- 1Department of Biotechnology, Lovely Professional University, Phagwara, Punjab, India
- 2Department of Bio and Nano technology, Guru Jambheshwar University of Science and Technology, Hisar, India
- 3International Center for Agriculture Research in the Dry Areas (ICARDA), Morocco
Wheat accounts for 19% of the total production of major cereal crops in the world. In view of ever increasing population and demand for global food production, there is an imperative need of 40–60% increase in wheat production to meet the requirement of developing world in coming 40 years. However, both biotic and abiotic stresses are major hurdles for attaining the goal. Among the most important diseases in wheat, fungal diseases pose serious threat for widening the gap between actual and attainable yield. Fungal disease management, mainly, depends on the pathogen detection, genetic and pathological variability in population, development of resistant cultivars and deployment of effective resistant genes in different epidemiological regions. Wheat protection and breeding of resistant cultivars using conventional methods are time-consuming, intricate and slow processes. Molecular markers offer an excellent alternative in development of improved disease resistant cultivars that would lead to increase in crop yield. They are employed for tagging the important disease resistance genes and provide valuable assistance in increasing selection efficiency for valuable traits via marker assisted selection (MAS). Plant breeding strategies with known molecular markers for resistance and functional genomics enable a breeder for developing resistant cultivars of wheat against different fungal diseases.
Introduction
Wheat is a major staple food for mankind in many parts of the world with 714 million tons produced during 2013 (http://www.agri-outlook.org). It is cultivated on 15.4% of the arable land in the world in almost all countries, except the humid and high-temperature areas in the tropics and high-latitude environments. Accounting for a fifth of humanity’s food, wheat is the second only to rice which provides 21% of the food calories and 20% of the protein for more than 4.5 billion people in 94 developing countries (Braun et al., 2010). It contributes 30% of the world’s edible dry matter and 60% of the daily calorie intake in several developing countries (FAOSTAT, 2015). Wheat is produced for a wide range of end-users and it is a critical staple food for a large proportion of the world’s poor farmers and consumers. Due to consistent increase in the world population, there is a need of 60% increase in wheat production to meet the requirement of developing world till 2050 (Singh and Trethowan, 2007; Singh et al., 2007; Rosegrant and Agcaoili, 2010).
Increasing wheat yield potential in the developing world is a primary aim for food security concern (Duveiller et al., 2007). Today, the most challenging task for wheat breeders is to increase grain yield as well as to improve the grain quality of crop for end products (Goutam et al., 2013). These two aspects must be cope up with the strategies employed for enhancing the tolerance against biotic (Keller et al., 2008; Todorovska et al., 2009) and abiotic stresses (Kamal et al., 2010) in addition to the enhanced capability to adapt to various climate changes (Olmstead and Rhode, 2011). Amongst the most important diseases in wheat (derived from fungi, virus, and bacteria), rust diseases (leaf, stem, and stripe) caused by fungus, powdery mildew and Karnal bunt have been reported to produce devastating consequences on wheat quality and production (Keller et al., 2008; Goyal and Prasad, 2010). Cereal rust fungi are highly variable for virulence and molecular polymorphism. Leaf rust, caused by Puccinia triticina is the most common rust of wheat on a worldwide basis (Kolmer, 2013). Leaf rust has potential to cause losses of up to 50% and because of its more frequent and widespread occurrence, leaf rust probably results in greater total annual losses worldwide than stem and stripe rusts (Huerta-Espino et al., 2011). However, management of fungal diseases using conventional plant protection and breeding strategies is quite easy and effective tool, but, it results into different types of environmental pollutions as it involves the use of various eco-hazardous chemicals. Identification and selection of resistant genes through breeding practices is also time-consuming and slow process. Moreover, disease management by host resistance, employment of stable diseases resistance and development of homozygous and resistant cultivars are also time consuming methods (Sharma, 2003; Keller et al., 2008).
To overcome these problems, molecular marker technology is the novel genetic tool for developing high yielding disease resistant cultivars (Landjeva et al., 2007; Varshney et al., 2007). Molecular markers could tag the presence of important resistance genes and allow breeders to identify the resistance genes rapidly and accurately. They also provide significant assistance for increasing selection efficiency through indirect selection for valuable traits via marker assisted selection (MAS). Thus, MAS offers a potential tool for assisting conventional plant breeding approaches to select phenotypic traits for screening disease resistant crop plants (Todorovska et al., 2009). Therefore, existing plant breeding techniques along with available molecular markers (Gupta et al., 2010) and functional genomic tools (Gupta et al., 2008) can help a breeder for developing superior wheat cultivars resistant against fungal diseases in order to minimize yield losses (Goyal and Prasad, 2010). Different types of markers such as random DNA markers, gene targeted markers (Gupta et al., 2010) and functional markers (Liu et al., 2012) have been reported for facilitating identification of genes responsible for individual traits and for improving potential of using MAS in wheat breeding programs (Gupta et al., 2008). DNA-based molecular markers like RFLP (Hartl et al., 1993; Ma et al., 1993, 1994; Autrique et al., 1995; Paull et al., 1995; Nelson et al., 1997), RAPD (Penner et al., 1995; Procunier et al., 1995; Demeke et al., 1996; Qi et al., 1996; Dweikat et al., 1997; Dubcovsky et al., 1998; Shi et al., 1998), STS (Schachermayr et al., 1994, 1995, 1997; Feuillet et al., 1995; Dedryver et al., 1996; Naik et al., 1998; Prins et al., 2001), SSR (Peng et al., 2000; Raupp et al., 2001; Wang et al., 2002), CAPS (Helguera et al., 2000, 2003), AFLP (Hartl et al., 1998), and SCAR (Gold et al., 1999; Liu et al., 1999) have been commonly used for the molecular characterization of plant pathogen and mapping of disease resistance genes in wheat. The development of plant gene transfer systems enable us for the introgression of foreign genes into plant genomes for novel disease control strategies, thus providing a mechanism for broadening the genetic resources available to plant breeders (Zhu et al., 2012).
Fungal Diseases of Wheat
Worldwide, wheat diseases caused by fungal pathogens are more threatening for crop yields and grain quality than those caused by bacteria and viruses. Since, the fungal pathogens are very adaptable and can rapidly evolve into new strains that can infect earlier disease resistant plants. Infection of wheat fungal diseases are influenced by various factors viz., nature of pathogen, susceptibility of host, diversity of virulence, density of inoculums and temperature (Rajaram and Van Ginkel, 1996; McIntosh et al., 1998). The most important fungal diseases in wheat include different types of rust, powdery mildew and Karnal bunt.
Key concepts
(1) DNA marker
It is a gene or DNA sequence with a known location on a chromosome that can be used to identify individuals or species. A genetic marker may be a short DNA sequence, such as a sequence surrounding a single base-pair change (single nucleotide polymorphism, SNP), or a long one, like minisatellites.
(2) Fungal disease
An abnormal growth and/or dysfunction of a plant caused by fungi, which disturbs the normal life process of the plant.
(3) Marker assisted selection (MAS)
MAS is a process whereby a marker (morphological, biochemical or one based on DNA/RNA variation) is used for indirect selection of a genetic determinant or determinants of a trait of interest (e.g., productivity, disease resistance, abiotic stress tolerance, and quality).
(4) Wheat rust
Wheat rust is a destructive disease of wheat caused by fungus genus Puccinia, especially a destructive stem rust characterized by reddish blisters that turn black at the end of the growing season.
Wheat Rust
Wheat rust pathogens belong to genus Puccinia, family Pucciniaceae, order Uredinales and class Basidiomycetes. The rust diseases of wheat such as leaf rust, stem rust, and stripe rust have historically been among the major biotic constraints in the world (Saari and Prescott, 1985; Todorovska et al., 2009). The rusts of wheat is caused by fungal pathogens that can be disseminated thousands of kilometers by wind and are capable of causing considerable economic loss throughout the world (Kolmer, 2005; Goyal and Prasad, 2010). The importance of genetic resistance for the control of rust diseases was demonstrated by Biffen (1905). A prerequisite for developing cultivars with long term rust resistance is the availability of diverse resistance genes.
Leaf Rust
Leaf rust, also known as brown rust, is caused by fungus P. triticina Rob. Ex Desm. f. sp. tritici Eriks (syn. P. recondita). It is a wheat disease of major historical and economic importance. Leaf rust is the most prevalent amongst all the wheat rust diseases occurring around nearly in all wheat grown areas (Kolmer, 2005; Huerta-Espino et al., 2011; Vanzetti et al., 2011). Therefore, it is considered as a widespread and commonly occurring rust disease of wheat. The disease has caused serious epidemics in wheat growing regions of USA (Appel et al., 2009), North Western Mexico (Dubin and Torres, 1981; Singh, 1991; Singh et al., 2004), South America (German et al., 2004), Northern Africa (Abdel-Hak et al., 1980; Deghais et al., 1999), Russia (Volkova et al., 2009), India (Joshi et al., 1975; Nagarajan and Joshi, 1978), Pakistan (Hassan et al., 1973; Hussain et al., 1980), Australia (Watson and Luig, 1961; Keed and White, 1971; Rees and Platz, 1975; Murray and Brennan, 2009), South Africa (Terefe et al., 2009) and other parts of the world. Leaf rust is generally localized on the leaves, but occasionally affects the glumes and awns. Symptoms include circular or oval, orange pustules (urediniospores) on the upper surface of infected leaves. Later on, these pustules become darker due to the formation of black telliospores (Roberson and Luttrell, 1987). The loss in yield depends on several factors such as time of initial infection, crop development stages and relative resistance or susceptibility of the wheat cultivars. Higher yield losses materialized if the initial infection occurs early in the growing season before tillering. However, infection occurred after heading when grain filling is in progress, will cause lesser crop loss (Agrios, 1997). Wheat yield losses are caused due reduction in number of kernels per spike, and kernel weight. Depending on the severity and duration of infection, the losses can vary up to 50% in susceptible wheat cultivars (Knott, 1989; McIntosh et al., 1995).
