- 1Department of Sanitary Inspection, School of Public Health, University of South China, Hengyang, China
- 2State Key Laboratory of Pathogen and Biosecurity, Beijing Institute of Microbiology and Epidemiology, Beijing, China
- 3Animal Husbandry Base Teaching and Research Section, College of Animal Science and Technology, Hebei North University, Zhangjiakou, China
Yersinia pestis, the causative agent of plague, poses a serious health threat to rodents and human beings. TyrR is a transcriptional regulator (TyrR) that controls the metabolism of aromatic amino acids in Escherichia coli. In this paper, TyrR played an important role in Y. pestis virulence. Inactivation of tyrR did not seem to affect the in vitro growth of this organism, but resulted in at least 10,000-fold attenuation compared with the wild-type (WT) strain upon subcutaneous infection to mice. In addition, loads of tyrR mutant within mice livers and spleens significantly decreased compared with the WT strain. Transcriptome analysis revealed that TyrR, directly or indirectly, regulated 29 genes encoded on Y. pestis chromosome or plasmids under in vitro growth condition. Similar to the regulatory function of this protein in E. coli, five aromatic-pathway genes (aroF-tyrA, aroP, aroL, and tyrP) were significantly reduced upon deletion of the tyrR gene. Two genes (glnL and glnG) that encode sensory histidine kinase and regulator in a two-component regulatory system involved in nitrogen assimilation were downregulated in the tyrR mutant. Several genes encoding type III secretion proteins were transcribed by 2.0–4.2-fold in a tyrR mutant relative to the WT strain. Interestingly, the acid-stressed genes, hdeB and hdeD, were downregulated, and such downregulation partly accounted for the decrease in tolerance of the tyrR mutant under acidic conditions. In conclusion, regulation of TyrR in Y. pestis is similar to, but distinct from, that in E. coli. TyrR is a metabolic virulence determinant in Y. pestis that is important for extracellular survival and/or proliferation.
Introduction
Yersinia pestis can cause fatal infections in rodents and humans and is usually transmitted by flea biting (Perry and Fetherston, 1997). A few virulence determinants have been defined based on their contributions to flea transmission colonization, invasion, intracellular growth or extracellular proliferation (Charnetzky and Shuford, 1985; Lindler et al., 1990; Galyov et al., 1991; Lahteenmaki et al., 1998). Numerous efforts to determine the mechanism of Yersinia pathogenesis are mainly focused on a few previously established virulence determinants (Perry and Fetherston, 1997). Many other genes encoding two-component systems or global transcriptional regulators have been proven to be implicated in the regulatory networks involved in Y. pestis pathogenicity (Oyston et al., 2000; Cathelyn et al., 2006; Zhan et al., 2008; Geng et al., 2009). More than 30 genes responsible for critical metabolic pathways functioned during Y. pestis fitness in vivo (Palace et al., 2014). Y. pestis lacking ~47-kb DNA fragment that contains more than 40 genes was obtained by our laboratory. Surprisingly, we observed the significantly attenuated virulence of this mutant via subcutaneous infection in mice. This observation prompted us to investigate which gene or operon was responsible for such attenuation. After careful tracing via gene knockout within the 47-kb region, a transcriptional regulator (TyrR) was confirmed to be mainly responsible for the virulence phenotype. A number of DNA-binding regulators could control gene expression common to many bacterial species. TyrR is responsible for aromatic amino acids metabolism in Escherichia coli, which is evolutionarily acquired and is only present in γ-proteobacteria (Panina et al., 2001; Song et al., 2005). Subsequent evidence showed that the recruitment of structure genes by TyrR is dynamic and is evolving further (Pittard et al., 2005). Here we showed that TyrR is required for Y. pestis pathogenesis and extracellular survival/proliferation. Transcriptome analysis demonstrated that the regulation of TyrR in Y. pestis is similar to, but distinct from, that in E. coli.
Experimental Procedure
Bacterial Strains
Y. pestis wild-type (WT) strain 201, 47-kb fragment and tyrR deletion mutant were used in this study. Strain 201 was isolated from Microtus brandti in Inner Mongolia, China. Strain 201 is supposed to be avirulent to humans, but highly lethal to mice (Fan et al., 1995). The deletion mutant of Y. pestis was constructed by replacing the entire target gene with the kan cassette by using λ-Red homologos recombination. To obtain a strain in which ΔtyrR is complemented, plasmid pACYC184, which contains a PCR fragment covering a region from 400 bp fragment upstream to 100 bp downstream of the tyrR gene, was introduced into Y. pestis ΔtyrR.
