- 1Department of Microbiology and Microbial Engineering, School of Life Sciences, Fudan University, Shanghai, China
- 2Key Laboratory of Parasite and Vector Biology of Ministry of Health, National Institute of Parasitic Diseases, Chinese Center for Diseases Control and Prevention, Shanghai, China
Schistosomiasis, caused by dioecious flatworms in the genus Schistosoma, is torturing people from many developing countries nowadays and frequently leads to severe morbidity and mortality of the patients. Praziquantel based chemotherapy and morbidity control for this disease adopted currently necessitate viable and efficient diagnostic technologies. Fortunately, those “-omics” researches, which rely on high-throughput experimental technologies to produce massive amounts of informative data, have substantially contributed to the exploitation and innovation of diagnostic tools of schistosomiasis. In its first section, this review provides a concise conclusion on the progresses pertaining to schistosomal “-omics” researches to date, followed by a comprehensive section on the diagnostic methods of schistosomiasis, especially those innovative ones based on the detection of antibodies, antigens, nucleic acids, and metabolites with a focus on those achievements inspired by “-omics” researches. Finally, suggestions about the design of future diagnostic tools of schistosomiasis are proposed, in order to better harness those data produced by “-omics” studies.
Schistosomiasis or “Bilharziasis” refers to the parasitic diseases caused by dioecious flatworms in the genus Schistosoma. The main pathogenic species of human schistosomiasis comprise Schistosoma mansoni, S. japonicum, and S. haematobium. Free living larvae of these pathogens released by variant kinds of snails in the fresh water, i.e., the cercariae can penetrate the skin of their definite hosts and subsequently bring about either egg-induced chronic responses, known as intestinal schistosomiasis (S. mansoni and S. japonicum) and urinary schistosomiasis (S. haematobium; Hu et al., 2004), or, less frequently, some acute reactions like toxemia or cytokine-induced shock (Wynn et al., 2004). These highly debilitating symptoms account for the severe morbidity and mortality incurred by schistosomiasis and render it a key poverty contributor for some developing countries in the tropical and subtropical regions. Moreover, schistosomiasis has also become an emerging threat in non-endemic areas due to the mounting numbers of immigrants and tourists (Patz et al., 2000; Enk et al., 2010). Currently, praziquantel (PZQ) based chemotherapy and morbidity control have become the predominantly adopted strategy for the treatment and control of schistosomiasis worldwide (Qing-Wu et al., 2002; Zhou et al., 2011b). Consequently, monitoring disease transmission, seeking patients for treatment in endemic areas and the evaluation of remedies entail viable and efficient diagnostic technologies, in order to optimize the extant control and prevention project for schistosomiasis (Balog et al., 2010).
Status Quo of the Current Schistosomal “-Omics” Researches
“-omics” is a suffix labeling a series of biological subjects that adopt high-throughput experimental technologies to produce a large body of holistic and informative data. Since the launching of the Schistosoma Genome Project propelled by the WHO in 1994 (Oliveira et al., 2008), colossal progresses have been made in various Schistosoma -omics arenas including genomics, transcriptomics, proteomics, metabonomics (Ju et al., 2010), to name but a few. Besides an augmented understanding of schistosome biology and pathogenesis, those knowledge acquired by Schistosoma -omics researches also substantially contribute to the exploitation and innovation of drugs, vaccines as well as diagnostic tools for schistosomiasis. This review aims to display the representative achievements of those schistosomal -omics researches and conclude the diagnostic inventions derived from or at least inspired by those -omics studies.
Transcripomics
Schistosoma transcriptomics research commenced from the establishment of complementary DNA (cDNA) libraries which generated expressed sequence tags (ESTs) from nearly all stages of S. mansoni and S. japonicum (Fan et al., 1998; Fung et al., 2002; Hu et al., 2003; Peng et al., 2003; Shabaan et al., 2003; Verjovski-Almeida et al., 2003; Merrick et al., 2009). After the obtainment of numerous ESTs data, microarray and serial analysis of gene expression (SAGE) technology have been used to profile the transcripts of schistosomes in different stages and/or under distinct conditions. In particular, Geoffrey Gobert classified these transcriptomics applications into four main categories (Gobert, 2010), i.e., (a) characterizing individual cell/tissue types, (Jones et al., 2007; Gobert et al., 2009a), (b) profiling the intact organism and lifecycle (Hoffmann et al., 2002; Fitzpatrick et al., 2004, 2005, 2009; Hoffmann and Fitzpatrick, 2004; Chai et al., 2006; Dillon et al., 2006, 2008; Gobert et al., 2006, 2009b; Moertel et al., 2006; Vermeire et al., 2006; Jolly et al., 2007; Ojopi et al., 2007; Williams et al., 2007; Hu et al., 2009; Taft et al., 2009), (c) parasite–host interactions and effect of therapies on parasite (Hoffmann et al., 2001; Alger and Williams, 2002; Sandler et al., 2003; Aragon et al., 2008, 2009; Gobert and Jones, 2008; de Moraes Mourão et al., 2009; You et al., 2009; Burke et al., 2010; Gobert et al., 2010), and (d) gene expression differences between geographical isolates or species (Le et al., 2002; Fitzpatrick et al., 2004; Moertel et al., 2006). Not until 2012 was the transcriptome of the adult and egg stages of S. haematobium profiled along with its genome (Young et al., 2012).
MicroRNA (miRNA) Genomics
Unlike messenger RNAs (mRNAs), small non-coding RNAs (sncRNAs) represent a group of untranslatable transcripts which are approximately 18–30 nt in length and serve as critical regulators to silence or activate specific target genes in a variety of organisms (Bartel, 2004; Molnar et al., 2010). Endogenous-small interfering (Endo-siRNAs), miRNAs, and Piwi-interacting RNA (piRNAs) are three main components of sncRNAs (Kim, 2005). Using protocols similar to the conventional transcriptomic researches, which can be outlined as RNA isolation, library construction, and sequencing (Cheng and Jin, 2012), vast numbers of schistosomal miRNAs and endo-siRNAs have been successfully detected in S. mansoni (Copeland et al., 2009; de Souza Gomes et al., 2011; Simões et al., 2011) and S. japonicum (Xue et al., 2008; Copeland et al., 2009; Hao et al., 2010; Wang et al., 2010b; Cai et al., 2011). What’s more, recently Cai et al. (2012) adopted a totally novel method, i.e., the immunoprecipitation of SjAgo2, a key factor in sncRNAs biogenesis with monoclonal antibodies (mAbs) to identify and characterize the associated small RNAs. Further classification steps showed that endo-siRNAs derived from transposable elements (TEs) were prominent among those conjugated sncRNAs.
Proteomics
Since the correlation between transcriptional level and translational level of one expressed gene is not necessarily straightforward, proteomic analyses are also of great importance aiming at the characterization and comparison of functions, abundance, and subcellular localization (Ju et al., 2010) of numerous gene products derived from single or multiple samples. In a typical proteomics research, schistosomal specimen would be separated by one dimensional-(1D-) or 2D-polyacrylamide gel electrophoresis (PAGE) in the beginning. Subsequently, the isolated bands or spots would be subjected to digestion and either Matrix-assisted laser desorption/ionization time-of-flight mass spectrometer (MALDI-TOF-MS) or liquid chromatography-tandem mass spectrometry (LC-MS/MS) according to the separation method. Finally, protein identities within the sample would be acquired through comparing the mass spectrometry (MS) results to the theoretical masses stocked in a particular database (van Hellemond et al., 2007). Based on the purpose of proteomics researches, schistosomal samples ought to be prepared in different ways. So far, schistosomal proteomics has been applied to the investigation and comparison of protein compositions in various developmental stages (Curwen et al., 2004; Liu et al., 2006) or between different genders (Cheng et al., 2005; Liu et al., 2006) and the worm proteins might be pre-fractioned accordingly, e.g., soluble membrane protein (Curwen et al., 2004; Cheng et al., 2005), tegumental fractions (Van Balkom et al., 2005; Braschi et al., 2006; Braschi and Wilson, 2006; Liu et al., 2006; Mulvenna et al., 2010; Castro-Borges et al., 2011), secreted antigens (Knudsen et al., 2005; Curwen et al., 2006; Liu et al., 2009), gut contents (Delcroix et al., 2007), etc.
Immunomics
Immunomics is a study that combines proteomics with serology and aims at ascertaining the interaction between host immune system and exotic antigens after pathogen invasion. Early schistosomal immunomics work utilized combined 2D Western blotting and MS to identify and compare proteins recognized by serum samples from S. haematobium exposed patients before and after PZQ treatment (Mutapi et al., 2005) or with different ages and infection intensities (Mutapi et al., 2008). Afterwards, an immunomics protein microarray technology was also applied in an attempt to seek prospective vaccine targets (Driguez et al., 2010). This advanced technology consists of several steps including genes selection, cell-free expression, chip printing, and serological probing.
Glycomics
In view of the critical role glycans played in the induction of innate immune responses during schistosome–host interplay, glycomics, the profiling of the schistosomal protein- and lipid-conjugated glycan or glycan elements was addressed even before the screening of schistosomal transcriptomics and proteomics (Hokke et al., 2007). So far, the patterns of glycoconjugates expressed by multiple life stages of S. mansoni have been precisely elucidated by either anticarbohydrate mAbs identification or MS-based method (Khoo et al., 1995, 1997a,b, 2001; van Remoortere et al., 2000, 2003; Wuhrer et al., 2000, 2002, 2006a,b; Huang et al., 2001; Nyame et al., 2003; Robijn et al., 2005, 2007a,b).
Genomics
Under the continual efforts from two consortia of researchers, the genome of S. mansoni and S. japonicum were deciphered and published simultaneously in 2009 (Berriman et al., 2009; The Schistosoma japonicum Genome Sequencing and Functional Analysis Consortium, 2009). Three years later, the genome information of S. haematobium, the pathogen causing urogenital schistosomiasis also became accessible (Young et al., 2012). Overall, the genome of the preceding three main causative human schistosomiasis pathogens are organized into eight chromosomes, including seven pairs of autosomes and a pair of ZW type sex chromosomes. The genome of S. mansoni (363 Mb), S. japonicum (397 Mb), and S. haematobium (385 Mb) encompass 13,184, 13,469, and 13,073 protein-coding genes, respectively, and a considerable proportion of them were mapped into gene ontology (GO) categories. Apart form those protein coding genes, more than 40% of these three genomes are taken up by repetitive elements, which contains retrotransposons, satellites, DNA transposons and other kinds of unknown repeats. As a whole, the relationship between S. mansoni and S. haematobium are more intimate than either of them with S. japonicum (Young et al., 2012).
