- Genetic Engineering and Fermentation Technology, Facultad de Ciencias Bioquímicas y Farmacéuticas, Universidad Nacional de Rosario-Conicet, Rosario, Argentina
The efficient production of functional proteins in heterologous hosts is one of the major bases of modern biotechnology. Unfortunately, many genes are difficult to express outside their original context. Due to their apparent “silent” nature, synonymous codon substitutions have long been thought to be trivial. In recent years, this dogma has been refuted by evidence that codon replacement can have a significant impact on gene expression levels and protein folding. In the past decade, considerable advances in the speed and cost of gene synthesis have facilitated the complete redesign of entire gene sequences, dramatically improving the likelihood of high protein expression. This technology significantly impacts the economic feasibility of microbial-based biotechnological processes by, for example, increasing the volumetric productivities of recombinant proteins or facilitating the redesign of novel biosynthetic routes for the production of metabolites. This review discusses the current applications of this technology, particularly those regarding the production of small molecules and industrially relevant recombinant enzymes. Suggestions for future research and potential uses are provided as well.
Introduction
Microorganisms are at the core of the production of pharmaceuticals, industrial enzymes, and fine chemicals. In many cases, heterologous expression of genes is required to meet commercial-level demands of target proteins and/or metabolites. In this context, variation in codon usage is considered as one of the major factors affecting protein expression levels, since the presence of rare codons can reduce the translation rate and induce translation errors with a significant impact on the economics of recombinant microbe-based production processes (Ikemura, 1981; Gustafsson et al., 2004). The generation of massive genome sequencing data and cost-effective custom DNA synthesis are foundational technologies for synthetic biology, an emerging discipline that aims to create novel organisms containing designer genetic circuits for the production of drugs, industrial enzymes, biofuels, and chemicals (Endy, 2005; McDaniel and Weiss, 2005; Heinemann and Panke, 2006; Leonard et al., 2008). These circuits are built from standard biological parts, including vectors, promoters, ribosomal binding sites (RBSs), transcriptional terminators, and other gene expression regulatory elements. These parts were initially borrowed from nature and nowadays engineered, to adapt their performance to a particular application, or combined to create sophisticated devices (Shiue and Prather, 2012).
Over the past decade, synthetic biology has contributed to significantly reduce the cost of many products manufactured in microbial systems where only one gene needs to be over-expressed. In many cases, the production of a target protein can be boosted by several orders of magnitude by replacing a native sequence with its optimized counterpart (Gustafsson et al., 2004, 2012). This seemingly simple adjustment is of remarkable importance, since many of these products are now traded as commodities and thus there is a continuous need to reduce manufacturing costs in order to remain competitive in the global markets (Menzella, 2011). The ambitious next step of synthetic biology is to further reduce the cost and time involved in developing recombinant organisms by using pre-assembled parts that provide stable, predictable protein expression (Dellomonaco et al., 2010; Nielsen and Keasling, 2011).
So far, most of the progress made in synthetic biology was achieved in Escherichia coli, a preferred host for the production of recombinant proteins because it combines fast growth rate, inexpensive fermentation media and well understood genetics (Burgess-Brown et al., 2008; Welch et al., 2009; Menzella, 2011). However, efforts have been recently expanded to other hosts including Streptomyces species (Medema et al., 2011), Corynebacterium glutamicum (Becker and Wittmann, 2012), yeast (Krivoruchko et al., 2011; Siddiqui et al., 2012; Furukawa and Hohmann, 2013), and algae (Wang et al., 2012; Gimpel et al., 2013). This expanded landscape seeks to take advantage of the natural capabilities to synthesize precursors and cofactors required to produce a particular target, exploit secretion abilities, or utilize natural tolerance to over-accumulated metabolites (Zhu et al., 2012). In this review we summarize the current state of the technology for the expression of codon optimized genes in microbial systems. Examples of its application for the production of small molecules and recombinant enzymes of industrial interest are presented, and suggestions for future research and uses are provided.
Gene Design
Choosing a gene for optimal expression requires selection from a large number of sequences. For example, a protein with an average size of 30 kDa may, in theory, be encoded by 10100 possible DNA sequences (Welch et al., 2009). Historically, two approaches have been used for codon optimization. The first, designated “one amino acid-one codon,” uses the most abundant codon of the host to encode all occurrences of a given amino acid in the optimized sequence (Fuglsang, 2003; Gao et al., 2004; Supek and Vlahovicek, 2004; Villalobos et al., 2006; Feng et al., 2010; Marlatt et al., 2010; Wang et al., 2010). This simple strategy, the most popular in the early days of gene synthesis technology, has a major drawback: a strongly transcribed mRNA from a gene with this design will contain a high concentration of a subset of codons, resulting in an imbalance in the tRNA pool, which in turn may reduce growth due to tRNA depletion (Gong et al., 2006; Villalobos et al., 2006).
The second approach, named “codon randomization,” uses translation tables based on the frequency distribution of the codons in an entire genome or a subset of highly expressed genes. These tables attach weights to each codon, thus codons are assigned randomly with a probability given by the weights (Kodumal et al., 2004; Jayaraj et al., 2005; Menzella et al., 2005; Welch et al., 2009; Wang et al., 2010). This strategy was shown to be superior and was quickly adopted by the synthetic biology community. In addition to improving the yield of the desired product, the “codon randomization” strategy offers many further advantages. For example, flexibility in codon selection facilitates gene design by avoiding: (i) repetitive elements that may lead to gene deletions; (ii) internal RBSs, polyadenylation signals, or transcriptional terminators; (iii) secondary mRNA structures (Luisi et al., 2013); and (iv) by facilitating elimination of unwanted restriction sites to assist the assembly of larger constructs (Villalobos et al., 2006). Several large-scale systematic studies describing variations on this strategy have been conducted in recent years to provide data on the effect of sequence variables (Kudla et al., 2009; Welch et al., 2009; Allert et al., 2010).
