- 1Department of Microbiology and Immunology, McGill University, Montreal, QC, Canada
- 2Department of Medicine, McGill University, Montreal, QC, Canada
- 3McGill International TB Centre, Montreal, QC, Canada
The genus Mycobacterium is comprised of more than 150 species that reside in a wide variety of habitats. Most mycobacteria are environmental organisms that are either not associated with disease or are opportunistic pathogens that cause non-transmissible disease in immunocompromised individuals. In contrast, a small number of species, such as the tubercle bacillus, Mycobacterium tuberculosis, are host-adapted pathogens for which there is no known environmental reservoir. In recent years, gene disruption studies using the host-adapted pathogen have uncovered a number of “virulence factors,” yet genomic data indicate that many of these elements are present in non-pathogenic mycobacteria. This suggests that much of the genetic make-up that enables virulence in the host-adapted pathogen is already present in environmental members of the genus. In addition to these generic factors, we hypothesize that molecules elaborated exclusively by professional pathogens may be particularly implicated in the ability of M. tuberculosis to infect, persist, and cause transmissible pathology in its host species, Homo sapiens. One approach to identify these molecules is to employ comparative analysis of mycobacterial genomes, to define evolutionary events such as horizontal gene transfer (HGT) that contributed M. tuberculosis-specific genetic elements. Independent studies have now revealed the presence of HGT genes in the M. tuberculosis genome and their role in the pathogenesis of disease is the subject of ongoing investigations. Here we review these studies, focusing on the hypothesized role played by HGT loci in the emergence of M. tuberculosis from a related environmental species into a highly specialized human-adapted pathogen.
Introduction
Through modification of their genome content, bacteria can evolve to exploit different ecological niches. While vertical events such as gene duplication, chromosomal rearrangement and gene decay can affect the shape and structure of a genome (Ventura et al., 2007), horizontal gene transfer (HGT) is an important mechanism for bacteria to acquire novel genetic material into their genomes (Lercher and Pal, 2008; Price et al., 2008), subsequently facilitating adaptation and diversification (Treangen and Rocha, 2011). HGT can be mediated by transformation (acquisition of naked DNA), transduction (DNA transfer via a bacteriophage), and conjugation (fusion of two bacterial cells enabling a unidirectional transfer of a plasmid or mobile element; Frost et al., 2005). HGT has been shown to profoundly impact the prokaryotic genome plasticity, allowing the acquisition of antibiotic resistance elements (Mohd-Zain et al., 2004; Palmer et al., 2010; Gray et al., 2013), virulence genes (Rosas-Magallanes et al., 2006; Li et al., 2012) and new metabolic pathways (Wilson et al., 2003; Baldwin et al., 2004; Chouikha et al., 2006; Noda-Garcia et al., 2013).
Mycobacterium, a genus of Actinobacteria, comprises mostly non-pathogenic species. Exceptionally, this genus contains a number of host-adapted pathogens, including the leprosy bacillus, Mycobacterium leprae, and the Johne’s bacillus, M. avium subspecies paratuberculosis, the latter defined by the presence of at least six genomic islands that were likely acquired by HGT (Alexander et al., 2009). In this review, we focus on the Mycobacterium tuberculosis complex (MTBC), agents of tuberculosis (TB) in their respective mammalian hosts. Among the various subspecies of the MTBC, M. tuberculosis sensu stricto is the cause of human TB, which infects over 2 billion people and causes an estimated 1.3 million deaths annually (World Health Organization, 2013).
When contrasting the genome of MTBC organisms with the most closely related environmental mycobacteria, M. marinum and M. kansasii, independent studies have identified M. tuberculosis-specific genetic factors putatively acquired by HGT (Becq et al., 2007; Veyrier et al., 2009; Supply et al., 2013), evidenced by the presence of clustering, vehicles of HGT (phage, transposons, toxin-antitoxin genes) and an aberrant GC content in their DNA (Zaneveld et al., 2008). For example, the M. tuberculosis genome codes for 55 proteins absent from M. kansasii, M. marinum and all other sequenced mycobacterial genomes (Veyrier et al., 2009). As 87% of these M. tuberculosis-specific genes are found in clusters, it has been postulated that these clusters may be pathogenicity islands that contribute to the unique virulence of M. tuberculosis (Hacker et al., 1997; Veyrier et al., 2009). As several of these M. tuberculosis-specific genes have been linked to host adaptation (Sassetti and Rubin, 2003; Pethe et al., 2004), this provides further support for the notion that HGT may have played a crucial role in the emergence of this pathogen. At the ecological level, M. tuberculosis uses humans as its sole known reservoir while environmental mycobacteria such as M. kansasii can be found in various aquatic habitats (McSwiggan and Collins, 1974; Steadham, 1980; Sartori et al., 2013; Thomson et al., 2013), further highlighting the impact of genome remodeling on bacterial biology.
In this review, we briefly describe the early interplay between M. tuberculosis and the host during an infection, followed by bioinformatic data supporting the evidence for HGT and its potential contribution to the host-adapted lifestyle of this pathogen. To illustrate the relationships between various mycobacteria including the species discussed in this manuscript, in Figure 1 we present an un-rooted phylogenic tree based on 20 randomly selected genes conserved across eight mycobacteria, including seven slow-growing species (M. tuberculosis, M. canetti, M. kansasii, M. marinum, M. ulcerans, M. avium subsp. hominssuis, M. avium subsp. paratuberculosis) and a rapid-growing species (M. smegmatis) as the out-group. It is worth noting that the topology of this independently generated tree is congruent with the tree built from housekeeping genes (Veyrier et al., 2009), providing additional support for the evolutionary relationships between these species. The genes used for tree generation are provided in the figure legend. The organization of each M. tuberculosis-specific locus discussed is illustrated in Figure 2, in comparison with the flanking genomic regions in the related organisms M. kansasii and M. marinum.
