- 1Laboratory for Molecular Microbiology, Institute of Molecular Genetics and Genetic Engineering, University of Belgrade, Belgrade, Serbia
- 2Faculty of Pharmacy, University of Belgrade, Belgrade, Serbia
Enterococci represent the most controversial group of dairy bacteria. They are found to be the main constituent of many traditional Mediterranean dairy products and contribute to their characteristic taste and flavor. On the other hand, during the last 50 years antibiotic-resistant enterococci have emerged as leading causes of nosocomial infections worldwide. The aim of this study was to determine the diversity, technological properties, antibiotic susceptibility and virulence traits of 636 enterococci previously isolated from 55 artisan dairy products from 12 locations in the Western Balkan countries (WBC) of Serbia, Croatia and Bosnia and Herzegovina. All strains were identified both by microbiological and molecular methods. The predominant species was Enterococcus durans, followed by Enterococcus faecalis and Enterococcus faecium. Over 44% of the isolates were resistant to ciprofloxacin and erythromycin, while 26.2% of the isolates were multi-resistant to three or more antibiotics belonging to different families. 185 isolates (29.1%) were susceptible to all 13 of the antibiotics tested. The antibiotic-susceptible isolates were further tested for possible virulence genes and the production of biogenic amines. Finally, five enterococci isolates were found to be antibiotic susceptible with good technological characteristics and without virulence traits or the ability to produce biogenic amines, making them possible candidates for biotechnological application as starter cultures in the dairy industry.
Introduction
Bacteria of the genus Enterococcus, or enterococci, are considered lactic acid bacteria (LAB) (Schleifer and Ludwig, 1995). The usual ecological niche for Enterococcus species is the gastrointestinal tracts of humans and animals (Garg and Mital, 1991). However, enterococci are widely distributed in large numbers in foods, especially those of animal origin, such as various dairy and meat products, and some are also found in soil, water and on plants.
Enterococcus faecium, Enterococcus faecalis, and Enterococcus durans are the most prevalent species of enterococci in raw milk cheeses (Terzic-Vidojevic et al., 2007, 2009a,b, 2013, 2014a,b; Golić et al., 2013). They break down lactose and citrate during cheese ripening, which leads to the production of various volatile compounds, such as acetaldehyde, acetoin, diacetyl, and ethanol, which are responsible for the formation of the unique aroma and flavor of the final product (Andrighetto et al., 2001; Sarantinopoulos et al., 2001; Giraffa, 2002; Abeijón et al., 2006; Foulquié-Moreno et al., 2006). Due to their interesting metabolic and biotechnological traits (the ability to metabolize citrate, proteolytic and esterolytic activities, bacteriocin production, and their probiotic characteristics), enterococci may be utilized in food fermentation as commercial starter cultures (Tsakalidou et al., 1993; Centeno et al., 1999; Giraffa, 2003; Menéndez et al., 2004). On the other hand, enterococci have recently been found to be extremely important in clinical microbiology. Food-derived enterococci isolates are known to be resistant to a wide variety of antibiotics and to possess virulence genes (Giraffa, 2002; Saavedra et al., 2003; Veljović et al., 2014). These features contribute to the pathogenicity of enterococci, making them opportunistic pathogens. A major concern is the emergence of vancomycin-resistant enterococci (VRE), since this antibiotic is considered the last alternative for the treatment of multiple resistant infections (Teuber et al., 1999). Ideally, each strain that is intended to be used in a starter culture should be tested individually before any use in food or medicine, must not possess a single virulence factor and should be susceptible to relevant clinical antibiotics (Domig et al., 2003; Franz et al., 2003).
Due to their controversial status enterococci have been studied extensively (Galgano et al., 2001; Gelsomino et al., 2001; Morandi et al., 2006; Psoni et al., 2006; Gomes et al., 2008; Franz et al., 2011; Macedo et al., 2011; Ducková et al., 2014). Preliminary research on autochthonous LAB isolated from artisan dairy products of the Western Balkan region has indicated that about one third of all isolated LAB are Enterococcus species (Terzic-Vidojevic et al., 2007, 2009a,b, 2013, 2014a,b; Golić et al., 2013). However, a comprehensive study regarding their safe use as starter cultures in this region is lacking.
This paper evaluates the genetic diversity and antibiotic susceptibility of 636 Enterococcus spp. isolates from a laboratory collection including strains previously isolated from various artisan dairy products such as cheese, sour cream and kajmak from Serbia, Croatia and Bosnia and Herzegovina. The genetic diversity was explored by rep-PCR using the (GTG)5 primer, combined with 16S rDNA sequencing. In antibiotic susceptible strains the presence of virulence factors and production of biogenic amines were also determined. We aimed to provide the most complete information about the phenotypic and genotypic diversity, as well as the safety status, of natural enterococcal isolates from traditional dairy products of the Western Balkan region for their potential biotechnological application as starter cultures in the dairy industry.
Materials and Methods
Bacterial Strains, Media, and Growth Conditions
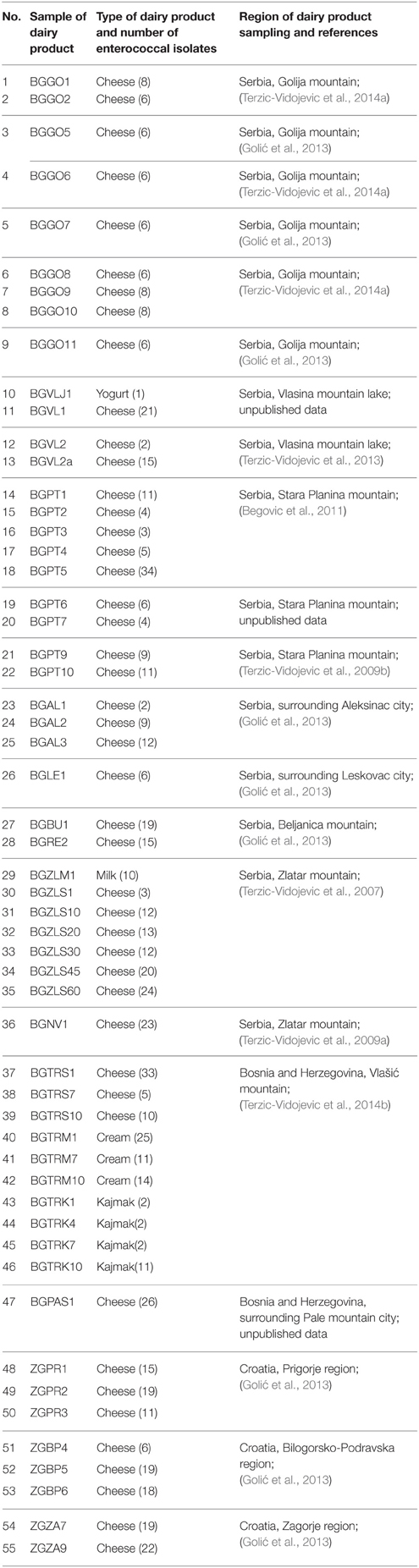
All enterococci strains used in this study were isolated from 55 samples of different artisan dairy products collected from specific rural locations in Serbia, Bosnia and Herzegovina and Croatia in the period from 2003 to 2011. The samples of dairy products, together with their sources of isolation, are listed in Table 1. In total, 636 natural isolates of enterococci from the laboratory collection, initially identified by physiological tests, were used for the investigation of genetic and phenotypic diversity and antibiotic susceptibility.