Metabonomics
Schistosomes-related metabonomics keeps tabs on the overall changes of metabolites within tissues and biofluids of the hosts before and after blood fluke infection. Metabonomics-based schistosomiasis biomarker discoveries were pioneered in 2004 by Wang et al. (2004), who used 1H nuclear magnetic resonance (NMR) spectroscopy and multivariate pattern recognition to analyze the metabolic signature of urine samples from S. mansoni infected mice. Subsequently, using various spectroscopic methods, e.g., NMR spectroscopy, MS, and capillary electrophoresis (CE), coupled with various data-mining technologies (Legido-Quigley, 2010), several metabonomics related to schistosome infection were profiled, such as urine samples from mice infected with S. mansoni (Garcia-Perez et al., 2008, 2009, 2010) and hamsters infected with S. japonicum (Wang et al., 2006), as well as blood and multiple tissue samples from schistosome-alone infected (Li et al., 2009) or two pathogens co-infected rodents (Wu et al., 2010). It was not until early this decade that one metabonomic investigation started to survey human Schistosoma spp. infection (Balog et al., 2011). Corresponding works will be reviewed in the next section.
The “New Generation” Diagnostic Methods for Schistosomiasis and their Progresses Impelled by the “-Omics” Achievements
Traditional, Etiological Diagnostic Methods
Etiological diagnosis of schistosomiasis, i.e., the microscopic detection of schistosome eggs in urine (S. haematobium) or stool (S. mansoni and S. japonicum) samples is the most straightforward and widely adopted diagnostic approach to investigate the infection of schistosomes (Utzinger et al., 2011). Representative methods of etiological diagnosis include Kato–Katz (KK) thick smear (Katz et al., 1972), miracidium hatching assay (Jurberg et al., 2008), egg floating technique (FLOTAC; Cringoli, 2006), nuclepore urine filtration (Mott et al., 1982), etc. Those etiological methods are deemed as the “gold standard” for the detection of cases particularly in high endemicity settings and in the early stage of a control program. However, after mass drug administration or in low transmission settings, these traditional methods show limited sensitivity and accuracy (Zhao et al., 2012). In addition, multiple stool examinations or combined techniques, although reduced the false-negative rate to some extent, are quite time-consuming and labor-intensive, which hindered their application in large-scale epidemiological surveys (Zhou et al., 2011b). Thus, they have been gradually replaced by immunological techniques based on antigen or antibody detection, as well as PCR-based molecular tools, metabolic profiling, i.e., those so-called “new generation” diagnostic methods. Meanwhile, those novel diagnostic methods have benefited continuously from the previously introduced “-omics” researches (Table 1).
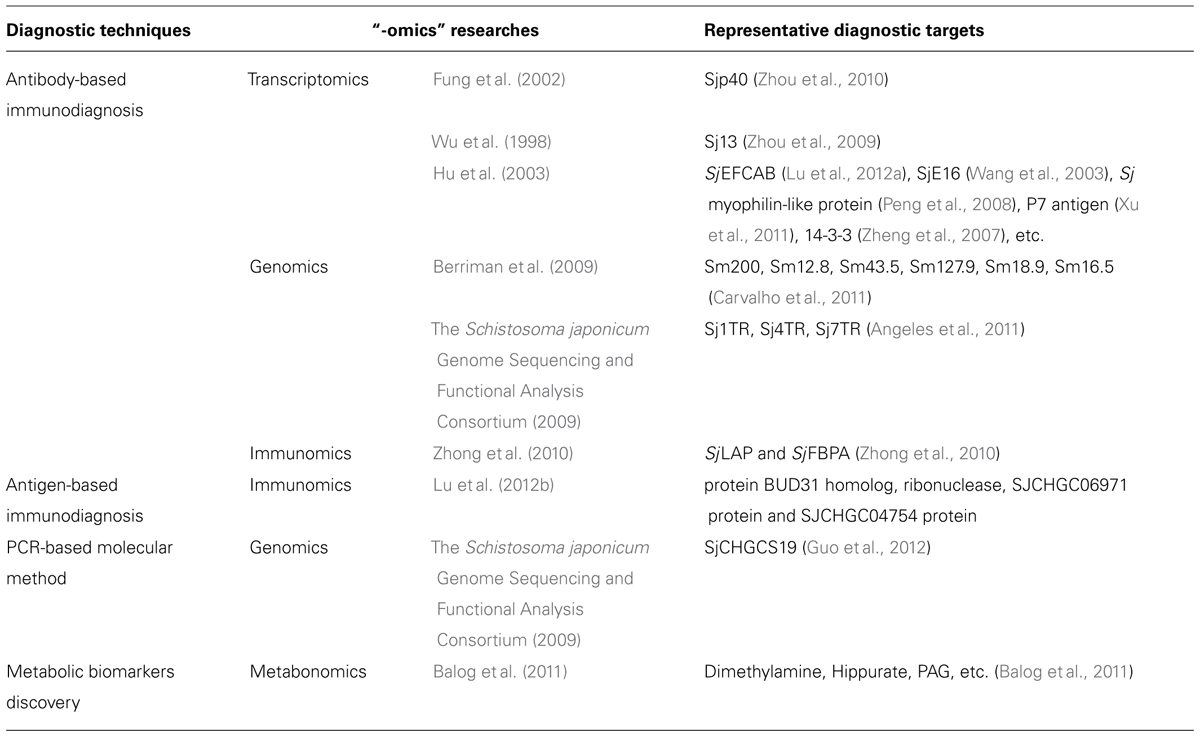
TABLE 1. An overview of “-omics” researches’ contribution to the revelation of novel schistosomiasis diagnostic targets.
Antibody-Based, Indirect Immunological Techniques
Indirect immunodiagnostic assays use a range of immunological methods, such as circumoval precipitin test (COPT), indirect hemagglutination assay (IHA), dipstick dye immunoassay (DDIA), and enzyme-linked immonosorbent assay (ELISA; Zhu, 2005), etc. in order to detect schistosome-specific antibodies. Immunodiagnosis based on the detection of antibodies is relatively repeatable, sensitive, tractable, and time-saving compared with traditional etiological methods. Early indirect immunodiagnosis mainly relied on the crude extracts of worm components, like microsomal extract, gut-associated polysaccharide, non-fractionated extracts of eggs, etc. (Doenhoff et al., 2004) as diagnostic antigens. However, the usage of crude antigens frequently poses a cross-reaction problem because of those components within the crude antigens shared with other, irrelevant pathogens (Jin et al., 2010). Besides, antibody-based serological assays are not quantitative and usually fail to discriminate between previous exposure and current infection. Although it is difficult to eradicate all of these innate drawbacks of these diagnostic techniques, they could be alleviated to some extent by the use of selected recombinant antigens (Doenhoff et al., 2004). Thanks to the enormous amount of gene sequence information procured by the transcriptomics and genomics researches, together with those screening methods including proteomics, immunomics as well as some bioinformatics tools, many promising diagnostic antigens have been identified and their immunogenicity have been testified, vastly enriching the diagnostic antigen pools for schistosomiasis.
Complementary DNA libraries represent a sort of precious transcriptomic resources. Under combined efforts of serological screening and immunological verification, schistosomal antigens with high immunogenicity could be identified in a cDNA library and finally applied to antibody-based immunodiagnosis of schistosomiasis. Zhou et al. (2010) used rabbit sera collected on day 21 post-infection in an antigenic screening of a previously constructed cercariae cDNA library of S. japonicum (Fung et al., 2002). From the identified positive clones, the authors finally selected Sjp40 as an antigen candidate and used time-resolved fluoroimmunoassay (TRFIA) to profile the levels of anti-Sjp40 IgG in the sera of rabbits with different days post-infection. The result showed that compared with the controls at each time interval, since 21 days post-infection, the titer of circulatory anti-Sjp40 IgG in the infected groups started to significantly increase. Thus, it was considered that Sjp40 was a potential antigen for early diagnosis of schistosomiasis.
Likewise, another group of researchers (Zhou et al., 2009) used human saliva, instead of rabbit sera, to screen a cDNA library of S. japonicum eggs, which resulted in the detection of Sj13. The following ELISA assay exhibited 92.50% sensitivity and 92.11% specificity of salivary IgG detection with recombinant Sj13. Given the easy-accessibility and non-invasiveness of the saliva samples, saliva/Sj13 was asserted as a promising alternative to serological test for schistosomiasis. In addition to these two examples, numerous antigens, sorted from the cDNA library established in 2003 (Hu et al., 2003), like SjEFCAB (Lu et al., 2012a), SjE16 (Wang et al., 2003), Sj myophilin-like protein (Peng et al., 2008), P7 antigen (Xu et al., 2011), etc. have been cloned and expressed in E. coli prokaryotic expression system even earlier and their immunogenicity and diagnostic potential were also investigated. Despite of their considerable sensitivity and specificity as diagnostic antigens, since most of them were published in Chinese journals, their significance thereby waned.
Besides cDNA libraries, schistosomal genomes stand for another kind of resources to search potential indirect diagnostic antigens for schistosomiasis. Using bioinformatics screening, another group succeeded in identifying several novel diagnostic antigen candidates from the previously sequenced S. mansoni genome (Berriman et al., 2009). Overall, four criteria were used in the in silico analysis strategy, including expression in all parasite life stage within the definitive host, extracellular or plasmatic membrane localization, low similarity to human and other helminthic proteins and presence of predicted B cell epitopes (Carvalho et al., 2011). As a result, six promising diagnostic antigens (Sm200, Sm12.8, Sm43.5, Sm127.9, Sm18.9, Sm16.5) were elicited from 13,197 transcripts described for S. mansoni (Berriman et al., 2009). Ensuing verification steps including 1D- and 2D-Western blotting using schistosomula antigen preparation, adult worm preparation and sera from infected mice indicated the good immunogenicity of those predicted antigens.
