- Unité de Recherche sur les Maladies Infectieuses et Tropicales Emergentes, UM 63, CNRS 7278, IRD 198, Inserm 1095, Faculté de Médecine, Aix Marseille Université, Marseille, France
Laboratory experiments have revealed many active mechanisms by which bacteria can inhibit the growth of other organisms. Bacteriocins are a diverse group of natural ribosomally synthesized antimicrobial peptides produced by a wide range of bacteria and which seem to play an important role in mediating competition within bacterial communities. In this study, we have identified and established the structural classification of putative bacteriocins encoded by 317 microbial genomes in the human intestine. On the basis of homologies to available bacteriocin sequences, mainly from lactic acid bacteria, we report the widespread occurrence of bacteriocins across the gut microbiota: 175 bacteriocins were found to be encoded in Firmicutes, 79 in Proteobacteria, 34 in Bacteroidetes, and 25 in Actinobacteria. Bacteriocins from gut bacteria displayed wide differences among phyla with regard to class distribution, net positive charge, hydrophobicity and secondary structure, but the α-helix was the most abundant structure. The peptide structures and physiochemical properties of bacteriocins produced by the most abundant bacteria in the gut, the Firmicutes and the Bacteroidetes, seem to ensure low antibiotic activity and participate in permanent intestinal host defense against the proliferation of harmful bacteria. Meanwhile, the potentially harmful bacteria, including the Proteobacteria, displayed highly effective bacteriocins, probably supporting the virulent character of diseases. These findings highlight the eventual role played by bacteriocins in gut microbial competition and their potential place in antibiotic therapy.
Introduction
Bacteriocins are natural antimicrobial peptides that are ribosomally synthesized by many bacteria and some Archaea (Cotter et al., 2005; Zacharof and Lovitt, 2012; Yang et al., 2014). Bacteriocins have been shown to successfully inhibit pathogenic bacteria such as Enterobacter, Salmonella, Klebsiella, Escherichia, Listeria, Enterococcus, and Clostridium (Hechard and Sahl, 2002; Snelling, 2005; Kirkup, 2006; Gillor et al., 2008). Therefore, bacteriocins have become increasingly of interest as viable alternative antibiotics to bacteriocins (Cotter et al., 2013). When they were discovered (Gratia, 1925), bacteriocins were thought to be secreted by certain bacteria for the purpose of eliminating competition in the medium. Currently, probiotics are used to enhance the ratio of beneficial to undesirable bacteria in human gastrointestinal microbiota (Fooks and Gibson, 2002) probably via the production of bacteriocins. Probiotics have been proposed to ensure in situ production of bacteriocins in order to fight against intestinal infections (Dobson et al., 2012; O’Shea et al., 2012; Schuijt et al., 2013). Altogether, bacteriocins may play a crucial role in determining the composition of gut microbiota (Corr et al., 2007; Angelakis et al., 2013; Million et al., 2013). They are presumed to actively participate in gastrointestinal host defense mechanisms by inhibiting one ecosystem, encouraging another, and offering a competitive advantage to bacteria in the intestinal tract.
A number of web-based software programs including the UniProtKB/Swiss-Prot, BAGEL (de Jong et al., 2006; van Heel et al., 2013) and BACTIBASE (Hammami et al., 2007, 2010) allow the detection and characterization of bacteriocins. Some 300 different bacteriocins have been identified and are available in public databases. They can be classified into three main categories on the basis of common characteristics such as molecular weight, heat stability and primary peptide structure. Class I, lanthionine-containing lantibiotics, are heat-stable polycyclic peptides (<5 kDa). Class II, non-lanthionine-containing bacteriocins are small heat-stable bacteriocins (<10 kDa), are subdivided into subclass IIa (pediocin-like bacteriocins), IIb (two-peptide bacteriocins), IIc (circular bacteriocins), and IId (bacteriocins that do not match the other three categories). Class III are large, heat-labile bacteriocins (>30 kDa) including bacteriolysins (murein hydrolases), colicins and linocins (Klaenhammer, 1993; van Belkum and Stiles, 2000; Kemperman et al., 2003; Cotter et al., 2005, 2013). Considering the potential ecological relevance of bacteriocins in microbial communities, the number of recognized bacteriocins seems to be relatively low. Further studies are needed to assess the production of bacteriocin in microbial communities. Since function prediction is essentially based on homology, the large sequence variability and very small length of certain bacteriocins (less than 30 amino acids) make computational identification of bacteriocins very hard (Whisstock and Lesk, 2003). Thus, bacteriocin databases are most likely incomplete and several genes encoding for bacteriocins are probably annotated as hypothetical proteins.
Here, we focused our analysis on the human gut microbiota. We constructed the most exhaustive database for bacteriocins (‘BUR’ standing for Bacteriocins of the URMITE database) by collecting all currently available sequences from the databases and from NCBI. Protein sequences from this database allowed putative bacteriocins from human gut microbiota to be identified using BLASTp methodology.