Enterococci and lactococci used as indicator strains for the analysis of antimicrobial activity of the enterococci strains were grown in M17 broth (Merck, GmbH, Darmstadt, Germany) supplemented with glucose (0.5% w/v) (GM17) at 30°C, while lactobacilli, used to screen the antimicrobial activity of Enterococcus sp. were incubated in MRS broth (Merck) at 30°C. Preliminary screening of 636 enterococci isolates for production of antimicrobial compounds was done by the deferred antagonism method using overnight cultures of isolates and various indicator strains. Briefly, soft GM17 and MRS agars (0.7% w/v) containing lactococci or lactobacilli indicator strains were overlaid onto GM17 and MRS plates, respectively. The plates were incubated overnight at the appropriate temperature (30 or 37°C) depending on the indicator strain. A clear zone of inhibition of indicator strain growth around the well was taken as a positive signal for production of antimicrobial compound.
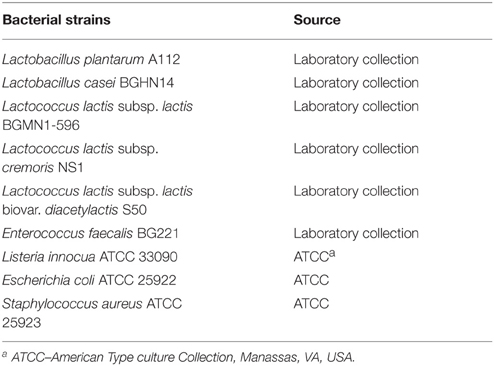
Escherichia coli and Staphylococcus aureus were cultivated in Luria broth (LB), containing 0.5% NaCl, 0.5% yeast extract (Torlak, Belgrade, Serbia), and 1% bacto tryptone (Torlak) at 37°C while Listeria innocua was cultivated on BHI medium (Difco, Detroit, MI, USA) at 37°C. Corresponding agar plates were prepared by adding agar (1.7% w/v, Torlak) into each broth. The indicator strains used in this study for screening enterococci antimicrobial activity are listed in Table 2. The isolates were stored at −80°C in GM17 broth (Merck) supplemented with glycerol (15% v/v) and revitalized in the same medium by overnight growth at 30°C.
Bacterial Identification
After microscopic examination, enterococci were identified to the genus level by Gram staining, catalase testing, arginine and bile-esculin hydrolysis, growth at 45°C and growth in NaCl 6.5% broth. All 636 enterococci isolates were subjected to the following tests: growth at 15°C, growth in broth with NaCl (8% w/v), production of CO2 from glucose, citrate utilization, acetoin and diacetyl production, time required for the formation of curd in reconstituted skim milk, exopolysaccharides (EPS) production, aggregation ability, and proteolytic and antimicrobial activity as described previously (Terzic-Vidojevic et al., 2007, 2009a,b, 2013, 2014a,b; Golić et al., 2013).
Identification of enterococci to the species level was performed according to Versalovic et al. (1994) using repetitive element palindromic-polymerase chain reaction (rep-PCR) analysis with (GTG)5 oligonucleotide (50-GTGGTGGTGGTGGTG-30). For this purpose the complete DNA from each of the 636 enterococci isolates was extracted as described by Hopwood et al. (1985) and details of the procedure has been reported by Terzic-Vidojevic et al. (2007). For sequencing of the 16S rRNA region, the complete DNA from certain enterococci isolates was used as a template for PCR amplification with UNI16SF (50-GAGAGTTTGATCCTGGC-30) and UNI16SR (50-AGG AGGTGATCCAGCCG-30) oligonucleotides (Jovcic et al., 2009). The PCR product obtained was purified by Qiagen (GmbH, Hilden, Germany) and sequenced (Macrogen, Amsterdam, the Netherlands and Seoul, South Korea). The BLAST algorithm was used to determine the most related sequences in the NCBI nucleotide sequence database (http://www.ncbi.nlm.nih.gov/BLAST).
Antibiotic Susceptibility Testing
The antibiotic resistance of enterococci isolates was determined by the disc diffusion method recommended by the Clinical and Laboratory Standards Institute (CLSI, 2012). The following antimicrobial drugs (Bio-Rad, Marnes-la-Coquette, France) were used: vancomycin (30 μg), teicoplanin (30 μg), ampicillin, (10 μg), erythromycin (15 μg), tetracycline (30 μg), minocycline (30 μg), quinupristin–dalfopristin (15 μg), ciprofloxacin (5 μg), chloramphenicol (30 μg), nitrofurantoin (300 μg), and linezolid (30 μg), and for high-level resistance (HLR) gentamicin (120 μg) and streptomycin (300 μg).
PCR Detection of Virulence Determinants
The complete DNA of 11 antibiotic-susceptible enterococci strains with the best technological characteristics was used in PCR reactions to detect the presence or absence of genes for the following virulence determinants: cytolysin (cylA, cylB, and cylM), aggregation factor (agg), gelatinase (gelE), enterococcal surface protein (esp), cell wall adhesions (efaAfs, and efaAfm), and sex-pheromones (cpd, cob, and ccf) according to Eaton and Gasson (2001), and collagen adhesin (ace) and hyaluronidase (hyl) as described by Vankerckhoven et al. (2004).
Biogenic Amines Determination
The ability of the 11 chosen enterococci strains to produce biogenic amines was qualitatively determined on an improved screening medium as described by Bover-Cid and Holzapfel (1999) using four precursor amino acids: histidine, lysine, ornithine, and tyrosine.
Statistical Analysis
Classical ecology indexes were used to obtain species richness (S), with the Shannon–Wiener index (H′) indicating general biodiversity and Simpson's index (D) evaluating dominance of the species in each cheese sample, as follows:
Where N is the number of species and pi is the number of isolates belonging to one species in the sample. Clustering was carried out in Statistica 7.0 for Windows (StatSoft Inc. USA) and in BioNumerics 6.5 using the algorithm “Unweighted Pair-Group Average Linkage Analysis.” Distances between the clusters were assessed using “Percent of disagreement.”
