- 1CAS Key Laboratory of Tropical Marine Bio-Resources and Ecology, RNAM Center for Marine Microbiology, South China Sea Institute of Oceanology, Chinese Academy of Sciences, Guangzhou, China
- 2College of Earth Science, University of Chinese Academy of Sciences, Beijing, China
Actinobacteria is a ubiquitous major group in coral holobiont. The diversity and spatial and temporal distribution of actinobacteria have been rarely documented. In this study, diversity of actinobacteria associated with mucus, tissue and skeleton of Porites lutea and in the surrounding seawater were examined every 3 months for 1 year on Luhuitou fringing reef. The population structures of the P. lutea-associated actinobacteria were analyzed using phylogenetic analysis of 16S rRNA gene clone libraries, which demonstrated highly diverse actinobacteria profiles in P. lutea. A total of 25 described families and 10 unnamed families were determined in the populations, and 12 genera were firstly detected in corals. The Actinobacteria diversity was significantly different between the P. lutea and the surrounding seawater. Only 10 OTUs were shared by the seawater and coral samples. Redundancy and hierarchical cluster analyses were performed to analyze the correlation between the variations of actinobacteria population within the divergent compartments of P. lutea, seasonal changes, and environmental factors. The actinobacteria communities in the same coral compartment tended to cluster together. Even so, an extremely small fraction of OTUs was common in all three P. lutea compartments. Analysis of the relationship between actinobacteria assemblages and the environmental parameters showed that several genera were closely related to specific environmental factors. This study highlights that coral-associated actinobacteria populations are highly diverse, and spatially structured within P. lutea, and they are distinct from which in the ambient seawater.
Introduction
Coral reef ecosystem is one of the most important tropical marine ecosystems, mainly distributed in the Indo-West Pacific, Eastern Pacific, Western Atlantic, and the Eastern Atlantic (Moberg and Folke, 1999). Corals provide habitats for numerous bacteria in their mucus layer, tissue, and calcium carbonate skeleton, as well as the dinoflagellates, fungi, archaea, and viruses (Rosenberg et al., 2007). Coral-associated bacteria not only take part in carbon, nitrogen, and sulfur biogeochemical cycles and provide necessary nutrient for coral, but also keep corals from being infected by pathogens (Rosenberg et al., 2007; Raina et al., 2009; Bourne and Webster, 2013).
Highly diverse and heterogeneous bacterial communities have been revealed in different coral species at various locations (Rohwer et al., 2002; Li et al., 2013). Actinobacteria is generally accepted as a ubiquitous major group in corals (Bourne and Munn, 2005; Carlos et al., 2013; Li et al., 2013, 2014a). Yang et al. (2013) detected 19 Actinobacteria genera in soft coral Alcyonium gracllimum and stony coral Tubastraea coccinea in the East China Sea through analysis of 16S rRNA gene clone libraries. Some actinobacterial genera were previously detected in corals by using the culture-dependent method (Lampert et al., 2006; Nithyanand and Pandian, 2009; Nithyanand et al., 2011b; Zhang et al., 2013; Li et al., 2014b). Among these culturable actinobacteria, Streptomyces, Verrucosispora, Rhodococcus, Micromonospora, Nocardia, Jiangella, Nocardiopsis, Pseudonocardia, and Salinispora showed antibacterial activities, which were considered to contribute to coral health (Ritchie, 2006; Nithyanand et al., 2011a; Krediet et al., 2013; Zhang et al., 2013; Li et al., 2014b).
Environmental conditions, coral species, colony physiology, and seasonal variation are considerable influencing factors on the coral-associated bacterial community (Hong et al., 2009). Moreover, due to various microhabitats provided by corals' biological structures, the spatial heterogeneity has been proved in bacterial communities associated with a single coral colony (Rohwer et al., 2002; Sweet et al., 2011; Li et al., 2014a). As a major coral-associated bacterial group, how actinobacteria is spatially and temporally organized in corals, and what is the connection between the actinobacteria communities in corals and in seawater remains poorly understood. Comprehensive investigation of the distribution of this ubiquitous group at spatial and temporal scales will help understanding the variation of coral associated bacteria and the potential function of actinobacteria, and will contribute a lot to bioprospect the actinobacteria resources for utilization as novel sources for bioactive natural products.
Coral reefs are widely distributed in the South China Sea (Liu et al., 2009; Wang et al., 2014). Porites lutea is the dominant, typical coral species in the Luhuitou fringing reef, which is located in the south end of the Hainan province (Zhao et al., 2008). In this study, the diversity and distribution of actinobacteria were investigated in coral P. lutea and in the surrounding seawater every 3 months for 1 year using culture-independent method for the first time. We aimed to reveal the coral-associated actinobacteria community structures in three divergent coral compartments in different months, compare the actinobacterial communities in the coral and in the surrounding seawater, and research the actinobacteria community variation responds to the environmental factors.
Materials and Methods
Sample Collection
The coral and surrounding sea water samples were collected in four different months (February, May, August, and November) in 2012 from the Luhuitou fringing reef (109°28′E, 18°13′N). Coral fragments (approximately 10 × 10 cm) were collected from the side of three healthy P. lutea colonies at the depth of 3–5 m each time using punch and hammer. Coral mucus, tissues and skeleton were separated and stored according to the method described previously (Li et al., 2014a). One liter of seawater adjacent to the coral colonies was collected, and filtered through 0.22 μm-pore-size filter membrane (Millipore). The filter membranes were stored at −80°C until DNA extraction. As the samples were collected at the same time, environmental parameters including water temperature, salinity, dissolved oxygen, pH value, ultraviolet radiation intensity, and rainfall were cited from the published data (Li et al., 2014a).
DNA Extraction and PCR Amplification
The coral tissue and skeleton samples were homogenized thoroughly in liquid nitrogen with sterile mortar and pestle before added to the PowerBead Tubes. The filter membranes with adsorbed microbial cells were cut into pieces, and then added to the PowerBead Tubes. Total DNA was extracted using the PowerSoil DNA Isolation Kit (MoBio, Solana Beach, CA, USA) according to the manufacturer's instruction.
