- Institute of Microbiology and College of Life Sciences, Zhejiang University, Hangzhou, China
Hydrogen sulfide (H2S) has been recognized as a physiological mediator with a variety of functions across all domains of life. In this study, mechanisms of endogenous H2S generation in Shewanella oneidensis were investigated. As a research model with highly diverse anaerobic respiratory pathways, the microorganism is able to produce H2S by respiring on a variety of sulfur-containing compounds with SirACD and PsrABC enzymatic complexes, as well as through cysteine degradation with three enzymes, MdeA, SO_1095, and SseA. We showed that the SirACD and PsrABC complexes, which are predominantly, if not exclusively, responsible for H2S generation via respiration of sulfur species, do not interplay with each other. Strikingly, a screen for regulators controlling endogenous H2S generation by transposon mutagenesis identified global regulator Crp to be essential to all H2S-generating processes. In contrast, Fnr and Arc, two other global regulators that have a role in respiration, are dispensable in regulating H2S generation via respiration of sulfur species. Interestingly, Arc is involved in the H2S generation through cysteine degradation by repressing expression of the mdeA gene. We further showed that expression of the sirA and psrABC operons is subjected to direct regulation of Crp, but the mechanisms underlying the requirement of Crp for H2S generation through cysteine degradation remain elusive.
Introduction
Hydrogen sulfide (H2S), traditionally recognized as a noxious gas, is now regarded as an important signaling molecule along with nitric oxide (NO) and carbon monoxide (CO), associated with beneficial functions in mammals from vasorelaxation, cardioprotection, and neurotransmission to anti-inflammation (Wang, 2012). In bacteria, investigations into its physiological roles just began. It has been recently elucidated that H2S confers bacterial cells defense against antibiotics by stimulating the cellular protection system against reactive oxygen species (ROS) (Shatalin et al., 2011). Beyond this, our understanding of the mechanism by which H2S affects biology is limited. In both eukaryotes and prokaryotes, H2S can be produced endogenously via assimilatory sulfate reduction and cysteine degradation. H2S of a very small amount formed via the former is rapidly assimilated into organic sulfur compounds such as sulfur-containing amino acids and is hardly released into extracellular environments. On the contrary, the latter, catalyzed by cystathionine β-synthase (CBS), cystathionine γ-lyase (CSE), and 3-mercaptopyruvate sulfurtransferase (3MST) together with cysteine aminotransferase (CAT), is responsible for H2S generation on a large scale, ensuring that the molecule can be functionally implicated in many physiological processes (Shatalin et al., 2011; Kimura, 2014). Most, if not all, of bacterial genomes encode some of these enzymes. For example, Bacillus anthracis, Pseudomonas aeruginosa, and Staphylococcus aureus have the CBS/CSE operon but not 3MST, whereas Escherichia coli carries 3MST, but not CBS/CSE (Shatalin et al., 2011).
Some prokaryotes have an additional route for H2S generation on a large scale, the dissimilatory sulfate/sulfite (SO2−4/SO2−3) reduction, which is best illustrated in sulfate-reducing bacteria (SRB) Desulfovibrio vulgaris (Bradley et al., 2011). The dissimilatory sulfite reductases, featuring siroheme and [4Fe-4S] prosthetic centers, are minimally composed of two subunits (DsrA and DsrB) in an α2β2 arrangement and catalyze the six-electron reduction of sulfite to sulfide (Oliveira et al., 2008). In addition, two other enzyme complexes, represented by PsrABC of Salmonella enterica Serovar Typhimurium and SirACD of Shewanella oneidensis, have been identified to be able to reduce various inorganic sulfur species to H2S (Heinzinger et al., 1995; Shirodkar et al., 2011). The former is able to use as electron acceptors (EAs) thiosulfate (S2O2−3), tetrathionate (S4O2−6), and elemental sulfur (S0) for respiration while the latter, which shares similarities with the formate-dependent nitrite reductase, catalyzes direct conversion of sulfite to sulfide. PsrC and SirD serve as quinol oxidases that transfer electrons stepwise via PsrB and SirC to the catalytic subunit PsrA and SirA for reduction of corresponding EAs, respectively (Jormakka et al., 2008; Cordova et al., 2011).
S.oneidensis, a facultative Gram-negative γ-proteobacterium, has been intensively studied owing to its exceptional metabolic flexibility and its potential use for the bioremediation of metal/radionuclide contaminants in the environment (Fredrickson et al., 2008). Although the microorganism neither belongs to SRB phylogenetically nor possesses analogs of DsrAB, under anaerobic conditions it generates H2S from various sulfur compounds including thiosulfate, sulfite, tetrathionate, and elemental sulfur, a feature firstly revealed more than two and half decades ago (Myers and Nealson, 1988). However, the enzymatic foundation for the reduction was not revealed until recently (Burns and DiChristina, 2009; Shirodkar et al., 2011). In addition to SirACD mentioned above, S.oneidensis also relies on a thiosulfate and polysulfide reductase (PsrABC), homologous to that from S. enterica Serovar Typhimurium (Heinzinger et al., 1995). Based on available genome sequences, it is clear that most of Shewanella species, if not all, are able to produce H2S endogenously via respiration of sulfur species (Gao et al., 2010a).
Most recently, endogenous H2S production has been linked to iron reduction in S. oneidensis and Sulfurospirillum deleyianum (Flynn et al., 2014; Lohmayer et al., 2014). In alkaline environments containing sulfur species and iron compounds, these bacteria first generates H2S (HS−), which subsequently reduces iron compounds abiotically (Flynn et al., 2014), indicating an essential role of H2S in bacterial metal reduction under certain conditions. In contrast to the beneficial role, Shewanellae are increasingly being implicated as human pathogens in individuals exposed through occupational or recreational activities to marine niches where these species thrive (Janda and Abbott, 2014). Additionally, Shewanella have been recognized as spoilage bacteria of food (especially marine products) and associated with foul odors, which are in part attributed to endogenous H2S (Janda and Abbott, 2014).
Hence, understanding endogenous H2S production would not only provide insights into biogeochemical redox processes related to metal reduction, but also help to deter and mitigate the emerging threat to human health. In this study, we made efforts to provide a relatively comprehensive understanding of endogenous H2S production and regulation in S.oneidensis. We found that the bacterium possesses a large set of enzymes catalyzing H2S generation through cysteine degradation in addition to anaerobic respiration of sulfur-containing compounds. While respiration predominantly, if not exclusively, relies on the SirACD and PsrABC complexes there is no interplay with each other. We showed that global regulator Crp (cyclic-AMP receptor protein) but not Fnr (fumarate nitrate regulator) or Arc (aerobic respiration control) is essential to H2S generation via anaerobic respiration. Additionally, we found that both Crp and Arc are involved in H2S generation through cysteine degradation. While it is clear that Arc exerts its impact on the process by repressing expression of the major contributor, the mechanisms underlying the requirement of Crp remain unknown.
