- 1Department of Environmental Sciences, University of California, Riverside, CA, USA
- 2Department of Biological Sciences, Faculty of Science, University of Alberta, Edmonton, AB, Canada
Sphingomonas wittichii RW1 has no reported denitrifying activity yet encodes nitrite and nitric oxide reductases. The aims of this study were to determine conditions under which S. wittichii RW1 consumes nitrite (NO−2) and produces nitrous oxide (N2O), examine expression of putative genes for N-oxide metabolism, and determine the functionality of chromosomal (ch) and plasmid (p) encoded quinol-dependent nitric oxide reductases (NorZ). Batch cultures of wildtype (WT) and a norZch mutant of S. wittichii RW1 consumed NO−2 and produced N2O during stationary phase. The norZch mutant produced N2O, although at significantly lower levels (c.a. 66–87%) relative to the WT. Rates of N2O production were 2–3 times higher in cultures initiated at low relative to atmospheric O2 per unit biomass, although rates of NO−2 consumption were elevated in cultures initiated with atmospheric O2 and 1 mM NaNO2. Levels of mRNA encoding nitrite reductase (nirK), plasmid-encoded nitric oxide dioxygenase (hmpp) and plasmid-encoded nitric oxide reductase (norZp) were significantly higher in the norZch mutant over a growth curve relative to WT. The presence of NO−2 further increased levels of nirK and hmpp mRNA in both the WT and norZch mutant; levels of norZp mRNA compensated for the loss of norZch expression in the norZch mutant. Together, the results suggest that S. wittichii RW1 denitrifies NO−2 to N2O and expresses gene products predicted to detoxify N-oxides. So far, only S. wittichii strains within four closely related taxa have been observed to encode both nirK and norZ genes, indicating a species-specific lateral gene transfer that may be relevant to the niche preference of S. wittichii.
Introduction
Denitrification is the sequential reduction of nitrate (NO−3) and nitrite (NO−2) to dinitrogen (N2) via the gaseous intermediates, nitric oxide (NO) and nitrous oxide (N2O) (Zumft, 1997). Respiratory denitrification is considered an anaerobic energy-generating metabolism; however, many bacteria can denitrify in the presence of O2 starting with NO−3 or NO−2 and terminating with N2O due to inhibition or absence of nitrous oxide reductase (Philippot et al., 2011; Stein, 2011; Chen and Strous, 2013). While some denitrifiers grow via hybrid respiration with O2 and NO−3 and/or NO−2, others respire N-oxides during late-log to stationary phase for detoxification and/or energy conservation, particularly under reduced O2 (Takaya et al., 2003; Stein, 2011; Chen and Strous, 2013). The collective role of denitrifying bacteria in transformation and release of highly reactive N-oxides is of critical importance because of the effects these molecules have on environmental and human health, atmospheric chemistry, and global warming (Fields, 2004; Galloway et al., 2008).
Sphingomonas wittichii RW1 was isolated from the River Elbe (Germany) as a model organism for studying the bioremediation of dioxin-containing compounds (Wittich et al., 1992; Wilkes et al., 1996; Yabuuchi et al., 2001; Keum et al., 2008). The complete genome sequence of S. wittichii RW1 revealed the presence of a single circular chromosome and two megaplasmids (Miller et al., 2010). Although not known to denitrify, S. wittichii RW1 encodes in its genome a copper-containing nitrite reductase (nirK) as the terminal member of a four-gene cluster with a NO-responsive NsrR regulator encoded upstream (Swit_1789-93). This gene cluster shares structural and sequence homology to ammonia- and nitrite-oxidizing bacteria in the Nitrosomonas and Nitrobacter genera, respectively (Cantera and Stein, 2007). S. wittichii RW1 also encodes a chromosomal (Swit_4614) and plasmid copy (Swit_5200) of quinol-linked nitric oxide reductase (norZ). NorZ is often expressed in non-denitrifying pathogenic bacteria for NO detoxification (Hendriks et al., 2000), but can also act alongside the terminal oxidase in the aerobic respiratory chain for energy conservation (Chen and Strous, 2013). The plasmid-encoded norZp (Swit_5200) is the terminal member of a four-gene cluster; the first member of which encodes a nitric oxide dioxygenase (hmpp; Swit_5203) with predicted function in NO oxidation to NO−3 or NO reduction to N2O depending on O2 concentration (Bonamore and Boffi, 2008). Nitric oxide dioxygenases are present in both denitrifying and non-denitrifying microorganisms to combat nitrosative and oxidative stresses (Bonamore and Boffi, 2008; Forrester and Foster, 2012). Although nitric oxide dioxygenases are usually conserved members of the NO-controlled NsrR transcriptional regulon in bacteria (Rodionov et al., 2005), the plasmid-encoded gene cluster in S. wittichii RW1 that includes both NorZ and nitric oxide dioxygenase is preceded by a CDS for the NO-responsive NnrR transcriptional regulator (Swit_5204). Aside from Swit_5203, S. wittichii RW1 encodes three other putative hmp genes, the plasmid-encoded Swit_5299 and the chromosomal Swit_1434 and 3173. A comparison of 51 genome-sequenced sphingomonad strains (encompassing the Sphingomonas, Sphingobium, Novosphingobium, and Sphingopyxis genera) by BLAST searches through the Integrated Microbial Genomes database (http://img.jgi.doe.gov) revealed that only the two strains of S. wittichii (RW1 and DP58) encode the complete nirK gene cluster, whereas eight sphingomonad genomes encode either one or two copies of norZ and 17 genomes encode one or more hmp genes whose translated sequences share >60% protein identity to Swit_5203. Hence, the potential for spingomonad bacteria to transform nitrogen oxides appears to be fairly restricted.