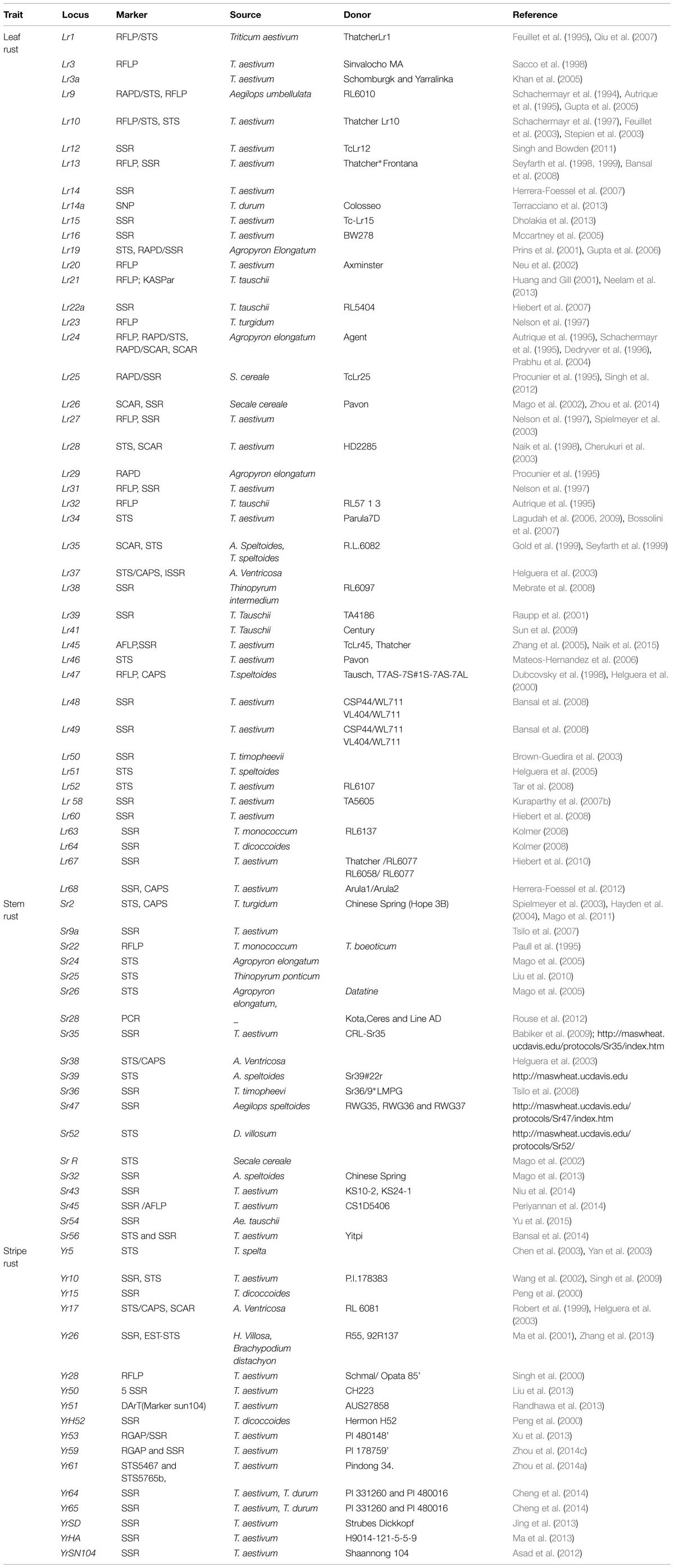
More than 60 leaf rust-resistance (Lr) genes have been identified in common wheat, durum wheat and diploid wheat species (McIntosh et al., 1995, 2008; Bansal et al., 2008; Chhuneja et al., 2008; Vida et al., 2009). Majority of the genes have been identified in the wild wheat relative Aegilops tauschii (Rowland and Kerber, 1974; Kerber, 1987; Gill et al., 1991; Cox et al., 1994; Huang and Gill, 2001; Raupp et al., 2001; Huang et al., 2003; Hiebert et al., 2007). Breeding for leaf rust resistance in wheat is the most challenging task for a breeder because resistance can be completely defeated by a shift in predominant pathogen race in a rust population. Therefore, use of genetic resistance is the comparatively promising option to combat rust epidemics in crop plants. Genetic resistance has two dimensions; one is monitoring dynamic changes of rust pathogen populations to identify new virulent races, and second is deploying resistance genes to defeat the new pathogen race. Molecular markers viz., RFLP, RAPD, STS, SCAR, CAPS, and SSR proves to be the best alternative for screening against leaf rust resistance (William et al., 2008). A wide range of markers are reported to be associated with Lr genes (Table 1). RFLP (Lr13-Seyfarth et al., 2000; Lr20-Neu et al., 2002; Lr21-Huang and Gill, 2001; Lr23, Lr27-Nelson et al., 1997; Lr24, Lr32-Autrique et al., 1995) and RAPD (Lr25, Lr29-Procunier et al., 1995) have been used to tag a variety of Lr genes in wheat. Moreover, the conversion of RFLPs and RAPDs into STS (Schachermayr et al., 1994, 1995, 1997; Feuillet et al., 1995; Helguera et al., 2005) or SCARs (Dedryver et al., 1996) provided a range of useful markers for Lr genes. STS or SCARs are the preferred DNA markers over RFLP, RAPD and AFLP. Lr1 (Feuillet et al., 1995), Lr9, Lr10 (Schachermayr et al., 1994, 1995, 1997), Lr19 (Prins et al., 2001; Cherukuri et al., 2003), Lr24 (Schachermayr et al., 1995; Dedryver et al., 1996), Lr28 (Naik et al., 1998), Lr35 (Gold et al., 1999; Seyfarth et al., 1999), LrX (Obert et al., 2005), Lr51 (Helguera et al., 2005) and Lr 26 (Zhou et al., 2014) are the different STS or SCAR markers associated to Lr genes. Lr67 (Hiebert et al., 2010) and Lr68 (Herrera-Foessel et al., 2012) are SSR linked Lr genes. A gene TaHIR3 has been characterized which encodes a hypersensitive-induced reaction (HIR) protein in response to pathogen attacks. Its expression profile at the DNA and protein levels suggested that TaHIR3 and its deduced protein play a significant role in wheat hypersensitive response caused by leaf rust pathogen (Yu et al., 2013). Validation of markers linked to resistance genes was done successfully in wheat germplasm worldwide. The 287 BC2F4 population of Hungarian wheat genotypes ‘Mv Emma’∗3/‘R.L.6010’ was tested for the presence of Lr (Lr9, Lr24, Lr 25, Lr 29, Lr35, and Lr37) genes. SCAR markers were used for screening of Lr24, Lr 25, and Lr 37 genes (Robert et al., 1999), whereas, STS and RAPD markers were used to validate the presence of Lr 9, Lr 35, and Lr 29, respectively (Vida et al., 2009). Prabhu et al. (2003) used RAPD and SSR marker to study presence of Lr 32 and Lr 28, respectively, in 10 elite near-isogenic lines (NILs) of Indian bread wheat genotypes. To identify the resistance genes in 23 hexaploid Russian spring wheat, STS markers linked to the known leaf rust resistance genes Lr1, Lr9, Lr10, Lr21, Lr24, Lr28, Lr35, Lr37, and Lr39 were used (Gajnullin et al., 2007). Gene-specific markers to the seedling resistance genes (Lr1, Lr10, and Lr21) and Adult plant resistance gene (Lr34) were utilized for molecular screening of 275 wheat accessions from 42 countries (Dakouri et al., 2013). Imbaby et al. (2014) conducted study to identify Lr13, Lr19, Lr24, Lr26, Lr34, Lr35, Lr36, Lr37, Lr39, and Lr46 in 15 Egyptian wheat cultivars using various types of molecular markers.
Cloning of resistance genes is an important approach for providing molecular insights and increasing resistance durability against rust resistance (Ellis et al., 2014; Jonathan et al., 2014). Lawrence et al. (1995) cloned first rust resistance gene L6 from flax (linseed). In case of cereal, Rp1-d was the first rust resistance gene to be cloned by Collins et al. (1999) from corn. More than 30 resistance genes have been cloned in common wheat including Lr10, Lr1, Lr21 for leaf rust (Huang et al., 2003; Cloutier et al., 2007; Loutre et al., 2009; Liu et al., 2012). The resistance genes are ineffective individually to the upcoming pathotypes of rusts in the world, thus pyramiding different resistance genes to breed multiline cultivars may increase the durability of resistance (Wen et al., 2008). Two highly effective genes for leaf rust resistance viz., Lr24, Lr28 and a stripe rust resistance gene Yr15 were selected for pyramiding in the susceptible but high yielding Indian bread wheat variety HD2877 (Revathi et al., 2010). Three highly effective leaf rust resistance genes, Lr 24, Lr 28, and Lr 9 were selected for pyramiding in the bread wheat variety HD 2329 of India (Charpe et al., 2012). Vanzetti et al. (2011) reported that combinations of Lr16, Lr47, Lr19, Lr41, Lr21, Lr25, and Lr29, with Lr34, SV2, Lr46 provide durable and effective resistance to leaf rust. An alternative and efficient strategy to detect quantitative trait loci (QTL) is association mapping (AM) or linkage disequilibrium (LD)-based mapping, in which genotype–phenotype relationships are explored in genetically diverse germplasm (Flint-Garcia et al., 2003; Zhu et al., 2008). AM has proved to be an efficient approach for both tetraploid and hexaploid wheat, by which enhancing previously available QTL information for MAS (Breseghello and Sorrells, 2006; Maccaferri et al., 2011). For leaf rust, QTLs were identified in 164 elite durum wheat accessions from different countries using AM approach (Maccaferri et al., 2010).
Stem Rust
Stem or black rust is a major disease caused by fungus P. graminis f. sp. tritici. Wheat, durum wheat, barley, triticale, barley grasses (Hordeum sp.) and common wheat grass (Agropyron scabrum) are among the most commonly infected crops by stem rust. The Italians Fontana and Tozzetti independently provided the first report on stem rust in wheat in 1767. In large areas of the world, the life cycle of P. graminis consists of continual uredinial generations. The disease either spreads via airborne spores or occasionally from local-wild susceptible barberry (Berberis sp.) plants (Eversmeyer, 2000). Wheat (primary host) and barberry (secondary host) are required to complete the life cycle of fungus (Leonard and Szabo, 2005). Five types of spores (pycniospores, aeciospores, urediniospores, teliospores, and basidiospores) occur in the life cycle of fungus at different developmental stages (Leonard, 2001). Warm temperature (15–30°C) and dew are the two important factors favoring the crop infection by stem rust. Stem rust usually occurs on the stem, and can also occur on the leaves (both sides), leaf sheaths or in severe infections on the head. Uredia pustules on stem and leaf sheaths are the main symptoms of disease spreading (Leonard, 2001). Reddish brown color and oval or spindle-shaped pustules are seen on the stem and leaf sheath. Pustules would change to black in color at the end of the season when infection is too old (Todorovska et al., 2009) and can cause severe crop loss in a short span of time at the end of the season.
In the early to mid 1950s; stem rust epidemics caused approximately 50% yield losses of wheat in North America (Leonard, 2001). During 1950s, Norman Borlaug and other scientists started developing high-yielding wheat varieties that were resistant to stem rust and other diseases in North America and throughout the world (Singh et al., 2006). Resistant plants exhibit no or less number of uredia surrounded by chlorosis or necrosis as compared to susceptible plants. A new race of stem rust (Ug99) causing a high level of infection on wheat genotypes was found in 1999 in Uganda (Pretorius et al., 2000). Heavy stem rust infections were observed in International Center for Wheat and Maize Improvement (CIMMYT)-derived lines of wheat in Kenya in 2004 (Kolmer, 2005; Todorovska et al., 2009). This race has spread to major wheat growing regions of the world such as Iran, Afghanistan, India, Pakistan, Turkmenistan, Uzbekista, Kazakhstan, USA, and Canada (Todorovska et al., 2009). Therefore it is necessary to develop a resistant germplasm to overcome the spreading of infection in these regions.
Since, breeding program in wheat for developing stem rust resistance is a challenging task for a breeder; therefore, acquisition of genetic resistance is the best alternative for controlling rust epidemics. Currently, about fifty stem rust resistance (Sr) genes have been identified. Moreover, mapping of few genes and their close relatives on different chromosomes of wheat has also been achieved (McIntosh et al., 1998). PCR (STS) and non-hybridization based (RFLP) markers are available for screening the genotypes which are resistant to stem rust disease (William et al., 2008). The molecular markers associated with Sr genes known so far are summarized in (Table 1). RFLP (Sr22-Paull et al., 1995) and STS (Sr2-Hayden et al., 2004; Sr24, Sr26- Mago et al., 2005; SrR-Mago et al., 2002; Sr39-Mas wheat ucdavis), STS/SSR (Sr56-Bansal et al., 2014), SSR/AFLP (Sr45- Periyannan et al., 2014) STS/CAPS (Sr38-Helguera et al., 2003) and SSR (Sr32- Mago et al., 2013; Sr43-Niu et al., 2014; Sr54- Yu et al., 2015) markers have been reported to be associated with different Sr genes in wheat. Sr2 is one of the non-race specific genes which have resulted in successful acquisition of durable rust resistance to slow rusting adult (Singh et al., 2004). It has been widely used by CIMMYT, Mexico in its wheat program for improvement of stem rust resistance and also in USA for hard winter wheat breeding program. Above all, the Sr2 complex when used in combination with other resistance genes has shown remarkable protection against Ug99 (Singh, 1993). CIMMYT and International center for agricultural research in the dry areas (ICARDA) started the global rust initiative (Later in 2008, BGRI- Borlaug global rust initiative) to coordinate efforts to track and study Ug99 and develop resistant varieties of wheat (Stokstad, 2007). Some genes like Sr33 and Sr35 for stem rust resistance were cloned with the objective to increase resistance (Periyannan et al., 2013; Saintenac et al., 2013) Various studies have been conducted to confirm the presence of Sr genes in wheat cultivars. A recombinant inbred line (RIL) population of 83 lines (developed from a cross from Indian wheat cultivars VL404 and WL711) was screened to identify Sr28 gene using SSR markers (Bansal et al., 2012). Haile et al. (2013) screened 58 tetraploid wheat accessions of Ethiopian wheat cultivars for the presence of 30 Sr genes using SSR and STS markers. 88 spring soft wheat of Kazakhstan were studied for presence of Sr genes (Sr2, Sr22, Sr24, Sr36, and Sr46) which are effective against Ug99 (Kokhmetova and Atishova, 2012). Thirty-seven lines of American cultivars with known stem rust resistance genes and five genetic background cultivars were used to further validate the six co-dominant STS markers for Sr25 and Sr26 (Liu et al., 2010). Mago et al. (2011) used DNA markers to check the presence of Sr24, Sr26, SrR, and Sr31 in wheat-rye recombinant T6-1. These Sr genes provide resistance against all strains of stem rust that are prevalent in Australia. However, Sr26 and SrR are effective outside Australia against strain Ug99. 104 F2:3 population of Gabo 56 with susceptible cultivar Chinese Spring were screened to check the presence of Sr9h using SSR markers. Minor stem rust resistance gene Sr2 was pyramided with two major stem rust resistance genes Sr24 and Sr36 in Indian wheat varieties ‘Lok-1’ and ‘Sonalika’ (Nisha et al., 2015). AM study for response to stem rust was conducted on 183 Ethiopian durum wheat accessions and 276 wheat lines from Kenya (Yu et al., 2011; Letta et al., 2013).