Besides codon optimization, other parameters need to be considered to design a gene for efficient translation, including the global GC content (Gustafsson, 2009), local context of a given codon (Villalobos et al., 2006), the presence of mRNA sequence motifs (Pertzev and Nicholson, 2006), and the sequence of the region including the first 10 codons (Goodman et al., 2013). Many web-based free softwares, with features ranging from basic to advanced, were created for gene design during the last decade. Examples include: DNA Works (Hoover and Lubkowski, 2002), GeMS (Jayaraj et al., 2005); Optimizer (Puigbo et al., 2007); Synthetic Gene Designer (Wu et al., 2006); and Gene Designer (Villalobos et al., 2006). Currently, the majority of synthetic DNA suppliers (including GenScript, DNA2.0, GeneArt and Genewiz) offer sequence optimization services using proprietary algorithms at no additional cost.
Parts and Vectors
The application of synthetic DNA technology in engineered microorganisms is not restricted to redesigned genes. Classic expression vectors widely used in strain engineering derive from natural sources and were never optimized for robust production. Recently, great interest has arisen in the systematic engineering and standardization of gene expression parts such as promoters, translation initiation signals, transcriptional terminators, selectable markers, and replication origins to allow fast and predictable combination of these elements.
Some applications, such as metabolic engineering, require optimal levels of each enzyme to maximize production. This is typically achieved by modulating gene expression by, for example, varying transcription or translation levels. Synthetic biology can offer collections of promoters and RBSs capable of providing different levels of gene expression for this purpose (Boyle and Silver, 2012; Meng et al., 2013; Vogl et al., 2013). So far, most of the available promoters have been taken from the natural sequences driving the expression of highly expressed genes. Typical examples are the widely used AOX promoter from Pichia pastoris (Tschopp et al., 1987) for yeast and the bacteriophage T7 promoter for E. coli (Studier and Moffatt, 1986), which provide high transcription levels. Nowadays, synthetic promoter libraries for tunable gene expression are available for many industrially relevant microorganisms including E. coli (Wu et al., 2013), P. pastoris (Hartner et al., 2008; Ruth et al., 2010; Vogl et al., 2013), C. glutamicum (Yim et al., 2013), and Bacillus subtilis (Hansen et al., 2009). Likewise, synthetic RBSs can be used to regulate gene expression (Basu et al., 2005; Pfleger et al., 2006). Furthermore a novel method for automatic design of artificial RBSs to control gene expression has been recently described, expanding the toolbox of artificial sequences to be used in custom genetic circuits (Salis et al., 2009).
Despite current efforts, accurate predictions of the response of any given promoter or RBS have often remained elusive. It is possible that unknown interactions among isolated components may significantly affect the optimal level of gene expression needed to achieve a particular flux through a biosynthetic pathway (Keasling, 2012). In a recent work, Kosuri et al. (2013) provided an alternative strategy to screen the behavior of gene expression regulatory elements. They synthesized 12,563 combinations of common promoters and RBSs and simultaneously measured DNA, RNA, and protein levels from the entire library. They found that RNA and protein expression were within twofold of expected levels 80 and 64% of the time, respectively, and that the worst 5% of constructs deviated from prediction by 13-fold on average, which could hinder large-scale genetic engineering projects. This comprehensive study provides a means to test standard part combinations to optimize production of a particular target molecule.
Genes are usually introduced into production microorganisms using plasmid vectors (Figure 1). Synthetic biology provides the means to speed up this process by using designer plasmid vectors, where all the components are synthesized with standard formats to facilitate exchange and testing of parts, as well as the assembly of multi-gene constructs (Leonard et al., 2008; Shetty et al., 2008). Several designs for the construction of synthetic plasmids and for the assembly of parts have been proposed (Menzella et al., 2005, 2007; Reisinger et al., 2006; Shetty et al., 2008; Sarrion-Perdigones et al., 2011). The most popular format among the synthetic biology community was created by Knight and co-workers (Shetty et al., 2011). They proposed the BioBrick standard, where all parts are flanked by a common set of restriction sites that allow the joining, combination, and rapid assembly of genetic parts to create functional gene expression units.
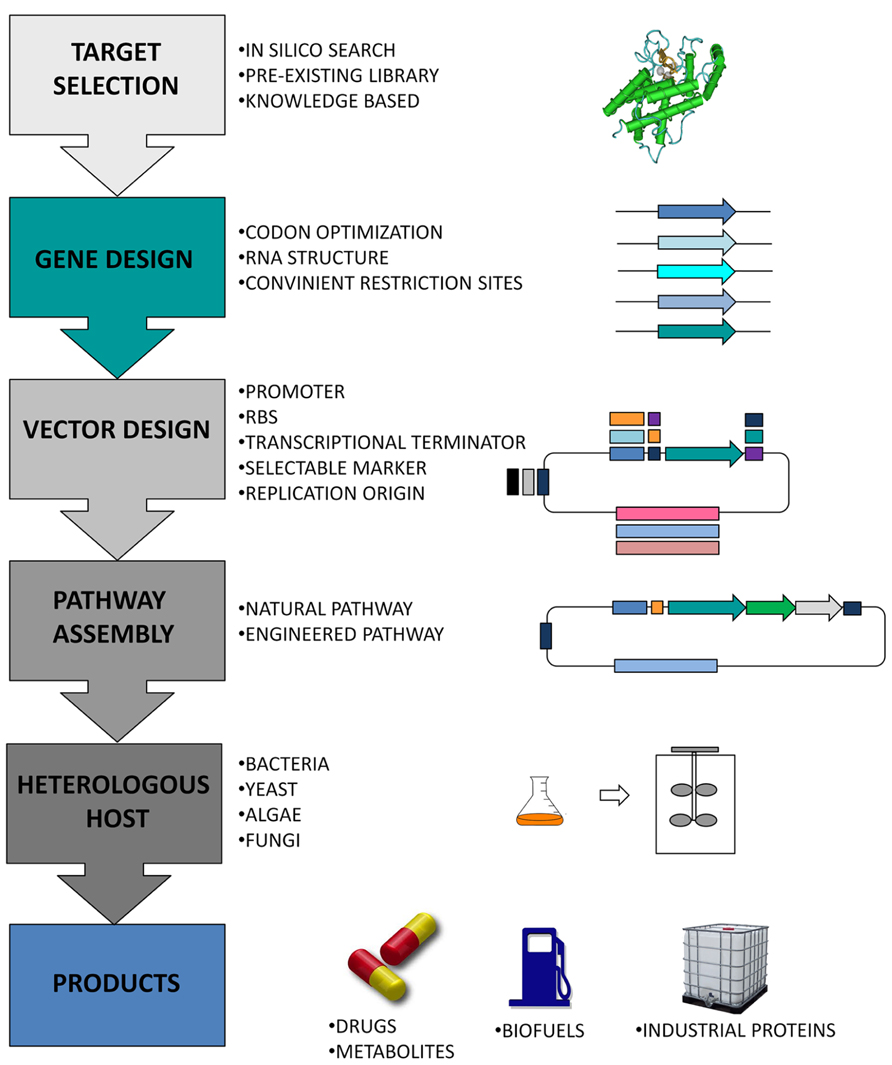
FIGURE 1. Schematic representation of a synthetic biology network applied to the design of novel bio-based parts and devices, as well as to the engineering of existing natural biological systems for the development of target products.