Previous studies in Neisseria and Synechococcus demonstrated that disruption of norZ expression resulted in increased NO sensitivity, diminished NO consumption and N2O production, and decreased growth under anoxia (Householder et al., 2000; Busch et al., 2002). Interestingly, Ralstonia eutropha H16 also possesses two independent quinol-linked nitric oxide reductases. Deletion of either gene in R. eutropha H16 resulted in no phenotypic change under aerobic or anaerobic growth at the expense of NO−3 or NO−2 (Cramm et al., 1997). Therefore, in the present study we tested the hypothesis that the norZ genes in S. wittichii RW1 are similarly isofunctional.
The overarching hypothesis of the present study is that S. wittichii RW1 reduces NO−2 to N2O and thus can be classified as a denitrifying strain. The ability of S. wittichii RW1 to denitrify from NO−3 was not investigated as the genome of S. wittichii RW1 encodes only the alpha subunit of assimilatory nitrate reductase (Swit_1709) and no features of dissimilatory nitrate reductases. Furthermore, this strain tested negative for reduction of NO−3 to NO−2 (Yabuuchi et al., 2001). There is no identifiable sequence in the genome with similarity to nitrous oxide reductase; hence, this strain is predicted to denitrify only NO−2 to N2O. To provide support for S. wittichii RW1 as a denitrifier, objectives were to: (a) determine whether and when S. wittichii RW1 produces N2O at the expense of NO−2 (b) investigate the regulation of putative N-oxide metabolism genes in response to varying NO−2, and (c) determine whether the chromosomal- and plasmid-encoded norZ genes in S. wittichii RW1 are isofunctional.
Materials and Methods
Culture Maintenance
Sphingomonas wittichii RW1 was provided as a gift from Dr. Rolf Halden. Cultures were grown in 5 mL Luria-Bertani Broth (LB) in sterilized 15 mL capped-polystyrene tubes in a rotary shaker (180 r.p.m.) at 28°C. Cultures were periodically streaked and grown on LB agar plates for single colony isolation to maintain culture purity.
Construction of norZch Mutant of S. wittichii RW1
The region from bp 203 to 776 of the norZch gene was PCR-amplified from S. wittichii RW1 genomic DNA with primers 203F 5′ aactggaacaggccgatg 3′ and 776R 5′ cgatcgccttcatcttcg 3′ to make use of an internal BclI restriction site [Primer3 Input 0.4.0 software (Rozen and Skaletsky, 2000)]. The amplification product was purified and ligated to the pGEM®-T Vector according to manufacturers' instructions (Promega Corp., Madison, WI). The ligation mixture was transformed into dam−/dcm− competent E. coli cells (New England BioLabs Inc., Ipswich, MA) and transformants were selected via blue-white screening on LB agar plates containing 0.5 mM IPTG, 80 μg/mL X-Gal, and 100 μg/mL ampicillin. Plasmids from positive transformants were purified using Wizard® Plus SV Minipreps DNA Purification System kit (Promega Corp., Madison, WI) and digested with the BclI restriction enzyme (New England BioLabs Inc., Ipswich MA). The digest was run on a 0.8% agarose gel and linearized vector was gel-purified using the Wizard® SV Gel and PCR Clean-Up System kit (Promega Corp., Madison, WI).
A gentamycin-resistance cassette (871 bp) was digested from the pUCGM vector (gift from N. Hommes) using the BamHI restriction enzyme (New England BioLabs Inc., Ipswich MA). The digest was gel-purified from a 0.8% agarose gel and ligated to the previously BclI-digested pGEM-T-norZ vector. The ligation mixture was transformed into competent E. coli JM109 cells. Transformed cells were plated onto LB agar containing 100 μg/mL ampicillin and 10 μg/mL gentamycin. Positive transformants were verified by PCR and Sanger sequencing using the BigDye Terminator Cycle Sequencing Kit (Applied Biosystems, Foster City USA). Plasmids containing the correct inserts were purified as described above and electroporated into S. wittichii RW1 cells using an E. coli Pulser™ Transformation Apparatus (BioRad Laboratories, Hercules, CA). Competent S. wittichii RW1 cells were prepared by harvesting in exponential phase, washing three times with 20 mL ice-cold and nuclease-free water, washing twice with 2 mL ice-cold 10% glycerol, and resuspended in 10% glycerol to a final volume of 100 μL. Electroporated cells were plated onto LB agar containing 10 μg/mL gentamycin. The norZch mutant strain was checked by PCR using additional primers: 45F 5′ agagacccaggaccacgac 3′, 854R 5′ tcaccgtcatggaatattgg 3′, pUCGM173F 5′ tgcctcgggcatccaagcagca 3′, pUCGM514R 5′ gagagcgccaacaaccgcttct 3′ and pUCGM519F 5′ cttacgttctgcccaggttt 3′. PCR products were purified and validated by Sanger sequencing. The norZch mutant strain was maintained on LB media with 50 μg/mL gentamycin.
