- 1Molecular Microbiology and Infection Biology, Centro de Investigaciones Biológicas, Consejo Superior de Investigaciones Científicas, Madrid, Spain
- 2Faculty of Medicine and Health Sciences, Universiti Sultan Zainal Abidin, Kuala Terengganu, Terengganu, Malaysia
- 3Department of Molecular Microbiology, National Medicines Institute, Warsaw, Poland
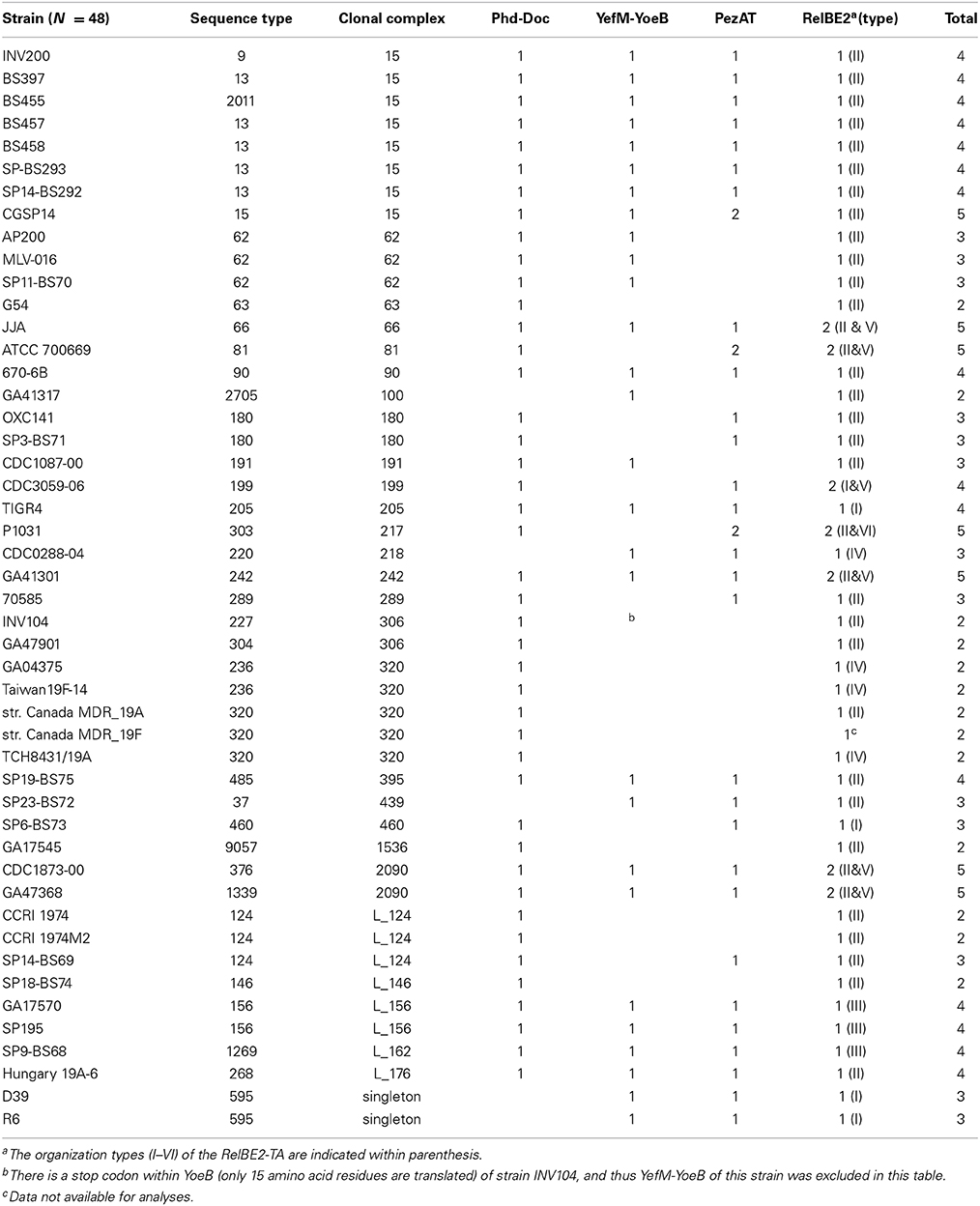
Bacterial toxin-antitoxin (TAs) loci usually consist of two genes organized as an operon, where their products are bound together and inert under normal conditions. However, under stressful circumstances the antitoxin, which is more labile, will be degraded more rapidly, thereby unleashing its cognate toxin to act on the cell. This, in turn, causes cell stasis or cell death, depending on the type of TAs and/or time of toxin exposure. Previously based on in silico analyses, we proposed that Streptococcus pneumoniae, a pathogenic Gram-positive bacterium, may harbor between 4 and 10 putative TA loci depending on the strains. Here we have chosen the pneumococcal strain Hungary19A-6 which contains all possible 10 TA loci. In addition to the three well-characterized operons, namely relBE2, yefM-yoeB, and pezAT, we show here the functionality of a fourth operon that encodes the pneumococcal equivalent of the phd-doc TA. Transcriptional fusions with gene encoding Green Fluorescent Protein showed that the promoter was slightly repressed by the Phd antitoxin, and exhibited almost background values when both Phd-Doc were expressed together. These findings demonstrate that phd-doc shows the negative self-regulatory features typical for an authentic TA. Further, we also show that the previously proposed TAs XreA-Ant and Bro-XreB, although they exhibit a genetic organization resembling those of typical TAs, did not appear to confer a functional behavior corresponding to bona fide TAs. In addition, we have also discovered new interesting bioinformatics results for the known pneumococcal TAs RelBE2 and PezAT. A global analysis of the four identified toxins-antitoxins in the pneumococcal genomes (PezAT, RelBE2, YefM-YoeB, and Phd-Doc) showed that RelBE2 and Phd-Doc are the most conserved ones. Further, there was good correlation among TA types, clonal complexes and sequence types in the 48 pneumococcal strains analyzed.
Introduction
Toxins-antitoxins (TAs) were discovered in the early 1980s and, later on, their cellular function was extensively studied especially in the past two decades. However, until now the world of TAs is still perplexing. TAs are found abundantly in prokaryotes, especially in free-living bacteria, as well as in archaea, and probably in fungi but so far, not in other eukaryotes (Pandey and Gerdes, 2005). In general, a TA locus consists of two genes organized as an operon, with the antitoxin gene, which is generally precedes the toxin gene, encoding either an untranslated RNA or a labile protein that neutralizes the toxicity of its cognate toxin, whereas the toxin gene encodes a toxin protein that binds to the cellular targets and halts essential cell processes. Under certain circumstances such as stress, the system would be triggered by a rapid degradation of the antitoxin, thus liberating the toxin to act on its target. Most known toxins function as RNases (Christensen et al., 2001; Nariya and Inouye, 2008; Jørgensen et al., 2009; Yamaguchi and Inouye, 2009), whereas other toxins target essential cellular components such as DNA gyrase (Van Melderen, 2002), cell wall (Mutschler et al., 2011), and EF-Tu elongation factor (Castro-Roa et al., 2013). TAs have been classified into five types, depending on how the antitoxin counteracts the toxin. Type II TAs, in which the antitoxin is a protein that binds avidly to the toxin protein to form an inert complex, are the most widely studied (Makarova et al., 2009; Leplae et al., 2011). When found on plasmids, TAs function primarily in the stable maintenance of the plasmid, post-segregationally killing off any plasmid-free daughter cells that developed. The functions of the chromosomally-encoded bacterial TAs are still intriguing. They might not be essential, but they have been reported to be involved in a variety of cellular processes related to global stress responses (Christensen et al., 2001), programmed cell death (Engelberg-Kulka and Glaser, 1999), maintenance of mobilomes (Rowe-Magnus et al., 2003; Szekeres et al., 2007), persistence (Gerdes and Maisonneuve, 2012), biofilm formation (Harrison et al., 2009; Soo and Wood, 2013), niche colonization (Norton and Mulvey, 2012), virulence (Ren et al., 2012), phage abortive infection system (Fineran et al., 2009; Dy et al., 2014), and other important cellular processes (Goeders and Van Melderen, 2014). Some bacterial genomes are bountiful in TAs, e.g., Mycobacterium tuberculosis encodes 30 functional TAs (Ramage et al., 2009), which makes their intricacy bewildering. The ability of TAs to impart growth control may attribute to the slow growth and dormant state, which are the hallmarks of latent tuberculosis infection (Schifano and Woychik, 2014), but the mechanisms involved, and how many TAs may be responsible for these phenotypic traits are still unknown. It is still unclear whether the TAs within the same host could cross-talk. However, some studies have shown that expression of one TA might activate another (Garcia-Pino et al., 2008). Whether numerous TAs would provide cumulative or synergistic effect to the cells is something worth to ponder.