Materials and Methods
Bacteriocin Database
Bacteriocin peptide sequences were collected from the BAGEL (482 sequences) and BACTIBASE (228 sequences) bacteriocin databases (de Jong et al., 2006; Hammami et al., 2007) as well as from the NCBI database (834 bacteriocins). A thesaurus-search of the NCBI Entrez Gene database using bacteriocin, sakacin, microcin, colicin, and plantaricin and other keywords related to bacteriocins, followed by manual examination of the linked Pubmed article information allowed all the sequences of bacteriocins reported in the literature to be retrieved. The multi-FASTA text file containing protein sequences was manually curated to remove all redundant sequences. Similar bacteriocin sequences from different strains were conserved.
Strain Sequences Collected
From NCBI and our sequencing platform, we retrieved a total of 641 available genomes for organisms from the gastrointestinal tract (307 whole genomes and 334 draft genomes) belonging to 199 different bacterial genera, including Lactobacillus (65 strains), Streptococcus (32 strains), Clostridium (32 strains), and Bacillus (30 strains). Among these genomes, 398 were Gram-positive bacteria, and 243 were Gram-negative bacteria.
Bacteriocins Prediction
A bidirectional protein BLAST (BLASTp) was performed to identify bacteriocins in the gut genomes. Protein sequences encoding for bacteriocins were used as search strings in a BLASTp series [27] against gut genomes. All primary BLAST hits returning with identity over 50% and coverage over 70% were then subjected to BLASTp against the BUR database using same thresholds.
Web Interface
The BUR database runs on a Windows NT platform (Microsoft Windows 2000) with an Apache web server (version 2.0.55). The web server and all parts of the database are hosted at the URMITE in France. The web interface is accessible from the ‘IHU Méditerranée Infection’ page in the database section http://www.mediterranee-infection.com/article.php?larub=143&titre=base-de-donnees (Figure 1A). The web interface consists of two main web pages: one homepage with general information about bacteriocins and another page allowing users to search the database using a BLAST algorithm (protein-protein BLAST), version 2.2.13. Sequences that are homologous to the submitted sequences are shown as multiple alignments.
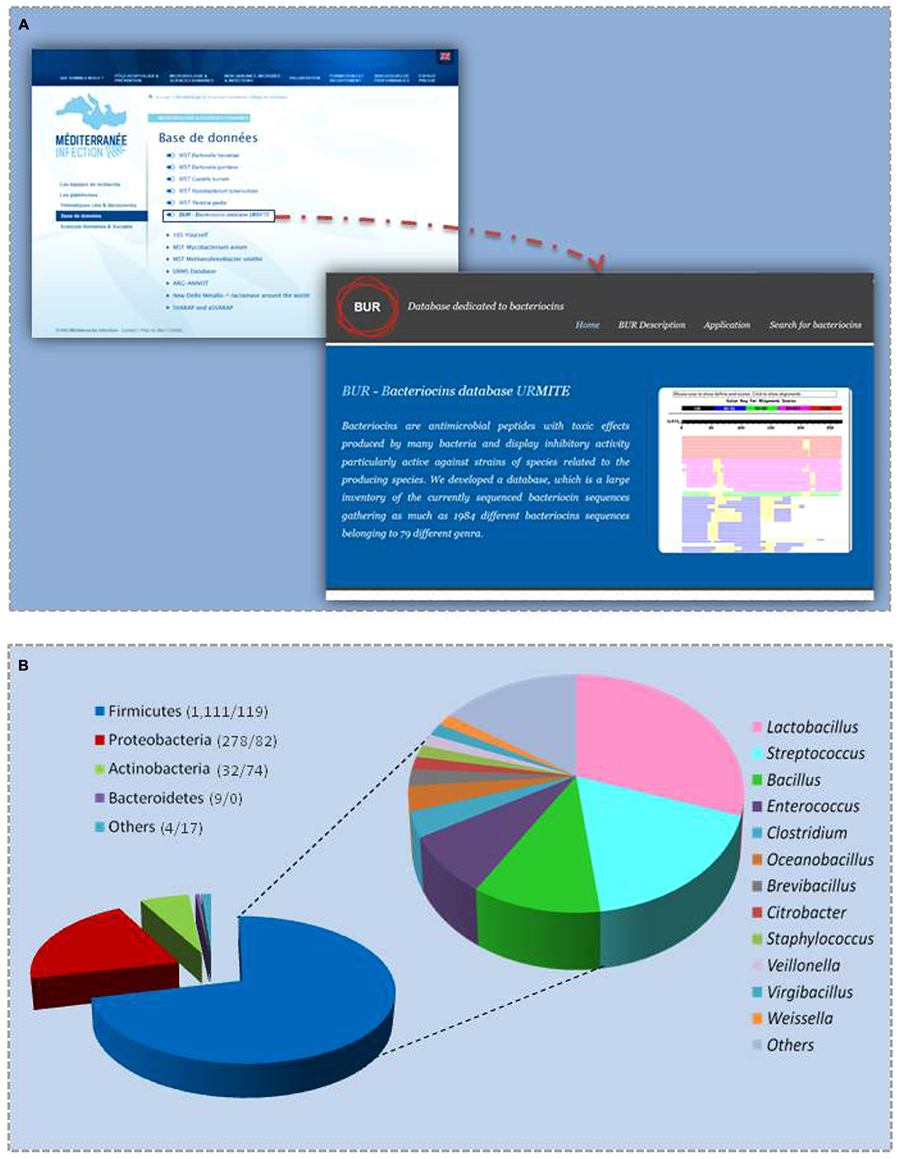
FIGURE 1. (A) Home page access and (B) phylum distribution of BUR bacteriocin sequences (numbers in brackets correspond to the number of bacteriocins found in gut bacteria and in other bacteria).