So far, most of the work to create synthetic vectors reported in the literature has been done in E. coli. Recently, we created a plasmid-based platform for the rapid engineering of C. glutamicum, a microorganism of great industrial interest. The approach uses reporter genes to examine and classify promoters and RBSs and permits the easy assembly of operons and genes clusters for co-expression of heterologous genes to facilitate metabolic engineering. Similarly, Constante and co-workers described a platform to engineer eukaryotic hosts by using the BioBrick principle. Interestingly, the system contains a variety of novel parts and implements a recombinase-mediated DNA insertion, allowing chromosomal site-directed exchange of genes in eukaryotic cell lines (Constante et al., 2011).
Practical Applications
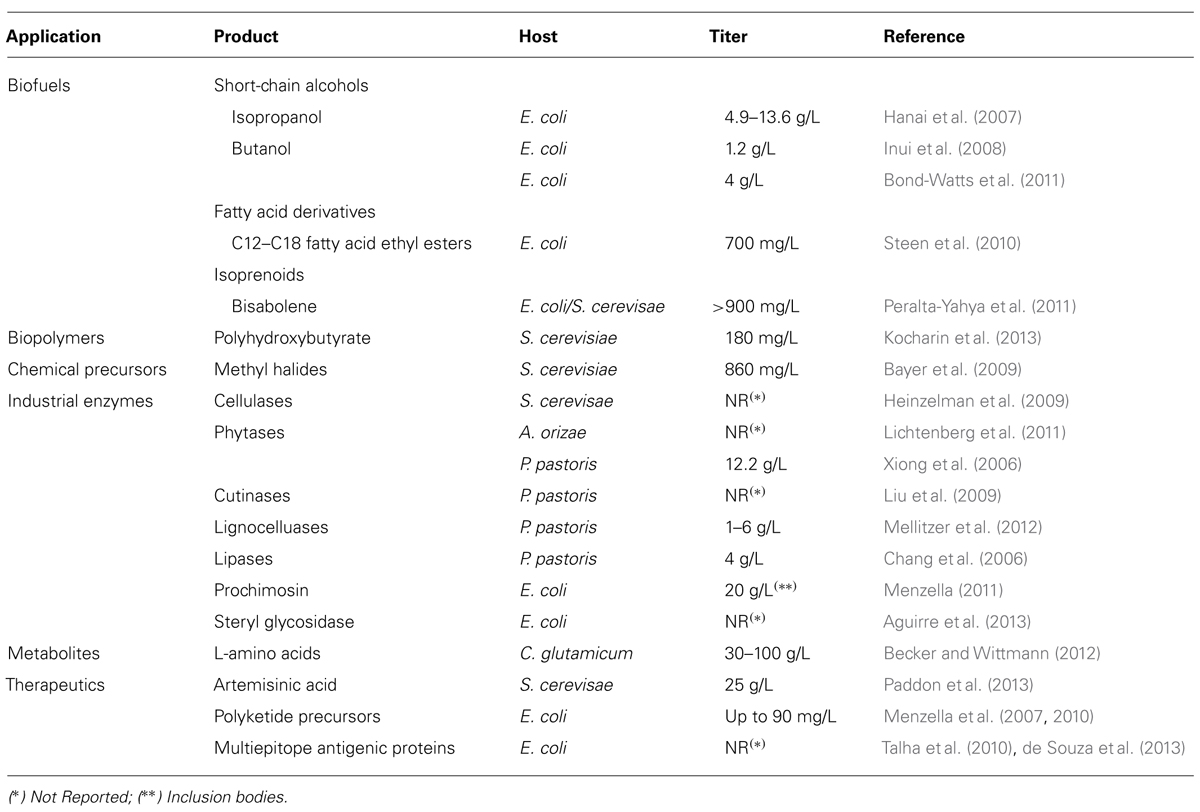
The list of products obtained by the expression of codon optimized genes in microorganisms is constantly growing and includes biofuels, pharmaceuticals, novel bio-based materials and chemicals, industrial enzymes, amino acids, and other metabolites (Table 1).
Production of novel biofuels is one of the most attractive applications for synthetic biology. Fuels like ethanol, biodiesel, butanol, and terpenoid compounds are currently produced using engineered microbes (Table 1). In fact, the main obstacle for the production of these molecules at commercial level is the development of robust microbes and processes (Fischer et al., 2008). Synthetic biology provides tools to achieve optimal expression of pathway genes to ensure the efficient conversion of feedstock materials to target molecules, which is critical to the success of any metabolic engineering strategy. There has been considerable progress recently in the production of different biofuels, and some of the processes have reached promising yields. Hanai and co-workers combined enzymes from Clostridium acetobutylicum (Thl, CtfAB, and ADC), Clostridium beijerinckii (ADH), and E. coli (AtoAD) to assemble a fermentative pathway in E. coli that resulted in production of isopropanol at titers ranging from 4.9 to 13.6 g/L (Hanai et al., 2007). Butanol production was achieved in E. coli using the biosynthetic pathway from C. acetobutylicum and other related clostridial species, reaching titers up to 1.2 g/L (Inui et al., 2008). This was further improved to more than 4 g/L of butanol production by replacing enzymes that are naturally reversible with those that drive the reaction toward butanol formation, expressed from codon optimized genes from different bacterial species (Bond-Watts et al., 2011).