Determination of Bacterial Growth Curves in vitro
Y. pestis strains were grown in LB medium at 26°C to exponential phase (OD620 ≈ 1.0). The bacterial cultures were diluted 1:20 in LB medium with the indicated pH value and incubated at 26°C. Bacterial growth was monitored by measuring absorbance at OD620. The experiments were performed in three independent cultures. Results were expressed as the mean percentage ± standard deviation from the three independent experiments.
Mouse Infection
Y. pestis strains were grown in brain-heart infusion (BHI) broth at 26°C to OD620 ≈ 1.0. Bacterial cells were harvested by centrifugation, washed twice and resuspended in phosphate-buffered saline (PBS). Groups of five or six female BALB/c mice (6-weeks-old) were injected subcutaneously with serial dilutions of bacterial cultures. Mortality was recorded daily for 14 d. The 50% lethal dose (LD50) was calculated by the Reed-Muench equation (Reed and Muench, 1938).
To monitor bacterial growth dynamics in vivo, Y. pestis WT strain 201 and the tyrR deletion mutant grown to exponential phase were washed and diluted to 600 CFU/mL in PBS. A 1:1 mixture (100 μL) of the two bacterial strains was used to infect eight BALB/c mice intravenously. For competitive index (CI) determination, the infected mice were sacrificed at 48 and 72 h postinfection, and livers and spleens were removed for homogenization. Bacterial loads per organ were determined by calculating the number of viable bacteira of the resulting homogenates plated on Hottinger agar with and without kanamycin. The CI value was calculated as the ratio of the number of mutant/WT bacteria recovered. The data were analyzed by Student's t-test, with P < 0.05 considered statistically significant. All mouse experiments were carried out according to the Guidelines for the Welfare and Ethics of Laboratory Animals of Beijing.
Transcription Analysis by Using RNA-Seq and Real-Time PCR
Y. pestis WT strain 201 and the ΔtyrR mutant were grown at 26°C in BHI medium to middle exponential phase and then transferred to 37°C for 3 h. Total RNA was extracted using the TRIzol Reagent according to the manufacturer's protocol (Invitrogen). These two RNA samples were subjected to cDNA library construction and deep sequencing performed by LC Sciences LLC, USA, as previously described (Yan et al., 2013). RPKM was used to estimate the expression levels of mRNA transcripts encoded by Y. pestis chromosome and plasmids. Those genes with cDNA coverage below 20 in both samples were removed from further analysis. The fold change of mRNA expression levels between WT and tyrR mutant were calculated. Two-fold was used as threshold to determine the differentially regulated genes.
For quantitative RT-PCR, cDNA was generated using 5 μg of total RNA and 3 μg of random hexamer primers with the Superscript II system. All primer pairs produced a 150–200 nt amplicon when Y. pestis genomic DNA was used as the template for PCR. Real-time PCR was performed in duplicate for each RNA preparation using the LightCycler system (Roche) with an appropriate dilution of cDNA as a template. Based on the standard curve of 16S rRNA, the relative mRNA level was determined by calculating the threshold cycle (ΔCt) of each gene by the classic ΔCt method. Quantification of 16S rRNA was also used to normalize the values of all the other genes in the RT-PCR experiment.
Measurement of Secreted LcrV Protein by Using Western Blotting
Y. pestis WT strain, the ΔtyrR mutant and the tyrR complementary strain were grown at 26°C in TMH medium without calcium to OD620 = 1.0 and then transferred to 37°C for 3 h. Secreted proteins from bacterial supernatants were precipitated with trichloroacetic acid (TCA). Proteins from equal amount of bacteria were separated on SDS–PAGE and immunoblotted with LcrV polyclonal antibody followed by detection using the Odyssey Infrared Imaging System.