Information Related to Bacteriocins and Statistical Analysis
General data, such as peptide name, class and microbial data (producer organism, phylum, and Gram staining), were collected for each peptide. Additional physicochemical data including length, mass, amino acid composition, charged residues, hydrophobic (alanine, phenylalanine, glycine, isoleucine, leucine, methionine, proline, valine, and tryptophan) and hydrophilic (cysteine, asparagine, glutamine, serine, threonine, and tyrosine) residues were provided using CLC Genomics v7 software (CLC bio, Aarhus, Denmark). The prediction of bacteriocin potency and spectrum of activity was obtained using Phyre 2 software1 (Kelley and Sternberg, 2009). Principal component analysis was performed within the R statistical package2 to infer relationships between physiochemical properties and secondary structures within the different bacteriocins classes. SYSTAT 13 software3 allowed us to perform statistical analysis using a Chi-squared test. A p-value of <0.05 was considered significant.
Results
The generated database holds 1,359 bacteriocin sequences including 1,050 bacteriocins from Gram-positive bacteria and 309 from Gram-negative bacteria. Of these, 962 are produced by Firmicutes (mainly Streptococcus and Lactobacillus), 292 by Proteobacteria, 87 by Actinobacteria, eight by Thermotogae, four by Euryarchaeota, three by Aquificae, two by Spirochaetes, and one by Bacteroidetes. The bacteriocin sequences differ greatly in size and composition, with sequences ranging from 10 amino acids (bacteriocins produced by Lactobacillus curvatus) to 2,064 amino acids (a bacteriocin produced by Xylella fastidiosa strain 9a5c), with average length and weight of 160 amino acids and 18 kDa, respectively. These bacteriocin sequences can be classified into the three classes as follows: 243 in the Class I, 492 in the Class II and 624 in the Class III. Alanine constitutes the most abundant amino acid in the bacteriocin sequences. All these bacteriocins contain an average of 17 glycine residues but no leucine or asparagine residues. Bacteriocin sequences showed a low similarity score of 15% within classes, probably in accordance with the heterogeneity of modes of action and antimicrobial spectra.
When using the available 1,359 sequences of non-redundant protein sequences, we found 367 genes encoding different types of bacteriocins in 317 studied genomes of bacteria from the gastrointestinal tract (Supplementary File 1). Most of these genes were previously annotated as hypothetical proteins in the NCBI database. The 367 bacteriocin sequences identified were added to the BUR database, giving a final database composed of 1,726 peptide sequences (Figure 1A). The majority of bacteriocins in the BUR database were produced by Firmicutes (71%), Proteobacteria (21%), Actinobacteria (6%), and Bacteroidetes (0.5%; a total of 1,705 bacteriocins; Figure 1B). We observed that 40% of Firmicutes bacteria encode more than three bacteriocins, whereas bacteria from other phyla did not contain more than three bacteriocins. Lactobacillus, Enterococcus, and Streptococcus spp. encoded one to 24, one to 12 and one to 10 bacteriocins, respectively. Lactobacillus plantarum WCFS1, with 24 genes, encoded the largest number of bacteriocins, followed by L. plantarum ZJ316 with 23 genes and L. plantarum 16 with 21 bacteriocins.