Fatty acid derivatives are other promising biofuel candidates, due to their high energy density and low water solubility. Stenn et al. engineered E. coli to produce C12–C18 fatty acid ethyl esters (FAEEs) directly from glucose at a titer of ~700 mg/L (Steen et al., 2010). Five engineering strategies were combined to achieve this titer, including the elimination of the β-oxidation pathway and the expression of several synthetic genes from different microorganisms. Monoterpene and sesquiterpene hydrocarbons such as limonene, pinene, and farnesene, are isoprenoid compounds with promising fuel applications that have been produced in E. coli and S. cerevisiae. Expression in E. coli of a codon-optimized bisabolene synthase from the fir tree Abies grandis, in conjunction with the introduction of an optimized heterologous mevalonate pathway, resulted in sesquiterpene bisabolene production of 900 mg/L.
A S. cerevisiae strain that overproduces farnesyl pyrophosphate also gave bisabolene titers higher than 900 mg/L using the same bisabolene synthase (Peralta-Yahya et al., 2011). The mevalonate pathway expression was further improved in E. coli by introducing codon-optimized versions of the mevalonate kinase and phosphomevalonate kinase genes after they were identified as potential pathway bottlenecks (Redding-Johanson et al., 2011).
Codon optimized genes have been extensively used to produce pharmaceuticals in microbial platforms. Polyketides are a class of natural products with a high number of well-established clinical applications. The development of a variety of methods for polyketide synthases (PKS) engineering (Menzella and Reeves, 2007; Peiru et al., 2009, 2010) led to a pioneer synthetic biology project conducted at Kosan Biosciences. The goal was to obtain polyketide precursors for the synthesis of novel drugs. First, a generic design for type I PKS genes was created to enable easy assembly and expression of chimeric enzymes (Kodumal et al., 2004; Menzella et al., 2005). The sequences of the synthetic genes were then redesigned with custom made software to optimize codon usage in order to maximize expression in E. coli and provide a standard set of restriction sites to allow combinatorial assembly into unnatural enzymes. Next, more than three million bases of PKS genes were tested to validate the platform. These efforts produced a variety of novel valuable compounds (Menzella et al., 2007; Menzella et al., 2010).
Another remarkable contribution of synthetic biology is the microbial production of artemisinin, a sesquiterpene endoperoxide used to treat malaria (Paddon et al., 2013). This compound is naturally produced by the plant Artemisia annua, but the production of plant-derived artemisinin is expensive; which limits its access to many patients. Recently, Paddon and coworkers engineered strains of S. cerevisiae for production of artemisinic acid, a precursor of artemisinin by fermentation. The simultaneous co-expression of synthetic genes provided an efficient biosynthetic route to artemisinic acid, with fermentation titers of 25 g/L.
Production of proteins for therapeutic use also takes advantage of the use of synthetic genes; a comprehensive review describing progress in this field has been recently published by Mitchell (2011). An elegant synthetic biology approach was used to create designer antigenic proteins for immunoassay-based diagnosis. By designing synthetic genes encoding tandem combinations of epitopes joined by flexible peptide linkers, chimeric proteins were obtained for the detection of antibodies in sera with higher sensitivity and specificity (Talha et al., 2010; de Souza et al., 2013).
The global market for industrial enzymes exceeded $4 billion in 2012 and is therefore an attractive target for cost reduction using synthetic biology tools (Zhou et al., 2004; Menzella, 2011). The use of codon optimized genes allowed notable increases in the production of many enzymes in a variety of hosts, including cellulases in S. cerevisiae (Heinzelman et al., 2009), phytases in Aspergillus oryzae (Lichtenberg et al., 2011), cutinases (Liu et al., 2009), lignocellulases (Mellitzer et al., 2012), and lipases (Chang et al., 2006) in P. pastoris and calf prochymosin in E. coli (Menzella, 2011). In the last example, a strain developed in our laboratory harboring a codon optimized gene produced 70% more prochymosin than that obtained with the wild type sequence, with the concomitant reduction in production costs.
In addition to the contribution to achieve more competitive production processes, synthetic genes provide an attractive alternative for the discovery of enzymes for new applications. For example, in order to search for thermostable enzymes to hydrolyze steryl glucosides (major contaminants of oil-derived biodiesel), we screened a library of archeal genes by retrieving the sequences in silico, synthesizing codon optimized genes for expression in E. coli and assessing their activity against the target. The approach was very successful and resulted in excellent candidates for industrial use (Aguirre et al., 2013). Other products of commercial interest recently obtained from strains carrying codon optimized genes include L-amino acids in C. glutamicum and E. coli (Becker and Wittmann, 2012), and polyhydroxybutyrate and methyl halides in S. cerevisiae (Bayer et al., 2009; Kocharin et al., 2013).
Conclusion and Future Perspectives
The benefits of using codon optimized genes in industrial biotechnology have been extensively demonstrated during the past decade and this technology is being rapidly adopted by strain developers in order to remain competitive in the current market. In the examples presented here, just one or a few synthetic genes need to be introduced into a host to generate novel products or to dramatically reduce the cost of producing existing ones. The cost of synthetic genes has been constantly decreasing during the last decade; and technologies to assemble large fragments of DNA and to make multiple simultaneous changes to wild type genomes are becoming available (Montague et al., 2012). Thus, we can envision a future where custom-made microorganisms can be designed for a particular application (Gibson et al., 2010).
One of the fields where these new technologies can make a dramatic contribution is the production of commodity chemicals in microbes. Initial steps toward this ambitious goal have already been taken by industry. For example, an E. coli strain has been engineered to produce 1,3-propanediol, where in addition to the introduction of the pathway for the production of this target from glycerol, several changes were made in the genome to increase the final yield (Nakamura and Whited, 2003).