Results
TyrR is Required for Y. pestis Infection Upon Subcutaneous Inoculation to Mice
We observed the strong attenuation of a deletion mutant of ~47-kb DNA fragment containing more than 40 genes in BALB/c mice upon subcutaneous inoculation. To trace which gene(s) or operon is responsible for the attenuated phenotype, DNA fragments within the 47-kb region were knockout one by one from the WT strain. The resulting mutants were subject to mice infection, respectively. Finally, a ~8-kb fragment containing tyrR and its flanking sequences were confirmed to be the main reason for the virulence phenotype. Therefore, we decided to assess the roles of TyrR protein in Y. pestis pathogenesis. Mice were infected subcutaneously (s.c.) with increasing numbers of WT, tyrR mutant, and complementary strain to estimate the virulence by calculating LD50. The LD50 of both the WT strain 201 and the tyrR complementary strain was, <8 CFU, but up to about 8 × 104 cells of the tyrR mutant was not lethal to s.c. inoculated mice (Figure 1). The observation suggested that the significantly attenuated virulence of the tyrR mutant is due to the lack of TyrR protein rather than to polar effects caused by the insertion of a kanamycin resistance cassette.
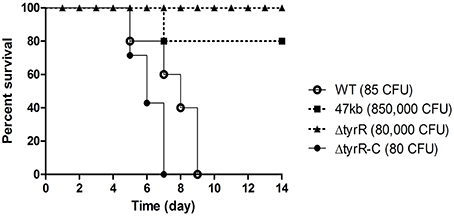
Figure 1. Effect of the tyrR gene on Y. pestis pathogenesis in mice. Groups of five or seven BALB/c mice were inoculated by subcutaneous injection with appropriated dose of Y. pestis strain 201 (open circles), 47-kb mutant (filled diamonds), tyrR (filled triangles) or tyrR complementary strain (filled circles). Mice mortality was monitored for 14 d.
TyrR Contributes to Growth Dynamics in vivo
Growth of the tyrR mutant was not retarded relative to that of WT strain under the condition used in vitro (Figure 2), indicating that the mutations do not cause any defect in the growth ability. Therefore, the differences in virulence could be due to the specific involvement of this protein under in vivo conditions. We next determine the kinetics of in vivo growth to examine the fitness of Y. pestis WT and tyrR mutant. CI assays were performed by intravenously inoculating the bacterial mixture into mice. The results showed that load burden of WT strain can achieved 107 CFU in the spleen or liver after infection for 48 and 72 h. However, compared with the WT strain, a relatively lower amount bacteria of the tyrR mutant strain was recovered from the spleen or liver. The mean CI values in the spleens were 0.043 at 48 h and 0.003 at 72 h (Figure 3A), and similar CI values were obtained in the livers (0.017 at 48 h and 0.005 at 72 h) (Figure 3B). Clearly, the tyrR mutant was nearly overwhelmed by the population of WT Y. pestis in vivo, indicating that this mutant was much less competitive in vivo than its parental strain.
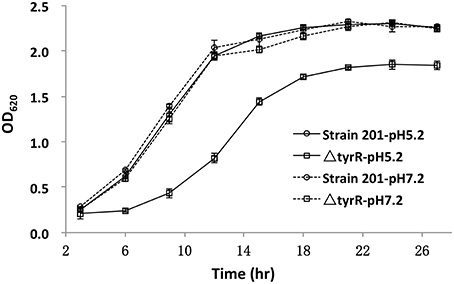
Figure 2. Growth curves of the WT and tyrR mutant strains under two different conditions. Overnight cultures of the WT strain 201 and tyrR mutant were used to inoculate fresh LB broth with pH 5.2 or 7.0. The OD620 values were recorded at fixed time points. Each time point is the average of two measurements and error bars represent standard deviations. The vertical arrows indicate the time point at which samples were removed for RNA extraction.
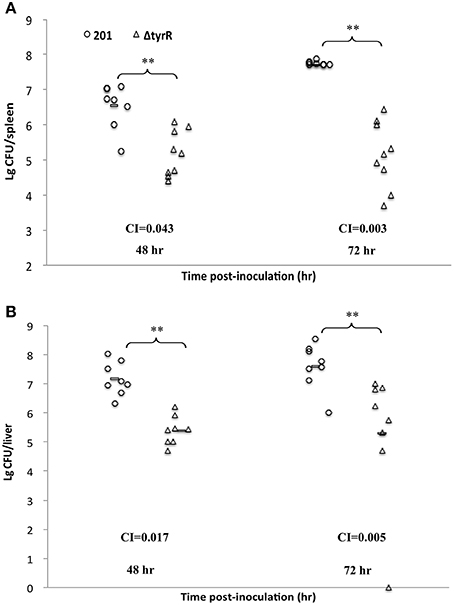
Figure 3. Growth dynamics of Y. pestis upon inoculation by intraveneous route. Groups of eight mice were inoculated intravenously with bacterial mixture of Y. pestis strain 201 and the tyrR mutant. Bacterial loads of WT strain (open circles) and tyrR mutant (open triangles) in spleen (A) and liver (B) were determined at 48 h and 72 h post-inoculation. Horizontal bars indicate geometric means. The indicated P values were determined using the Student's t-test.