Results
The Diversity of Enterococci Isolates
The general index of enterococci species diversity (H′) was calculated on the basis of the number of different enterococci species among the 636 isolates. The results showed that the dairy samples analyzed contained five Enterococcus species: E. durans, E. faecalis, E. faecium, E. italicus, and E. avium. The highest diversity and the lowest index of dominance of enterococci species were scored by samples from cheeses from the Prigorje region, Croatia (H′ = 1.61; D = 0.36) and Pirot region, Serbia (H′ = 1.5; D = 0.34), where four of five enterococci species were identified (E. durans, E. faecalis, E. faecium, E. italicus). In contrast, the lowest diversity (H′ = 0.99) and the highest dominance index (D = 0.62) were scored by samples from cheese manufactured in Pale, Bosnia and Herzegovina, where almost all isolates (20 out of 26) belonged to E. faecium. E. durans was the most abundant species in dairy products from Travnik, Bosnia and Herzegovina (84 out of 115), while E. faecalis was the most abundant in dairy samples from Zlatar Mountain, Serbia (84 out of 117). Interestingly, only 12 out of 636 isolates belonged to E. italicus and they were isolated from five out of 12 regions, while only one out of 636 isolates was identified as E. avium and was found in the Zlatar region, Serbia (Table 3).
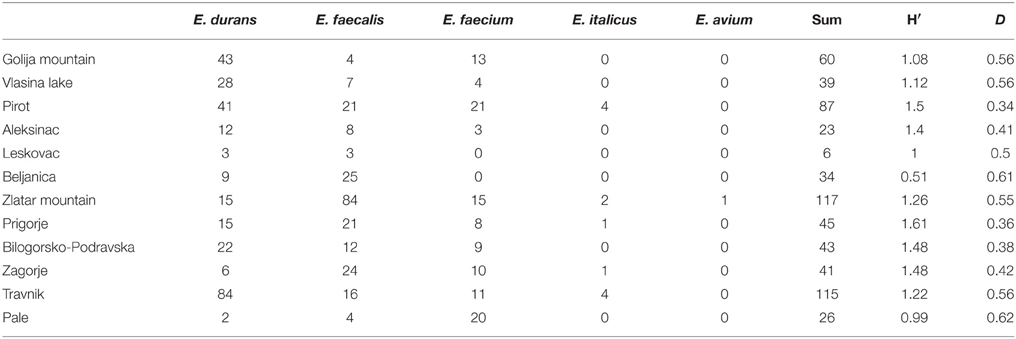
Table 3. The diversity of Enterococcus isolates from autochthonous dairy products collected at various geographic locations of the Western Balkan Countries.
Genotypic Characterization of Enterococci
All 636 enterococci isolates were subjected to (GTG)5-fingerprint analysis. The results revealed that 278 out of 636 isolates (43.71%) differ among each other and have unique (GTG)5-fingerprint profiles, indicating great diversity of the isolates. Among them 124 isolates belonged to E. durans, 74 to E. faecium, 68 to E. faecalis, 11 to E. italicus and 1 to E. avium species. The fingerprint profiles obtained by (GTG)5-PCR of the enterococci isolates are presented in Figure S1. Interestingly, four main clusters have been determined. According to (GTG)5-fingerprint patterns, Cluster 1, comprising 46 isolates, includes the isolates with less genetic distance, mostly isolated from dairy products sampled in close geographical regions situated in the western part of Serbia. Cluster 2, comprising 124 isolates, includes more heterogeneous isolates, mostly isolated from dairy products sampled from southern Serbia, but also from Bosnia and Herzegovina. Cluster 3, comprising 10 isolates, includes only enterococci isolated from Bosnia and Herzegovina, specifically from milk and cheese sampled in the same household. Cluster 4 is the most heterogeneous according to (GTG)5-fingerprint profiles and encompasses the isolates originating from various geographical locations. It can be further divided into groups 4a and 4b. Group 4a is mostly comprised of the isolates originating from Golija Mountain, while group 4b is further divided to subgroups comprising isolates originating from western Serbia and Bosnia and Herzegovina. Hence, the results show that the corresponding groups of strains are partly correlated with the sources of isolation. Interestingly, a subdivision on the basis of households in the same locality was seen within the groups of isolates (e.g., the isolates from dairy products sampled in Croatia are scattered throughout the dendrogram), and isolates from the same households are divided among separate clusters. Finally, the isolates that were undistinguished by rep-PCR were classified on the basis of phenotypic characteristics.
Phenotypic Characterization of Enterococci
Apart from general and genotypic diversity, the ultimate goal in exploring the diversity of natural isolates from various ecological niches is to determine their technological and functional potential. Hence, detailed phenotypic characterization of the enterococci strains with regard to acetoin (VP+) and diacetyl (D+) production, citrate utilization (C+), time of milk curdling (TMC), antimicrobial properties (Bac+), production of proteinases (Prt+), aggregation ability (Agg+), and EPS production was performed (Table 4). A large number of enterococci produced acetoin and diacetyl, 54.6 and 22.3% respectively. In addition, 40.9% of enterococci could utilize citrate as their only carbon source. On the other hand, only 4.2% of all the enterococci curdled milk within 6 h at 37°C (Table 4). Examination of the proteolytic activity of all 636 strains revealed that 17.5% of enterococci exhibited proteolytic activity as evaluated from β-casein hydrolysis by whole cells after 3 h of incubation. Almost 25% of all enterococci isolates showed the ability to produce antimicrobial compounds and most of those were enterococci that belonged to the E. durans species (29.3% of all 280 E. durans strains). Experiments with pronase E revealed the proteinaceous nature of the antimicrobial compounds, indicating that they could be bacteriocin-like substances (BLIS). Thirteen of 280 E. durans, three of 114 E. faecium, two of 229 E. faecalis strains and one E. avium strain had aggregation ability. Furthermore, one E. durans and one E. italicus strain were EPS producers. Some isolates exhibited two, three, or even four of the tested characteristics (Table 4).
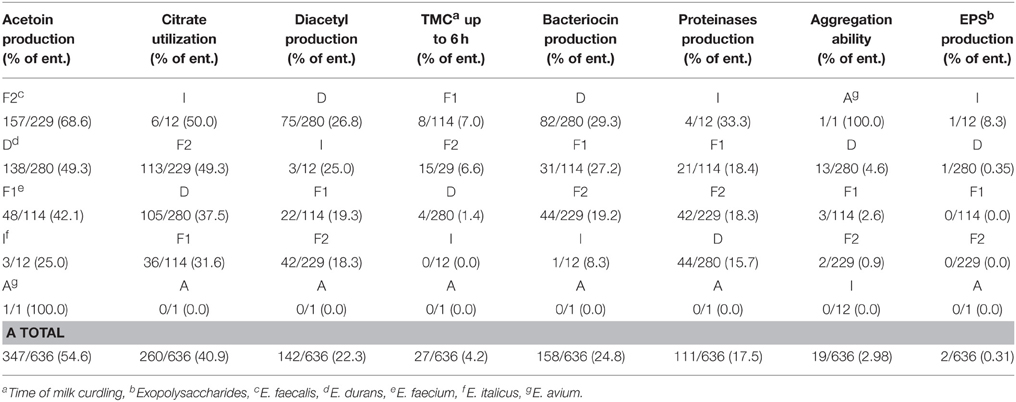
Table 4. Different technological activities in Enterococcus isolates from autochthonous dairy products.
