- 1Laboratory of Veterinary Pharmacology, College of Veterinary Medicine, South China Agricultural University, Guangzhou, China
- 2Jiangsu Co-Innovation Centre for Prevention and Control of Important Animal Infectious Diseases and Zoonoses, Yangzhou, China
The purpose of this study was to characterize a collection of 103 multidrug resistance IncF plasmids recovered from Escherichia coli of food producing and companion animals between 2003 and 2012. A total of 103 incF plasmids were characterized using an established PCR-based IncF replicon sequence typing (RST) system to identify FII, FIA, and FIB (FAB) groups. Plasmids were also analyzed using-restriction fragment length polymorphism (RFLP). Antibiotic Resistance determinants blaCTX-M, plasmid-mediated quinolone resistance (PMQR) genes and rmtB and plasmid addiction systems (PAS) were identified by PCR screening. A total of 20 different RSTs from 103 IncF plasmids were identified. The groups F2 and F33 with the RST formulae A-: B- were the most frequently encountered types (63.1%). The antibiotic resistance genes (ARGs) blaCTX-M, rmtB, and oqxB were carried by 82, 37, and 34 IncF plasmids, respectively. Most of these plasmids carried more than one resistance gene (59.2%, 61/103). The IncF plasmids also had a high frequency of addiction systems (mean 2.54) and two antisense RNA-regulated systems (hok–sok and srnBC) and a protein antitoxin-regulated system (pemKI) were the most prevalent. Not surprisingly, RFLP profiles among the IncF plasmids were diverse even though some shared identical IncF-RSTs. This is the first extensive study of IncF plasmid-positive E. coli isolates from animals in China. Our results demonstrate that IncF is the most prevalent plasmid family in E. coli plasmids and they commonly carry multiple resistance determinants that render them resistant to different antibiotic classes simultaneously. IncF plasmids also harbor addiction systems, promoting their stability and maintenance in the bacterial host, under changing environmental conditions.
Introduction
Increasing antibiotic resistance in bacteria has potentially disastrous consequences for human and animal health around the world. Recent studies have demonstrated that plasmids act as efficient vehicles for the spread of antibiotic resistance genes (ARGs) more frequently than previously believed (Taylor et al., 2004; García-Fernández et al., 2009; Dolejska et al., 2011; Accogli et al., 2013; Carattoli, 2013; Dahmen et al., 2013b). Thus far, 27 different plasmid incompatibility (Inc.) groups are recognized in the Enterobacteriaceae (Couturier et al., 1988; Carattoli et al., 2005; Villa et al., 2010). A smaller number of particular plasmid families play a major role in the diffusion of specific resistance genes among Enterobacteriaceae and are the most frequently encountered. The IncF plasmids represent one of the most prevalent incompatibility types, and have been identified worldwide in Enterobacteriaceae from different origins and sources (Carattoli, 2009; Mathers et al., 2015). Currently, a total of 924 plasmids have been deposited at the plasmid multilocus sequence typing (pMLST) database (http://pubmlst.org/plasmid/, 30 January, 2015). Of these, 214 plasmids belong to IncF group and 158 of them (73.8%) recovered from Escherichiacoli could be classified in 90 different subgroups on the basis of their FII, FIA and FIB (FAB) alleles.
IncF plasmids have systems which guarantee their autonomous replication but also encode addiction systems frequently based on toxin-antitoxin factors. These systems contribute to control their copy number and ensurethe instable inheritance during cell division (Cohen, 1976; Kado, 1998; Hayes, 2003). In our previous studies, we found that IncF plasmids could integrate a wide range of genes conferring resistance to all major classes of antimicrobials including β-lactams, aminoglycosides, tetracyclines, chloramphenicol, and quinolones (Liao et al., 2013; Liu et al., 2013). The extended-spectrum β-lactamases (ESBLs), especially those of the CTX-M type, are often associated with plasmid-mediated quinolone resistance (PMQR) and aminoglycoside resistance genes. Plasmids belonging to the IncF group have been the primary hosts for these traits (Boyd et al., 2004; Yao et al., 2011; Matsumura et al., 2013; Ogbolu et al., 2013).
The spread of such multidrug resistance plasmids among Enterobacteriaceae strains could also affect clinical treatments. For instance, IncF-blaCTX-M association was observed from both human and animal E. coli isolates such as the IncF-like plasmid R100 involved in the dissemination of blaCTX-M-14 in Hong Kong, United Kingdom and France (Woodford et al., 2009; Ho et al., 2012; Dahmen et al., 2013a). The genes rmtB, qepA, qnr, fosA3, and oqxAB have been recently identified on IncF plasmids in E. coli from China, Korea and Spain (Tamang et al., 2008; Li et al., 2012; Ruiz et al., 2012; Ho et al., 2013). All together, the IncF-related family has the potential to be a major contributor worldwide to the diffusion of different resistance genes.
In general, the IncF plasmids are not a homogeneous group and vary in size (50–200 kb) and replicon type (Garcillán-Barcia et al., 2009; de Been et al., 2014; Lanza et al., 2014). Most importantly, plasmids encoding virulence-associated traits fall almost exclusively within IncF group (Johnson and Nolan, 2009). Therefore, knowledge of the prevalence and IncF plasmid subtypes in resistant populations could be a useful aid in exploring novel plasmid-targeting strategiesto treat resistant bacteria (Osborn et al., 2000; DeNap and Hergenrother, 2005; Baquero et al., 2011). We investigated variations in IncF plasmids and the phylogenetic relationship among IncF plasmids from different sources using RST and RFLP analyses. This study characterized IncF plasmids according to resistance genes, genetic relatedness and addiction systems in 103 E. coli isolates from pigs, poultry and companion animals.