Identification of Differential Genes Regulated by TyrR
To obtain a representative image of TyrR protein in affecting gene expression throughout growth, we recovered RNA from cultures in vitro at different time points to determine the optimal time points. The abundance of the tyrR transcript was measured by qPCR (data not shown). The bacterial culture grown in BHI at 26°C for 9 h and then transferred to 37°C for 3 h was chosen for RNA recovery for RNA sequencing.
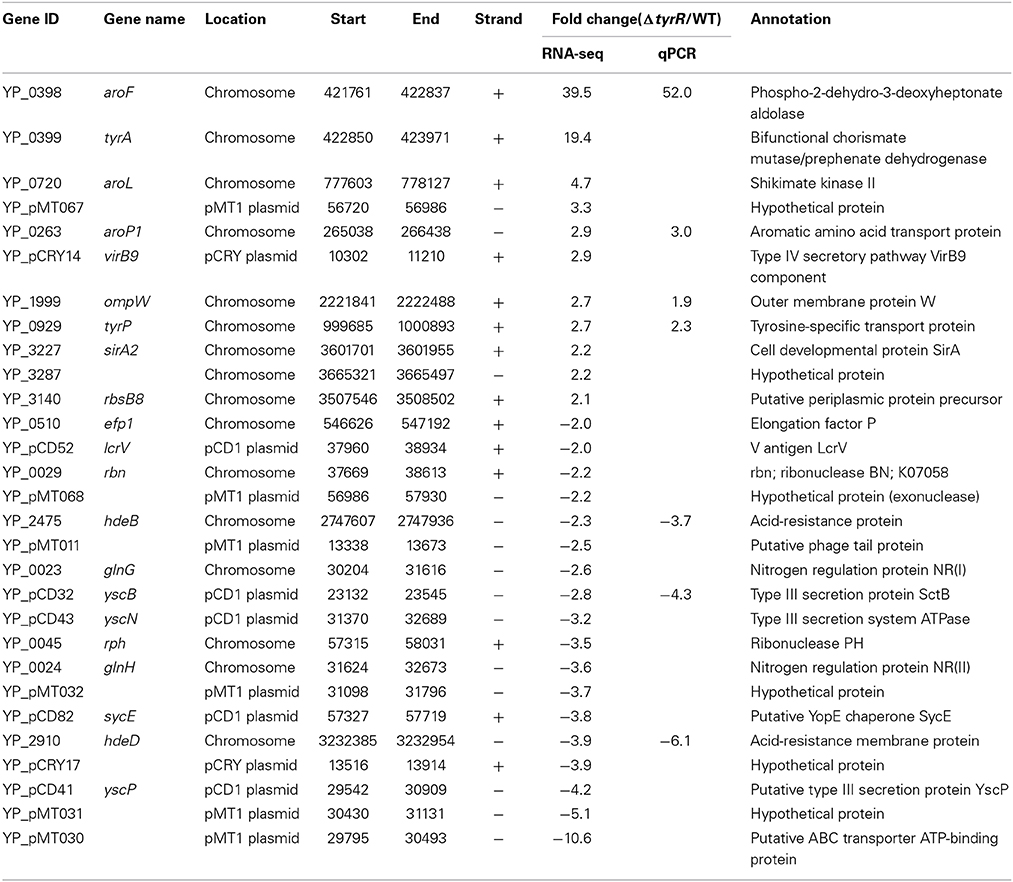
In total, 29 genes were shown differentially expressed between WT and tyrR mutant strain (Table 1). Of these genes, 11 were upregulated and 18 were downregulated in tyrR mutant compared with the WT strain. Intriguingly, approximately 45% (13/29) of the differentially regulated genes were derived from three plasmids (pCD1, pMT1, and pCRY), thereby suggesting that the laterally acquired genetic elements might be recruited to TyrR regulon during the process of evolution. Of which, five genes encoding type III secretion proteins in plasmid pCD1 (yscB, yscN, yscP, sycE, and lcrV) were downregulated 2.0–4.2- fold in tyrR mutant relative to WT strain.