16S rRNA genes were nest PCR amplified, the first PCR reactions using the combination of universal bacterial primers 27F (5′-AGAGTTTGATCMTGGCTCAG-3′) and 1492R (5′-TACGGYTACCTTGTTACGACTT-3′). PCR amplifications were performed in a Mastercycler pro (Eppendorf, Hamburg, Germany) in a final volume of 50 μL, containing 2 μL (10 μM) each primer, 1 μL (10–20 ng) template DNA and 25 μL premix Ex Taq mixture (Takara, Dalian). The PCR conditions were as follows: 94°C for 5 min; 30 cycles of 94°C for 30 s, 54°C for 30 s, 72°C for 90 s; followed by 72°C for 10 min. In the second PCR reactions, the actinobacteria-specific primer pairs, S-C-Ac-0325-a-S-20 (5′-CGCGCCTATCAGCTTGTTG-3′) and S-C-Act-0878-a-A-19 (5′-CCGTATCCCCAGGCGGGG-3′), were used to amplify the V3-V5 regions (about 640 bp) of the actinobacteria 16S rRNA gene (Stach et al., 2003). In the PCR reactions, 5 μL of 1: 10 dilution of the first round PCR product was used as DNA template, the PCR mixture (50 μL) contain 2 μL (10 μM) each primer, 25 μL premix Ex Taq mixture, the PCR conditions were as follows: 95°C for 5 min; 30 cycles of 95°C for 45 s, 68°C for 45 s, 72°C for 60 s; followed by 72°C for 10 min. Each genomic DNA sample was amplified in triplicate PCR reactions. Amplicons from the same sample were pooled and purified using the E.Z.N.A.® Gel Extraction Kit (Omega Bio-Tek, China).
Gene Library Construction and Sequencing
Sixteen clone libraries of actinobacterial 16S rRNA genes were constructed using the pMD18-T Vector Cloning Kit and E. coli DH5α competent cells (Takara, Dalian) following the manufacturer′s instructions. The positive clones from each library inoculated on MacConkey agar with ampicillin (100 μg/ml) were randomly picked and sequenced using M13F (−47) primer on ABI 3730xl capillary sequencers (Applied Biosystems, USA).
Libraries Analysis
The vector sequences were screened by the VecScreen tool provided in NCBI (http://www.ncbi.nlm.nih.gov/tools/vecscreen/). Chimeras were checked by running chimera.uchime packaged in Mothur (Schloss et al., 2009), and potential chimeras were removed. All valid sequences were deposited in GenBank (accession numbers were shown in Data S1). All qualified sequences were identified by using the classify.seqs command in Mothur with Silva reference alignment database (http://www.mothur.org/wiki/Silva_reference_files, Release 119) at a confidence level of 80%. The sequences, which do not belong to Actinobacteria, were removed from further analysis. Sequences were clustered into operational taxonomic units (OTUs) with a 97% threshold using the cluster command in Mothur. The relationships among actinobacterial communities associated with different coral compartments and in the ambient seawater in different months were analyzed by hierarchical cluster analysis. Based on Bray-Curtis similarity estimated from the OTU matrix, clustering was generated by using the complete linkage method with the PRIMER 5 software (Clarke, 1993). The shared OTUs were determined by using the online tool venny (Oliveros, 2007–2015, http://bioinfogp.cnb.csic.es/tools/venny/index.html).
The correlations between Actinobacteria assemblages of coral samples and the environmental factors were analyzed by using the software package CANOCO 4.5.1 (ter Braak and Šmilauer, 2002). Redundancy analysis (RDA) was carried out to determine the relationship between the actinobacteria community and the environmental factors including temperature, salinity, dissolved oxygen, pH value, rainfall, and UV radiation and in combination with two nominal variables including the coral divergent compartments and the different sampling months. The significance of the relation between the explanatory variables and the actinobacterial community compositions was tested using Monte Carlo permutation tests (9999 unrestricted permutations, P < 0.05).
Results
Coral-associated Actinobacteria Diversity
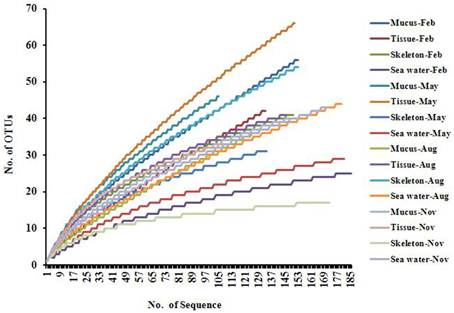
A total of 2403 sequences were obtained from sixteen 16S rRNA gene clone libraries, resulting in 395 OTUs (stringency at 97%). The rarefaction analysis result showed that most of the curves did not flatten to asymptote, but climbed less steeply (Figure 1). The coverages ranged from 0.69 to 0.97 in 16 libraries, and the average coverage was 0.83 (Table 1). The highest number of OTUs was found in the tissue collected in May, while the lowest OTUs was found in the skeleton collected in November (Table 1). The Shannon indices in mucus collected in different months ranged from 2.32 to 3.44, from 2.45 to 3.55 in tissues, from 1.82 to 3.35 in skeleton, and from 1.53 to 2.82 in sea water (Table 1), and the diversity in the actinobacterial community associated with P. lutea was higher than which in the surrounding sea water (P = 0.045).
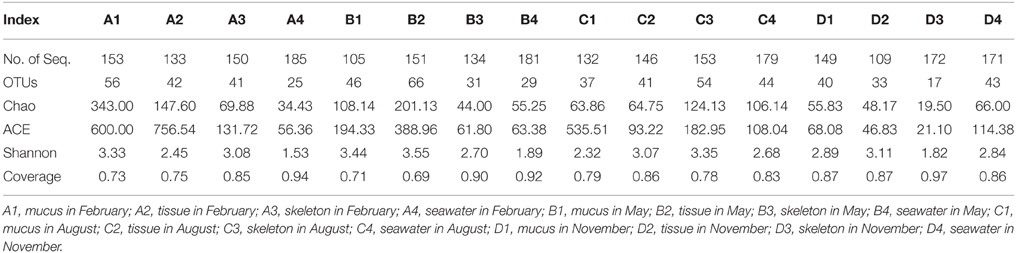
Table 1. Number of sequences and OTUs (97%) and diversity estimates of the Actinobacteria libraries in P. lutea and in the ambient seawater.