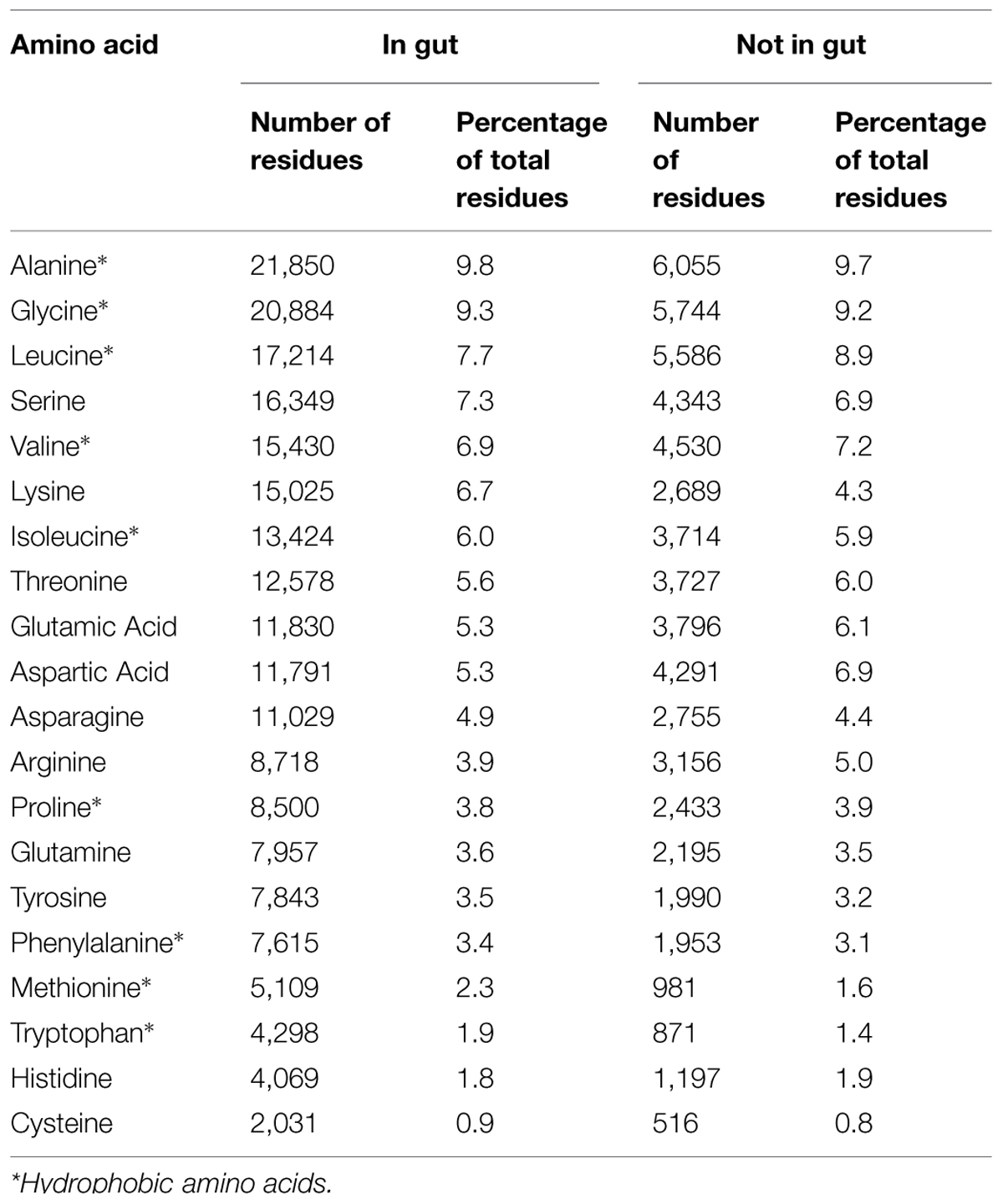
Bacteriocins produced by bacteria found in the gut are very different from those produced by other bacteria with regards to amino acid composition and length. Thus, bacteriocins produced by gut bacteria had significantly smaller percentages of aspartic acid, leucine, arginine, or glutamic acid but higher percentages of lysine and methionine than other bacteriocins (Table 1) and they were shorter in length (155 amino acid vs. 298 amino acid, p = 0.00). Overall, gut bacteria possessed more class I bacteriocins (44%) than class II (38.6%) or class III (17.3%), whereas other bacteria possessed more class III bacteriocins (59.2%) than class II (27.7%) or class I (13%; p = 0.00). Moreover, average hydrophobic residue content and net positive charge of the different classes of bacteriocins from the BUR database were similar (52.5% and +4, respectively).
Analysis of bacteriocins only from gut bacteria revealed significant differences in the structure and physiochemical properties of bacteriocins among the different phyla. Bacteriocins from Firmicutes were distributed fairly evenly across the different classes with 20% in class I, 45% in class II, and 35% in class III. Bacteriocins from Actinobacteria and Proteobacteria belonged primarily to class III (58 and 94%, respectively), and bacteriocins from Bacteroides were exclusively in class III. Proteobacteria had the highest net positive charge (+6), whereas Bacteroidetes had the lowest net positive charge (+3; p = 0.01; Figure 2A). Although the distribution of the percentage of hydrophobic residues varied widely within the different phyla (Figure 2B), bacteriocins from Firmicutes, Proteobacteria, and Actinobacteria all had an average of 53% hydrophobic residues, while Bacteroidetes contained 60% hydrophobic residues. Using homology models, we determined the 2D structure and activity domain for 731 bacteriocins from gut bacteria. Bacteriocins from Firmicutes displayed 31% α-helix, 31% β-strand, and 31% disordered structures, whereas Actinobacteria and Bacteroidetes contained a majority of β-strand structures (32 and 31%, respectively). Bacteriocins from Proteobacteria displayed a significantly higher proportion of α-helix (49%) and disordered structures (35%; p = 0.00). Overall, the most abundant peptides were those forming α-helices, with 33% on average (p = 0.00; Figure 2C). Finally, the principal component analysis revealed that bacteriocin classes were not homogeneous (Figure 3). Indeed, even within a class, we observed that bacteriocins displayed different features, except for the length criterion, which was the lowest for class I and the highest for class III.