Eight genes were selected for qPCR analysis to validate the RNA-seq data. A high correlation (R2 = 0.988) was observed between expression values obtained by RNA-seq and qPCR (Figure 4). Aromatic-pathway genes were confirmed to be regulated by TyrR in E. coli (Pittard et al., 2005). As expected, the aroF-tyrA operon responsible for aromatic biosynthesis and aroL, aroP and tyrP for aromatic transport were most strongly upregulated. Two genes responsible for acid stress, hdeB and hdeD, were confirmed to be downregulated upon deletion of tyrR gene, and this finding was consistent with the observation on the compromised tolerance to acid stress in in vitro assays (Figure 1).
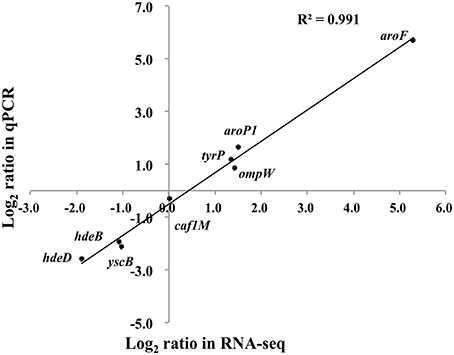
Figure 4. Transcription measurements of eight genes were chosen for quantitative RT-PCR validation. The real-time PCR log2 values were plotted against the RNA-seq data log2 values. The coefficient of determination (R2) for comparison of the two datasets is 0.991.
Use of Conserved TyrR Motif to Identify Direct TyrR -Regulated Genes
DNA-binding regulator usually regulates its target genes by directly binding to a dyad DNA consensus sequence. To predict which genes identified by RNA-seq might be under the direct control of TyrR, we extracted the putative TyrR-binding motif sequence (TGTAAA-N6-TTTACA) from RegulonDB and conducted a research in the upstream 300 nt of the differentially regulated genes identified by RNA-seq experiment. Four genes were identified as candidates for direct TyrR regulation. As expected, conserved DNA motifs were found in the promoter region of three known aromatic-pathway transcripts (aroF, tyrP, and aroP1). Another candidate targeted by TyrR is the hdeD gene, which encodes an acid stress membrane protein, because TyrR box was found ~140 nt upstream of the translational start site of hdeD.
Synthesis of Secreted V Antigen not Influenced by TyrR in Y. pestis
Y. pestis LcrV is secreted via T3SS machinery during infection and can be exploited as a protective antigen known as V antigen (Skrzypek and Straley, 1995). Abrogation of LcrV expression render Y. pestis avirulent (Burrows, 1956). Since LcrV protein is responsible for extracellular survival and dissemination in Y. pestis, the down-regulation of lcrV might lead to the reduced capability of organ colonization. Y. pestis V antigen is maximally expressed and secreted under calcium-restricted conditions at 37°C in vitro. Therefore, we compared the levels of secreted V antigen synthesis by using Western blot method in WT and tyrR mutant grown in calcium-deficiency TMH medium at 37°C. Unexpectedly, the results showed that the expression of V antigen was not obviously changed upon the deletion of tyrR (Supplementary Figure 1). TyrR might have failed to affect LcrV expression at least in Y. pestis grown in vitro.
Discussion
Aromatic amino acid biosynthesis is required for the intracellular replication of Listeria monocytogenes and Shigella flexneri (Cersini et al., 1998; Stritzker et al., 2004). Coincidently, two aromatic-pathway genes (aroE and aroA) play a crucial role in Y. pestis fitness in deep tissues during infection (Palace et al., 2014). Y. pestis lacking tyrR was significantly less virulent than the WT strain, and complementation of tyrR restored WT virulence in the deletion strain, thereby suggesting that TyrR plays an important role in Y. pestis virulence. The regulation of aromatic amino acid metabolism has been assigned to the transcription factor, TyrR. This is the first report to demonstrate the effect of this protein on Y. pestis pathogenesis. TyrR is not required for growth in vitro, but is required for full virulence of Y. pestis. Whether the availability of aromatic amino acids in vivo might have an effect during the process of infection unknown. Intriguingly, we observed that tyrR mutant efficiently protects mice against Y. pestis infection as an attenuated strain (data unpublished). The potential of TyrR as a new therapeutic target is worth exploring.