Yellow Rust or Stripe Rust
Stripe or yellow rust, caused by P. striiformis f. sp. tritici, mainly infects wheat, but can also cause infection in barley, rye, and triticale. It was first reported in USA (Carleton, 1915) and outbreaks were reported in the Western states in 1960s (Boyd, 2005). Later on, the infections were also reported from other parts of the of world including USA, East Asia (China north-west and southwest), South Asia (India, Pakistan, and Nepal), Oceania (Australia, New Zealand), East Africa (Ethiopia, Kenya), the Arabian Peninsula (Yemen) and Western Europe (Wellings, 2011). Presently, more than 35% of area under wheat cultivation is affected by stripe rust disease (Singh et al., 2004). Cool and wet weather is favorable for the development of yellow rust. Pustules are light yellow and occur on leaves in distinct straight-sided stripes about 1/16 inches wide and of regular length. The spores are yellow to orange in color. Reduced dry matter production, root growth, plant height, size and number of flowering spikes, and the size and number of grains are the parameters affected by infection. These effects were more pronounced with infection beginning at the seedling stage, although infections initiated at anthesis were also associated with reduced root weight and grain yield (Wellings, 2011).
Breeding efforts for stripe rust resistance has been made in the past. Breeding approaches involves developing several crosses with careful phenotypic selection which makes it difficult for a breeder to achieve the desired objective. About 52 permanently named and more than 40 temporarily designated genes or QTL for stripe rust resistance have been reported (Chen, 2005; McIntosh et al., 2011; Ren et al., 2012). Among the permanently named resistance genes, Yr11, Yr12, Yr13, Yr14, Yr16, Yr18, Yr29, Yr30, Yr34, Yr36, Yr39, Yr46, Yr48, and Yr52, confer adult plant or high temperature adult plant (HTAP) resistance genes, whereas the others confer all-stage resistance. The identification and use of the resistant genes is the only way to conquer the impact of disease on wheat production. Till date, 65 (Yr1–Yr65) yellow rust resistance genes have been characterized and designated in wheat (McIntosh et al., 1995; Singh et al., 2004; Boyd, 2005; McIntosh et al., 2008). A wide range of markers are reported to be associated with Yr genes (Table 1). RFLP (Yr28-Singh et al., 2000), SSR (Yr10-Wang et al., 2002; Yr15, Yr26, YrH52-Peng et al., 2000), STS/CAPS (Y17-Robert et al., 1999; Helguera et al., 2003; YrMoro-Smith et al., 2002), STS (Yr61-Zhou et al., 2014a), DArt (Yr51-Randhawa et al., 2014), RGAP/SSR (Yr59-Zhou et al., 2014c) and SSR (YrSN104-Asad et al., 2012; Yr 50- Liu et al., 2013; Yr64 and Yr65-Cheng et al., 2014) markers have been reported to be associated with different Yr genes in wheat. Most of the identified yellow rust resistant Yr genes have been characterized as the race specific ones and are responsible for acquiring resistance against the isolates of P. striiformis f. sp. tritici only, which carries the corresponding avirulence (avr) gene. Various stripe rust resistant genes have been transferred into hexaploid wheat from different wild species (Kuraparthy et al., 2007a,b; Singh et al., 2007; Chhuneja et al., 2008). With the help of molecular marker a study reveals that recent Canadian wheat varieties have the strip rust resistant genes Yr 10, Yr17, Yr18, and Yr 36 (Randhawa et al., 2012). Further, a highly stripe rust resistant gene, namely Yr36 has been used for positional cloning. Yr36 gene, derived from wild emmer wheat, carries broad spectrum resistance for stripe rust races (Fu et al., 2009). A total of 54 wheat genotypes representing breeding lines and current grown cultivars in the western US were tested with race PST-100 and the Yr53-flanking markers, XLRRrev/NLRRrev350, Xgwm441 and the STS marker (STS2F/1R219) developed from RGAP marker, Ptokin2/Xa1NBSF234 (Xu et al., 2013).
Four Gatersleben wheat microsatellite (GWM) markers were used to identify non-specific adult plant disease resistance genes against stripe rust in 160 F2 plants from the cross of UK/German wheat cultivars Lgst.7/Winzi (Khlestkina et al., 2007). To identify genes for stripe rust in 181 plants from one segregating F3 line of Xiaoyan/Mingxian cross. SSR primers were used to identify molecular markers flanking Yrxy2, whereas for Yrxy1 RGAP and SSR markers both were used (Zhou et al., 2011). Naz et al. (2012) done QTL analysis by using a genetic map based on 118 SSR markers in 150 back cross lines of German wheat cultivars Zentos and Syn86L. To identify genes for stripe rust resistance in 179 F2 population of Wuhan 2/Mingxian 169 cross against races CYR30 and CYR31 using RGAP and SSR markers (Zhou et al., 2014b). Yaniv et al. (2015) concluded from their findings that SSR markers from Yr15 region are efficient tools for MAS and for introgression of Yr15 into wheat from T. dicoccoides. In case of stripe rust resistance genes, Yr17, Yr18, and Yr36 were amongst the successfully cloned genes (Helguera et al., 2003; Lagudah et al., 2009; Fu et al., 2009). Stripe rust response for adult plants was evaluated using AM in 192 genotypes including 181 synthetic hexaploid wheat (SHW) and 11 bread wheat cultivars from different countries (Zegeye et al., 2014). Similar studies were performed using 402 wheat varieties and 1000 spring wheat accessions from USA (Naruoka et al., 2015; Maccaferri et al., 2015).
Recent Trends
Recently the new technologies are being used for sequencing of cereal crops, but the storage of data and analysis are difficult due to its vast size. Single nucleotide polymorphism (SNP) genotyping offers a solution to this problem and accelerates the crop improvement by providing insights into their genetic constitution. It has number of advantages over conventional marker system such as rapid processing of large populations, abundance of markers and varieties of genotyping system (Thomson, 2014). In quantitative trait locus (QTL) mapping experiments and genome-wide association studies (GWAS), SNP data is frequently used to detect marker-trait associations (Zhao et al., 2011; Cook et al., 2012). Discovery of SNPs using complete genome is facilitated by recent advances in next-generation sequencing (Berkman et al., 2012; Chia et al., 2012; Xu et al., 2012). Genetic studies of number of economically important crops have been successfully done by the application of high-density SNP arrays (Wiedmann et al., 2008; Ganal et al., 2011; Zhao et al., 2011; Sim et al., 2012; Song et al., 2013). 44K SNP genotyping chip was employed for GWAS of diverse rice accessions and identified number of alleles responsible for governing morphological and agronomic traits (Zhao et al., 2011). Similarly, the genetic control of maize kernel composition in a nested AM panel was studied by the use of 50K maize SNP chip (Cook et al., 2012; Hufford et al., 2012). Moreover, the genomic regions targeted by breeding in wheat were detected by 9K SNP wheat (Cavanagh et al., 2013). The most challenging task is to analyze the genotypic data of durum [T. turgidum subsp. durum (Desf.) Husnot] and bread wheat (T. aestivum L.) genome using SNP genotyping platforms (Akhunov et al., 2009). The use of wheat SNP iSelect array has proven to be a promising tool to infer detailed haplotype structure in polyploid wheat and will serve as an invaluable resource for diversity studies and investigating the genetic basis of trait variation in wheat. A combination of eight mapping populations was used to genetically map 46,977 SNPs using wheat 90K array (Wang et al., 2014).
Conclusion
Due to global food security and consistent increase in world population, there is an immediate need to increase wheat yield considerably. Fungal diseases continue to cause huge losses and pose a great challenge for wheat production. Novel genetic tools based on molecular marker technologies provide a good alternative for developing improved resistant cultivars. Development of molecular markers such as RFLPs, SSRs, AFLPs, SNPs, and DArT in last more than two decades has revolutionized wheat genomics. Marker assisted breeding and functional genomics tools are effective strategies to develop resistant cultivars against fungal diseases in wheat for achieving estimated production paradigm. In future, functional genomics approaches such as TILLING, RNAi and epigentics etc. are needed to strengthen the development of resistant varieties. Mutagenesis-derived broad-spectrum disease resistance may lead to a better understanding of the regulation of defense response networks in wheat. Large-scale genome sequencing and associated bioinformatics are becoming widely accepted research tools for accelerating the analysis of wheat genome structure and function. Currently, functional markers are being increasingly adopted in wheat breeding. These markers are needed for important traits such as disease and stress resistance in order to strengthen the application of molecular markers in breeding programs. The collaborative effort (MASwheat: http://maswheat.ucdavis.edu/index.htm) by United States Department of Agriculture (USDA), National Institute of Food and Agriculture (NIFA) and Borlaug Global Rust Initiative (BGRI) has given the platform for transferring new developments in wheat genomics and biotechnology to increase wheat production. Many traits such as the disease/pest resistance and end-use quality which has increased the competitiveness of wheat breeding programs through MAS were included. Triticeae Coordinated Agricultural Project (T-CAP) focused on studying the effects of climate change on crop yields by identification and incorporation of genetic loci for enhancing tolerance in crops. For improving the barley and wheat germplasm, gene variants for disease resistance, water and nitrogen use efficiency and yield improvement are being identified, along with molecular markers to tag them and accelerate breeding. The International Wheat Genome Sequencing Consortium (IWGSC) will put the foundation to accelerate wheat improvement for wheat growers, scientists, and breeders. The ultimate goal leads to obtain high quality annotation of the genome and thus complete sequencing of the common wheat genome.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Abdel-Hak, T. M., El-Sherif, N. A., Bassiouny, A. A., Shafik, E. L., and Dauadi, Y. (1980). “Control of wheat leaf rust by systemic fungicides,” in Proceeding 5th European and Mediterranean Cereal rusts Conference, Bari, 255–266.
Agrios, G. N. (ed.) (1997). “Control of plant diseases,” in Plant Pathology, 4th Edn, (London: Academic Press), 635.
Akhunov, E., Nicolet, C., and Dvorak, J. (2009). Single nucleotide polymorphism genotyping in polyploid wheat with the Illumina Golden Gate assay. Theor. App. Genet. 119, 507–517. doi: 10.1007/s00122-009-1059-5
Appel, J. A., DeWolf, E., Bockus, W. W., and Todd, T. (2009). Kansas Cooperative Plant Disease Survey Report. Preliminary Kansas Wheat Disease Loss Estimates. Available at: http://agriculture.ks.gov/docs/default-source/PP-Disease-Reports-2014/2014-ks-wheat-disease-loss-estimates.pdf [accessed August 11, 2009].
Asad, M. A., Xia, X., Wang, C., and He, Z. (2012). Molecular mapping of stripe rust resistance gene YrSN104 in Chinese wheat line Shaannong 104. Hereditas 149, 146–152. doi: 10.1111/j.1601-5223.2012.02261.x
Autrique, E., Singh, R. P., Tanksley, S. D., and Sorrells, M. E. (1995). Molecular markers for four leaf rust resistance genes introgressed into wheat from wild relatives. Genome 38, 75–83. doi: 10.1139/g95-009
Babiker, E., Ibrahim, A. M. H., Yen, Y., and Stein, J. (2009). Identification of a microsatellite marker associated with stem rust resistance gene Sr35 in wheat. Aust. J. Crop Sci. 3, 195–200.