Streptococcus pneumoniae (the pneumococcus) is a Gram-positive bacterium, pathogenic for humans and responsible for both infections in respiratory tracts as well as invasive infections, and associated with significant morbidity and mortality (Chan et al., 2012). Furthermore, clinical isolates of pneumococci show a high degree of variability, probably due to the recombinogenic nature of this bacterium (Claverys et al., 2006; Baquero, 2009). We had previously identified up to 10 putative Type II pneumococcal TAs based on in silico data mining (Chan et al., 2012), and three of them had already been proven to be functional, namely relBE2, yefM-yoeB, and pezAT (Chan et al., 2013). Here, we have undertaken the validation and characterization of the rest of the putative pneumococcal TAs. We show that S. pneumoniae encodes a fourth functional TA operon, phd-doc, which corresponds to a bona fide TA. Further, we experimentally ruled out some of the predicted pneumococcal gene pairs as typical TAs, since some of these putative toxins turned out to be either non-toxic, or possibly toxic but lacking a cognate antitoxin gene.
Materials and Methods
Bacterial Strains and Growth Conditions
S. pneumoniae R6 (Hoskins et al., 2001) and Escherichia coli Top10 [F¯mcrAΔ(mrr-hsdRMS-mcrBC) Φ 80lacZΔM15 ΔlacX74 recA1 araD139 Δ(araAleu)7697 galUgalKrpsL(Strr) endA1 nupG] were used as homologous and heterologous hosts for verification of putative TAs from S. pneumonia Hungary19A-6 (McGee et al., 2001) via overexpression assays. S. pneumoniae R6 was grown in AGCH medium (Lacks, 1968) with 0.3 sucrose and 0.2% yeast extract at 37°C in static water bath, and supplemented with erythromycin (1 μg/ml) or tetracycline (1 μg/ml) when the cells harbored recombinant plasmids pLS1ROM-MCS (Lacks et al., 1986; Ruiz-Masó et al., 2012), or pAST (Ruiz-Cruz et al., 2010), respectively. E. coli cultures were grown in TY medium (Sambrook and Russel, 2001), at 37°C with agitation at 250 rpm (for liquid medium), and supplemented with kanamycin (50 μg/ml) when the cells harbored plasmid pFUS2 (Lemonnier et al., 2003) or its derivatives.
DNA Manipulations, Sequencing, and Sequence Data Analysis
DNA manipulations and other molecular biology techniques were done following standard protocols (Sambrook and Russel, 2001). S. pneumoniae genomic DNA was extracted using the Bacterial Genomic DNA Isolation Kit (Norgen Biotech Corp.). Plasmid DNA from both S. pneumoniae and E. coli was isolated using the High Pure Plasmid Isolation kit (Roche). However, for S. pneumoniae, the plasmid extraction protocols were slightly modified as described (Ruiz-Cruz et al., 2010). DNA fragments and PCR products were purified with the NZYGelpure from NZYTech. DNA was sent for automated Sanger sequencing in Secugen S.L., Centro de Investigaciones Biológicas, CSIC, Madrid. DNA sequences were analyzed using the BioEdit Sequence Alignment Editor version 7.0.4.1 (Hall, 1999).
Construction of Recombinant Plasmids with S. pneumoniae Hungary19A-6 Putative TAs
All plasmids and primers used in this study are shown in Table 1, and the details are described below.
pLS1ROM-MCS derivatives
Seven putative pneumococcal TA genes from strain Hungary19A-6 (GI: 169832377) (Chan et al., 2012) were verified in the S. pneumoniae homologous host. Plasmid pLS1ROM-MCS (Lacks et al., 1986; Ruiz-Masó et al., 2012), which was derived from the streptococcal promiscuous plasmid pMV158 (Espinosa, 2013), harbors a tight maltose-inducible promoter PM. The pneumococcal putative toxin genes along with their Shine-Dalgano (SD) sequences were cloned downstream of promoter PM, and then transformed (Lacks, 1968) into S. pneumoniae R6 for overexpression assays. The seven putative toxin genes were: relE1 (GI: 169832900), hicA (GI: 169834067), genes COG2856CA (GI: 169832377 and 169833989), gene COG2856B (GI: 169832735), doc (GI: 169833045), ant (antirepressor) (GI: 169833276; previously known as bro1, Chan et al., 2012) and bro (baculovirus repeated orfs) (GI: 169834001; previously known as bro2, Chan et al., 2012). In the case of the doc gene, its SD sequences were not apparent and thus we decided to use the SD sequences of its cognate antitoxin phd. Consequently, those which exhibited toxicity to the cells, i.e., doc, ant and bro, were further analyzed by cloning their cognate putative antitoxins phd (GI: 169832994), xreA (GI: 169832832; previously known as xre, Chan et al., 2012) and xreB (GI: 169834410; previously known as xre, Chan et al., 2012), as well as both putative TA operons along with their SD, respectively, into pLS1ROM-MCS, and then subjected to overexpression assays.
pFUS2SD and derivatives
The arabinose-inducible plasmid pFUS2 (Lemonnier et al., 2003) was used to construct recombinant plasmids with the seven putative toxins from S. pneumoniae Hungary19A-6 (as described above) to validate their functionality in E. coli heterologous host. Since TA operons are usually co-transcribed and some of the toxin genes which are located downstream of the antitoxin genes do not have an obvious SD sequence, we had constructed plasmid pFUS2SD (this study) by inserting an SD consensus sequence (AGGAGG) downstream of the arabinose-inducible PBAD promoter of pFUS2 by reverse PCR and then self-ligation. This recombinant plasmid was subsequently used for the cloning of the seven putative toxin genes from Hungary19A-6 strain. In addition, the known toxin yoeB (GI: 15903627) from S. pneumoniae R6 was also constructed in pFUS2SD to serve as a positive control (Chan et al., 2011). All these genes were inserted at the EcoRI restriction site downstream of the SD sequence, and thus the start codons of all these genes were 6 bp apart from the SD sequence. The recombinant plasmids were transformed into E. coli Top10 (Sambrook and Russel, 2001) and used to conduct overexpression assays.
pAST derivatives
To verify the promoter activities and to characterize the transcriptional regulation of the promoter by pneumococcal Phd and Phd-Doc, transcriptional fusions were constructed to measure Green Fluorescent Protein (GFP) fluorescence. Plasmid pAST (Ruiz-Cruz et al., 2010), which harbors a promoter-less gfp gene was used for this study. The putative promoter of the phd-doc operon, the phd gene along with its promoter and the phd-doc genes along with their promoter were inserted upstream of the gfp gene of pAST plasmid, respectively. All the recombinant plasmids were transformed into S. pneumoniae R6 and then to evaluate the promoter activities by measuring GFP fluorescence.
Overexpression Assays of Putative TAs in S. Pneumoniae and E. Coli
Cell cultures of S. pneumoniae R6 harboring the different recombinant plasmids were grown in AGCH medium supplemented with 0.3 sucrose and 0.2% yeast extract (Lacks et al., 1986) until optical density (OD)650 ~ 0.3 and subsequently subcultured to OD650 of ~0.02 in the same medium but with addition of 0.2% maltose to induce promoter PM. OD of the cultures was measured every hour for 8 h. E. coli Top10 with different recombinant plasmids were inoculated in TY medium (Maniatis et al., 1982) supplemented with 0.4% glucose to repress expression from the PBAD promoter, and the cells were allowed to grow until OD600 of ~0.3. Cells were harvested and washed twice with TY medium, followed by suspension in TY medium to OD600 of ~0.02, and then induced with 0.4% arabinose. The assays were repeated three times and the mean values and the standard errors were calculated.
GFP Fluorescence Assays
To characterize the promoter activities of pneumococcal phd-doc, fluorescence assays were conducted as previously described (Ruiz-Cruz et al., 2010). Briefly, S. pneumoniae carrying recombinant plasmids with various gfp-fusions were grown until OD650 ~ 0.3. Various volumes of cells (25 μl to 1 ml) were harvested and resuspended in 200 μl 1 × PBS buffer in a 96-well black plate, respectively. Fluorescence was measured with a Varioskan Flash Spectral Scanning Multimode Reader (Thermoscientific) by excitation at 488 nm with a bandwidth of 12 nm and detection of emission at 515 nm. Three independent cultures were analyzed and results of 1 ml of cultures were chosen to show in this manuscript. The cells were also subsequently used for microscopy examination.
Cell Morphology Examination with Phase-Contrast and Fluorescence Microscopy
Cells from overexpression assays were harvested at 4 h and 8 h after induction. Cells were washed twice with 1 × PBS, resuspended in 4% paraformaldehyde and fixed at room temperature for 30 min. Cells were again washed once with 1 × PBS buffer and resuspended in the same buffer. The fixed cells were placed on polylysine coated microscope slides, air-dried and mounted with Vectashield (Vector Laboratories) before being covered with cover slides, and sealed with nail polish. The cells were examined under microscope (Olympus CKX41), and pictures were taken at 100 × magnification. To examine pneumococcal cells harboring various gpf-fused promoter-phd-doc recombinant plasmids, 10 μl of cells in 1 × PBS buffer from Section GFP Fluorescence Assays were viewed under the phase-contrast and fluorescence microscopy. Pictures were taken at 100 × magnification, 200 ms.