Materials and Methods
Transconjugants and Antibiotic Susceptibility Testing
A total of non-duplicate 795 E. coli were isolated from animals between 2002 and 2012, according to the data published in our previous study (Liao et al., 2013; Liu et al., 2013, 2014). Among them, 696 E. coli were isolated from liver, heart or faces samples of diseased food-producing animals including 318 from avian (177 from ducks, 110 from chicken and 31 from geese) and 378 from pigs. The food-producing animals originated from more than 80 farms all over Guangdong province. The remaining 99 isolates were from rectal swab samples of pets including 71 dogs and 28 cats in five animal hospitals in Guangdong province. Further information about these animals, the underlying disease and possible antimicrobial pretreatment were unfortunately not available. One suspected colony with typical E. coli morphology and size was selected from all agar plates of each sample and was identified by Matrix Assisted Laser Desorption/lionization Time-of-Flight mass spectrometry (MALDI-TOF VITEK MS RUO System) using the Saramis™ database (bioMérieux, France). The genetic relatedness among partial E. coli isolates randomly selected were determined by pulsed-field gel electrophoresis (PFGE) with XbaI according to a protocol described previously (Gautom, 1997). Isolates encoding blaCTX−M, rmtB and/or PMQR gene (535/795), were selected for conjugation experiments by the broth-mating method using streptomycin-resistance E. coli C600 as the recipient. Transconjugants were selected on MacConkey agar plates containing streptomycin (1000 mg/L) and cefotaxime (2 mg/L)/amikacin (32 mg/L)/olaquindox (64 mg/L), respectively. The transconjugants harboring blaCTX-M, rmtB, and/or PMQR gene mentioned above were confirmed by PCR as previously.
One hundred and three transconjugants, each carrying one single IncF plasmid, were selected from a collection of clonally unrelated E. coli strains (15 pets, 43 avian, and 45 pigs) in our study. Table S1 contains detailed information for this population. Within this group 28 IncF plasmids had been previously characterized (Liao et al., 2013; Liu et al., 2013, 2014). Antimicrobial susceptibility tests were determined by agar dilution method on Mueller-Hinton agar plates. The antimicrobial drugs studied included quinolones (nalidixicacid), fluoroquinolones (ciprofloxacin, enrofloxacin, and levofloxacin), ampicillin, third-generation cephalosporins (ceftiofur, cefotaxime and ceftazidime), aminoglycoside (streptomycin, amikacin, kanamycin, and gentamycin) and other antimicrobials (olaquindox, chloramphenicol, florfenicol, and tetracyclines). The susceptibility tests were carried out and evaluated according to the guidelines of the Clinical and Laboratory Standards Institute (CLSI). The breakpoints for each antimicrobial were those recommended by the CLSI (M100-S19), CLSI (Vet01-A4/Vet01-S2), and DANMAP 98 (olaquindox) (Clinical and Laboratory Standards Institute, 2008, 2013). E. coli ATCC 25922 was used for quality control purposes.
Detection of Antimicrobial Resistance Genes
The total DNA of transconjugants harvested from LB broth were extracted by TIANPrep Plasmid Mini Kit (TIANGEN Biotech, Beijing, China), according to the manufacturer's instructions. The blaCTX-M, rmtB, and PMQR genes were screened by PCR using primers as described previously (Cano et al., 2009; Wang et al., 2012). The blaCTX-M, rmtB, and oqxB-positive transconjugants were also evaluated for the presence of other PMQR genes (qnrA, qnrB, qnrC, qnrD, qnrS, and qepA). PCR products were sequenced and compared with the reported sequences from GenBank (www.ncbi.nlm.nih.gov/genbank/).
IncF Plasmid Analysis
Plasmid DNA was extracted by QIAGENPrep Plasmid Midi Kit (QIAGEN, Hilden, Germany), according to the manufacturer's instructions. PCR-base replicon typing (PBRT) was performed on all transconjugants carrying a single plasmid, as previously described (Carattoli et al., 2005). To better characterize IncF, replicon sequencing typing (RST) was performed according to protocols described previously (Villa et al., 2010), and alleles were assigned by comparing the amplicon sequence to the plasmid MLST database (http://pubmlst.org/plasmid/).
Plasmid sizes in the transconjugants were determined by pulsed-field gel electrophoresis using nuclease S1 (TaKaRa Biotechnology, Dalian, China) digestion of plasmid DNA prepared in agarose blocks (S1-PFGE). DNA fragments were separated by PFGE on a CHEF-DR III apparatus (Bio-Rad, Hercules, CA, USA) for 15 h at 6 V/cm at 14°C with an initial pulse time of 1 s and a final pulse time of 12 s. Southern hybridization was performed following S1-PFGE using blaCTX-M, rmtB, and oqxB-specific digoxigen in (DIG)-labeled probes. Hybridization was detected using the DIG Nucleic Acid Detection Kit (Roche Diagnostics). All plasmids were also analyzed by restriction fragment length polymorphism (RFLP) as following: Plasmid DNA was digested with EcoRI and electrophoresed in 1.0% agarose at 5 V/cm for 2 h. DNA was visualized by EtBr staining. Cluster analysis of digestion patterns generated dendrograms and plasmid similarity was measured using BioNumerics software (Applied Maths, Sint-Martens-Latem, Belgium) using the Dice Similarity Index (DSI) (tolerance 1.5% and optimization 1.5%. Plasmids with a DSI ≥80% were assigned to the same cluster (designated with numbers 1, 2, 3, and 4). Letters were used to discriminate RFLP patterns assigned to the same clusterthat differed by one or two restriction bands (i.e., 1a, 1b, 1c etc.).
Identification of Plasmid Addiction Systems
Plasmid-mediated addiction systems were identified using primers and amplification conditions were previously described (Mnif et al., 2010). We screened for 8 major addiction systems: ccdA-ccdB (involved in cell division), pemK-pemI (for plasmid emergency maintenance), relB-relE (relaxed control of stable RNA), parD-parE (DNA replication), vagC-vagD (virulence-associated protein), hok-hok (host-killing), snrB-snrC (RNA stability), and pndA-pndC (promotion of nucleic acid).
Statistical Analysis
Statistical significance for the comparison of prevalence data and proportions was determined by the x2-test. p-values less than 0.05 were deemed to be statistically significant.