Although tremendous progress has been made, in order to fully harness the potential of synthetic biology we need a deeper understanding of the underlying molecular principles of living systems and further development of bioinformatic tools to assist in the modeling of synthetic genomes behavior. These advances are expected to arrive from the interactions among many scientific disciplines.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The Associate Editor declares that despite being affiliated to the same institution as the authors, the review process was handled objectively and no conflict of interest exists.
References
Aguirre, A., Peiru, S., Eberhardt, F., Vetcher, L., Cabrera, R., and Menzella, H. G. (2013). Enzymatic hydrolysis of steryl glucosides, major contaminants of vegetable oil-derived biodiesel. Appl. Microbiol. Biotechnol. doi: 10.1007/s00253-013-5345-4 [Epub ahead of print].
Allert, M., Cox, J. C., and Hellinga, H. W. (2010). Multifactorial determinants of protein expression in prokaryotic open reading frames. J. Mol. Biol. 402, 905–918. doi: 10.1016/j.jmb.2010.08.010
Basu, S., Gerchman, Y., Collins, C. H., Arnold, F. H., and Weiss, R. (2005). A synthetic multicellular system for programmed pattern formation. Nature 434, 1130–1134. doi: 10.1038/nature03461
Bayer, T. S., Widmaier, D. M., Temme, K., Mirsky, E. A., Santi, D. V., and Voigt, C. A. (2009). Synthesis of methyl halides from biomass using engineered microbes. J. Am. Chem. Soc. 131, 6508–6515. doi: 10.1021/ja809461u
Becker, J., and Wittmann, C. (2012). Bio-based production of chemicals, materials and fuels-Corynebacterium glutamicum as versatile cell factory. Curr. Opin. Biotechnol. 23, 631–640. doi: 10.1016/j.copbio.2011.11.012
Bond-Watts, B. B., Bellerose, R. J., and Chang, M. C. (2011). Enzyme mechanism as a kinetic control element for designing synthetic biofuel pathways. Nat. Chem. Biol. 7, 222–227. doi: 10.1038/nchembio.537
Boyle, P. M., and Silver, P. A. (2012). Parts plus pipes: synthetic biology approaches to metabolic engineering. Metab. Eng. 14, 223–232. doi: 10.1016/j.ymben.2011.10.003
Burgess-Brown, N. A., Sharma, S., Sobott, F., Loenarz, C., Oppermann, U., and Gileadi, O. (2008). Codon optimization can improve expression of human genes in Escherichia coli: a multi-gene study. Protein Expr. Purif. 59, 94–102. doi: 10.1016/j.pep.2008.01.008
Constante, M., Grunberg, R., and Isalan, M. (2011). A biobrick library for cloning custom eukaryotic plasmids. PLoS ONE 6:e23685. doi: 10.1371/journal.pone.0023685
Chang, S. W., Lee, G. C., and Shaw, J. F. (2006). Codon optimization of Candida rugosa lip1 gene for improving expression in Pichia pastoris and biochemical characterization of the purified recombinant LIP1 lipase. J. Agric. Food Chem. 54, 815–822. doi: 10.1021/jf052183k
Dellomonaco, C., Fava, F., and Gonzalez, R. (2010). The path to next generation biofuels: successes and challenges in the era of synthetic biology. Microb. Cell Fact. 9, 3. doi: 10.1186/1475-2859-9-3
de Souza, M. Q., Galdino, A. S., Dos Santos, J. C., Soares, M. V., De Nobrega, Y. C., Alvares Ada, C., et al. (2013). A recombinant multiepitope protein for hepatitis B diagnosis. Biomed. Res. Int. 2013, 148317. doi: 10.1155/2013/148317
Feng, Z., Lanwei. Z., Han, X., and Zhang, Y. (2010). Codon optimization of the calf prochymosin gene and its expression in Kluyveromyces lactis. World J. Microbiol. Biotechnol. 26, 895–901. doi: 10.1007/s11274-009-0249-2
Fischer, C. R., Klein-Marcuschamer, D., and Stephanopoulos, G. (2008). Selection and optimization of microbial hosts for biofuels production. Metab. Eng. 10, 295–304. doi: 10.1016/j.ymben.2008.06.009
Fuglsang, A. (2003). Codon optimizer: a freeware tool for codon optimization. Protein Expr. Purif. 31, 247–249. doi: 10.1016/S1046-5928(03)00213-4
Furukawa, K., and Hohmann, S. (2013). Synthetic biology: lessons from engineering yeast MAPK signalling pathways. Mol. Microbiol. 88, 5–19. doi: 10.1111/mmi.12174
Gao, W., Rzewski, A., Sun, H., Robbins, P. D., and Gambotto, A. (2004). UpGene: application of a web-based DNA codon optimization algorithm. Biotechnol. Prog. 20, 443–448. doi: 10.1021/bp0300467
Gibson, D. G., Glass, J. I., Lartigue, C., Noskov, V. N., Chuang, R. Y., Algire, M. A., et al. (2010). Creation of a bacterial cell controlled by a chemically synthesized genome. Science 329, 52–56. doi: 10.1126/science.1190719
Gimpel, J. A., Specht, E. A., Georgianna, D. R., and Mayfield, S. P. (2013). Advances in microalgae engineering and synthetic biology applications for biofuel production. Curr. Opin. Chem. Biol. 17, 489–495. doi: 10.1016/j.cbpa.2013. 03.038
Gong, M., Gong, F., and Yanofsky, C. (2006). Overexpression of tnaC of Escherichia coli inhibits growth by depleting tRNA2Pro availability. J. Bacteriol. 188, 1892–1898. doi: 10.1128/JB.188.5.1892-1898.2006
Goodman, D. B., Church, G. M., and Kosuri, S. (2013). Causes and effects of N-terminal codon bias in bacterial genes. Science 342, 475–479. doi: 10.1126/science.1241934
Gustafsson, C. (2009). “Tools designed to regulate translational efficiency,” in The Metabolic Pathway Engineering Handbook: Tools and Applications, ed. C. Smolke (Boca Raton, FL: CRC Press), 678.