Growth Experiments
S. wittichii RW1 wildtype (WT) and norZch mutant cells from exponentially growing cultures were inoculated into LB media (500 μL into 100 mL) containing 0, 0.3, or 1 mM NaNO2 into glass serum bottles (160 mL), which were then crimp-sealed with rubber septa and aluminum seals. Incubations of norZch mutant cells contained 50 μg/mL gentamycin. Triplicate incubations of each control condition included the same concentrations of NaNO2 plus: (1) heat-inactivated cells, (2) no cells, or (3) live cells in bottles purged of O2 by sparging the medium with N2. All control incubations were treated identically to the experimental incubations to determine whether chemical decomposition of NO−2 contributed to NO−2 loss or N2O accumulation. Gas headspace (60 mL) was either left unchanged (atmospheric O2) or, for WT cells, sparged with N2 and injected with pure O2 prior to inoculation (ca. 3% O2 in gas headspace as validated by gas chromatography; GC-TCD, Shimadzu, Kyoto, Japan; Molecular Sieve 6A column, Alltech, Deerfield IL). Experimental and control bottles were incubated in a rotary shaker (180 r.p.m.) at 28°C. Starting at t = 0 h, 2 mL samples were extracted every 4 h using a sterile 1 mL needle and syringe. Growth was determined by measuring OD 600 nm using a Spectronic 20 Genesys spectrophotometer (Thermo Fisher Scientific, Inc., Waltham, MA). Cells were immediately treated with 500 μL RNAprotect™ Bacteria Reagent (Qiagen, Valencia, CA), and kept at −80°C. Experiments consisted of five independent trials performed on different days for both strains and under every condition.
Nucleic Acid Extraction
Genomic DNA was isolated using the Wizard® SV Genomic DNA Purification System kit (Promega Corp., Madison, WI). Total RNA was extracted using the Aurum™ Total RNA Mini kit (Bio-Rad Laboratories, Hercules, CA). Nucleic acid concentration was determined using a NanoDrop® ND-1000 Spectrophotometer (Nanodrop Technologies, Inc., Wilmington, DE). DNA and RNA samples were kept at −20 and −80°C, respectively.
Dot-Blot Hybridization
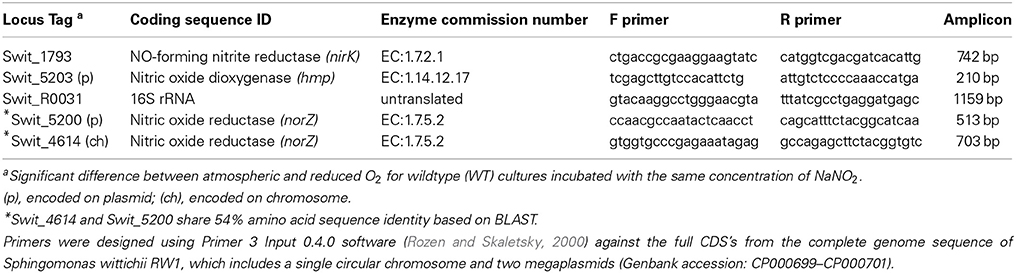
Gene-specific primers were designed from CDS's of selected genes from the S. wittichii RW1 genome sequence (Genbank accession: CP000699 to CP000701) using Primer3 Input 0.4.0 software (Rozen and Skaletsky, 2000) (Table 1). PCR reactions included standard reagents for Taq polymerase and genomic DNA as template in 25 μL reactions (Sambrook and Russell, 2001). Thermal cycler (iCycler, BioRad, Hercules, CA) amplification conditions were: 95°C for 5 min, 30 cycles of 95°C for 40 s, 55°C for 40 s and 72°C for 50 s, with an additional extension cycle of 72°C for 7 min. PCR products were checked by agarose gel (1%) to verify single products of appropriate size. Amplification products were purified using the Wizard® SV Gel and PCR Clean-Up System kit (Promega Corp., Madison, WI). Amplification products were labeled using the Prime-a-Gene labeling system (Promega Corp., Madison, WI) with [α-32P]-dCTP (3000 Ci mmol−1; Perkin-Elmer Inc., Waltham, MA) and random hexamers. The dynamic range of detection for each probe was tested using a concentration series of specific mRNA from 0.1 to 3 μg from control incubations (0 mM NaNO2). The r2 values for the slope of hybridization intensity/μg mRNA was from 0.94 to 1.0 for all probes.