Results
Antibiotic Resistance Phenotypes
A total of 103 transconjugants were examined in this study and 96% exhibited multi-drug resistance phenotypes with to ampicillin and streptomycin. More than half of the isolates also showed resistance tocefotaxime, ceftriaxone, gentamicin, kanamycin and ceftiofur (Table S1). The antimicrobial resistance rates to other antibiotics tested as follows: amikacin (43.7%), olaquindox (42.7%), tetracycline (40.8%), chloramphenicol (34.0%), and florfenicol (26.2%). Most of these isolates were still susceptible to levofloxacin, enrofloxacin and ciprofloxacin.
Detection of Resistance Genes
Results of screening for resistance genes among 103 transconjugants are shown in Table S1. The predominant resistance genes identified were blaCTX-M, rmtB, and oqxB found in 82, 37, and 34 IncF-type plasmids, respectively. Interestingly, there was a range of blaCTX-M genotypes including M55 (n = 30), M14 (n = 20), M27 (n = 14), M65 (n = 13), M15 (n = 3) and M24 and M3 (1 each). A smaller subset also harbored qnrS, qepA, and qnrB genes (11, 6, and 3, respectively). Sixty-one plasmids carried more than one resistance gene, and the most prevalent combinations were blaCTX-M-rmtB (n = 28) and blaCTX-M-oqxAB (n = 15). The qnrA, qnrC, and qnrD loci were not found in any of the plasmids tested.
Characterization of IncF Plasmids
S1-PFGE and Southern blot hybridization analysis showed that all the 103 multi-drug resistant strains contained plasmids ranging in size from 60 to 194 kb (Table 1). This group contained very diverse IncF-RST patterns. A total of 20 IncF-RSTs were found that had replicon FII alone or in combination with FIA or FIB. Various alleles were observed for each of FII (F2, F14, F16, F18, F31, F33, F35, F36, F43, F46, and F58), FIA (A-, A1, A3, and A4) and FIB (B-, B1, B8, B10, and B24) replicons. Meanwhile, we found identical combinations were detected in a number of different plasmids; F2: A-: B- (n = 35) and F33: A-: B- (n = 30) were the most common. Curiously, more than half of F2: A-: B- positive E. coli strains (54.8%, 19/35) were recovered from pigs, while F33: A-: B- (46.7%, 14/30) and F18: A-: B- (50.0%, 5/10) were mainly found in E. coli strains recovered from avians.
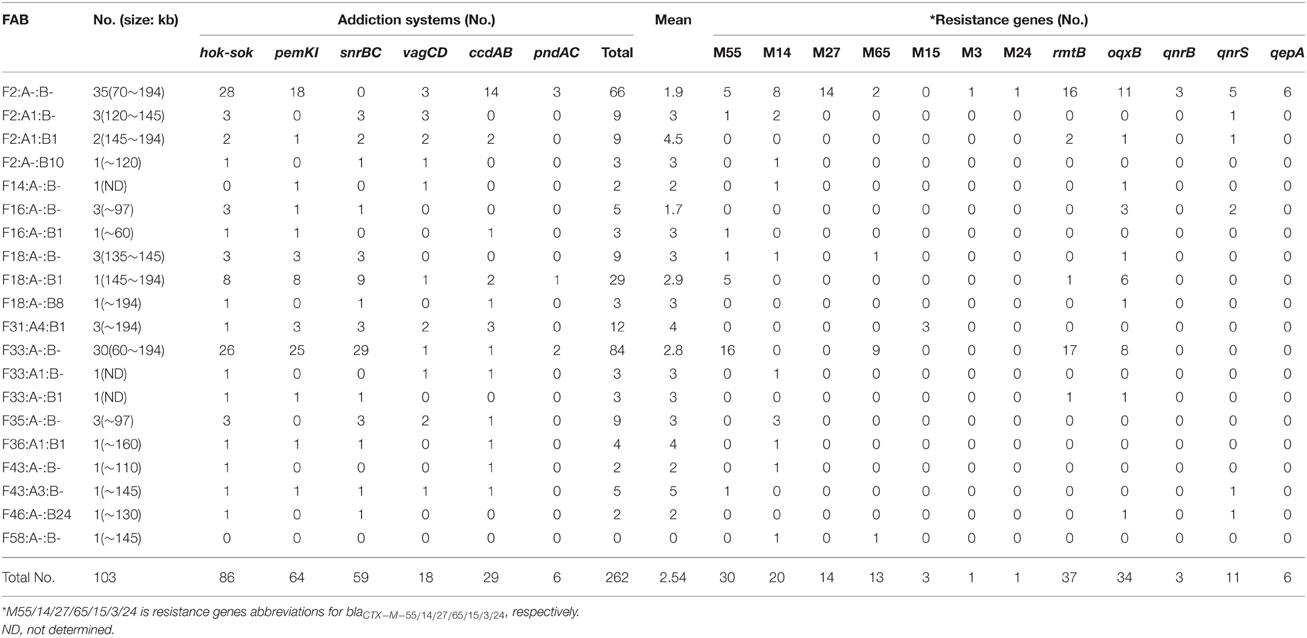
Table 1. The distribution of addiction systems and resistance genes according to FAB type identified in the 103 multi-resistance plasmids.
Further analysis of IncF plasmids bearing multi-resistance genes was performed to see whether a particular resistance gene was associated with a specific plasmid backbone defined by replicon type and addiction system content. Notably, there was higher prevalence of resistance genes located on F2: A- B- and F33: A-: B- plasmids than on the remaining replicon types (Table 1). For instance, blaCTX-M-27, qepA, and qnrS were preferentially associated with the F2: A-: B-. The blaCTX-M-55, which was the most common identified β-lactamase in our study, combined with F33: A-: B-; while rmtB and oqxB were mainly found in F2: A-: B- and F33: A-: B-.