Gustafsson, C., Govindarajan, S., and Minshull, J. (2004). Codon bias and heterologous protein expression. Trends Biotechnol. 22, 346–353. doi: 10.1016/j.tibtech.2004.04.006
Gustafsson, C., Minshull, J., Govindarajan, S., Ness, J., Villalobos, A., and Welch, M. (2012). Engineering genes for predictable protein expression. Protein Expr. Purif. 83, 37–46. doi: 10.1016/j.pep.2012.02.013
Hanai, T., Atsumi, S., and Liao, J. C. (2007). Engineered synthetic pathway for isopropanol production in Escherichia coli. Appl. Environ. Microbiol. 73, 7814–7818. doi: 10.1128/AEM.01140-07
Hansen, M. E., Wangari, R., Hansen, E. B., Mijakovic, I., and Jensen, P. R. (2009). Engineering of Bacillus subtilis 168 for increased nisin resistance. Appl. Environ. Microbiol. 75, 6688–6695. doi: 10.1128/AEM.00943-09
Hartner, F. S., Ruth, C., Langenegger, D., Johnson, S. N., Hyka, P., Lin-Cereghino, G. P., et al. (2008). Promoter library designed for fine-tuned gene expression in Pichia pastoris. Nucleic Acids Res. 36, e76. doi: 10.1093/nar/gkn369
Heinemann, M., and Panke, S. (2006). Synthetic biology – putting engineering into biology. Bioinformatics 22, 2790–2799. doi: 10.1093/bioinformatics/btl469
Heinzelman, P., Snow, C. D., Wu, I., Nguyen, C., Villalobos, A., Govindarajan, S., et al. (2009). A family of thermostable fungal cellulases created by structure-guided recombination. Proc. Natl. Acad. Sci. U.S.A. 106, 5610–5615. doi: 10.1073/pnas.0901417106
Hoover, D. M., and Lubkowski, J. (2002). DNAWorks: an automated method for designing oligonucleotides for PCR-based gene synthesis. Nucleic Acids Res. 30, e43. doi: 10.1093/nar/30.10.e43
Ikemura, T. (1981). Correlation between the abundance of Escherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes. J. Mol. Biol. 146, 1–21. doi: 10.1016/0022-2836(81)90363-6
Inui, M., Suda, M., Kimura, S., Yasuda, K., Suzuki, H., Toda, H., et al. (2008). Expression of Clostridium acetobutylicum butanol synthetic genes in Escherichia coli. Appl. Microbiol. Biotechnol. 77, 1305–1316. doi: 10.1007/s00253-007-1257-5
Jayaraj, S., Reid, R., and Santi, D. V. (2005). GeMS: an advanced software package for designing synthetic genes. Nucleic Acids Res. 33, 3011–3016. doi: 10.1093/nar/gki614
Keasling, J. D. (2012). Synthetic biology and the development of tools for metabolic engineering. Metab. Eng. 14, 189–195. doi: 10.1016/j.ymben.2012.01.004
Kocharin, K., Siewers, V., and Nielsen, J. (2013). Improved polyhydroxybutyrate production by Saccharomyces cerevisiae through the use of the phosphoketolase pathway. Biotechnol. Bioeng. 110, 2216–2224. doi: 10.1002/bit.24888
Kodumal, S. J., Patel, K. G., Reid, R., Menzella, H. G., Welch, M., and Santi, D. V. (2004). Total synthesis of long DNA sequences: synthesis of a contiguous 32-kb polyketide synthase gene cluster. Proc. Natl. Acad. Sci. U.S.A. 101, 15573–15578. doi: 10.1073/pnas.0406911101
Kosuri, S., Goodman, D. B., Cambray, G., Mutalik, V. K., Gao, Y., Arkin, A. P., et al. (2013). Composability of regulatory sequences controlling transcription and translation in Escherichia coli. Proc. Natl. Acad. Sci. U.S.A. 110, 14024–14029. doi: 10.1073/pnas.1301301110
Krivoruchko, A., Siewers, V., and Nielsen, J. (2011). Opportunities for yeast metabolic engineering: lessons from synthetic biology. Biotechnol. J. 6, 262–276. doi: 10.1002/biot.201000308
Kudla, G., Murray, A. W., Tollervey, D., and Plotkin, J. B. (2009). Coding-sequence determinants of gene expression in Escherichia coli. Science 324, 255–258. doi: 10.1126/science.1170160
Leonard, E., Nielsen, D., Solomon, K., and Prather, K. J. (2008). Engineering microbes with synthetic biology frameworks. Trends Biotechnol. 26, 674–681. doi: 10.1016/j.tibtech.2008.08.003
Lichtenberg, J., Pedersen, P. B., Elvig-Joergensen, S. G., Skov, L. K., Olsen, C. L., and Glitsoe, L. V. (2011). Toxicological studies on a novel phytase expressed from synthetic genes in Aspergillus oryzae. Regul. Toxicol. Pharmacol. 60, 401–410. doi: 10.1016/j.yrtph.2011.05.010
Liu, Z., Gosser, Y., Baker, P. J., Ravee, Y., Lu, Z., Alemu, G., et al. (2009). Structural and functional studies of Aspergillus oryzae cutinase: enhanced thermostability and hydrolytic activity of synthetic ester and polyester degradation. J. Am. Chem. Soc. 131, 15711–15716. doi: 10.1021/ja9046697
Luisi, I., Pavan, S., Fontanive, G., Tossi, A., Benedetti, F., Savoini, A., et al. (2013). An albumin-derived peptide scaffold capable of binding and catalysis. PLoS ONE 8:e56469. doi: 10.1371/journal.pone.0056469
Marlatt, N. M., Spratt, D. E., and Shaw, G. S. (2010). Codon optimization for enhanced Escherichia coli expression of human S100A11 and S100A1 proteins. Protein Expr. Purif. 73, 58–64. doi: 10.1016/j.pep.2010.03.015
McDaniel, R., and Weiss, R. (2005). Advances in synthetic biology: on the path from prototypes to applications. Curr. Opin. Biotechnol. 16, 476–483. doi: 10.1016/j.copbio.2005.07.002