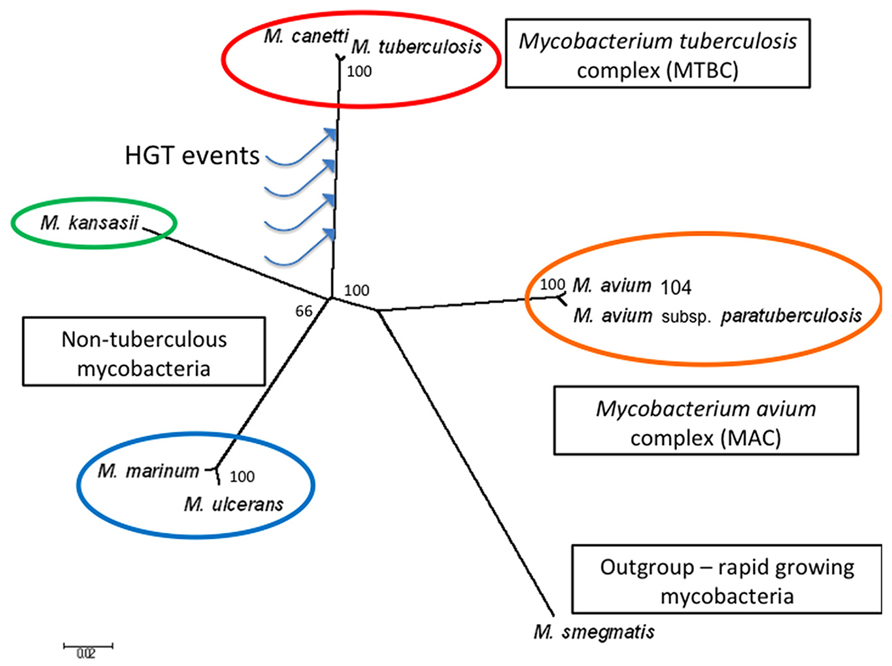
FIGURE 1. Phylogeny of M. tuberculosis and closely related Mycobacterium species. The un-rooted phylogenetic tree was generated by MEGA6.0 using 20 randomly selected genes conserved across eight Mycobacterium species (Schwab et al., 2009). The blue arrows schematically represent where putative HGT events may have occurred, resulting in M. tuberculosis-specific genomic islands. The scale bar indicates 0.02 substitutions per nucleotide position, and the bootstrap values calculated using the neighbor-joining method (expressed as a percentage of 1000 replicates) are shown at the branch points. The fast growing species M. smegmatis is used as the out-group. Genes used are listed below (represented as M. tuberculosis genes): Rv0001-dnaA, Rv0041-leuS, Rv0236A–Rv0236A, Rv0248c–Rv0248c, Rv0285-PE5, Rv0287-esxG, Rv0288-esxH, Rv1085c–Rv1085c, Rv0197–Rv0197, Rv1304-atpB, Rv1305-atpE, Rv1894c–Rv1894c, Rv2172c–Rv2172c, Rv2392-cysH, Rv2440c-obg, Rv2477c–Rv2477c, Rv3019c-esxR, Rv3045-adhC, Rv3392c-cmaA1, Rv3502c- hsd4A.
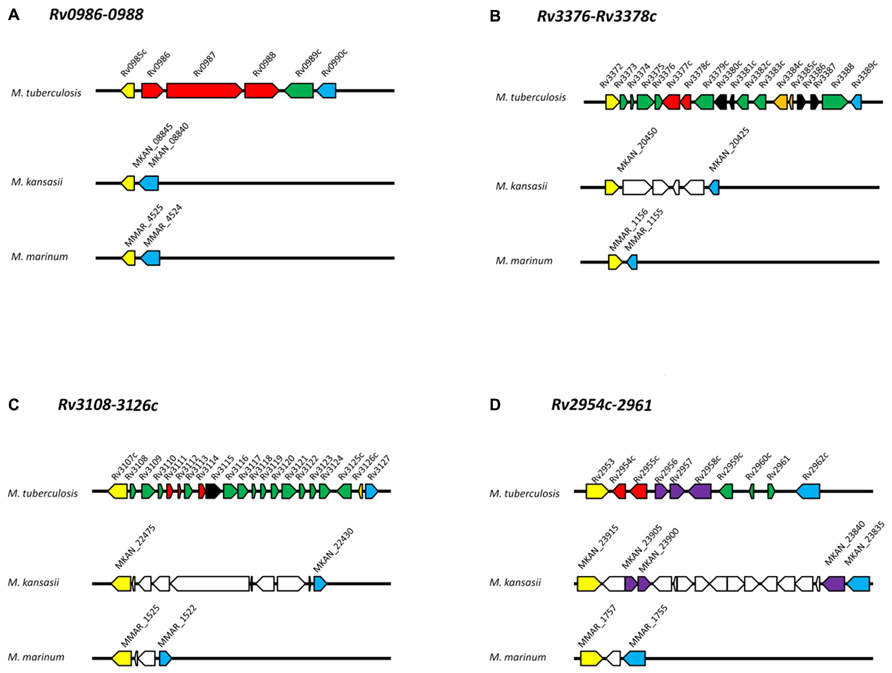
FIGURE 2. Genomic organization of operons in M. tuberculosis, M. kansasii, and M. marinum. (A) Rv0986-0988; (B) Rv3376-3378c; (C) Rv3108-3126c; (D) Rv2954c-2961. Protein-coding genes are represented by arrows and orthologous genes are indicated by arrows of the same color. Yellow and blue arrows mark the “boundary” of each M. tuberculosis-specific locus. Red arrows indicate genes discussed in this review. Dark green arrows indicate M. tuberculosis genes with no orthologs within the corresponding M. kansasii and M. marinum genomic regions. White arrows in M. kansasii and M. marinum genomes indicate genes present but not orthologs to M. tuberculosis genes. Black arrows indicate transposases; orange arrows indicate toxin-antitoxin genes. Genome organizations for M. tuberculosis, M. kansasii, M. marinum and gene clusters were obtained from the Kyoto Encyclopedia of Genes and Genomes (http://www.kegg.jp/) based on databases available at the Sanger Institute, Tuberculist, and McGill University, respectively. Orthologs were verified by comparing each predicted protein against the H37Rv genome using the BLAST program. Only proteins with >50% coverage, 60% identity and E-value <e-20 were used.