Coral-associated Actinobacterial Community Composition
At a confidence threshold of 80%, 2403 qualified reads were assigned to four classes, i.e., Acidimicrobiia, Actinobacteria, Thermoleophilia, and KIST-JJY010. Among them, Acidimicrobiia and Actinobacteria were ubiquitous and dominant in P. lutea and in the seawater samples. Thermoleophilia was not detected in corals collected in February, in the mucus and seawater in May, and in the mucus in August, while accounted for 0.5–48.8% in all other samples. KIST-JJY010 was detected only in the mucus in November (0.6%), and in the skeleton in August (2.6%).
Twenty-five described families and 10 unnamed families were detected in the 16 libraries (Figure 2). OM1_clade and Propionibacteriaceae (genera Friedmanniella and Propionibacterium) were ubiquitous, major groups in P. lutea. Meanwhile, OM1_clade was not detected in the seawater in February and May, and rare in the other two seawater libraries, and Propionibacteriaceae was absent in all the seawater libraries. Micromonosporaceae was the most abundant group in the tissue in February (47.4%) and in the mucus in August (46.2%), in which most of the reads were affiliated with an unclassified group. Nonetheless, Micromonosporaceae was absent in all other coral and seawater samples. Sva0996_marine_group was detected in all coral samples (5.2–50%) except in the skeleton collected in November, and which also was abundant in the ambient sea water (21.9–80%). Micrococcaceae was absent in the coral skeleton collected in August and in November, and in the sea water samples. Group 480-2 was abundant in the coral tissue in August (24.7%), as well as in the skeleton in May (26.9%) and in November (48.8%), but it was nearly absent in the surrounding seawater. In reverse, Microbacteriaceae and Ilumatobacter were major groups in sea water, while they were less abundant in P. lutea.
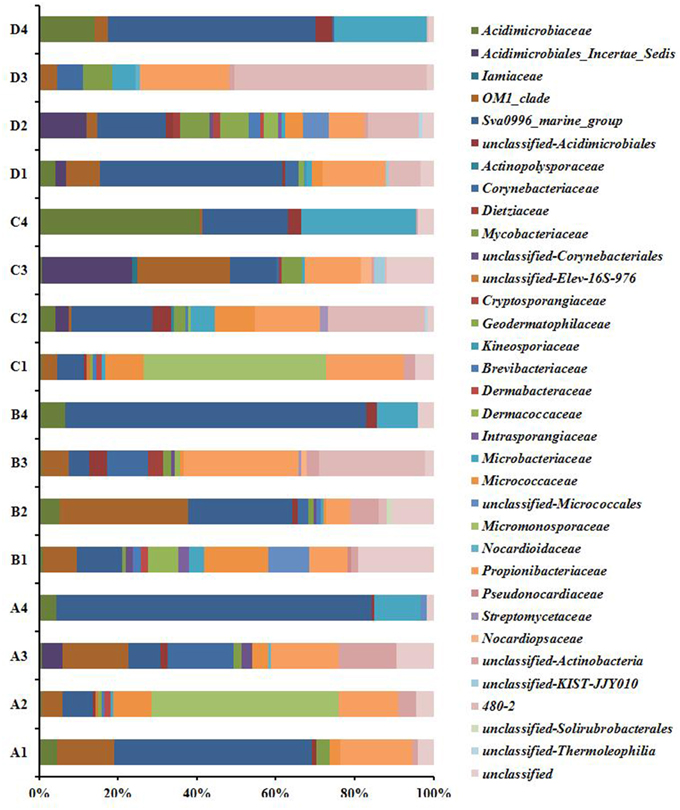
Figure 2. Actinobacteria composition profiles. Taxonomic classification of actinobacteria sequences in to family identified by using the classify.seqs command in Mothur using Silva reference alignment database (http://www.mothur.org/wiki/Silva_reference_files, Release 119) with a confidence level of 80% were applied for classification. A1, mucus in February; A2, tissue in February; A3, skeleton in February; A4, seawater in February; B1, mucus in May; B2, tissue in May; B3, skeleton in May; B4, seawater in May; C1, mucus in August; C2, tissue in August; C3, skeleton in August; C4, seawater in August; D1, mucus in November; D2, tissue in November; D3, skeleton in November; D4, seawater in November.
Spatial and Temporal Distribution of P. lutea-associated Actinobacteria
Results of hierarchical cluster analysis showed that the actinobacteria communities were significantly different between in the coral and in the surrounding seawater samples (p = 0.01, R = 0.993). The actinobacterial communities associated with the same coral compartments tended to cluster together (Figure 3). The season factor did not significantly influence the variation in the actinobacteria communities. The RDA results indicated that 38.9% of the total variance in the coral-associated actinobacterial composition was explained by the environmental, spatial and temporal factors (Figure 4). The first and second axes differentiated the actinobacteria assemblages in the distinct coral compartments (Figure 4, Table S1). This result was consistent with the hierarchical cluster analysis. None of the environment parameters analyzed in this study was determined as the significant influencing factor in the variation of the P. lutea associated actinobacteria communities. A triplot map illustrated the relationship between major actinobacterial groups, with abundance more than 1%, and the environmental parameters (Figure 4). Friedmanniella and Micrococcus were positively related with the salinity. Microbacterium, Propionibacterium, and group 480-2 were positively correlated with seawater temperature, but negatively correlated with dissolved oxygen.