Methods and Materials
Bacterial Strains, Plasmids, and Culture Conditions
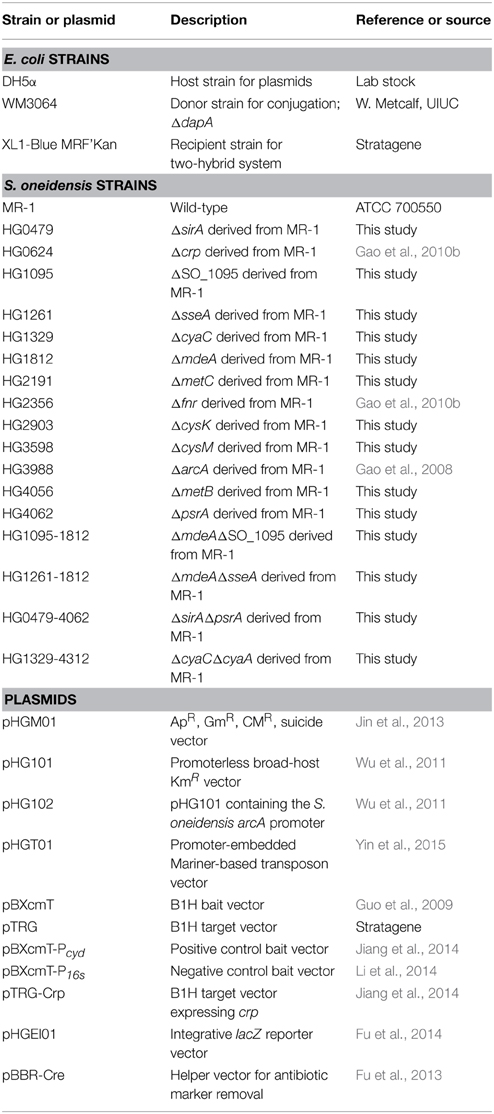
A list of all bacterial strains and plasmids used in this study is given in Table 1. All chemicals were acquired from Sigma Co. (Shanghai, China) unless otherwise noted. Information for primers used in this study is available upon request. For genetic manipulation, E. coli and S. oneidensis strains under aerobic conditions were grown in Lysogeny Broth (LB, Difco, Detroit, MI) medium (Bertani, 2004), which was modified to contain tryptone (10 g/L), yeast extract (5 g/L), and NaCl (5 g/L), at 37 and 30°C, respectively. When needed, the growth medium was supplemented with chemicals at the following concentrations: 2,6-diaminopimelic acid (DAP), 0.3 mM; ampicillin sodium, 50 μg/ml; kanamycin sulfate, 50 μg/ml; and gentamycin sulfate; 15 μg/ml.
Physiological Characterization of S. oneidensis Strains
M1 defined medium containing 0.02%(w/v) of vitamin free casamino acids and 20 mM lactate was used as described previously (Gao et al., 2008). Growth of S. oneidensis strains under aerobic or anaerobic conditions was determined by recording optical densities of 600 nm (OD600) of cultures. For aerobic growth, fresh media were inoculated to ~0.01 of OD600 with overnight cultures grown from a single colony. For anaerobic growth, exponential phase cultures grown aerobically were centrifuged, washed with fresh medium twice, purged in nitrogen and suspended in fresh medium to ~0.01 of OD600 in an anaerobic glove box. To cultivate Δcrp and ΔcyaC under anaerobic conditions, Trimethylamine N-oxide (TMAO) was used as growth supporting EA. EAs were used at the concentration of 10 mM unless otherwise noted.
Mutagenesis and Genetic Complementation
S. oneidensis in-frame deletion strains were constructed using the att-based Fusion PCR method (Jin et al., 2013). In brief, two fragments flanking the target gene were generated by PCR with primers containing attB and the gene specific sequence, and then were joined by a second round of PCR. The fusion fragments were introduced into plasmid pHGM01 by site-specific recombination using the BP Clonase (Invitrogen) according to the manufacturer's instruction. The resulting mutagenesis vectors were transformed into E. coli WM3064 and the verified ones were transferred into proper S. oneidensis strains via conjugation. Integration of the mutagenesis constructs into the chromosome were selected by resistance to gentamycin and confirmed by PCR. Verified transconjugants were grown in LB broth in the absence of NaCl and plated on LB supplemented with 10% sucrose. Gentamycin-sensitive and sucrose-resistant colonies were screened by PCR for the intended deletion. All mutations were verified by sequencing the mutated regions.
Mutants from previous studies were successfully complemented by using either multiple-copy or single-copy (integrative) vectors (Table 1). In this study, the same complementation constructs were used and similar results were obtained as indicated in the figure legends. For newly constructed mutants, plasmids pHG101 and pHG102 were utilized for genetic complementation (Wu et al., 2011). For genes adjacent to their promoter, a fragment containing the gene of interest and its native promoter was generated by PCR and cloned into pHG101. For others, the gene of interest was amplified and inserted into the multiple-cloning site (MCS) of pHG102 under the control of the arcA promoter, which is constitutively active (Gao et al., 2010b). After verified by sequencing, the resulting complementation vector was transferred into its relevant mutant strains via conjugation.
H2S Detection
H2S generation in S. oneidensis strains was monitored by using a lead acetate detection method (Shatalin et al., 2011). Paper strips saturated by 2% of Pb(Ac)2 were affixed to the inner wall of a cultural tube, above the level of the liquid culture. Overnight cultures were used to inoculate fresh LB media to ~0.01 of OD600 and incubated for ~20 h at 30°C with aeration. Stained paper strips were scanned and quantified with a UVP HR410 Imaging System (UVP). The results were normalized to OD600 readings. H2S concentrations in liquid cultures grown under aerobic and anaerobic conditions were quantified by using the methylene blue formation assay (Siegel, 1965). In brief, properly diluted aliquots (1.6 ml) were mixed with 0.2 ml DPD (20 mM N, N-dimetyl-p-phenylenediamine sulfate in 7.2 M HCl) and 0.2 ml FeCl3 (30 mM FeCl3 in 1.2 M HCl). After 30 min at 25°C, the absorbance at 667 nm was measured and related to sulfide concentration using calibration curves generated with NaHS.
Transposon Mutagenesis
A random transposon-insertion library was constructed with pHGT01, a mariner-based transposon vector which carries an embedded promoter within the transposable fragment (Fu et al., 2013; Yin et al., 2015). When mutants were grown in LB containing SO2−3, S2O2−3, and cysteine at 2 mM to the late exponential phase in 96-well plates, the lids of the plates were removed to let out gaseous H2S, and 1 h later the H2S levels of cultures in each well were visualized with paper strips saturated by 2% of Pb(Ac)2. Mutants exhibiting significant reduction in H2S generation were saved and then cultured in individual tubes for verification. The confirmed mutants were subjected to arbitrary PCR for mapping the transposon insertion sites (Das et al., 2005).
B1H Assay
Bacterial one-hydrid system was used to investigate DNA-protein interaction in vivo in E. coli cells (Guo et al., 2009). Briefly, plasmid constructs were created by cloning the bait DNA and target Crp into the pBXcmT and pTRG vectors, respectively, and verified by sequencing. The resultant plasmids were used to co-transform BacterioMatch II Validation Reporter Competent Cells on M9 salt agar plates containing 25 mg/ml chloramphenicol and 12.5 mg/ml tetracycline with or without 3-AT. A pair of pBXcmT-Pcyd/pTRG-Crp were used as positive control, and a 300 bp DNA fragment of the 16s rRNA gene promoter (pBXcmT-P16s) was used as negative control (Jiang et al., 2014; Li et al., 2014). The plates were incubated for 24 h and then moved to room temperature for an additional 16 h (colonies indicating positive interaction usually appeared between 18 and 24 h). The positive interactions were confirmed by streaking colonies on plates containing both 3-AT and streptomycin (12.5 mg/ml).
Expression Analyses
To prepare samples for expression analyses, cultures at the mid-log phase were aliquotted; an aliquot was used as the untreated control and other aliquots were added with chemicals indicated in the text, and then collected 30 min post-addition. In total, four biological replicates under each condition were collected for the analyses. Levels of gene expression were determined by using an integrative lacZ-reporter system and β-Galactosidase activity assay, as performed essentially the same as before (Fu et al., 2014; Jiang et al., 2014). In addition, levels of mRNAs were also assayed using Quantitative RT-PCR (qRT-PCR) with an ABI7300 96-well qRT-PCR system (Applied Biosystems) as described previously (Yuan et al., 2011).
Other Analyses
DNA and protein sequence similarity searches were performed using the BLAST program. Sequences of proteins of interest for alignments were obtained from Genbank. Alignments were performed using Clustal Omega (http://www.ebi.ac.uk/Tools/msa/clustalo). Promoter prediction for genes of interest was performed by using the Neural Network Promoter Prediction program (Reese, 2001). Experimental values are subjected to statistical analyses and presented as means ± SD (standard deviation). Student's t-test was performed for pairwise comparisons of groups.