Furthermore, bioinformatics techniques also served as screening tools to seek antigen candidates for indirect immunological diagnosis in the S. japonicum genome (Angeles et al., 2011). A group of antigens termed tandem repeat (TR) proteins were targeted in this study because of their role in humoral responses known previously (Kim et al., 2001; Goto et al., 2010). Specifically speaking, TR genes were identified from S. japonicum genome by a program named Tandem Repeats Finder. A total of 12,657 gene sequences were analyzed and 134 genes were found to have TR regions. Four TR genes, i.e.,Sj1TR, Sj2TR, Sj4TR, and Sj7TR were selected for further research from the top 20 hits elicited by in silico screening based on their biochemical properties, conservation with other organisms, expression evidence, etc. Their recombinant proteins in expected size were successfully expressed and purified and ELISA was performed using sera from healthy people in endemic or non-endemic countries, post-treated individuals, stool-confirmed patients and patients of other parasitic diseases. Sj7TR had the highest sensitivity (85.71%), while both Sj1TR and Sj7TR had 100% specificity. More importantly, no cross-reaction with sera from patients infected by other pathogens was detected among those TR proteins. In contrast, the crude antigen of S. japonicum, SEA, had a higher sensitivity of 97.14% compared with the recombinant antigens but its specificity was poor (71.76%) and cross-reaction emerged with Paragonimus westermani, Opisthorchis viverrini, and Entamoeba histolytica-positive samples. In conclusion, Sj7TR was a promising candidate antigen for diagnosis purpose found by this research.
In another study, immunomics methods were also employed to uncover prospective diagnosis antigens for schistosomiasis (Zhong et al., 2010). In the beginning, more than 30 immunodominant spots were recognized by pooled sera from infected rabbits with Western blotting, 10 of which were precisely matched to the homologous 2D-gel. LC/MS-MS was then adopted to identify those 10 spots and they were found to correspond to four distinct proteins. Two of the four identified proteins that had not been investigated before, namely, SjLAP, a complex of metallopeptidases and SjFBPA, a central glycolytic enzyme were successfully cloned and the recombinant protein products were further applied in the diagnosis of Schistosomiasis japonica by ELISA, which yielded sensitivities of 98.1 and 87.8% for acute and chronic schistosomiasis with rSjLAP and 100 and 84.7% with rSjFBPA, respectively, whereas the specificities were 96.7% for both antigens. Moreover, both antigen ELISA assays showed more than 80% sensitivity in diagnosis of chronic schistosomiasis with a low intensity infection and significantly declined antibody titers after the treatment with PZQ. Thus, rSjLAP and rSjFBPA proved to be useful diagnostic antigens for S. japonicum infection.
Antigen-Based Direct Immunological Diagnosis
Early immunoassays have shown that schistosome-derived antigens, such as adult worm antigen (AWA), soluble egg antigen (SEA), and circulating antigen (CA; Zhao et al., 2012) could be released into the host circulatory system/excreta by schistosomes, which facilitates the researches on the direct immunological diagnosis of schistosomiasis. Circulating anodic antigen (CAA) and circulating cathodic antigen (CCA) are by far the most extensively used antigens for the antigen-based immunodiagnosis of schistosomiasis and methods for the detection of CAs encompass sandwich ELISA (De Jonge et al., 1988), IHA (Deelder et al., 1989), time-resolved immunofluorometric assay (TR-IFMA; De Jonge et al., 1989), magnetic bead immunoassay (Gundersen et al., 1992), reagent strips (Van Etten et al., 1994), and liquid-phase piezoelectric immunosensor (LP-PEIS; Wen et al., 2011). Compared with the two foregoing diagnostic methods, the detection of CA is more suitable for drug efficacy trials owing to its high specificity and the ability to discriminate between previous exposure and current infection. Moreover, CAA and CCA can be readily detected in urine (Van Dam et al., 2004), which is easier and less invasive to be collected than blood required for antibody detection. Nonetheless, low sensitivity (Van Lieshout et al., 1995, 2000), failing to distinguish different species (Agnew et al., 1995) are disadvantages associated with this method.
Research conducted by Lu et al. (2012b) becomes the first and currently the sole immunomics-based study in an attempt to profile CAs in Schistosoma spp. In this study, AWA was initially prepared using S. japonicum worms collected from infected rabbits and then employed to subcutaneously immunize the hyline hens so as to produce anti-AWA IgY. The IgY antibody contained in the egg yolk was purified by water-dilution and ammonium sulfate precipitation approach and characterized by ELISA and Western blotting. Afterwards, purified IgY was immobilized onto aldehyde-activated beaded agarose resin and served as a capture antibody to immunoprecipitate and enrich CAs within the sera of patients. At last, the obtained CAs were separated by 1D electrophoresis and analyzed by LC-MS/MS, which lead to the identification of four proteins, i.e., protein BUD31 homolog, ribonuclease, SJCHGC06971 protein, and SJCHGC04754 protein. The following analysis indicated that those four CAs belonged to neither CAA nor CCA and had not been reported from previous proteomics researches. CAs revealed by corresponding -omics researches, just like these four, ought to be cloned and expressed in the future and the produced mAbs could be applied for antigen detection through sandwich ELISA which stands for a promising way to overcome the shortcomings of the current direct immunodiagnostic methods.
In addition to this finished work, one recent review about proteomics at the schistosome–mammalian host interface (Wilson, 2012) also proposed several candidate antigens that can be potentially used for CA detection, including serpin, α2 macroglobulin, Am200, LMWP, as well as some MEG-2 and MEG-3 proteins secreted by eggs.
Nucleic Acid Amplification-Based Molecular Methods
Given the existence of schistosomal DNA in the serum and other tissue samples of the host derived from dead worms, tegument shedding or inactive eggs (Xu et al., 2013), PCR-based molecular diagnosis has become a promising alternative to overcome the innate shortcomings of etiological and immunological diagnosis for schistosomiasis. Since the publication of the proof-of-concept study in 2002 (Pontes et al., 2002), PCR-based molecular diagnosis has been applied to the detection of schistosomes in multiple diagnostic field works (Pontes et al., 2003; Obeng et al., 2008; Enk et al., 2012) and several novel technologies, such as real time PCR (RT-PCR; Lier et al., 2006), PCR-ELISA (Gomes et al., 2010), and loop-mediated isothermal amplification (LAMP; Xu et al., 2010), etc. have been added to the tool assemblies in addition to conventional PCR in order to boost the detection sensitivity. Empirical evidence showed that molecular diagnosis of schistosomiasis has both high sensitivity and high specificity, enables detection during larval stage or at least before egg spawning and can use manifold materials, e.g., feces, serum (Pontes et al., 2002), plasma (Wichmann et al., 2009), and urine (Sandoval et al., 2006) as templates, all of which make it superior to the conventional diagnostic approaches. Factors that impact the result of molecular diagnosis are various and the choice of amplified products is definitely among them. However, confined by the inaccessibility to the schistosomal genome sequence, in the pre-genomic era, researchers could only select targets for PCR reactions within limited candidates. Representative examples include two different 121-bp repetitive elements in S. mansoni (Hamburger et al., 1991) or S. haematobium (Hamburger et al., 2001), S. japonicum 18s rDNA (Zhou et al., 2011a), highly repetitive retrotransposon SjR2 (Laha et al., 2002), mitochondrial NADH1 gene (nad1) of S. mansoni (Jannotti-Passos et al., 1997) and S. japonicum (Lier et al., 2008), etc.
As early as 2009, 25 novel retrotransposons had been identified along with the decoding of the genome of S. japonicum (The Schistosoma japonicum Genome Sequencing and Functional Analysis Consortium, 2009). Two years later, a large scale selection had been conducted in order to seek suitable target sequences among those 25 retrotransposons to achieve the sensitive and specific diagnosis of S. japonica (Guo et al., 2012). To begin with, primer pairs were designed for the 25 candidates with SjR2, a previously described molecular diagnostic target (Xia et al., 2009; Xu et al., 2010), serving as a positive control. A series of diluted genomic DNA of S. japonicum were used as the templates in a PCR assay to amplify the target fragments and judge the sensitivity of those 25 candidates. The result showed that in addition to SjR2, a new non-LTR retrotransposon named SjCHGCS19, with the PCR product of 303-bp in length, had high sensitivity for detecting S. japonicum DNA. Coincidentally, bioinformatics research showed that both SjCHGCS19 and SjR2 have higher genome proportions, repetitive complete copies and partial copies and more active ESTs compared with other candidates in the S. japonicum genome, which may also outline the general features of ideal targets for PCR diagnosis of schistosomiasis. Besides, further researches showed that SjCHGCS19 has good specificity except for a cross-reaction with S. mansoni and it is also an ideal PCR product for the early diagnosis of schistosomiasis as well as the evaluation of chemotherapy. At last, the clinical utility of SjCHGCS19 was also tested by diagnosing 43 human serum samples from patients of S. japonica and 51 serum samples from healthy individuals. As a result, the sensitivity of 97.67% and the specificity of 96.07% fully validated the effectiveness of the SjCHGCS19-based molecular diagnosis of schistosomiasis.
Parasite-derived miRNAs, comparable to genomic DNA, are also present in multiple biofluids of the host, such as plasma, serum, urine, and saliva (Cortez et al., 2011), which implies their potential as biomarkers for the molecular diagnosis of schistosomiasis. Besides, in response to the invasion of pathogens, some characteristic changes of the host’s miRNAs profile might appear (Delić et al., 2011), which can also serve as hints of infection event. Nowadays, the available sequences of schistosomal miRNAs, in association with those approaches established for the profiling of miRNAs, e.g., stem-loop RT-PCR, miRNA array, and high-throughput sequencing (Vaz et al., 2010) have paved the way for the future miRNA-based molecular diagnosis of schistosomiasis. For detailed information about the diagnostic and therapeutic value of parasite miRNA, see review (Manzano-Román and Siles-Lucas, 2012).
Metabonomics-Based Biomarker Discovery
As mentioned above, early schistosomal metabonomics focused on the dynamics of biochemical responses associated with S. japonicum or S. mansoni infection in multiple biofluids of rodent models. Corresponding progresses made thus far have been reviewed in several papers (Legido-Quigley, 2010; Wang et al., 2010a; Utzinger et al., 2011). To sum up, the metabolic fingerprint of rodents in response to schistosomes infection can be concluded as: reduced levels of the tricarboxylic acid cycle intermediates, stimulated glycolysis, disturbed amino acid metabolism, disturbances in the gut microbiota as well as a phenomenon unique to S. japonicum infection, i.e., the inhibition of short-chain fatty acids. Despite the considerable progresses achieved in rodent models, clinical diagnosis of schistosomiasis based on metabonomics still lags behind other -omics methods, because after all, the physiological mechanisms between rodents in the laboratories and humans are far from similar. However, the newly published metabonomic work about human schistosomiasis (Balog et al., 2011) has laid a solid foundation for the clinical application of metabonomic outcomes in the future. Prospective molecular signatures of human S. mansoni infection found by this study were primarily related to changes in gut microflora, energy metabolism, and liver function, in line with the earlier findings in experimental animals.