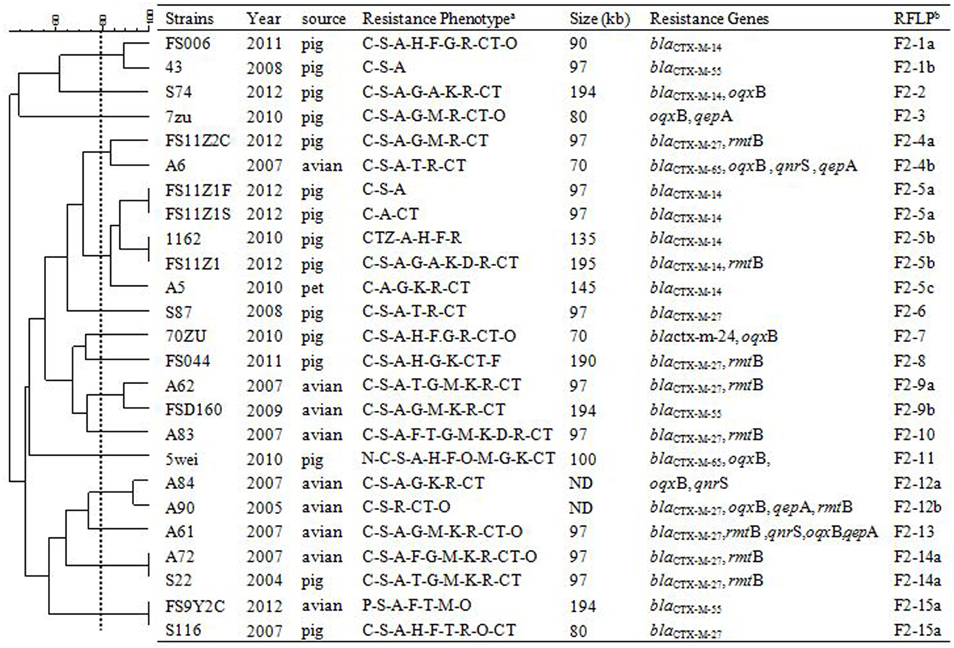
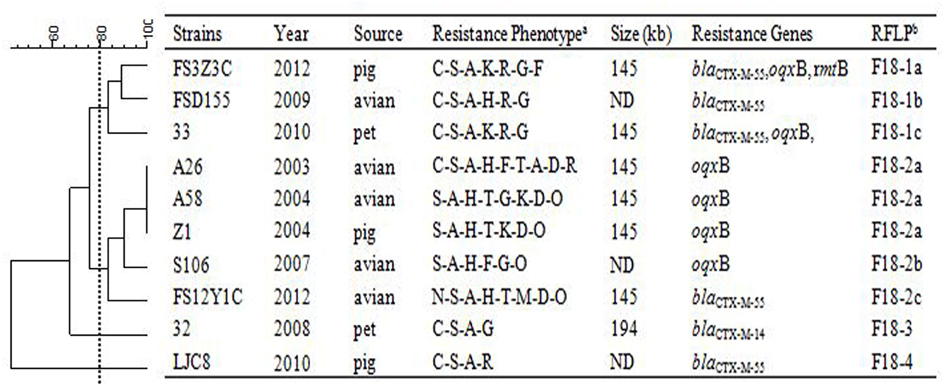
In order to assess relationships among the different replicon types of 103 IncF plasmids encoding variable resistance determinants, RFLP analysis was conducted which allowed visualization of variable clustering. However, our RFLP results showed diverse plasmid profiles even among identical IncF-RST groups. For instance, among 27/30 F33: A-: B- and 25/35 F2: A-: B- plasmids, 12 and 15 RFLP patterns were identified, respectively. In addition, antimicrobial susceptibility, plasmid size and antimicrobial resistance genes varied (Figures 1–3).
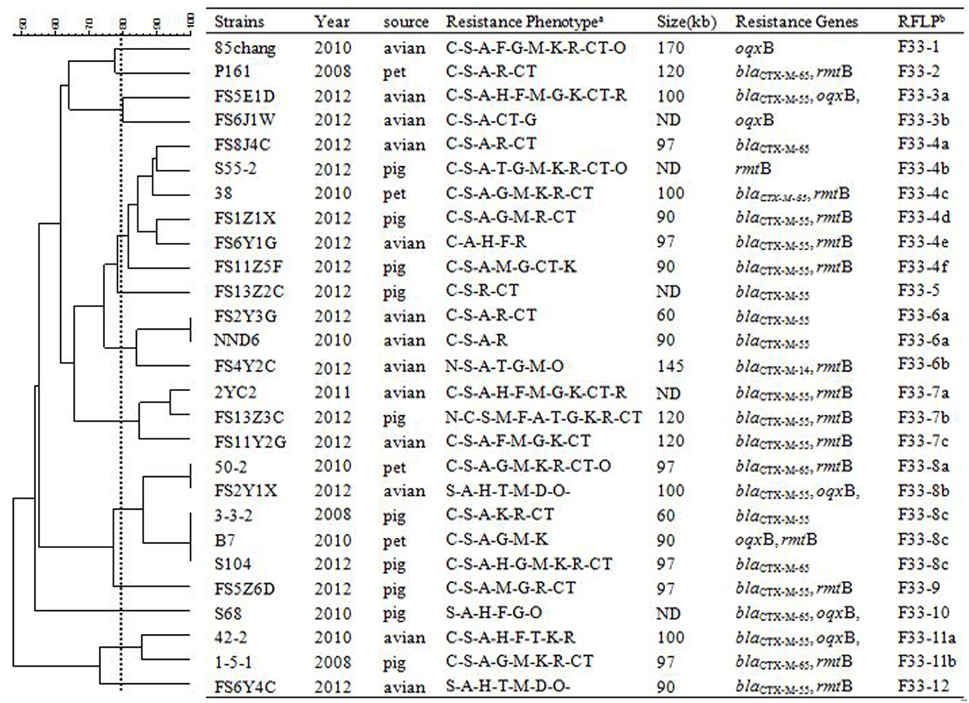
Figure 3. RFLP patterns and characterization of multi-resistance 27 F33: A-: B- plasmids. The scale on the top left of the figure indicates the percentage of similarity for the EcoRI restriction profiles of the IncF plasmids. aAntimicrobial for which the plasmids MICs fell within the resistant range; Antimicrobial abbreviations: A, ampicillin; S, streptomycin; M, amikacin; K, kanamycin; G, gentamycin; R, ceftriaxone; C, cefotaxime; CT, ceftiofur; N, nalidixic acid; P, ciprofloxacin; E, enrofloxacin; L, levoflox-acin; O, olaquindox; T, tetracycline; H, chloramphenicol and F, florfenicol. bPlasmids with a DSI ≥80% were assigned to the same cluster (designated with numbers 1, 2, 3, 4 etc.). Letters were used to discriminate RFLP patterns assigned to the same cluster, but differing by one or two restriction bands (i.e., 1a, 1b, 1c etc.). ND, not determined.