When Y. pestis enters invasion sites, most of the bacteria are engulfed and killed by the polymorphonuclear leukocytes (PMNs) that are attracted to these invasion sites. However, a few bacilli survive and proliferate within phagolysosomes of tissue macrophages during the initial infection stage and are then released and to elicit the systematic infection (Perry and Fetherston, 1997). The growth defect of Y. pestis tyrR mutant in BHI with pH 5.2 (Figure 1) was a hint that TyrR might be associated with intracellular replication, because acidic pH is a hypothesized prevailing condition in phagolysosomal microenvironments.
Based on the results of RNA-seq-based transcriptional profiling, less than 1% of Y. pestis genes were affected by TyrR, thereby indicating that this protein is likely to function as a local regulator. However, it was unexpected that a regulatory defect in a minority of non-essential genes has as much effect on pathogenesis in vivo as the removal of a single critical virulence factor. We supposed that the regulation of T3SS encoded by pCD1 plasmid might partially account for the virulence variations between WT and tyrR-deletion strains. However, the pCD1 plasmid-encoded genes are downregulated ranging from 2.0 to 4.2 fold, which are generally much lower than that for the aromatic amino acid metabolism genes (2.7 ~ 39.5 fold). In addition, no predicted TyrR-binding sites exist in the regulated genes of T3SS, indicating perhaps TyrR weakly affects transcription of these genes due to lacking exact DNA binding motifs on these plasmids. Thus, we speculated that regulation of T3SS by TyrR is likely to be indirect.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This study was funded by the National Natural Science Foundation of China (31171248) and the National Basic Research Program of China (2014CB744405).
Supplementary Material
The Supplementary Material for this article can be found online at: http://www.frontiersin.org/journal/10.3389/fmicb.2015.00110/abstract
References
Burrows, T. W. (1956). An antigen determining virulence in Pasteurella pestis. Nature 177, 426–7. doi: 10.1038/177426b0
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Cathelyn, J. S., Crosby, S. D., Lathem, W. W., Goldman, W. E., and Miller, V. L. (2006). RovA, a global regulator of Yersinia pestis, specifically required for bubonic plague. Proc. Natl. Acad. Sci. U.S.A. 103, 13514–13519. doi: 10.1073/pnas.0603456103
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Cersini, A., Salvia, A. M., and Bernardini, M. L. (1998). Intracellular multiplication and virulence of Shigella flexneri auxotrophic mutants. Infect. Immun. 66, 549–557.
Charnetzky, W. T., and Shuford, W. W. (1985). Survival and growth of Yersinia pestis within macrophages and an effect of the loss of the 47-megadalton plasmid on growth in macrophages. Infect. Immun. 47, 234–241.
Fan, Z., Luo, Y., Wang, S., Jin, L., Zhou, X., Liu, J., et al. (1995). Microtus brandti plague in the Xilin Gol Grassland was inoffensive to humans. Chin. J. Control Endem. Dis. 10, 56–7.
Galyov, E. E., Karlishev, A. V., Chernovskaya, T. V., Dolgikh, D. A., Smirnov, O., Volkovoy, K. I., et al. (1991). Expression of the envelope antigen F1 of Yersinia pestis is mediated by the product of caf1M gene having homology with the chaperone protein PapD of Escherichia coli. FEBS Lett. 286, 79–82. doi: 10.1016/0014-5793(91)80945-Y
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Geng, J., Song, Y., Yang, L., Feng, Y., Qiu, Y., Li, G., et al. (2009). Involvement of the post-transcriptional regulator Hfq in Yersinia pestis virulence. PLoS ONE 4:e6213. doi: 10.1371/journal.pone.0006213
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Lahteenmaki, K., Virkola, R., Saren, A., Emody, L., and Korhonen, T. K. (1998). Expression of plasminogen activator pla of Yersinia pestis enhances bacterial attachment to the mammalian extracellular matrix. Infect. Immun. 66, 5755–5762.
Lindler, L. E., Klempner, M. S., and Straley, S. C. (1990). Yersinia pestis pH 6 antigen: genetic, biochemical, and virulence characterization of a protein involved in the pathogenesis of bubonic plague. Infect. Immun. 58, 2569–2577.