Medema, M. H., Breitling, R., and Takano, E. (2011). Synthetic biology in Streptomyces bacteria. Methods Enzymol. 497, 485–502. doi: 10.1016/B978-0-12-385075-1.00021-4
Mellitzer, A., Weis, R., Glieder, A., and Flicker, K. (2012). Expression of lignocellulolytic enzymes in Pichia pastoris. Microb. Cell Fact. 11, 61. doi: 10.1186/1475-2859-11-61
Meng, H., Wang, J., Xiong, Z., Xu, F., Zhao, G., and Wang, Y. (2013). Quantitative design of regulatory elements based on high-precision strength prediction using artificial neural network. PLoS ONE 8:e60288. doi: 10.1371/journal.pone.0060288
Menzella, H. G. (2011). Comparison of two codon optimization strategies to enhance recombinant protein production in Escherichia coli. Microb. Cell Fact. 10, 15. doi: 10.1186/1475-2859-10-15
Menzella, H. G., Carney, J. R., Li, Y., and Santi, D. V. (2010). Using chemobiosynthesis and synthetic mini-polyketide synthases to produce pharmaceutical intermediates in Escherichia coli. Appl. Environ. Microbiol. 76, 5221–5227. doi: 10.1128/AEM.02961-09
Menzella, H. G., Carney, J. R., and Santi, D. V. (2007). Rational design and assembly of synthetic trimodular polyketide synthases. Chem. Biol. 14, 143–151. doi: 10.1016/j.chembiol.2006.12.002
Menzella, H. G., and Reeves, C. D. (2007). Combinatorial biosynthesis for drug development. Curr. Opin. Microbiol. 10, 238–245. doi: 10.1016/j.mib.2007.05.005
Menzella, H. G., Reid, R., Carney, J. R., Chandran, S. S., Reisinger, S. J., Patel, K. G., et al. (2005). Combinatorial polyketide biosynthesis by de novo design and rearrangement of modular polyketide synthase genes. Nat. Biotechnol. 23, 1171–1176. doi: 10.1038/nbt1128
Mitchell, W. (2011). Natural products from synthetic biology. Curr. Opin. Chem. Biol. 15, 505–515. doi: 10.1016/j.cbpa.2011.05.017
Montague, M. G., Lartigue, C., and Vashee, S. (2012). Synthetic genomics: potential and limitations. Curr. Opin. Biotechnol. 23, 659–665. doi: 10.1016/j.copbio.2012.01.014
Nakamura, C. E., and Whited, G. M. (2003). Metabolic engineering for the microbial production of 1,3-propanediol. Curr. Opin. Biotechnol. 14, 454–459. doi: 10.1016/j.copbio.2003.08.005
Nielsen, J., and Keasling, J. D. (2011). Synergies between synthetic biology and metabolic engineering. Nat. Biotechnol. 29, 693–695. doi: 10.1038/nbt.1937
Paddon, C. J., Westfall, P. J., Pitera, D. J., Benjamin, K., Fisher, K., Mcphee, D., et al. (2013). High-level semi-synthetic production of the potent antimalarial artemisinin. Nature 496, 528–532. doi: 10.1038/nature12051
Peiru, S., Gramajo, H., and Menzella, H. G. (2009). Design and synthesis of pathway genes for polyketide biosynthesis. Methods Enzymol. 459, 319–337. doi: 10.1016/S0076-6879(09)04614-X
Peiru, S., Gramajo, H. C., and Menzella, H. G. (2010). Recombinant approaches to large polyketide molecules as potential drugs. Drug Discov. Today Technol. 7, e95–e146. doi: 10.1016/j.ddtec.2010.10.002
Peralta-Yahya, P. P., Ouellet, M., Chan, R., Mukhopadhyay, A., Keasling, J. D., and Lee, T. S. (2011). Identification and microbial production of a terpene-based advanced biofuel. Nat. Commun. 2, 483. doi: 10.1038/ncomms1494
Pertzev, A. V., and Nicholson, A. W. (2006). Characterization of RNA sequence determinants and antideterminants of processing reactivity for a minimal substrate of Escherichia coli ribonuclease III. Nucleic Acids Res. 34, 3708–3721. doi: 10.1093/nar/gkl459
Pfleger, B. F., Pitera, D. J., Smolke, C. D., and Keasling, J. D. (2006). Combinatorial engineering of intergenic regions in operons tunes expression of multiple genes. Nat. Biotechnol. 24, 1027–1032. doi: 10.1038/nbt1226
Puigbo, P., Guzman, E., Romeu, A., and Garcia-Vallve, S. (2007). OPTIMIZER: a web server for optimizing the codon usage of DNA sequences. Nucleic Acids Res. 35, W126–W131. doi: 10.1093/nar/gkm219
Redding-Johanson, A. M., Batth, T. S., Chan, R., Krupa, R., Szmidt, H. L., Adams, P. D., et al. (2011). Targeted proteomics for metabolic pathway optimization: application to terpene production. Metab. Eng. 13, 194–203. doi: 10.1016/j.ymben.2010.12.005
Reisinger, S. J., Patel, K. G., and Santi, D. V. (2006). Total synthesis of multi-kilobase DNA sequences from oligonucleotides. Nat. Protoc. 1, 2596–2603. doi: 10.1038/nprot.2006.426
Ruth, C., Zuellig, T., Mellitzer, A., Weis, R., Looser, V., Kovar, K., et al. (2010). Variable production windows for porcine trypsinogen employing synthetic inducible promoter variants in Pichia pastoris. Syst. Synth. Biol. 4, 181–191. doi: 10.1007/s11693-010-9057-0
Salis, H. M., Mirsky, E. A., and Voigt, C. A. (2009). Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 27, 946–950. doi: 10.1038/nbt.1568
Sarrion-Perdigones, A., Falconi, E. E., Zandalinas, S. I., Juarez, P., Fernandez-Del-Carmen, A., Granell, A., et al. (2011). GoldenBraid: an iterative cloning system for standardized assembly of reusable genetic modules. PLoS ONE 6:e21622. doi: 10.1371/journal.pone.0021622
Shetty, R., Lizarazo, M., Rettberg, R., and Knight, T. F. (2011). Assembly of BioBrick standard biological parts using three antibiotic assembly. Methods Enzymol. 498, 311–326. doi: 10.1016/B978-0-12-385120-8.00013-9