Bioinformatics Analyses
Genome databases in NCBI (http://www.ncbi.nlm.nih.gov/) and Pfam (http://pfam.sanger.ac.uk/) were used for bioinformatics analyses. Sequence alignments were conducted using web tool Clustal Omega (http://www.ebi.ac.uk/Tools/msa/clustalo/) and the BioEdit Sequence Alignment analytical software, whereas neighbor-joining phylogenetic trees were generated with Composition Vector Tree Version 2 (http://tlife.fudan.edu.cn/cvtree/) and MEGA6.05. A web-based resource for bacterial integrative and conjugative elements (ICEs) termed ICEberg (http://db-mml.sjtu.edu.cn/ICEberg/) (Bi et al., 2012) was used to detect the presence of TAs in pneumococal ICEs. PHAST (PHAge Search Tool) (http://phast.wishartlab.com/) (Zhou et al., 2011) was used to identify TAs in prophage sequences within pneumococcal genomes. In the case of strains, whose sequence types (STs) were not directly available, sequences of seven loci used in the multilocus sequence typing (MLST) scheme (Enright and Spratt, 1998) were retrieved from genomic data and submitted to the online database http://pubmlst.org/spneumoniae/ (28th August 2014, date last accessed) to obtain allele numbers and STs. Position of the STs was then analyzed for all strains within the population snapshot, constructed based on the content of the whole MLST database (28th August 2014, date last accessed) using the eBURST approach (Feil et al., 2004) at http://eburst.mlst.net/ and resulting clonal complexes (CCs) were named based on the predicted founder of the group. The largest observed CC, including ST156 as a proposed primary founder, was additionally divided into lineages named according to the predicted secondary founders.
Results and Discussion
Verification of the Putative TAs from S. Pneumoniae Hungary19A-6 in S. Pneumoniae R6 and in E. Coli Top10
Between 4 and 10 putative TAs were found in our previous in silico analysis of S. pneumoniae genomes (Chan et al., 2012). Genes of the putative TAs analyzed here were derived from the Hungary19A-6 strain. It is a pathogenic strain, which harbors all 10 putative TA pairs, thus covering most (if not all) of the putative pneumococcal TAs reported to date. Three out of the 10 TA operons, namely the R6 strain relBE2, yefM-yoeB and pezAT, had been previously shown to encode functional TAs in S. pneumoniae and also in E. coli (Nieto et al., 2006, 2007; Khoo et al., 2007). Thus, these three pairs were not included here, with the exception of the yoeB toxin gene that was used as a positive control for the overexpression assays in the E. coli host.
The functionality of the seven putative toxins was first analyzed in their natural host S. pneumoniae and the ones that retarded cell growth were further analyzed with their cognate antitoxins. In the case of the pneumococcal doc gene, the construction of plasmid pLS1ROM-MCS harboring doc had a mutation at the SD in which ′AGGAGG′ was changed to ′AGGCGG′. However, this mutation did not hinder Doc protein to exhibit its toxicity to the cell. Overproduction of Doc had strongly retarded the growth of S. pneumoniae despite the growth resumed after 6 h (Figure 1A). Nonetheless, its toxicity could be clearly neutralized by co-expression of its cognate antitoxin Phd in cis (Figure 1B). This finding demonstrated that the pneumococcal phd-doc pair constitutes a bona fide functional TA. For Ant and Bro, cell growth was also inhibited when the proteins were overproduced, respectively, albeit less toxic than Doc (Figure 1A). Growth of S. pneumoniae cells resumed after a few hours of induction of the toxins (Figure 1A), indicating the toxicity was temporal. Unexpectedly, co-expression of their presumptive cognate antitoxins XreA and XreB did not neutralize the toxicity of Ant and Bro, respectively (Figure 1B). Expression of XreA and XreB alone was not toxic to the cells, but intriguingly, the pairs XreA-Ant and Bro-XreB seemed to be slightly more toxic to the cells, compared to the Ant and Bro toxins alone (Figure 1B). Conversely, overproduction of the rest of the putative pneumococcal toxins HicA, RelE1, COG2856CA, and COG2856B had no effect on the growth of S. pneumoniae (Figure 1A).
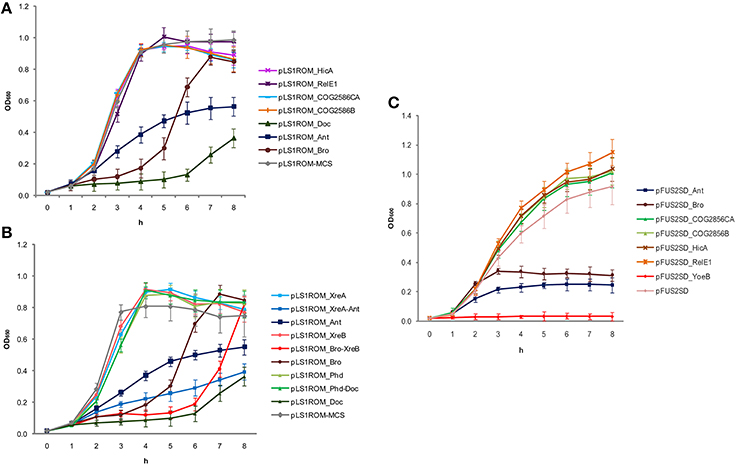
Figure 1. Overexpression of pneumococcal putative TAs. (A) Overexpression of pneumococcal putative toxins in homologous host S. pneumoniae had shown that HicA, RelE1, COG2856CA, and COG2856C were not harmful to the cells; whereas overexpression of Doc, Ant and Bro had thwarted cell growth but resumed a few hours albeit slowly. (B) Since Doc, Ant and Bro were toxic to the cells, they were thus further analyzed to ensure if their putative cognate antitoxins were functional as well in S. pneumoniae host. For Doc, co-expression of its cognate antitoxin Phd in cis had neutralized the Doc toxicity to the cell, signifying Phd-Doc is a functional TA. On the other hand, overexpression of Ant and Bro repressed the growth of the cells but co-expression of their cognate putative antitoxins XreA and XreB, respectively, were not able to neutralize the toxic effect of the toxins, and instead more toxic to the cells. Note that XreA or XreB alone was inert to the cells. Thus, we have ruled out that XreA-Ant and Bro-XreB are typical TAs. (C) Overexpression assays in heterologous host E. coli. Similar to the natural host S. pneumoniae, overexpression of pneumococcal putative toxins HicA, RelE1, COG2856CA, and COG2856C did not affect cell growth. Unfortunately we were not able to construct doc toxin in pFUS2SD plasmid likely due to its high toxicity. YoeB from strain R6, which is a known functional pneumococcal toxin (Nieto et al., 2007; Chan et al., 2011) was used as a positive control (pFUS2SD_YoeB), whereas pFUS2SD without insert served as a negative control. As with the pneumococcal YoeB toxin, overexpression of pneumococcal Ant and Bro inhibited growth of E. coli and no recovery was observed even after 8 h of incubation, which was in contrary to the results in S. pneumoniae, in which cell growth was resumed a few hours after toxins overproduction.
In the case of the heterologous host E. coli, as in their native host S. pneumoniae, overexpression of HicA, RelE1, COG2856CA, and COG2856B were inert to the cells (Figure 1C). We were unable to obtain the recombinant plasmid of pFUS2SD with the putative pneumococcal toxin doc gene, most likely because of its high toxicity to the host. As it was observed for YoeB, overexpression of the Ant and Bro proteins led to suppression of the growth of E. coli; moreover, in contrast to S. pneumoniae, no indication that the cultures resumed growth was found even after 8 h of incubation, indicating that both proteins were very toxic to the host (Figure 1C).
We conclude that out of the seven gene pairs that we had investigated here, only Phd-Doc was demonstrated to be a bona fide pneumococcal TA. Concerning Ant and Bro, overexpression of each of these proteins inhibited growth of both S. pneumoniae and E. coli; however, the products of their adjacent genes XreA and XreB were unable to counteract their toxicity, respectively. Therefore, we do not classify them as typical TA pairs. Thus, we have re-evaluated the number of functional bona fide TAs for the 48 pneumococcal strains whose genomic sequences are available in the NCBI genome databases (Table 2, see below).