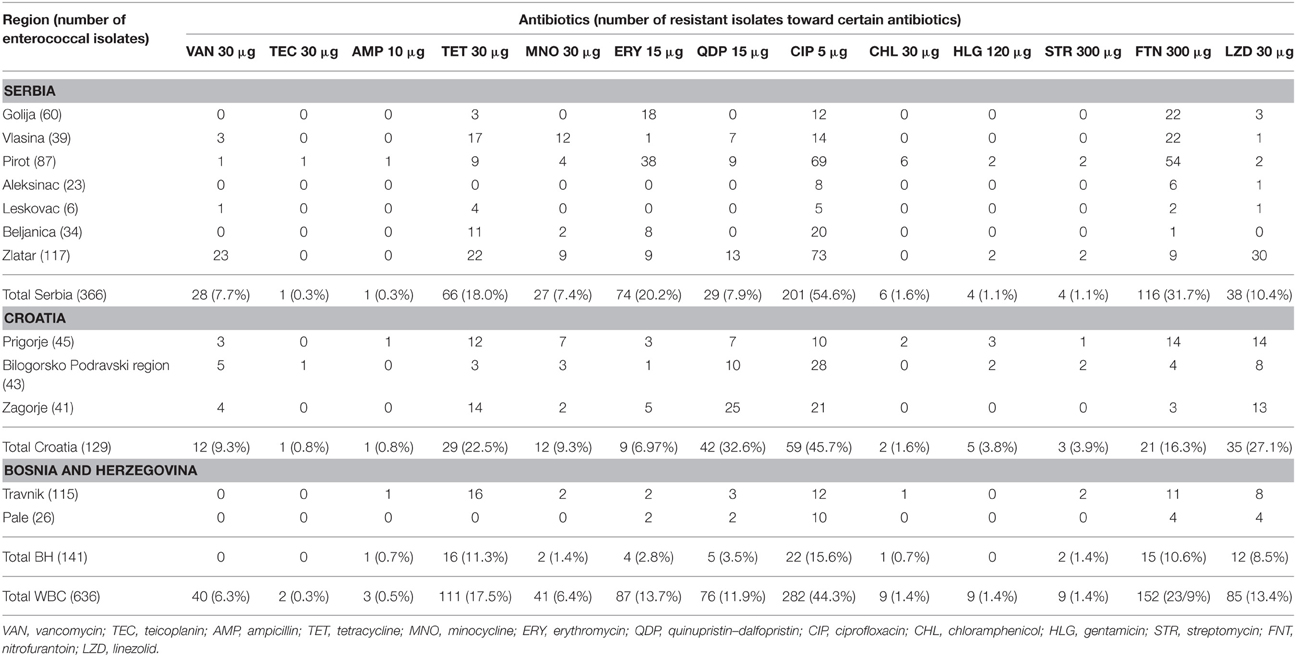
Antibiotic Susceptibility and Resistance
An important issue in the selection of strains for safe use in food production is the characterisation of their antibiotic susceptibility in order to avoid uncontrolled spreading of the antibiotic resistance genes through horizontal gene transfer. Susceptibility of all 636 enterococci isolates was examined by the agar disc diffusion method using 13 antibiotics: vancomycin, teicoplanin, ampicillin, erythromycin, tetracycline, minocycline, quinupristin–dalfopristin, ciprofloxacin, chloramphenicol, nitrofurantoin, linezolid, gentamicin (HLR), and streptomycin (HLR) (Table 5). The results of antibiotic susceptibility revealed that a total of 451 out of 636 isolates (70.9%) were resistant to at least one of the tested antibiotics. One hundred eighty-two (28.6%) isolates were resistant to one antibiotic (151 to two, 77 to three, 26 isolates to four, 7 isolates to five, 4 isolates to six, 3 isolates to seven antibiotics, while 1 isolate was resistant to even eight antibiotics). It is worthwhile to note that the most resistant isolates belonged to E. faecalis species. In general, the highest percent of the enterococci isolates shown to have multiple resistances to various antibiotics originated from dairy products sampled from Pirot, Zlatar Mountain and Vlasina Lake in Serbia and the Bilogorsko-Podravski region and Zagorje, Croatia. On the other hand, a high number of the strains were susceptible to all tested antibiotics (185/636). Eighty nine of them belonged to E. durans species, originating mainly from Golija Mountain, Serbia and Travnik, Bosnia and Herzegovina. Thirteen of 31 antibiotic sensitive E. faecalis strains were isolated from the region of Zlatar Mountain, Serbia, while 11 of 28 and 9 of 28 antibiotic-sensitive E. faecium strains were isolated from the Pale and Travnik regions, Bosnia and Herzegovina. In addition, four strains of E. italicus species were also sensitive to all tested antibiotics (data not shown).
Virulence Determinants and Biogenic Amines Production
Finally, 136 out of 185 antibiotic-susceptible isolates, with unique (GTG)5-fingerprint profiles, were further analyzed in order to choose the candidates with the best technological properties. Based on the phenotypic characteristics, 10 E. durans strains [BGGO6-15 (D+, C+, VP+), BGAL3-19 (Bac+, C+, VP±), BGTRM1-52 (Bac+, Prt+, VP+, D+, TMC 6.5 h), BGTRM7-39 (Bac+, D+, C+, VP±), BGTRS1-10 (Bac+, Prt+, D±), BGTRS7-54 (Bac+, Prt+, VP+, TMC 6.5 h), BGTRS10-42 (Bac+, D+, VP±), BGTRK10-29 (C+, VP+, D±), BGPAS1-80 (VP+, TMC 6 h), and ZGPR2-1 (Bac+, D+, C±)] and one E. italicus [BGTRK4-42 (Bac+, D+, VP±)] were chosen.
The PCR analysis for detection of virulence genes (see PCR Detection of Virulence Determinants for details) revealed that none of the virulence genes was found in any of the analyzed strains. Moreover, the results obtained in this study indicate that five out of the 11 chosen enterococci strains (BGAL3-19, BGTRS7-54, BGTRM7-39, BGTRS10-42, and ZGPR2-1) were not able to produce biogenic amines in the presence of the precursor amino acids histidine, lysine, ornithine, and tyrosine (Table 6).
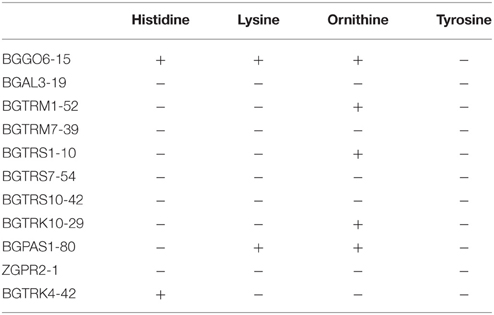
Table 6. Production of biogenic amines in the presence of the precursor amino acids histidine, lysine, ornithine, and tyrosine.