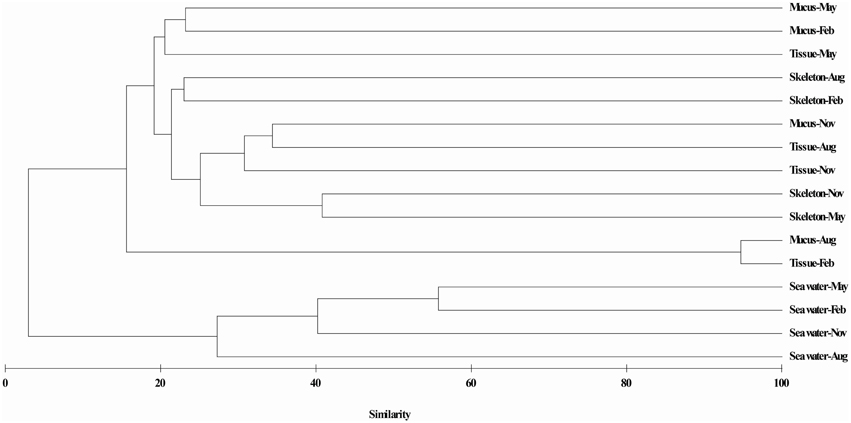
Figure 3. Hierarchical cluster analysis of actinobacteria communities associated with P. lutea. Clustering was based on Bray-Curtis similarity estimated from the OTUs matrix by using the complete linkage method.
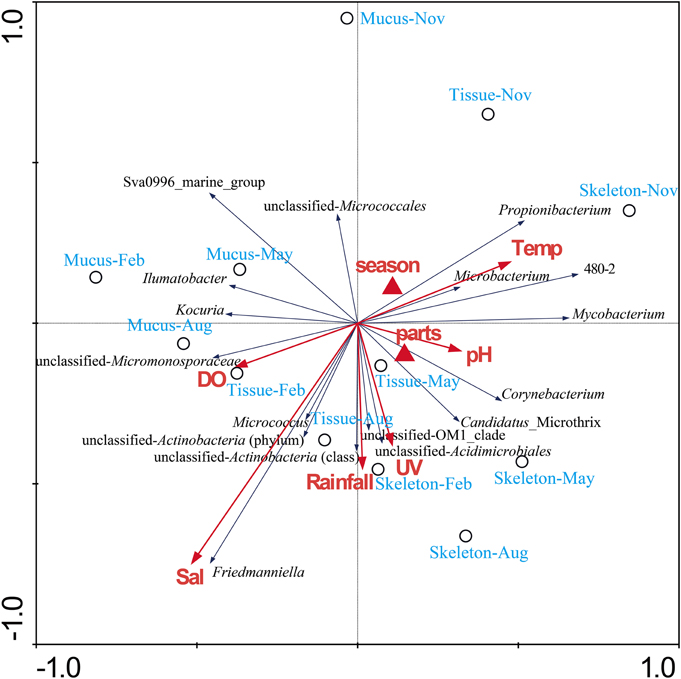
Figure 4. RDA ordination triplot showing the relationship among the environmental variables, coral samples, and actinobacterial components. Correlations between environmental variables and the first two RDA axes are represented by the lengths and angles of the arrows (environmental-factor vectors). Only abundant actinobacterial groups (>1%) were showed in the triplot. UV, ultraviolet radiation intensity; Temp, seawater temperature; DO, dissolved oxygen.
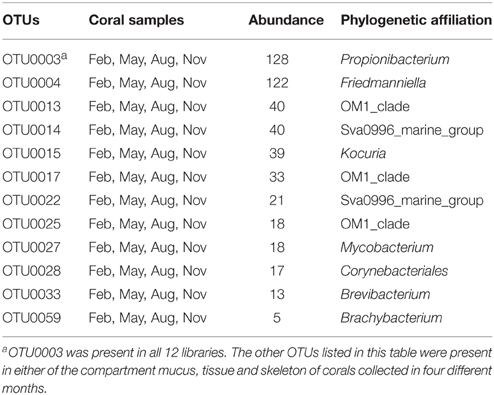
To investigate the distribution of OTUs in the three divergent coral compartments (mucus, tissue, and skeleton) and in the surrounding seawater, a venn diagram was constructed. The results showed that only 5 OTUs were present in all of P. lutea mucus, tissue and skeleton, and in sea water, which were identified as Sva0996_marine_group, Ilumatobacter, Corynebacterium, OM1_clade and Microbacterium (Table 2, Figure S1A). Another 17 OTUs, which were identified as Candidatus_Microthrix, Corynebacteriales, Friedmanniella, Micrococcus, Mycobacterium, OM1_clade, Propionibacterium, Sva0996_marine_group, Yonghaparkia and 480-2 were common in mucus, tissue, and skeleton (Table 2, Figure S1A). Twelve OTUs distributed in Propionibacterium, Friedmanniella, OM1_clade, Sva0996_marine_group, Kocuria, Mycobacterium, Corynebacteriales, Brevibacterium, and Brachybacterium were present in coral libraries in all four different months (Table 3, Figure S1B). The most abundant OTU0003, which was classified as Propionibacterium, was present in all coral samples with a high abundance (128 out of total 1687 reads in the coral libraries, 7.6%). The secondary abundance OTU0004 affiliated with Friedmanniella was present in all libraries except in skeleton collected in November.
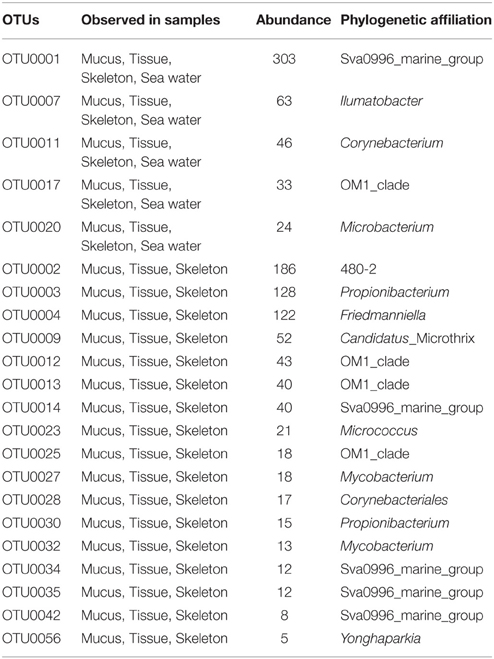
Table 2. OTUs presented in all of the coral and seawater libraries, or presented in all three divergent compartments of P. lutea.