Future Diagnostic Tools of Schistosomiasis and Advice to Better Harness -Omics Achievements
In our view, future diagnostic tools of schistosomiasis ought to follow three main developmental trends: (1) Field-applicability, test kit protocols should be robust, easy to follow, and produce results within a few minutes without the aid of extra instruments. (2) Non-invasiveness, diagnostic tools should detect specific biomarkers from easy accessible biofluids, e.g., sweat, urea, or saliva. (3) High-throughput, diagnostic kits in the next generation should integrate multiple probes and enable parallel identification of different pathogens.
“-Omics” studies result in a treasure trove of molecular data on genes, proteins, and metabolites. However, identification of effective diagnostic biomarkers from these data pools requires bioinformatics tools based on certain criteria. Further summarization of the general features shared by those currently available diagnostic targets will undoubtedly help us optimize the algorithms for biomarker selection and thereby better utilize the information generated by -omics studies. Besides, some molecular candidates of great diagnostic value are able to be sorted directly by -omics tools based on either their positive interactions with certain biofluids of patients or the changes of their presences or concentrations in response to parasite infection. Schistosomal samples and/or biofluids of patients used in such researches should be chosen and combined according to the prospective utility of a diagnostic tool. Immunomics and metabonomics studies using distinct biological samples should be rigorously designed and carried out in future, in an attempt to seek more diagnostic biomarkers that satisfy variant requirements, e.g., to distinguish infection by different Schistosoma species and/or with different worm burden, to discriminate between current infection and previous exposure, etc. Among all available body fluids to date, priority should be given to those that can be sampled without invasive procedures in order to achieve a rapid and safe diagnosis. Last but not least, we should also pay attention to the achievements of those newly established -omics tools, so as to enrich our arsenal of schistosomiasis diagnosis with innovative biomarkers.
Progress in the control and prevention of parasitic diseases is expected to be more rapid and efficient thanks to the achievements of “-omics” researches, benefitting the health of humans and animals alike.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Agnew, A., Fulford, A., De Jonge, N., Krijger, F., Rodriguez-Chacon, M., Gutsmann, V.,et al. (1995). The relationship between worm burden and levels of a circulating antigen (CAA) of five species of Schistosoma in mice. Parasitology 111, 67–76. doi: 10.1017/S0031182000064611
Alger, H. M., and Williams, D. L. (2002). The disulfide redox system of Schistosoma mansoni and the importance of a multifunctional enzyme, thioredoxin glutathione reductase. Mol. Biochem. Parasitol. 121, 129–139. doi: 10.1016/S0166-6851(02)00031-2
Angeles, J. M., Goto, Y., Kirinoki, M., Leonardo, L., Tongol-Rivera, P., Villacorte, E.,et al. (2011). Human antibody response to thioredoxin peroxidase-1 and tandem repeat proteins as immunodiagnostic antigen candidates for Schistosoma japonicum infection. Am. J. Trop. Med. Hyg. 85, 674–679. doi: 10.4269/ajtmh.2011.11-0245
Aragon, A. D., Imani, R. A., Blackburn, V. R., and Cunningham, C. (2008). Microarray based analysis of temperature and oxidative stress induced messenger RNA in Schistosoma mansoni. Mol. Biochem. Parasitol. 162, 134–141. doi: 10.1016/j.molbiopara.2008.08.004
Aragon, A. D., Imani, R. A., Blackburn, V. R., Cupit, P. M., Melman, S. D., Goronga, T.,et al. (2009). Towards an understanding of the mechanism of action of praziquantel. Mol. Biochem. Parasitol. 164, 57–65. doi: 10.1016/j.molbiopara.2008.11.007
Balog, C. I., Alexandrov, T., Derks, R. J., Hensbergen, P. J., Van Dam, G. J., Tukahebwa, E. M.,et al. (2010). The feasibility of MS and advanced data processing for monitoring Schistosoma mansoni infection. Proteomics Clin. Appl. 4, 499–510. doi: 10.1002/prca.200900158
Balog, C. I., Meissner, A., Göraler, S., Bladergroen, M. R., Vennervald, B. J., Mayboroda, O. A.,et al. (2011). Metabonomic investigation of human Schistosoma mansoni infection. Mol. Biosyst. 7, 1473–1480. doi: 10.1039/C0MB00262C
Bartel, D. P. (2004). MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297. doi: 10.1016/S0092–8674(04)00045-5
Berriman, M., Haas, B. J., Loverde, P. T., Wilson, R. A., Dillon, G. P., Cerqueira, G. C.,et al. (2009). The genome of the blood fluke Schistosoma mansoni. Nature 460, 352–358. doi: 10.1038/nature08160
Braschi, S., Curwen, R. S., Ashton, P. D., Verjovski-Almeida, S., and Wilson, A. (2006). The tegument surface membranes of the human blood parasite Schistosoma mansoni: a proteomic analysis after differential extraction. Proteomics 6, 1471–1482. doi: 10.1002/pmic.200500368
Braschi, S., and Wilson, R. A. (2006). Proteins exposed at the adult schistosome surface revealed by biotinylation. Mol. Cell. Proteomics 5, 347–356. doi: 10.1074/mcp.M500287-MCP200
Burke, M. L., Mcmanus, D. P., Ramm, G. A., Duke, M., Li, Y., Jones, M. K.,et al. (2010). Temporal expression of chemokines dictates the hepatic inflammatory infiltrate in a murine model of schistosomiasis. PLoS Negl. Trop. Dis. 4:e598. doi: 10.1371/journal.pntd.0000598
Cai, P., Hou, N., Piao, X., Liu, S., Liu, H., Yang, F.,et al. (2011). Profiles of small non-coding RNAs in Schistosoma japonicum during development. PLoS Negl. Trop. Dis. 5:e1256. doi: 10.1371/journal.pntd.0001256
Cai, P., Piao, X., Hou, N., Liu, S., Wang, H., and Chen, Q. (2012). Identification and characterization of argonaute protein, Ago2 and its associated small RNAs in Schistosoma japonicum. PLoS Negl. Trop. Dis. 6:e1745. doi: 10.1371/journal.pntd.0001745
Carvalho, G. B. F., Silva-Pereira, R. A. D., Pacífico, L. G. G., and Fonseca, C. T. (2011). Identification of Schistosoma mansoni candidate antigens for diagnosis of schistosomiasis. Mem. Inst. Oswaldo Cruz 106, 837–843. doi: 10.1590/S0074–02762011000700009
Castro-Borges, W., Dowle, A., Curwen, R. S., Thomas-Oates, J., and Wilson, R. A. (2011). Enzymatic shaving of the tegument surface of live schistosomes for proteomic analysis: a rational approach to select vaccine candidates. PLoS Negl. Trop. Dis. 5:e993. doi: 10.1371/journal.pntd.0000993
Chai, M., Mcmanus, D., Mcinnes, R., Moertel, L., Tran, M., Loukas, A.,et al. (2006). Transcriptome profiling of lung schistosomula, in vitro cultured schistosomula and adult Schistosoma japonicum. Cell Mol. Life Sci. 63, 919–929. doi: 10.1007/s00018-005-5578-1
Cheng, G., and Jin, Y. (2012). MicroRNAs: potentially important regulators for schistosome development and therapeutic targets against schistosomiasis. Parasitology 139, 669–679. doi: 10.1017/S0031182011001855
Cheng, G. F., Lin, J. J., Feng, X. G., Fu, Z. Q., Jin, Y. M., Yuan, C. X.,et al. (2005). Proteomic analysis of differentially expressed proteins between the male and female worm of Schistosoma japonicum after pairing. Proteomics 5, 511–521. doi: 10.1002/pmic.200400953
Copeland, C., Marz, M., Rose, D., Hertel, J., Brindley, P., Santana, C.,et al. (2009). Homology-based annotation of non-coding RNAs in the genomes of Schistosoma mansoni and Schistosoma japonicum. BMC Genomics 10:464. doi: 10.1186/1471–2164-10-464
Cortez, M. A., Bueso-Ramos, C., Ferdin, J., Lopez-Berestein, G., Sood, A. K., and Calin, G. A. (2011). MicroRNAs in body fluids–the mix of hormones and biomarkers. Nat. Rev. Clin. Oncol. 8, 467–477. doi: 10.1038/nrclinonc.2011.76
Cringoli, G. (2006). FLOTAC, a novel apparatus for a multivalent faecal egg count technique. Parassitologia 48, 381–384.
Curwen, R. S., Ashton, P. D., Johnston, D. A., and Wilson, R. A. (2004). The Schistosoma mansoni soluble proteome: a comparison across four life-cycle stages. Mol. Biochem. Parasitol. 138, 57–66. doi: 10.1016/j.molbiopara.2004.06.016
Curwen, R. S., Ashton, P. D., Sundaralingam, S., and Wilson, R. A. (2006). Identification of novel proteases and immunomodulators in the secretions of schistosome cercariae that facilitate host entry. Mol. Cell. Proteomics 5, 835–844. doi: 10.1074/mcp.M500313-MCP200
De Jonge, N., Boerman, O., and Deelder, A. (1989). Time-resolved immunofluorometric assay (TR-IFMA) for the detection of the schistosome circulating anodic antigen. Trans. R. Soc. Trop. Med. Hyg. 83, 659–663. doi: 10.1016/0035-9203(89)90391-X
De Jonge, N., Gryseels, B., Hilberath, G. W., Polderman, A. M., and Deelder, A. M. (1988). Detection of circulating anodic antigen by ELISA for seroepidemiology of Schistosomiasis mansoni. Trans. R. Soc. Trop. Med. Hyg. 82, 591–594. doi: 10.1016/0035-9203(88)90523-8.
de Moraes Mourão, M., Dinguirard, N., Franco, G. R., and Yoshino, T. P. (2009). Role of the endogenous antioxidant system in the protection of Schistosoma mansoni primary sporocysts against exogenous oxidative stress. PLoS Negl. Trop. Dis. 3:e550. doi: 10.1371/journal.pntd.0000550
de Souza Gomes, M., Muniyappa, M. K., Carvalho, S. G., Guerra-Sa, R., and Spillane, C. (2011). Genome-wide identification of novel microRNAs and their target genes in the human parasite Schistosoma mansoni. Genomics 98, 96–111. doi: 10.1016/j.ygeno.2011.05.007
Deelder, A. M., De Jonge, N., Fillié, Y. E., Kornelis, D., Helaha, D., Qian, Z.-L.,et al. (1989). Quantitative determination of circulating antigens in human Schistosomiasis mansoni using an indirect hemagglutination assay. Am. J. Trop. Med. Hyg. 40, 50–54.