Results
MdeA is the Major Contributor for H2S Generation Under Aerobic Conditions
Given that the assimilatory reduction of sulfite usually does not release H2S into the environment, endogenous H2S at physiologically relevant levels in cultures is predominantly produced by reductases reducing sulfur-containing compounds and orthologs of mammalian H2S generation enzymes (cysteine degradation), including CBS, CSE, and 3MST (Shatalin et al., 2011; Keseler et al., 2013) (Figure S1). In S. oneidensis, thiosulfate and polysulfite reductase PsrABC and sulfite reductase SirACD are known to be responsible for H2S generation under anaerobic conditions (Burns and DiChristina, 2009; Shirodkar et al., 2011). In contrast, the pathways for H2S generation through the cysteine degradation pathway remain veiled. To identify orthologs of mammalian cysteine degradation enzymes, we performed BLASTp screening using E. coli 3MST and P. aeruginosa CBS and CSE against the S. oneidensis proteome. Results showed that the genome encodes a set of high confidence hits for these three proteins (Table S1 and Figure S2). Multiple orthologs of high confidence for CBS (CysM and CysK) and CSE (MetB, MdeA, SO_1095, and MetC) are present while there is only one for 3MST (SseA). This is not surprising given that the E. coli genome also encodes several proteins homologous to these three enzymes (Shatalin et al., 2011). However, it is common that only a portion of the homologous proteins are enzymes involved in H2S generation, based on the findings from E. coli, B. anthracis, P. aeruginosa, and S. aureus (Shatalin et al., 2011). To assess the contribution of the predicted enzymes in H2S generation through cysteine degradation under aerobic conditions, we inactivated each set of enzymes chemically using their specific inhibitors: amino-oxyacetate (AOAA) for CBS, asparate (Asp) for 3MST, and DL-propargylglycine (PAG) for CSE (Shatalin et al., 2011). H2S generation was detected using lead acetate [Pb(Ac)2], which can specifically react with H2S to form brown-colored precipitate of lead sulfide [PbS], whose amount is proportional to the concentration of H2S. As shown in Figure 1A, effects of these inhibitors on H2S generation varied significantly. Compared to the non-inhibitor control culture, addition of 7.5 mM PAG nearly eliminated PbS staining (<10%) and Asp had a rather mild impact on H2S generation, manifesting contribution of CSE and 3MST in H2S generation. In contrast, addition of AOAA increased H2S generation significantly, implicating that the CBS homologs do not function as cystathionine β-synthase. To confirm that CSE is the major source of H2S generation, we assessed the effects of PAG of various concentrations. As expected, the amounts of H2S produced were inversely proportional to the concentrations of the inhibitor (Figure 1B). In parallel, H2S generation was also quantified in liquid static media in 24-well plates by using the methylene blue formation assay (Siegel, 1965). We found that data from these two methods were comparable in general. In the rest of the study, the methylene blue formation assay was chosen because it is more sensitive and easier for quantification.
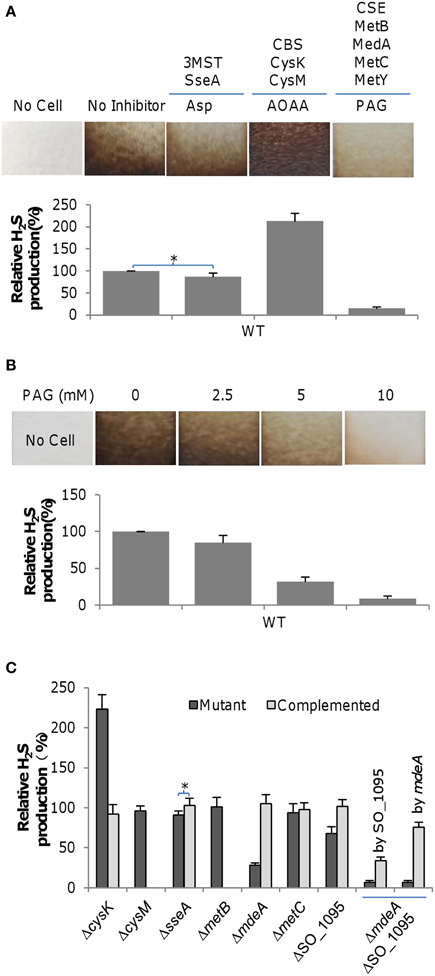
Figure 1. H2S generation by S. oneidensis. (A) Effects of Asp (3 mM), AOAA (50 μM), and PAG (7.5 mM), inhibitors of 3MST, CBS, and CSE respectively, on H2S generation by the S. oneidensis wild-type. (B) Effects of PAG, the CSE inhibitor, at varying concentrations on H2S generation by the S. oneidensis wild-type. (C) H2S generation by indicated S. oneidensis mutants. Mutants showing a significant defect in H2S generation were subjected to genetic complementation. The double mutant (ΔmdeAΔSO_1095) was complemented by either gene separately. Lead acetate–soaked paper strips show a PbS brown or black stain as a result of reaction with H2S. Strips were affixed to the inner wall of a culture tube, above the level of the liquid cultures for ~20 h. Levels of the wild-type were averaged and set to be 100%, and used to normalize averaged H2S levels of all other samples. In (C), the methylene blue formation assay was used for quantification of H2S. Data are presented as the mean ± SD from at least five independent experiments. Asterisks indicate statistically significant difference (*, p < 0.05).
To decisively determine roles of these enzymes in H2S generation under aerobic conditions, we constructed a set of mutants that lack one of these enzymes and monitored H2S production in the resulting mutants (Figure 1C). All mutants were indistinguishable from the wild-type with respect to growth under aerobic or anaerobic conditions with fumarate as the sole EA (data not shown). Among all mutants, the ΔcysK strain stood out as it produced substantially more H2S than the wild-type. This phenotype, similar to that resulting from the addition of AOAA, was confidently attributed to the cysK mutation after successful genetic complementation. The result, however, is not surprising because CysK primarily catalyzes the synthesis of L-cysteine from O-acetyl-L-serine and H2S although it has also been implicated in H2S generation (Byrne et al., 1988) (Figure S1). Consistent with the effect of Asp, a mutational analysis of the sseA gene revealed that the protein had a negligible role in H2S generation. In the case of CSE analogs, MetB and MetC appeared to be dispensable for H2S generation, a scenario agreeing with their annotated functions (Figure S1). In contrast, loss of either MdeA or SO_1095 resulted in drastic reduction in H2S generation, indicating that these two proteins are major contributors for releasing endogenous H2S. Apparently, MdeA is the most important enzyme for the task in S. oneidensis, accountable for at least 70% of H2S generation under test conditions. The involvement of MdeA, a methionine γ-lyase, in the process is reasonable because it converts L-cysteine and H2O to pyruvate, NH3, and H2S (Figure S1) (Sato and Nozaki, 2009). However, it is not immediately evident why SO_1095 contributes to H2S generation given that its predicted role as O-acetylhomoserine (thiol)-lyase is not related to any known pathway for H2S generation. The observed phenotypes of reduced H2S generation resulting from the mdeA and SO_1095 deletions were corrected by their expression in trans, validating the linkage between the phenotypes and corresponding mutations. To further confirm that MdeA and SO_1095 are enzymes predominantly responsible for endogenous H2S generation in S. oneidensis, we constructed a strain lacking both the mdeA and SO_1095 genes. Not surprisingly, the double removal further compromised the capacity of H2S generation (Figure 1C). These data, collectively, indicate that in S. oneidensis MdeA and SO_1095 dictate endogenous H2S generation.