Addiction Systems
Of the eight addiction systems, the most frequently represented systems were hok-sok (n = 86), followed by pemKI (n = 64), snrBC (n = 59), ccdAB (n = 29), vagCD (n = 18), and pndAC (n = 6). None of the IncF-type plasmids harbored parDE and relBE. The average mean of addiction system was 2.54 with a range of 0–5 per plasmid. Interestingly, when the replicon FII was associated with FIA and FIB, the isolate also carried more addiction systems than in the absence of this association (Table 1). For instance, between the groups F2: A1: B1 and F2: A-: B-, there was a mean of 4.5 vs. 1.9, and the combination of F16: A-: B1 and F16: A-: B- with the mean of 3.0 vs. 1.7.
The number of different types of addiction systems also varied between the most prevalence replicon types in this study, F2: A-: B- and F33: A-: B-. The number of snrBC loci on F33: A-: B- was significantly higher than on F2: A-: B- (P < 0.01). In contrast, this number was reversed in the case of the ccdAB system (Table 1).
Discussion
In the present study, the prevalence of the resistance determinants blaCTX-M, PMQR genes and rmtB in IncF plasmids from farm animals and pets in China was investigated and the characteristics of IncF plasmids were elucidated. One-hundred and three IncF plasmids were recovered, the majority from pigs and avians (85.4%, 88/103), and the remaining obtained from pets (14.6%, 15/103), demonstrating that IncF plasmids are prevalent among multi-drug resistant E. coli strains from animals in China. The IncF plasmids identified in our study were associated with multidrug resistance profiles that conferred resistance to different classes of antimicrobials simultaneously. Most of IncF plasmids harbored blaCTX-M and, frequently carried other resistance genes including qnr, qepA, oqxB, and rmtB. These results are in agreement with previous findings that the rmtB and PMQR genes are often linked with blaCTX-M-located on the same plasmid (Dhanji et al., 2011; Li et al., 2012; Matsumura et al., 2013; Chen et al., 2014). Together, these data suggest that the spread of multiple ARGs on the same plasmid has been an important dissemination mechanism of multidrug resistance (Liebert et al., 1999).
The most impressive finding in this study was the diversity of IncF plasmids. The result is similar to previous reports that showed IncF plasmids are not a homogeneous group (Osborn et al., 2000; Coque et al., 2008; Villa et al., 2010; Shin et al., 2012). Moreover, heterogeneous RFLP patterns among identical IncF-RST groups indicate that the cargos such as resistance genes are also likely associated with the diversity of IncF plasmids. Interestingly, we identified some particular resistance genes were associated with the specific plasmid backbone defined by replicon type. For instance, blaCTX-M-3 was located on F2: A-: B-, whereas it has been previously on detected IncL/M plasmids (Golebiewski et al., 2007). As for blaCTX-M-14, which was primarily found on F2: A-: B- in our study, but in Spain and France, this gene was presented on the IncK plasmids (Diestra et al., 2009; Valverde et al., 2009). Although the movements of these resistance genes were attributed to transposable elements, it is certain that the epidemic plasmids may accelerate their widespread dissemination.
Interestingly, although RFLP patterns were diverse among the plasmids, some with the same IncF-RST grouping did show similar RFLP patterns with minor variations. These data agree with previous studies reporting that IncF plasmids form a very heterogeneous group (Dahmen et al., 2013b). Thus, the extensive diversity of RFLP patterns among IncF plasmids from the collection of E. coli used in this study may be due to frequent recombination and acquisition resistance genes between the plasmids (Partridge et al., 2011; Wiedenbeck and Cohan, 2011; Cain and Hall, 2012).
In contrast to RFLP patterns, the content of addiction systems seemed to differ by replicon type. For instance, the snrBC system was widely represented in F33: A-: B- but absent in F2: A-: B-, while ccdAB was associated with F2: A-: B-. This finding indicates a linkage between addiction systems and specific plasmid backbones, but this must be investigated further. As previously reported, plasmid addiction systems (PAS) contribute to the stability and maintenance of plasmids and they have been shown to facilitate plasmid dissemination even in the absence of antibiotic selection (Hayes, 2003; Moritz and Hergenrother, 2007). Together, these data suggest a hypothesis that the diversity of IncF plasmids can be driven by a combination of acquired multiple resistance genes and addiction modules. In this way, host maintenance would be facilitated by their spread in the E. coli population.
In conclusion, we identified extensive diversity of RSTs, plasmid sizes, addiction systems and RFLP patterns among IncF plasmids, carrying blaCTX-M or PMQR or aminoglycoside resistance genes. Considering the fact that the IncF plasmids have been common reservoirs for diffusion of resistance genes, its wide dissemination among Enterobacteriaceae especially among E. coli could pose a serious threat to public health.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by the Program for Changjiang Scholars and Innovative Research Team in University of Ministry of Education of China (Grant No. IRT13063), the National Natural Science Foundation and Natural Science Foundation of Guangdong Province, China (Grant No. U1201214), the Natural Science Foundation of Guangdong Province (Grant No. S2012030006590) and the National Natural ScienceFunds of China (Grant No. 31402247).