The Pneumococcal Phd-Doc: The Fourth Functional TA Pair in S. Pneumoniae
As mentioned above, the pneumococcal Doc was shown to be toxic to its natural host, and co-expression of the cognate Phd antitoxin was able to counteract the toxicity of Doc. The pneumococcal phd-doc gene pair was present in a single copy in almost all the genomes of pneumococcal strains examined (43 out of 48 strains; Table 2) (Chan et al., 2012). The phd-doc genes are structured as an operon, which was consistently surrounded by genes encoding Type I restriction-modification systems; further, a gene that encodes a putative integrase/recombinase was also conserved downstream of doc (Figure 2A). However, the genomic context of phd-doc was not entirely conserved since, in some strains, transposases were also evident further downstream. The pneumococcal phd antitoxin and doc toxin genes overlap by four nucleotides and both genes are likely co-transcribed from a single promoter located upstream of phd (Figure 2A). An inverted repeat of 8 nt (GTAA·TTAC) was identified, which partly covered the −35 consensus sequences (Figure 2A). This differs from the promoter of phd-doc of bacteriophage P1, in which two palindrome sequences (TGTGT·ACACA and CGAGT·ACACG) were found to be located between the −10 consensus sequence and the start codon of Phd, with one of the palindromes overlapping the SD (Magnuson et al., 1996). We postulate that the inverted repeat would function as the operator where the pneumococcal Phd antitoxin binds to repress its own transcription, as in other TAs (Magnuson et al., 1996; Tian et al., 2001; Kedzierska et al., 2007). To test the regulation of transcription of pneumococcal Phd-Doc, we constructed a set of pneumococcal plasmids based on the pAST promoter-probe vector that harbors the gfp gene (Ruiz-Cruz et al., 2010). Thus, transcriptional fusions allowed us to measure the phd-doc promoter. In addition, and to demonstrate whether Phd antitoxin and the Phd-Doc pair regulate their transcription, either the antitoxin or the TA pair was fused to the promoter-less gene encoding GFP. Measurement of the levels of fluorescence showed that indeed the promoter region resulted in GFP expression of ~8 arbitrary units, which were slightly repressed by synthesis of the antitoxin (~6 arbitrary units) and repressed further to ~1 arbitrary units (down to nearly basal levels ~0.1 arbitrary units) by the joint production of the Phd-Doc pair (Figure 2B). In agreement to these results, observation of these cells under phase-contrast and fluorescence microscopy also demonstrated that the fluorescence was not much different for cells harboring promoter-gfp and promoter-phd-gfp; whereas the fluorescence of cells harboring promoter-phd-doc-gfp was visibly lower (Figure 2C). Employment of the GFP reporter to detect TA activation was previously shown for the Salmonella sehAB operon (De La Cruz et al., 2013). In our case, it has turned out to be a simple and reliable way to show auto-repression of the phd-doc operon. Further, it could be used for any of the TA to be tested and also to perform in vivo studies on conditions that trigger any particular TA. To our understanding, this is the first time that this strategy is performed for a Gram-positive bacterium.
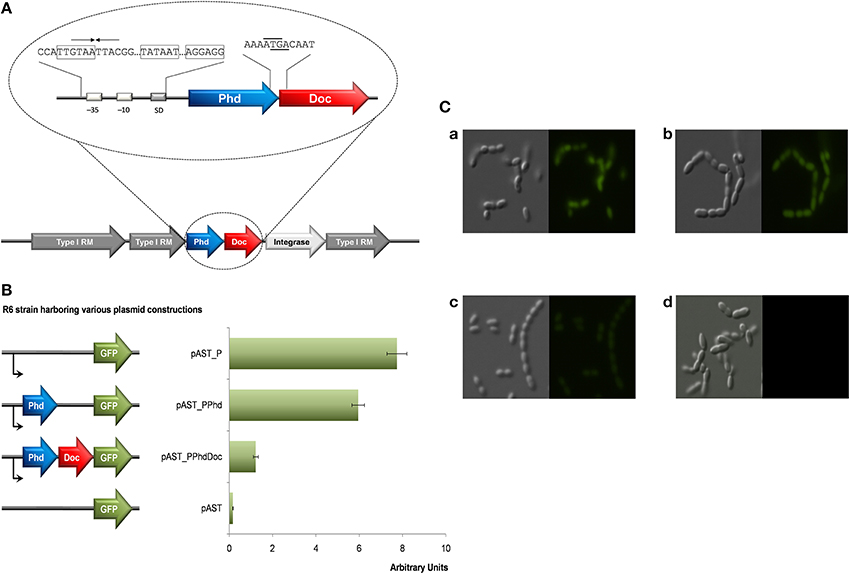
Figure 2. Gene organization and expression of the pneumococcal phd-doc locus. (A) The pneumococcal phd-doc is surrounded by genes that encode Type I restriction-modification (RM) proteins and integrase/recombinase. The pneumococcal phd antitoxin gene is located upstream of its cognate toxin gene doc, and both genes overlap by 4 nucleotides. A promoter and a Shine-Dalgano (SD) were identified upstream of phd, and an inverted repeat (denoted by arrows) was found overlapping the −35 sequences (boxed); the −10 region of the putative promoter and the SD sequences are also boxed. The stop codon of Phd is underlined and the start codon of Doc is indicated in overline. (B) Expression and self-regulation of the phd-doc operon. Cultures (1 ml) of S. pneumoniae R6 cells harboring recombinant plasmids were grown until an OD650 ~0.3. Cells were harvested, resuspended in 1 × PBS buffer and their fluorescence was measured. Plasmids used were: pAST_P, harboring only the promoter of the operon; pAST_PPhd, harboring the promoter and the phd gene; pAST_PPhdDoc, harboring the entire phd-doc operon, and the promoter-less gfp plasmid vector, pAST, used as control. SE (average of three independent experiments) are indicated. (C) Cells examination under phase-contrast and fluorescence microscopy. The GFP level for cells harboring pAST_P (a) and pAST_PPhd (b) did not show much difference; whereas GFP level for pAST_PPhdDoc (c) was visibly lower, which is coincided with the fluorescence measurement. Cells harboring plasmid pAST served as negative control.
Like other Doc homologs, the pneumococcal Doc has a Fic (filamentation induced by cyclic AMP) domain (Utsumi et al., 1982) at the N-terminal moiety. In general, the conserved Fic motif, HxFx(D/E)GNGRxxR (Engel et al., 2012; Goepfert et al., 2013), possesses adenylylation (or “AMPylation”) activity (Itzen et al., 2011; Woolery et al., 2012). By comparison, the consensus residues of the Fic motifs in all Doc toxins are slightly different, HxFx(D/N)(A/G)NKR (Cruz et al., 2014), as is the pneumococcal Fic motif HVFANANKR. The target of the Doc toxin from bacteriophage P1 (Magnuson and Yarmolinksy, 1998) was revised recently, in which Doc was reported to be a new type of protein kinase [instead of AMPylating like other Fic proteins (Kinch et al., 2009) or obstructing ribosomes (Liu et al., 2008)] that inhibits bacterial translation by phosphorylating the conserved threonine (Thr382) of the translation elongation factor EF-Tu, thereby rendering it unable to bind aminoacylated tRNAs (Castro-Roa et al., 2013; Cruz et al., 2014). Interestingly, it was also demonstrated that the Phd antitoxin can only block the de novo phosphorylation of EF-Tu, but cannot revert it, and dephosphorylation of EF-Tu was probably catalyzed by the Doc toxin itself (Castro-Roa et al., 2013). This finding coincided with our observation, in which the inhibition of growth mediated by the pneumococcal Doc toxin was slowly relieved a few hours after overexpression (Figure 1B). It had been shown that ectopic overexpression of Doc in E. coli might lead to Lon-dependent activation of RelE to exhibit its mRNA cleavage activity (Garcia-Pino et al., 2008). Since S. pneumoniae harbors both RelBE and Phd-Doc TAs, it would be interesting to find out whether both TAs could cross-talk as described by Garcia-Pino et al. (2008).
Morphology of E. Coli and S. Pneumoniae Cells after Toxin Overexpression
In many cases where overexpressed genes may cause a toxic effect, the bacterial cells change their morphology and may exhibit a phenotype of either filamentation or other gross changes in the cell shape (Scott et al., 2010). To test whether overexpression of the various toxins analyzed here had any influence on the cell morphology of S. pneumoniae and E. coli, cultures were subjected to toxin overexpression under the above conditions, and cells were observed using phase-contrast microscopy. In the case of S. pneumoniae no prominent morphological changes were observed for any cells that overexpressed the different toxins (Supplementary Figure S1). On the other hand, for E. coli, filamentation was observed when Ant or Bro were overexpressed, indicating that the cells were not dividing properly and thus cell growth was impaired (Figures 3C,D). The filamentation was not reversible for at least 8 h as inhibition of cell growth by Ant or Bro did not show any sign of resuming even after 8 h (Figure 1C). Overexpression of YoeB toxin, which likely an endoribonuclease as its E. coli homolog (Christensen et al., 2004; Zhang and Inouye, 2009), did not result in very obvious differences in morphology when compared to the wild type (Figures 3A,B).