Discussion
Highly Diverse Actinobacteria Associated with P. lutea
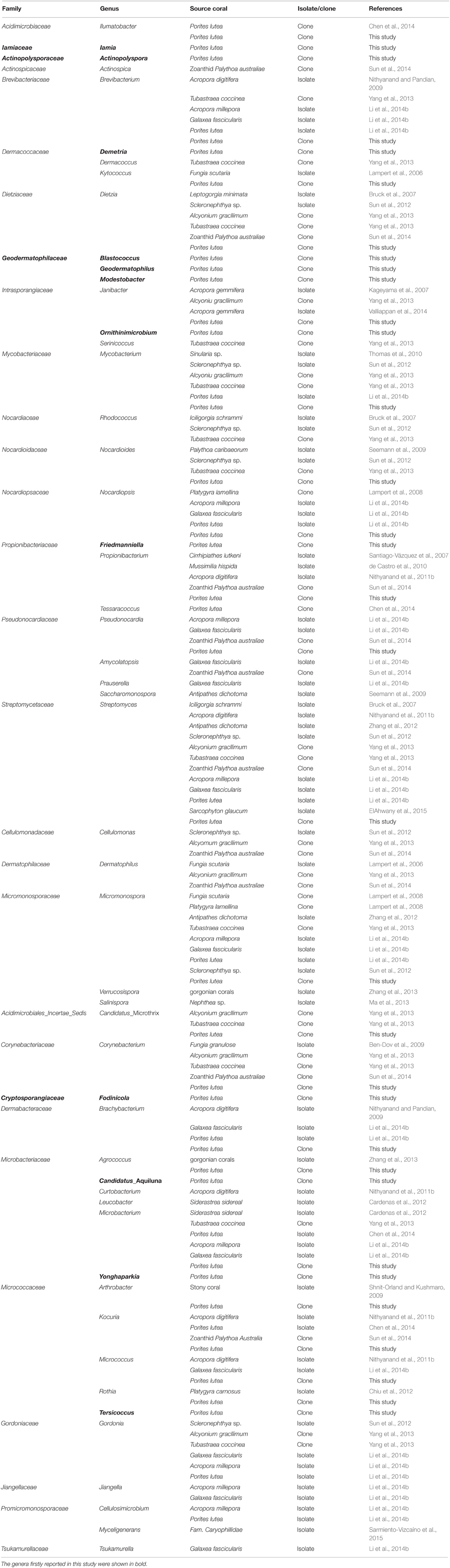
In comparison with previously reported results (Lampert et al., 2006, 2008; Bruck et al., 2007; Kageyama et al., 2007; Santiago-Vázquez et al., 2007; Ben-Dov et al., 2009; Nithyanand and Pandian, 2009; Seemann et al., 2009; Shnit-Orland and Kushmaro, 2009; de Castro et al., 2010; Thomas et al., 2010; Nithyanand et al., 2011a,b; Cardenas et al., 2012; Chiu et al., 2012; Sun et al., 2012, 2014; Zhang et al., 2012, 2013; Yang et al., 2013; Chen et al., 2014; Li et al., 2014a,b; EIAhwany et al., 2015; Sarmiento-Vizcaíno et al., 2015), 12 genera including Actinopolyspora, Blastococcus, Candidatus_Aquiluna, Demetria, Fodinicola, Friedmanniella, Geodermatophilus, Iamia, Modestobacter, Ornithinimicrobium, Tersicoccus, and Yonghaparkia were firstly detected in corals in this study (Table 4). Furthermore, many unclassified groups were detected in P. lutea, including even the group at the class taxon level. These results suggested that highly diverse and abundant known actinobacteria were associated with P. lutea as well as unknown groups. It was also noticed that many actinobacterial groups were only detected by the culture-independent method (Table 4), and some of them were ubiquitous and abundant, such as Friedmanniella, Ilumatobacter, and OM1_clade. Their physiological properties and ecological significance are worthy of deep research. For this purpose, the development and innovation of the isolation and cultivation methods in order to obtain pure cultures from the coral holobiont is particularly important.
According to our summary (Table 4), genera Agrococcus, Amycolatopsis, Arthrobacter, Brachybacterium, Brevibacterium, Candidatus_Microthrix, Corynebacterium, Cellulosimicrobium, Cellulomonas, Dermatophilus, Dietzia, Gordonia, Janibacter, Jiangella, Kocuria, Kytococcus, Microbacterium, Micromonospora, Micrococcus, Mycobacterium, Nocardioides, Nocardiopsis, Propionibacterium, Pseudonocardia, Rhodococcus, Rothia, and Streptomyces were detected in diverse coral species including scleractinian corals, such as Acropora digitifera (Nithyanand and Pandian, 2009; Nithyanand et al., 2011b), P. lutea (Li et al., 2014b; Sun et al., 2014) and Galaxea fascicularis (Li et al., 2014b), and gorgonian corals, Siderastrea sidereal (Cardenas et al., 2012) and Platygyra carnosus (Chiu et al., 2012). Most of them were present also in other marine organisms, such as sponges (Kim and Fuerst, 2006; Zhang et al., 2006; Selvin et al., 2009; Abdelmohsen et al., 2010, 2014; Schneemann et al., 2010; Sun et al., 2010; Webster and Taylor, 2012; Vicente et al., 2013), mollusks (Romanenko et al., 2008; Peraud et al., 2009), fishes (Sheeja et al., 2011), seaweeds (Lee, 2008; Singh and Reddy, 2013), seagrasses (Ravikumar et al., 2012), and sea cucumber (Kurahashi et al., 2009). Moreover, some of these widely distributed groups were considered as the bioactive compounds producers (Fiedler et al., 2005; Tabares et al., 2011; Margassery et al., 2012; Vicente et al., 2013; Manivasagan et al., 2014; Valliappan et al., 2014; EIAhwany et al., 2015), and probably take part in nitrogen (Su et al., 2013) and phosphorus (Sabarathnam et al., 2010) biogeochemical cycles. Whether they play these functional roles in corals in situ need to be further investigated.