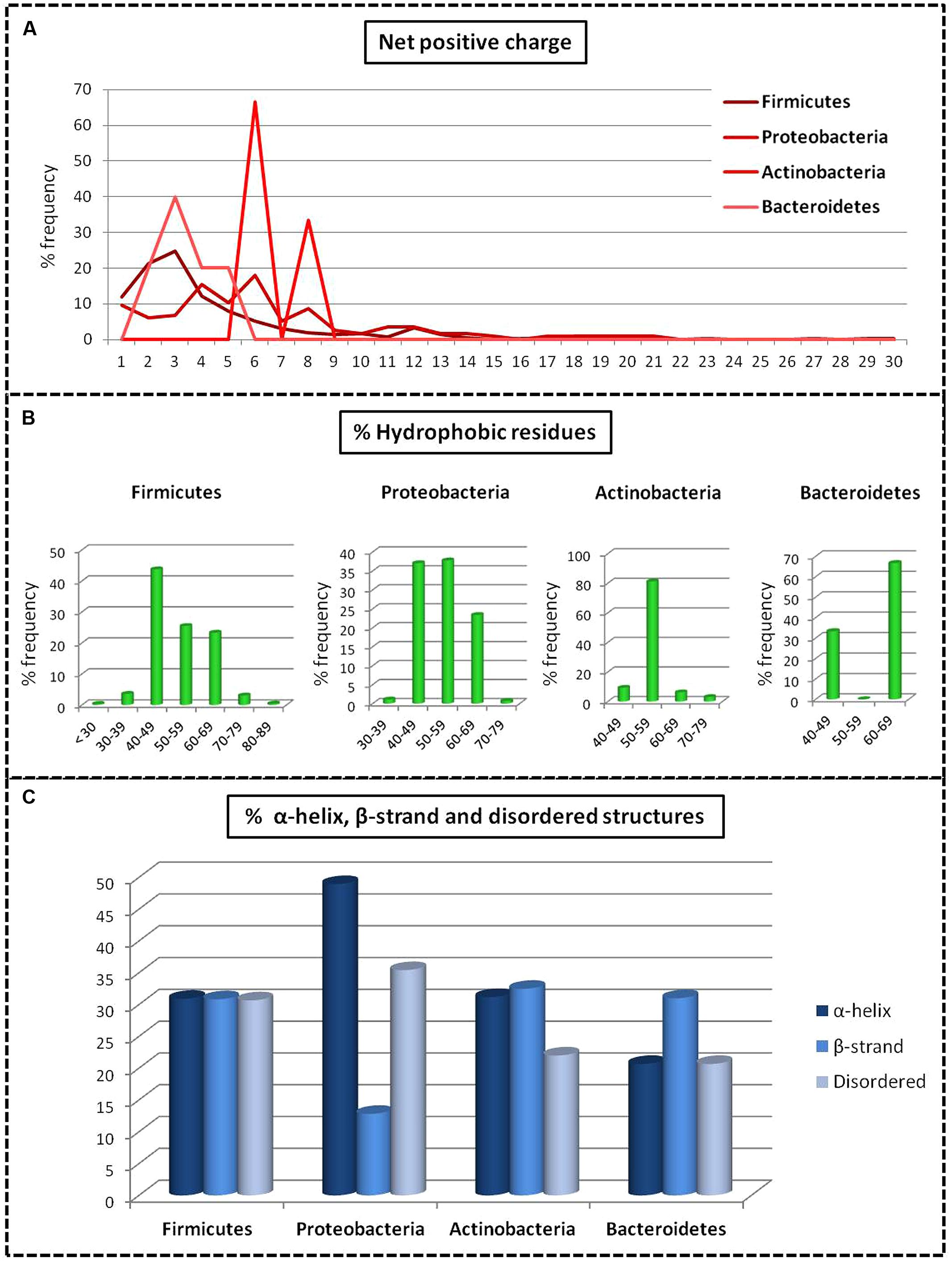
FIGURE 2. Comparative analysis of the structures of bacteriocins from gut bacteria produced by the main phyla. (A) Net positive charge. (B) Composition of hydrophobic residues. (C) Structural statistics (representation of the mean data).
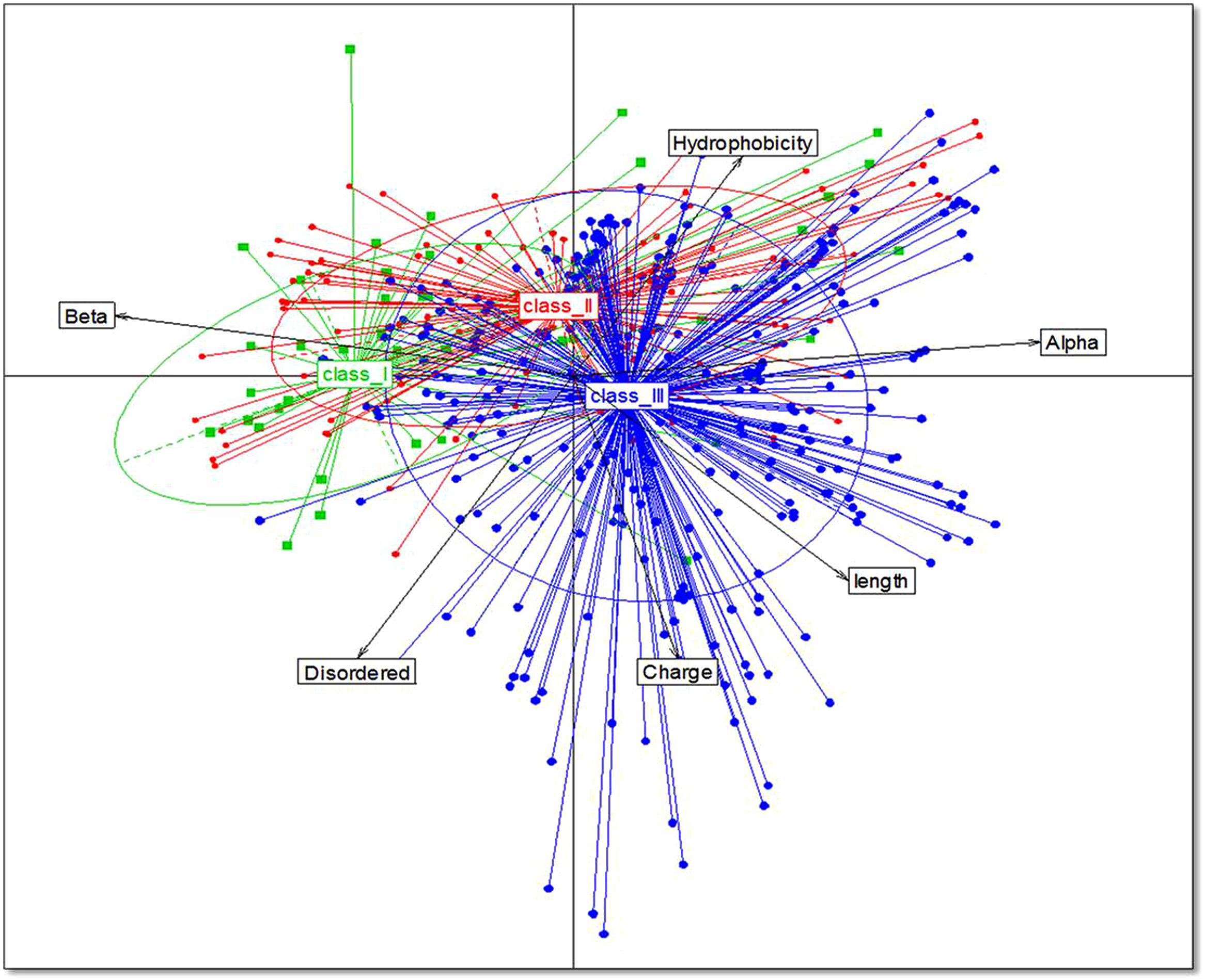
FIGURE 3. Relationships between bacteriocins from the different classes. Data from 1,434 bacteriocins are projected on the first two principal component analysis (PCA) axes.
Discussion
The present study led to the creation of the largest collection of bacteriocin sequences to date and revealed the abundance of bacteriocins in gastrointestinal tract genomes. These findings help to shed light on the underlying factors that shape the microbial composition of the gut. Indeed, in the highly competitive microbial ecosystem, bacteria seem to foray over each other, producing many signals and substances including bacteriocins (Lahti et al., 2014).
When comparing bacteriocins produced by bacteria found in the gut with other bacteriocins, we showed that they have different amino acid composition (higher levels of lysine and methionine) and also have smaller length, significantly less positive charge, and significantly less hydrophobic residues. However, the α-helical structure was consistent across all bacteriocins. The α-helical peptides are known to be essential for antimicrobial activity (Giangaspero et al., 2001) and have been found to have the broadest spectrum of activity (Dathe and Wieprecht, 1999). Physico-chemical parameters, including size, residue arrangement, charge and hydrophobicity can influence bacteriocin potency and spectrum of activity of α-helical peptides (Giangaspero et al., 2001; Zelezetsky and Tossi, 2006). It has been shown that increasing the charge or hydrophobicity arrangement of a peptide results in potent, broad-range antimicrobial activity (Giangaspero et al., 2001; Zelezetsky and Tossi, 2006).
We found that the Firmicutes and Bacteroidetes, which are some of the most predominant bacterial phyla in the human gut (Eckburg et al., 2005), produce the largest number of bacteriocins. Bacteriocins seem to offer the bacteria within the intestinal tract a competitive advantage (Schuijt et al., 2013). However, these bacteriocins seem to have low antibiotic