- Section of Eukaryotic Biotechnology, Department of Systems Biology, Technical University of Denmark, Kongens Lyngby, Denmark
Aspergillus, Penicillium, and Talaromyces are among the most chemically inventive of all fungi, producing a wide array of secondary metabolites (exometabolites). The three genera are holophyletic in a cladistic sense and polythetic classes in an anagenetic or functional sense, and contain 344, 354, and 88 species, respectively. New developments in classification, cladification, and nomenclature have meant that the species, series, and sections suggested are natural groups that share many extrolites, including exometabolites, exoproteins, exocarbohydrates, and exolipids in addition to morphological features. The number of exometabolites reported from these species is very large, and genome sequencing projects have shown that a large number of additional exometabolites may be expressed, given the right conditions (“cryptic” gene clusters for exometabolites). The exometabolites are biosynthesized via shikimic acid, tricarboxylic acid cycle members, nucleotides, carbohydrates or as polyketides, non-ribosomal peptides, terpenes, or mixtures of those. The gene clusters coding for these compounds contain genes for the biosynthetic building blocks, the linking of these building blocks, tailoring enzymes, resistance for own products, and exporters. Species within a series or section in Aspergillus, Penicillium, and Talaromyces have many exometabolites in common, seemingly acquired by cladogenesis, but some the gene clusters for autapomorphic exometabolites may have been acquired by horizontal gene transfer. Despite genome sequencing efforts, and the many breakthroughs these will give, it is obvious that epigenetic factors play a large role in evolution and function of chemodiversity, and better methods for characterizing the epigenome are needed. Most of the individual species of the three genera produce a consistent and characteristic profile of exometabolites, but growth medium variations, stimulation by exometabolites from other species, and variations in abiotic intrinsic and extrinsic environmental factors such as pH, temperature, redox potential, and water activity will add significantly to the number of biosynthetic families expressed in anyone species. An example of the shared exometabolites in a natural group such as Aspergillus section Circumdati series Circumdati is that most, but not all species produce penicillic acids, aspyrones, neoaspergillic acids, xanthomegnins, melleins, aspergamides, circumdatins, and ochratoxins, in different combinations.
Introduction
The genera Aspergillus sensu lato and Penicillium sensu lato contain a high number of very diverse species. These species produce a large number of exometabolites, also known as secondary metabolites. Exometabolites are small molecules produced during morphological and chemical differentiation that are outward directed, i.e., secreted or deposited in or on the cell wall, and accumulated in contrast to endometabolites (primary metabolites), that are fluctuating in concentration (the fluxome), and either transformed into other endometabolites or feeding into exometabolites, exoproteins, exopolysaccharides, and morphological structures. While endometabolites can be found in almost all species of fungi (and most other kinds of organisms), exometabolites, exoproteins, and exopolysaccharides are taxonomically restricted, being produced in species-specific profiles. Some metabolites can occur both as endo- and exometabolites, for example citric acid. When citric acid is part of the mitochondrial fluxome, it should be regarded as an endometabolite, but when citric acid is secreted and accumulated (Goldberg et al., 2006; Andersen et al., 2011; Poulsen et al., 2012), as in Aspergillus niger, it must be regarded as an exometabolite. Accumulation of citric acid requires that there is a reductive pathway for it in the cytosol and that it can be secreted to the surroundings via an exporter. Thus the transport from the mitochondria to the cytosol, the cytosolic reduction, and the secretion requires a dedicated gene cluster. Such a gene cluster has been found in for example A. terreus that is coding for accumulating and secreting itaconic acid (van der Straat et al., 2014), but the gene cluster for citric acid accumulation has not been described yet. Some species related to Aspergillus and Penicillium, such as Xeromyces bisporus, are predominantly stress-selected (S-selected) and the lack of any competitors at very low water activities will have the consequence that X. bisporus produces no exometabolites (Leong et al., 2014). In Aspergillus most species produce a large number of exometabolites, but some stress selected species, such as A. penicillioides and A. restrictus, have only been reported to produce asperglaucide and cristatin A, and the related arestrictin A and B (Itabashi et al., 2006). However, the closely related xerotolerant/xerophilic species in the Aspergillus subgenus Aspergillus (formerly Eurotium) produce a high number of exometabolites in the ascomata, making them chemically very diverse (Slack et al., 2009).
Aspergillus and Penicillium Taxonomy and Nomenclature
Because of their importance, species of Aspergillus and Penicillium have been taxonomically treated several times, but the monographs by Raper and Thom (1949) for Penicillium and Raper and Fennell (1965) for Aspergillus are still regarded as cornerstones in the taxonomy of these fungi. In the period between these two monographs, however, several authors (Benjamin, 1955; Malloch and Cain, 1971) suggested to use names for the sexual state of the Aspergilli and Penicillia, whenever possible, to adhere to the Botanical Code in a nomenclatural sense. The use of Penicillium and Aspergillus for species that had not yet been found to produce a sexual state could keep their Penicillium and Aspergillus names, because of a special nomenclatural “exception” in the Botanical Code (Art. 59) that allowed to use two names for a specific fungal species, one for the asexual states (the “anamorph”) and one for the sexual state (the “teleomorph”). Despite this it was recommended to use the sexual name for the whole fungus (the “holomorph”), whenever a sexual state had been found. For this reason many species in Penicillium were renamed Eupenicillium or Talaromyces (Pitt, 1980) and many Aspergillus species were renamed Chaetosartorya, Emericella, Eurotium, Fennellia, Hemicarpenteles, Hemisartorya, Neocarpenteles, Neopetromyces, Neosartorya, Petromyces, Saitoa, Sclerocleista, or Warcupiella (Rai and Chowdhery, 1975; Rajendran and Muthappa, 1980; Gams and Samson, 1986). To give one example of the name changes one can mention a fungus that was originally described as A. fischeri. Since this fungus was described including the sexual state, it could not be used for the asexual state anymore, so the correct name for the fungus according the nomenclatural rules before 2011 was Neosartorya fischeri, while A. fischeri had to be renamed A. fischerianus if one wanted only to refer to the asexual state. A full monographic revision of Aspergillus according to the Botanical Code has not been written, but lists of accepted Aspergillus and Penicillium species have been made (Pitt et al., 2000) and several revisions of the individual genera have been published.