Delcroix, M., Medzihradsky, K., Caffrey, C. R., Fetter, R. D., and Mckerrow, J. H. (2007). Proteomic analysis of adult S. mansoni gut contents. Mol. Biochem. Parasitol. 154, 95–97. doi: 10.1016/j.molbiopara.2007.03.008
Delić, D., Dkhil, M., Al-Quraishy, S., and Wunderlich, F. (2011). Hepatic miRNA expression reprogrammed by Plasmodium chabaudi malaria. Parasitol. Res. 108, 1111–1121. doi: 10.1007/s00436-010-2152-z
Dillon, G. P., Feltwell, T., Skelton, J., Coulson, P. S., Wilson, R. A., and Ivens, A. C. (2008). Altered patterns of gene expression underlying the enhanced immunogenicity of radiation-attenuated schistosomes. PLoS Negl. Trop. Dis. 2:e240. doi: 10.1371/journal.pntd.0000240
Dillon, G. P., Feltwell, T., Skelton, J. P., Ashton, P. D., Coulson, P. S., Quail, M. A.,et al. (2006). Microarray analysis identifies genes preferentially expressed in the lung schistosomulum of Schistosoma mansoni. Int. J. Parasitol. 36, 1–8. doi: 10.1016/j.ijpara.2005.10.008
Doenhoff, M. J., Chiodini, P. L., and Hamilton, J. V. (2004). Specific and sensitive diagnosis of schistosome infection: can it be done with antibodies? Trends Parasitol. 20, 35–39. doi: 10.1016/j.pt.2003.10.019
Driguez, P., Doolan, D. L., Loukas, A., Felgner, P. L., and Mcmanus, D. P. (2010). Schistosomiasis vaccine discovery using immunomics. Parasit. Vectors 3:4. doi: 10.1186/1756-3305-3–4
Enk, M. J., Amaral, G. L., Costa E Silva, M. F., Silveira-Lemos, D., Teixeira-Carvalho, A., Martins-Filho, O. A.,et al. (2010). Rural tourism: a risk factor for schistosomiasis transmission in Brazil. Mem. Inst. Oswaldo Cruz 105, 537–540. doi: 10.1590/S0074-02762010000400032
Enk, M. J., E Silva, G. O., and Rodrigues, N. B. (2012). Diagnostic accuracy and applicability of a PCR system for the detection of Schistosoma mansoni DNA in human urine samples from an endemic area. PLoS ONE 7:e38947. doi: 10.1371/journal.pone.0038947
Fan, J., Minchella, D. J., Day, S. R., Mcmanus, D. P., Tiu, W. U., and Brindley, P. J. (1998). Generation, identification, and evaluation of expressed sequence tags from different developmental stages of the Asian blood fluke Schistosoma japonicum. Biochem. Biophys. Res. Commun. 252, 348–356. doi: 10.1006/bbrc.1998.9491
Fitzpatrick, J. M., Johansen, M. V., Johnston, D. A., Dunne, D. W., and Hoffmann, K. F. (2004). Gender-associated gene expression in two related strains of Schistosoma japonicum. Mol. Biochem. Parasitol. 136, 191–209. doi: 10.1016/j.molbiopara.2004.03.014
Fitzpatrick, J. M., Johnston, D. A., Williams, G. W., Williams, D. J., Freeman, T. C., Dunne, D. W.,et al. (2005). An oligonucleotide microarray for transcriptome analysis of Schistosoma mansoni and its application/use to investigate gender-associated gene expression. Mol. Biochem. Parasitol. 141, 1–13. doi: 10.1016/j.molbiopara.2005.01.007
Fitzpatrick, J. M., Peak, E., Perally, S., Chalmers, I. W., Barrett, J., Yoshino, T. P.,et al. (2009). Anti-schistosomal intervention targets identified by lifecycle transcriptomic analyses. PLoS Negl. Trop. Dis. 3:e543. doi: 10.1371/journal.pntd.0000543
Fung, M. C., Lau, M. T., and Chen, X. G. (2002). Expressed sequence tag (EST) analysis of a Schistosoma japonicum cercariae cDNA library. Acta Trop. 82, 215–224. doi: 10.1016/S0001-706X(02)00013-X
Garcia-Perez, I., Angulo, S., Utzinger, J., Holmes, E., Legido-Quigley, C., and Barbas, C. (2010). Chemometric and biological validation of a capillary electrophoresis metabolomic experiment of Schistosoma mansoni infection in mice. Electrophoresis 31, 2338–2348. doi: 10.1002/elps.200900523
Garcia-Perez, I., Couto Alves, A., Angulo, S., Li, J., Utzinger, J., Ebbels, T.,et al. (2009). Bidirectional correlation of NMR and capillary electrophoresis fingerprints: a new approach to investigating Schistosoma mansoni infection in a mouse model. Anal. Chem. 82, 203–210. doi: 10.1021/ac901728w
Garcia-Perez, I., Whitfield, P., Bartlett, A., Angulo, S., Legido-Quigley, C., Hanna-Brown, M.,et al. (2008). Metabolic fingerprinting of Schistosoma mansoni infection in mice urine with capillary electrophoresis. Electrophoresis 29, 3201–3206. doi: 10.1002/elps.200800031
Gobert, G. N. (2010). Applications for profiling the schistosome transcriptome. Trends Parasitol. 26, 434–439. doi: 10.1016/j.pt.2010.04.009
Gobert, G. N., and Jones, M. K. (2008). Discovering new schistosome drug targets: the role of transcriptomics. Curr. Drug Targets 9, 922–930. doi: 10.2174/138945008786786136
Gobert, G. N., Mcinnes, R., Moertel, L., Nelson, C., Jones, M. K., Hu, W.,et al. (2006). Transcriptomics tool for the human Schistosoma blood flukes using microarray gene expression profiling. Exp. Parasitol. 114, 160–172. doi: 10.1016/j.exppara.2006.03.003
Gobert, G. N., Mcmanus, D. P., Nawaratna, S., Moertel, L., Mulvenna, J., and Jones, M. K. (2009a). Tissue specific profiling of females of Schistosoma japonicum by integrated laser microdissection microscopy and microarray analysis. PLoS Negl. Trop. Dis. 3:e469. doi: 10.1371/journal.pntd.0000469
Gobert, G. N., Moertel, L., Brindley, P. J., and Mcmanus, D. P. (2009b). Developmental gene expression profiles of the human pathogen Schistosoma japonicum. BMC Genomics 10:128. doi: 10.1186/1471-2164-10-128
Gobert, G. N., Tran, M. H., Moertel, L., Mulvenna, J., Jones, M. K., Mcmanus, D. P.,et al. (2010). Transcriptional changes in Schistosoma mansoni during early schistosomula development and in the presence of erythrocytes. PLoS Negl. Trop. Dis. 4:e600. doi: 10.1371/journal.pntd.0000600
Gomes, L. I., Dos Santos Marques, L. H., Enk, M. J., De Oliveira, M. C., Coelho, P. M. Z., and Rabello, A. (2010). Development and evaluation of a sensitive PCR-ELISA system for detection of Schistosoma infection in feces. PLoS Negl. Trop. Dis. 4:e664. doi: 10.1371/journal.pntd.0000664
Goto, Y., Carter, D., Guderian, J., Inoue, N., Kawazu, S.-I., and Reed, S. G. (2010). Upregulated expression of B-cell antigen family tandem repeat proteins by Leishmania amastigotes. Infect. Immun. 78, 2138–2145. doi: 10.1128/IAI.01102-09
Gundersen, S., Haagensen, I., Jonassen, T., Figenschau, K., De Jonge, N., and Deelder, A. (1992). Magnetic bead antigen capture enzyme-linked immunoassay in microtitre trays for rapid detection of schistosomal circulating anodic antigen. J. Immunol. Methods 148, 1–8. doi: 10.1016/0022-1759(92)90151-I
Guo, J.-J., Zheng, H.-J., Xu, J., Zhu, X.-Q., Wang, S.-Y., and Xia, C.-M. (2012). Sensitive and specific target sequences selected from retrotransposons of Schistosoma japonicum for the diagnosis of schistosomiasis. PLoS Negl. Trop. Dis. 6:e1579. doi: 10.1371/journal.pntd.0001579
Hamburger, J., Abbasi, I., Ramzy, R. M., Jourdane, J., and Ruppel, A. (2001). Polymerase chain reaction assay based on a highly repeated sequence of Schistosoma haematobium: a potential tool for monitoring schistosome-infested water. Am. J. Trop. Med. Hyg. 65, 907-911.