M. tuberculosis and the Host Environment
When M. tuberculosis enters the pulmonary alveoli via the aerosol route, it is thought to first encounter alveolar macrophages. Following phagocytosis by these macrophages, the bacterium finds itself in the phagosomal compartment which is, among other attributes, iron limiting, carbon poor, hypoxic, nitrosative, and oxidative (Schnappinger et al., 2003). M. tuberculosis is able to resist these bactericidal effects by synthesizing antioxidants, repairing DNA and proteins, maintaining intracellular pH and cell wall integrity (Buchmeier et al., 2000; Master et al., 2002; Boshoff et al., 2003; Darwin and Nathan, 2005; Vandal et al., 2008, 2009; Colangeli et al., 2009). Moreover, M. tuberculosis can cope with hypoxic and other growth-limiting environments using a number of tactics such as activating the dormancy regulon, promoting alternate metabolic pathways and iron metabolism (Leistikow et al., 2010; Marrero et al., 2010; Ryndak et al., 2010; Griffin et al., 2011). In addition, M. tuberculosis is able to prevent the fusion of phago-lysosome (Sun et al., 2010; Wong et al., 2011), permeabilize the phagosomal membrane (Manzanillo et al., 2012), and escape into the cytosol. The cytosol likely offers a less hostile, thus more permissive environment, where the bacteria can replicate and induce the infected macrophages to undergo necrosis instead of apoptosis, a strategy that allows the bacteria to infect neighboring cells, thereby enabling the perpetuation of the infection process (van der Wel et al., 2007; Divangahi et al., 2009; Behar et al., 2010).
Rv0986–0988
In the seminal work that first described evidence of HGT in the M. tuberculsosis genome, Becq et al. (2007) detected a 5.6 kb M. tuberculosis-specific Island with a reduced GC content (53%) compared to the average for the M. tuberculosis genome (65.6%; Cole et al., 1998). Further molecular and in silico analyses demonstrated that this operon is present in other members of the MTBC, including M. bovis, M. africanum, and M. microti (Rosas-Magallanes et al., 2006). Based on phylogenetic analyses, it was proposed that three genes within this locus, Rv0986–8, had been acquired from phylogenetically distant γ-proteobacteria via plasmid transfer. The fact that the orthologs of these three genes are consistently together suggest that one single HGT event occurred during the acquisition of this operon by the ancestor of M. tuberculosis (Rosas-Magallanes et al., 2006).
Rv0986 is predicted to encode an adenosine triphosphate (ATP)-binding protein that is orthologous to the Agrobacterium tumefaciens attE polypeptide, and form an ABC transporter with Rv0987 (Braibant et al., 2000; Rosas-Magallanes et al., 2006). The A. tumefaciens attE gene is located on a plasmid which harbors the attE-H operon. This operon has been proposed to encode an ABC transporter that secretes a host cell adhesion factor (Matthysse et al., 1996, 2000). Intriguingly, the N- and C- terminal sequences of Rv0987 share 40% similarity with attF and attG, and the neighboring Rv0988 shows 50% similarity with attH (Rosas-Magallanes et al., 2006).
The Rv0986–8 operon has been implicated in M. tuberculosis virulence as mutants with disruption in Rv0986 and 0987 exhibit reduced ability to inhibit phagosome acidification in macrophages (Pethe et al., 2004). Furthermore, these mutants had impaired binding to host cells, and this phenotype could be rescued by complementing with a cosmid carrying M. tuberculosis DNA encompassing the Rv0986–8 operon (Rosas-Magallanes et al., 2007). Although Rv0986 and Rv0987 mutants were not shown to be attenuated in mouse lungs and spleens (Rosas-Magallanes et al., 2007), the Rv0986 mutant was subsequently shown to be less virulent in the context of central nervous system infection (Be et al., 2008). Recently Rv0986–8 have been found to be regulated by EspR, a transcription factor that also regulates the ESX-1 secretion system (Blasco et al., 2012), a major virulence mediator of M. tuberculosis (Brodin et al., 2006).
Rv3376–Rv3378c
Another genomic island potentially acquired by HGT is the 3.1 kb region encompassing Rv3376–8c (Becq et al., 2007; Veyrier et al., 2009). This island exhibits a reduced GC content (54.7%) and is associated with the presence of transposases, known to mediate HGT events (Becq et al., 2007; Veyrier et al., 2009). The closest genera harboring such genes are Agrobacterium and Rhizobium (Becq et al., 2007). More recently, Mann and Peters, (2012) speculated that, while Rv3377c amino acid sequence shares homology with proteins from another actinomycete, Micromonospora, Rv3378c has no ortholog in any other organism with the exception of a hypothetical protein in amoeba. These observations suggest that these MTBC-specific genes originated in different sources (Mann and Peters, 2012).
Biochemical characterization has revelated that Rv3377c and Rv3378c encode a halimadienyl diphosphate (HPP) synthase and a diterpene synthase, respectively. In a step-wise fashion, these enzymes catalyze the cyclization of the precursor, geranylgeranyl diphosphate (GGPP), and the hydrolysis of the HPP intermediate to produce isotuberculosinol (isoTB), a diterpene species (Nakano et al., 2005; Mann et al., 2009a, b). Terpenes are one of the most widespread and chemically diverse compounds found in nature. They are hydrocarbons made up of five, or multiples of five, carbon units (Simion, 2005). In plants and fungi, they are commonly found as essential as well as secondary metabolites involved in signaling and defense (Buckingham, 1994). While members of the Actinobcteria group synthesize a plethora of natural products (Baltz, 2008), very few are known to encode diterpene (C20) synthases (Dairi et al., 2001; Hamano et al., 2002; Dürr et al., 2006; Smanski et al., 2011).
During macrophage infection, M. tuberculosis mutants with disruption in either Rv3377c or Rv3378c showed marked defect in arresting phagosome acidification as well as intracellular survival, suggesting that these genes are involved in the modulation of early infection process (Pethe et al., 2004). Intriguingly, these genes are not transcriptionally altered during macrophage infection (Stewart et al., 2005; Rohde et al., 2007; Waddell and Butcher, 2007), implying that the synthesis of isoTB is regulated at the protein level, potentially triggered by ambient magnesium levels (Mann et al., 2009a, 2011). In addition, isoTB has shown to inhibit phagosome acidification by 0.5 pH units as well as proteolytic activity (Mann et al., 2009b). Recently Rv3378c has been characterized as a tuberculosinyl transferase that converts isoTB into the proposed end product, tuberculosinyladenosine (TbAd; Layre et al., 2014). The cellular mechanism by which Rv3377–8c modifies phagosome function thus remains to be further investigated.
Rv3108–3126c
Rv3108–3126c is a 15.1 kb, MTBC-specific genomic island that has a reduced GC content (56.7%) and contains genes encoding insertion sequences and a transposase, features typical of HGT (Becq et al., 2007). The proposed donor species include Burkholderia, Corynebacterium, and Pseudomonas. This island contains two potential virulence genes; Rv3111 and Rv3114.