Two μg total RNA from each sample was blotted onto a Zeta-Probe® GT nylon membrane (Bio-Rad Laboratories, Hercules, CA) using a Minifold® microsample filtration manifold (Dot-Blot System, Schleicher & Schuell, Keene, NH) following the Zeta-Probe® protocol. Membranes were allowed to dry overnight and UV-crosslinked (FB-UVXL-1000, Fisher Scientific, Pittsburgh, PA). Prehybridization, hybridization, and washing of Zeta-Probe® nylon membranes were done according to manufacturer's instructions at 30°C. To allow re-probing, membranes were stripped of radioactivity by washing twice in a 0.1× SSC/0.5% SDS solution at 95–100°C for 20 min. All blots were hybridized to gene-specific probes to normalize hybridization signals to the 16S rRNA pool. Hybridization intensity was analyzed using a Typhoon Phosphorimager and Imagequant software (Amersham, Piscataway, New Jersey).
Data Analysis
Background and signal from non-specific binding was subtracted, after which the relative hybridization intensity of specific probes was normalized by dividing gene-specific signal by signal from 16S rRNA probe hybridizations. The fold difference in levels of mRNA for each gene and time point was determined by dividing hybridization intensities from dot blots of RNA extracted from NO−2 amended by those from unamended cultures. Student's t-test (p < 0.05) was performed to determine significant differences between treatments.
Analytical Measurements
Nitrite and ammonium were measured colorimetrically using standard methods (Clesceri et al., 1998). Nitrate was measured using a Standard Range Lab Nitrate Test kit (NECi, Lake Linden, Michigan). O2 and N2O were measured from the gas headspace of sample bottles by GC-TCD (Shimadzu, Kyoto Japan; Molesieve 5A and Hayesep Q columns, Alltech, Deerfield IL). Concentrations were determined by comparing to standard curves generated for each reagent and gas within the limits of detection.
Results
Effect of O2 and NO−2 on Growth of WT and NorZch Mutant Strains of S. wittichii RW1
S. wittichii RW1 is an aerobic heterotrophic bacterium; hence, the doubling time (calculated from 12 to 20 h growth) and final yields of non-mutagenized cells were significantly faster and higher, respectively, for cultures initiated under atmospheric compared to reduced (ca. 3%) O2 levels (Figure 1 and Table 2). Doubling times of the norZch mutant were significantly shorter than those of the WT during exponential growth; thus, even though the norZch mutant exhibited a longer lag phase, the cell density of the cultures were equivalent in stationary phase (Figure 1 and Table 2). The addition of NaNO2 to cultures initiated at atmospheric O2 only significantly increased the doubling time of WT cultures, but significantly reduced the final yields of both WT and norZch mutant cultures (Table 2).
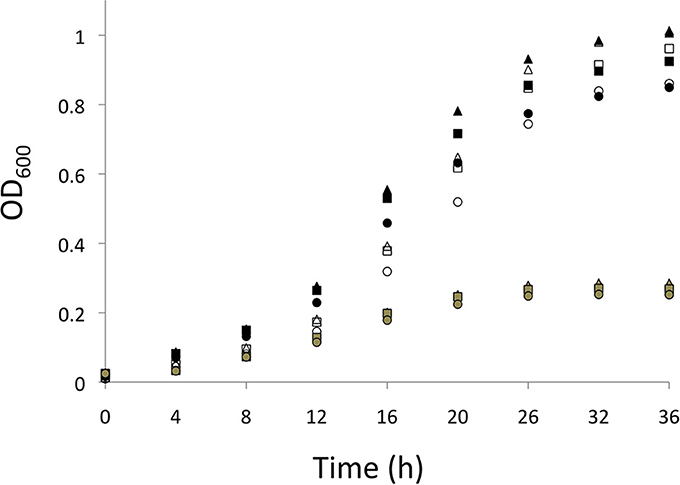
Figure 1. Growth (OD600) measured over time (h) for S. wittichii RW1 WT cultures initiated at atmospheric O2 (black symbols) or reduced O2 (gray symbols), and for norBch mutant S. wittichii RW1 cultures initiated at atmospheric O2 (white symbols). Triangles = 0 mM added NaNO2, squares = 0.3 mM added NaNO2, and circles = 1 mM added NaNO2. Points on the growth curves represent averaged values from 5 independent experiments for each incubation condition.
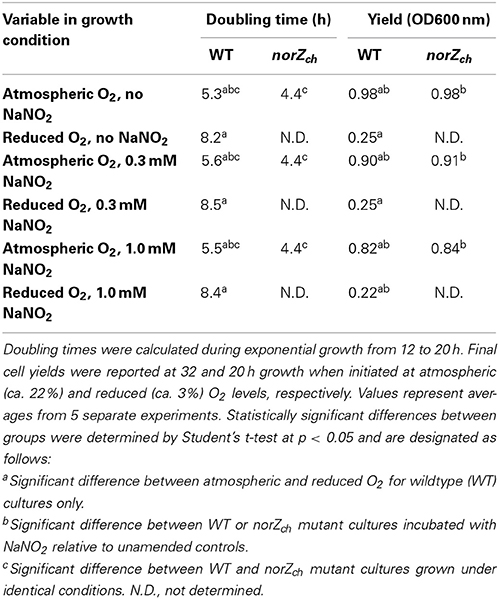
Table 2. Growth of WT and norZch mutant strains of S. wittichii RW1 at variable NaNO2 and O2 concentrations.