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2015.00964
References
Accogli, M., Fortini, D., Giufrè, M., Graziani, C., Dolejska, M., Carattoli, A., et al. (2013). IncI1 plasmids associated with the spread of CMY-2, CTX-M-1 and SHV-12 in Escherichia coli of animal and human origin. Clin. Microbiol. Infect. 19, E238–E240. doi: 10.1111/1469-0691.12128
Baquero, F., Coque, T. M., and de la Cruz, F. (2011). Ecology and evolution as targets: the need for novel eco-evo drugs and strategies to fight antibiotic resistance. Antimicrob. Agents Chemother. 55, 3649–3660. doi: 10.1128/AAC.00013-11
Boyd, D. A., Tyler, S., Christianson, S., McGeer, A., Muller, M. P., Willey, B. M., et al. (2004). Complete nucleotide sequence of a 92-kilobase plasmid harboring the CTX-M-15 extended-spectrum beta-lactamase involved in an outbreak in long-term-care facilities in Toronto, Canada. Antimicrob. Agents Chemother. 48, 3758–3764. doi: 10.1128/AAC.48.10.3758-3764.2004
Cain, A. K., and Hall, R. M. (2012). Evolution of a multiple antibiotic resistance region in IncHI1 plasmids: reshaping resistance regions in situ. J. Antimicrob. Chemother. 67, 2848–2853. doi: 10.1093/jac/dks317
Cano, M. E., Rodriguez-Martinez, J. M., Aguero, J., Pascual, A., Calvo, J., Garcia-Lobo, J. M., et al. (2009). Detection of plasmid-mediated quinolone resistance genes in clinical isolates of Enterobacter spp. in Spain. J. Clin. Microbiol. 47, 2033–2039. doi: 10.1128/JCM.02229-08
Carattoli, A. (2009). Resistance plasmid families in Enterobacteriaceae. Antimicrob. Agents Chemother. 53, 2227–2238. doi: 10.1128/AAC.01707-08
Carattoli, A. (2013). Plasmids and the spread of resistance. Int. J. Med. Microbiol. 303, 298–304. doi: 10.1016/j.ijmm.2013.02.001
Carattoli, A., Bertini, A., Villa, L., Falbo, V., Hopkins, K. L., and Threlfall, E. J. (2005). Identification of plasmids by PCR-based replicon typing. J. Microbiol. Methods 63, 219–228. doi: 10.1016/j.mimet.2005.03.018
Chen, X., He, L., Li, Y., Zeng, Z., Deng, Y., Liu, Y., et al. (2014). Complete sequence of a F2:A-:B- plasmid pHN3A11 carrying rmtB and qepA, and its dissemination in China. Vet Microbiol. 174, 267–271. doi: 10.1016/j.vetmic.2014.08.023
Clinical Laboratory Standards Institute. (2008). Performance Standards for Antimi-crobial Susceptibility Testing; Eighteenth Informational Supplement. Document M100-S18. Wayne, PA: CLSI.
Clinical Laboratory Standards Institute. (2013). Performance Standards for Antimi-crobial Disk and Dilution Susceptibility Tests for Bacteria Isolated from Animals; Approved Standard-fourth Edition and Supplement. Documents VET01-A4E and VET01-S2E. Wayne, PA: CLSI.
Cohen, S. N. (1976). Transposable genetic elements and plasmid evolution. Nature 263, 731–738. doi: 10.1038/263731a0
Coque, T. M., Novais, A., Carattoli, A., Poirel, L., Pitout, J., Peixe, L., et al. (2008). Dissemination of clonally related Escherichia coli strains expressing extended-spectrum beta-lactamase CTX-M-15. Emerg. Infect. Dis. 14, 195–200. doi: 10.3201/eid1402.070350
Couturier, M., Bex, F., Bergquist, P. L., and Maas, W. K. (1988). Identification and classification of bacterial plasmids. Microbiol. Rev. 52, 375–395.
Dahmen, S., Madec, J. Y., and Haenni, M. (2013a). F2: A-: B- plasmid carrying the extended-spectrum beta-lactamase bla(CTX−M−55∕57) gene in Proteus mirabilis isolated from a primate. Int. J. Antimicrob. Agents 41, 594–595. doi: 10.1016/j.ijantimicag.2013.02.004
Dahmen, S., Métayer, V., Gay, E., Madec, J. Y., and Haenni, M. (2013b). Characterization of extended-spectrum beta-lactamase (ESBL)-carrying plasmids and clones of Enterobacteriaceae causing cattle mastitis in France. Vet Microbiol. 162, 793–799. doi: 10.1016/j.vetmic.2012.10.015
de Been, M., Lanza, V. F., de Toro, M., Scharringa, J., Dohmen, W., Du, Y., et al. (2014). Dissemination of cephalosporin resistance genes between Escherichia coli strains from farm animals and humans by specific plasmid lineages. PLoS Genet. 10:e1004776. doi: 10.1371/journal.pgen.1004776
DeNap, J. C., and Hergenrother, P. J. (2005). Bacterial death comes full circle: targeting plasmid replication in drug-resistant bacteria. Org. Biomol. Chem. 3, 959–966. doi: 10.1039/b500182j
Dhanji, H., Murphy, N. M., Akhigbe, C., Doumith, M., Hope, R., Livermore, D. M., et al. (2011). Isolation of fluoroquinolone-resistant O25b:H4-ST131 Escherichia coli with CTX-M-14 extended-spectrum beta-lactamase from UK river water. J. Antimicrob. Chemother. 66, 512–516. doi: 10.1093/jac/dkq472
Diestra, K., Juan, C., Curiao, T., Moyá, B., Mirá, E., Oteo, J., et al. (2009). Characterization of plasmids encoding blaESBL and surrounding genes in Spanish clinical isolates of Escherichia coli and Klebsiella pneumoniae. J. Antimicrob. Chemother. 63, 60–66. doi: 10.1093/jac/dkn453
Dolejska, M., Duskova, E., Rybarikova, J., Janoszowska, D., Roubalova, E., Dibdakova, K., et al. (2011). Plasmids carrying blaCTX−M−1 and qnr genes in Escherichia coli isolates from an equine clinic and a horseback riding centre. J. Antimicrob. Chemother. 66, 757–764. doi: 10.1093/jac/dkq500
García-Fernández, A., Fortini, D., Veldman, K., Mevius, D., and Carattoli, A. (2009). Characterization of plasmids harbouring qnrS1, qnrB2 and qnrB19 genes in Salmonella. J. Antimicrob. Chemother. 63, 274–281. doi: 10.1093/jac/dkn470
Garcillán-Barcia, M. P., Francia, M. V., and de la Cruz, F. (2009). The diversity of conjugative relaxases and its application in plasmid classification. FEMS Microbiol. Rev. 33, 657–687. doi: 10.1111/j.1574-6976.2009.00168.x
Gautom, R. K. (1997). Rapid pulsed-field gel electrophoresis protocol for typing of Escherichia coli O157:H7 and other gram-negative organisms in 1 day. J. Clin. Microbiol. 35, 2977–2980.