Discussion
Enterococci are found to be a normal part of the microbiota of artisan dairy products, especially in the Mediterranean region (Saavedra et al., 2003; Morandi et al., 2006; Psoni et al., 2006). However, their use as starter cultures in the dairy industry is still controversial since they have traditionally been considered indicators of fecal contamination. Moreover, enterococci are found to be involved in food spoilage (Franz et al., 1999) and food poisoning (Gardin et al., 2001), as well as in the spread of antibiotic resistance (Giraffa, 2002). Also, their role in the etiology of nosocomial infections cannot be neglected (Franz et al., 2003; Giraffa, 2003; Kayser, 2003). Hence, the aim of this paper was to evaluate the technological potential and safety issues for the use of natural dairy isolates of enterococci as starter cultures in the dairy industry. For that reason, in this study we have analyzed 636 natural isolates of Enterococcus sp. originating from various artisan dairy products collected in Western Balkan countries (WBC).
The results obtained in this study revealed the huge diversity among dairy enterococci isolates. The highest enterococci species diversity was recorded in fresh soft cheeses (1–10 days old) sampled from the Prigorje region, Croatia and Pirot, Serbia. Our previous results showed that enterococci and Leuconostoc sp. were the dominant species in the fresh soft cheeses of the Prigorje region, while lower diversity was found among other LAB species, indicating that enterococci were probably metabolically active and important for ripening of the artisan cheeses in that region (Golić et al., 2013).
Taking into account the diverse origins of the isolates, considerable genotypic heterogeneity was observed. The results showed that (GTG)5-fingerprint analysis was very effective in grouping and discriminating among enterococci strains, partly correlating with the sources of isolation. Hence, the results indicate that variable conditions, such as local vegetation or specific climates in the regions where dairy products are manufactured, significantly contribute to the diversity among enterococci strains. Interestingly, isolates belonging to the same species are found to be located in different clusters, indicating that identification by sequencing of 16S rDNA is not completely reliable in the case of natural isolates originating from complex communities such as dairy products. Possibly, horizontal transfer among bacteria living in the same ecological niche lead to their convergent/divergent evolution.
In order to explore the technological and functional potential of the natural enterococci isolates of artisan dairy origin, phenotypic variability was characterized. Due to various environmental pressures, natural isolates are usually shown to exhibit phenotypic variability (Giraffa et al., 2000, 2004). Enterococci play an important role in cheese ripening and contribute to the formation of the distinctive flavor of dairy products. Their ability to synthesize volatile compounds, such as diacetyl and acetoin, their proteolytic activity, and certainly antimicrobial activity are the important technological characteristics which make enterococci good candidates for starter and functional cultures for the dairy industry (Asteri et al., 2009; Nieto-Arribas et al., 2011). Enterococci usually exhibit weak proteolytic activities (Suzzi et al., 2000), and the best ability for casein degradation is shown by E. faecalis strains (Sarantinopoulos et al., 2001; Veljovic et al., 2009). Although the strains E. faecalis BGPT1-10P and BGPT1–78 were shown previously to have high activity in milk (Veljovic et al., 2009), in this study we found that only 17.5% of the analyzed strains exhibited proteolytic activity and it was equally distributed among E. durans (44/280), E. faecium (21/114), and E. faecalis (42/229) strains. According to the results of Morea et al. (1999) the degree of milk acidification by enterococci strains depends on the origin of the strain, hence it is highly likely that enterococci isolates of dairy origin are adapted to growth in milk. On the other hand, synthesis of diacetyl, acetoin from glucose, and citrate in the process of metabolic degradation by enterococci are shown to be very important for flavor formation in dairy products (Nieto-Arribas et al., 2011). Our analysis of enterococci dairy isolates from the WBC region showed that 347 out of 636 strains produced acetoin (54.6%), and 260/636 strains utilized citrate (40.9%), while 142/636 strains synthesized diacetyl (22.3%).
In addition, the excellent antibacterial activity shown by natural enterococci isolates make them promising candidates for food preservation and may contribute to the prevention of food spoilage (Viedma et al., 2009; Ananou et al., 2010). Bacteriocins produced by enterococci, enterocins, are very diverse and widely distributed among isolates. Production of enterocins in combination with a wide range of tolerance to high temperatures, dryness and increased salinity enables enterococci to become the dominant microbiota in fermented products (Franz et al., 1999). Our results support previous reports and indicate that the antimicrobial activity in natural dairy isolates from the WBC region showed a great effect on a number of pathogenic and non-pathogenic strains. Antimicrobial activity detected after treatment with protease suggests the proteinaceous nature of the bacteriocin activity. In our previous work it was shown that a number of E. faecalis strains exhibited an antimicrobial effect on L. innocua and Listeria monocytogenes (Veljovic et al., 2009). In particular, strains BGPT1-10P and BGPT1-78 showed antimicrobial activity against the Gram-negative strain Pseudomonas sp. PA17 and therefore could be used for the production of food biopreservatives (Veljovic et al., 2009). Apart from that, a number of E. durans and E. faecium strains exhibiting antimicrobial activity were found in this study.
Finally, with the aim of the safe use of enterococci as starter cultures in functional food, the frequency of virulence determinants and antibiotic resistance, as well as the synthesis of biogenic amines, was analyzed. The results showed that 185 out of 636 isolates (29.1%) were susceptible to the tested antibiotics, and as many as 59.6% of those isolates were resistant to two or more antibiotics. Interestingly, the presence of virulence determinants and antibiotic resistance is strain-dependent and region-specific. A high number of isolates from localities in Serbia (Zlatar Mountain, Pirot, and Vlasina Lake) had multiple antibiotic resistances. The results implicate the uncontrolled use of antibiotics in these regions, leading to antibiotic residues in feed or foods, which contribute to the occurrence of acquired and/or mutational antibiotic resistance mechanisms. More importantly, the further dissemination of antibiotic resistance genes among food-associated bacteria could cause the loss of natural isolates suitable for application in the dairy industry.
In order to eliminate the enterococci with pathogenic potential from among the possible candidates for use in dairy food production, and to avoid further transfer of virulence genes to other bacteria in the environment, identification of virulence factors is essential (Franz et al., 2003; Mannu et al., 2003). Fortunately, these important features are strain-specific, not species-specific. For this reason, each enterococcal strain intended for use in the dairy industry should be thoroughly tested for the absence of any pathogenic property. Ideally, a strain proposed for use in food production should not possess a single virulence factor and must be sensitive to relevant clinical antibiotics (Domig et al., 2003; Franz et al., 2003).