In 2010 it was suggested to introduce a new nomenclatural system in which one fungus had only one name (Hawksworth et al., 2011; Hawksworth, 2012). This suggestion was adopted by the Botanical Congress in Melbourne (McNeill et al., 2012), and thus hereafter any species in fungi will only have one official name. The selection of those names is encouraged to take place by consensus among international experts in the group of fungi under consideration. The International Commission of Penicillium and Aspergillus (ICPA) has decided to use Penicillium for the monophyletic clade that includes Penicillium subgenera Aspergilloides, Furcatum, and Penicillium sensu Pitt (1980), Eupenicillium, Chromocleista, Thysanophora and Eladia, and Talaromyces for the monophyletic clade that includes Talaromyces itself and Penicillium subgenus Biverticillium sensu Pitt (1980). For Aspergillus sensu lato, a cladistic study using DNA sequence data, showed that most known Aspergillus species were included in a monophyletic clade (Houbraken and Samson, 2011), while a few rare species, such as A. zonatus and A. clavatoflavus were more closely related to other genera in the Eurotiomycetes. The nomenclatural consequence of this is to call all the species in the monophyletic clade Aspergillus (Houbraken et al., 2014; Samson et al., 2014) or retypify the genus Aspergillus with for example A. niger, and then subdivide Aspergillus into the genera Aspergillus, Neosartorya, Emericella, Eurotium, and Chaetosartorya (Pitt and Taylor, 2014). This would have the consequence that the name Aspergillus would only be used for a paraphyletic weakly supported clade representing subgenus Aspergillus and that the genus Neosartorya would be polyphyletic as it includes Dichotomomyces. Even though a majority of ICPA members voted for the Aspergillus solution, which includes mentioning the sexual state informally, for example A. fischeri (neosartorya-morph present), general consensus has not yet been reached. In this review Aspergillus names will be used, as suggested by Samson et al. (2014), as the name Aspergillus can be confidently used for the monophyletic clade that includes the genera listed above (Houbraken et al., 2014). All species formerly included in Dichotomomyces, Cristaspora, Phialosimplex, Polypaecilum, in addition to Penicillium inflatum, have been formally combined into Aspergillus (Samson et al., 2014), while A. crystallinus, A. malororatus, and A. paradoxus (Hemicarpenteles paradoxus) have been combined into Penicillium, as P. crystallinum, P. malodoratum, and P. paradoxum (Visagie et al., 2014b). This means that the presence of aspergilla in an isolate does not necessarily mean that the isolate belongs to Aspergillus sensu stricto, and the presence of penicilli in an isolates does not necessarily mean the species belong in Penicillium sensu stricto. However, in the majority of cases aspergilla or penicilli indicates that the species belong to Aspergillus and Penicillium, respectively.
Pitt and Taylor (2014) suggested to use Aspergillus for the paraphyletic subgenus Circumdati only (after potential re-typification of Aspergillus with A. niger), stating that this restricted use of the genus Aspergillus would make this genus phenotypically different from the closely related Aspergillus subgenus Nidulantes and therefore suggested the name Emericella for the latter monophyletic clade. However, there are many phenotypic traits in common between section Circumdati and Nidulantes, including the presence of hülle cells and the exometabolites kojic acid, aflatoxins, and sterigmatocystins in both subgenera (Raper and Fennell, 1965; Wiley and Simmons, 1973; Frisvad and Samson, 2004a; Frisvad et al., 2005; Zalar et al., 2008). Subgenus Circumdati includes species with both multiple cleistothecia in sclerotia (Petromyces, Neopetromyces, Saitoa; Udagawa et al., 1994; Yaguchi et al., 1994; Frisvad and Samson, 2000; Horn et al., 2013) and pseudoparenchymatous multiple ascomata in hyphal masses with or without hülle cells (Fennellia and the perfect state of A. terreus; Wiley and Simmons, 1973; Locquin-Linard, 1990; Yaguchi et al., 1994; Samson et al., 2011a; Arabatsis and Velegraki, 2013), while several species in Aspergillus section Nidulantes produce pseudoparenchymatous single ascomata and hülle cells.
Assistance in choosing between Aspergillus (Samson et al., 2014) versus the genera Eurotium, Emericella, Neosarotrya and Chaetosartorya, Phialosimplex, Polypaecilum, Dichotomomyces, and Cristaspora (Pitt and Taylor, 2014) can be sought from scientific databases. It is very clear that while Aspergillus has been used in 56178 publications the other genera, when all added, have only been used in 1093 publications (approximately 2%; Table 1).