Golebiewski, M., Kern-Zdanowicz, I., Zienkiewicz, M., Adamczyk, M., Zylinska, J., Baraniak, A., et al. (2007). Complete nucleotide sequence of the pCTX-M-3 plasmid and its involvement in spread of the extended-spectrum beta-lactamase gene blaCTX−M−3. Antimicrob. Agents Chemother. 51, 3789–3795. doi: 10.1128/AAC.00457-07
Hayes, F. (2003). Toxins-antitoxins: plasmid maintenance, programmed cell death, and cell cycle arrest. Science 301, 1496–1499. doi: 10.1126/science.1088157
Ho, P. L., Chan, J., Lo, W. U., Law, P. Y., and Chow, K. H. (2013). Plasmid-mediated fosfomycin resistance in Escherichia coli isolated from pig. Vet. Microbiol. 162, 964–967. doi: 10.1016/j.vetmic.2012.09.023
Ho, P. L., Lo, W. U., Yeung, M. K., Li, Z., Chan, J., Chow, K. H., et al. (2012). Dissemination of pHK01-like incompatibility group IncFII plasmids encoding CTX-M-14 in Escherichia coli from human and animal sources. Vet Microbiol. 158, 172–179. doi: 10.1016/j.vetmic.2012.02.004
Johnson, T. J., and Nolan, L. K. (2009). Pathogenomics of the virulence plasmids of Escherichia coli. Microbiol. Mol. Biol. Rev. 73, 750–774. doi: 10.1128/MMBR.00015-09
Kado, C. I. (1998). Origin and evolution of plasmids. Antonie Van Leeuwenhoek 73, 117–126. doi: 10.1023/A:1000652513822
Lanza, V. F., de Toro, M., Garcillán-Barcia, M. P., Mora, A., Blanco, J., Coque, T. M., et al. (2014). Plasmid flux in Escherichia coli ST131 sublineages, analyzed by plasmid constellation network (PLACNET), a new method for plasmid reconstruction from whole genome sequences. PLoS Genet. 10:e1004766. doi: 10.1371/journal.pgen.1004766
Li, D. X., Zhang, S. M., Hu, G. Z., Wang, Y., Liu, H. B., Wu, C. M., et al. (2012). Tn3-associated rmtB together with qnrS1, aac(6′)-Ib-cr and blaCTX−M−15 are co-located on an F49:A-:B- plasmid in an Escherichia coli ST10 strain in China. J. Antimicrob. Chemother. 67, 236–238. doi: 10.1093/jac/dkr428
Liao, X. P., Liu, B. T., Yang, Q. E., Sun, J., Li, L., Fang, L. X., et al. (2013). Comparison of plasmids coharboring 16s rrna methylase and extended-spectrum beta-lactamase genes among Escherichia coli isolates from pets and poultry. J. Food Prot. 76, 2018–2023. doi: 10.4315/0362-028X.JFP-13-200
Liebert, C. A., Hall, R. M., and Summers, A. O. (1999). Transposon Tn21, flagship of the floating genome. Microbiol. Mol. Biol. Rev. 63, 507–522.