Virulence factors are commonly detected in clinical isolates of enterococci. Studies by Franz et al. (2001) and Eaton and Gasson (2001) have indicated the presence of virulence factors in food isolates, especially among strains of E. faecalis, which generally carry more virulence factors than strains of E. faecium. Our previous results showed that dairy strains E. faecalis BGPT1-10P and BGPT1-78 carry the genE gene and have gelatinase activity (Veljović et al., 2014). Moreover, in the strain BGPT1–10P a set of three genes of the cytolysin operon (cylM, cylB, and cylA) was detected, although two additional genes, cylL1 and cylL2, required for the expression of hemolytic activity, were not identified in this strain (Veljović et al., 2014). The genes for hemolytic activity (esp and efaA) are found in high frequency in all tested E. faecalis strains (Veljović et al., 2014). However, in this study we demonstrated that none of the 11 selected E. durans dairy strains had any of the tested virulence genes, making them good candidates for use in the dairy industry.
Besides the analysis of virulence determinants and antibiotic resistance, an analysis of biogenic amine synthesis was performed. Biogenic amines are organic bases with aliphatic, aromatic, and heterocyclic structures produced through decarboxylation of the corresponding amino acid (Giraffa, 2002). The capacity for amino acid decarboxylation restricts the use of the strains in the dairy industry. The previous results of Valenzuela et al. (2009) showed that E. faecalis BGPT1-10P has the ability of tyrosine and ornithine decarboxylation. The strains tested in this study did not have the ability of amino acid decarboxylation.
Conclusion
The results of this study reveal that autochthonous dairy products in the Western Balkan region are a rich source of new and diverse enterococci strains with considerable genetic, metabolic and technological potential. Since the role of enterococci in food spoilage and opportunistic infections is well known, before recommending the use of a particular strain as a starter culture it is necessary to characterize each strain in detail. This study reveals that out of 636 natural dairy enterococci isolates, only five strains belonging to E. durans have good technological potential and meet the safety criteria for use in the dairy industry. This finding points out the necessity for detailed characterization of enterococci isolated from dairy food (as a food of animal origin), since they could be reservoirs of antibiotic resistance and virulence genes, as well as producers of biogenic amines.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was funded by the Ministry of Education and Science of the Republic of Serbia, grant No. 173019. We are grateful to Nathaniel Aaron Sprinkle, native English editor for the proofreading of the manuscript.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2015.00954
Figure S1. Dendrogram based on statistical analysis of the (GTG)5-PCR fingerprints of Enterococcus sp. isolated from autochthonous dairy products from Serbia, Bosnia and Herzegovina and Croatia. The distances between the clusters were performed using “percent disagreement.” The algorithm “unweighted pair-group with average linkage” was used. The enterococci identified by 16S rDNA sequencing are given in bold.
References
Abeijón, M. C., Medina, R. B., Katz, M. B., and González, S. N. (2006). Technological properties of Enterococcus faecium isolated from ewe's milk and cheese with importance for flavour development. Can. J. Microbiol. 52, 237–245. doi: 10.1139/W05-136
Ananou, S., Garriga, M., Jofré, A., Aymerich, T., Gálvez, A., Maqueda, M., et al. (2010). Combined effect of enterocin AS-48 and high hydrostatic pressure to control food-borne pathogens inoculated in low acid fermented sausages. Meat Sci. 84, 594–600. doi: 10.1016/j.meatsci.2009.10.017
Andrighetto, C., Knijff, E., Lombardi, A., Torriani, S., Vancanneyt, M., Kersters, K., et al. (2001). Phenotypic and genetic diversity of enterococci isolated from Italian cheeses. J. Dairy Res. 68, 303–316. doi: 10.1017/S0022029901004800
Asteri, I. A., Robertson, N., Kagkli, D. M., Andrewes, P., Nychas, G., Coolbear, T., et al. (2009). Technological and flavour potential of cultures isolated from traditional Greek cheese-A pool of novel species and starters. Int. Dairy J. 19, 595–604. doi: 10.1016/j.idairyj.2009.04.006
Begovic, J., Brandsma, J. B., Jovcic, B., Tolinacki, M., Veljovic, K., Meijer, W. C., et al. (2011). Analysis of dominant lactic acid bacteria from artisanal raw milk cheeses produced on the mountain Stara Planina, Serbia. Arch. Biol. Sci. 63, 11–20. doi: 10.2298/ABS1101011B
Bover-Cid, S., and Holzapfel, W. H. (1999). Improved screening procedure for biogenic amine production by lactic acid bacteria. Int. J. Food Microbiol. 53, 33–41. doi: 10.1016/S0168-1605(99)00152-X
Centeno, J. A., Menéndez, S., Hermida, M. A., and Rodríguez-Otero, J. L. (1999). Effects of the addition of Enterococcus faecalis in Cebreiro cheese manufacture. Int. J. Food Microbiol. 48, 97–111. doi: 10.1016/S0168-1605(99)00030-6
CLSI. (2012). Performance Standards for Antimicrobial Susceptibility Testing, Vol. 32 (Clinical and Laboratory Standards Institute, Wayne, PA), 22nd Informational Supplement M100–S22.