H2S Generation Pathways with MdeA and SseA are Cysteine-inducible
The primary source of endogenous H2S is L-cysteine; as a consequence, many processes leading to H2S generation are subjected to induction by the amino acid (Shatalin et al., 2011). To determine which enzymes are cysteine-inducible, we measured the H2S generation capacity of S. oneidensis strains with cysteine. In the presence of 10 mM cysteine, there was an increase of ~2.8-fold in H2S generation in the wild-type (Figure 2A). Similar results were obtained from mutant strains in which mutations do not show any noticeable impact on H2S generation, confirming that these genes are not involved in the process. Additionally, induction to comparable levels by L-cysteine was also observed in the ΔsseA and ΔSO_1095, supporting that MdeA is the enzyme responsible for increased H2S generation. In both the ΔmdeA and ΔmdeAΔSO_1095 strains, the induction decreased to similar levels, implicating that SO_1095 may not be subjected to induction by L-cysteine. Rather, this induction is likely attributed to SseA. To test it, we generated an mdeA and sseA double mutant (ΔmdeAΔsseA) and assayed its capacity of H2S generation (Figure 2A). In the absence of cysteine, the mutant generated H2S at a level similar to that from the mdeA single mutant. Addition of cysteine did not result in any increase in H2S generation, indicating that the mutant is reluctant to respond to exogenous cysteine. These data manifest that SseA becomes a significant contributor for H2S generation under induced conditions.
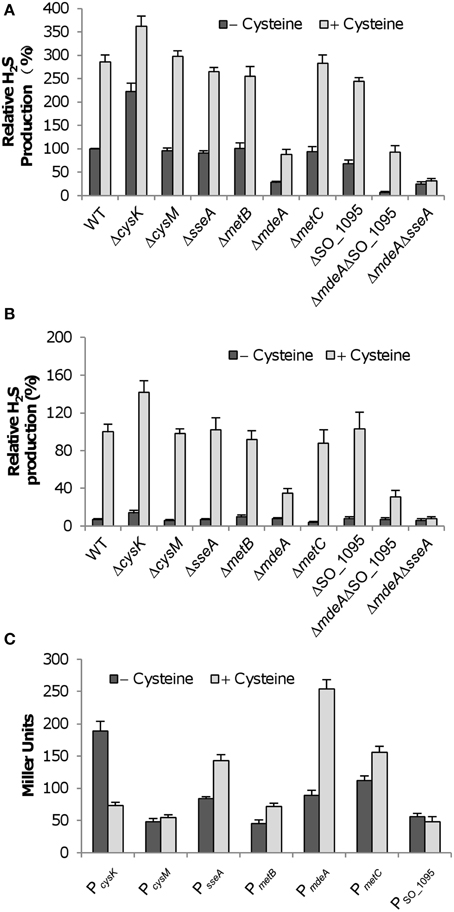
Figure 2. Effect of cysteine on S. oneidensis strains. (A) Effect of 10 mM cysteine on H2S generation by S. oneidensis strains under aerobic conditions. Averaged H2S level of the wild-type in the absence of cysteine was set to 100% for subsequent normalization. (B) Effect of 10 mM cysteine on H2S generation by S. oneidensis strains under anaerobic conditions. Fumarate was used as the sole EA. Averaged H2S level of the wild-type in the presence of cysteine was set to 100% for subsequent normalization because H2S levels were extremely low without induction. (C) Effect of 10 mM cysteine on activities of indicated promoters. For each test gene, the promoter-lacZ (E. coli) construct was introduced into the chromosome of the wild-type and the activities of the promoter as single copy were assessed by β-galactosidase assay. Data are presented as the mean ± SD from at least three independent experiments.
H2S generation through cysteine degradation is supposed to be independent of EAs for respiration. We therefore reasoned that the induction of H2S generation by L-cysteine should be observed under anaerobic conditions as well. To test this, the wild-type and mutant strains were assayed for H2S generation with fumarate as the sole EA. As shown in Figure 2B, H2S was hardly generated in any of these strains in the absence of L-cysteine, most likely because levels of metabolically synthesized substrates for cysteine degradation enzymes are too low to elicit a significant influence on overall H2S generation during growth. When cysteine was supplemented, levels of H2S in cultures of all test strains increased drastically (Figure 2B). Compared to the data obtained from the aerobic cultures, a similar trend was observed, confirming that the activities of MdeA and SseA are EA-independent and cysteine-inducible.
To further investigate into the effect of L-cysteine on H2S generation in S. oneidensis, we employed an integrative lacZ-reporter system pHGEI01 to assess expression of these genes (Fu et al., 2014). The reporter system is characterized by its ability to integrate into the chromosome and a removable antibiotic marker, and thus allows us to avoid complications of antibiotics on growth and of copy numbers of plasmids on lacZ expression. All of these genes are immediately adjacent to the promoters of their residing operons (data not shown). According to the promoter prediction, promoters of the highest confidence for all genes were located within 400 bp upstream of coding sequences (data not shown). We therefore amplified a fragment of ~ 400 bp upstream of each gene and inserted it into the multiple cloning site (MCS) before the E. coli lacZ gene in pHGEI01. After integration and antibiotic marker removal, mid-log phase cultures (~0.3 of OD600) were prepared for the β-galactosidase assay. Cysteine-treated samples were collected 30 min after cysteine addition. As shown in Figure 2C, lacZ expression driven by the cysM and SO_1095 promoters fluctuated within a narrow range, indicating that impacts of the addition of cysteine on their expression were negligible. In contrast, expression levels of the cysK, sseA, metB, metC, and mdeA genes were significantly different in cultures between the normal and cysteine-induced conditions. Given that cysteine is the product of CysK, reduction in cysK expression is expected upon the addition of the amino acid. The remaining four genes showed increases in their expression levels to varying extent, depending on the gene. Consistent with the data presented in Figure 2A, the sseA and mdeA genes were expressed substantially higher in the treated samples, confirming that SseA and MdeA are cysteine-inducible enzymes involved in H2S generation in S. oneidensis. In the case of the metB and metC genes, the induction is not surprising because cysteine is a substrate of the pathway in which they play a role (Figure S1). Moreover, similar results were obtained from an independent examination of expression levels of these genes by using qRT-PCR (Figure S3). Taken together, these data conclude that MdeA is the predominant enzyme for H2S generation from cysteine and SseA makes a significant contribution when cysteine is abundant.
SirACD and PsrABC are Independent of Each Other
With SO2−3, S0, and S2O2−3 as the sole EAs, S. oneidensis is able to generate H2S by using the SirACD and PsrABC complexes, respectively (Burns and DiChristina, 2009; Shirodkar et al., 2011). However, whether these two complexes interplay with each other is worth studying given that many sulfur species are enzymatically convertible from one another (Kimura, 2014). To this end, we constructed ΔsirA, ΔpsrA, and ΔsirAΔpsrA strains and compared their capacities of H2S generation to that of the wild-type. When grown on fumarate, H2S generation in the ΔsirA, ΔpsrA, and ΔsirAΔpsrA strains was extremely low, resembling that in the wild-type (Figure 3), implicating that loss of SirACD, PsrABC, or both has no effect on the cysteine degradation pathway under the condition. In the presence of SO2−3, H2S levels in cultures of the wild-type and ΔpsrA strains increased dramatically (Figure 3), an observation in excellent agreement with the results of a previous study (Shirodkar et al., 2011). Similarly, robust H2S generation was observed from the wild-type and ΔsirA strains with S2O2−3 as the sole EA. In both cases, the ΔsirAΔpsrA strain produced H2S at levels comparable to that of the wild-type grown on fumarate. In the presence of cysteine, H2S generation was further enhanced in the wild-type, suggesting an additive effect from respiration of sulfur species and cysteine degradation, which was also evident in ΔsirA with S2O2−3 and in ΔpsrA with SO2−3. Expression of each of these genes in trans in its corresponding mutant restored H2S generation to the wild-type levels, confirming that the observed phenotypes were due to the intended mutations (Figure S4). These results indicate that the SirACD and PsrABC complexes are fully responsible for dissimilatory H2S generation, and appear functionally independent of each other.