Liu, B. T., Li, L., Fang, L. X., Sun, J., Liao, X. P., Yang, Q. E., et al. (2014). Characterization of plasmids carrying oqxAB in bla-negative Escherichia coli isolates from food-producing animals. Microb. Drug Resist. 20, 641–650. doi: 10.1089/mdr.2014.0022
Liu, B. T., Yang, Q. E., Li, L., Sun, J., Liao, X. P., Fang, L. X., et al. (2013). Dissemination and characterization of plasmids carrying oqxAB-blaCTX−M genes in Escherichia coli isolates from food-producing animals. PLoS ONE 8:e73947. doi: 10.1371/journal.pone.0073947
Mathers, A. J., Peirano, G., and Pitout, J. D. (2015). The role of epidemic resistance plasimds and international high-risk clones in the spread of multidrug-resistant in Enterobacteriaceae. Clin. Microbiol. Rev. 28, 565–591. doi: 10.1128/CMR.00116-14
Matsumura, Y., Yamamoto, M., Nagao, M., Ito, Y., Takakura, S., Ichiyama, S., et al. (2013). Association of fluoroquinolone resistance, virulence genes, and IncF plasmids with extended-spectrum-beta-lactamase-producing Escherichia coli sequence type 131 (ST131) and ST405 clonal groups. Antimicrob. Agents Chemother. 57, 4736–4742. doi: 10.1128/AAC.00641-13
Mnif, B., Vimont, S., Boyd, A., Bourit, E., Picard, B., Branger, C., et al. (2010). Molecular characterization of addiction systems of plasmids encoding extended-spectrum beta-lactamases in Escherichia coli. J. Antimicrob. Chemother. 65, 1599–1603. doi: 10.1093/jac/dkq181
Moritz, E. M., and Hergenrother, P. J. (2007). Toxin-antitoxin systems are ubiquitous and plasmid-encoded in vancomycin-resistant enterococci. Proc. Natl. Acad. Sci. U.S.A. 104, 311–316. doi: 10.1073/pnas.0601168104
Ogbolu, D. O., Daini, O. A., Ogunledun, A., Terry Alli, O. A., and Webber, M. A. (2013). Dissemination of IncF plasmids carrying beta-lactamase genes in Gram-negative bacteria from Nigerian hospitals. J. Infect. Dev. Ctries. 7, 382–390. doi: 10.3855/jidc.2613
Osborn, A. M., da Silva Tatley, F. M., Steyn, L. M., Pickup, R. W., and Saunders, J. R. (2000). Mosaic plasmids and mosaic replicons: evolutionary lessons from the analysis of genetic diversity in IncFII-related replicons. Microbiology 146, 2267–2275. doi: 10.1099/00221287-146-9-2267
Partridge, S. R., Zong, Z., and Iredell, J. R. (2011). Recombination in IS26 and Tn2 in the evolution of multi-resistance regions carrying blaCTX-M-15 on conjugative IncF pla-smids from Escherichia coli. Antimicrob. Agents Chemother. 55, 4971–4978. doi: 10.1128/AAC.00025-11
Ruiz, E., Sáenz, Y., Zarazaga, M., Rocha-Gracia, R., Martínez-Martinez, L., Arlet, G., et al. (2012). qnr, aac(6′)-Ib-cr and qepA genes in Escherichia coli and Klebsiella spp.: genetic environments and plasmid and chromosomal location. J. Antimicrob. Chemother. 67, 886–897. doi: 10.1093/jac/dkr548
Shin, J., Choi, M. J., and Ko, K. S. (2012). Replicon sequence typing of IncF plasmids and the genetic environments of blaCTX−M−15 indicate multiple acquisitions of blaCTX−M−15 in Escherichia coli and Klebsiella pneumoniae isolates from South Korea. J. Antimicrob. Chemother. 67, 1853–1857. doi: 10.1093/jac/dks143
Tamang, M. D., Seol, S. Y., Oh, J. Y., Kang, H. Y., Lee, J. C., Lee, Y. C., et al. (2008). Plasmid-mediated quinolone resistance determinants qnrA, qnrB, and qnrS among clinical isolates of Enterobacteriaceae in a Korean hospital. Antimicrob. Agents Chemother. 52, 4159–4162. doi: 10.1128/AAC.01633-07
Taylor, D. E., Gibreel, A., Lawley, T. D., and Tracz, D. M. (2004). “Antibiotic resistance plasmids,” in Plasmid Biology, eds B. Funnell and G. Phillips (Washington, DC: ASM Press), 473–491.
Valverde, A., Cantón, R., Garcillán-Barcia, M. P., Novais, A., Galán, J. C., Alvarado, A., et al. (2009). Spread of bla(CTX-M-14) is driven mainly by IncK plasmids disseminated among Escherichia coli phylogroups A, B1, and D in Spain. Antimicrob. Agents Chemother. 53, 5204–5212. doi: 10.1128/AAC.01706-08
Villa, L., García-Fernández, A., Fortini, D., and Carattoli, A. (2010). Replicon sequence typing of IncF plasmids carrying virulence and resistance determinants. J. Antimicrob. Chemother. 65, 2518–2529. doi: 10.1093/jac/dkq347
Wang, Y., He, T., Han, J., Wang, J., Foley, S. L., Yang, G., et al. (2012). Prevalence of ESBLs and PMQR genes in fecal Escherichia coli isolated from the non-human primates in six zoos in China. Vet Microbiol. 159, 53–59. doi: 10.1016/j.vetmic.2012.03.009
Wiedenbeck, J., and Cohan, F. M. (2011). Origins of bacterial diversity through horizontal genetic transfer and adaptation to new ecological niches. FEMS Microbiol. Rev. 35, 957–976. doi: 10.1111/j.1574-6976.2011.00292.x
Woodford, N., Carattoli, A., Karisik, E., Underwood, A., Ellington, M. J., and Livermore, D. M. (2009). Complete nucleotide sequences of plasmids pEK204, pEK499, and pEK516, encoding CTX-M enzymes in three major Escherichia coli lineages from the United Kingdom, all belonging to the international O25:H4-ST131 clone. Antimicrob. Agents Chemother. 53, 4472–4482. doi: 10.1128/AAC.00688-09
Keywords: Escherichia coli, IncF plasmid, multi-resistance, addiction systems, RFLP
Citation: Yang Q-E, Sun J, Li L, Deng H, Liu B-T, Fang L-X, Liao X-P and Liu Y-H (2015) IncF plasmid diversity in multi-drug resistant Escherichia coli strains from animals in China. Front. Microbiol. 6:964. doi: 10.3389/fmicb.2015.00964
Received: 18 April 2015; Accepted: 31 August 2015;
Published: 22 September 2015.
Edited by:
Alexandre Gonçalves, University of Trás-os-Montes e Alto Douro, PortugalReviewed by:
Teresa M. Coque, Hospital Universitario Ramón y Cajal, SpainLei Dai, Iowa State University, USA
Maria Jose Gosalbes, CIBER, Spain
Copyright © 2015 Yang, Sun, Li, Deng, Liu, Fang, Liao and Liu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ya-Hong Liu, Laboratory of Veterinary Pharmacology, College of Veterinary Medicine, South China Agricultural University, 483 Wushan Street, Tianhe District, Guangzhou 510642, China,Z2FsZUBzY2F1LmVkdS5jbg==
†Present Address: Bao-Tao Liu, College of Animal Science and Technology, Qingdao Agricultural University, Qingdao, China
‡These authors have contributed equally to this work.
 Qiu-E. Yang1‡
Qiu-E. Yang1‡ Jian Sun
Jian Sun Liang Li
Liang Li Bao-Tao Liu
Bao-Tao Liu Xiao-Ping Liao
Xiao-Ping Liao