Domig, K. J., Mayer, H. K., and Kneifel, W. (2003). Methods used for the isolation, enumeration, characterisation and identification of Enterococcus spp.: 1. Media for isolation and enumeration. Int. J. Food Microbiol. 88, 147–164. doi: 10.1016/S0168-1605(03)00177-6
Ducková, V., Èanigová, M., and Kroèko, M. (2014). Enterococci and their resistance to antibiotics and thyme essential oil. J. Microbiol. Biotechnol. Food Sci. 3, 1–4. Available online at: http://www.jmbfs.org/wp-content/uploads/2014/01/1_jmbfs_duckova_2014_m.pdf
Eaton, T. J., and Gasson, M. J. (2001). Molecular screening of Enterococcus virulence determinants and potential for genetic exchange between food and medical isolates. Appl. Environ. Microbiol. 67, 1628–1635. doi: 10.1128/AEM.67.4.1628-1635.2001
Foulquié-Moreno, M. R., Sarantinopoulos, P., Tsakalidou, E., and De Vuyst, L. (2006). The role of application of enterococci in food and health. Int. J. Food Microbiol. 106, 1–24. doi: 10.1016/j.ijfoodmicro.2005.06.026
Franz, C. M., Holzapfel, W. H., and Stiles, M. E. (1999). Enterococci at the crossroads of food safety? Int. J. Food Microbiol. 47, 1–24. doi: 10.1016/S0168-1605(99)00007-0
Franz, C. M., Huch, M., Abriouel, H., Holzapfel, W., and Gálvez, A. (2011). Enterococci as probiotics and their implications in food safety. Int. J. Food Microbiol. 151, 125–140. doi: 10.1016/j.ijfoodmicro.2011.08.014
Franz, C. M. A. P., Muscholl-Silberhorn, N., Yousif, M. K., Vancanneyt, M., Swings, J., and Holzapfel, W. H. (2001). Incidence of virulence factors and antibiotic resistance among enterococci isolated from food. Appl. Environ. Microbiol. 67, 4385–4389. doi: 10.1128/AEM.67.9.4385-4389.2001
Franz, C. M. A. P., Stiles, M. E., Schleifer, K. H., and Holzapfel, W. H. (2003). Enterococci in foods–a conundrum for food safety. Int. J. Food Microbiol. 88, 105–122. doi: 10.1016/S0168-1605(03)00174-0
Galgano, F., Suzzi, G., Favati, F., Caruso, M., Martuscelli, M., Gardini, F., et al. (2001). Biogenic amines during ripening in “Semicotto Caprino” cheese: role of enterococci. Int. J. Food Sci. Technol. 36, 153–160. doi: 10.1046/j.1365-2621.2001.00443.x
Gardin, F., Martuscelli, M., Caruso, M. C., Galgano, F., Crudele, M. A., Favati, F., et al. (2001). Effects of pH, temperature and NaCl concentration on the growth kinetics, proteolytic activity and biogenic amine production of Enterococcus faecalis. Int. J. Food Microbiol. 64, 105–117. doi: 10.1016/S0168-1605(00)00445-1
Garg, S. K., and Mital, B. K. (1991). Enterococci in milk and milk products. Crit. Rev. Microbiol. 18, 15–45. doi: 10.3109/10408419109113508
Gelsomino, R., Vancanneyt, M., Condon, S., Swings, J., and Cogan, T. M. (2001). Enterococcal diversity in the environment of an Irish Cheddar-type cheesemaking factory. Int. J. Food Microbiol. 171, 177–188. doi: 10.1016/S0168-1605(01)00620-1
Giraffa, G. (2002). Enterococci from foods. FEMS Microbiol. Rev. 26, 163–171. doi: 10.1111/j.1574-6976.2002.tb00608.x
Giraffa, G. (2003). Functionality of enterococci in dairy products. Int. J. Food Microbiol. 88, 215–222. doi: 10.1016/S0168-1605(03)00183-1
Giraffa, G., Andrighetto, C., Antonello, C., Gatti, M., Lazzi, C., Marcazzan, G., et al. (2004). Genotypic and phenotypic diversity of Lactobacillus delbrueckii subsp. lactis strains of dairy origin. Int. J. Food Microbiol. 91, 129–139. doi: 10.1016/S0168-1605(03)00368-4
Giraffa, G., Gatti, M., Rossetti, L., Senini, L., and Neviani, E. (2000). Molecular diversity within Lactobacillus helveticus as revealed by genotypic characterization. Appl. Environ. Microbiol. 66, 1259–1265. doi: 10.1128/AEM.66.4.1259-1265.2000
Golić, N., Cadež, N., Terzić-Vidojević, A., Suranská, H., Beganović, J., Lozo, J., et al. (2013). Evaluation of lactic acid bacteria and yeast diversity in traditional white pickled and fresh soft cheeses from the mountain regions of Serbia and lowland regions of Croatia. Int. J. Food Microbiol. 166, 294–300. doi: 10.1016/j.ijfoodmicro.2013.05.032
Gomes, B. C., Esteves, C. T., Palazzo, I. C., Darini, A. L., Felis, G. E., Sechi, L. A., et al. (2008). Prevalence and characterization of Enterococcus spp. isolated from Brazilian foods. Food Microbiol. 25, 668–675. doi: 10.1016/j.fm.2008.03.008
Hopwood, D. A., Bibb, M. J., Chater, K. F., Kieser, T., Bruton, C. J., Kieser, H. M., et al. (1985). Genetic Manipulation of Streptomyces: A Laboratory Manual. Norwich: John Innes Foundation.
Jovcic, B., Begovic, J., Lozo, J., Topisirovic, L., and Kojic, M. (2009). Dynamic of sodium dodecyl sulfate utilization and antibiotic susceptibility of strain Pseudomonas sp. ATCC19151. Arch. Biol. Sci. 61, 159–165. doi: 10.2298/ABS0902159J
Kayser, F. H. (2003). Safety aspects of enterococci from the medical point of view. Int. J. Food Microbiol. 88, 255–262. doi: 10.1016/S0168-1605(03)00188-0
Macedo, A. S., Freitas, A. R., Abreu, C., Machado, E., Peixe, L., Sousa, J. C., et al. (2011). Characterization of antibiotic resistant enterococci isolated from untreated waters for human consumption in Portugal. Int. J. Food Microbiol. 145, 315–319. doi: 10.1016/j.ijfoodmicro.2010.11.024
Mannu, L., Paba, A., Daga, E., Comunian, R., Zanetti, S., Dupré, I., et al. (2003). Comparison of the incidence of virulence determinants and antibiotic resistance between Enterococcus faecium strains of dairy, animal and clinical origin. Int. J. Food Microbiol. 88, 291–304. doi: 10.1016/S0168-1605(03)00191-0
Menéndez, S., Godinez, R., Hermida, M., Centeno, J. A., and Rodríguez-Otero, J. L. (2004). Characteristics of “Tetilla” pasteurized milk cheese manufactured with the addition of autochthonous cultures. Food Microbiol. 21, 97–104. doi: 10.1016/S0740-0020(03)00014-5
Morandi, S., Brasca, M., Andrighetto, C., Lombardi, A., and Lodi, R. (2006). Technological and molecular characterisation of enterococci isolated from north–west Italian dairy products. Int. Dairy J. 16, 867–875. doi: 10.1016/j.idairyj.2005.09.005
Morea, M., Baruzzi, F., and Cocconcelli, P. S. (1999). Molecular and physiological characterization of dominant bacterial populations in traditional Mozzarella cheese processing. J. Appl. Microbiol. 87, 574–582. doi: 10.1046/j.1365-2672.1999.00855.x
Nieto-Arribas, P., Seseña, S., Poveda, J. M., Chicón, R., Cabezas, L., and Palop, L. (2011). Enterococcus populations in artisanal Manchego cheese: Biodiversity, technological and safety aspects. Food Microbiol. 28, 891–899. doi: 10.1016/j.fm.2010.12.005
Psoni, L., Kotzamanides, C., Andrighetto, C., Lombardi, A., Tzanetakis, N., and Litopoulou-Tzanetaki, E. (2006). Genotypic and phenotypic heterogeneity in Enterococcus isolates from Batzos, a raw goat milk cheese. Int. J. Food Microbiol. 109, 109–120. doi: 10.1016/j.ijfoodmicro.2006.01.027
Saavedra, L., Taranto, M. P., Sesma, F., and Valdez, G. F. (2003). Homemade traditional cheeses for the isolation of probiotic Enterococcus faecium strains. Int. J. Food Microbiol. 88, 241–245. doi: 10.1016/S0168-1605(03)00186-7
Sarantinopoulos, P., Andrigheto, C., Georgalaki, M. D., Rea, M. C., Lombardi, A., Cogan, T. M., et al. (2001). Biochemical properties of enterococci relevant to their technological performance. Int. Dairy J. 11, 621–647. doi: 10.1016/S0958-6946(01)00087-5
Schleifer, K. H., and Ludwig, W. (1995). “Phylogenetic relationships of lactic acid bacteria,” in The Genera of Lactic Acid Bacteria, Vol. 2, eds. B. J. B. Wood and W. H. Holzapfel (London: Blackie Academic & Professional), 7–18.