Oyston, P. C., Dorrell, N., Williams, K., Li, S. R., Green, M., Titball, R. W., et al. (2000). The response regulator PhoP is important for survival under conditions of macrophage-induced stress and virulence in Yersinia pestis. Infect. Immun. 68, 3419–3425. doi: 10.1128/IAI.68.6.3419-3425.2000
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Palace, S. G., Proulx, M. K., Lu, S., Baker, R. E., and Goguen, J. D. (2014). Genome-wide mutant fitness profiling identifies nutritional requirements for optimal growth of Yersinia pestis in deep tissue. Am. Soc. Microbiol. 5:e01385-14. doi: 10.1128/mBio.01385-14
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Panina, E. M., Vitreschak, A. G., Mironov, A. A., and Gelfand, M. S. (2001). Regulation of aromatic amino acid biosynthesis in gamma-proteobacteria. J. Mol. Microbiol. Biotechnol. 3, 529–543.
Perry, R. D., and Fetherston, J. D. (1997). Yersinia pestis–etiologic agent of plague. Clin. Microbiol. Rev. 10, 35–66.
Pittard, J., Camakaris, H., and Yang, J. (2005). The TyrR regulon. Mol. Microbiol. 55, 16–26. doi: 10.1111/j.1365-2958.2004.04385.x
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Reed, L. J., and Muench, H. (1938). A simple method for estimating fifty percent endpoints. Am. J. Hygiene. 27, 493–497.
Skrzypek, E., and Straley, S. C. (1995). Differential effects of deletions in lcrV on secretion of V antigen, regulation of the low-Ca2+ response, and virulence of Yersinia pestis. J. Bacteriol. 177, 2530–2542.
Song, J., Bonner, C. A., Wolinsky, M., and Jensen, R. A. (2005). The TyrA family of aromatic-pathway dehydrogenases in phylogenetic context. BMC Biol. 3:13. doi: 10.1186/1741-7007-3-13
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Stritzker, J., Janda, J., Schoen, C., Taupp, M., Pilgrim, S., Gentschev, I., et al. (2004). Growth, virulence, and immunogenicity of Listeria monocytogenes aro mutants. Infect. Immun. 72, 5622–5629. doi: 10.1128/IAI.72.10.5622-5629.2004
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Yan, Y., Su, S., Meng, X., Ji, X., Qu, Y., Liu, Z., et al. (2013). Determination of sRNA Expressions by RNA-seq in Yersinia pestis Grown In Vitro and during Infection. PLoS ONE 8:e74495. doi: 10.1371/journal.pone.0074495
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Zhan, L., Han, Y., Yang, L., Geng, J., Li, Y., Gao, H., et al. (2008). The cyclic AMP receptor protein, CRP, is required for both virulence and expression of the minimal CRP regulon in Yersinia pestis biovar microtus. Infect. Immun. 76, 5028–5037. doi: 10.1128/IAI.00370-08
Pubmed Abstract | Pubmed Full Text | CrossRef Full Text | Google Scholar
Keywords: Yersinia pestis, TyrR, aromatic amino acid metabolism, pathogenesis
Citation: Deng Z, Liu Z, He J, Wang J, Yan Y, Wang X, Cui Y, Bi Y, Du Z, Song Y, Yang R and Han Y (2015) TyrR, the regulator of aromatic amino acid metabolism, is required for mice infection of Yersinia pestis. Front. Microbiol. 6:110. doi: 10.3389/fmicb.2015.00110
Received: 17 December 2014; Accepted: 29 January 2015;
Published online: 12 February 2015.
Edited by:
Yi-Cheng Sun, Chinese Academy of Medical Sciences and Peking Union Medical College, ChinaReviewed by:
Yan Ling, Beijing Institute of Biotechnology, ChinaChristopher Francis Bosio, National Institutes of Health, USA
Copyright © 2015 Deng, Liu, He, Wang, Yan, Wang, Cui, Bi, Du, Song, Yang and Han. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ruifu Yang and Yanping Han, No. 20 Dongdajie, Fengtai, Beijing 100071, China e-mail:cnVpZnV5YW5nQGdtYWlsLmNvbQ==;eWFucGluZ2hhbkBnbWFpbC5jb20=
†These authors have contributed equally to this work.
 Zhongliang Deng
Zhongliang Deng Zizhong Liu
Zizhong Liu Junming He
Junming He Jing Wang2,3
Jing Wang2,3 Yanfeng Yan
Yanfeng Yan Xiaoyi Wang
Xiaoyi Wang Yujun Cui
Yujun Cui Yujing Bi
Yujing Bi Zongmin Du
Zongmin Du Yajun Song
Yajun Song Ruifu Yang
Ruifu Yang Yanping Han
Yanping Han