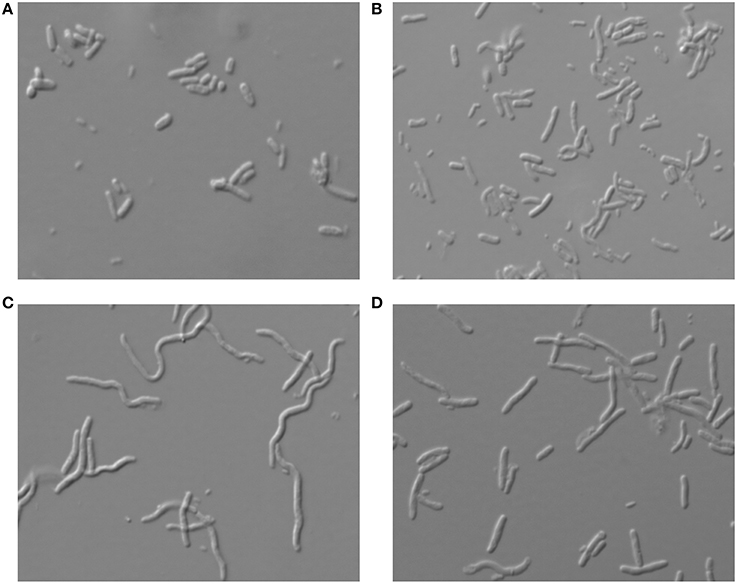
Figure 3. Morphology of E. coli cells under microscope at 100 × magnification after overexpression of the respective pneumococcal toxins. Morphology of the cells which harbor (A) the negative control pFUS2SD, (B) the positive control pFUS2SD_YoeB, (C) pFUS2SD_Ant and (D) pFUS2SD_Bro, were examined after 8 h of overexpression. Filamented cells were observed in cultures overexpressing Ant or Bro, but no prominent differences were found for cells overexpressing YoeB when compared to wild type.
Other TAs in S. Pneumoniae and their Genetic Organization
The relBE1 operon
A previous study showed that RelE1 of S. pneumoniae had no RNase activity (Christensen and Gerdes, 2003), but the cell growth profile after overexpression of RelE1 was not assessed. Here, we confirmed that expression of RelE1 was not detrimental to its homologous host or to the heterologous host E. coli (Figure 1) even though RelE1 shared 52% similarity with RelE2 (in the Hungary19A-6 strain) which had been demonstrated to be functional (Nieto et al., 2006). In spite of the high conservation of both relB1 and relE1 in all the strains (not shown), two (R65 and R85) out of five crucial residues responsible for RelE toxicity in Pyrococcus horikoshii were missing in the pneumococcal RelE1 (Takagi et al., 2005; Nieto et al., 2010; Chan et al., 2012). In addition, truncation of relE1 in some strains, such as ATCC 700669 and 70585 was also observed. In general, the gene arrangement around the relBE1 genes of all the strains was similar, i.e., these genes were flanked upstream by genes that encode DNA polymerase III, elongation factor G and 30s ribosomal proteins, whereas downstream genes were found to be encoding a possible amino peptidase, an integral membrane protein and ribosomal small subunit pseudouridine synthase. In some instances, transposases were identified further upstream of relB1 (strains ATCC 700669 and INV104). Conservation of the relBE1 gene pair in many of the pneumococcal genomes could point to a function not related to type II TA, but this remains to be investigated.
The gene pairs xreA-ant and bro-xreB
These two gene pairs were previously proposed as putative TAs in bioinformatics searches of the S. pneumoniae genome (Makarova et al., 1999; Chan et al., 2012). Overexpression of the pneumococcal Ant and Bro proteins inhibited cell growth in both S. pneumoniae and E. coli, but co-expression of their putative respective cognate genes xreA and xreB did not counteract their toxicity. However, we still do not rule out the possibility that Ant and Bro are solitary toxins, similar to the MazF-mx of M. xanthus, which lacks the co-transcribed antitoxin (Nariya and Inouye, 2008). MrpC, which is expressed from another part of the M. xanthus genome, serves as an antitoxin by forming complexes with MazF-mx, and it also positively regulates the MazF-mx expression (Nariya and Inouye, 2008).
We made use of a web server designed to identify prophages (PHAST; Zhou et al., 2011) to search whether pneumococcal xreA-ant and bro-xreB gene pairs were located within pneumococcal prophages. We found that they are located within a 94.4 kb intact prophage termed Streptococcus phage MM1 of S. pneumoniae Hungary19A-6 strain (Supplementary material). Analysis with Pfam showed that both Ant and Bro have two domains, respectively: Ant (232 amino acids) has an AntA (AntA/AntB antirepressor) domain at the N-terminal moiety and a phage antirepressor KilAC domain at the C-terminal moiety (Figure 4A); whereas Bro (237 amino acids) has a Bro-N domain at the N-terminus and an ORF6C (C-terminal of bacteriophage bIL2850 ORF6) (Iyer et al., 2002) domain at the C-terminus (Figure 4B). Both KilA and Bro domains are widely prevalent in bacteriophages, bacteria, eukaryotic viruses of the nucleo-cytoplasmic large DNA virus and baculovirus classes (Iyer et al., 2002). A study by Kuan et al. showed that the protein ANT8 (ORF8) of lytic corynephage P1201 isolated from Corynebacterium glutamicum harbors a Bro domain at the N-terminus and an antirepressor domain at the C-terminal end (http://research.nchu.edu.tw/upfiles/ADUpload/oc_downmul2272130635.pdf). In this case, the DNA binding ability of ANT8 was contributed by the Bro domain, whereas overexpression of the antirepressor domain inhibited E. coli cell growth and also killed the cells as shown by a decrease in colony forming units. Under microscopic examination, the cells that overexpressed the ANT8 protein became filamentous, with irregular multiple nucleoids, indicating inhibition of cell division, and this result is also in agreement with our findings.
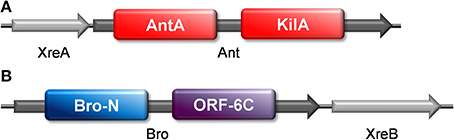
Figure 4. The domains of pneumococcal Ant and Bro toxins. Sequence analyses from Pfam showed that (A) pneumococcal Ant harbors two domains, with the AntA domain at the N-terminus and KilAC domain at the C-terminus; whereas (B) pneumococcal Bro harbors a Bro-N domain at the N-terminal and an ORF6C domain at the C-terminal.
Genomic Context of the Functional Pneumococcal TAs relBE2 and pezAT, and their Distribution in S. Pneumoniae Population
The distribution of functional TAs showed a very good correlation with pneumococcal STs and CCs (Table 2). As described in our previous report, RelBE2 is present in all the 48 pneumococcal database strains with seven out of them having two copies. Moreover, six different gene organizations were found for this TA (Chan et al., 2012) that fall into two groups, encompassing the previously defined Types I-IV and Types V-VI (Chan et al., 2012; Figure 5A). The latter are flanked by xre upstream and COG2856B downstream, an arrangement that is not observed in Types I-IV. RelE2 of Types V and VI were also clustered together in the phylogenetic analyses (Figure 5B). In addition, in strains with two copies of RelBE2, one copy belonged to Types I-IV whereas the second copy belonged to either Type V or VI. Integrases and IS elements were evident in Types III-VI. The particular gene arrangement showed also a very good agreement with STs and CCs, with a single exception in the Taiwan19F-14 international clone (Table 2). All these observations suggest that the second copy of relBE2 may likely be horizontally acquired by some strains rather than represent a product of gene duplication.
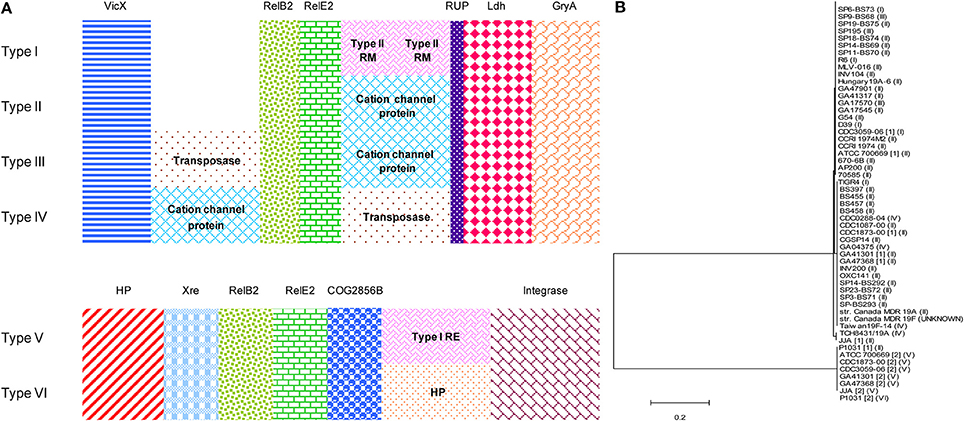
Figure 5. Gene organization and phylogenetic analyses of pneumococcal RelE2 homologs. (A) Polymorphisms associated with the gene organization of relBE2 and their neighboring genes found in 48 pneumococcal strains. Six variations of gene organizations were found (Type I–VI). In general, the relBE2 operon was flanked upstream by vicX (metal-dependent hydrolase), whereas a RUP (repeat unit of pneumococcus), element, ldh (lactate dehydrogenease), and gyrA (the A subunit of DNA gyrase) are located downstream for Type I–IV. For Type V–VI, the relBE2 cassette is flanked upstream by xre-like protein and COG2856B downstream. Mobile elements are seen in Types III–VI. Other abbreviations used: restriction-modification (RM); restriction-endonuclease (RE); hypothetical protein (HP). (B) Phylogenetic analyses of RelE2 homologs with neighbor-joining algorithm, in which the determined start codon of relE2 was considered (Moreno-Córdoba et al., 2012), had shown that Type V and Type VI gene organizations of RelE2 cluster together, indicating RelE2 from both types are of more similar in sequences. The type of relBE2 gene organization was indicated. For the strain Canada MDR 19F, the type of gene arrangement is unknown due to data unavailability in the NCBI databases.