Hamburger, J., Turetski, T., Kapeller, I., and Deresiewicz, R. (1991). Highly repeated short DNA sequences in the genome of Schistosoma mansoni recognized by a species-specific probe. Mol. Biochem. Parasitol. 44, 73–80. doi: 10.1016/0166-6851(91)90222-R
Hao, L., Cai, P., Jiang, N., Wang, H., and Chen, Q. (2010). Identification and characterization of microRNAs and endogenous siRNAs in Schistosoma japonicum. BMC Genomics 11:55. doi: 10.1186/1471-2164-11-55
Hoffmann, K. F., and Fitzpatrick, J. M. (2004). Gene expression studies using self-fabricated parasite cDNA microarrays. Methods Mol. Biol. 270, 219–236. doi: 10.1385/1-59259-793-9:219
Hoffmann, K. F., Johnston, D. A., and Dunne, D. W. (2002). Identification of Schistosoma mansoni gender-associated gene transcripts by cDNA microarray profiling. Genome Biol. 3:41. doi: 10.1186/gb-2002-3-8-research0041
Hoffmann, K. F., Mccarty, T. C., Segal, D. H., Chiaramonte, M., Hesse, M., Davis, E. M.,et al. (2001). Disease fingerprinting with cDNA microarrays reveals distinct gene expression profiles in lethal type 1 and type 2 cytokine-mediated inflammatory reactions. FASEB J. 15, 2545–2547. doi: 10.1096/fj.01-0306fje
Hokke, C. H., Fitzpatrick, J. M., and Hoffmann, K. F. (2007). Integrating transcriptome, proteome and glycome analyses of Schistosoma biology. Trends Parasitol. 23, 165–174. doi: 10.1016/j.pt.2007.02.007
Hu, S., Law, P. K., and Fung, M. C. (2009). Microarray analysis of genes highly expressed in cercarial stage of Schistosoma japonicum and the characterization of the antigen Sj20H8. Acta Trop. 112, 26–32. doi: 10.1016/j.actatropica.2009.06.001
Hu, W., Brindley, P. J., Mcmanus, D. P., Feng, Z., and Han, Z. G. (2004). Schistosome transcriptomes: new insights into the parasite and schistosomiasis. Trends Mol. Med. 10, 217–225. doi: 10.1016/j.molmed.2004.03.002
Hu, W., Yan, Q., Shen, D. K., Liu, F., Zhu, Z. D., Song, H. D.,et al. (2003). Evolutionary and biomedical implications of a Schistosoma japonicum complementary DNA resource. Nat. Genet. 35, 139–147. doi: 10.1038/ng1236
Huang, H.-H., Tsai, P.-L., and Khoo, K.-H. (2001). Selective expression of different fucosylated epitopes on two distinct sets of Schistosoma mansoni cercarial O-glycans: identification of a novel core type and Lewis X structure. Glycobiology 11, 395–406. doi: 10.1093/glycob/11.5.395
Jannotti-Passos, L., Vidigal, T., Dias-Neto, E., Pena, S., Simpson, A., Dutra, W.,et al. (1997). PCR amplification of the mitochondrial DNA minisatellite region to detect Schistosoma mansoni infection in Biomphalaria glabrata snails. J. Parasitol. 83, 395–399. doi: 10.2307/3284400
Jin, Y. M., Lu, K., Zhou, W. F., Fu, Z. Q., Liu, J. M., Shi, Y. J.,et al. (2010). Comparison of recombinant proteins from Schistosoma japonicum for schistosomiasis diagnosis. Clin. Vaccine Immunol. 17, 476–480. doi: 10.1128/cvi.00418-09
Jolly, E., Chin, C.-S., Miller, S., Bahgat, M., Lim, K., Derisi, J.,et al. (2007). Gene expression patterns during adaptation of a helminth parasite to different environmental niches. Genome Biol. 8:R65. doi: 10.1186/gb-2007-8-4-r65
Jones, M. K., Higgins, T., Stenzel, D. J., and Gobert, G. N. (2007). Towards tissue specific transcriptomics and expression pattern analysis in schistosomes using laser microdissection microscopy. Exp. Parasitol. 117, 259–266. doi: 10.1016/j.exppara.2007.06.004
Ju, C., Feng, Z., Brindley, P. J., Mcmanus, D. P., Han, Z., Peng, J.-X.,et al. (2010). Our wormy world: genomics, proteomics and transcriptomics in East and Southeast Asia. Adv. Parasitol. 73, 327–371. doi: 10.1016/S0065-308X(10)73011-6
Jurberg, A. D., Oliveira, A. D., Lenzi, H. L., and Coelho, P. M. Z. (2008). A new miracidia hatching device for diagnosing schistosomiasis. Mem. Inst. Oswaldo Cruz 103, 112–114. doi: 10.1590/S0074-02762008005000005
Katz, N., Chaves, A., and Pellegrino, J. (1972). A simple device for quantitative stool thick-smear technique in Schistosomiasis mansoni. Rev. Inst. Med. Trop. Sao Paulo 14, 397–400.
Khoo, K.-H., Chatterjee, D., Caulfield, J. P., Morris, H. R., and Dell, A. (1997a). Structural mapping of the glycans from the egg glycoproteins of Schistosoma mansoni and Schistosoma japonicum: identification of novel core structures and terminal sequences. Glycobiology 7, 663–677. doi: 10.1093/glycob/7.5.663
Khoo, K. H., Chatterjee, D., Caulfield, J. P., Morris, H. R., and Dell, A. (1997b). Structural characterization of glycosphingolipids from the eggs of Schistosoma mansoni and Schistosoma japonicum. Glycobiology 7, 653–661. 10.1093/glycob/7.5.653
Khoo, K.-H., Huang, H.-H., and Lee, K.-M. (2001). Characteristic structural features of schistosome cercarial N-glycans: expression of Lewis X and core xylosylation. Glycobiology 11, 149–163. doi: 10.1093/glycob/11.2.149
Khoo, K.-H., Sarda, S., Xu, X., Caulfield, J. P., Mcneil, M. R., Homans, S. W.,et al. (1995). A unique multifucosylated-3GalNAcβ14GlcNAcβ13Galα1-motif constitutes the repeating unit of the complex O-glycans derived from the cercarial glycocalyx of Schistosoma mansoni. J. Biol. Chem. 270, 17114–17123. doi: 10.1074/jbc.270.29.17114
Kim, T. Y., Kang, S.-Y., Ahn, I.-Y., Cho, S.-Y., and Hong, S.-J. (2001). Molecular cloning and characterization of an antigenic protein with a repeating region from Clonorchis sinensis. Korean J. Parasitol. 39, 57–66. doi: 10.3347/kjp.2001.39.1.57
Knudsen, G. M., Medzihradszky, K. F., Lim, K.-C., Hansell, E., and Mckerrow, J. H. (2005). Proteomic analysis of Schistosoma mansoni cercarial secretions. Mol. Cell. Proteomics 4, 1862–1875. doi: 10.1074/mcp.M500097-MCP200
Laha, T., Brindley, P. J., Smout, M. J., Verity, C. K., Mcmanus, D. P., and Loukas, A. (2002). Reverse transcriptase activity and untranslated region sharing of a new RTE-like, non-long terminal repeat retrotransposon from the human blood fluke, Schistosoma japonicum. Int. J. Parasitol. 32, 1163–1174. doi: 10.1016/S0020-7519(02)00063-2
Le, T., Blair, D., and Mcmanus, D. (2002). Revisiting the question of limited genetic variation within Schistosoma japonicum. Ann. Trop. Med. Parasitol. 96, 155–164. doi: 10.1179/000349802125000268
Legido-Quigley, C. (2010). Metabolite-biomarker investigations in the life cycle of and infection with Schistosoma. Parasitology 137:1425. doi: 10.1017/S0031182010000545
Li, J. V., Holmes, E., Saric, J., Keiser, J., Dirnhofer, S., Utzinger, J.,et al. (2009). Metabolic profiling of a Schistosoma mansoni infection in mouse tissues using magic angle spinning-nuclear magnetic resonance spectroscopy. Int. J. Parasitol. 39, 547–558. doi: 10.1016/j.ijpara.2008.10.010
Lier, T., Johansen, M. V., Hjelmevoll, S., Vennervald, B. J., and Simonsen, G. (2008). Real-time PCR for detection of low intensity Schistosoma japonicum infections in a pig model. Acta Trop. 105, 74–80. doi: 10.1016/j.actatropica.2007.10.004
Lier, T., Simonsen, G., Haaheim, H., Hjelmevoll, S., Vennervald, B., and Johansen, M. (2006). Novel real-time PCr for detection of Schistosoma japonicum in stool. Southeast Asian J. Trop. Med. Public Health 37, 257–264.
Liu, F., Cui, S.-J., Hu, W., Feng, Z., Wang, Z.-Q., and Han, Z.-G. (2009). Excretory/secretory proteome of the adult developmental stage of human blood fluke, Schistosoma japonicum. Mol. Cell. Proteomics 8, 1236–1251. doi: 10.1074/mcp.M800538-MCP200
Liu, F., Lu, J., Hu, W., Wang, S. Y., Cui, S. J., Chi, M.,et al. (2006). New perspectives on host–parasite interplay by comparative transcriptomic and proteomic analyses of Schistosoma japonicum. PLoS Pathog. 2:e29. doi: 10.1371/journal.ppat.0020029
Lu, Y., Xu, B., Ju, C., Mo, X., Chen, S., Feng, Z.,et al. (2012a). Cloning, expression and immunodiagnostic analysis of Schistosoma japonicum calcium-binding EF-hand domain containing protein. Chin. J. Parasitol. Parasit. Dis. 30, 165–169.
Lu, Y., Xu, B., Ju, C., Mo, X., Chen, S., Feng, Z.,et al. (2012b). Identification and profiling of circulating antigens by screening with the sera from Schistosomiasis japonica patients. Parasit. Vectors 5, 115. doi: 10.1186/1756-3305-5-115
Manzano-Román, R., and Siles-Lucas, M. (2012). MicroRNAs in parasitic diseases: Potential for diagnosis and targeting. Mol. Biochem. Parasitol. 186, 81–86. doi: 10.1016/j.molbiopara.2012.10.001
Merrick, J. M., Osman, A., Tsai, J., Quackenbush, J., Loverde, P. T., and Lee, N. H. (2009). The Schistosoma mansoni gene index: gene discovery and biology by reconstruction and analysis of expressed gene sequences. J. Parasitol. 89, 261–269. doi: 10.1645/0022-3395 (2003)089[0261:TSMGIG]2.0.CO;2
Moertel, L., Mcmanus, D. P., Piva, T. J., Young, L., Mcinnes, R. L., and Gobert, G. N. (2006). Oligonucleotide microarray analysis of strain- and gender-associated gene expression in the human blood fluke, Schistosoma japonicum. Mol. Cell. Probes 20, 280–289. doi: 10.1016/j.mcp.2006.02.002
Molnar, A., Melnyk, C. W., Bassett, A., Hardcastle, T. J., Dunn, R., and Baulcombe, D. C. (2010). Small silencing RNAs in plants are mobile and direct epigenetic modification in recipient cells. Sci. Signal. 328, 872–875. doi: 10.1126/science.1187959
Mott, K., Baltes, R., Bambagha, J., and Baldassini, B. (1982). Field studies of a reusable polyamide filter for detection of Schistosoma haematobium eggs by urine filtration. Tropenmed. Parasitol. 33, 227–228.