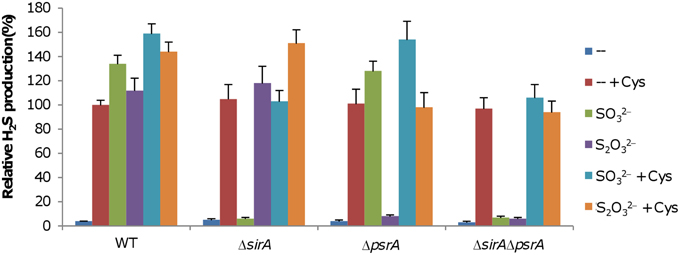
Figure 3. Characteristics of H2S generation by S. oneidensis strains under anaerobic conditions. Fumarate of 5 mM was used as EA to support growth to ~0.2 of OD600, which was then added with either SO2−3 or S2O2−3. H2S generation was measured 8 h after the addition. Averaged H2S level of the wild-type in the presence of cysteine was set to 100% for subsequent normalization as stated in Figure 2. “EA + Cys” represents the indicated EA plus cysteine. Data are presented as the mean ± SD from at least four independent experiments.
Crp is a Global Regulator Essential for Endogenous H2S Generation
Given that many enzymes participate in H2S generation simultaneously, we became interested in whether there is a regulator mediating all of the processes in which these enzymes are involved, not necessarily in a direct manner. In our effort to identify such a regulator, we took advantage of transposon-based random mutagenesis. A random mutant library was generated using pHGT01, which allows screening for active operons as well as cryptic ones because of an embedded promoter within the transposable fragment (Fu et al., 2013; Yin et al., 2015). The medium was supplemented with proper amounts of cysteine, SO2−3 and S2O2−3, such that one of the significant contributors alone can produce a sufficient amount of H2S for being a H2S-plus strain. In approximately 10,000 gentamycin-resistant isolates screened, we found ~200 showing substantial reduction in H2S generation by using Pb(Ac)2-socked paper strips. However, many of them turned out to be false positive during subsequent validation. Among the remaining, two mutants were completely negative in H2S generation whereas three generated H2S at levels just detectable. Insertions in the former two were mapped into the crp gene and in the latter three were mapped into the cyaC gene. In S. oneidensis, Crp is the most important global regulator for both aerobic and anaerobic respiration and is activated by binding to cAMP, whose production mainly depends on membrane-bound adenylate cyclase CyaC (Saffarini et al., 2003; Charania et al., 2009). The insertions found in both crp and cyaC genes strongly suggest that the Crp regulatory system likely plays a crucial role in endogenous H2S generation in this microorganism.
To confirm this, we constructed a cyaC in-frame deletion strain (ΔcyaC) and examined the H2S generation capacity of the resulting strain, along with a crp deletion strain (Δcrp). The H2S generation assay revealed that the Δcrp strain completely lost the ability to produce H2S whereas the ΔcyaC strain retained residual capacity, approximately 17% relative to the wild-type when grown with sulfite and thiosulfate anaerobically (Figure 4A). This scenario can be readily explained by the presence of other adenylate cyclases, CyaA and CyaB, the former of which displays a detectable, albeit weak, ability to synthesize cAMP (Charania et al., 2009). Additional removal of the cyaA gene from the cyaC− background further reduced H2S generation to a level comparable to that resulting from the crp deletion. Similar results were obtained with cultures grown with 2 mM cysteine under aerobic conditions (Figure 4B), manifesting that Crp is also required for H2S generation from cysteine degradation. The essentiality of Crp to H2S generation from both processes was further supported by similar findings observed from cultures grown with sulfur, thiosulfate, and 2 mM cysteine under anaerobic conditions (Figure S5). The observed phenotypes resulting from the crp or cyaC deletions were corrected by their expression in trans, confirming that the defect in H2S synthesis was due to the intended mutations (Figure 4).
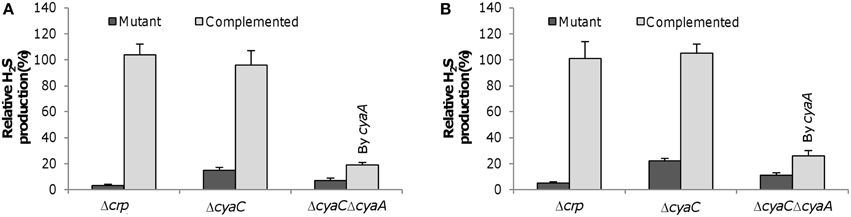
Figure 4. Crp-cAMP is essential to H2S generation by S. oneidensis. (A) H2S levels in anaerobic cultures of indicated mutants with SO2−3 and S2O2−3 at 2 mM each were measured and normalized to values of the wild-type. TMAO of 5 mM was used as EA to support growth to ~0.2 of OD600, which was then added with SO2−3 and S2O2−3. (B) H2S levels in aerobic cultures of indicated mutants with cysteine at 2 mM were measured and normalized to values from the wild-type. In both (A,B) The double mutant (ΔcyaCΔcyaA) was complemented by the cyaA gene. Data are presented as the mean ± SD from at least five independent experiments.
Crp Directly Controls Transcription of psrA and sirA but Not mdeA, SO_1095, or sseA
Given that Crp is the master regulator for anaerobic respiration and directly controls transcription of many genes encoding terminal reductases and their immediate regulators (Saffarini et al., 2003; Dong et al., 2012; Zhou et al., 2013), we predicted a similar scenario for the sirACD and psrABC genes, or at least some of them. The psrABC genes are predicted to be co-transcribed but the sirA and sirCD genes belong to two separate operons. To test whether these operons are under control of Crp, the promoters for these operons were predicted and fragments covering the promoter sequences were generated and placed in the front of the E. coli lacZ gene within pHGEI01 for β-galactosidase assay. In the Δcrp strain, PsirA and PpsrA were not responsive to SO2−3 and S2O2−3, respectively (Figure 5). However, the loss of Crp did not significantly affect the activity of PsirCD. We then assessed the impact of the crp deletion on expression of the mdeA, SO_1095, and sseA genes. Surprisingly, none of these genes was negatively affected by the loss of Crp with respective to expression, even in the presence of cysteine (Figure 5).
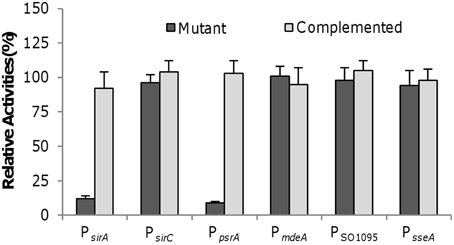
Figure 5. Crp on activities of indicated promoters. For each test gene, the promoter-lacZ construct was introduced into the chromosome of the Δcrp strain and the activities of the promoter as single copy were assessed by β-galactosidase assay. The same assay was performed with the complemented mutant for comparison. Data are presented as the mean ± SD from at least four independent experiments.
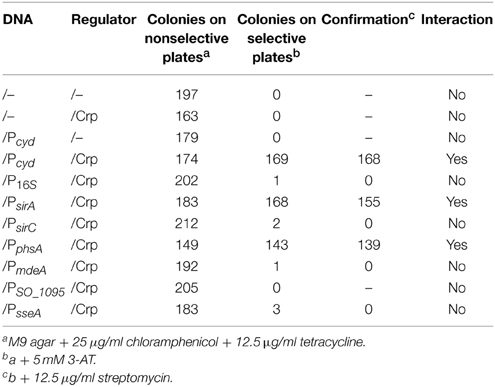
In a previous study, operons under direct control of Crp were predicted by screening the S. oneidensis genome to identify those with conserved Crp-binding motifs in their upstream region (Gao et al., 2010b). The presence of such motifs in front of both the sirA and psrABC operons is in line with our expression data presented above. To confirm there is direct interaction between Crp and the upstream region of the sirA and psrABC operons, we performed bacterial one-hybrid (B1H) analysis. The B1H assay, a robust technique applicable to a wide variety of different transcriptional factor families, used here is derived from the BacterioMatch II two-hybrid system (Guo et al., 2009). To detect interaction, vectors containing “bait” (DNA) and “target” (DNA-binding regulator) are co-transformed into BacterioMatch II Validation Reporter Competent Cells, of which those having positive DNA–protein interactions are able to grow on 3-amino-1,2,4-triazole (3-AT). To prepare the bait, a ~300 bp fragment centered by the predicted Crp-binding motif for each test gene was cloned into pBXcmT, which was paired with pTRG carrying the crp gene for co-transformation. Positive interactions from PsirA/Crp and PpsrA/Crp were detected and confirmed by growth on plates containing both 3-AT and streptomycin (12.5 mg ml−1), contrasting all other test genes that failed to produce any colonies in 40 h on the selective or confirmation plates (Table 2). In summary, these data manifest that Crp, likely in a direct manner, activates transcription of the sirA and psrABC operons but has little impact on the genes involved in cysteine degradation.