Comparison of Actinobacterial Communities in the Corals and in the Ambient Seawater
Comparing the actinobacteria communities between in P. lutea and in the surrounding seawater will help us to understand the source of coral associated actinobacteria, and the interaction between the bacteria in sea water and in corals. Consisted with previous study on bacteria communities (Li et al., 2014a), the P. lutea associated actinobacteria communities were significantly different from which in the ambient seawater (Figure 3). Groups such as Propionibacteriaceae, Micromonosporaceae, and Micrococcaceae, were specifically associated with the corals rather than in the ambient seawater, where they originated from should be in doubt. Whether the wide distributed groups such as Sva0996_marine_group, OM1_clade, Microbacteriaceae and Ilumatobacter travel between the ambient seawater and the corals need to be investigated.
When researchers make a general observation of the whole bacterial communities, which were observed significantly different in coral mucus, tissue, and skeleton (Rohwer et al., 2002; Bourne and Munn, 2005; Sweet et al., 2011; Lee et al., 2012). However, it is unclear whether actinobacteria has a similar distribution pattern. In this study, both the hierarchical cluster analysis (Figure 3) and the RDA analysis (Figure 4) showed that the actinobacteria communities from the same compartment tended to cluster together. The distinct physiochemical microenvironments provided by corals probably is one of the causes (Le Tissier, 1990; Brown and Bythell, 2005; Sweet et al., 2011; Tremblay et al., 2011). Only a small fraction of OTUs (22 out of 299 OTUs in the coral libraries) was common in the coral mucus, tissue, and skeleton libraries in this study (Table 2). This result suggested that these members might have capabilities to adapt to different micro-environments in divergent compartments of P. lutea. A large amount of the OTUs was specifically associated with a certain coral compartment. Whether and how the properties of distinct actinobacteria assemblages in different coral compartments actually contribute to the close relationship constructed between these associates and corals should be addressed from a functional perspective.
Relationship of environmental factors and the P. lutea-associated Actinobacteria
It is different from previous conclusion of the distribution of coral-associated bacteria (Chen et al., 2011; Li et al., 2014a), actinobacteria associated with P. lutea did not show the apparent seasonal dynamic variations. We suggest that the actinobacteria compositions are relatively stable in distinct compartments in P. lutea. In addition, none of the environmental factors analyzed in this study was determined as the most significant influence on the actinobacteria communities. Even so, some genera were found closely correlated with specific environmental factors. For instance, Propionibacterium showed negatively correlation with dissolved oxygen, probably due to its capability of living in the anaerobic conditions (Patrick and McDowell, 2012). Moreover, the OTUs0003 and 0004 affiliated with Propionibacteriaceae was present in almost all 12 clone libraries with a very high abundance. Whether they are true symbionts, and what functions they play are worth further research.
Conclusion
The diversity and distribution of coral-associated actinobacteria were first comprehensively investigated in this study. Highly diverse actinobacteria was revealed in the 16S rRNA gene clone libraries of scleractinian coral P. lutea in the South China Sea. Twelve Actinobacteria genera were detected in corals for the first time as well as a large number of unclassified groups. The actinobacterial community compositions were distinct in P. lutea and in the surrounding seawater. Furthermore, the higher similarity of actinobacteria composition was observed in the same compartment (i.e., mucus, tissue, or skeleton) of P. lutea. This study will help attracting the attentions on the ecological role of actinobacteria in corals besides the natural products bioprospecting.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We would like to thank the Tropical Marine Biological Research Station in Hainan for help with sample collection. This research was supported by the Key Research Program of the Chinese Academy of Sciences (No. KSCX2-EW-B-13), National Natural Science Foundation of China (No. 41106139, 41230962) and Pearl River Nova Program of Guangzhou (No. 2014J2200075), Administration of Ocean and Fisheries of Guangdong Province (No. GD2012-D01-002), and the Knowledge Innovation Program of the Chinese Academy of Sciences (No. SQ201301).
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2015.01094
References
Abdelmohsen, U. R., Bayer, K., and Hentschel, U. (2014). Diversity, abundance and natural products of marine sponge-associated actinomycetes. Nat. Prod. Rep. 31, 381–399. doi: 10.1039/c3np70111e
Abdelmohsen, U. R., Pimentel-Elardo, S. M., Hanora, A., Radwan, M., Abou-El-Ela, S. H., Ahmed, S., et al. (2010). Isolation, phylogenetic analysis and anti-infective activity screening of marine sponge associated actinomycetes. Mar. Drugs 8, 399–412. doi: 10.3390/md8030399
Ben-Dov, E., Ben Yosef, D. Z., Pavlov, V., and Kushmaro, A. (2009). Corynebacteriummaris sp. nov., a marine bacterium isolated from the mucus of the coral Fungia granulose. Int. J. Syst. Evol. Microbiol. 59, 2458–2463. doi: 10.1099/ijs.0.007468-0
Bourne, D. G., and Munn, C. B. (2005). Diversity of bacteria associated with the coral Pocillopora damicornis from the Great Barrier Reef. Environ. Microbiol. 7, 1162–1174. doi: 10.1111/j.1462-2920.2005.00793.x
Bourne, D. G., and Webster, N. S. (2013). “Coral reef bacterial communities,” in The Prokaryotes, eds E. Rosenberg, E. F. DeLong, S. Lory, E. Stackebrandt, and F. Thompson (Heidelberg: Springer-Verlag), 163–187.