A transposon mutant of Rv3111 (moaC1) was first shown to be attenuated in replicating in macrophages (Rosas-Magallanes et al., 2007). In a more recent high-throughput genetic screen, mutants disrupted for the genes moaC1/moaD1(Rv3112), implicated in molybdenum cofactor biosynthesis, were found to be trafficked to acidified intracellular compartments rapidly (Brodin et al., 2010), potentially providing an explanation for the impaired intracellular growth. Molybdopterin is the main building block of the molybdenum cofactor (MoCo) and can be found in enzymes that catalyze redox reactions in carbon, nitrogen, and sulfur utilization (Williams et al., 2011). moeB1, a homologous gene potentially involved in MoCo biosynthesis, has also been shown to be required for arresting phagosome acidification (MacGurn and Cox, 2007). moaC1 mutant itself has exhibited reduced virulence in macrophages as well as primate lungs (Rosas-Magallanes et al., 2007; Dutta et al., 2010). Further investigation of how MoCo-mediated redox reactions alter the intraphagosomal environment should provide more insight on the cellular processes employed by M. tuberculosis to adapt to the mammalian host environment.
Rv3114 has been shown to be required for M. tuberculosis persistence in the mouse spleen (Sassetti and Rubin, 2003) and is temporally regulated during infection (Talaat et al., 2004). It encodes a putative nucleoside deaminase involved in nucleotide metabolism (Akhter et al., 2008).
Rv2954c–2961
Rv2954c–2961 make-up a 7 kb genomic island with a low GC content (53.6%) and a transposase gene (Becq et al., 2007). A phylogenetic analysis of multiple mycobacterial genome sequences proposed a step-wise acquisition of the genes within this locus. Specifically, some genes are present in slow-growing, but not rapid-growing mycobacteria, suggesting that they were acquired by the common ancestor of the slow-growing species. Conversely, other genes in this island are specific to M. tuberculosis and are therefore inferred to have been acquired after the common ancestor with M. kansasii and M. marinum (Veyrier et al., 2009).
Genes within this island are involved in the synthesis and modification of phenolic glycolipids (PGLs), complex lipids located in the outermost layer of the mycobacterial cell envelope. PGLs are composed of long-chain fatty acid backbones with a phenol ring and methylated sugars, including two rhamnosyl and a terminal fucosyl residue (Onwueme et al., 2005). PGLs are only produced by the members of the MTBC and related slow-growing mycobacteria, yet even among the mycobacteria that make PGL; there are species-specific modifications in the carbohydrate moiety [Daffe and Laneelle, 1988; Onwueme et al., 2005; schematically illustrated in (Veyrier et al., 2011)]. The PGLs have been implicated in mycobacterial pathogenicity such as oxidative stress resistance (Chan et al., 1989), cell tropism (Ng et al., 2000; Rambukkana, 2001), and immunomodulation (Reed et al., 2004; Guenin-Mace et al., 2009; Cambier et al., 2014).
The major type of PGLs produced by M. tuberculosis is denoted PGL-tb (Simeone et al., 2007). While most genes involved in the synthesis of the lipid core and carbohydrates are characterized, the enzymes responsible for O-methylating the fucosyl residue remained elusive until recently. It is now known that Rv2954c–Rv2956 code for the methyltransferases that are responsible for O-methylation of the terminal fucosyl residue (Veyrier et al., 2009; Simeone et al., 2013). These three proteins catalyze the O-methylation of the hydroxyl groups of the terminal fucosyl residue of PGL-tb in a sequential process (Simeone et al., 2013).
In other pathogenic mycobacteria that lack the Rv2954c, Rv2955c, and Rv2956 orthologs such as M. marinum and M. leprae, their PGLs do not contain the terminal O-methylated fucosyl residue (Onwueme et al., 2005). Although M. kansasii does not possess Rv2954c or 2955c, it does encode an enzyme that is highly similar to Rv2956 (84%), and its PGL contains four sugar residues (Riviere et al., 1987; Onwueme et al., 2005). Interestingly, Rv2954c and Rv2955c have been found to be virulence genes during macrophage infection (Rosas-Magallanes et al., 2007), and Rv2954c was induced upon exposure to lung surfactant (Schwab et al., 2009).
As the enzymes encoded within this island have been observed to catalyze the transfer of functional groups from one molecule to another, they may play an important role at “decorating” existing mycobacterial products and fine-tuning host responses toward the organism to optimize its intracellular survival (Veyrier et al., 2011).
Conclusion
Phylogenetic analyses have been used as robust and reliable tools for identifying potential HGT loci in M. tuberculosis and other pathogenic mycobacteria. However, the biological relevance of most of these genomic regions remains to be delineated. In this review we examine four examples of how such putative HGT genes can affect the physiology of the pathogen and its interaction with the host. The functional characterization of these and other putative HGT-associated genes will allow us to understand whether and how HGT events have contributed to the pathogenesis of M. tuberculosis, ultimately guiding the development of new diagnostic tests and vaccines against this particularly successful pathogen.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
Joyce Wang is supported by a studentship award from the Fonds de la Recherche en Sante du Quebec (FRSQ). Marcel A. Behr is a Chercheur-National of the FRSQ and a William Dawson Scholar of McGill University. Mycobacterial genomic work in the Behr lab is funded by an operating grant from the Canadian Institutes for Health Research, MOP-97813.
References
Akhter, Y., Yellaboina, S., Farhana, A., Ranjan, A., Ahmed, N., and Hasnain, S. E. (2008). Genome scale portrait of cAMP-receptor protein (CRP) regulons in mycobacteria points to their role in pathogenesis. Gene 407, 148–158. doi: 10.1016/j.gene.2007.10.017
Alexander, D. C., Turenne, C. Y., and Behr, M. A. (2009). Insertion and deletion events that define the pathogen Mycobacterium avium subsp paratuberculosis. J. Bacteriol. 191, 1018–1025. doi: 10.1128/JB.01340-08
Baldwin, A., Sokol, P. A., Parkhill, J., and Mahenthiralingam, E. (2004). The Burkholderia cepacia epidemic strain marker is part of a novel genomic island encoding both virulence and metabolism-associated genes in Burkholderia cenocepacia. Infect. Immun. 72, 1537–1547. doi: 10.1128/IAI.72.3.1537-1547.2004
Baltz, R. H. (2008). Renaissance in antibacterial discovery from actinomycetes. Curr. Opin. Pharmacol. 8, 557–563. doi: 10.1016/j.coph.2008.04.008
Be, N. A., Lamichhane, G., Grosset, J., Tyagi, S., Cheng, Q. J., Kim, K. S., et al. (2008). Murine model to study the invasion and survival of Mycobacterium tuberculosis in the central nervous system. J. Infect. Dis. 198, 1520–1528. doi: 10.1086/592447
Becq, J., Gutierrez, M. C., Rosas-Magallanes, V., Rauzier, J., Gicquel, B., Neyrolles, O., et al. (2007). Contribution of horizontally acquired genomic islands to the evolution of the tubercle bacilli. Mol. Biol. Evol. 24, 1861–1871. doi: 10.1093/molbev/msm111
Behar, S. M., Divangahi, M., and Remold, H. G. (2010). Evasion of innate immunity by Mycobacterium tuberculosis: is death an exit strategy? Nat. Rev. Microbiol. 8, 668–674.