Suzzi, G., Caruso, M., Gardini, F., Lombardi, A., Vannini, L., Guerzoni, M. E., et al. (2000). A survey of the enterococci isolated from an artisanal Italian goat's cheese (semicotto caprino). J. Appl. Microbiol. 89, 267–274. doi: 10.1046/j.1365-2672.2000.01120.x
Terzic-Vidojevic, A., Lozo, J., and Topisirovic, L. J. (2009b). Dominant lactic acid bacteria in artisanal Pirot cheeses of different ripening period. Genetika 41, 341–352. doi: 10.2298/GENSR0903341T
Terzic-Vidojevic, A., Mihajlović, S., Uzelac, G., Golić, N., Fira, Ð., Kojić, M., et al. (2014a). Identification and characterization of lactic acid bacteria isolated from artisanal white brined Golija cows' milk cheeses. Arch. Biol. Sci. 66, 179–192. doi: 10.2298/ABS1401179T
Terzic-Vidojevic, A., Mihajlovic, S., Uzelac, G., Veljovic, K., Tolinacki, M., Nikolic, M., et al. (2014b). Characterization of lactic acid bacteria isolated from artisanal Travnik young cheeses, sweet creams and sweet kajmaks over four seasons. Food Microbiol. 39, 27–38. doi: 10.1016/j.fm.2013.10.011
Terzic-Vidojevic, A., Tolinacki, M., Nikolic, M., Veljovic, K., Jovanovic, S., Macej, O., et al. (2013). Artisanal Vlasina raw goat's milk cheeses: evaluation and selection of autochthonous lactic acid bacteria as starter cultures. Food Technol. Biotechnol. 51, 554–563. Available online at: http://www.ftb.com.hr/images/pdfarticles/2013/October-december/ftb_51-4_554-563.pdf
Terzic-Vidojevic, A., Veljovic, K., Tolinacki, M., Nikolic, M., Ostojic, M., and Topisirovic, L. J. (2009a). Characterization of lactic acid bacteria isolated from artisanal Zlatar cheeses produced at two different geographical location. Genetika 41, 117–136. doi: 10.2298/GENSR0901117T
Terzic-Vidojevic, A., Vukasinovic, M., Veljovic, K., Ostojic, M., and Topisirovic, L. (2007). Characterization of microflora in homemade semi-hard white Zlatar cheese. Int. J. Food Microbiol. 114, 36–42. doi: 10.1016/j.ijfoodmicro.2006.10.038
Teuber, M., Meile, L., and Schwarz, F. (1999). Acquired antibiotic resistance in lactic acid bacteria from food. Antonie Van Leeuwenhoek 76, 115–137. doi: 10.1023/A:1002035622988
Tsakalidou, E., Manolopoulou, E., Tsilibari, V., Georgalaki, M., and Kalantzopoulos, G. (1993). Esterolytic activities of Enterococcus durans and Enterococcus faecium strains isolated from Greek cheese. Neth. Milk Dairy J. 47, 145–150.
Valenzuela, A. S., Ben Omar, N., Abriouel, H., López, R. L., Veljovic, K., Cañamero, M. M., et al. (2009). Virulence factors, antibiotic resistance, and bacteriocins in enterococci from artisan foods of animal origin. Food Control 20, 381–385. doi: 10.1016/j.foodcont.2008.06.004
Vankerckhoven, V., Autgaerden, T. V., Vael, C., Lammens, C., Chapelle, S., Rossi, R., et al. (2004). Development of a multiplex PCR for the detection of asa 1, gelE, cylA, esp, and hyl genes in enterococci and survey for virulence determinants among European hospital isolates of Enterococcus faecium. J. Clin. Microbiol. 42, 4473–4479. doi: 10.1128/JCM.42.10.4473-4479.2004
Veljovic, K., Fira, D., Terzic-Vidojevic, A., Abriouel, H., Galvez, A., and Topisirovic, L. (2009). Evaluation of antimicrobial and proteolytic activity of enterococci isolated from fermented products. Eur. Food Res. Technol. 230, 63–70. doi: 10.1007/s00217-009-1137-6
Veljović, K., Terzić-Vidojević, A., Tolinaèki, M., Mihajlović, S., Vukotić, G., Golić, N., et al. (2014). “Molecular characterization of natural dairy isolates of Enterococcus faecalis and evaluation of their antimicrobial potential,” in Enterococcus faecalis Molecular Characteristics, Role in Nosocomial Infections and Antimicrobial Effects. Bacteriology Research Developments, ed. H. L. Mack (New York, NY: Nova Publishers, Inc), 123–135.
Versalovic, J., Schneider, M., De Bruijn, F. J., and Lupski, J. R. (1994). Genomic fingerprinting of bacteria using repetitive sequence-based polymerase chain reaction. Method Mol. Cell. Biol. 5, 25–40.
Keywords: Enterococcus sp., artisan dairy products, diversity, antibiotic susceptibility, virulence
Citation: Terzić-Vidojević A, Veljović K, Begović J, Filipić B, Popović D, Tolinački M, Miljković M, Kojić M and Golić N (2015) Diversity and antibiotic susceptibility of autochthonous dairy enterococci isolates: are they safe candidates for autochthonous starter cultures? Front. Microbiol. 6:954. doi: 10.3389/fmicb.2015.00954
Received: 15 July 2015; Accepted: 28 August 2015;
Published: 09 September 2015.
Edited by:
Fausto Gardini, University of Bologna, ItalyReviewed by:
Stella Maris Reginensi Rivera, Universidad de la República, UruguayChiara Montanari, CIRI Agroalimentare, Italy
Copyright © 2015 Terzić-Vidojević, Veljović, Begović, Filipić, Popović, Tolinački, Miljković, Kojić and Golić. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Amarela Terzić-Vidojević, Laboratory for Molecular Microbiology, Institute of Molecular Genetics and Genetic Engineering, University of Belgrade, Vojvode Stepe 444a, PO Box 23, 11010 Belgrade, Serbia,YW1hcmVsYUBpbWdnZS5iZy5hYy5ycw==;bGFiNkBpbWdnZS5iZy5hYy5ycw==website: http://www.imgge.bg.ac.rs
 Amarela Terzić-Vidojević
Amarela Terzić-Vidojević Katarina Veljović
Katarina Veljović Jelena Begović1
Jelena Begović1 Brankica Filipić
Brankica Filipić Dušanka Popović
Dušanka Popović Maja Tolinački
Maja Tolinački Marija Miljković
Marija Miljković Milan Kojić
Milan Kojić Nataša Golić
Nataša Golić