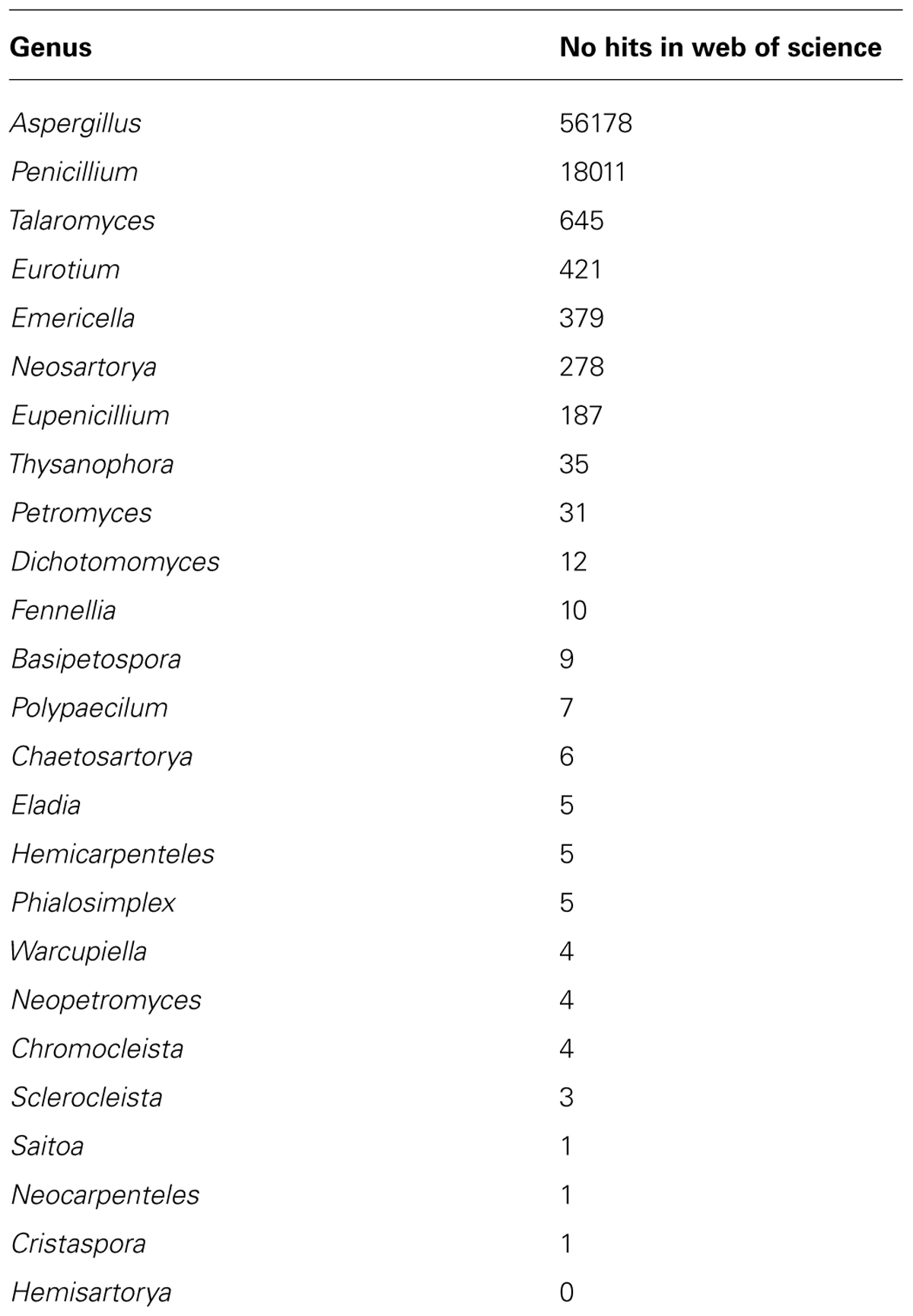

TABLE 1. References to Aspergillus, Penicillium, Talaromyces, and associated genera (Web of Science, as of 18 October, 2014).
A classification of isolates into sections and series in Penicillium, Talaromyces, and Aspergillus based on phenotypic characters will show that these supraspecific taxa are natural polythetic classes (Beckner, 1959) in exometabolite, ecophysiological, and morphological characters. For example in Aspergillus subgenus Circumdati section Circumdati (the former A. ochraceus group) most species, but not all, produce aspyrones, penicillic acids, xanthomegnins, ochratoxins, melleins, circumdatins, neoaspergillic acids, and aspergamides/stephacidins (Frisvad et al., 2004a,b; Finefield et al., 2012; Visagie et al., 2014a). In addition individual species produces exometabolites that are only accumulated by few species in the section. A. westerdijkiae and A. ochraceus can both produce all the exometabolites listed above, but in addition A. westerdijkiae produces preussin and mellamide, not produced by A. ochraceus.
At present Aspergillus comprises 344 species (Samson et al., 2014), Penicillium 354 species (Visagie et al., 2014b), and Talaromyces 88 species (Yilmaz et al., 2014). These genera include species that have been reported to produce large numbers of exometabolites (Table 2).
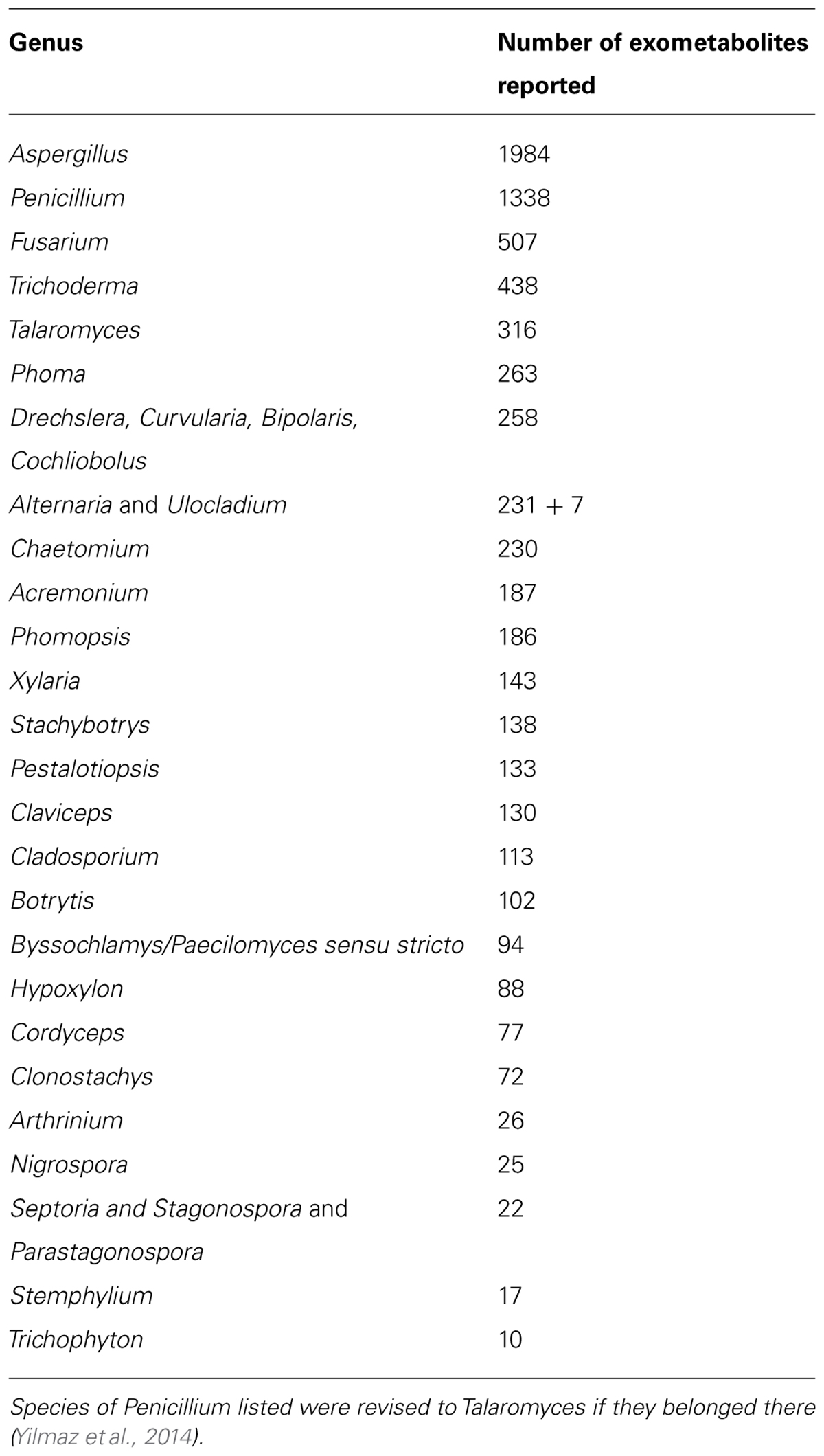
TABLE 2. Individual exometabolites produced by important genera of filamentous fungi ranked according to highest number of exometabolites reported (according to AntiBase).