Mulvenna, J., Moertel, L., Jones, M. K., Nawaratna, S., Lovas, E. M., Gobert, G. N.,et al. (2010). Exposed proteins of the Schistosoma japonicum tegument. Int. J. Parasitol. 40, 543–554. doi: 10.1016/j.ijpara.2009.10.002
Mutapi, F., Burchmore, R., Mduluza, T., Foucher, A., Harcus, Y., Nicoll, G.,et al. (2005). Praziquantel treatment of individuals exposed to Schistosoma haematobium enhances serological recognition of defined parasite antigens. J. Infect. Dis. 192, 1108–1118. doi: 10.1086/432553
Mutapi, F., Burchmore, R., Mduluza, T., Midzi, N., Turner, C. M. R., and Maizels, R. M. (2008). Age-related and infection intensity-related shifts in antibody recognition of defined protein antigens in a schistosome-exposed population. J. Infect. Dis. 198, 167–175. doi: 10.1086/589511
Nyame, A. K., Lewis, F. A., Doughty, B. L., Correa-Oliveira, R., and Cummings, R. D. (2003). Immunity to schistosomiasis: glycans are potential antigenic targets for immune intervention. Exp. Parasitol. 104, 1–13. doi: 10.1016/S0014–4894(03)00110-3
Obeng, B. B., Aryeetey, Y. A., De Dood, C. J., Amoah, A. S., Larbi, I. A., Deelder, A. M.,et al. (2008). Application of a circulating-cathodic-antigen (CCA) strip test and real-time PCR, in comparison with microscopy, for the detection of Schistosoma haematobium in urine samples from Ghana. Ann. Trop. Med. Parasitol. 102, 625–633. doi: 10.1179/136485908X337490
Ojopi, E. P., Oliveira, P. S., Nunes, D. N., Paquola, A., Demarco, R., Gregório, S. P.,et al. (2007). A quantitative view of the transcriptome of Schistosoma mansoni adult-worms using SAGE. BMC Genomics 8:186. doi: 10.1186/1471–2164-8-186
Oliveira, G., Franco, G., and Verjovski-Almeida, S. (2008). The Brazilian contribution to the study of the Schistosoma mansoni transcriptome. Acta Trop. 108, 179–182. doi: 10.1016/j.actatropica.2008.04.022
Patz, J. A., Graczyk, T. K., Geller, N., and Vittor, A. Y. (2000). Effects of environmental change on emerging parasitic diseases. Int. J. Parasitol. 30, 1395–1405. doi: 10.1016/S0020–7519(00)00141-7
Peng, H., Song, K., Huang, C., Ye, S., Song, H., Hu, W.,et al. (2008). Expression, immunolocalization and serodiagnostic value of a myophilin-like protein from Schistosoma japonicum. Exp. Parasitol. 119, 117–124. doi: 10.1016/j.exppara.2008.01.017
Peng, H. J., Chen, X. G., Wang, X. Z., and Lun, Z. R. (2003). Analysis of the gene expression profile of Schistosoma japonicum cercariae by a strategy based on expressed sequence tags. Parasitol. Res. 90, 287–293. doi: 10.1007/s00436-003-0843-4
Pontes, L. A., Dias-Neto, E., and Rabello, A. (2002). Detection by polymerase chain reaction of Schistosoma mansoni DNA in human serum and feces. Am. J. Trop. Med. Hyg. 66, 157–162.
Pontes, L. A., Oliveira, M. C., Katz, N., Dias-Neto, E., and Rabello, A. (2003). Comparison of a polymerase chain reaction and the Kato–Katz technique for diagnosing infection with Schistosoma mansoni. Am. J. Trop. Med. Hyg. 68, 652–656.
Qing-Wu, J., Li-Ying, W., Jia-Gang, G., Ming-Gang, C., Xiao-Nong, Z., and Engels, D. (2002). Morbidity control of schistosomiasis in China. Acta Trop. 82, 115–125. doi: 10.1016/S0001-706X(02)00006-2
Robijn, M., Koeleman, C., Hokke, C. H., and Deelder, A. M. (2007a). Schistosoma mansoni eggs excrete specific free oligosaccharides that are detectable in the urine of the human host. Mol. Biochem. Parasitol. 151, 162–172. doi: 10.1016/j.molbiopara.2006.10.018
Robijn, M., Koeleman, C., Wuhrer, M., Royle, L., Geyer, R., Dwek, R. A.,et al. (2007b). Targeted identification of a unique glycan epitope of Schistosoma mansoni egg antigens using a diagnostic antibody. Mol. Biochem. Parasitol. 151, 148–161. doi: 10.1016/j.molbiopara.2006.10.019
Robijn, M., Wuhrer, M., Kornelis, D., Deelder, A., Geyer, R., and Hokke, C. (2005). Mapping fucosylated epitopes on glycoproteins and glycolipids of Schistosoma mansoni cercariae, adult worms and eggs. Parasitology 130, 67–77. doi: 10.1017/S0031182004006390
Sandler, N. G., Mentink-Kane, M. M., Cheever, A. W., and Wynn, T. A. (2003). Global gene expression profiles during acute pathogen-induced pulmonary inflammation reveal divergent roles for Th1 and Th2 responses in tissue repair. J. Immunol. 171, 3655–3667. doi: 10.4049/jimmunol.171.7.3655
Sandoval, N., Siles-Lucas, M., Perez-Arellano, J., Carranza, C., Puente, S., Lopez-Aban, J.,et al. (2006). A new PCR-based approach for the specific amplification of DNA from different Schistosoma species applicable to human urine samples. Parasitology 133, 581–587. doi: 10.1017/S0031182006000898
Shabaan, A. M., Mohamed, M. M., Abdallah, M. S., Ibrahim, H. M., and Karim, A. M. (2003). Analysis of Schistosoma mansoni genes using the expressed sequence tag approach. Acta Biochim. Pol. 50, 259–268.
Simões, M. C., Lee, J., Djikeng, A., Cerqueira, G. C., Zerlotini, A., Da Silva-Pereira, R. A.,et al. (2011). Identification of Schistosoma mansoni microRNAs. BMC Genomics 12:47. doi: 10.1186/1471–2164-12-47
Taft, A., Vermeire, J., Bernier, J., Birkeland, S., Cipriano, M., Papa, A.,et al. (2009). Transcriptome analysis of Schistosoma mansoni larval development using serial analysis of gene expression (SAGE). Parasitology 136, 469–485. doi: 10.1017/S0031182009005733
The Schistosoma japonicum Genome Sequencing and Functional Analysis Consortium. (2009). The Schistosoma japonicum genome reveals features of host–parasite interplay. Nature 460, 345–351. doi: 10.1038/nature08140
Utzinger, J., N’goran, E. K., Caffrey, C. R., and Keiser, J. (2011). From innovation to application: social–ecological context, diagnostics, drugs and integrated control of schistosomiasis. Acta Trop. 120, S121–S137. doi: 10.1016/j.actatropica.2010.08.020
Van Balkom, B. W., Van Gestel, R. A., Brouwers, J. F., Krijgsveld, J., Tielens, A. G., Heck, A. J.,et al. (2005). Mass spectrometric analysis of the Schistosoma mansoni tegumental sub-proteome. J. Proteome Res. 4, 958–966. doi: 10.1021/pr050036w
Van Dam, G., Wichers, J., Ferreira, T. F., Ghati, D., Van Amerongen, A., and Deelder, A. (2004). Diagnosis of schistosomiasis by reagent strip test for detection of circulating cathodic antigen. J. Clin. Microbiol. 42, 5458–5461. doi: 10.1128/JCM.42.12.5458–5461.2004
Van Etten, L., Folman, C. C., Eggelte, T. A., Kremsner, P. G., and Deelder, A. M. (1994). Rapid diagnosis of schistosomiasis by antigen detection in urine with a reagent strip. J. Clin. Microbiol. 32, 2404–2406.
van Hellemond, J. J., Van Balkom, B. W., and Tielens, A. G. (2007). Schistosome biology and proteomics: progress and challenges. Exp. Parasitol. 117, 267–274. doi: 10.1016/j.exppara.2007.05.004
Van Lieshout, L., Gangaram Panday, U., De Jonge, N., Krijger, F., Oostburg, B., Polderman, A.,et al. (1995). Immunodiagnosis of schistosomiasis mansoni in a low endemic area in Surinam by determination of the circulating antigens CAA and CCA. Acta Trop. 59, 19–29. doi: 10.1016/0001-706X(94)00084-E
van Lieshout, L., Polderman, A. M., and Deelder, A. M. (2000). Immunodiagnosis of schistosomiasis by determination of the circulating antigens CAA and CCA, in particular in individuals with recent or light infections. Acta Trop. 77, 69–80. doi: 10.1016/S0001-706X(00)00115-7
van Remoortere, A., Bank, C. M., Nyame, A. K., Cummings, R. D., Deelder, A. M., and Van Die, I. (2003). Schistosoma mansoni-infected mice produce antibodies that cross-react with plant, insect, and mammalian glycoproteins and recognize the truncated biantennary N-glycan Man3GlcNAc2-R. Glycobiology 13, 217–225. doi: 10.1093/glycob/cwg025
van Remoortere, A., Hokke, C. H., Van Dam, G. J., Van Die, I., Deelder, A. M., and Van Den Eijnden, D. H. (2000). Various stages of Schistosoma express Lewisx, LacdiNAc, GalNAcβ1–4 (Fucα1–3) GlcNAc and GalNAcβ1–4 (Fucα1–2Fucα1–3) GlcNAc carbohydrate epitopes: detection with monoclonal antibodies that are characterized by enzymatically synthesized neoglycoproteins. Glycobiology 10, 601–609. doi: 10.1093/glycob/10.6.601
Vaz, C., Ahmad, H. M., Sharma, P., Gupta, R., Kumar, L., Kulshreshtha, R.,et al. (2010). Analysis of microRNA transcriptome by deep sequencing of small RNA libraries of peripheral blood. BMC Genomics 11:288. doi: 10.1186/1471-2164-11-288
Verjovski-Almeida, S., Demarco, R., Martins, E. A., Guimaraes, P. E., Ojopi, E. P., Paquola, A. C.,et al. (2003). Transcriptome analysis of the acoelomate human parasite Schistosoma mansoni. Nat. Genet. 35, 148–157. doi: 10.1038/ng1237
Vermeire, J. J., Taft, A. S., Hoffmann, K. F., Fitzpatrick, J. M., and Yoshino, T. P. (2006). Schistosoma mansoni: DNA microarray gene expression profiling during the miracidium-to-mother sporocyst transformation. Mol. Biochem. Parasitol. 147, 39–47. doi: 10.1016/j.molbiopara.2006.01.006
Wang, Y., Holmes, E., Nicholson, J. K., Cloarec, O., Chollet, J., Tanner, M.,et al. (2004). Metabonomic investigations in mice infected with Schistosoma mansoni: an approach for biomarker identification. Proc. Natl. Acad. Sci. U.S.A. 101, 12676–12681. doi: 10.1073/pnas.0404878101
Wang, Y., Li, J. V., Saric, J., Keiser, J., Wu, J., Utzinger, J.,et al. (2010a). Advances in metabolic profiling of experimental nematode and trematode infections. Adv. Parasitol. 73, 373–404. doi: 10.1016/S0065-308X(10)73012-8
Wang, Z., Xue, X., Sun, J., Luo, R., Xu, X., Jiang, Y.,et al. (2010b). An “in-depth” description of the small non-coding RNA population of Schistosoma japonicum schistosomulum. PLoS Negl. Trop. Dis. 4:e596. doi: 10.1371/journal.pntd.0000596
Wang, Y., Utzinger, J., Xiao, S.-H., Xue, J., Nicholson, J. K., Tanner, M.,et al. (2006). System level metabolic effects of a Schistosoma japonicum infection in the Syrian hamster. Mol. Biochem. Parasitol. 146, 1–9. doi: 10.1016/j.molbiopara.2005.10.010
Wang, Z. J., Hu, W., Shen, D. K., Wu, X. W., Wang, J. J., Han, Z. G.,et al. (2003). Prokaryotic expression of gene encoding Schistosoma japonicum SjE16 and its potential application in immunodiagnosis. Zhongguo Ji Sheng Chong Xue Yu Ji Sheng Chong Bing Za Zhi 21, 76–79.