Consumption of NO−2 and Production of N2O
Cultures of WT and norZch mutant S. wittichii RW1 were incubated in the presence of NaNO2 to assess whether expression of norZch was required for aerobic denitrifying activity. Amounts of remaining NO−2, remaining O2, and headspace N2O levels were compared over stationary phase (Table 3). NO−2 was consumed nearly to completion in both WT and norZch mutant cultures by 96 h incubation. Neither WT nor norZch mutant cultures consumed O2 to complete anoxia and headspace O2 levels remained largely stable following 72 h incubation, even with continuous shaking at 180 rpm. N2O was measurable in the gas headspace starting after 48 h of incubation and continued to accumulate proportionally with the amount of added NaNO2 (Table 3). The norZch mutant cultures produced significantly less N2O than the WT cultures (66–87% of WT levels) at both NO−2 concentrations. Nitrate production was not observed, which would be an expected aerobic activity of Hmp. NH+4 concentrations also did not vary between treatment groups, which would be expected if S. wittichii RW1 reduced NO−2 directly to NH+4 and allowed its accumulation prior to assimilation (data not shown). N2 was not measured. Control incubations containing heat-inactivated cells, no cells, or live cells inoculated into bottles sparged of O2 with N2 gas showed no consumption of NO−2 and no production of N2O.
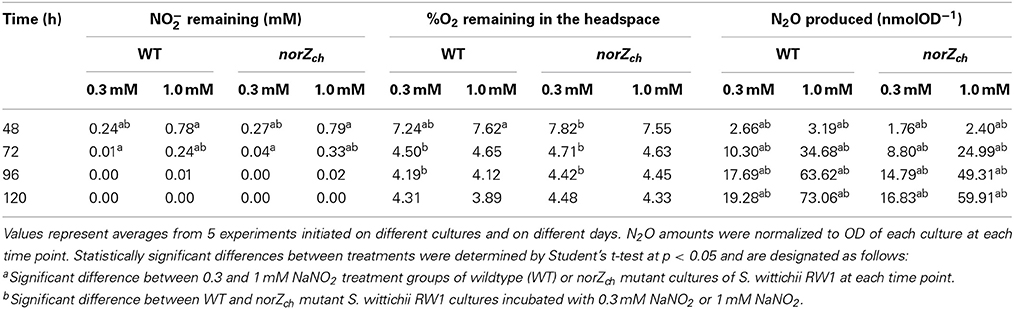
Table 3. Consumption of nitrite and oxygen and production of nitrous oxide by wild-type and norZch mutant cultures of S. wittichii RW1 initiated at atmospheric oxygen headspace and with 0.3 or 1 mM NaNO2.
We next tested whether lower oxygen had an effect on the rates of NO−2 or O2 consumption or N2O production in non-mutated S. wittichii RW1. To address this question, S. wittichii RW1 cultures were inoculated with 0, 0.3, or 1 mM NaNO2 at either atmospheric or reduced (ca. 3%) O2 levels. Cultures initiated at atmospheric O2 and 1 mM NaNO2 consumed O2 and NO−2 significantly faster than cultures initiated at reduced O2 and 1 mM NaNO2, yet the rate of N2O production was 2–3 times faster for cultures initiated at reduced relative to atmospheric O2 levels (Table 4). The N2O-N measured in the gas headspace of the cultures was orders of magnitude lower than the amount of NO−2 consumed per unit biomass (i.e., nmol N2O produced from μmol NO−2 consumed). Even though N2O is highly soluble, the vast difference between NO−2 consumption and N2O production implies conversion of NO−2 into a product other than N2O; however, NO−3 was undetectable and NH+4 levels did not vary in any culture at any time point (data not shown).
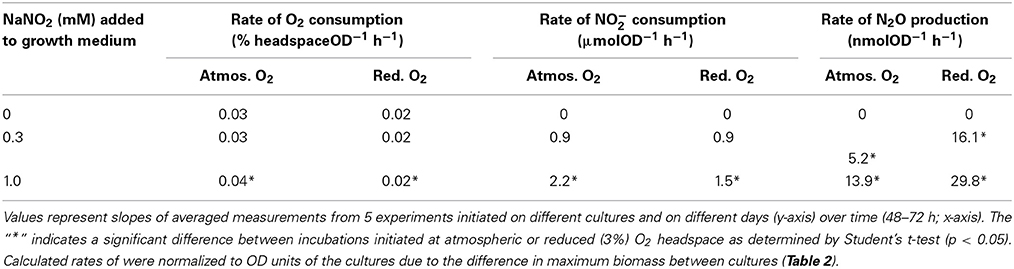
Table 4. Maximum rates of nitrite and oxygen consumption and nitrous oxide production by stationary phase S. wittichii RW1 wildtype cultures grown at atmospheric (ca. 22%) or reduced (ca. 3%) O2 headspace.