Bansal, U., Bariana, H., Wong, D., Randhawa, M., Wicker, T., Hayden, M., et al. (2014). Molecular mapping of an adult plant stem rust resistance gene Sr56 in winter wheat cultivar Arina. Theor. Appl. Genet. 127, 1441–1448. doi: 10.1007/s00122-014-2311-1
Bansal, U. K., Hayden, M. J., Venkata, B. P., Khanna, R., Saini, R. G., and Bariana, H. S. (2008). Genetic mapping of adult plant leaf rust resistance genes Lr48 and Lr49 in common wheat. Theor. Appl. Genet. 117, 307–312. doi: 10.1007/s00122-008-0775-6
Bansal, U. K., Zwart, R., Bhavani, S., Wanyera, R., Gupta, V., and Bariana, H. S. (2012). Microsatellite mapping identifies TTKST-effective stem rust resistance gene in wheat cultivars VL404 and Janz. Mol. Breed. 30, 1757–1765. doi: 10.1007/s11032-012-9759-y
Berkman, P. J., Lai, K., Lorenc, M. T., and Edwards, D. (2012). Next-generation sequencing applications for wheat crop improvement. Am. J. Bot. 99, 365–371. doi: 10.3732/ajb.1100309
Biffen, R. H. (1905). Mendel’s laws of inheritance and wheat breeding. J. Agric. Sci. 1, 4–48. doi: 10.1017/S0021859600000137
Bossolini, E., Wicker, T., Knobel, P. A., and Keller, B. (2007). Comparison of orthologous loci from small grass genomes Brachypodium and rice: implications for wheat genomics and grass genome annotation. Plant J. 49, 704–717. doi: 10.1111/j.1365-313X.2006.02991.x
Boyd, L. A. (2005). Can Robigus defeat an old enemy? – Yellow rust of wheat. J. Agric. Sci. 143, 233–243. doi: 10.1017/S0021859605005095
Braun, H. J., Atlin, G., and Payne, T. (2010). “Multi-location testing as a tool to identify plant response to global climate change,” in Climate change and Crop Production, ed. C. R. P. Reynolds (London: CABI).
Breseghello, F., and Sorrells, M. E. (2006). Association mapping of kernel size and milling quality in wheat (Triticum aestivum L.) cultivars. Genetics 172, 1165–1177. doi: 10.1534/genetics.105.044586
Brown-Guedira, G. L., Singh, S., and Fritz, K. (2003). Performance and mapping of leaf rust resistance transferred to wheat from Triticum timopheecii subsp. armeniacum. Phytopathology 93, 784–789. doi: 10.1094/PHYTO.2003.93.7.784
Cavanagh, C. R., Chao, S., Wang, S., Huang, B. E., Stephen, S., Kiani, S., et al. (2013). Genome-wide comparative diversity uncovers multiple targets of selection for improvement in hexaploid wheat landraces and cultivars. Proc. Natl Acad. Sci. U.S.A. 110, 8057–8062. doi: 10.1073/pnas.1217133110
Charpe, A., Koul, S., Gupta, S. K., Singh, A., Pallavi, J. K., and Prabhu, K. V. (2012). Marker assisted gene pyramiding of leaf rust resistance genes Lr 9, Lr 24 and Lr 28 in a bread wheat cultivar HD 2329. J. Wheat Res. 4, 20–28.
Chen, X. M. (2005). Epidemiology and control of stripe rust [Puccinia striiformis f. sp. tritici] on wheat. Can. J. Plant Pathol 27, 314–337.
Chen, X., Soria, M. A., Yan, G., Sun, J., and Dubcovsky, J. (2003). Development of sequence tagged site and cleaved amplified polymorphic sequence markers for wheat stripe rust resistance gene Yr5. Crop Sci. 43, 2058–2064. doi: 10.2135/cropsci2003.2058
Cheng, P., Xu, L. S., Wang, M. N., See, D. R., and Chen, X. M. (2014). Molecular mapping of genes Yr64 and Yr65 for stripe rust resistance in hexaploid derivatives of durum wheat accessions PI 331260 and PI 480016. Theor. Appl. Genet. 127, 2267–2277. doi: 10.1007/s00122-014-2378-8
Cherukuri, D. P., Gupta, S. K., Charpe, A., Koul, S., Prabhu, K. V., Singh, R. B., et al. (2003). Identification of a molecular marker linked to an Agropyron elongatum-derived gene Lr19 for leaf rust resistance in wheat. Plant Breed. 122, 204–208. doi: 10.1046/j.1439-0523.2003.00846.x
Chhuneja, P., Kaur, S., Garg, T., Ghai, M., Kaur, S., Prashar, M., et al. (2008). Mapping of adult plant stripe rust resistance genes in diploid: a genome wheat species and their transfer to bread wheat. Theor. Appl. Genet. 116, 313–324. doi: 10.1007/s00122-007-0668-0
Chia, J. M., Song, C., Bradbury, P. J., Costich, D., De-Leon, N., Doebley, J., et al. (2012). Maize HapMap2 identifies extant variation from a genome in flux. Nat. Genet. 44, 803–807. doi: 10.1038/ng.2313
Cloutier, S., McCallum, B. D., Loutre, C., Banks, T. W., Wicker, T., Feuillet, C., et al. (2007). Leaf rust resistance gene Lr1, isolated from bread wheat (Triticum aestivum L.) is a member of the large psr567 gene family. Plant Mol. Biol. 65, 93–106. doi: 10.1007/s11103-007-9201-8
Collins, N., Drake, J., Ayliffe, M., Sun, Q., Ellis, J., Hulbert, S., et al. (1999). Molecular characterization of the maize Rp1- D rust resistance haplotype and its mutants. Plant Cell 11, 1365–1376. doi: 10.2307/3870755
Cook, J. P., McMullen, M. D., Holland, J. B., Tian, F., Bradbury, P., Ross-Ibarra, J., et al. (2012). Genetic architecture of maize kernel composition in the nested association mapping and inbred association panels. Plant Physiol. 158, 824–834. doi: 10.1104/pp.111.185033
Cox, T. S., Raupp, W. J., and Gill, B. (1994). Leaf Rust-Resistance Genes Lr41. Crop Sci. 34, 339–343.
Dakouri, A., Callum, B. D., Radovanovic, N., and Cloutier, S. (2013). Molecular and phenotypic characterization of seedling and adult plant leaf rust resistance in a world wheat collection. Mol. Breed. 32, 663–677. doi: 10.1007/s11032-013-9899-8
Dedryver, F., Jubier, M. F., Thouvenin, J., and Goyeau, H. (1996). Molecular markers linked to the leaf rust resistance gene Lr24 in different wheat cultivars. Genome 39, 830–835. doi: 10.1139/g96-105
Deghais, M., El-Faleh, M., and Gharbi, M. S. (1999). Acquis de la recherche en amelioration des cereales en Tunisie. Ann. INRAT 26, 33–40.
Demeke, T., Laroche, A., and Gaudet, D. A. (1996). A DNA marker for the Bt-10 common bunt resistance gene in wheat. Genome 39, 51–55. doi: 10.1139/g96-007
Dholakia, B. B., Rajwade, A. V., Hosmani, P., Khan, R. R., Chavan, S., Reddy, D. M. R., et al. (2013). Molecular mapping of leaf rust resistance gene Lr15 in hexaploid wheat. Mol. Breed. 31, 743–747. doi: 10.1007/s11032-012-9813-9
Dubcovsky, J., Lukaszewski, A. J., Echaide, M., Antonelli, E. F., and Porter, D. R. (1998). Molecular characterisation of two Triticum speltoides interstitial translocations carrying leaf rust and greenbug resistance genes. Crop Sci. 38, 1655–1660. doi: 10.2135/cropsci1998.0011183X003800060040x
Dubin, H. J., and Torres, E. (1981). Causes and Consequences of the 1976–77 wheat leaf rust epidemic in northwest Mexico. Annual Rev. Phytopathol. 19, 44–49. doi: 10.1146/annurev.py.19.090181.000353
Duveiller, E., Singh, R. P., and Nicol, J. M. (2007). The challenges of maintaining wheat productivity: pests, diseases, and potential epidemics. Euphytica 157, 417–443. doi: 10.1007/s10681-007-9380-z
Dweikat, I., Ohm, H., Patterson, F., and Cambron, S. (1997). Identification of RAPD markers for 11 Hessian fly resistance genes in wheat. Theor. Appl. Genet. 94, 419–423. doi: 10.1007/s001220050431
Ellis, J. G., Lagudah, E. S., Spielmeyer, W., and Dodds, P. N. (2014). The past, present and future of breeding rust resistant wheat. Front. Plant Sci. 5:641. doi: 10.3389/fpls.2014.00641
Eversmeyer, M. G. (2000). Epidemology of wheat leaf and stem rust in the central great plains of the USA. Ann. Rev. Phytopathol. 38, 491–513. doi: 10.1146/annurev.phyto.38.1.491
FAOSTAT. (2015). Organisation Des Nations Unies Pour L’alimentation Et L’agriculture. Avaialbe at: http://faostat.fao.org
Feuillet, C., Messmer, M., Schachermayr, G., and Keller, B. (1995). Genetic and physical characterization of the Lr1 leaf rust resistance locus in wheat (Triticum aestivum L.). Mol. Gen. Genet. 248, 553–562. doi: 10.1007/BF02423451
Feuillet, C., Travella, S., Stein, N., Albar, L., Nublat, A., and Keller, B. (2003). Map-based isolation of the leaf rust disease resistance gene Lr10 from the hexaploid wheat (Triticum aestivum L.) genome. Proc. Natl. Acad. Sci. U.S.A. 100, 15253–15258. doi: 10.1073/pnas.2435133100
Flint-Garcia, S. A., Thornsberry, J. M., and Buckler, E. S. (2003). Structure of linkage disequilibrium in plants. Annu. Rev. Plant Biol. 54, 357–374. doi: 10.1146/annurev.arplant.54.031902.134907
Fu, D. L., Cristobal, U., Assaf, D., Ann, B., Lynn, E., Chen, X. M., et al. (2009). A Kinase-START gene confers temperature dependent resistance to wheat stripe rust. Science 323, 1357–1360. doi: 10.1126/science.1166289
Gajnullin, N. R., Lapochkina, I. F., Zhemchuzhina, A. I., Kiseleva, M. I., Kolomiets, T. M., and Kovalenko, E. D. (2007). Phytopathological and molecular genetic identification of leaf rust resistance genes in common wheat accessions with alien genetic material. Russ. J. Genet. 43, 875–881. doi: 10.1134/S1022795407080078
Ganal, M. W., Durstewitz, G., Polley, A., Bérard, A., Buckler, E. S., Charcosset, A., et al. (2011). A large maize (Zea mays L.) SNP genotyping array: development and germplasm genotyping, and genetic mapping to compare with the B73 reference genome. PLoS ONE 6:e28334. doi: 10.1371/journal.pone.0028334
German, S., Kohli, M., Chaves, M., and Barcellos, A. (2004). “Breakdown of resistance of wheat cultivars and estimated losses caused by recent changes in the leaf rust population in South America,” in Proceeing of the 11th International Cereal Rusts and Powdery Mildews Conference abstract, (Norwich: John Innes Centre), A2.21.
Gill, K. S., Lubbers, E. L., Gill, B. S., Raupp, W. J., and Cox, T. S. (1991). A genetic linkage map of Triticum tauschii (DD) and its relationship to the D genome of bread wheat (AABBDD). Genome 34, 362–374. doi: 10.1139/g91-058
Gold, J., Harder, D., Townley-Smith, F., Aung, T., and Procunier, J. (1999). Development of a molecular marker for rust resistance genes Sr39 and Lr35 in wheat breeding lines. Electron. J. Biotechnol. 2:1. doi: 10.2225/vol2-issue1-fulltext-1
Goutam, U., Kukreja, S., Tiwari, R., Chaudhary, A., Gupta, R. K., Dholakia, B. B., et al. (2013). Biotechnological approaches for grain quality improvement in wheat: present status and future possibilities. Aust. J. Cereal Sci. 7, 469–483.
Goyal, A., and Prasad, R. (2010). “Some important fungal diseases and their impact on wheat production,” in Management of Fungal Plant Pathogens, Vol. 11, eds A. Arya and A. E. V. Perelló (London: CABI), 362.
Gupta, P. K., Landridge, P., and Mir, R. R. (2010). Marker-assisted wheat breeding: present status and future possibilities. Mol. Breed. 26, 145–161.
Gupta, P. K., Mir, R. R., Mohan, A., and Kumar, J. (2008). Wheat genomics: present status and future prospects. Int. J. Plant Genome 2008:896451.