Wen, Z., Wang, S., Wu, Z., and Shen, G. (2011). A novel liquid-phase piezoelectric immunosensor for detecting Schistosoma japonicum circulating antigen. Parasitol. Int. 60, 301–306. doi: 10.1016/j.parint.2011.05.001
Wichmann, D., Panning, M., Quack, T., Kramme, S., Burchard, G.-D., Grevelding, C.,et al. (2009). Diagnosing schistosomiasis by detection of cell-free parasite DNA in human plasma. PLoS Negl. Trop. Dis. 3:e422. doi: 10.1371/journal.pntd.0000422
Williams, D. L., Sayed, A. A., Bernier, J., Birkeland, S. R., Cipriano, M. J., Papa, A. R.,et al. (2007). Profiling Schistosoma mansoni development using serial analysis of gene expression (SAGE). Exp. Parasitol. 117, 246–258. doi: 10.1016/j.exppara.2007.05.001
Wilson, R. A. (2012). Proteomics at the schistosome–mammalian host interface: any prospects for diagnostics or vaccines? Parasitology 139, 1178–1194. doi: 10.1017/S0031182012000339
Wu, H. W., Chen, S. Z., Zhang, Z. S., Wang, R. Z., and Wu, G. L. (1998). Construction of Schistosoma japonicum (Chinese Mailand Strain) egg cDNA library. Chin. J. Parasit. Dis. Control 11, 302–304.
Wu, J.-F., Holmes, E., Xue, J., Xiao, S.-H., Singer, B. H., Tang, H.-R.,et al. (2010). Metabolic alterations in the hamster co-infected with Schistosoma japonicum and Necator americanus. Int. J. Parasitol. 40, 695–703. doi: 10.1016/j.ijpara.2009.11.003
Wuhrer, M., Balog, C. I., Catalina, M., Jones, F. M., Schramm, G., Haas, H.,et al. (2006a). IPSE/alpha-1, a major secretory glycoprotein antigen from schistosome eggs, expresses the Lewis X motif on core-difucosylated N-glycans. FEBS J. 273, 2276–2292. doi: 10.1111/j.1742-4658.2006.05242.x
Wuhrer, M., Koeleman, C. A., Fitzpatrick, J. M., Hoffmann, K. F., Deelder, A. M., and Hokke, C. H. (2006b). Gender-specific expression of complex-type N-glycans in schistosomes. Glycobiology 16, 991–1006. doi: 10.1093/glycob/cwl020
Wuhrer, M., Dennis, R. D., Doenhoff, M. J., Lochnit, G., and Geyer, R. (2000). Schistosoma mansoni cercarial glycolipids are dominated by Lewis X and pseudo-Lewis Y structures. Glycobiology 10, 89–101. doi: 10.1093/glycob/10.1.89
Wuhrer, M., Kantelhardt, S. R., Dennis, R. D., Doenhoff, M. J., Lochnit, G., and Geyer, R. (2002). Characterization of glycosphingolipids from Schistosoma mansoni eggs carrying Fuc (α1–3) GalNAc-, GalNAc (β1–4)[Fuc (α1–3)] GlcNAc- and Gal (β1–4)[Fuc (α1–3)] GlcNAc- (Lewis X) terminal structures. Eur. J. Biochem. 269, 481–493. doi: 10.1046/j.0014-2956.2001.02673.x
Wynn, T. A., Thompson, R. W., Cheever, A. W., and Mentink-Kane, M. M. (2004). Immunopathogenesis of schistosomiasis. Immunol. Rev. 201, 156–167. doi: 10.1111/j.0105-2896.2004.00176.x
Xia, C.-M., Rong, R., Lu, Z.-X., Shi, C.-J., Xu, J., Zhang, H.-Q.,et al. (2009). Schistosoma japonicum: a PCR assay for the early detection and evaluation of treatment in a rabbit model. Exp. Parasitol. 121, 175–179. doi: 10.1016/j.exppara.2008.10.017
Xu, B., Duan, X. W., Lu, Y., Chen, S. B., Feng, Z., and Hu, W. (2011). Cloning, expression and stage-specific analysis of Schistosoma japonicum P7 antigen and evaluation of its value in early diagnosis. Zhongguo Ji Sheng Chong Xue Yu Ji Sheng Chong Bing Za Zhi 29, 161–166.
Xu, J., Liu, A.-P., Guo, J.-J., Wang, B., Qiu, S.-J., Sun, H.,et al. (2013). The sources and metabolic dynamics of Schistosoma japonicum DNA in serum of the host. Parasitol. Res. 112, 129–133. doi: 10.1007/s00436–012-3115-3
Xu, J., Rong, R., Zhang, H., Shi, C., Zhu, X., and Xia, C. (2010). Sensitive and rapid detection of Schistosoma japonicum DNA by loop-mediated isothermal amplification (LAMP). Int. J. Parasitol. 40, 327–331. doi: 10.1016/j.ijpara.2009.08.010
Xue, X., Sun, J., Zhang, Q., Wang, Z., Huang, Y., and Pan, W. (2008). Identification and characterization of novel microRNAs from Schistosoma japonicum. PLoS ONE 3:e4034. doi: 10.1371/journal.pone.0004034
You, H., Zhang, W., Moertel, L., Mcmanus, D. P., and Gobert, G. N. (2009). Transcriptional profiles of adult male and female Schistosoma japonicum in response to insulin reveal increased expression of genes involved in growth and development. Int. J. Parasitol. 39, 1551–1559. doi: 10.1016/j.ijpara.2009.06.006
Young, N. D., Jex, A. R., Li, B., Liu, S., Yang, L., Xiong, Z.,et al. (2012). Whole-genome sequence of Schistosoma haematobium. Nat. Genet. 44, 221–225. doi: 10.1038/ng.1065
Zhao, G. H., Li, J., Blair, D., Li, X. Y., Elsheikha, H. M., Lin, R. Q.,et al. (2012). Biotechnological advances in the diagnosis, species differentiation and phylogenetic analysis of Schistosoma spp. Biotechnol. Adv. 30, 1381–1389. doi: 10.1016/j.biotechadv.2012.02.008
Zheng, M. J., Li, M., Li, C. L., Chu, D. Y., Wang, Z. C., Luo, F.,et al. (2007). Secreted expression of signaling protein 14-3-3 of Schistosoma japonicum in Pichia pastoris system with primary evaluation on its antigenicity. Zhongguo Ji Sheng Chong Xue Yu Ji Sheng Chong Bing Za Zhi 25, 12–16.
Zhong, Z.-R., Zhou, H.-B., Li, X.-Y., Luo, Q.-L., Song, X.-R., Wang, W.,et al. (2010). Serological proteome-oriented screening and application of antigens for the diagnosis of Schistosomiasis japonica. Acta Trop. 116, 1–8. doi: 10.1016/j.actatropica.2010.04.014
Zhou, L., Tang, J., Zhao, Y., Gong, R., Lu, X., Gong, L.,et al. (2011a). A highly sensitive TaqMan real-time PCR assay for early detection of Schistosoma species. Acta Trop. 120, 88–94. doi: 10.1016/j.actatropica.2011.06.006
Zhou, Y. B., Zheng, H. M., and Jiang, Q. W. (2011b). A diagnostic challenge for Schistosomiasis japonica in China: consequences on praziquantel-based morbidity control. Parasit Vectors 4, 194. doi: 10.1186/1756-3305-4-194
Zhou, X.-H., Wu, J.-Y., Huang, X.-Q., Kunnon, S. P., Zhu, X.-Q., and Chen, X.-G. (2010). Identification and characterization of Schistosoma japonicum Sjp40, a potential antigen candidate for the early diagnosis of schistosomiasis. Diagn. Microbiol. Infect. Dis. 67, 337–345. doi: 10.1016/j.diagmicrobio.2010.03.003
Zhou, Y.-P., Wu, Z.-D., Yang, L.-L., Sun, X., You, X., Yu, X.-B.,et al. (2009). Cloning, molecular characterization of a 13-kDa antigen from Schistosoma japonicum, Sj13, a putative salivary diagnosis candidate for Schistosomiasis japonica. Parasitol. Res. 105, 1435–1444. doi: 10.1007/s00436-009-1575-x
Keywords: Schistosoma, -omics, diagnosis, biomarkers, parasite
Citation: Wang S and Hu W (2014) Development of “-omics” research in Schistosoma spp. and -omics-based new diagnostic tools for schistosomiasis. Front. Microbiol. 5:313. doi: 10.3389/fmicb.2014.00313
Received: 12 February 2014; Accepted: 09 June 2014;
Published online: 26 June 2014.
Edited by:
Paul J. Brindley, The George Washington University, USAReviewed by:
Estela Castillo, Universidad de la República, UruguayRudi Grams, Thammasat University, Thailand
Copyright © 2014 Wang and Hu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Wei Hu, Department of Microbiology and Microbial Engineering, School of Life Sciences, Fudan University, Shanghai 200433, China e-mail:aHV3QGZ1ZGFuLmVkdS5jbg==
 Shuqi Wang
Shuqi Wang Wei Hu
Wei Hu