In our previous bioinformatics search, 28 out of the 48 annotated pneumococcal strains harbored a single copy of pezAT; whereas three strains harbored two copies (Chan et al., 2012). The two copies of pezAT within the same strain were almost identical (similarities of 96–97%), and the catalytically important residues (Meinhart et al., 2003) were also conserved. The gene organization of pezAT among the various S. pneumoniae strains is even more varied than relBE2. This TA is flanked by a number of genes that contribute to site-specific recombination and even genes that play roles in conjugative transfer. The pezAT locus is known to be present on a genomic island known as the pneumococcal pathogenicity island 1 (Meinhart et al., 2003; Nieto et al., 2006) but interestingly, a search using a web-based resource for bacterial ICEs known as ICEberg database led us to discover the presence of a previously unreported pezAT operon in a putative pneumococcal ICE designated Tn5253 (Ayoubi et al., 1991). This was corroborated when a recent analysis of the complete sequence of Tn5253 from S. pneumoniae DP1322 indicated the presence of two copies of pezAT (Iannelli et al., 2014) which comprised part of a 1169 bp direct repeat that flanks the Ω cat(pC194) element within Tn5253. Experimental evidence showed that excision of Ω cat(pC194) from Tn5253 occurred by a recombination event within the pezAT-containing repeat segment leaving one copy in Tn5253 and another copy in the newly-excised plasmid pC194 (Iannelli et al., 2014). Besides that, ectopic integration of Ω cat(pC194) to other regions of the S. pneumoniae genome was also detected, likely by homologous recombination involving the pezAT-containing direct repeat segments (Iannelli et al., 2014). Thus, pezAT appeared to play a direct role in the transfer of the Ω cat(pC194) element and may also help in the stable maintenance of this element in its hosts. It is therefore not surprising that phylogenetic analyses showed that PezAT found in Tn5253 formed a separate cluster from PezAT located in the pneumococcal pathogenicity island 1 (Supplementary Figure S2 and Table S1).
Conclusions
Previous searches for putative type II TAs in S. pneumoniae (Chan et al., 2012, 2013) led us to conclude that their number was greatly undervalued. However, our present in vivo validation has shown that, indeed, only four of the TAs, namely RelBE2 (Nieto et al., 2006), YefM-YoeB (Nieto et al., 2007; Chan et al., 2011), PezAT (Khoo et al., 2007; Mutschler et al., 2011) and now, Phd-Doc can be considered as bona fide TAs in the “classical” sense that the harmful effects of the toxins must be counteracted by their cognate antitoxins and that their two genes should be organized as an operon (Gerdes, 2013). Bioinformatics search approaches have helped us to identify various putative TA homologs; however, the actual number of bona fide TAs might be overestimated as in our case e.g., pneumococcal relBE2 is functional TA but pneumococcal relBE1 is apparently not. In the case of M. tuberculosis, of the 88 putative TAs identified, only 30 are functional (Ramage et al., 2009). Other approaches like shotgun cloning as presented by Sberro et al. (2013) could be another alternative and more thorough approach to identify TAs. However, there might be more TAs than these four if we consider Ant and Bro as possible solo toxins whose respective antitoxins might be located elsewhere in the pneumococcal chromosome. The finding that these two novel proteins might be associated with lysogenic bacteriophages is, indeed, appealing and merits an in-depth study of their features.
Author Contributions
Manuel Espinosa, Chew Chieng Yeo, Ewa Sadowy and Wai Ting Chan designed the study. Wai Ting Chan performed laboratory work and data analysis. Manuel Espinosa, Chew Chieng Yeo, Ewa Sadowy and Wai Ting Chan wrote the manuscript. All authors read and approved the final manuscript.
Conflict of Interest Statement
The reviewer Ramon Diaz Orejas declares that, despite being affiliated at the same institution as the authors Wai Ting Chan and Manuel Espinosa, the review process was handled objectively and no conflict of interest exists. The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
Thanks are also due to Dr. F. Gasset-Rosa and D. Lobato-Márquez for their help with the microscope analyses. Research financed by Ministry of Economy and Competitiveness, grant CSD-2008/00013 (INTERMODS) to Manuel Espinosa.
Supplementary Material
The Supplementary Material for this article can be found online at: http://www.frontiersin.org/journal/10.3389/fmicb.2014.00677/abstract
Abbreviations
TAs, toxins-antitoxins; SD, Shine-Dalgarno; ant, antirepressor; bro, baculovirus repeated orfs; GFP, Green Fluorescent Protein; OD, optical density; STs, sequence types; ICEs, integrative and conjugative elements; MLST, multilocus sequence typing; CCs, clonal complexes; Fic, filamentation induced by cyclic AMP.
References
Ayoubi, P., Kilic, A. O., and Vijayakumar, M. N. (1991). Tn5253, the pneumococcal omega (cat tet) BM6001 element, is a composite structure of two conjugative transposons, Tn5251 and Tn5252. J. Bacteriol. 173, 1617–1622.
Baquero, F. (2009). Environmental stress and evolvability in microbial systems. Clin. Microbiol. Infect. 15, 5–10. doi: 10.1111/j.1469-0691.2008.02677.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Bi, D., Xu, Z., Harrison, E. M., Tai, C., Wei, Y., He, X., et al. (2012). ICEberg: a web-based resource for integrative and conjugative elements found in Bacteria. Nucleic Acids Res. 40, D621–D626. doi: 10.1093/nar/gkr846
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Castro-Roa, D., Garcia-Pino, A., De Gieter, S., Van Nuland, N. A. J., Loris, R., and Zenkin, N. (2013). The Fic protein Doc uses an inverted substrate to phosphorylate and inactivate EF-Tu. Nat. Chem. Biol. 9, 811–817. doi: 10.1038/nchembio.1364
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Chan, W. T., Moreno-Córdoba, I., Yeo, C. C., and Espinosa, M. (2012). Toxin-antitoxin genes of the gram-positive pathogen Streptococcus pneumoniae: so few and yet so many. Microbiol. Mol. Biol. Rev. 76, 773–791. doi: 10.1128/MMBR.00030-12
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Chan, W. T., Moreno-Córdoba, I., Yeo, C. C., and Espinosa, M. (2013). “Toxin-antitoxin loci in Streptococcus pneumoniae,” in Prokaryotic toxin-antitoxins, ed K. Gerdes (Berlin: Springer-Verlag Berlin Heidelberg), 315–339. doi: 10.1007/978-3-642-33253-1_18
Chan, W. T., Nieto, C., Harikrishna, J. A., Khoo, S. K., Yasmin Othman, R., Espinosa, M., et al. (2011). Genetic regulation of the yefM-yoeBSpn toxin-antitoxin locus of Streptococcus pneumoniae. J. Bacteriol. 193, 4612–4625. doi: 10.1128/JB.05187-11
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Christensen, K. S., Maenhauf-Michel, G., Mine, N., Gothesman, S., Gerdes, K., and Van Melderen, L. (2004). Overproduction of the Lon protease triggers inhibition of translation in Escherichia coli: involvement of the yefM-yoeB toxin-antitoxin system. Mol. Microbiol. 51, 1705–1717. doi: 10.1046/j.1365-2958.2003.03941.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Christensen, S. K., and Gerdes, K. (2003). RelE toxins from bacteria and archaea cleave mRNAs on translating ribosomes, which are rescued by tmRNA. Mol. Microbiol. 48, 1389–1400. doi: 10.1046/j.1365-2958.2003.03512.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Christensen, S. K., Mikkelsen, M., Pedersen, K., and Gerdes, K. (2001). RelE, a global inhibitor of translation, is activated during nutritional stress. Proc. Natl. Acad. Sci. U.S.A. 98, 14328–14333. doi: 10.1073/pnas.251327898
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Claverys, J. P., Prudhomme, M., and Martin, B. (2006). Induction of competence regulons as general stress responses in Gram-positive bacteria. Annu. Rev. Microbiol. 60, 451–475. doi: 10.1146/annurev.micro.60.080805.142139
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Cruz, J. W., Rothenbacher, F. P., Maehigashi, T., Lane, W. S., Dunham, C. M., and Woychik, N. A. (2014). Doc toxin is a kinase that inactivates elongation factor Tu. J. Biol. Chem. 289, 7788–7798. doi: 10.1074/jbc.M113.544429
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
De La Cruz, M. A., Zhao, W., Farenc, C., Gimenez, G., Raoult, D., Cambillau, C., et al. (2013). A toxin-antitoxin module of Salmonella promotes virulence in mice. PLoS Pathog. 9:e1003827. doi: 10.1371/journal.ppat.1003827