ArcA Directly Represses Transcription of the mdeA Gene
In S. oneidensis, the function of major global regulators, Crp, Arc, and Fnr, is substantially altered compared to their respective E. coli counterparts (Gao et al., 2010b). In E. coli, Fnr and Arc two-component system (TCS) are primarily responsible for the switch between aerobic and anaerobic metabolism and Crp is the decisive regulator in the process of carbon repression (Green and Paget, 2004; Deutscher et al., 2006). Interestingly, the S. oneidensis counterpart of E. coli Fnr has no significant role in these processes and Arc appears to be important in aerobic respiration and plays a limited role in anaerobiosis (Gralnick et al., 2005; Gao et al., 2008, 2010b), whereas Crp becomes the most important regulator mediating aerobic and anaerobic respiration (Saffarini et al., 2003; Dong et al., 2012; Zhou et al., 2013). Given the functional switches among these regulators, we intended to examine whether Fnr or Arc influences H2S synthesis in S. oneidensis. Under aerobic and anaerobic conditions, loss of Fnr did not show any noticeable effect on H2S generation, consistent with its overall dispensable role in respiration (Figure 6A). However, Arc, although negligible for H2S generation from anaerobic respiration, apparently was involved in regulation of cysteine degradation. In line with that Arc functions as a repressor in general, the loss of the system resulted in overproduction of H2S (Figure 6A).
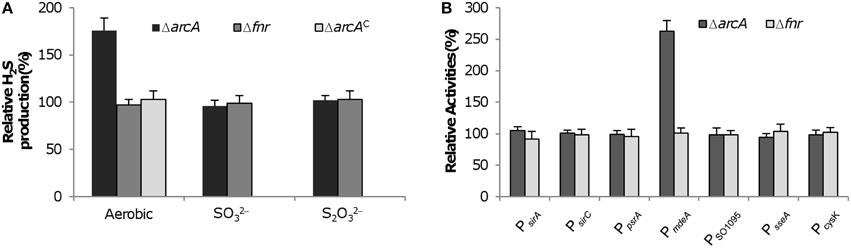
Figure 6. Impact of loss of on H2S generation in S. oneidensis. (A) ArcA and Fnr on H2S generation. H2S levels in cultures of indicated mutants grown aerobically or anaerobically with 10 mM SO2−3 or S2O2−3 as the sole EA were measured and normalized to values from the wild-type. Complementation of the ΔarcA mutant (ΔarcAC) was conducted with aerobic growth. (B) ArcA and Fnr on activities of indicated promoters. For each test gene, the promoter-lacZ construct was introduced into the chromosome of the Δcrp strain and the activities of the promoter as single copy were assessed by β-galactosidase assay. Data are presented as the mean ± SD from at least four independent experiments.
To determine which genes are under influence of Arc, we measured the activity of promoters for all genes involved in H2S generation (Figure 6B). Compared to the wild-type, the strain missing the fnr gene was indistinguishable in expression of all test genes, supporting that Fnr has no role in regulation of H2S generation. In the case of Arc, although expression levels of the sirA, sirCD, psrABC, SO_1095, and sseA operons were barely altered in the arcA deletion strain, the mdeA gene was expressed significantly higher compared to the wild-type. This repression by Arc is probably a result of direct binding to the upstream region of the operon because it contains an ArcA-binding motif (Gao et al., 2008; Wang et al., 2008). Moreover, we assessed the impact of Arc on expression of the cysK gene as its loss leads to a H2S overproduction phenotype, similar to that resulting from the loss of ArcA. However, expression levels of the cysK gene were comparable between the arcA positive and negative strains (Figure 6B), eliminating a possibility that Arc plays a role in cysteine biosynthesis. These results conclude that Crp is essential for H2S generation and Arc specifically regulates transcription of the mdeA gene whereas Fnr is dispensable in S. oneidensis.
Discussion
The purpose of this study was to identify predominant sources of H2S inside S. oneidensis. Previous studies on anaerobic respiration of sulfur-containing substances had demonstrated that PsrABC and SirACD enzyme complexes could catalyze H2S generation, provided that the corresponding EA was present (Burns and DiChristina, 2009; Shirodkar et al., 2011). PsrC and SirD belong to the NrfD/PsrC family of integral membrane proteins, which exclusively function as menaquinone oxidase/reductase (Simon and Kern, 2008). We provided evidence that these two complexes are predominantly, if not exclusively, accountable for H2S generation via respiration of sulfur species. Interestingly, there is no interplay between the complexes detected in this study, indicating that they do not release intermediates for each other during respiration.
In parallel, the genome of S. oneidensis encodes many homologs of enzymes in cysteine degradation, whose counterparts are accountable for H2S generation in mammals (Wang, 2012; Shatalin et al., 2011). By a combination of chemical and biological analyses, we determined that the predominant route for H2S generation is through the MdeA-based cysteine degradation. As a methionine γ-lyase, MdeA is highly homologous to P. aeruginosa CSE, able to use L-cysteine as a substrate, together with H2O, to produce pyruvate, NH3, and H2S (Sato and Nozaki, 2009). Additionally, we found that another CSE homolog and a 3MST homolog, SO_1095 and SseA respectively, contributed to H2S generation in S. oneidensis. Under aerobic growth conditions, MdeA accounts for at least two-thirds of H2S that is generated endogenously, regardless of the presence of cysteine. SO_1095, which is surprisingly not subjected to cysteine induction, catalyzes the reaction that releases H2S of ~20% in the absence of additional cysteine. On the contrary, SseA is highly inducible by cysteine but without the inducer its contribution is negligible. S. oneidensis probably does not possess functional CBS as CBS homologs appear to be cysteine synthases, suggesting that the bacterium, similar to those studied before such as E. coli, lacks at least one of enzymes for H2S generation via cysteine degradation (Shatalin et al., 2011).
In E. coli, the transcriptional regulator Fnr and the Arc TCS together control switches between aerobic and anaerobic respiration whereas Crp dictates carbon metabolism (Green and Paget, 2004; Deutscher et al., 2006). While Fnr of S. oneidensis has been reported to have a limited role in respiration of certain EAs (Maier and Myers, 2001; Cruz-Garcia et al., 2011), the data documented here, together with those presented previously, support the notion that the physiological significance of the regulator is marginal (Gao et al., 2010b; Fu et al., 2013; Zhou et al., 2013). The S. oneidensis Arc system is atypical, consisting of three components, ArcS, HptA and ArcA (Gralnick et al., 2005; Lassak et al., 2010; Shroff et al., 2010). Moreover, the regulons of the E. coli and S. oneidensis Arc systems have few components in common, suggesting a major difference in their physiological roles (Gao et al., 2008; Wang et al., 2008; Yuan et al., 2012). Several lines of evidence manifest that this three-component system functions primarily under aerobic conditions although it is required for dimethyl sulfoxide respiration, a phenomenon presumably arising from adventitious appearance of an ArcA-binding motif in front of the operon (Gao et al., 2008; Wang et al., 2008; Dong et al., 2014).
Instead, S. oneidensis employs Crp to control expression of most of respiratory reductases and, in some cases, their immediate regulators (Saffarini et al., 2003; Dong et al., 2012). Therefore, it is conceivable that expression of the psrA and sirA operons is mediated by Crp. By B1H analysis, we detected positive interactions between Crp and the upstream sequences of these two operons, manifesting that the regulation is carried out by Crp directly. Notably, the regulator of S. oneidensis resembles its E. coli counterpart in cAMP-dependent activation despite the substantial difference in Crp regulons between these two organisms (Charania et al., 2009; Zhou et al., 2013). This gains further support from the finding that loss of CyaC, the major cAMP synthetase, results in a phenotype similar to that caused by the Crp removal as shown here and before (Charania et al., 2009).