activity, which may result in moderate control, allowing for a substantial quantity of other bacteria to develop and maintain a fairly balanced diversity within the gut. In contrast, bacteriocins produced by Proteobacteria seem to have higher potential antibacterial activity thanks to cationic charges and α-helices. The relative abundance of disordered structures in Proteobacteria enhances their capacity for interaction depending on the medium and thus ensures potent antibiotic activity in different environments (Zelezetsky and Tossi, 2006). Overall, bacteria from the gut seem to produce a large number of bacteriocins with low activity and small amounts of highly effective bacteriocins. Bacteriocins participate in the establishment of an equilibrium of strength between the predominant commensal bacteria of the microflora and potentially pathogenic opportunistic bacteria.
The majority of gut bacteria were found to produce class I bacteriocins, whereas bacteria not found in the gut were more likely to produce class II bacteriocins. However, we found that bacteriocins displayed different features, even within a class. Considering the relationship between the structure and antimicrobial activity, this finding likely reflects the fact that bacteriocin activity patterns differ considerably within classes. It has been claimed that there is no universal consensus on a method for subdividing the classes of bacteriocins (Ennahar et al., 2000; Reddy et al., 2004). Our findings confirm that classifying bacteriocins into existing classes seems to be uninformative. Indeed, the primary structure of bacteriocins, when there is an appropriate balance between hydrophobicity and net positive charge renders the bacteriocins active toward bacteria. However, selective activity also depends on other parameters; hence, the secondary and tertiary structures of bacteriocins appear to be important for the insertion of peptides through the outer layer of the bacteria and into the phospholipid membrane, due to their contribution to the oligomeric state and volume of the antimicrobial peptides. Overall, the secondary and tertiary criteria may be useful for classifying bacteriocins, for characterizing novel bacteriocins, or for designing novel peptides with high potency against pathogens or with broad antimicrobial spectra.