Shetty, R. P., Endy, D., and Knight, T. F. Jr. (2008). Engineering BioBrick vectors from BioBrick parts. J. Biol. Eng. 2, 5. doi: 10.1186/1754-1611-2-5
Shiue, E., and Prather, K. L. J. (2012). Synthetic biology devices as tools for metabolic engineering. Biochem. Eng. J. 65, 82–89. doi: 10.1016/j.bej.2012.04.006
Siddiqui, M. S., Thodey, K., Trenchard, I., and Smolke, C. D. (2012). Advancing secondary metabolite biosynthesis in yeast with synthetic biology tools. FEMS Yeast Res. 12, 144–170. doi: 10.1111/j.1567-1364.2011.00774.x
Steen, E. J., Kang, Y., Bokinsky, G., Hu, Z., Schirmer, A., Mcclure, A., et al. (2010). Microbial production of fatty-acid-derived fuels and chemicals from plant biomass. Nature 463, 559–562. doi: 10.1038/nature08721
Studier, F. W., and Moffatt, B. A. (1986). Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J. Mol. Biol. 189, 113–130. doi: 10.1016/0022-2836(86)90385-2
Supek, F., and Vlahovicek, K. (2004). INCA: synonymous codon usage analysis and clustering by means of self-organizing map. Bioinformatics 20, 2329–2330. doi: 10.1093/bioinformatics/bth238
Talha, S. M., Salminen, T., Chugh, D. A., Swaminathan, S., Soukka, T., Pettersson, K., et al. (2010). Inexpensive designer antigen for anti-HIV antibody detection with high sensitivity and specificity. Clin. Vaccine Immunol. 17, 335–341. doi: 10.1128/CVI.00283-09
Tschopp, J. F., Brust, P. F., Cregg, J. M., Stillman, C. A., and Gingeras, T. R. (1987). Expression of the lacZ gene from two methanol-regulated promoters in Pichia pastoris. Nucleic Acids Res. 15, 3859–3876. doi: 10.1093/nar/15.9.3859
Villalobos, A., Ness, J. E., Gustafsson, C., Minshull, J., and Govindarajan, S. (2006). Gene designer: a synthetic biology tool for constructing artificial DNA segments. BMC Bioinformatics 7:285. doi: 10.1186/1471-2105-7-285
Vogl, T., Ruth, C., Pitzer, J., Kickenweiz, T., and Glieder, A. (2013). Synthetic core promoters for Pichia pastoris. ACS Synth. Biol. doi: 10.1021/sb400091p [Epub ahead of print].
Wang, B., Wang, J., Zhang, W., and Meldrum, D. R. (2012). Application of synthetic biology in cyanobacteria and algae. Front. Microbiol. 3:344. doi: 10.3389/fmicb.2012.00344
Wang, X., Li, X., Zhang, Z., Shen, X., and Zhong, F. (2010). Codon optimization enhances secretory expression of Pseudomonas aeruginosa exotoxin A in E. coli. Protein Expr. Purif. 72, 101–106. doi: 10.1016/j.pep.2010.02.011
Welch, M., Govindarajan, S., Ness, J. E., Villalobos, A., Gurney, A., Minshull, J., et al. (2009). Design parameters to control synthetic gene expression in Escherichia coli. PLoS ONE 4:e7002. doi: 10.1371/journal.pone.0007002
Wu, G., Bashir-Bello, N., and Freeland, S. J. (2006). The synthetic gene designer: a flexible web platform to explore sequence manipulation for heterologous expression. Protein Expr. Purif. 47, 441–445. doi: 10.1016/j.pep.2005.10.020
Wu, Y., Zhang, Y., Tu, R., Liu, H., and Wang, Q. (2013). Construction of synthetic promoters for Escherichia coli and application in the biosynthesis of cis, cis-muconic acid. Sheng Wu Gong Cheng Xue Bao 29, 760–771.
Xiong, A. S., Yao, Q. H., Peng, R. H., Zhang, Z., Xu, F., Liu, J. G., et al. (2006). High level expression of a synthetic gene encoding Peniophora lycii phytase in methylotrophic yeast Pichia pastoris. Appl. Microbiol. Biotechnol. 72, 1039–1047. doi: 10.1007/s00253-006-0384-8
Yim, S. S., An, S. J., Kang, M., Lee, J., and Jeong, K. J. (2013). Isolation of fully synthetic promoters for high-level gene expression in Corynebacterium glutamicum. Biotechnol. Bioeng. 110, 2959–2969. doi: 10.1002/bit.24954
Zhou, Z., Schnake, P., Xiao, L., and Lal, A. A. (2004). Enhanced expression of a recombinant malaria candidate vaccine in Escherichia coli by codon optimization. Protein Expr. Purif. 34, 87–94. doi: 10.1016/j.pep.2003.11.006
Keywords: synthetic biology, gene design, codon optimization, strain engineering, microbial systems
Citation: Elena C, Ravasi P, Castelli ME, Peirú S and Menzella HG (2014) Expression of codon optimized genes in microbial systems: current industrial applications and perspectives. Front. Microbiol. 5:21. doi: 10.3389/fmicb.2014.00021
Received: 10 December 2013; Paper pending published: 17 December 2013;
Accepted: 14 January 2014; Published online: 04 February 2014.
Edited by:
Eduardo A. Ceccarelli, Universidad Nacional de Rosario, ArgentinaReviewed by:
Blaine Pfeifer, The State University of New York at Buffalo, USAChristopher Desmond Reeves, Amyris, USA
Copyright © 2014 Elena, Ravasi, Castelli, Peirú and Menzella. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Hugo G. Menzella, Genetic Engineering and Fermentation Technology, Facultad de Ciencias Bioquímicas y Farmacéuticas, Universidad Nacional de Rosario-Conicet, Suipacha 590, Rosario 2000, Argentina e-mail:aG1lbnplbGxhQGZiaW95Zi51bnIuZWR1LmFy
 Claudia Elena
Claudia Elena Pablo Ravasi
Pablo Ravasi María E. Castelli
María E. Castelli