Chemodiversity
Species of Aspergillus, Penicillium, and Talaromyces are extraordinarily productive concerning exometabolites. A comparison with other genera shows that most exometabolites have been reported from Aspergillus (1984), next-most from Penicillium (1338), and fifth-most by Talaromyces, (316), with only Fusarium (507) and Trichoderma (438) producing more exometabolites in toto (Table 1). The number of exometabolites pr species is 5.77 for Aspergillus, 3.77 for Penicillium, and 3.58 for Talaromyces. These number per species are clearly underestimates as some exometabolites are produced by more than one species in a genus, in addition to the fact that many species have not been examined and that some exometabolites are only expressed under unique circumstances and thus may remain undetected (Sanchez et al., 2012; Brakhage, 2013; Scherlach et al., 2013; Takahashi et al., 2013; Bertrand et al., 2014; Marmann et al., 2014). Light, pH, redox potential, temperature, water activity, carbon sources, nitrogen sources, iron starvation, and exometabolites from other species can all have a regulatory effect on the regulatory proteins for exometabolite expression in a fungus (Brakhage, 2013). A majority of the exometabolites produced by Penicillium and Aspergillus are only found sporadically in other genera, but a large number of exometabolites are in common between Aspergillus and Penicillium. On the other hand exometabolites from Talaromyces are nearly all unique to that genus (Samson et al., 2011b), or only shared with few other species.
The same exometabolite may be produced by widely different species. For example aflatoxin is produced by the species listed in Aspergillus section Flavi (15 spp.), Aspergillus section Nidulantes (3 spp.), Aspergillus section Ochraceorosei (2 spp.), and Aschersonia (2 spp.; Frisvad and Samson, 2004a; Frisvad et al., 2005; Zalar et al., 2008; Varga et al., 2009, 2011; Kornsakulkarn et al., 2012, 2013; Massi et al., 2014). Three species in Aspergillus section Flavi and all the seven species outside section Flavi listed above only produce aflatoxins of the B type. It is surprising that aflatoxins have never been found in Penicillium, but they have been found in the unrelated scale insect fungi Aschersonia coffea and Aschersonia marginata (Kornsakulkarn et al., 2012, 2013). However, the precursor sterigmatocystin, although end-product for some species, has been found in a large number of unrelated genera (Rank et al., 2011), suggesting that this complicated gene cluster has been horizontally transfered between species in widely different genera, as shown by Slot and Rokas (2011) for Podospora anserina and A. nidulans. Fungal species are specifically associated to certain habitats or few plant, animal, or other kind of organisms (Filtenborg et al., 1996), and will therefore produce exometabolites in reponse to the challenges in the particular habitat. For example P. herquei was thought to be a soil fungus saprophyte (Kwaśna, 2004), but recent studies have shown tha the leaf-rolling weevil (Euops chinensis) have developed mycangia to inoculates leaves with P. herquei conidia to protect the weevil eggs (Li et al., 2012). P. herquei produce a species specific profile of exometabolites, of which several are antibiotically active (Petit et al., 2009; Tansakul et al., 2014). Thus the specificity in both association of fungal species to other species and the profile of exometabolites are factors that have boosted the evolution of so many exometabolites.
Dichotomomyces cejpii was transferred to A. cejpii by Samson et al. (2014), and this new combination is supported by chemotaxonomic evidence. D. cejpii was reported to produce gliotoxin, xanthocillin X monomethylether, tryptoquivalones, JBIR-03, emindole SB, emindole SB beta-mannoside, and 27-O-methylasporyzin (Varga et al., 2007; Harms et al., 2014). While gliotoxin, tryptoquivalones, and xanthocillins (Frisvad et al., 2009; Zuck et al., 2011) indicates a relationship to A. fumigatus and tryptoquivalones a close relationship to A. clavatus, as supported by DNA sequences (Varga et al., 2007), production of emindole SB indicates a relationship to Aspergillus section Nidulantes. The report of emindole SB, emeniveol, asporyzin A-C, and JBIR-03 from a marine-derived A. oryzae (Qiao et al., 2010), indicates what they identified as “A. oryzae” is a fungus related to A. cejpii or a species in Nidulantes rather than A. oryzae, however.