One of the most striking findings in this study is that Crp is also absolutely essential for H2S generation via cysteine degradation in S. oneidensis. We do not know yet the mechanism underlying this essentiality. Expression of the mdeA, SO_1095, and sseA genes is not affected by the Crp removal, indicating that Crp influences biological processes beyond these proteins. Given that the loss of Crp abolishes cysteine induction, we speculate that the regulator may play an essential role in controlling intracellular cysteine levels. Unlike Crp, the involvement of the Arc system in H2S generation via cysteine degradation is clear. The TCS represses expression of the mdeA gene likely in a direct control manner. To date, little is known about the environmental cues for S. oneidensis ArcS, but it is clearly different from the E. coli paradigm as the atypical system functions under both aerobic and anaerobic conditions (Gralnick et al., 2005; Gao et al., 2008; Yuan et al., 2012). We are working to decipher the mechanism by which Crp dictates H2S generation via cysteine degradation and to identify the signal triggering the Arc system.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This research was supported by National Natural Science Foundation of China (31270097, 41476105), Major State Basic Research Development Program (973 Program: 2010CB833803), and Doctoral Fund of Ministry of Education of China (20130101110142).
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2015.00374/abstract
References
Bertani, G. (2004). Lysogeny at mid-twentieth century: P1, P2, and other experimental systems. J. Bacteriol. 186, 595–600. doi: 10.1128/JB.186.3.595-600.2004
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Bradley, A. S., Leavitt, W. D., and Johnston, D. T. (2011). Revisiting the dissimilatory sulfate reduction pathway. Geobiology 9, 446–457. doi: 10.1111/j.1472-4669.2011.00292.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Burns, J. L., and DiChristina, T. J. (2009). Anaerobic respiration of elemental sulfur and thiosulfate by Shewanella oneidensis MR-1 requires psrA, a homolog of the phsA gene of Salmonella enterica Serovar Typhimurium LT2. Appl. Environ. Microbiol. 75, 5209–5217. doi: 10.1128/AEM.00888-09
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Byrne, C. R., Monroe, R. S., Ward, K. A., and Kredich, N. M. (1988). DNA sequences of the cysK regions of Salmonella typhimurium and Escherichia coli and linkage of the cysK regions to ptsH. J. Bacteriol. 170, 3150–3157.
Charania, M. A., Brockman, K. L., Zhang, Y., Banerjee, A., Pinchuk, G. E., Fredrickson, J. K., et al. (2009). Involvement of a membrane-bound class III adenylate cyclase in regulation of anaerobic respiration in Shewanella oneidensis MR-1. J. Bacteriol. 191, 4298–4306. doi: 10.1128/JB.01829-08
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Cordova, C. D., Schicklberger, M. F., Yu, Y., and Spormann, A. M. (2011). Partial functional replacement of CymA by SirCD in Shewanella oneidensis MR-1. J. Bacteriol. 193, 2312–2321. doi: 10.1128/JB.01355-10
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Cruz-Garcia, C., Murray, A. E., Rodrigues, J. L., Gralnick, J. A., McCue, L. A., Romine, M. F., et al. (2011). Fnr (EtrA) acts as a fine-tuning regulator of anaerobic metabolism in Shewanella oneidensis MR-1. BMC Microbiol. 11:64. doi: 10.1186/1471-2180-11-64
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Das, S., Noe, J. C., Paik, S., and Kitten, T. (2005). An improved arbitrary primed PCR method for rapid characterization of transposon insertion sites. J. Microbiol. Methods 63, 89–94. doi: 10.1016/j.mimet.2005.02.011
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Deutscher, J., Francke, C., and Postma, P. W. (2006). How phosphotransferase system-related protein phosphorylation regulates carbohydrate metabolism in bacteria. Microbiol. Mol. Biol. Rev. 70, 939–1031. doi: 10.1128/MMBR.00024-06
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Dong, Y., Wan, F., Yin, J., and Gao, H. (2014). Ecological roles of Arc signal transduction system revealed by evolutionary genetic analysis. J. Bacteriol. Mycol. 1:8. Available online at: http://austinpublishinggroup.com/bacteriology/fulltext/bacteriology-v1-id1007.php
Dong, Y., Wang, J., Fu, H., Zhou, G., Shi, M., and Gao, H. (2012). A Crp-dependent two-component system regulates nitrate and nitrite respiration in Shewanella oneidensis. PLoS ONE 7:e51643. doi: 10.1371/journal.pone.0051643
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Flynn, T. M., O'Loughlin, E. J., Mishra, B., DiChristina, T. J., and Kemner, K. M. (2014). Sulfur-mediated electron shuttling during bacterial iron reduction. Science 344, 1039–1042. doi: 10.1126/science.1252066
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Fredrickson, J. K., Romine, M. F., Beliaev, A. S., Auchtung, J. M., Driscoll, M. E., Gardner, T. S., et al. (2008). Towards environmental systems biology of Shewanella. Nat. Rev. Micro. 6, 592–603. doi: 10.1038/nrmicro1947
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Fu, H., Chen, H., Wang, J., Zhou, G., Zhang, H., Zhang, L., et al. (2013). Crp-dependent cytochrome bd oxidase confers nitrite resistance to Shewanella oneidensis. Environ. Microbiol. 15, 2198–2212. doi: 10.1111/1462-2920.12091
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Fu, H., Jin, M., Ju, L., Mao, Y., and Gao, H. (2014). Evidence for function overlapping of CymA and the cytochrome bc1 complex in the Shewanella oneidensis nitrate and nitrite respiration. Environ. Microbiol. 16, 3181–3195. doi: 10.1111/1462-2920.12457
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Gao, H., Barua, S., Liang, Y., Wu, L., Dong, Y., Reed, S., et al. (2010a). Impacts of Shewanella oneidensis c-ype cytochromes on aerobic and anaerobic respiration. Microb. Biotechnol. 3, 455–466. doi: 10.1111/j.1751-7915.2010.00181.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Gao, H., Wang, X., Yang, Z. K., Chen, J., Liang, Y., Chen, H., et al. (2010b). Physiological roles of ArcA, Crp, and EtrA and their interactive control on aerobic and anaerobic respiration in Shewanella oneidensis. PLoS ONE 5:e15295. doi: 10.1371/journal.pone.0015295
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Gao, H., Wang, X., Yang, Z., Palzkill, T., and Zhou, J. (2008). Probing regulon of ArcA in Shewanella oneidensis MR-1 by integrated genomic analyses. BMC Genomics 9:42. doi: 10.1186/1471-2164-9-42
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Gralnick, J. A., Brown, C. T., and Newman, D. K. (2005). Anaerobic regulation by an atypical Arc system in Shewanella oneidensis. Mol. Microbiol. 56, 1347–1357. doi: 10.1111/j.1365-2958.2005.04628.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Green, J., and Paget, M. S. (2004). Bacterial redox sensors. Nat. Rev. Micro. 2, 954–966. doi: 10.1038/nrmicro1022
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Guo, M., Feng, H., Zhang, J., Wang, W., Wang, Y., Li, Y., et al. (2009). Dissecting transcription regulatory pathways through a new bacterial one-hybrid reporter system. Genome Res. 19, 1301–1308. doi: 10.1101/gr.086595.108
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Heinzinger, N. K., Fujimoto, S. Y., Clark, M. A., Moreno, M. S., and Barrett, E. L. (1995). Sequence analysis of the phs operon in Salmonella typhimurium and the contribution of thiosulfate reduction to anaerobic energy metabolism. J. Bacteriol. 177, 2813–2820.