Expression Levels of Putative Aerobic Denitrification Genes
Levels of specific mRNAs encoding nirK, hmpp, norZch, and norZp, as normalized to levels of 16S rRNA, were compared between WT and norZch mutant S. wittichii RW1 from mid-log and into stationary phase (24–66 h). This period of time covers the interval over which consumption of NO−2 and production of N2O is measurably active. Expression of norZp substituted for norZch in the norZch mutant strain and the levels of respective norZ transcript remained relatively high in both cell lines over time (Figure 2). Levels of nirK and hmpp transcript were significantly higher in the norZch mutant than in the WT strain at nearly all time points. Whereas nirK and hmpp transcript levels increased between 24 and 66 h in WT cells, both transcript levels remained relatively high in the norZch mutant over the full time course (Figure 2).
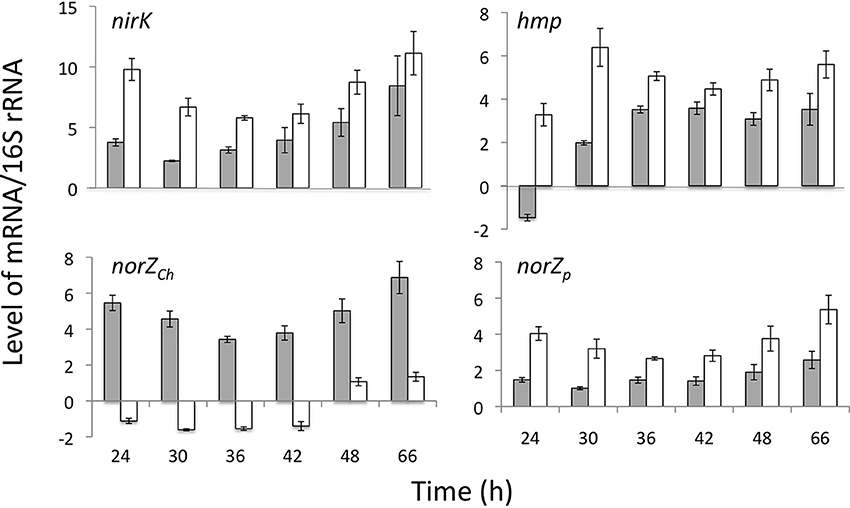
Figure 2. Levels of mRNA of select genes in S. wittichii RW1 WT (gray bars) and norZch mutant (white bars) in batch cultures over time. Error bars represent standard error for 5 replicate experiments performed with separate cultures on different days.
Finally, the effect of NO−2 on transcript levels was examined in norZch mutant and WT cultures, and for non-mutated cultures initiated under atmospheric and reduced O2 levels. Each hybridization signal was normalized to that for 16S rRNA, after which the ratio of hybridization intensity between NaNO2-treated and untreated sample was calculated for every culture and each transcript pool. There was no significant effect of NO−2 treatment on any transcript level for any culture until late log phase (i.e., 24 h for WT and norZch mutant cultures and 20 h for WT cultures initiated at reduced O2). At least a two-fold increase between NaNO2-treated and untreated cells was considered a significant effect; thus, nirK and hmpp were positively responsive to NaNO2 in both WT and norZch mutant cultures (Table 5). For both norZ genes, only transcription levels of norZp in the norZch mutant were responsive to 1 mM NaNO2 treatment.
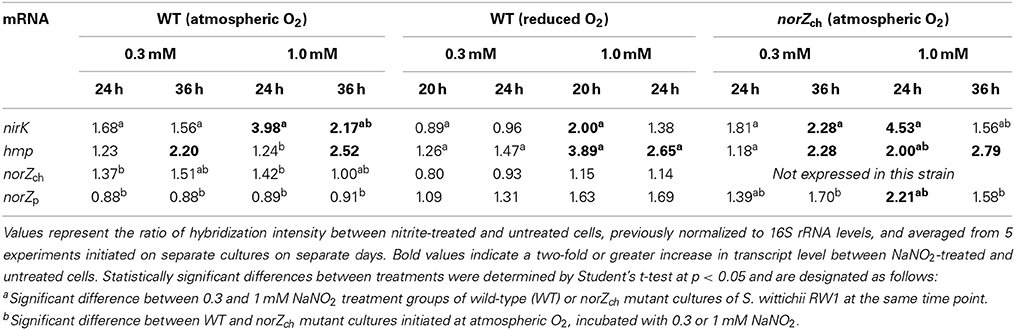
Table 5. Effect of nitrite and O2 on levels of specific transcripts of S. wittichii RW1 from mid-log and early stationary phase cultures.