Blasco, B., Chen, J. M., Hartkoorn, R., Sala, C., Uplekar, S., Rougemont, J., et al. (2012). Virulence regulator EspR of Mycobacterium tuberculosis is a nucleoid-associated protein. PLoS Pathog. 8:e1002621. doi: 10.1371/journal.ppat.1002621
Boshoff, H. I., Reed, M. B., Barry, C. E. III, and Mizrahi, V. (2003). DnaE2 polymerase contributes to in vivo survival and the emergence of drug resistance in Mycobacterium tuberculosis. Cell 113, 183–193. doi: 10.1016/S0092-8674(03)00270-8
Braibant, M., Gilot, P., and Content, J. (2000). The ATP binding cassette (ABC) transport systems of Mycobacterium tuberculosis. FEMS Microbiol. Rev. 24, 449–467. doi: 10.1111/j.1574-6976.2000.tb00550.x
Brodin, P., Majlessi, L., Marsollier, L., de Jonge, M. I., Bottai, D., Demangel, C., et al. (2006). Dissection of ESAT-6 system 1 of Mycobacterium tuberculosis and impact on immunogenicity and virulence. Infect. Immun. 74, 88–98. doi: 10.1128/IAI.74.1.88-98.2006
Brodin, P., Poquet, Y., Levillain, F., Peguillet, I., Larrouy-Maumus, G., Gilleron, M., et al. (2010). High content phenotypic cell-based visual screen identifies Mycobacterium tuberculosis acyltrehalose-containing glycolipids involved in phagosome remodeling. PLoS Pathog. 6:e1001100. doi: 10.1371/journal.ppat.1001100
Buchmeier, N., Blanc-Potard, A., Ehrt, S., Piddington, D., Riley, L., and Groisman, E. A. (2000). A parallel intraphagosomal survival strategy shared by Mycobacterium tuberculosis and Salmonella enterica. Mol. Microbiol. 35, 1375–1382. doi: 10.1046/j.1365-2958.2000.01797.x
Cambier, C. J., Takaki, K. K., Larson, R. P., Hernandez, R. E., Tobin, D. M., Urdahl, K. B., et al. (2014). Mycobacteria manipulate macrophage recruitment through coordinated use of membrane lipids. Nature 505, 218–222. doi: 10.1038/nature12799
Chan, J., Fujiwara, T., Brennan, P., McNeil, M., Turco, S. J., Sibille, J. C., et al. (1989). Microbial glycolipids: possible virulence factors that scavenge oxygen radicals. Proc. Natl. Acad. Sci. U.S.A. 86, 2453–2457. doi: 10.1073/pnas.86.7.2453
Chouikha, I., Germon, P., Brée, A., Gilot, P., Moulin-Schouleur, M., and Schouler, C. (2006). A selC-associated genomic island of the extraintestinal avian pathogenic Escherichia coli strain BEN2908 is involved in carbohydrate uptake and virulence. J. Bacteriol. 188, 977–987. doi: 10.1128/JB.188.3.977-987.2006
Colangeli, R., Haq, A., Arcus, V. L., Summers, E., Magliozzo, R. S., McBride, A., et al. (2009). The multifunctional histone-like protein Lsr2 protects mycobacteria against reactive oxygen intermediates. Proc. Natl. Acad. Sci. U.S.A. 106, 4414–4418. doi: 10.1073/pnas.0810126106
Cole, S. T., Brosch, R., Parkhill, J., Garnier, T., Churcher, C., Harris, D., et al. (1998). Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393, 537–544. doi: 10.1038/31159
Daffe, M., and Laneelle, M. A. (1988). Distribution of phthiocerol diester, phenolic mycosides and related compounds in mycobacteria. J. Gen. Microbiol. 134, 2049–2055.
