1635–1636. doi: 10.1128/JB.06814-11
Inskeep, W. P., Macur, R. E., Hamamura, N., Warelow, T. P., Ward, S. A., and Santini, J. M. (2007). Detection, diversity and expression of aerobic bacterial arsenite oxidase genes. Environ. Microbiol. 9, 934–943. doi: 10.1111/j.1462-2920.2006.01215.x
Jochimsen, B., Lolle, S., McSorley, F. R., Nabi, M., Stougaard, J., and Zechel, D. L. (2011). Five phosphonate operon gene products as components of a multi-subunit complex of the carbon-phosphorus lyase pathway. Proc. Natl. Acad. Sci. U.S.A.. 108, 11393–11398. doi: 10.1073/pnas.1104922108
Kang, Y. S., Bothner, B., Rensing, C., and McDermott, T. R. (2012). Involvement of RpoN in regulating bacterial arsenite oxidation. Appl. Environ. Microbiol. 78, 5638–5645. doi: 10.1128/AEM.00238-12
Kashyap, D. R., Botero, L. M., Franck, W. L., Hassett, D. J., and McDermott, T. R. (2006). Complex regulation of arsenite oxidation in Agrobacterium tumefaciens. J. Bacteriol. 188, 1081–1088. doi: 10.1128/JB.188.3.1081-1088.2006
Koechler, S., Cleiss-Arnold, J., Proux, C., Sismeiro, O., Dillies, M. A., and Goulhen-Chollet, F. (2010). Multiple controls affect arsenite oxidase gene expression in Herminiimonas arsenicoxydans. BMC Microbiol. 10:53–65. doi: 10.1186/1471-2180-10-53
Li, X., Hu, Y., Gong, J., Lin, Y., Johnstone, L., Rensing, C., et al. (2012). Genome sequence of the highly efficient arsenite-oxidizing bacterium Achromobacter arsenitoxydans SY8. J. Bacteriol. 194, 1243–1244. doi: 10.1128/JB.06667-11
Lieutaud, A., van Lis, R., Duval, S., Capowiezl, L., Muller, D., and Lebrun, R. (2010). Arsenite oxidase from Ralstonia sp. 22: characterization of the enzyme and its interaction with soluble cytochromes. J. Biol. Chem. 285, 20433–20441. doi: 10.1074/jbc.M110.113761
Lin, Y., Fan, H., Hao, X., Johnstone, L., Hu, Y., Wei, G., et al. (2012). Draft genome sequence of Halomonas sp. strain HAL1, a moderately halophilic arsenite-oxidizing bacterium isolated from gold-mine soil. J. Bacteriol. 194, 199–200. doi: 10.1128/JB.06359-11
Liu, G., Liu, M., Kim, E. H., Matty, W., Bothner, B., Lei, B., et al. (2012). A periplasmic arsenite-binding protein involved in regulating arsenite oxidation. Environ. Microbiol. 14, 1624–1634. doi: 10.1111/j.1462-2920.2011.02672.x
Moreno-Sanchez, D., Hernandez-Ruiz, L., Ruiz, F. A., and Docampo, R. (2012). Polyphosphate is a novel pro-inflammatory regulator of mast cells and is located in acidocalcisomes. J. Biol. Chem. 287, 28435–28444. doi: 10.1074/jbc.M112.385823
Muller, D., Médigue, C., Koechler, S., Barbe, V., Barakat, M., and Talla, E. (2007). A tale of two oxidation states: bacterial colonization of arsenic-rich environments. PLoS Genet. 3:e53. doi: 10.1371/journal.pgen.0030053
Phung, L. T., Trimble, W. L., Meyer, F., Gilbert, J. A., and Silver, S. (2012). Draft genome sequence of Alcaligenes faecalis subsp. faecalis NCIB 8687 (CCUG 2071). J. Bacteriol. 194, 5153. doi: 10.1128/JB.01185-12
Quéméneur, M., Heinrich-Salmeron, A., Muller, D., Lièvremont, D., Jauzein, M., and Bertin, P. N. (2008). Diversity surveys and evolutionary relationships of aoxB genes in aerobic arsenite-oxidizing bacteria. Appl. Environ. Microbiol. 74, 4567–4573. doi: 10.1128/AEM.02851-07
Richey, C., Chovanec, P., Hoeft, S. E., Oremland, R. S., Basu, P., and Stolz, J. F. (2009). Respiratory arsenate reductase as a bidirectional enzyme. Biochem. Biophys. Res. Commun. 382, 298–302. doi: 10.1016/j.bbrc.2009.03.045