Gupta, S. K., Charpe, A., and Koul, S. (2005). Development and validation of molecular markers linked to an Aegilops umbellulata-derived leaf rust- resistance gene. Genome 48, 823–830.
Gupta, S. K., Charpe, A., Prabhu, K. V., and Haque, Q. M. R. (2006). Identification and validation of molecular markers linked to the leaf rust resistance gene Lr19 in wheat. Theor. Appl. Genet. 113, 1027–1036. doi: 10.1007/s00122-006-0362-7
Haile, J. K., Hammer, K., Badebo, A., Nachit, M. M., and Roder, M. S. (2013). Genetic diversity assessment of Ethiopian tetraploid wheat landraces and improved durum wheat varieties using microsatellites and markers linked with stem rust resistance. Genet. Resour. Crop Evol. 60, 513–527. doi: 10.1007/s10722-012-9880-0
Hartl, L., Mori, S., and Schwizer, G. (1998). “Identification of a diagnostic molecular marker for the powdery mildew resistance gene Pm4b based on fluorescence labelled AFLPs,” in Proceedings 9th International Wheat Genetics Symposium, University of Saskatchewan, Vol. 3, ed. A. E. Slinkard (Saskatoon, SK: Extension Press), 111–113.
Hartl, L., Weiss, H., Zeller, F. J., and Jahoor, A. (1993). Use of RFLP markers for the identification of alleles of the Pm3 locus conferring powdery mildew resistance in wheat (Triticum aestivum L.). Theor. Appl. Genet. 86, 959–963. doi: 10.1007/BF00211048
Hassan, S. F., Hussain, M., and Rizvi, S. A. (1973). “Wheat disease situation in Pakistan,” in Proceedings of the National Farmers and wheat Research Production, Islamabad, 231–234.
Hayden, M. J., Kuchel, H., and Chalmers, K. J. (2004). Sequence tagged microsatellites for the Xgwm533 loci provide new diagnostic markers to select for the presence of stem rust resistance gene Sr2 in bread wheat (Triticum aestivum L). Theor. Appl. Genet. 109, 1641–1647. doi: 10.1007/s00122-004-1787-5
Helguera, M., Khan, I. A., and Dubcovsky, J. (2000). Development of PCR markers for the wheat leaf rust resistance gene Lr47. Theor. Appl. Genet. 100, 1137–1143. doi: 10.1007/s001220051397
Helguera, M., Khan, I. A., Kolmer, J., Lijavetzky, D., and Zhong-qi, L. (2003). PCR assays for the Lr37-Yr17-Sr38 cluster of rust resistance genes and their use to develop isogenic hard red spring wheat lines. Crop Sci. 43, 1839–1847. doi: 10.2135/cropsci2003.1839
Helguera, M., Vanzetti, L., Soria, M., Khan, I. A., and Kolmer, J. (2005). PCR Markers for Triticum speltoides leaf rust resistance gene Lr51 and their use to develop isogenic hard red spring wheat lines. Crop Sci. 45, 728–734. doi: 10.2135/cropsci2005.0728
Herrera-Foessel, S. A., Singh, R. P., Huerta-Espino, J., Rosewarne, G. M., Periyannan, S. K., Viccar, L., et al. (2012). Lr68: a new gene conferring slow rusting resistance to leaf rust in wheat. Theor. Appl. Genet. 124, 1475–1486. doi: 10.1007/s00122-012-1802-1
Herrera-Foessel, S. A., Singh, R. P., Huerta-Espino, J., William, H. M., Rosewarne, G., Djurle, A., et al. (2007). Identification and mapping of Lr3 and a linked leaf rust resistance gene in durum wheat. Crop Sci. 47, 1459–1466. doi: 10.1007/s00122-012-1788-8
Hiebert, C., Thomas, J., Somers, D., Mccallum, B., and Fox, S. (2007). Microsatellite mapping of adult-plant leaf rust resistance gene Lr22a in wheat. Theor. Appl. Genet. 115, 877–884. doi: 10.1007/s00122-007-0623-0
Hiebert, C. W., Thomas, J. B., McCallum, B. D., Humphreys, D. G., DePauw, R. M., Hayden, M. J., et al. (2010). An introgression on wheat chromosome 4DL in RL6077 (Thatcher*6/PI 250413) confers adult plant resistance to stripe rust and leaf rust (Lr67). Theor. Appl. Genet. 121, 1083–1091. doi: 10.1007/s00122-010-1373-y
Hiebert, C. W., Thomas, J. B., Mccallum, B. D., and Somers, D. J. (2008). Genetic mapping of the wheat leaf rust resistance gene Lr60 (LrW2). Crop Sci. 48, 1020–1026. doi: 10.2135/cropsci2007.08.0480
Huang, L., Brooks, S. A., Li, W. L., Fellers, J. P., Trick, H. N., and Gill, B. S. (2003). Map-based cloning of leaf rust resistance gene Lr21 from the large and polyploid genome of bread wheat. Genetics 164, 655–664.
Huang, L., and Gill, B. S. (2001). An RGA – like marker detects all known Lr21 leaf rust resistance gene family members in Aegilops tauschii and wheat. Theor. Appl. Genet. 103, 1007–1013. doi: 10.1007/s001220100701
Huerta-Espino, J., Singh, R. P., German, S., McCallum, B. D., Park, R. F., Chen, W. Q., et al. (2011). Global status of wheat leaf rust caused by Puccinia triticina. Euphytica 179, 143–160. doi: 10.1007/s10681-011-0361-x
Hufford, M. B., Xu, X., Van-Heerwaarden, J., Pyhäjärvi, T., Chia, J. M., Cartwright, R. A., et al. (2012). Comparative population genomics of maize domestication and improvement. Nat. Genet. 44, 808–811. doi: 10.1038/ng.2309
Hussain, M., Hassan, S. F., and Kirmani, M. A. S. (1980). “Virulence in Puccinia recondita Rob. ex. Desm. f. sp. tritici in Pakistan during 1978 and 1979,” in Proceedings of the 5th European and Mediterranean Cereal Rust Conference, Bari, 179–184.
Imbaby, I. A., Mahmoud, M. A., Hassan, M. E. M., and Abd-El-Aziz, A. R. M. (2014). Identification of leaf rust resistance genes in selected egyptian wheat cultivars by molecular markers. Sci. World J. 2014:574285. doi: 10.1155/2014/574285
Jing, F., Jiao-Jiao, X., Rin-Ming, L., Yue-Qiu, H., and Shi-Chang, X. (2013). Genetic analysis and location of gene for resistance to stripe rust in wheat international differential host Strubes Dickkopf. J. Genet. 92, 267–272. doi: 10.1007/s12041-013-0260-0
Jonathan, D. G. J., Witek, K., Walter, V., Florian, J., David, C., Stephen, D., et al. (2014). Elevating crop disease resistance with cloned genes. Philos. Trans. R. Soc. B Biol. Sci. 369, 20130087. doi: 10.1098/rstb.2013.0087
Joshi, L. M., Srivastava, K. D., and Ramanujam, K. (1975). An analysis of brown rust epidemics of 1971–72 and 1972–73. Indian Phytopathol. 28, 138.
Kamal, M. H. A., Kim, H. K., Shin, H. K., Choi, S. J., Baik, K. B., Tsujimoto, H., et al. (2010). Abiotic stress responsive proteins of wheat grain determined using proteomics technique. Am. J. Crop Sci. 4, 196–208.
Keed, B. R., and White, N. H. (1971). Quantitative effects of leaf and stem rust on yield and quality of wheat. Austr. J. Exp. Agric. 11, 550–555. doi: 10.1071/EA9710550
Keller, B., Krattinger, S., Yahiaoui, N., Brunner, S., Kaur, N., Cloutre, C., et al. (2008). “Molecular analaysis of fungal disease resistance in wheat,” in Proceedings of the 11th International Wheat Genetics Symposium, Vol. 1, eds R. Appels, R. Eastwood, E. Lagudah, P. Langridge, and M. Mackay (Sydney, NSW: Sydney University Press Lynne McIntyre and Peter Sharp).
Kerber, E. R. (1987). Resistance to leaf rust in hexaploid wheat: lr32 a third gene derived from Triticum tauschii. Crop Sci. 27, 204–206. doi: 10.2135/cropsci1987.0011183X002700020013x
Khan, R. R., Bariana, H. S., Dholakia, B. B., Naik, S. V., Lagu, M. D., Rathjen, A. J., et al. (2005). Molecular mapping of stem and leaf rust resistance in wheat. Theor. Appl. Genet. 111, 846–850. doi: 10.1007/s00122-005-0005-4
Khlestkina, E. K., Roder, M. S., Unger, O., Meinel, A., and Borner, A. (2007). More precise map position and origin of a durable non-specific adult plant disease resistance against stripe rust (Puccinia striiformis) in wheat. Euphytica 153, 110.
Knott, D. R. (1989). “The wheat rust – breeding for resistance,” in Monographs on Theorical and Applied Genetics, eds M. Grossman, H. F. Linskens, P. Maliga, and R. Riley (Berlin: Springer-Verlag), 12.
Kokhmetova, A. M., and Atishova, M. N. (2012). Identification of sources of resistance to wheat stem using molecular markers. Russ. J. Genet. Appl. Res. 2, 486–493. doi: 10.1134/S2079059712060081
Kolmer, J. A. (2005). Tracking wheat rust on a continental scale. Curr. Opin. Plant Biol. 8, 441–449. doi: 10.1016/j.pbi.2005.05.001
Kolmer, J. (2008). “Lr63, Lr64,” in Catalogue of Gene Symbols for Wheat: 2009 Supplement, ed. X. C. Xia 271. (Reference 10550, p 273).
Kolmer, J. (2013). Leaf rust of wheat: pathogen biology, variation and host resistance. Forests 4, 70–84. doi: 10.3390/f4010070
Kuraparthy, V., Chhuneja, P., Dhaliwal, H. S., Kaur, S., Bowden, R. L., and Gill, B. S. (2007a). Characterization and mapping of cryptic alien introgression from Aegilops geniculata with novel leaf rust and stripe rust resistance genes Lr57 and Yr40 in wheat. Theor. Appl. Genet. 114, 1379–1389. doi: 10.1007/s00122-007-0524-2
Kuraparthy, V., Sood, S., Chhuneja, P., Dhaliwal, H. S., Kaur, S., Bowden, R. L., et al. (2007b). A cryptic wheat-Aegilops triuncialis translocation with leaf rust resistance gene Lr58. Crop Sci. 47, 1995–2003. doi: 10.2135/cropsci2007.01.0038
Lagudah, E. S., Krattinger, S. G., Herrera-Foessel, S., Singh, R. P., Huerta-Espino, J., Spielmeyer, W., et al. (2009). Gene-specific markers for the wheat gene Lr34/Yr18/Pm38 which confers resistance to multiple fungal pathogens. Theor. Appl. Genet. 119, 889–898. doi: 10.1007/s00122-009-1097-z
Lagudah, E. S., McFadden, H., Singh, R. P., Huerta-Espino, J., Bariana, H. S., and Spielmeyer, W. (2006). Molecular genetics characterization of the Lr34/Yr18 slow rusting resistance gene region in wheat. Theor. Appl. Genet. 114, 21–30. doi: 10.1007/s00122-006-0406-z
Landjeva, S., Korzun, V., and Borner, A. (2007). Molecular markers: actual and potential contributions to wheat genome characterization and breeding. Euphytica 15, 271–296. doi: 10.1007/s10681-007-9371-0
Lawrence, G. J., Finnegan, E. J., Ayliffe, M. A., and Ellis, J. G. (1995). The L6 gene for flax rust resistance is related to the Arabidopsis bacterial resistance gene RPS2 and the tobacco viral resistance gene N. Plant Cell 7, 1195–1206. doi: 10.2307/3870095
Leonard, K. J. (2001). “Stem rust–future enemy,” in Stem Rust of Wheat: From Ancient Enemy to Modern Foe, ed. P. D. Peterson (St. Paul, MN: APS Press), 119–146.