Dairi, T., Hamano, Y., Kuzuyama, T., Itoh, N., Furihata, K., and Seto, H. (2001). Eubacterial diterpene cyclase genes essential for production of the isoprenoid antibiotic terpentecin. J. Bacteriol. 183, 6085–6094. doi: 10.1128/JB.183.20.6085-6094.2001
Darwin, K. H., and Nathan, C. F. (2005). Role for nucleotide excision repair in virulence of Mycobacterium tuberculosis. Infect. Immun. 73, 4581–4587. doi: 10.1128/IAI.73.8.4581-4587.2005
Divangahi, M., Chen, M., Gan, H., Desjardins, D., Hickman, T. T., Lee, D. M., et al. (2009). Mycobacterium tuberculosis evades macrophage defenses by inhibiting plasma membrane repair. Nat. Immunol. 10, 899–906. doi: 10.1038/ni.1758
Dürr, C., Schnell, H. J., Luzhetskyy, A., Murillo, R., Weber, M., Welzel, K., et al. (2006). Biosynthesis of the terpene phenalinolactone in Streptomyces sp. Tu6071: analysis of the gene cluster and generation of derivatives. Chem. Biol. 13, 365–377. doi: 10.1016/j.chembiol.2006.01.011
Dutta, N. K., Mehra, S., Didier, P. J., Roy, C. J., Doyle, L. A., Alvarez, X., et al. (2010). Genetic requirements for the survival of tubercle bacilli in primates. J. Infect. Dis. 201, 1743–1752. doi: 10.1086/652497
Frost, L. S., Leplae, R., Summers, A. O., and Toussaint, A. (2005). Mobile genetic elements: the agents of open source evolution. Nat. Rev. Microbiol. 3, 722–732. doi: 10.1038/nrmicro1235
Gray, T. A., Krywy, J. A., Harold, J., Palumbo, M. J., and Derbyshire, K. M. (2013). Distributive conjugal transfer in mycobacteria generates progeny with meiotic-like genome-wide mosaicism, allowing mapping of a mating identity locus. PLoS Biol. 11:e1001602. doi: 10.1371/journal.pbio.1001602
Griffin, J. E., Gawronski, J. D., Dejesus, M. A., Ioerger, T. R., Akerley, B. J., and Sassetti, C. M. (2011). High-resolution phenotypic profiling defines genes essential for mycobacterial growth and cholesterol catabolism. PLoS Pathog. 7:e1002251. doi: 10.1371/journal.ppat.1002251
Guenin-Mace, L., Simeone, R., and Demangel, C. (2009). Lipids of pathogenic mycobacteria: contributions to virulence and host immune suppression. Transbound. Emerg. Dis. 56, 255–268. doi: 10.1111/j.1865-1682.2009.01072.x
Hacker, J., Blum-Oehler, G., Mühldorfer, I., and Tschäpe, H. (1997). Pathogenicity islands of virulent bacteria: structure, function and impact on microbial evolution. Mol. Microbiol. 23, 1089–1097. doi: 10.1046/j.1365-2958.1997.3101672.x
Hamano, Y., Kuzuyama, T., Itoh, N., Furihata, K., Seto, H., and Dairi, T. (2002). Functional analysis of eubacterial diterpene cyclases responsible for biosynthesis of a diterpene antibiotic, terpentecin. J. Biol. Chem. 277, 37098–37104. doi: 10.1074/jbc.M206382200
Layre, E., Lee, H. J., Young, D. C., Jezek Martinot, A., Buter, J., Minnaard, A. J., et al. (2014). Molecular profiling of Mycobacterium tuberculosis identifies tuberculosinyl nucleoside products of the virulence-associated enzyme Rv3378c. Proc. Natl. Acad. Sci. U.S.A. 111, 2978–2983. doi: 10.1073/pnas.1315883111
Leistikow, R. L., Morton, R. A., Bartek, I. L., Frimpong, I., Wagner, K., and Voskuil, M. I. (2010). The Mycobacterium tuberculosis DosR regulon assists in metabolic homeostasis and enables rapid recovery from nonrespiring dormancy. J. Bacteriol. 192, 1662–1670. doi: 10.1128/JB.00926-09
Lercher, M. J., and Pal, C. (2008). Integration of horizontally transferred genes into regulatory interaction networks takes many million years. Mol. Biol. Evol. 25, 559–567. doi: 10.1093/molbev/msm283
Li, M., Du, X., Villaruz, A. E., Diep, B. A., Wang, D., Song, Y., et al. (2012). MRSA epidemic linked to a quickly spreading colonization and virulence determinant. Nat. Med. 18, 816–819. doi: 10.1038/nm.2692
MacGurn, J. A., and Cox, J. S. (2007). A genetic screen for Mycobacterium tuberculosis mutants defective for phagosome maturation arrest identifies components of the ESX-1 secretion system. Infect. Immun. 75, 2668–2678. doi: 10.1128/IAI.01872-06
Mann, F. M., and Peters, R. J. (2012). Isotuberculosinol: the unusual case of an immunomodulatory diterpenoid from Mycobacterium tuberculosis. Medchemcomm 3, 899–904. doi: 10.1039/c2md20030a
Mann, F. M., Prisic, S., Hu, H., Xu, M., Coates, R. M., and Peters, R. J. (2009a). Characterization and inhibition of a class II diterpene cyclase from Mycobacterium tuberculosis: implications for tuberculosis. J. Biol. Chem. 284, 23574–23579. doi: 10.1074/jbc.M109.023788
Mann, F. M., Xu, M., Chen, X., Fulton, D. B., Russell, D. G., and Peters, R. J. (2009b). Edaxadiene: a new bioactive diterpene from Mycobacterium tuberculosis. J. Am. Chem. Soc. 131, 17526–17527. doi: 10.1021/ja9019287
Mann, F. M., VanderVen, B. C., and Peters, R. J. (2011). Magnesium depletion triggers production of an immune modulating diterpenoid in Mycobacterium tuberculosis. Mol. Microbiol. 79, 1594–1601. doi: 10.1111/j.1365-2958.2011.07545.x
Manzanillo, P. S., Shiloh, M. U., Portnoy, D. A., and Cox, J. S. (2012). Mycobacterium tuberculosis activates the DNA-dependent cytosolic surveillance pathway within macrophages. Cell Host Microbe 11, 469–480. doi: 10.1016/j.chom.2012.03.007
Marrero, J., Rhee, K. Y., Schnappinger, D., Pethe, K., and Ehrt, S. (2010). Gluconeogenic carbon flow of tricarboxylic acid cycle intermediates is critical for Mycobacterium tuberculosis to establish and maintain infection. Proc. Natl. Acad. Sci. U.S.A. 107, 9819–9824. doi: 10.1073/pnas.1000715107
Master, S. S., Springer, B., Sander, P., Boettger, E. C., Deretic, V., and Timmins, G. S. (2002). Oxidative stress response genes in Mycobacterium tuberculosis: role of ahpC in resistance to peroxynitrite and stage-specific survival in macrophages. Microbiology 148(Pt 10), 3139–3144.
Matthysse, A. G., Yarnall, H. A., and Young, N. (1996). Requirement for genes with homology to ABC transport systems for attachment and virulence of Agrobacterium tumefaciens. J. Bacteriol. 178, 5302–5308.