San Martin-Uriz, P., Gomez, M. J., Arcas, A., Bargiela, R., and Amils, R. (2011). Draft genome sequence of the Electricigen Acidiphilium sp. strain PM (DSM 24941). J. Bacteriol. 193, 5585–5586. doi: 10.1128/JB.05386-11
Sardiwal, S., Santini, J. M., Osborne, T. H., and Djordjevic, S. (2010). Characterization of a two-component signal transduction system that controls arsenite oxidation in the chemolithoautotroph NT-26. FEMS Microbiol. Lett. 313, 20–28. doi: 10.1111/j.1574-6968.2010.02121.x
Shingler, V. (2011). Signal sensory systems that impact σ54-dependent transcription. FEMS Microbiol. Rev. 35, 425–440. doi: 10.1111/j.1574-6976.2010.00255.x
Silver, S., and Phung, L. T. (2005). Genes and enzymes involved in bacterial oxidation and reduction of inorganic arsenic. Appl. Environ. Microbiol. 71, 599–608. doi: 10.1128/AEM.71.2.599-608.2005
Tamura, K., Dudley, J., Nei, M., and Kumar, S. (2007). MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24, 1596–1599. doi: 10.1093/molbev/msm092
Thompson, J. D., Gibson, T. J., Plewniak, F., Jeanmougin, F., and Higgins, D. G. (1997). The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876–4882. doi: 10.1093/nar/25.24.4876
Trimble, W. L., Phung, L. T., Meyer, F., Gilbert, J. A., and Silver, S. (2012a). Draft genome sequence of Agrobacterium albertimagni strain AOL15. J. Bacteriol. 194, 6986–6987. doi: 10.1128/JB.01912-12
Trimble, W. L., Phung, L., Meyer, F., Silver, S., and Gilbert, J. A. (2012b). Draft genome sequence of Achromobacter piechaudii strain HLE. J. Bacteriol. 194, 6355. doi: 10.1128/JB.01660-12
Tuffin, I. M., de Groot, P., Deane, S. M., and Rawlings, D. E. (2005). An unusual Tn21-like transposon containing an ars operon is present in highly arsenic-resistant strains of the biomining bacterium Acidithiobacillus caldus. Microbiology 151, 3027–3039. doi: 10.1099/mic.0.28131-0
Zargar, K., Conrad, A., Bernick, D. L., Lowe, T. M., Stolc, V., and Hoeft, S. (2012). ArxA, a new clade of arsenite oxidase within the DMSO reductase family of molybdenum oxidoreductases. Environ. Microbiol. 14, 1635–1645. doi: 10.1111/j.1462-2920.2012.02722.x
Zargar, K., Hoeft, S., Oremland, R., and Saltikov, C. W. (2010). Identification of a novel arsenite oxidase gene, arxA, in the haloalkaliphilic, arsenite-oxidizing bacterium Alkalilimnicola ehrlichii strain MLHE-1. J. Bacteriol. 192, 3755–3762. doi: 10.1128/JB.00244-10
Keywords: arsenic islands, arsenite oxidase, AioBA, phosphorus, synteny
Citation: Li H, Li M, Huang Y, Rensing C and Wang G (2013) In silico analysis of bacterial arsenic islands reveals remarkable synteny and functional relatedness between arsenate and phosphate. Front. Microbiol. 4:347. doi: 10.3389/fmicb.2013.00347
Received: 29 July 2013; Accepted: 31 October 2013; Published online: 20 November 2013.
Edited by:
Partha Basu, Duquesne University, USAReviewed by:
Simon Silver, UIC, Cayman IslandsWolfgang Nitschke, Centre National de la Recherche Scientifique, France
Copyright © 2013 Li, Li, Huang, Rensing and Wang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Christopher Rensing, Department of Plant and Environmental Science, University of Copenhagen, Thorvaldsensvej 40, Frederiksberg DK-1871, Denmark e-mail:Y3JlbnNpbmc5NEBnbWFpbC5jb20=;
Gejiao Wang, State Key Laboratory of Agricultural Microbiology, College of Life Sciences and Technology, Huazhong Agricultural University, Shizishan Street No. 1, Wuhan 430070, P. R. of China e-mail:Z2VqaWFvQG1haWwuaHphdS5lZHUuY24=
† Joint first authors.
 Hang Li
Hang Li Mingshun Li1†
Mingshun Li1† Yinyan Huang
Yinyan Huang Christopher Rensing
Christopher Rensing