Discussion
Sphingomonas wittichii RW1 Denitrifies NO−2 to N2O
Rapid consumption of NO−2 by S. wittichii RW1 occurred only once the cells reached stationary phase (Table 3), suggesting that S. wittichii RW1 performs this process for detoxification or maintenance metabolism rather than for generating proton motive force for cellular growth. During growth under reduced O2, an increased rate of NO−2 conversion to N2O (Table 4) relative to cultures initiated at atmospheric O2 implies that O2 limitation must be reached for denitrifying activity to commence as would be commonly expected (Zumft, 1997). It is interesting that a faster rate of NO−2 consumption occurred for cultures initiated at atmospheric than at reduced O2in the presence of 1 mM NaNO2 as this implies an additional process from denitrification for NO−2 loss. Although a substantial quantity of NO−2 consumed by S. wittichii RW1 was converted to N2O, there was a considerable pool of transformed NO−2 that could not be accounted for in NH+4 or NO−3 pools. There is no homolog for nitrous oxide reductase (nosZ) in the genome sequence of S. wittichii RW1; hence, denitrification to N2 is unlikely. Sphingomonads are also not known to produce N-storage polymers, but S. wittichii RW1 does encode an assimilatory nitrite reductase (nirBD; Swit_1707-8). Thus, the fate of the remaining NO−2-N remains unknown.
Genes for Nitrogen Oxide Transformations are Expressed in S. wittichii RW1, and the norz Genes are Isofunctional
Levels of nirK and hmpp and either norZch (WT) or norZp (norZch mutant) transcripts remained relatively high through stationary phase of S. wittichii RW1 (Figure 2), supporting the stationary phase onset of denitrifying activity (Table 3). The absence of norZch expression in S. wittichii RW1 had the effects of increasing the exponential growth rate and preventing slowed growth upon exposure to NO−2 (Figure 1 and Table 1). This phenotype may be in part due to increased expression of genes for handling nitrosative stress, that is nirK, hmpp, and norZp, in the norZch mutant compared to the WT (Figure 2). The increase in transcript pools corresponded to a decrease in the amount of NO−2 converted to N2O (Table 3), further suggesting that the norZch mutant cells were not as susceptible to nitrosative stress as the WT. While other unexamined genetic factors were likely at play in mediating these phenotypes of the norZchmutant, the present data clearly show that the loss of norZch expression was compensated for by expression of norZp; hence, the norZ genes of S. wittichii RW1 are isofunctional. As with the WT cells, expression of both nirK and hmpp genes were positively affected by exposure to NO−2 in the norZch mutant (Table 5). This increased expression was potentially a function of nirK and hmpp genes being regulated by NsrR (Swit_1789) and NnrR (Swit_5204) NO responsive regulators, respectively. In addition, the increased level of norZp transcript in the norZch mutant upon exposure to 1 mM NaNO2 (Table 5), suggests a conditional co-regulation of hmp-orf1-orf2-norZ genes when norZp expression is required.
Conclusions
Results from this study confirm the ability of S. wittichii RW1 to reduce NO−2 to N2O and also to transform excess NO−2 via another mechanism. This metabolic capability may be restricted to the Spingomonas wittichii species of the sphingomonads based on the limited co-occurrence of nirK and norZ genes in their genomes. This denitrification module was likely acquired by S. wittichii strains by lateral gene transfer as a function of ecological niche and need for N-oxide detoxification. As meta-'omic studies often rely on correlating functional genes to 16S rRNA phylotypes, this study sheds light on the complication of relatively rare LGT events that can confer biogeochemically important functions to individual species of broadly distributed bacterial families.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by the Kearny Foundation for Soil Science and the Environmental Toxicology Program at the University of California, Riverside.
References
Bonamore, A., and Boffi, A. (2008). Flavohemoglobin: structure and reactivity. IUBMB Life 60, 19–28. doi: 10.1002/iub.9
Busch, A., Friedrich, B., and Cramm, R. (2002). Characterization of the norB gene, encoding nitric oxide reductase, in the nondenitrifying cyanobacterium Synechocystis sp strain PCC6803. Appl. Environ. Microbiol. 68, 668–672. doi: 10.1128/AEM.68.2.668-672.2002
Cantera, J. J. L., and Stein, L. Y. (2007). Molecular diversity of nitrite reductase (nirK) genes in nitrifying bacteria. Environ. Microbiol. 9, 765–776. doi: 10.1111/j.1462-2920.2006.01198.x
Chen, J. W., and Strous, M. (2013). Denitrification and aerobic respiration, hybrid electron transport chains and co-evolution. Biochim. Biophys. Acta Bioenerg. 1827, 136–144. doi: 10.1016/j.bbabio.2012.10.002
Clesceri, L. S., Greenberg, A. E., and Eaton, A. D. (1998). Standard Methods for the Examination of Water and Wastewater. Washington, DC: American Public Health Assoc.
Cramm, R., Siddiqui, R. A., and Friedrich, B. (1997). Two isofunctional nitric oxide reductases in Alcaligenes eutrophus H16. J. Bacteriol. 179, 6769–6777.