Leonard, K. J., and Szabo, L. J. (2005). Stem rust of small grains and grasses caused by Puccinia graminis. Mol. Plant Path. 6, 99–111. doi: 10.1111/j.1364-3703.2005.00273.x
Letta, T., Maccaferri, M., Ammar, K., Badebo, A., Ricci, A., Crossa, J., et al. (2013). Searching for novel sources of field resistance to Ug99 and Ethiopian stem rust races in durum wheat via association mapping. Theor. Appl. Genet. 126, 1237–1256. doi: 10.1007/s00122-013-2050-8
Liu, J., Chang, Z., Zhang, X., Yang, Z., Li, X., Jia, J., et al. (2013). Putative Thinopyrum intermedium-derived stripe rust resistance gene Yr50 maps on wheat chromosome arm 4BL. Theor. Appl. Genet. 126, 265–274. doi: 10.1007/s00122-012-1979-3
Liu, S., Yu, L. X., Singh, R. P., Jin, Y., Sorrells, M. E., and Anderson, J. A. (2010). Diagnostic and co-dominant PCR markers for wheat stem rust resistance genes Sr25 and Sr26. Theor. Appl. Genet. 120, 691–697. doi: 10.1007/s00122-009-1186-z
Liu, Y., He, Z., Apples, R., and Xia, X. (2012). Functional markers in wheat: current status and future prospects. Theor. Appl. Genet. 1, 1–10. doi: 10.1007/s00122-012-1829-3
Liu, Z. Y., Sun, Q. X., Ni, Z. F., and Yang, T. M. (1999). Development of SCAR markers linked to the Pm21 gene conferring resistance to powdery mildew in common wheat. Plant Breed. 118, 215–219. doi: 10.1046/j.1439-0523.1999.118003215.x
Loutre, C., Wicker, T., Travella, S., Galli, P., Scofield, S., Fahima, T., et al. (2009). Two different CC-NBS-LRR genes are required for Lr10-mediated leaf rust resistance in tetraploid and hexaploid wheat. Plant J. 60, 1043–1054. doi: 10.1111/j.1365-313X.2009.04024.x
Ma, D., Zhou, X., Hou, L., Bai, Y., Li, Q., Wang, H., et al. (2013). Genetic analysis and molecular mapping of a stripe rust resistance gene derived from Psathynrostachys huashanica Keng in wheat line H9014-121-5-5-9. Mol. Breed. 32, 365–372. doi: 10.1007/s11032-013-9876-2
Ma, J. X., Zhou, R. H., Dong, Y. S., and Wang, L. F. (2001). Molecular mapping and detection of the yellow rust resistance gene Yr26 in wheat transferred from Triticum turgidum L. using microsatellite markers. Euphytica 120, 219–226. doi: 10.1023/A:1017510331721
Ma, Z. Q., Gill, B. S., Sorrells, M. E. and Tanksley, S. D. (1993). RELP markers linked to two Hessian fly-resistance genes in wheat (Triticum aestivum L.) from Triticum tauschii (Coss.) Schmal. Theor. Appl. Genet. 85, 750–754.
Ma, Z. Q., Sorrells, M. E., and Tanksley, S. D. (1994). RELP markers linked to powdery mildew resistance genes Pm1, Pm2, Pm3, and Pm4 in wheat. Genome 37, 871–875.
Maccaferri, M., Sanguineti, M. C., Demontis, A., El-Ahmed, A., Moral, L. G., Maalouf, F. et al. (2010). Association mapping in durum wheat grown across a broad range of water regimes. J. Exp. Bot. 62, 409–438. doi: 10.1093/jxb/erq287
Maccaferri, M., Sanguineti, M. C., Moral, L. F. G., Demontis, A., El-Ahmed, A., Maalouf, F., et al. (2011). Association mapping in durum wheat grown across a broad range of water regimes and yield potential. J. Exp. Bot 62, 409–438. doi: 10.1093/jxb/erq287
Maccaferri, M., Zhang, J., Bulli, P., Abate, Z., Chao, S., Cantu, D., et al. (2015). A genome-wide association study of resistance to stripe rust (Puccinia striiformis f. sp. tritici) in a worldwide collection of hexaploid spring wheat (Triticum aestivum L.). G3 (Bethesda) 5, 449–465. doi: 10.1534/g3.114.014563
Mago, R., Bariana, H. S., Dundas, I. S., Spielmeyer, W., and Lawrence, G. L. (2005). Development of PCR markers for the selection of wheat stem rust resistance genes Sr24 and Sr26 in diverse wheat germplasm. Theor. Appl. Genet. 111, 496–504. doi: 10.1007/s00122-005-2039-z
Mago, R., Brown-Guedira, G., Dreisigacker, S., Breen, J., Jin, Y., Singh, R., et al. (2011). An accurate DNA marker assay for stem rust resistance gene Sr2 in wheat. Theor. Appl. Genet. 122, 735–744. doi: 10.1007/s00122-010-1482-7
Mago, R., Spielmeyer, W., Lawrence, G. L., Lagudah, E. S., and Ellis, G. J. (2002). Identification and mapping of molecular markers linked to rust resistance genes located on chromosome 1RS of rye using wheat-rye translocation lines. Theor. Appl. Genet. 104, 1317–1324. doi: 10.1007/s00122-002-0879-3
Mago, R., Verlin, D., Zhang, P., Bansal, U., Bariana, H., Jin, Y., et al. (2013). Development of wheat-Aegilops speltoides recombinants and simple PCR-based markers for Sr32 and a new stem rust resistance gene on the 2S#1 chromosome. Theor. Appl. Genet. 126, 2943–2955. doi: 10.1007/s00122-013-2184-8
Mateos-Hernandez, M., Singh, R., Hulbert, S. H., Bowden, R. L., Huerta-Espino, J., Gill, B. S., et al. (2006). Targeted mapping of ESTs linked to the adult plant resistance gene Lr46 in wheat using synteny with rice. Funct. Integr. Genomics 6, 122–131. doi: 10.1007/s10142-005-0017-9
Mccartney, C. A., Somers, D. J., Mccallum, B. D., Thomas, J., Humphreys, D. G., Menzies, J. G., et al. (2005). Microsatellite tagging of the leaf rust resistance gene Lr16 on wheat chromosome 2BSc. Mol. Breed. 15, 329–337. doi: 10.1007/s11032-004-5948-7
McIntosh, R., Dubcovsky, J., Rogers, J., Morris, C., Appels, R., and Xia, X. (2011). Catalogue of gene symbols for wheat: 2011 supplement. Annu. Wheat Newsl. 56, 273–282.
McIntosh, R. A., Hart, G. E., Devos, K. M., Gale, M. D., and Rogers, W. J. (1998). “Catalog of gene symbols for wheat,” in Proceedings of the 9th International Wheat Genetics Symposium, (Saskatoon, SK), 1–235.
McIntosh, R. A., Wellings, C. R., and Park, R. F. (1995). Wheat Rusts: An Atlas of Resistance Genes. Melbourne, VIC: CSIRO Publications.
McIntosh, R. A., Yamazaki, Y., Dubcovsky, J., Rogers, J., Morris, C., Somers, D. J., et al. (2008). “Catalogue of gene symbols for wheat,” in Proceedings of the11th International Wheat Genetics Symposium, Brisbane, QLD.
Mebrate, S. A., Oerke, E. C., Dehne, H. W., and Pillen, K. (2008). Mapping of the leaf rust resistance gene Lr38 on wheat chromosome arm 6DL using SSR markers. Euphytica 162, 457–466. doi: 10.1007/s10681-007-9615-z
Murray, G. M., and Brennan, J. P. (2009). The Current and Potential Costs from Diseases of Wheat in Australia. Canberra: Grains Research and Development Corporation.
Nagarajan, S., and Joshi, L. M. (1978). Epidemiology of brown and yellow rusts of wheat in north India. II. Associated meteorology conditions. Plant Dis. Reptr. 62, 186–188.
Naik, B. K., Sharma, J. S., Sivasamy, M., Prabhu, K. V., Tomar, R. S., and Tomar, S. M. S. (2015). Molecular mapping and validation of the microsatellite markers linked to the Secale cereale-derived leaf rust resistance gene Lr45 in wheat. Mol. Breed. 35, 61. doi: 10.1007/s11032-015-0234-4
Naik, S., Gill, K. S., Prakasa, R. V. S., Gupta, V. S., Tamhankar, S. A., Pujar, S., et al. (1998). Identification of a STS marker linked to the Aegilops speltoides derived leaf rust resistance gene Lr28 in wheat. Theor. Appl. Genet. 97, 535–540. doi: 10.1007/s001220050928
Naruoka, Y., Garland-Campbell, K. A., and Carter, A. H. (2015). Genome-wide association mapping for stripe rust (Puccinia striiformis F. sp. tritici) in US Pacific Northwest winter wheat (Triticum aestivum L.). Theor. Appl. Genet. 128, 1083–1101. doi: 10.1007/s00122-015-2492-2
Naz, A. A., Kunert, A., Flath, K., Pillen, K., and Leon, J. (2012). Advanced backcross quantitative trait locus analysis in winter wheat: dissection of stripe rust seedling resistance and identification of favorable exotic alleles originated from a primary hexaploid wheat (Triticum turgidum ssp. dicoccoides 3 Aegilops tauschii). Mol. Breed. 30, 1219–1229.
Neelam, K., Brown-Guedira, G., and Huang, L. (2013). Development and validation of a breeder-friendly KASPar marker for wheat leaf rust resistance locus Lr21. Mol. Breed. 31, 233–237. doi: 10.1007/s11032-012-9773-0
Nelson, J. C., Singh, R. P., Autrique, J. E., and Sorrells, M. E. (1997). Mapping genes conferring and suppressing leaf rust resistance in wheat. Crop Sci. 37, 1928–1935. doi: 10.2135/cropsci1997.0011183X003700060043x
Neu, C., Stein, N., and Keller, B. (2002). Genetic mapping of the Lr20–Pm1 resistance locus reveals suppressed recombination on chromosome arm 7AL in hexaploid wheat. Genome 45, 737–744. doi: 10.1139/g02-040
Nisha, R., Sivasamy, M., Gajalakshmi, K., Shajitha, P., Vikas, V. K., Peter, J., et al. (2015). Pyramiding of stem rust resistance genes to develop durable and multiple disease resistant wheat varieties through marker aided selection. International J Ext Res. 5, 1–9.
Niu, Z., Klindworth, D. L., Yu, G., Friesen, T., Chao, S., Jin, Y., et al. (2014). Development and characterization of wheat lines carrying stem rust resistance gene Sr43 derived from Thinopyrum ponticum. Theor. Appl. Genet. 127, 969–980. doi: 10.1007/s00122-014-2272-4
Obert, D. E., Fritz, A. K., Moran, J. L., Sukhwinder, S., Rudd, J. C., and Menz, M. A. (2005). Identification and molecular tagging of a gene from PI 289824 conferring resistance to leaf rust (Puccinia triticina) in wheat. Theor. Appl. Genet. 110, 1439–1444. doi: 10.1007/s00122-005-1974-z
Olmstead, A. L., and Rhode, P. W. (2011). Adapting North American wheat production to climatic challenges, 1839–2009. Proc. Natl. Acad. Sci. U.S.A. 108, 480–485. doi: 10.1073/pnas.1008279108
Paull, J. G., Pallotta, M. A., Langridge, P., and The, T. T. (1995). RFLP markers associated with Sr22 and recombination between chromosome 7A of bread wheat and the diploid species Triticum boeoticum. Theor. Appl. Genet. 89, 1039–1045. doi: 10.1007/BF00224536
Peng, J. H., Fahima, T., Roeder, M. S., Huang, Q. Y., and Dahan, A. (2000). A High-density molecular map of chromosome region harboring stripe-rust resistance genes YrH52 and Yr15 derived from wild emmer wheat, Triticum dicoccoides. Genetics 109, 199–210.