Chemoconsistency and OSMAC
The abbreviation OSMAC (one strain many compounds) was introduced by Bode et al. (2002) where the authors showed, among several examples, that a strain of A. westerdijkiae produced a series of exometabolites that could be ordered into different biosynthetic families. Furthermore, by using several media a more full profile of these exometabolites could be revealed. The idea that one strain can produce several exometabolites was already introduced by Frisvad (1981) and Frisvad and Filtenborg (1983, 1989). These authors showed that terverticillate penicillia produced a unique profile of different exometabolites and also that certain media, such as Czapek yeast autolysate (CYA) agar and yeast extract sucrose (YES) agar were very efficient for production of a large number of different exometabolites, while further media may increase the number of exometabolites expressed (Bills et al., 2008; Nielsen et al., 2011; Frisvad, 2012). Furthermore they showed that these profiles of exometabolites were species specific and consistent from isolate to isolate, i.e., the isolates in anyone fungal species were chemo consistent (Larsen et al., 2005). One of the original terms for exometabolites or secondary metabolites was idiolites, the latter indicating that production of exometabolites was strain specific, however, exometabolite profiles are clearly species specific (Frisvad et al., 2008). However, a single mutation in a gene in an exometabolite gene-cluster will often be sufficient for loss of phenotypic expression (Susca et al., 2014), and this may be the reason some authors call the production of certain exometabolites “strain-specific” (i.e., Engel et al., 1982). The ability to produce mycophenolic acid in P. roqueforti is retained in most strains, however, (Frisvad and Filtenborg, 1989; Geisen et al., 2001; Frisvad and Samson, 2004b), but is not as consistent as in P. brevicompactum, where a non-producing strain has never been found (Frisvad and Filtenborg, 1989; Frisvad and Samson, 2004b). Reasons for observing “unusual “ or “unexpected” exometabolites in a species may be horizontal gene transfer of a gene cluster in only one or few strains, hybridization (which is not common in filamentous fungi), or epigenetic priming. Raper and Thom (1949) mentioned a strain of P. citrinum (NRRL 822, their group III in a subdivision of P. citrinum “transitional toward P. chrysogenum series”) produced both citrinin, known from this species, and penicillin, known from P. chrysogenum. They also mentioned that their strain had the cultural appearance of both P. citrinum and P. chrysogenum. We have re-examined this strain, and indeed it had characters of both species, and appeared to be a (rare) hybrid. However, exometabolites from co-occurring species from the same habitat may stimulate the epigenome by acting as inhibitors of histone acetylation or methylation, and this exometabolite stimulation will be one of many ways of having silent exometabolite gene clusters in filamentous fungi expressed (Bertrand et al., 2014). It was recently shown that A. niger could produce sclerotia with many hitherto not expressed aflavinins in them (Frisvad et al., 2014) simply by stimulating A. niger with whole fruits or rice. Whether this stimulation is caused by extrolites from those whole fruits or rice or from a physical stimulation is not yet known. Furthermore variations in the growth medium and ecophysiological factors such as pH, temperature, and water activity will obviously also stimulate expression of gene clusters of exometabolites that were initially thought to be silent.
In conclusion Aspergillus, Penicillium, and Talaromyces contain species that produce a very large number of species-specific exometabolites with a high degree of chemoconsistency. The chemodiversity of the many species in these three genera is extremely high and many more bioactive compounds from the species will be found in the future. Both ecological and genetic/molecular approaches are needed to fully explore this treasure-trove of natural products.
Conflict of Interest Statement
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Andersen, M. R., Salazar, M. P., Schaap, P. J., van de Vondervoort, P. J. I., Culley, D., Thykaer, J.,et al. (2011). Comparative genomics of citric-acid producing Aspergillus niger ATCC 1015 versus enzyme-producing CBS 513.88. Genome Res. 21, 885–897. doi: 10.1101/gr.112169.110
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Arabatsis, M., and Velegraki, A. (2013). Sexual reproduction cycle in the opportunistic human pathogen Aspergillus terreus. Mycologia 105, 71–79. doi: 10.3852/11-426
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Benjamin, C. R. (1955). Ascocarps of Aspergillus and Penicillium. Mycologia 47, 669–687. doi: 10.2307/3755578
Bertrand, S., Bohni, N., Schnee, S., and Schumpp, O. (2014). Metabolite induction via microorganism co-culture: a potential way to enhance chemical diversity for drug discovery. Biotechnol. Adv. 32, 1180–1204. doi: 10.1016/j.biotechadv.2014.03.001
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Bills, G. F., Platas, G., Fillola, A., Jiménez, M. R., Collado, J., Vicente, F.,et al. (2008). Enhancement of antibiotic and secondary metabolite detection from filamentous fungi by growth on nutritional arrays. J. Appl. Microbiol. 104, 1644–1658. doi: 10.1111/j.1365-2672.2008.03735.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Bode, H. B., Bether, B., Hofs, K., and Zeeck, A. (2002). Big effects from small changes: possible ways to explore nature’s chemical diversity. Chembiochem 3, 619–627. doi: 10.1002/1439-7633
Brakhage, A. A. (2013). Regulation of fungal secondary metabolism. Nat. Rev. Microbiol. 11, 21–32. doi: 10.1038/nrmicro2916
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Engel, G., von Milczewski, K. E., Prokopek, D., and Teuber, M. (1982). Strain-specific synthesis of mycophenolic acid by Penicillium roqueforti in blue-veined cheeses. Appl. Environ. Microbiol. 43, 1034–1040.
Filtenborg, O., Frisvad, J. C., and Thrane, U. (1996). Moulds in food spoilage. Int. J. Food Microbiol. 33, 85–102. doi: 10.1016/0168-1605(96)01153-1
Finefield, J. M., Frisvad, J. C., Sherman, D. H., and Williams, R. M. (2012). Fungal origins of the bicyclo[2.2.2]diazaoctane ring system of prenylated indol alkaloids. J. Nat. Prod. 75, 812–833. doi: 10.1021/np200954v
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Frisvad, J. C. (1981). Physiological criteria and mycotoxin production as aids in identification of common asymmetric penicillia. Appl. Environ. Microbiol. 41, 568–579.
Frisvad, J. C. (2012). “Media and growth conditions for induction of secondary metabolites,” in Fungal Secondary Metabolism: Methods and Protocols, eds N. P. Keller and G. Turner (New York: Humana Press), 944, 47–58. doi: 10.1007/978-1-62703-122-6_3
Frisvad, J. C., Andersen, B., and Thrane, U. (2008). The use of secondary metabolite profiling in fungal taxonomy. Mycol. Res. 112, 231–240. doi: 10.1016/j.mycres.2007.08.018
Frisvad, J. C., and Filtenborg, O. (1983). Classification of terverticillate penicillia based on profiles of mycotoxins and other secondary metabolites. Appl. Environ. Microbiol. 46, 1301–1310.