Brown, B. E., and Bythell, J. C. (2005). Perspectives on mucus secretion in reef corals. Mar. Ecol. Prog. Ser. 296, 291–309. doi: 10.3354/meps296291
Bruck, T. B., Bruck, W. M., Santiago-Vázquez, L. Z., McCarthy, P. J., and Kerr, R. G. (2007). Diversity of the bacterial communities associated with the azooxanthellate deep water octocorals Leptogorgia minimata, Iciligorgia schrammi, and Swiftia exertia. Mar. Biotechnol. 9, 561–576. doi: 10.1007/s10126-007-9009-1
Cardenas, A., Rodriguez-R, L. M., Pizarro, V., Cádavid, L. F., and Arevalo- Ferro, C. (2012). Shifts in bacterial communities of two Caribbean reef- building coral species affected by white plague disease. ISME J. 6, 502–512. doi: 10.1038/ismej.2011.123
Carlos, C., Torres, T. T., and Ottoboni, L. M. (2013). Bacterial communities and species-specific associations with the mucus of Brazilian coral species. Sci. Rep. 3:1624. doi: 10.1038/srep01624
Chen, C. P., Tseng, C. H., Chen, C. A., and Tang, S. L. (2011). The dynamics of microbial partnerships in the coral Isopora palifera. ISME J. 5, 728–740. doi: 10.1038/ismej.2010.151
Chen, Q., Long, L. J., Zhang, S., Dong, J. D., and Li, J. (2014). Diversity of actinobacteria associated with coral Porites lutea and Galaxea fascicularis. Microbiol. China 41, 691–698. doi: 10.13344/j.microbiol.china.130230
Chiu, J. M. Y., Li, S., Li, A., Po, B., Zhang, R., and Shin, P. K. S. (2012). Bacteria associated with skeletal tissue growth anomalies in the coral Platygyra carnosus. FEMS Microbiol. Ecol. 79, 380–391. doi: 10.1111/j.1574-6941.2011.01225.x
Clarke, K. R. (1993). Non-parametric multivariate analyses of changes in community structure. Aust. J. Ecol. 18, 117–143. doi: 10.1111/j.1442-9993.1993.tb00438.x
de Castro, A. P., Araújo, S. D. Jr., Reis, A. M., Moura, R. L., Francini-Filho, R. B., Pappas, G. Jr., et al. (2010). Bacterial community associated with healthy and diseased reef coral Mussismilia hispida from eastern Brazil. Microb. Ecol. 59, 658–667. doi: 10.1007/s00248-010-9646-1
EIAhwany, A. M. D., Ghozlan, H. A., ELSharif, H. A., and Sabry, S. A. (2015). Phylogenetic diversity and antimicrobial activity of marine bacteria associated with the soft coral Sarcophyton glaucum. J. Basic. Microb. 55, 2–10. doi: 10.1002/jobm.201300195
Fiedler, H. P., Bruntner, C., Bull, A. T., Ward, A. C., Goodfellow, M., Potterat, O., et al. (2005). Marine actinomycetes as a source of novel secondary metabolites. Anton. Leeuw. Int. J. G. 87, 37–42. doi: 10.1007/s10482-004-6538-8
Hong, M. J., Yu, Y. T., Chen, C. A., Chiang, P. W., and Tang, S. L. (2009). Influence of species specificity and other factors on bacteria associated with the coral Stylophora pistillata in Taiwan. Appl. Environ. Microbiol. 75, 7797–7806. doi: 10.1128/AEM.01418-09
Kageyama, A., Takahashi, Y., Yasumoto-Hirose, M., Kasai, H., Shizuri, Y., and Omura, S. (2007). Janibacter corallicola sp. nov., isolated from coral in Palau. J. Gen. Appl. Microbiol. 53, 185–189. doi: 10.2323/jgam.53.185
Kim, T. K., and Fuerst, J. A. (2006). Diversity of polyketide synthase genes from bacteria associated with the marine sponge Pseudoceratina clavata: culture-dependent and culture-independent approaches. Environ. Microbiol. 8, 1460–1470. doi: 10.1111/j.1462-2920.2006.01040.x
Krediet, C. J., Ritchie, K. B., Alagely, A., and Teplitski, M. (2013). Members of native coral microbiota inhibit glycosidases and thwart colonization of coral mucus by an opportunistic pathogen. ISME J. 7, 980–990. doi: 10.1038/ismej.2012.164
Kurahashi, M., Fukunaga, Y., Sakiyama, Y., Harayama, S., and Yokota, A. (2009). Iamia majanohamensis gen. nov., sp. nov., an actinobacterium isolated from sea cucumber Holothuria edulis, and proposal of Iamiaceae fam. nov. Int. J. Syst. Evol. Microbiol. 59, 869–873. doi: 10.1099/ijs.0.005611-0
Lampert, Y., Kelman, D., Dubinsky, Z., Nitzan, Y., and Hill, R. T. (2006). Diversity of culturable bacteria in the mucus of the Red Sea coral Fungia scutaria. FEMS Microbiol. Ecol. 58, 99–108. doi: 10.1111/j.1574-6941.2006.00136.x
Lampert, Y., Kelman, D., Nitzan, Y., Dubinsky, Z., Behar, A., and Hill, R. T. (2008). Phylogenetic diversity of bacteria associated with the mucus of Red Sea corals. FEMS Microb. Ecol. 64, 187–198. doi: 10.111/j.1574-6941.2008.00458.x
Lee, O. O., Yang, J., Bougouffa, S., Wang, Y., Batang, Z., Tian, R. M., et al. (2012). Spatial and species variations in bacterial communities associated with corals from the Red Sea as revealed by pyrosequencing. Appl. Environ. Microbiol. 78, 7173–7184. doi: 10.1128/AEM.01111-12
Lee, S. D. (2008). Agrococcus jejuensis sp. nov., isolated from dried seaweed. Int. J. Syst. Evol. Microbiol. 58, 2297–2300. doi: 10.1099/ijs.0.65731-0
Le Tissier, M. D. A. (1990). The ultrastructure of the skeleton and skeletogenic tissues of the temperate coral Caryophyllia smithii. J. Mar. Biol. Assoc. UK. 70, 295–310. doi: 10.1017/S0025315400035414
Li, J., Chen, Q., Long, L. J., Dong, J. D., Yang, J., and Zhang, S. (2014a). Bacterial dynamics within the mucus, tissue and skeleton of the coral Porites lutea during different seasons. Sci. Rep. 4:07320. doi: 10.1038/srep07320
Li, J., Chen, Q., Zhang, S., Huang, H., Yang, J., Tian, X. P., et al. (2013). Highly heterogeneous bacterial communities associated with the South China Sea reef corals Porites lutea, Galaxea fascicularis and Acropora millepora. PLoS ONE 8:e71301. doi: 10.1038/nrmicro1635
Li, J., Dong, J. D., Yang, J., Luo, X. M., and Zhang, S. (2014b). Detection of polyketide synthase and nonribosomal peptide synthetase biosynthetic genes from antimicrobial coral associated actinomycetes. Anton. Leeuw. Int. J. 106, 623–635. doi: 10.1007/s10482-014-0233-1
Liu, Z. F., Huang, W., Li, J. R., Wang, P. X., Wang, R. J., Yu, K. F., et al. (2009). “Sedimentology,” in The South China Sea: Paleoceanography and Sedimentology, eds P. X. Wang and Q. Y. Li (Heidelberg; London; New York, NY: Springer Science+Bussiness Media B. V: Dordrecht), 229–236.