Matthysse, A. G., Yarnall, H., Boles, S. B., and McMahan, S. (2000). A region of the Agrobacterium tumefaciens chromosome containing genes required for virulence and attachment to host cells. Biochim. Biophys. Acta 1490, 208–212. doi: 10.1016/S0167-4781(99)00250-X
McSwiggan, D. A., and Collins, C. H. (1974). The isolation of M. kansasii and M. xenopi from water systems. Tubercle 55, 291–297. doi: 10.1016/0041-3879(74)90038-5
Mohd-Zain, Z., Turner, S. L., Cerdeño-Tárraga, A. M., Lilley, A. K., Inzana, T. J., Duncan, A. J., et al. (2004). Transferable antibiotic resistance elements in Haemophilus influenzae share a common evolutionary origin with a diverse family of syntenic genomic islands. J. Bacteriol. 186, 8114–8122. doi: 10.1128/JB.186.23.8114-8122.2004
Nakano, C., Okamura, T., Sato, T., Dairi, T., and Hoshino, T. (2005). Mycobacterium tuberculosis H37Rv3377c encodes the diterpene cyclase for producing the halimane skeleton. Chem. Commun. (Camb.) 1016–1018. doi: 10.1039/B415346D
Ng, V., Zanazzi, G., Timpl, R., Talts, J. F., Salzer, J. L., Brennan, P. J., et al. (2000). Role of the cell wall phenolic glycolipid-1 in the peripheral nerve predilection of Mycobacterium leprae. Cell 103, 511–524. doi: 10.1016/S0092-8674(00)00142-2
Noda-Garcia, L., Camacho-Zarco, A. R., Medina-Ruíz, S., Gaytán, P., Carrillo-Tripp, M., Fülöp, V., et al. (2013). Evolution of substrate specificity in a recipient’s enzyme following horizontal gene transfer. Mol. Biol. Evol. 30, 2024–2034. doi: 10.1093/molbev/mst115
Onwueme, K. C., Vos, C. J., Zurita, J., Ferreras, J. A., and Quadri, L. E. (2005). The dimycocerosate ester polyketide virulence factors of mycobacteria. Prog. Lipid Res. 44, 259–302. doi: 10.1016/j.plipres.2005.07.001
Palmer, K. L., Kos, V. N., and Gilmore, M. S. (2010). Horizontal gene transfer and the genomics of enterococcal antibiotic resistance. Curr. Opin. Microbiol. 13, 632–639. doi: 10.1016/j.mib.2010.08.004
Pethe, K., Swenson, D. L., Alonso, S., Anderson, J., Wang, C., and Russell, D. G. (2004). Isolation of Mycobacterium tuberculosis mutants defective in the arrest of phagosome maturation. Proc. Natl. Acad. Sci. U.S.A. 101, 13642–13647. doi: 10.1073/pnas.0401657101
Price, M. N., Dehal, P. S., and Arkin, A. P. (2008). Horizontal gene transfer and the evolution of transcriptional regulation in Escherichia coli. Genome Biol. 9:R4. doi: 10.1186/gb-2008-9-1-r4
Rambukkana, A. (2001). Molecular basis for the peripheral nerve predilection of Mycobacterium leprae. Curr. Opin. Microbiol. 4, 21–27. doi: 10.1016/S1369-5274(00)00159-4
Reed, M. B., Domenech, P., Manca, C., Su, H., Barczak, A. K., Kreiswirth, B. N., et al. (2004). A glycolipid of hypervirulent tuberculosis strains that inhibits the innate immune response. Nature 431, 84–87. doi: 10.1038/nature02837
Riviere, M., Fournie, J. J., and Puzo, G. (1987). A novel mannose containing phenolic glycolipid from Mycobacterium kansasii. J. Biol. Chem. 262, 14879–14884.
Rohde, K. H., Abramovitch, R. B., and Russell, D. G. (2007). Mycobacterium tuberculosis invasion of macrophages: linking bacterial gene expression to environmental cues. Cell Host Microbe 2, 352–364. doi: 10.1016/j.chom.2007.09.006
Rosas-Magallanes, V., Deschavanne, P., Quintana-Murci, L., Brosch, R., Gicquel, B., and Neyrolles, O. (2006). Horizontal transfer of a virulence operon to the ancestor of Mycobacterium tuberculosis. Mol. Biol. Evol. 23, 1129–1135. doi: 10.1093/molbev/msj120
Rosas-Magallanes, V., Stadthagen-Gomez, G., Rauzier, J., Barreiro, L. B., Tailleux, L., Boudou, F., et al. (2007). Signature-tagged transposon mutagenesis identifies novel Mycobacterium tuberculosis genes involved in the parasitism of human macrophages. Infect. Immun. 75, 504–507. doi: 10.1128/IAI.00058-06
Ryndak, M. B., Wang, S., Smith, I., and Rodriguez, G. M. (2010). The Mycobacterium tuberculosis high-affinity iron importer, IrtA, contains an FAD-binding domain. J. Bacteriol. 192, 861–869. doi: 10.1128/JB.00223-09
Sartori, F. G., Leandro, L. F., Montanari, L. B., de Souza, M. G., Pires, R. H., Sato, D. N., et al. (2013). Isolation and identification of environmental mycobacteria in the waters of a hemodialysis center. Curr. Microbiol. 67, 107–111. doi: 10.1007/s00284-013-0341-6
Sassetti, C. M., and Rubin, E. J. (2003). Genetic requirements for mycobacterial survival during infection. Proc. Natl. Acad. Sci. U.S.A. 100, 12989–12994. doi: 10.1073/pnas.2134250100
Schnappinger, D., Ehrt, S., Voskuil, M. I., Liu, Y., Mangan, J. A., Monahan, I. M., et al. (2003). Transcriptional adaptation of Mycobacterium tuberculosis within macrophages: insights into the phagosomal environment. J. Exp. Med. 198, 693–704. doi: 10.1084/jem.20030846
Schwab, U., Rohde, K. H., Wang, Z., Chess, P. R., Notter, R. H., and Russell, D. G. (2009). Transcriptional responses of Mycobacterium tuberculosis to lung surfactant. Microb. Pathog. 46, 185–193. doi: 10.1016/j.micpath.2008.12.006
Simeone, R., Constant, P., Guilhot, C., Daffé, M., and Chalut, C. (2007). Identification of the missing trans-acting enoyl reductase required for phthiocerol dimycocerosate and phenolglycolipid biosynthesis in Mycobacterium tuberculosis. J. Bacteriol. 189, 4597–4602. doi: 10.1128/JB.00169-07
Simeone, R., Huet, G., Constant, P., Malaga, W., Lemassu, A., Laval, F., et al. (2013). Functional characterisation of three o-methyltransferases involved in the biosynthesis of phenolglycolipids in Mycobacterium tuberculosis. PLoS ONE 8:e58954. doi: 10.1371/journal.pone.0058954
Simion, G. (2005). “Terpenoids,” in Encyclopedia of Chromatography, 2nd Edn, ed. J. Cazes, CRC Press. 1676–1681. doi: 10.1201/NOE0824727857.ch368
Smanski, M. J., Yu, Z., Casper, J., Lin, S., Peterson, R. M., Chen, Y., et al. (2011). Dedicated ent-kaurene and ent-atiserene synthases for platensimycin and platencin biosynthesis. Proc. Natl. Acad. Sci. U.S.A. 108, 13498–13503. doi: 10.1073/pnas.1106919108
Steadham, J. E. (1980). High-catalase strains of Mycobacterium kansasii isolated from water in Texas. J. Clin. Microbiol. 11, 496–498.