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Dy, R. L., Przybilski, R., Semeijn, K., Salmond, G. P. C., and Fineran, P. C. (2014). A widespread bacteriophage abortive infection system functions through a Type IV toxin–antitoxin mechanism. Nucleic Acids Res. 42, 4590–4605. doi: 10.1093/nar/gkt1419
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Engel, P., Goepfert, A., Stanger, F. V., Harms, A., Schmidt, A., Schirmer, T., et al. (2012). Adenylylation control by intra- or intermolecular active-site obstruction in Fic proteins. Nature 482, 107–110. doi: 10.1038/nature10729
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Engelberg-Kulka, H., and Glaser, G. (1999). Addiction modules and programmed cell death and antideath in bacterial cultures. Annu. Rev. Microbiol. 53, 43–70. doi: 10.1146/annurev.micro.53.1.43
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Enright, M. C., and Spratt, B. G. (1998). A multilocus sequence typing scheme for Streptococcus pneumoniae: identification of clones associated with serious invasive disease. Microbiology 144, 3049–3060. doi: 10.1099/00221287-144-11-3049
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Espinosa, M. (2013). Plasmids as models to study macromolecular interactions: the pMV158 paradigm. Res. Microbiol. 164, 199–204. doi: 10.1016/j.resmic.2013.01.006
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Feil, E. J., Li, B. C., Aanensen, D. M., Hanage, W. P., and Spratt, B. G. (2004). eBURST: inferring patterns of evolutionary descent among clusters of related bacterial genotypes from multilocus sequence typing data. J. Bacteriol. 186, 1518–1530. doi: 10.1128/JB.186.5.1518-1530.2004
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Fineran, P. C., Blower, T. R., Foulds, I. J., Humphreys, D. P., Lilley, K. S., and Salmond, G. P. (2009). The phage abortive infection system, ToxIN, functions as a protein-RNA toxin-antitoxin pair. Proc. Natl. Acad. Sci. U.S.A. 106, 894–899. doi: 10.1073/pnas.0808832106
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Garcia-Pino, A., Christensen-Dalsgaard, M., Wyns, L., Yarmolinsky, M. B., Magnuson, R. D., Gerdes, K., et al. (2008). Doc of prophage P1 is inhibited by its antitoxin partner Phd through fold complementation. J. Biol. Chem. 283, 30821–30827. doi: 10.1074/jbc.M805654200
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Gerdes, K. (ed.). (2013). Prokaryotic Toxin-Antitoxins. Berlin-Heidelberg: Springer. doi: 10.1007/978-3-642-33253-1
Gerdes, K., and Maisonneuve, E. (2012). Bacterial persistence and toxin-antitoxin loci. Annu. Rev. Microbiol. 66, 103–123. doi: 10.1146/annurev-micro-092611-150159
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Goeders, N., and Van Melderen, L. (2014). Toxin-antitoxin systems as multilevel interaction systems. Toxins 6, 304–324. doi: 10.3390/toxins6010304
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Goepfert, A., Stanger, F. V., Dehio, C., and Schirmer, T. (2013). Conserved inhibitory mechanism and competent ATP binding mode for adenylyltransferases with Fic fold. PLoS ONE 8:e64901. doi: 10.1371/journal.pone.0064901
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Hall, T. A. (1999). BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41, 95–98.
Harrison, J. J., Wade, W. D., Akierman, S., Vacchi-Suzzi, C., Stremick, C. A., Turner, R. J., et al. (2009). The chromosomal toxin yafQ is a determinant of multidrug tolerance for Escherichia coli growing in a biofilm. Antimicrob. Agents Chemother. 53, 2253–2258. doi: 10.1128/AAC.00043-09
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Hoskins, J., Alborn, W. E. Jr., Arnold, J., Blaszczak, L. C., Burgett, S., Dehoff, B. S., et al. (2001). Genome of the bacterium Streptococcus pneumoniae strain R6. J. Bacteriol. 183, 5709–5717. doi: 10.1128/JB.183.19.5709-5717.2001
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Iannelli, F., Santoro, F., Oggioni, M. R., and Pozzi, G. (2014). Nucleotide sequence analysis of integrative conjugative element Tn5253 of Streptococcus pneumoniae. Antimicrob. Agents Chemother. 58, 1235–1239. doi: 10.1128/AAC.01764-13
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Itzen, A., Blankenfeldt, W., and Goody, R. S. (2011). Adenylylation: renaissance of a forgotten post-translational modification. Trends Biochem. Sci. 36, 221–228. doi: 10.1016/j.tibs.2010.12.004
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Iyer, L. M., Koonin, E. V., and Aravind, L. (2002). Extensive domain shuffling in transcription regulators of DNA viruses and implications for the origin of fungal APSES transcription factors. Genome Biol. 3, RESEARCH0012. doi: 10.1186/gb-2002-3-3-research0012
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Jørgensen, M. G., Pandey, D. P., Jaskolska, M., and Gerdes, K. (2009). HicA of Escherichia coli defines a novel family of translation-independent mRNA interferases in bacteria and archaea. J. Bacteriol. 191, 1191–1199. doi: 10.1128/JB.01013-08
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Kedzierska, B., Lian, L. Y., and Hayes, F. (2007). Toxin–antitoxin regulation: bimodal interaction of YefM–YoeB with paired DNA palindromes exerts transcriptional autorepression. Nucleic Acids Res. 35, 325–339. doi: 10.1093/nar/gkl1028
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Khoo, S. K., Loll, B., Chan, W. T., Shoeman, R. L., Ngoo, L., Yeo, C. C., et al. (2007). Molecular and structural characterization of the PezAT chromosomal toxin-antitoxin system of the human pathogen Streptococcus pneumoniae. J. Biol. Chem. 282, 19606–19618. doi: 10.1074/jbc.M701703200
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Kinch, L. N., Yarbrough, M. L., Orth, K., and Grishin, N. V. (2009). Fido, a novel AMPylation domain common to fic, doc, and AvrB. PLoS ONE 4:e5818. doi: 10.1371/journal.pone.0005818
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Lacks, S. (1968). Genetic regulation of maltosaccharide utilization in Pneumococcus. Genetics 60, 685–706.
Lacks, S. A., López, P., Greenberg, B., and Espinosa, M. (1986). Identification and analysis of genes for tetracycline resistance and replication functions in the broad-host-range plasmid pLS1. J. Mol. Biol. 192, 753–765. doi: 10.1016/0022-2836(86)90026-4
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Lemonnier, M., Ziegelin, G., Reick, T., Muñoz Gómez, A., Díaz-Orejas, R., and Lanka, E. (2003). P1 Ban protein is a hexameric DNA helicase that interacts with and substitutes for Escherichia coli DnaB. Nucleic Acid Res. 31, 3918–3928. doi: 10.1093/nar/gkg463
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Leplae, R., Geeraerts, D., Hallez, R., Guglielmini, J., Drèze, P., and Van Melderen, L. (2011). Diversity of bacterial type II toxin–antitoxin systems: a comprehensive search and functional analysis of novel families. Nucleic Acids Res. 39, 5513–5525. doi: 10.1093/nar/gkr131
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Liu, M., Zhang, Y., Inouye, M., and Woychik, N. A. (2008). Bacterial addiction module toxin Doc inhibits translation elongation through its association with the 30S ribosomal subunit. Proc. Natl. Acad. Sci. U.S.A. 105, 5885–5890. doi: 10.1073/pnas.0711949105
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Magnuson, R., Lehnherr, H., Mukhopadhyay, G., and Yarmolinsky, M. B. (1996). Autoregulation of the plasmid addiction operon of bacteriophage P1. J. Biol. Chem. 271, 18705–18710. doi: 10.1074/jbc.271.31.18705
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Magnuson, R., and Yarmolinksy, M. B. (1998). Corepression of the P1 addiction operon by Phd and Doc. J. Bacteriol. 180, 6342–6351.
Makarova, K. S., Aravind, L., Galperin, M. Y., Grishin, N. V., Tatusov, R. L., Wolf, Y. I., et al. (1999). Comparative genomics of the Archaea (Euryarchaeota): evolution of conserved protein families, the stable core, and the variable shell. Genome Res. 9, 608–628.
Makarova, K., Wolf, Y. I., and Koonin, E. V. (2009). Comprehensive comparative-genomic analysis of type 2 toxin-antitoxin systems and related mobile stress response systems in prokaryotes. Biol. Direct 4:19. doi: 10.1186/1745-6150-4-19
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Maniatis, T., Fritsch, E. F., and Sambrook, J. (1982). Molecular Cloning: A Laboratory Manual. New York, NY: Cold Spring Harbor Laboratory Press.