Conclusion
Our genome-mining approach allowed the description of the overall production of bacteriocins in the gut microbiota among which Firmicutes that account for a high proportion of the overall population, represent a major group of bacteriocin producers. The physico-chemical characteristics of these bacteriocins predict a low antibiotic activity that may intervene in intestinal host defense against harmful organisms. More investigations of the in vitro efficacy, will accurately test for the antibiotic potential and help to elucidate the role played by bacteriocins in the modeling of gut microbial composition.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors would like to thank Dr. Lucile Pinault for her help with the structural analysis. VM was supported by a Chairs of Excellence program from the Centre National de Recherche Scientifique (CNRS). This study was funded by the Mediterranee Infection Foundation.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2015.00441/abstract
Abbreviations
BLAST, Basic Local Alignment Search Tool; BUR, bacteriocins of the URMITE; NCBI, National Center for Biotechnology Information; URMITE, Unité de Recherche sur lesMaladies Infectieuses et Tropicales Émergentes.
Footnotes
References
Angelakis, E., Merhej, V., and Raoult, D. (2013). Related actions of probiotics and antibiotics on gut microbiota and weight modification. Lancet Infect. Dis. 13, 889–99. doi: 10.1016/S1473-3099(13)70179-8
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Corr, S. C., Li, Y., Riedel, C. U., O’Toole, P. W., Hill, C., and Gahan, C. G. (2007). Bacteriocin production as a mechanism for the antiinfective activity of Lactobacillus salivarius UCC118. Proc. Natl. Acad. Sci. U.S.A. 104, 7617–7621. doi: 10.1073/pnas.0700440104
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Cotter, P. D., Hill, C., and Ross, R. P. (2005). Bacteriocins: developing innate immunity for food. Nat. Rev. Microbiol. 3, 777–788. doi: 10.1038/nrmicro1273
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Cotter, P. D., Ross, R. P., and Hill, C. (2013). Bacteriocins – a viable alternative to antibiotics? Nat. Rev. Microbiol. 11, 95–105. doi: 10.1038/nrmicro2937
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Dathe, M., and Wieprecht, T. (1999). Structural features of helical antimicrobial peptides: their potential to modulate activity on model membranes and biological cells. Biochim. Biophys. Acta 1462, 71–87. doi: 10.1016/S0005-2736(99)00201-1
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
de Jong, A., van Hijum, S. A., Bijlsma, J. J., Kok, J., and Kuipers, O. P. (2006). BAGEL: a web-based bacteriocin genome mining tool. Nucleic Acids Res. 34, W273–W279. doi: 10.1093/nar/gkl237
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Dobson, A., Cotter, P. D., Ross, R. P., and Hill, C. (2012). Bacteriocin production: a probiotic trait? Appl. Environ. Microbiol. 78, 1–6. doi: 10.1128/AEM.05576-11
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Eckburg, P. B., Bik, E. M., Bernstein, C. N., Purdom, E., Dethlefsen, L., Sargent, M., et al. (2005). Diversity of the human intestinal microbial flora. Science 308, 1635–1638. doi: 10.1126/science.1110591
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Ennahar, S., Sashihara, T., Sonomoto, K., and Ishizaki, A. (2000). Class IIa bacteriocins: biosynthesis, structure and activity. FEMS Microbiol. Rev. 24, 85–106. doi: 10.1111/j.1574-6976.2000.tb00534.x
Fooks, L. J., and Gibson, G. R. (2002). Probiotics as modulators of the gut flora. Br. J. Nutr. 88(Suppl. 1), S39–S49. doi: 10.1079/BJN2002628
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Giangaspero, A., Sandri, L., and Tossi, A. (2001). Amphipathic alpha helical antimicrobial peptides. Eur. J. Biochem. 268, 5589–5600. doi: 10.1046/j.1432-1033.2001.02494.x
Gillor, O., Etzion, A., and Riley, M. A. (2008). The dual role of bacteriocins as antiand probiotics. Appl. Microbiol. Biotechnol. 81, 591–606. doi: 10.1007/s00253-008-1726-5
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Gratia, A. (1925). Sur un remarquable exemple d’antagonisme entre deux souches de colibacille. C. R. Soc. Biol. 93, 1040–1041.