Stewart, G. R., Patel, J., Robertson, B. D., Rae, A., and Young, D. B. (2005). Mycobacterial mutants with defective control of phagosomal acidification. PLoS Pathog. 1:269–278. doi: 10.1371/journal.ppat.0010033
Sun, J., Wang, X., Lau, A., Liao, T. Y., Bucci, C., and Hmama, Z. (2010). Mycobacterial nucleoside diphosphate kinase blocks phagosome maturation in murine RAW 264.7 macrophages. PLoS ONE 5:e8769. doi: 10.1371/journal.pone.0008769
Supply, P., Marceau, M., Mangenot, S., Roche, D., Rouanet, C., Khanna, V., et al. (2013). Genomic analysis of smooth tubercle bacilli provides insights into ancestry and pathoadaptation of Mycobacterium tuberculosis. Nat. Genet. 45, 172–179. doi: 10.1038/ng.2517
Talaat, A. M., Lyons, R., Howard, S. T., and Johnston, S. A. (2004). The temporal expression profile of Mycobacterium tuberculosis infection in mice. Proc. Natl. Acad. Sci. U.S.A. 101, 4602–4607. doi: 10.1073/pnas.0306023101
Thomson, R. M., Carter, R., Tolson, C., Coulter, C., Huygens, F., and Hargreaves, M. (2013). Factors associated with the isolation of Nontuberculous mycobacteria (NTM) from a large municipal water system in Brisbane, Australia. BMC Microbiol. 13:89. doi: 10.1186/1471-2180-13-89
Treangen, T. J., and Rocha, E. P. C. (2011). Horizontal transfer, not duplication, drives the expansion of protein families in prokaryotes. PLoS Genet. 7:e1001284. doi: 10.1371/journal.pgen.1001284
Vandal, O. H., Pierini, L. M., Schnappinger, D., Nathan, C. F., and Ehrt, S. (2008). A membrane protein preserves intrabacterial pH in intraphagosomal Mycobacterium tuberculosis. Nat. Med. 14, 849–854. doi: 10.1038/nm.1795
Vandal, O. H., Roberts, J. A., Odaira, T., Schnappinger, D., Nathan, C. F., and Ehrt, S. (2009). Acid-susceptible mutants of Mycobacterium tuberculosis share hypersusceptibility to cell wall and oxidative stress and to the host environment. J. Bacteriol. 191, 625–631. doi: 10.1128/JB.00932-08
van der Wel, N., Hava, D., Houben, D., Fluitsma, D., van Zon, M., Pierson, J., et al. (2007). M. tuberculosis and M. leprae translocate from the phagolysosome to the cytosol in myeloid cells. Cell 129, 1287–1298. doi: 10.1016/j.cell.2007.05.059
Ventura, M., Canchaya, C., Tauch, A., Chandra, G., Fitzgerald, G. F., Chater, K. F., et al. (2007). Genomics of Actinobacteria: tracing the evolutionary history of an ancient phylura. Microbiol. Mol. Biol. Rev. 71, 495–548. doi: 10.1128/MMBR.00005-07
Veyrier, F. J., Dufort, A., and Behr, M. A. (2011). The rise and fall of the Mycobacterium tuberculosis genome. Trends Microbiol. 19, 156–161. doi: 10.1016/j.tim.2010.12.008
Veyrier, F., Pletzer, D., Turenne, C., and Behr, M. A. (2009). Phylogenetic detection of horizontal gene transfer during the step-wise genesis of Mycobacterium tuberculosis. BMC Evol. Biol. 9:196. doi: 10.1186/1471-2148-9-196
Waddell, S. J., and Butcher, P. D. (2007). Microarray analysis of whole genome expression of intracellular Mycobacterium tuberculosis. Curr. Mol. Med. 7, 287–296. doi: 10.2174/156652407780598548
Williams, M. J., Kana, B. D., and Mizrahi, V. (2011). Functional analysis of molybdopterin biosynthesis in mycobacteria identifies a fused molybdopterin synthase in Mycobacterium tuberculosis. J. Bacteriol. 193, 98–106. doi: 10.1128/JB.00774-10
Wilson, M. S., Herrick, J. B., Jeon, C. O., Hinman, D. E., and Madsen, E. L. (2003). Horizontal transfer of phnAc dioxygenase genes within one of two phenotypically and genotypically distinctive naphthalene-degrading guilds from adjacent soil environments. Appl. Environ. Microbiol. 69, 2172–2181. doi: 10.1128/AEM.69.4.2172-2181.2003
Wong, D., Bach, H., Sun, J., Hmama, Z., and Av-Gay, Y. (2011). Mycobacterium tuberculosis protein tyrosine phosphatase (PtpA) excludes host vacuolar-H+-ATPase to inhibit phagosome acidification. Proc. Natl. Acad. Sci. U.S.A. 108, 19371–19376. doi: 10.1073/pnas.1109201108
World Health Organization. (2013). Global Tuberculosis Report 2013. Geneva, Switzerland: WHO/HTM/TB/2013.11.
Keywords: Mycobacterium tuberculosis, Mycobacterium kansasii, M. tuberculosis-specific genes, horizontal gene transfer, comparative genomics
Citation: Wang J and Behr MA (2014) Building a better bacillus: the emergence of Mycobacterium tuberculosis. Front. Microbiol. 5:139. doi: 10.3389/fmicb.2014.00139
Received: 17 December 2013; Accepted: 18 March 2014;
Published online: 03 April 2014.
Edited by:
Frank T. Robb, University of California, USAReviewed by:
David L. Bernick, University of California at Santa Cruz, USAStephen Techtmann, University of Tennessee at Knoxville, USA
Copyright © 2014 Wang and Behr. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Marcel A. Behr, McGill University Health Centre, A5.156, 1650 Cedar Avenue, Montreal, QC H3G 1A4, Canada e-mail:bWFyY2VsLmJlaHJAbWNnaWxsLmNh
 Joyce Wang
Joyce Wang Marcel A. Behr
Marcel A. Behr