Penner, G. A., Clarke, J., Bezte, L. J., and Leisle, D. (1995). Identification of RAPD markers linked to a gene governing cadmium uptake in durum wheat. Genome 38, 543–547. doi: 10.1139/g95-070
Periyannan, S., Bansal, U., Bariana, H., Deal, K., Luo, M. C., Dvorak, J., et al. (2014). Identification of a robust molecular marker for the detection of the stem rust resistance gene Sr45 in common wheat. Theor. Appl. Genet. 127, 947–955. doi: 10.1007/s00122-014-2270-6
Periyannan, S., Moore, J., Ayliffe, M., Bansal, U., Wang, X., Huang, L., et al. (2013). The gene Sr33, an ortholog of barley mla genes, encodes resistance to wheat stem rust race Ug99. Science 341, 786–788. doi: 10.1126/science.1239028
Prabhu, K. V., Gupta, S. K., Charpe, A., and Koul, S. (2004). SCAR marker tagged to the alien leaf rust resistance gene Lr19 uniquely marking the Agropyron elongatum-derived gene Lr24 in wheat: a revision. Plant Breed. 123, 417–420. doi: 10.1111/j.1439-0523.2004.00971.x
Prabhu, K. V., Gupta, S. K., Charpe, A., Koul, S., Cherukuri, D. P., Dhaliwal, H. S., et al. (2003). Molecular markers detect redundancy and miss-identity in genetic stocks with alien leaf rust resistance genes Lr32 and Lr28 in bread wheat. J. Plant Biochem. Biot. 12, 123–129. doi: 10.1007/BF03263172
Pretorius, Z. A., Singh, R. P., Wagoire, W. W., and Payne, T. S. (2000). Detection of virulence to wheat stem rust resistance gene Sr31 in Puccinia graminis f. sp. tritici in Uganda. Plant Dis. 84, 203. doi: 10.1094/PDIS.2000.84.2.203B
Prins, R., Groenewald, J. Z., Marais, G. F., Snape, J. W., and Koebner, R. M. D. (2001). AFLP and STS tagging of Lr19, a gene conferring resistance to leaf rust in wheat. Theor. Appl. Genet. 103, 618–624. doi: 10.1007/PL00002918
Procunier, J. D., Townley-Smith, T. F., Prashar, S., Gray, M. A., Kim, W. K., Czarnecki, E., et al. (1995). PCR-based RAPD/DGGE markers linked to leaf rust resistance genes Lr29 and Lr25 in wheat (Triticum aestivum L.). J. Genet. Breed. 49, 87–92.
Qi, L., Cao, M., Chen, P., Li, W., and Liu, D. (1996). Identification, mapping and application of polymorphic DNA associated with resistance gene Pm21 of wheat. Genome 39, 191–197. doi: 10.1139/g96-025
Qiu, J. W., Schürch, A. C., Yahiaoui, N., Dong, L. L., Fan, H. J., Zhang, Z. J., et al. (2007). Physical mapping and identification of a candidate for the leaf rust resistance gene Lr1 of wheat. Theor. Appl. Genet. 115, 159–168. doi: 10.1007/s00122-007-0551-z
Rajaram, S., and Van Ginkel, M. (1996). A Guide to the CIMMYT Bread Wheat Section. In wheat special Report, No. 5. Mexico: International Maize and Wheat Improvement Center (CIMMYT).
Randhawa, H. S., Asif, M., Pozniak, C., Clarke, J. M., Graf, R. J., Fox, S. L., et al. (2013). Application of molecular markers to wheat breeding in Canada. Plant Breed. 132, 458–471. doi: 10.1111/pbr.12057
Randhawa, H., Puchalski, B. J., Frick, M., Goyal, A., Despins, T., Graf, R. J., et al. (2012). Stripe rust resistance among western Canadian spring wheat and triticale varieties. Can. J. Plant Sci. 92, 713–722. doi: 10.4141/cjps2011-252
Randhawa, M., Bansal, U., Valárik, M., Klocová, B., Doležel, J., and Bariana, H. (2014). Molecular mapping of stripe rust resistance gene Yr51 in chromosome 4AL of wheat. Theor. Appl. Genet. 127, 317–324. doi: 10.1007/s00122-013-2220-8
Raupp, W. J., Sukhwinder, S., Brown-Guedira, G. L., and Gill, B. S. (2001). Cytogenetic and molecular mapping of the leaf rust resistance gene Lr39 in wheat. Theor. Appl. Genet. 102, 347–352. doi: 10.1007/s001220051652
Rees, R. G., and Platz, G. J. (1975). Control of wheat leaf rust with 4-n-butyl-1, 2, 4-triazole. Austr. J. Exp. Agric. Anim. Husb. 15, 276–280. doi: 10.1071/EA9750276
Ren, R. S., Wang, M. N., Chen, X. M., and Zhang, Z. J. (2012). Characterization and molecular mapping of Yr52 for high-temperature adult-plant resistance to stripe rust in spring wheat germplasm PI 183527. Theor. Appl. Genet. 125, 847–857. doi: 10.1007/s00122-012-1877-8
Revathi, P., Tomar, S. M. S., and Singh, N. K. (2010). Marker assisted gene pyramiding of leaf rust resistance genes Lr24, Lr28 along with stripe rust resistance gene Yr15 in wheat (Triticum aestivum L.). Indian J. Genet. 70, 349–354.
Roberson, R. W., and Luttrell, E. S. (1987). Ultrastructure of teliospore ontogeny in Tilletia indica. Mycologia 79, 753–763. doi: 10.2307/3807828
Robert, O., Abelard, C., and Dedryve, F. (1999). Identification of molecular markers for the detection of the yellow rust resistance gene Yr17 in wheat. Mol. Breed. 5, 167–175. doi: 10.1023/A:1009672021411
Rosegrant, M. W., and Agcaoili, M. (2010). Global Food Demand, Supply and Food Prospects. Washington, DC: International food policy research Institute.
Rouse, M. N., Nava, I. C., Chao, S., Anderson, J. A., and Jin, Y. (2012). Identification of markers linked to the race Ug99 effective stem rust resistance gene Sr28 in wheat (Triticum aestivum L.). Theor. Appl. Genet. 3, 19–25. doi: 10.1007/s00122-012-1879-6
Rowland, G. G., and Kerber, E. R. (1974). Telocentric mapping in hexaploid wheat of genes for leaf rust resistance and other characters derived from Aegilops squarrosa. Can. J. Genet. Cytol. 16, 137–144.
Saari, E. E., and Prescott, J. M. (1985). “World distribution in relation to economic losses,” in The Cereal Rusts, eds A. P. Roelfs and W. R. Bushnell (Orlando, FL: Academic Press), 259–298.
Sacco, F., Suarez, E. Y., and Naranjo, T. (1998). Mapping of the leaf rust resistance gene Lr3 on chromosome 6B of Sinvalocho MA wheat. Genome 41, 686–690. doi: 10.1139/gen-41-5-686
Saintenac, C., Zhang, W., Salcedo, A., Rouse, M. N., Trick, H. N., Akhunov, E., et al. (2013). Identification of wheat gene Sr35 that confers resistance toUg99 stem rust race group. Science 341, 783–786. doi: 10.1126/science.1239022
Schachermayr, G. M., Feuillet, C., and Keller, B. (1997). Molecular markers for the detection of the wheat leaf rust resistance gene Lr10 in diverse genetic backgrounds. Mol. Breed. 3, 65–74. doi: 10.1023/A:1009619905909
Schachermayr, G. M., Messmer, M. M., Feuillet, C., Winzeler, H., Winzeler, M., and Keller, B. (1995). Identification of molecular markers linked to the Agropyron elongatum-derived leaf rust resistance gene Lr24 in wheat. Theor. Appl. Genet. 90, 982–990. doi: 10.1007/BF00222911
Schachermayr, G. M., Siedler, H., Gale, M. D., Winzeler, H., Winzeler, M., and Keller, B. (1994). Identification and localization of molecular markers linked to the Lr9 leaf rust resistance gene of wheat. Theor. Appl. Genet. 88, 110–115. doi: 10.1007/BF00222402
Seyfarth, R., Feuillet, C., and Keller, B. (1998). “Development of a molecular marker for the adult plant leaf rust resistance gene Lr13 and Lr35 in wheat,” in Proceedings of the 9th International Wheat Genetics Symposium, Vol. 3, (Saskatoon, SK: University Extension Press), 154–155.
Seyfarth, R., Feuillet, C., Schachermayr, G., Winzeler, M., and Keller, B. (1999). Development of a molecular marker for the adult plant leaf rust resistance gene Lr35 in wheat. Theor. Appl. Genet. 99, 554–560. doi: 10.1007/s001220051268
Seyfarth, R., Feuillt, C., Shachermayr, G., Messmer, M., Winzeler, M., and Keller, B. (2000). Molecular maping of the adult plant leaf rust resistance gene lr13 in wheat (Triticum aestivum L.). J.Gent. Breed. 54, 193–198.
Sharma, T. R. (2003). Molecular diagnosis and application of DNA markers in the management of fungal and bacterial plant diseases. Indian J. Biotechnol. 2, 99–109.
Shi, A. N., Leath, S., and Murphy, J. P. (1998). A major gene for powdery mildew resistance transferred to common wheat from wild einkorn wheat. Phytopathology 88, 144–147. doi: 10.1094/PHYTO.1998.88.2.144
Sim, S. C., Durstewitz, G., Plieske, J., Wieseke, R., Ganal, M. W., Van-Deynze, A., et al. (2012). Development of a large SNP genotyping array and generation of high-density genetic maps in tomato. PLoS ONE 7:e40563. doi: 10.1371/journal.pone.0040563
Singh, A., Pallavi, J. K., and Gupta, P. (2012). Identification of microsatellite markers linked to leaf rust resistance gene Lr25 in wheat. National Phytotron Facility, Indian Agricultural Research Institute, New Delhi, India. J. Appl. Genet. 53, 19–25. doi: 10.1007/s13353-011-0070-0
Singh, R. P. (1991). Pathogenicity variations of Puccinia recondite f. sp. tritici and P. graminis f. sp. tritici in wheat growing areas of Mexico during 1988 and 1989. Plant Dis. 75, 790–794. doi: 10.1094/PD-75-0790
Singh, R. P. (1993). Genetic association of gene Bdv1 for tolerance to barley yellow dwarf virus with genes Lr34 and Yr18 for adult plant resistance to rusts in bread wheat. Plant Dis. 77, 1103–1106. doi: 10.1094/PD-77-1103
Singh, R., Goutam, U., Gupta, R. K., Pandey, G. C., Shoran, J., and Tiwari, R. (2009). Allelic variations of functional markers for polyphenol oxidase (PPO) genes in Indian Wheat (Triticum aestivum L.) cultivars. J. Genet. 88, 325–329. doi: 10.1007/s12041-009-0047-5
Singh, R. P., Hodsonm, D. P., Jin, Y., Huerta-Espino, J., Kinyua, M. G., Wanyera, R., et al. (2006). Current status, likely migration and strategies to mitigate the threat to wheat production from race Ug99 (TTKS) of stem rust pathogen. CAB Rev. 54, 1–14.
Singh, R. P., Huerta-Espino, J., Pfeiffer, W., and Figueroa-Lopez, P. (2004). Occurrence and impact of a new leaf rust race on durum wheat in northwestern Mexico from 2001 to 2003. Plant Dis. 88, 703–708. doi: 10.1094/PDIS.2004.88.7.703
Singh, R. P., Huerta-Espino, J., Sharma, R., Joshi, A. K., and Trethowan, R. (2007). High yielding spring bread wheat germplasm for global irrigated and rainfed production systems. Euphytica 157, 351–363. doi: 10.1007/s10681-006-9346-6
Singh, R. P., Nelson, J. C., and Sorrels, M. E. (2000). Mapping Yr28 and other genes for resistance to stripe rust in wheat. Crop Sci. 40, 1148–1155. doi: 10.2135/cropsci2000.4041148x
Singh, R. P., and Trethowan, R. (2007). “Breeding spring bread wheat for irrigated and rainfed production systems of the developing world,” in Breeding Major Food Staples, eds M. Kang and P. M. Priyadarshan (Ames, IA: Blackwell Publishing), 109–140.
Singh, S., and Bowden, R. L. (2011). Molecular mapping of adult-plant race-specific leaf rust resistance gene Lr12 in bread wheat. Mol. Breed. 28, 137–142. doi: 10.1007/s11032-010-9467-4
Smith, P., Koebner, R., and Boyd, L. (2002). The development of a STS marker linked to a yellow rust resistance derived from the wheat cultivar Moro. Theor. Appl. Genet. 104, 1278–1282. doi: 10.1007/s00122-002-0895-3
Song, Q., Hyten, D. L., Jia, G., Quigley, C. V., Fickus, E. W., Nelson, R. L., et al. (2013). Development and evaluation of Soy SNP50K, a high-density genotyping array for soybean. PLoS ONE 8:e54985. doi: 10.1371/journal.pone.0054985
Spielmeyer, W., Sharp, P. J., and Lagudah, E. S. (2003). Identification and validation of markers linked to broad-spectrum stem rust resistance gene Sr2 in wheat (Triticum aestivum L.). Crop Sci. 43, 36. doi: 10.2135/cropsci2003.0333
Stepien, L., Golka, L., and Chelkowski, J. (2003). Leaf rust resistance genes of wheat: identification in cultivars and resistance sources. J. Appl. Genet. 44, 139–149.