Frisvad, J. C., and Filtenborg, O. (1989). Terverticillate penicillia: chemotaxonomy and mycotoxin production. Mycologia 81, 836–861. doi: 10.2307/3760103
Frisvad, J. C., Frank, J. M., Houbraken, J. A. M. P., Kuijpers, A. F. A., and Samson, R. A. (2004a). New ochratoxin producing species of Aspergillus section Circumdati. Stud. Mycol. 50, 23–43.
Frisvad, J. C., Smedsgaard, J., Larsen, T. O., and Samson, R. A. (2004b). Mycotoxins, drugs and other extrolites produced by species in Penicillium subgenus Penicillium. Stud. Mycol. 49, 201–241.
Frisvad, J. C., Petersen, L. M., Lyhne, E. K., and Larsen, T. O. (2014). Formation of sclerotia and production of indoloterpenes by Aspergillus niger and other species in section Nigri. PLoS ONE 9:e94857. doi: 10.13717/journal.pone.0094857
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Frisvad, J. C., Rank, C., Nielsen, K. F., and Larsen, T. O. (2009). Metabolomics of Aspergillus fumigatus. Med. Mycol. 47, S53–S71. doi: 10.1080/13693780802307720
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Frisvad, J. C., and Samson, R. A. (2000). Neopetromyces gen. nov. and an overview of teleomorphs of Aspergillus subgenus Circumdati. Stud. Mycol. 45, 201–207.
Frisvad, J. C., and Samson, R. A. (2004a). Emericella venezuelensis, a new species with stellate ascospores producing sterigmatocystin and aflatoxin B1. Syst. Appl. Microbiol. 27, 672–680. doi: 10.1078/0723202042369910
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Frisvad, J. C., and Samson, R. A. (2004b). Polyphasic taxonomy of Penicillium subgenus Penicillium. A guide to identification of the food and air-borne terverticillate penicillia and their mycotoxins. Stud. Mycol. 49, 1–173.
Frisvad, J. C., Skouboe, P., and Samson, R. A. (2005). Taxonomic comparison of three different groups of aflatoxin producers and a new efficient producer of aflatoxin B1, sterigmatocystin and 3-O-methylsterigmatocystin, Aspergillus rambellii sp. nov. Syst. Appl. Microbiol. 28, 442–453. doi: 10.1016/j.syapm.2005.02.012
Gams, W., and Samson, R. A. (1986). “Typification of Aspergillus and related teleomorph genera,” in Advances in Penicillium and Aspergillus Systematics, eds R. A. Samson and J. I. Pitt (New York: Plenum Press), 23–29. doi: 10.1007/978-1-4757-1856-0_3
Geisen, R., Cantor, M. D., Hansen, T. K., Holzapfel, W. H., and Jakobsen, M. (2001). Characterization of Penicillium roqueforti strains used as cheese starter cultures by RAPD typing. Int. J. Food Microbiol. 65, 181–191. doi: 10.1016/S0168-1605(00)00514-6
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Goldberg, I., Rokem, J. S., and Pines, O. (2006). Organic acids: old metabolites, new themes. J. Chem. Technol. Biotechnol. 81, 1601–1611. doi: 10.1002/jctb.1590
Harms, H., Rempel, V., Kehraus, S., Kaiser, M., Hufendick, P., Müller, C. E.,et al. (2014). Indoloterpenes from a marine-derived fungal strain of Dichotomomyces cejpii with an antagonistic activity at GPR18 and cannabinoid receptors. J. Nat. Prod. 77, 673–677. doi: 10.1021/np400850g
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Hawksworth, D. L. (2012). Managing and coping with names of pleomorphic fungi in a period of transition. IMA Fungus 3, 15–24. doi: 10.5598/imafungus.2012.03.01.03
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Hawksworth, D. L., Crous, P. W., Redhead, S. A., Reynolds, D. R., Samson, R. A., Seifert, K. A.,et al. (2011). The Amsterdam declaration on fungal nomenclature. IMA Fungus 2, 105–112. doi: 10.5598/imafungus.2011.02.01.14
Horn, B. W., Olarte, R. A., Peterson, S. W., and Carbone, I. (2013). Sexual reproduction in Aspergillus tubingensis from section Nigri. Mycologia 105, 1153–1163. doi: 10.3852/13-101
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Houbraken, J., de Vries, R. P., and Samson, R. A. (2014). Modern taxonomy of biotechnologically important Aspergillus and Penicillium species. Adv. Appl. Microbiol. 86, 199–249. doi: 10.1016/B978-0-12-800262-9.00004-4
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Houbraken, J., and Samson, R. A. (2011). Phylogeny of Pencillium and the segregation of Trichocomaceae into three families. Stud. Mycol. 70, 1–51. doi: 10.3114/sim.2011.70.01
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Itabashi, T., Matsuishi, N., Hosoe, T., Toyasaki, N., Usagawa, S., Imai, T.,et al. (2006). Two new dioxopiperazine derivatives, arestrictina A and B, isolated from Aspergillus restrictus and Aspergillus penicilloides. Chem. Pharm. Bull. 54, 1639–1641. doi: 10.1248/cpb.54.1639
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Kornsakulkarn, J., Saepua, S., Laksanacharoen, P., Rachtawee, P., and Thongpanchang, C. (2013). Xanthone and anthraquinone-type mycotoxins from the scale insect fungus Aschersonia marginata BCC 28721. Tetrahedron Lett. 54, 3813–3815. doi: 10.1016/j.tetlet.2013.05.036
Kornsakulkarn, J., Saepua, S., Srichomthong, K., Supothina, S., and Thongpanchang, C. (2012). New mycotoxins from the scale insect fungus Aschersonia coffeae Henn. BCC 28712. Tetrahedron 68, 8480–8486. doi: 10.1016/j.tet.2012.07.059
Kwaśna, H. (2004). Natural shifts in communities of rhizosphere fungi of common oak after felling. Plant Soil 264, 209–218. doi: 10.1023/B:PLSO.0000047752.41575.c7
Larsen, T. O., Smedsgaard, J., Nielsen, K. F., Hansen, M. E., and Frisvad, J. C. (2005). Phenotypic taxonomy and metabolite profiling in microbial drug discovery. Nat. Prod. Rep. 22, 672–695. doi: 10.1039/b404943h
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Leong, S. L., Lantz, H., Petterson, O. V., Frisvad, J. C., Thrane, U., Heipieper, H. J.,et al. (2014). Genome and physiology of the ascomycete filamentous fungus Xeromyces bisporus, the most xerophilic organism isolated to date. Environ. Microbiol. doi: 10.1111/1462-2920.12596 [Epub ahead of print].