Janda, J. M., and Abbott, S. L. (2014). The genus Shewanella: from the briny depths below to human pathogen. Crit. Rev. Microbiol. 40, 293–312. doi: 10.3109/1040841X.2012.726209
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Jiang, Y., Dong, Y., Luo, Q., Li, N., Wu, G., and Gao, H. (2014). Protection from oxidative stress relies mainly on derepression of OxyR-dependent KatB and Dps in Shewanella oneidensis. J. Bacteriol. 196, 445–458. doi: 10.1128/JB.01077-13
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Jin, M., Jiang, Y., Sun, L., Yin, J., Fu, H., Wu, G., et al. (2013). Unique organizational and functional features of the cytochrome c maturation system in Shewanella oneidensis. PLoS ONE 8:e75610. doi: 10.1371/journal.pone.0075610
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Jormakka, M., Yokoyama, K., Yano, T., Tamakoshi, M., Akimoto, S., Shimamura, T., et al. (2008). Molecular mechanism of energy conservation in polysulfide respiration. Nat. Struct. Mol. Biol. 15, 730–737. doi: 10.1038/nsmb.1434
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Keseler, I. M., Mackie, A., Peralta-Gil, M., Santos-Zavaleta, A., Gama-Castro, S., Bonavides-Martínez, C., et al. (2013). EcoCyc: fusing model organism databases with systems biology. Nucleic Acids Res. 41, D605–D612. doi: 10.1093/nar/gks1027
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kimura, H. (2014). Generation and physiological effects of hydrogen sulfide. Antioxid. Redox Signal. 20, 783–793. doi: 10.1089/ars.2013.5309
Lassak, J., Henche, A.-L., Binnenkade, L., and Thormann, K. M. (2010). ArcS, the cognate sensor kinase in an atypical Arc system of Shewanella oneidensis MR-1. Appl. Environ. Microbiol. 76, 3263–3274. doi: 10.1128/AEM.00512-10
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Li, N., Luo, Q., Jiang, Y., Wu, G., and Gao, H. (2014). Managing oxidative stresses in Shewanella oneidensis: intertwined roles of the OxyR and OhrR regulons. Environ. Microbiol. 16, 1821–1834. doi: 10.1111/1462-2920.12418
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lohmayer, R., Kappler, A., Lösekann-Behrens, T., and Planer-Friedrich, B. (2014). Sulfur species as redox partners and electron shuttles for ferrihydrite reduction by Sulfurospirillum deleyianum. Appl. Environ. Microbiol. 80, 3141–3149. doi: 10.1128/AEM.04220-13
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Maier, T. M., and Myers, C. R. (2001). Isolation and characterization of a Shewanella putrefaciens MR-1 electron transport regulator etrA mutant: reassessment of the role of EtrA. J. Bacteriol. 183, 4918–4926. doi: 10.1128/JB.183.16.4918-4926.2001
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Myers, C. R., and Nealson, K. H. (1988). Bacterial manganese reduction and growth with manganese oxide as the sole electron acceptor. Science 240, 1319–1321. doi: 10.1126/science.240.4857.1319
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Oliveira, T. F., Vonrhein, C., Matias, P. M., Venceslau, S. S., Pereira, I. A., and Archer, M. (2008). The crystal structure of Desulfovibrio vulgaris dissimilatory sulfite reductase bound to DsrC provides novel insights into the mechanism of sulfate respiration. J. Biol. Chem. 283, 34141–34149. doi: 10.1074/jbc.M805643200
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Reese, M. G. (2001). Application of a time-delay neural network to promoter annotation in the Drosophila melanogaster genome. Comput. Chem. 26, 51–56. doi: 10.1016/S0097-8485(01)00099-7
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Saffarini, D. A., Schultz, R., and Beliaev, A. (2003). Involvement of cyclic AMP (cAMP) and cAMP receptor protein in anaerobic respiration of Shewanella oneidensis. J. Bacteriol. 185, 3668–3671. doi: 10.1128/JB.185.12.3668-3671.2003
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Sato, D., and Nozaki, T. (2009). Methionine gamma-lyase: the unique reaction mechanism, physiological roles, and therapeutic applications against infectious diseases and cancers. IUBMB Life 61, 1019–1028. doi: 10.1002/iub.255
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Shatalin, K., Shatalina, E., Mironov, A., and Nudler, E. (2011). H2S: a universal defense against antibiotics in bacteria. Science 334, 986–990. doi: 10.1126/science.1209855
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Shirodkar, S., Reed, S., Romine, M., and Saffarini, D. (2011). The octahaem SirA catalyses dissimilatory sulfite reduction in Shewanella oneidensis MR-1. Environ. Microbiol. 13, 108–115. doi: 10.1111/j.1462-2920.2010.02313.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Shroff, N. P., Charania, M. A., and Saffarini, D. A. (2010). ArcB1, a homolog of Escherichia coli ArcB, regulates dimethyl sulfoxide reduction in Shewanella oneidensis MR-1. J. Bacteriol. 192, 3227–3230. doi: 10.1128/JB.01695-09
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Siegel, L. M. (1965). A direct microdetermination for sulfide. Anal. Biochem. 11, 126–132. doi: 10.1016/0003-2697(65)90051-5
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Simon, J., and Kern, M. (2008). Quinone-reactive proteins devoid of haem b form widespread membrane-bound electron transport modules in bacterial respiration. Biochem. Soc. Trans. 36, 1011–1016. doi: 10.1042/BST0361011
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Wang, R. (2012). Physiological implications of hydrogen sulfide: a whiff exploration that blossomed. Physiol. Rev. 92, 791–896. doi: 10.1152/physrev.00017.2011
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Wang, X., Gao, H., Shen, Y., Weinstock, G. M., Zhou, J., and Palzkill, T. (2008). A high-throughput percentage-of-binding strategy to measure binding energies in DNA-protein interactions: application to genome-scale site discovery. Nucleic Acids Res. 36, 4863–4871. doi: 10.1093/nar/gkn477
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Wu, L., Wang, J., Tang, P., Chen, H., and Gao, H. (2011). Genetic and molecular characterization of flagellar assembly in Shewanella oneidensis. PLoS ONE 6:e21479. doi: 10.1371/journal.pone.0021479
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Yin, J., Jin, M., Zhang, H., Ju, L., Zhang, L., and Gao, H. (2015). Regulation of nitrite resistance of the cytochrome cbb3 oxidase by cytochrome c ScyA in Shewanella oneidensis. Microbiologyopen 4, 84–99. doi: 10.1002/mbo3.224
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Yuan, J., Wei, B., Lipton, M. S., and Gao, H. (2012). Impact of ArcA loss in Shewanella oneidensis revealed by comparative proteomics under aerobic and anaerobic conditions. Proteomics 12, 1957–1969. doi: 10.1002/pmic.201100651
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Yuan, J., Wei, B., Shi, M., and Gao, H. (2011). Functional assessment of EnvZ/OmpR two-component system in Shewanella oneidensis. PLoS ONE 6:e23701. doi: 10.1371/journal.pone.0023701
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Zhou, G., Yin, J., Chen, H., Hua, Y., Sun, L., and Gao, H. (2013). Combined effect of loss of the caa3 oxidase and Crp regulation drives Shewanella to thrive in redox-stratified environments. ISME J. 7, 1752–1763. doi: 10.1038/ismej.2013.62
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Keywords: H2S, endogenous generation, regulation, Crp, Shewanella
Citation: Wu G, Li N, Mao Y, Zhou G and Gao H (2015) Endogenous generation of hydrogen sulfide and its regulation in Shewanella oneidensis. Front. Microbiol. 6:374. doi: 10.3389/fmicb.2015.00374
Received: 28 February 2015; Accepted: 12 April 2015;
Published: 28 April 2015.
Edited by:
Dongsheng Zhou, Beijing Institute of Microbiology and Epidemiology, ChinaReviewed by:
Atsushi Kouzuma, Tokyo University of Pharmacy and Life Sciences, JapanJürgen Lassak, Ludwig-Maximilians-Universität Munich, Germany
Copyright © 2015 Wu, Li, Mao, Zhou and Gao. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Haichun Gao, Institute of Microbiology and College of Life Sciences, Zhejiang University, Hangzhou, Zhejiang 310058, China,aGFpY2h1bmdAemp1LmVkdS5jbg==
 Genfu Wu
Genfu Wu Guangqi Zhou
Guangqi Zhou Haichun Gao
Haichun Gao