Stokstad, E. (2007). Deadly wheat fungus threatens world’s breadbaskets. Science 315, 1786–1787. doi: 10.1126/science.315.5820.1786
Sun, X., Bai, G., and Carver, B. (2009). Molecular markers for wheat leaf rust resistance gene Lr41. Mol. Breed. 23, 311–321. doi: 10.1007/s11032-008-9237-8
Tar, M., Purnhauser, L., and Csõsz, M. (2008). Identification and localization of molecular markers linked to the Lr52 leaf rust resistance gene of wheat. Cereal Res. Commun. 36, 409–415. doi: 10.1007/BF00222402
Terefe, T., Paul, I., Mebalo, J., Naicker, K., and Meyer, L. (2009). Occurrence and pathogenicity of Puccinia triticina on wheat in South Africa during 2007. S. Afr. J. Plant Soil 26, 51–54. doi: 10.1080/02571862.2009.10639933
Terracciano, I., Maccaferri, M., Bassi, F., Mantovani, P., Sanguineti, M. C., Salvi, S., et al. (2013). Development of COS-SNP and HRM markers for high throughput and reliable haplotype-based detection of Lr14a in durum wheat (Triticum durum Desf.). Theor. Appl. Genet. 126, 1077–1101. doi: 10.1007/s00122-012-2038-9
Thomson, M. J. (2014). High Throughput SNP Genotyping to accelerate crop improvement. Plant Breed. Biotechnol. 2, 195–212. doi: 10.9787/PBB.2014.2.3.195
Todorovska, E., Christov, N., Slavov, S., Christova, P., and Vassilev, D. (2009). Biotic stress resistance in wheat-Breeding and Genomic selection implications. Biotechnol. Biotechnol. Equip. 23, 1417–1426. doi: 10.2478/V10133-009-0006-6
Tsilo, J. T., Jin, Y., and Anderson, J. A. (2007). Microsatellite markers linked to stem rust resistance allele Sr9a in wheat. Crop Sci. 47, 2013–2020. doi: 10.2135/cropsci2007.02.0087
Tsilo, T. J., Jin, Y., and Anderson, J. A. (2008). Diagnostic microsatellite markers for the detection of stem rust resistance gene Sr36 in diverse genetic backgrounds of wheat. Crop Sci. 48, 253–261. doi: 10.2135/cropsci2007.04.0204
Vanzetti, L. S., Campos, P., Demichelis, M., Lombardo, L. A., Aurelia, P. A., Vaschetto, L. M., et al. (2011). Identification of leaf rust resistance genes in selected Argentinean bread wheat cultivars by gene postulation and molecular markers. Electronic J. Biotechnol. 14, 1–6. doi: 10.2225/vol14-issue3-fulltext-14
Varshney, R. K., Manhendar, T., Aggarwal, R. K., and Borner, A. (2007). “Genetic molecular markers in plants: development and applications,” in Genomics-Assisted Crop Improvement: Genomics Approaches and PlatformsVarshney, 1, eds R. K. Varshney and R. Tuberosa (Berlin: Springer), 13–29.
Vida, G., Gal, M., Uhrin, A., Veisz, O., Syed, N. H., Flavell, A. J., et al. (2009). Molecular markers for the identification of resistance genes and marker-assisted selection in breeding wheat for leaf rust resistance. Euphytica 170, 67–76. doi: 10.1007/s10681-009-9945-0
Volkova, G. V., Alekseeva, T. P., Anpilogova, L. K., Dobryanskaya, M. V., Vaganova, O. F., and Kolbin, D. A. (2009). Phytopathological characteristics of leaf rust resistance of new winter wheat varieties. Russ. Agric. Sci. 35, 168–171. doi: 10.3103/S1068367409030112
Wang, L., Ma, J., Zhou, R., Wang, X., and Jia, J. (2002). Molecular tagging of the yellow rust resistance gene Yr10 in common wheat, PI 178383 (Triticum aestivum L). Euphytica 124, 71–73. doi: 10.1023/A:1015689817857
Wang, S., Wong, D., Forrest, K., Allen, A., Chao, S., and Huang, B. E. (2014). Characterization of polyploid wheat genomic diversity using a high-density 90000 single nucleotide polymorphism array. Plant Biotechnol. J. 12, 787–796. doi: 10.1111/pbi.12183
Watson, I. A., and Luig, N. H. (1961). Leaf rust on wheat in Australia: a systematic scheme for the classification of strains. Proc. Linn. Soc. N.S.W. 86, 241–250.
Wellings, C. R. (2011). Global status of stripe rust: a review of historical and current threats. Euphytica 179, 129–141. doi: 10.1007/s10681-011-0360-y
Wen, Z., Zhao, T., Zheng, Y., Liu, S., Wang, C., Wang, F., et al. (2008). Association analysis of agronomic and quality traits with SSR markers in glycine max and glycine soja in China: II. Exploration of elite alleles. Acta Agron. Sin. 34, 1339–1349. doi: 10.3724/SP.J.1006.2008.01339
Wiedmann, R. T., Smith, T. P. L., and Nonneman, D. J. (2008). SNP discovery in swine by reduced representation and high throughput pyrosequencing. BMC Genet. 9:81. doi: 10.1186/1471-2156-9-81
William, M., Langridge, P., Trethowan, R., Dreisigacker, S., and Crouch, J. (2008). “Genomics of wheat, the basis of our daily bread,” in Genomics of Tropical Plants, Vol. 15, eds P. H. Moore and R. Ming (New York, NY: Springer-Verlag), 515–548.
Xu, L. S., Wang, M. N., Cheng, P., Kang, Z. S., Hulbert, S. H., and Chen, X. M. (2013). Molecular mapping of Yr53, a new gene for stripe rust resistance in durum wheat accession PI 480148 and its transfer to common wheat. Theor. Appl. Genet. 126, 523–533. doi: 10.1007/s00122-012-1998-0
Xu, X., Liu, X., Ge, S., Jensen, J. D., Hu, F., Li, X., et al. (2012). Resequencing 50 accessions of cultivated and wild rice yields markers for identifying agronomically important genes. Nat. Biotechnol. 30, 105–111. doi: 10.1038/nbt.2050
Yan, G. P., Chen, X. M., Line, R. F., and Wellings, C. R. (2003). Resistance gene analog polymorphism markers cosegregating with the Yr5 gene for resistance to wheat stripe rust have homology with plant disease resistance genes. Theor. Appl. Genet. 106, 636–643.
Yaniv, E., Raats, D., Ronin, Y., Korol, A. B., Grama, A., Bariana, H., et al. (2015). Evaluation of marker-assisted selection for the stripe rust resistance gene Yr15, introgressed from wild emmer wheat. Mol. Breed. 35, 43. doi: 10.1007/s11032-015-0238-0
Yu, G., Zhang, Q., Friesen, T. L., Rouse, M. N., Jin, Y., Zhong, S., et al. (2015). Identification and mapping of Sr46 from Aegilops tauschii accession CIae 25 conferring resistance to race TTKSK (Ug99) of wheat stem rust pathogen. Theor. Appl. Genet. 128, 431–443. doi: 10.1007/s00122-014-2442-4
Yu, L. X., Lorenz, A., Rutkoski, J., Singh, R. P., Bhavani, S., Huerta-Espino, J., et al. (2011). Association mapping and gene-gene interaction for stem rust resistance in CIMMYT spring wheat germplasm. Theor. Appl. Genet. 123, 1257–1268. doi: 10.1007/s00122-011-1664-y
Yu, X. M., Zhao, W. Q., Yang, W. X., Liu, F., Chen, J. P., Goyer, C., et al. (2013). characterization of a hypersensitive response-induced gene TaHIR3 from wheat leaves infected with leaf rust. Plant Mol. Biol. Rep. 2, 314–322. doi: 10.1007/s11105-012-0504-9
Zegeye, H., Rasheed, A., Makdis, F., Badebo, A., and Ogbonnaya, F. C. (2014). Genome-wide association mapping for seedling and adult plant resistance to stripe rust in synthetic hexaploid wheat. PLoS ONE 9:e105593. doi: 10.1371/journal.pone.0105593
Zhang, N., Yang, W., Yan, H., Liu, D., Chu, D., Meng, Q., et al. (2005). Molecular markers of wheat leaf rust resistance gene Lr45 based on AFLP. Sci. Agric. Sin. 38, 1364–1368.
Zhang, X., Han, D., Zeng, Q., Duan, Y., Yuan, F., Shi, J., et al. (2013). Fine mapping of wheat stripe rust resistance gene Yr26 based on collinearity of wheat with Brachypodium distachyon and rice. PLoS ONE 8:e57885. doi: 10.1371/journal.pone.0057885
Zhao, K., Tung, C. W., Eizenga, G. C., Wright, M. H., Ali, M. L., Price, A. H., et al. (2011). Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat. Commun. 2, 467–471. doi: 10.1038/ncomms1467
Zhou, X. L., Han, D. J., Chen, X. M., Gou, H. L., Guo, S. J., Rong, L., et al. (2014a). Characterization and molecular mapping of stripe rust resistance gene Yr61 in winter wheat cultivar Pindong 34. Theor. Appl. Genet. 127, 2349–2358. doi: 10.1007/s00122-014-2381-0
Zhou, X. L., Han, D. J., Gou, H. L., Wang, Q. L., Zeng, Q. D., Yuan, F. P., et al. (2014b). Molecular mapping of a stripe rust resistance gene in wheat cultivar Wuhan 2. Euphytica 196, 251–259. doi: 10.1094/PHYTO-99-10-1209
Zhou, X. L., Wang, M. N., Chen, X. M., Lu, Y., Kang, Z. S., and Jing, J. X. (2014c). Identification of Yr59 conferring high temperature adult plant resistance to stripe rust in wheat germplasm PI 178759. Theor. Appl. Genet. 127, 935–945. doi: 10.1007/s00122-014-2269-z
Zhou, X. L., Wang, W. L., Wang, L. L., Hou, D. Y., Jing, J. X., Wang, Y., et al. (2011). Genetics and molecular mapping of genes for high-temperature resistance to stripe rust in wheat cultivar Xiaoyan 54. Theor. Appl. Genet. 123, 431–438. doi: 10.1007/s00122-011-1595-7
Zhou, Y., Ren, Y., Lillemo, M., Yao, Z., Zhang, P., Xia, X., et al. (2014). QTL mapping of adult plant resistance to leaf rust in a RIL population derived from a cross of wheat cultivars Shanghai 3/Catbird and Naxos. Theor. Appl. Genet. 127, 1873–1883. doi: 10.1007/s00122-014-2346-3
Zhu, C., Gore, M., Buckler, E. S., and Yu, J. (2008). Status and prospects of association mapping in plants. Plant Gen. 1, 5–20. doi: 10.3835/plantgenome2008.02.0089
Keywords: MAS, molecular markers, R genes, wheat, wheat rust
Citation: Goutam U, Kukreja S, Yadav R, Salaria N, Thakur K and Goyal AK (2015) Recent trends and perspectives of molecular markers against fungal diseases in wheat. Front. Microbiol. 6:861. doi: 10.3389/fmicb.2015.00861
Received: 01 June 2015; Accepted: 06 August 2015;
Published: 25 August 2015.
Edited by:
Vincenzo Lionetti, Sapienza Università di Roma, ItalyReviewed by:
Juan B. Alvarez, Universidad de Córdoba, SpainAnna Maria Mastrangelo, Centro di Ricerca per la Cerealicoltura, Italy
Copyright © 2015 Goutam, Kukreja, Yadav, Salaria, Thakur and Goyal. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Aakash K. Goyal, International Center for Agriculture Research in the Dry Areas (ICARDA), Morocco,YWtncm95YWxAZ21haWwuY29t
 Umesh Goutam
Umesh Goutam Sarvjeet Kukreja
Sarvjeet Kukreja Rakesh Yadav
Rakesh Yadav Neha Salaria
Neha Salaria Kajal Thakur
Kajal Thakur Aakash K. Goyal
Aakash K. Goyal