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Li, X., Guo, W., and Ding, J. (2012). Mycangial fungus benefits the development of a leaf-rolling weevil, Euops chinensis. J. Insect Physiol. 58, 867–873. doi: 10.1016/j.jinsphys.2012.03011
Locquin-Linard, M. (1990). Fennellia monodii, nouvelle espèce d’ascomycete coprophile de zones aride Africaines. Mycotaxon 39, 9–15.
Malloch, D., and Cain, R. F. (1971). New species and combinations of cleistothecial Ascomycetes. Can. J. Bot. 50, 61–72. doi: 10.1139/b72-011
Marmann, A., Aly, A. H., Lin, W., Wang, B., and Proksch, P. (2014). Co-cultivation – a powerful emerging tool for enhancing the chemical diversity of microorganisms. Mar. Drugs 12, 1043–1062. doi: 10.3390/md12021043 d
Massi, F. P., Vieira, M. L. C., Sartori, D., Penha, R. E. S., Munhoz, C. D. F., Ferreira, J. M.,et al. (2014). Brazil nuts are subject to infection with B and G aflatoxin-producing fungus, Aspergillus pseudonomius. Int. J. Food Microbiol. 186, 14–21. doi: 10.1016/j.ijfoodmicro.2014.06.006
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
McNeill, J., Barrie, F. R., Buck, W. R., Demoulin, V., Greuter, W., Hawksworth, D. L.,et al. (2012). International Code of Nomenclature for Algae, Fungi and Plants (Melbourne Code) Adopted by the 18th International Botanical Congress, Melbourne, 2011. Koenigstein: Koeltz Scientific Books.
Nielsen, M. L., Nielsen, J. B., Rank, C., Klejnstrup, M. L., Holm, D. M. K., Brogaard, K. H.,et al. (2011). A genome-wide polyketide synthase deletion library uncovers novel genetic links to polyketides and meroterpenoids in Aspergillus nidulans. FEMS Microbiol. Lett. 321, 157–166. doi: 10.1111/j.1574-6968.2011.02327.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Petit, P., Lucas, E. M. F., Abreau, L. M., Pfennig, L. H., and Takahashi, J. A. (2009). Novel antimicrobial secondary metabolites from a Penicillium sp. isolated from Brazilian cerrado soil. Electronic J. Biotechnol. 12:9. doi: 10.2225/vol12-issue4-fultext-9
Pitt, J. I. (1980). The Genus Penicillium and its Teleomorphic State Eupenicillium and Talaromyces. London: Academic Press.
Pitt, J. I., Samson, R. A., and Frisvad, J. C. (2000). “List of accepted species and their synonyms in the family Trichomaceae,” in Integration of Modern Taxonomic Methods for Pencillium and Aspergillus Classification, eds R. A. Samson and J. I. Pitt (Amsterdam: Harwood Academic Publishers), 9–49.
Pitt, J. I., and Taylor, J. W. (2014). Aspergillus, its sexual states and the new international code of nomenclature. Mycologia 106, 1051–1062. doi: 10.3852/14-060
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Poulsen, L., Andersen, M. R., Lanz, A. E., and Thykaer, J. (2012). Identification of a transcription factor controlling pH-dependent organic acid response in Aspergillus niger. PLoS ONE 7:e50596. doi: 10.1371/journal.pone.0050596
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Qiao, M.-F., Ji, N.-Y., Liu, X.-H., Li, K., Zu, Q.-M., and Xue, Q.-Z. (2010). Indoloterpenes from an algicolous isolate of Aspergillus oryzae. Bioorg. Med. Chem. Lett. 20, 5677–5680. doi: 10.1016/j.bmcp.2010.08-024
Rai, J. N., and Chowdhery, H. J. (1975). Hemisartorya, a new genus of cleistothecial Ascomycetes with Aspergillus state. Kavaka 3, 73–76.
Rajendran, C., and Muthappa, B. N. (1980). Saitoa, a new genus of Plectomycetes. Proc. Indian Acad. Sci. 89, 185–191.
Rank, C., Nielsen, K. F., Larsen, T. O., Varga, J., Samson, R. A., and Frisvad, J. C. (2011). Distribution of sterigmatocystin in filamentous fungi. Fungal Biol. 115, 406–420. doi: 10.1016/j.funbio.2011.02.013
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Raper, K. B., and Fennell, D. I. (1965). The Genus Aspergillus. Baltimore, MD: Williams and Wilkins.