Hammami, R., Zouhir, A., Ben Hamida, J., and Fliss, I. (2007). BACTIBASE: a new web-accessible database for bacteriocin characterization. BMC Microbiol. 7:89. doi: 10.1186/1471-2180-7-89
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Hammami, R., Zouhir, A., Le Lay, C., Ben Hamida, J., and Fliss, I. (2010). BACTIBASE second release: a database and tool platform for bacteriocin characterization. BMC Microbiol. 10:22. doi: 10.1186/1471-2180-10-22
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Hechard, Y., and Sahl, H. G. (2002). Mode of action of modified and unmodified bacteriocins from Gram-positive bacteria. Biochimie 84, 545–557. doi: 10.1016/S0300-9084(02)01417-7
Kelley, L. A., and Sternberg, M. J. (2009). Protein structure prediction on the Web: a case study using the Phyre server. Nat. Protoc. 4, 363–371. doi: 10.1038/nprot.2009.2
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kemperman, R., Kuipers, A., Karsens, H., Nauta, A., Kuipers, O., and Kok, J. (2003). Identification and characterization of two novel clostridial bacteriocins, circularin A and closticin 574. Appl. Environ. Microbiol. 69, 1589–1597. doi: 10.1128/AEM.69.3.1589-1597.2003
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kirkup, B. Jr. (2006). Bacteriocins as oral and gastrointestinal antibiotics: theoretical considerations, applied research, and practical applications. Curr. Med. Chem. 13, 3335–3350. doi: 10.2174/092986706778773068
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Klaenhammer, T. R. (1993). Genetics of bacteriocins produced by lactic acid bacteria. FEMS Microbiol. Rev. 12, 39–86. doi: 10.1111/j.1574-6976.1993.tb00012.x
Lahti, L., Salojarvi, J., Salonen, A., Scheffer, M., and de Vos, W. M. (2014). Tipping elements in the human intestinal ecosystem. Nat. Commun. 5, 4344. doi: 10.1038/ncomms5344
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Million, M., Angelakis, E., Drissi, F., and Raoult, D. (2013). Occam’s razor and probiotics activity on Listeria monocytogenes. Proc. Natl. Acad. Sci. U.S.A. 110:E1. doi: 10.1073/pnas.1218418110
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
O’Shea, E. F., Cotter, P. D., Stanton, C., Ross, R. P., and Hill, C. (2012). Production of bioactive substances by intestinal bacteria as a basis for explaining probiotic mechanisms: bacteriocins and conjugated linoleic acid. Int. J. Food Microbiol. 152, 189–205. doi: 10.1016/j.ijfoodmicro
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Reddy, K. V., Yedery, R. D., and Aranha, C. (2004). Antimicrobial peptides: premises and promises. Int. J. Antimicrob. Agents 24, 536–547. doi: 10.1016/j.ijantimicag.2004.09.005
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Schuijt, T. J., van der Poll, T., de Vos, W. M., and Wiersinga, W. J. (2013). The intestinal microbiota and host immune interactions in the critically ill. Trends Microbiol. 21, 221–229. doi: 10.1016/j.tim.2013.02.001
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Snelling, A. M. (2005). Effects of probiotics on the gastrointestinal tract. Curr. Opin. Infect. Dis. 18, 420–426. doi: 10.1097/01.qco.0000182103.32504.e3
van Belkum, M. J., and Stiles, M. E. (2000). Nonlantibiotic antibacterial peptides from lactic acid bacteria. Nat. Prod. Rep. 17, 323–335. doi: 10.1039/a801347k
van Heel, A. J., de Jong, A., Montalbán-López, M., Kok, J., and Kuipers, O. P. (2013). BAGEL3: automated identification of genes encoding bacteriocins and (non-) bactericidal posttranslationally modified peptides. Nucleic Acids Res. 41, W448–W453. doi: 10.1093/nar/gkt391
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Whisstock, J. C., and Lesk, A. (2003). Prediction of protein function from protein sequence and structure. Q. Rev. Biophys. 36, 307–340. doi: 10.1017/S0033583503003901
Yang, S. C., Lin, C. H., Sung, C. T., and Fang, J. (2014). Antibacterial activities of bacteriocins: application in foods and pharmaceuticals. Front. Microbiol. 5:241. doi: 10.3389/fmicb.2014.00241
Zacharof, M., and Lovitt, R. (2012). Bacteriocins produced by lactic acid bacteria. APCBEE Procedia 2, 50–56. doi: 10.1016/j.apcbee.2012.06.010
Zelezetsky, I., and Tossi, A. (2006). Alpha-helical antimicrobial peptides–using a sequence template to guide structure-activity relationship studies. Biochim. Biophys. Acta 1758, 1436–1449. doi: 10.1016/j.bbamem.2006.03.021
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Keywords: bacteriocin, antimicrobial peptides, microbiota, gastrointestinal tract, BUR database
Citation: Drissi F, Buffet S, Raoult D and Merhej V (2015) Common occurrence of antibacterial agents in human intestinal microbiota. Front. Microbiol. 6:441. doi: 10.3389/fmicb.2015.00441
Received: 26 February 2015; Paper pending published 22 March 2015;
Accepted: 24 April 2015 Published online: 07 May 2015.
Edited by:
Eric Altermann, AgResearch Ltd., New ZealandReviewed by:
M. Andrea Azcarate-Peril, University of North Carolina, Chapel Hill, USAPaula Teixeira, Universidade Católica Portuguesa, Portugal
Giovanna Suzzi, Università degli Studi di Teramo, Italy
Copyright © 2015 Drissi, Buffet, Raoult and Merhej. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Vicky Merhej, Unité de Recherche sur les Maladies Infectieuses et Tropicales Emergentes, UM 63, CNRS 7278, IRD 198, Inserm 1095, Faculté de Médecine, Aix Marseille Université, 27 Boulevard Jean Moulin, 13385 Marseille Cedex 05, France,dmlja3lfbWVyaGVqQGhvdG1haWwuY29t
 Fatima Drissi
Fatima Drissi Sylvain Buffet
Sylvain Buffet Didier Raoult
Didier Raoult Vicky Merhej
Vicky Merhej