Fields, S. (2004). Global nitrogen - Cycling out of control. Environ. Health Perspect. 112, A556–A563. doi: 10.1289/ehp.112-a556
Forrester, M. T., and Foster, M. W. (2012). Protection from nitrosative stress: a central role for microbial flavohemoglobin. Free Rad. Biol. Med. 52, 1620–1633. doi: 10.1016/j.freeradbiomed.2012.01.028
Galloway, J. N., Townsend, A. R., Erisman, J. W., Bekunda, M., Cai, Z., Freney, J. R., et al. (2008). Transformation of the nitrogen cycle: recent trends, questions, and potential solutions. Science 320, 889–892. doi: 10.1126/science.1136674
Hendriks, J., Oubrie, A., Castresana, J., Urbani, A., Gemeinhardt, S., and Saraste, M. (2000). Nitric oxide reductases in bacteria. Biochim. Biophys. Acta Bioenerg. 1459, 266–273. doi: 10.1016/S0005-2728(00)00161-4
Householder, T. C., Fozo, E. M., Cardinale, J. A., and Clark, V. L. (2000). Gonococcal nitric oxide reductase is encoded by a single gene, norB, which is required for anaerobic growth and is induced by nitric oxide. Infect. Immunol. 68, 5241–5246. doi: 10.1128/IAI.68.9.5241-5246.2000
Keum, Y. S., Lee, Y. J., and Kim, J. H. (2008). Metabolism of nitrodiphenyl ether herbicides by dioxin-degrading bacterium Sphingomonas wittichii RW1. J. Agricult. Food Chem. 56, 9146–9151. doi: 10.1021/jf801362k
Miller, T. R., Delcher, A. L., Salzberg, S. L., Saunders, E., Detter, J. C., and Halden, R. U. (2010). Genome sequence of the dioxin-mineralizing bacterium Sphingomonas wittichii RW1. J. Bacteriol. 192, 6101–6102. doi: 10.1128/JB.01030-10
Philippot, L., Andert, J., Jones, C. M., Bru, D., and Hallin, S. (2011). Importance of denitrifiers lacking the genes encoding the nitrous oxide reductase for N2O emissions from soil. Global Change Biol. 17, 1497–1504. doi: 10.1111/j.1365-2486.2010.02334.x
Rodionov, D. A., Dubchak, I. L., Arkin, A. P., Alm, E. J., and Gelfand, M. S. (2005). Dissimilatory metabolism of nitrogen oxides in bacteria: comparative reconstruction of transcriptional networks. PLoS Comp. Biol. 1:e55. doi: 10.1371/journal.pcbi.0010055
Rozen, S., and Skaletsky, H. J. (2000). “Primer3 on the WWW for general users and for biologist programmers,” in Bioinformatics Methods and Protocols: Methods in Molecular Biology, eds S. Krawetz and S. Misener (Totowa, NJ: Humana Press), 365–386.
Sambrook, J., and Russell, D. W. (2001). Molecular Cloning: a Laboratory Manual. Cold Springs Harbor, NY: Cold Springs Harbor Laboratory Press.
Stein, L. Y. (2011). “Heterotrophic nitrification and nitrifier denitrification,” in Nitrification, eds B. B. Ward, D. J. Arp, and M. G. Klotz (Washington, DC: ASM Press), 95–114.
Takaya, N., Kuwazaki, S., Adachi, Y., Suzuki, S., Kikuchi, T., Nakamura, H., et al. (2003). Hybrid respiration in the denitrifying mitochondria of Fusarium oxysporum. J. Biochem. 133, 461–465. doi: 10.1093/jb/mvg060
Wilkes, H., Wittich, R. M., Timmis, K. N., Fortnagel, P., and Francke, W. (1996). Degradation of chlorinated dibenzofurans and dibenzo-p-dioxins by Sphingomonas sp. strain RW1. Appl. Environ. Microbiol. 62, 367–371.
Wittich, R. M., Wilkes, H., Sinnwell, V., Francke, W., and Fortnagel, P. (1992). Metabolism of dibenzo-para-dioxin by Sphingomonas sp. strain RW1 O. Appl. Environ. Microbiol. 58, 1005–1010.
Yabuuchi, E., Yamamoto, H., Terakubo, S., Okamura, N., Naka, T., Fujiwara, N., et al. (2001). Proposal of Sphingomonas wittichii sp. nov. for strain RW1T, known as a dibenzo-p-dioxin metabolizer. Int. J. Syst. Evol. Microbiol. 51, 281–292.
Keywords: Sphingomonas wittichii RW1, nitrous oxide, nitrite reductase, nitric oxide reductase, nitric oxide dioxygenase, denitrification
Citation: Cua LS and Stein LY (2014) Characterization of denitrifying activity by the alphaproteobacterium, Sphingomonas wittichii RW1. Front. Microbiol. 5:404. doi: 10.3389/fmicb.2014.00404
Received: 06 May 2014; Accepted: 17 July 2014;
Published online: 06 August 2014.
Edited by:
Martin G. Klotz, University of North Carolina at Charlotte, USAReviewed by:
Asa Frostegard, Norwegian University of Life Sciences, NorwayJames Moir, University of York, UK
Copyright © 2014 Cua and Stein. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Lisa Y. Stein, Department of Biological Sciences, Faculty of Science, University of Alberta, CW 405 Bio Sci Centre, Edmonton, AB, Canada e-mail:bGlzYS5zdGVpbkB1YWxiZXJ0YS5jYQ==
†Present address: Lynnie S. Cua, County of San Diego, Department of Environmental Health, Vector Disease and Diagnostic Laboratory, San Diego, CA, USA
 Lynnie S. Cua
Lynnie S. Cua Lisa Y. Stein
Lisa Y. Stein