Samson, R. A., Peterson, S. W., Frisvad, J. C., and Varga, J. (2011a). New species in Aspergillus section Terrei. Stud. Mycol. 69, 39–55. doi: 10.3114/sim.2011.69.04
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Samson, R. A., Yilmaz, N., Houbraken, J., Spierenburg, H., Seifert, K. A., Peterson, S. W.,et al. (2011b). Phylogeny and nomenclature of the genus Talaromyces and taxa accommodated in Penicillium subgenus Biverticillium. Stud. Mycol. 70, 159–184. doi: 10.3114/sim.2011.70.04
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Samson, R. A., Visagie, C. M., Houbraken, J., Hong, S.-B., Hubka, V., Klaasen, H. W.,et al. (2014). Phylogeny, identification and nomenclature of the genus Aspergillus. Stud. Mycol. 78, 141–173. doi: 10.1016/j.simyco.2014.07.004
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Sanchez, J. F., Somoza, A. D., Keller, N. P., and Wang, C. C. (2012). Advances in Aspergillus secondary metabolite research in the post-genomic era. Nat. Prod. Rep. 29, 351–371. doi: 10.1039/c2np00084a
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Scherlach, K., Graupner, K., and Hertweck, C. (2013). Molecular bacteria-fungi interactions: effects on environment, food and medicine. Annu. Rev. Microbiol. 67, 375–397. doi: 10.1146/annurev-micro-092412-155702
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Slack, G., Puniani, E., Frisvad, J. C., Samson, R. A., and Miller, J. D. (2009). Secondary metabolites from Eurotium species, A. calidoustus and A. insuetus common in Canadian homes with a review of their chemistry and biological activities. Mycol. Res. 113, 480–490. doi: 10.1016/j.mycres.2008.12.002
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Slot, J. C., and Rokas, A. (2011). Horizontal transfer of a large and highly toxic secondary metabolic gene cluster between fungi. Curr. Biol. 21, 134–139. doi: 10.1016/j.cub.2010.12.020
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Susca, A., Proctor, R. H., Butchko, R. A. E., Haidukowski, M., Stea, G., Logrieco, A.,et al. (2014). Variation in the fumonisin biosynthetic gene cluster in fumonisin–producing and nonproducing black aspergilli. Fungal Genet. Biol. 73, 39–52. doi: 10.1016/j.fgb.2014.09.009
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Takahashi, J. A., Teles, A. P. C., Bracarense, A. D. A. P., and Gomes, D. C. (2013). Classical and epigenetic approaches to metabolite diversification in filamerntous fungi. Phytochem. Rev. 12, 773–789. doi: 10.1007/s11101-013-9305-5
Tansakul, C., Rukachaisirikul, V., Maha, A., Kongpragan, T., Phongpaichit, S., Hutadilok-Totwatana, N.,et al. (2014). A new phenalenone derivative from the soil fungus Penicillium herquei PSU-RSPG93. Nat. Prod. Res. 28, 1718–1724. doi: 10.1080/14786419.2014.941363
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Udagawa, S., Uchiyama, S., and Kamiya, S. (1994). Petromyces muricatus, a new species with an Aspergillus anamorph. Mycotaxon 52, 207–214.
van der Straat, L., Vernooij, M., Lammers, M., vand en Berg, W., Schoneville, T., Cordewener, J.,et al. (2014). Expression of the Aspergillus terreus itaconic acid biosynthesis gene cluster in Aspergillus niger. Microb. Cell Fact. 13:11. doi: 10.1186/1475-2859-13-11
Varga, J., Due, M., Frisvad, J. C., and Samson, R. A. (2007). Taxonomic revision of Aspergillus section Clavati based on molecular, morphological and physiological data. Stud. Mycol. 59, 89–106. doi: 10.3114/sim.2007.59.11
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Varga, J., Frisvad, J. C., and Samson, R. A. (2009). A reappraisal of fungi producing aflatoxin. World Mycotoxin J. 2, 263–277. doi: 10.3920/WMJ2008.1094
Varga, J., Frisvad, J. C., and Samson, R. A. (2011). Two new aflatoxin producing species, and an overview of Aspergillus section Flavi. Stud. Mycol. 69, 57–80. doi: 10.3114/sim.2011.69.05
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Visagie, C., Varga, J., Houbraken, J., Meijer, M., Yilmaz, N., Seifert, K. A.,et al. (2014a). Ochratoxin production and taxonomy of the yellow aspergilli (Aspergillus section Circumdati). Stud. Mycol. 78, 1–61. doi: 10.1016/j.simyco.2014.07.001
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Visagie, C. M., Houbraken, J., Frisvad, J. C., Hong, S.-B., Klaassen, C. H. W., Perrone, G.,et al. (2014b). Identification and nomenclature of the genus Penicillium. Stud. Mycol. 78, 343–371. doi: 10.1016/j.simyco.2014.09.001
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Wiley, B. J., and Simmons, E. G. (1973). New species and a new genus of Plectomycetes with Aspergillus states. Mycologia 65, 934–938. doi: 10.2307/3758529
Yaguchi, T., Someya, A., and Udagawa, S. (1994). Fennellia flavipes and Neosartorya stramenia, two new records from Japan. Mycoscience 35, 175–178. doi: 10.1007/BF02318496
Yilmaz, N., Visagie, C. M., Houbraken, J., Frisvad, J. C., and Samson, R. A. (2014). Polyphasic taxonomy of the genus Talaromyces. Stud. Mycol. 78, 175–341. doi: 10.1016/j.simyco.2014.08.001
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Zalar, P., Frisvad, J. C., Gunde-Cimerman, N., Varga, J., and Samson, R. A. (2008). Four new species of Emericella from the Mediterranean region of Europe. Mycologia 100, 779–795. doi: 10.3852/08-078
Zuck, K. M., Shipley, S., and Newman, D. J. (2011). Induced production of N-formyl alkaloids from Aspergillus fumigatus by co-culture with Streptomyces peucetius. J. Nat. Prod. 74, 1653–1657. doi: 10.1021/np200255f
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Keywords: Aspergillus, Penicillium, Talaromyces, secondary metabolites, chemodiversity, chemoconsistency
Citation: Frisvad JC (2015) Taxonomy, chemodiversity, and chemoconsistency of Aspergillus, Penicillium, and Talaromyces species. Front. Microbiol. 5:773. doi: 10.3389/fmicb.2014.00773
Received: 09 November 2014; Paper pending published: 05 December 2014;
Accepted: 17 December 2014; Published online: 12 January 2015.
Edited by:
Jonathan Palmer, United States Department of Agriculture Forest Service, USAReviewed by:
Antonello Santini, University of Naples Federico II, ItalyRob Samson, CBS Fungal Biodiversity Centre, Netherlands
Copyright © 2015 Frisvad. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jens C. Frisvad, Section for Eukaryotic Biotechnology, Department of Systems Biology, Technical University of Denmark, Søltofts Plads B. 221, 2800 Kongens Lyngby, Denmark e-mail:amNmQGJpby5kdHUuZGs=
 Jens C. Frisvad
Jens C. Frisvad