McGee, L., McDougal, L., Zhou, J., Spratt, B. G., Tenover, F. C., George, R., et al. (2001). Nomenclature of major antimicrobial-resistant clones of Streptococcus pneumoniae defined by the pneumococcal molecular epidemiology network. J. Clin. Microbiol. 39, 2565–2571. doi: 10.1128/JCM.39.7.2565-2571.2001
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Meinhart, A., Alonso, J. C., Strater, N., and Saenger, W. (2003). Crystal structure of the plasmid maintenance system epsilon/zeta: functional mechanism of toxin zeta and inactivation by epsilon2zeta2 complex formation. Proc. Natl. Acad. Sci. U.S.A. 100, 1661–1666. doi: 10.1073/pnas.0434325100
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Moreno-Córdoba, I., Diago-Navarro, E., Barendregt, A., Heck, A. J. R., Alfonso, C., Díaz-Orejas, R., et al. (2012). The toxin-antitoxin proteins RelBE2Spn of Streptococcus pneumoniae: characterization and association to their DNA target. Proteins 80, 1834–1846. doi: 10.1002/prot.24081
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Mutschler, H., Gebhardt, M., Shoeman, R. L., and Meinhart, A. (2011). A novel mechanism of programmed cell death in bacteria by toxin-antitoxin systems corrupts peptidoglycan synthesis. PLoS Biol. 9:e1001033. doi: 10.1371/journal.pbio.1001033
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Nariya, H., and Inouye, M. (2008). MazF, an mRNA interferase, mediates programmed cell death during multicellular Myxococcus development. Cell 132, 55–66. doi: 10.1016/j.cell.2007.11.044
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Nieto, C., Cherny, I., Khoo, S. K., García De Lacoba, M., Chan, W. T., Yeo, C. C., et al. (2007). The yefM-yoeB toxin-antitoxin systems of Escherichia coli and Streptococcus pneumoniae: functional and structural correlation. J. Bacteriol. 189, 1266–1278. doi: 10.1128/JB.01130-06
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Nieto, C., Pellicer, T., Balsa, D., Christensen, S. K., Gerdes, K., and Espinosa, M. (2006). The chromosomal relBE2 toxin-antitoxin locus of Streptococcus pneumoniae: characterization and use of a bioluminescence resonance energy transfer assay to detect toxin-antitoxin interaction. Mol. Microbiol. 59, 1280–1296. doi: 10.1111/j.1365-2958.2006.05027.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Nieto, C., Sadowy, E., De La Campa, A. G., Hryniewicz, W., and Espinosa, M. (2010). The relBE2Spn toxin-antitoxin system of Streptococcus pneumoniae: role in antibiotic tolerance and functional conservation in clinical isolates. PLoS ONE 5:e11289. doi: 10.1371/journal.pone.0011289
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Norton, J. P., and Mulvey, M. A. (2012). Toxin-antitoxin systems are important for niche-specific colonization and stress resistance of uropathogenic Escherichia coli. PLoS Pathog. 8:e1002954. doi: 10.1371/journal.ppat.1002954
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Pandey, D. P., and Gerdes, K. (2005). Toxin-antitoxin loci are highly abundant in free-living but lost from host-associated prokaryotes. Nucleic Acids Res. 33, 966–976. doi: 10.1093/nar/gki201
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Ramage, H. R., Connolly, L. E., and Cox, J. S. (2009). Comprehensive functional analysis of Mycobacterium tuberculosis toxin-antitoxin systems: implications for pathogenesis, stress responses, and evolution. PLoS Genet. 5:e1000767. doi: 10.1371/journal.pgen.1000767
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Ren, D., Walker, A., and Daines, D. A. (2012). Toxin-antitoxin loci vapBC-1 and vapXD contribute to survival and virulence in nontypeable Haemophilus influenzae. BMC Microbiol. 12:263. doi: 10.1186/1471-2180-12-263
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Rowe-Magnus, D. A., Guerout, A. M., Biskri, L., Bouige, P., and Mazel, D. (2003). Comparative analysis of superintegrons: engineering extensive genetic diversity in the Vibrionaceae. Genome Res. 13, 428–442. doi: 10.1101/gr.617103
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Ruiz-Cruz, S., Solano-Collado, V., Espinosa, M., and Bravo, A. (2010). Novel plasmid-based genetic tools for the study of promoters and terminators in Streptococcus pneumoniae and Enterococcus faecalis. J. Microbiol. Methods 83, 156–163. doi: 10.1016/j.mimet.2010.08.004
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Ruiz-Masó, J. A., López-Aguilar, C., Nieto, C., Sanz, M., Burón, P., Espinosa, M., et al. (2012). Construction of a plasmid vector based on the pMV158 replicon for cloning and inducible gene expression in Streptococcus pneumoniae. Plasmid 67, 53–59. doi: 10.1016/j.plasmid.2011.09.001
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Sambrook, J., and Russel, D. W. (2001). Molecular Cloning: A Laboratory Manual. New York, NY: Cold Spring Harbor Laboratory Press.
Sberro, H., Leavitt, A., Kiro, R., Koh, E., Peleg, Y., Qimron, U., et al. (2013). Discovery of functional toxin/antitoxin systems in bacteria by shotgun cloning. Mol. Cell 50, 136–148. doi: 10.1016/j.molcel.2013.02.002
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Schifano, J. M., and Woychik, N. A. (2014). 23S rRNA as an a-Maz-ing new bacterial toxin target. RNA Biol. 11, 101–105. doi: 10.4161/rna.27949
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Scott, M., Gunderson, C. W., Mateescu, E. M., Zhang, Z., and Hwa, T. (2010). Interdependence of cell growth and gene expression: origins and consequences. Science 330, 1099–1102. doi: 10.1126/science.1192588
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Soo, V. W., and Wood, T. K. (2013). Antitoxin MqsA represses curli formation through the master biofilm regulator CsgD. Sci. Rep. 3, 3186. doi: 10.1038/srep03186
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Szekeres, S., Dauti, M., Wilde, C., Mazel, D., and Rowe-Magnus, D. A. (2007). Chromosomal toxin-antitoxin loci can diminish large-scale genome reductions in the absence of selection. Mol. Microbiol. 63, 1588–1605. doi: 10.1111/j.1365-2958.2007.05613.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Takagi, H., Kakuta, Y., Okada, T., Yao, M., Tanaka, I., and Kimura, M. (2005). Crystal structure of archaeal toxin-antitoxin RelE-RelB complex with implications for toxin activity and antitoxin effects. Nat. Struct. Mol. Biol. 12, 327–331. doi: 10.1038/nsmb911
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Tian, Q.-B., Ohnishi, M., Murata, T., Nakayama, K., Terawaki, Y., and Hayashi, T. (2001). Specific protein-DNA and protein-protein interaction in the hig gene system, a plasmid-borne proteic killer gene system of plasmid Rts1. Plasmid 45, 63–74. doi: 10.1006/plas.2000.1506
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Utsumi, R., Nakamoto, Y., Kawamukai, M., Himeno, M., and Komano, T. (1982). Involvement of cyclic AMP and its receptor protein in filamentation of an Escherichia coli fic mutant. J. of Bacteriol. 151, 807–812.
Van Melderen, L. (2002). Molecular interactions of the CcdB poison with its bacterial target, the DNA gyrase. Int. J. Med. Microbiol. 291, 537–544. doi: 10.1078/1438-4221-00164
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Woolery, A. R., Luong, P., Broberg, C. A., and Orth, K. (2012). AMPylation: something old is new again. Front Microbiol. 1:113. doi: 10.3389/fmicb.2010.00113
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Yamaguchi, Y., and Inouye, M. (2009). mRNA interferases, sequence-specific endoribonucleases from the toxin-antitoxin systems. Prog. Mol. Biol. Transl. Sci. 85, 467–500. doi: 10.1016/S0079-6603(08)00812-X
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Zhang, Y., and Inouye, M. (2009). The inhibitory mechanism of protein synthesis by YoeB, an Escherichia coli toxin. J. Biol. Chem. 284, 6627–6638. doi: 10.1074/jbc.M808779200
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Zhou, Y., Liang, Y., Lynch, K. H., Dennis, J. J., and Wishart, D. S. (2011). PHAST: a fast phage search tool. Nucleic Acids Res. 39, W347–W352. doi: 10.1093/nar/gkr485
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Keywords: Streptococcus pneumoniae, pneumococcal toxin-antitoxin loci, Phd-Doc, Green Fluorescent Protein, self-regulation
Citation: Chan WT, Yeo CC, Sadowy E and Espinosa M (2014) Functional validation of putative toxin-antitoxin genes from the Gram-positive pathogen Streptococcus pneumoniae: phd-doc is the fourth bona-fide operon. Front. Microbiol. 5:677. doi: 10.3389/fmicb.2014.00677
Received: 03 September 2014; Paper pending published: 12 November 2014;
Accepted: 19 November 2014; Published online: 05 December 2014.
Edited by:
John R. Battista, Louisiana State University, USAReviewed by:
Tatiana Venkova-Canova, The University of Texas Medical Branch at Galveston, USARamon Diaz Orejas, Consejo Superior de Investigaciones Científicas, Spain
Copyright © 2014 Chan, Yeo, Sadowy and Espinosa. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Wai Ting Chan, Centro de Investigaciones Biológicas, Consejo Superior de Investigaciones Científicas, Ramiro de Maeztu 9, 28040 Madrid, Spain e-mail:Y2hhbnl0aW5nQGhvdG1haWwuY29t
 Wai Ting Chan
Wai Ting Chan Chew Chieng Yeo
Chew Chieng Yeo Ewa Sadowy
Ewa Sadowy Manuel Espinosa
Manuel Espinosa