Ma, L., Zhang, W. J., Zhu, Y. G., Wu, Z. C., Saurav, K., Hang, H., et al. (2013). Isolation of Actinobacteria with antibiotic activity associated with soft coral Nephthea sp. Acta. Microbiol. Sinica. 53, 1063–1071.
Manivasagan, P., Venkatesan, J., Sivakumar, K., and Kim, S. K. (2014). Pharmaceutically active secondary metabolites of marine actinobacteria. Microbiol. Res. 169, 262–278. doi: 10.1016/j.micres.2013.07.014
Margassery, L. M., Kennedy, J., O'Gara, F., Dobson, A. D., and Morrissey, J. P. (2012). Diversity and antibacterial activity of bacteria isolated from the coastal marine sponges Amphilectus fucorum and Eurypon major. Lett. Appl. Microbiol. 55, 2–8. doi: 10.1111/j.1472-765X.2012.03256.x
Moberg, F., and Folke, C. (1999). Ecological goods and services of coral reef ecosystems. Ecol. Econ. 29, 215–233. doi: 10.1016/S0921-8009(99)00009-9
Nithyanand, P., Indhumathi, T., Ravi, A. V., and Pandian, S. K. (2011a). Culture independent characterization of bacteria associated with the mucus of the coral Acropora digitifera from the Gulf of Mannar. World J. Microb. Biot. 27, 1399–1406. doi: 10.1007/s11274-010-0591-4
Nithyanand, P., Manju, S., and Pandian, S. K. (2011b). Phylogenetic characterization of culturable actinomycetes associated with the mucus of the coral Acropora digitifera from Gulf of Mannar. FEMS Microbiol. Lett. 314, 112–118. doi: 10.1111/j.1574-6941.2009.00723.x
Nithyanand, P., and Pandian, S. K. (2009). Phylogenetic characterization of culturable bacterial diversity associated with the mucus and tissue of the coral Acropora digitifera from the Gulf of Mannar. FEMS Microbiol. Ecol. 69, 384–394. doi: 10.1111/j.1574-6941.2009.00723.x
Oliveros, J. C. (2007–2015). Venny. An interactive Tool for Comparing Lists with Venn's Diagrams. Available online at: http://bioinfogp.cnb.csic.es/tools/venny/index.html.
Patrick, S., and McDowell, A. (2012). “Propionibacterium Orla-Jensen 1909, 337AL emend. Charfreitag, Collins and Stackebrandt 1988, 356,” in Bergey's Manual of Systematic Bacteriology, The Actinobacteria, Part B, Vol. 5., eds M. Goodfellow, P. Kämpfer, H. Busse, M. E. Trujillo, K. Suzuki, W. Ludwig, and W. B. Whitman (New York, NY: Springer), 1138–1155.
Peraud, O., Biggs, J. S., Hughen, R. W., Light, A. R., Concepcion, G. P., and Olivera, B. (2009). Microhabitats within venomous cone snails yield diverse actinobacteria. Appl. Environ. Microbiol. 75, 6820–6826. doi: 10.1128/AEM.01238-09
Raina, J. B., Tapiolas, D., Willis, B. L., and Bourne, D. G. (2009). Coral-associated bacteria and their role in the biogeochemical cycling of sulfur. Appl. Environ. Microbiol. 75, 3492–3501. doi: 10.1128/AEM.02567-08
Ravikumar, S., Gnanadesigan, M., Saravanan, A., Monisha, N., Brindha, V., and Muthumari, S. (2012). Antagonistic properties of seagrass associated Streptomyces sp. RAUACT-1: a source for anthraquinone rich compound. Asian Pac. J. Trop. Med. 5, 887–890. doi: 10.1016/S1995-7645(12)60165-5
Ritchie, K. B. (2006). Regulation of microbial populations by coral surface mucus and mucus-associated bacteria. Mar. Ecol. Prog. Ser. 322, 1–14. doi: 10.3354/meps322001
Rohwer, F., Seguritan, V., Azam, F., and Knowlton, N. (2002). Diversity and distribution of coral-associated bacteria. Mar. Ecol. Prog. Ser. 243, 1–10. doi: 10.3354/meps243001
Romanenko, L. A., Uchino, M., Kalinovskaya, N. I., and Mikhailov, V. V. (2008). Isolation, phylogenetic analysis and screening of marine mollusc-associated bacteria for antimicrobial, hemolytic and surface activities. Microbiol. Res. 163, 633–644. doi: 10.1016/j.micres.2006.10.001
Rosenberg, E., Koren, O., Reshef, L., Efrony, R., and Ziber-Rosenberg, I. (2007). The role of microorganisms in coral health, disease and evolution. Nat. Rev. Microb. 5, 355–362. doi: 10.1038/nrmicro1635
Sabarathnam, B., Manilal, A., Sujith, S., Kiran, G. S., Selvin, J., Thomas, A., et al. (2010). Role of sponge associated actinomycetes in the marine phosphorus biogeochemical cycles. Am. Eur. J. Agric. Environ. Sci. 8, 253–256.
Santiago-Vázquez, L. Z., Brück, T. B., Brück, W. M., Duque-Alarcón, A. P., McCarthy, P. J., and Kerr, R. G. (2007). The diversity of the bacterial communities associated with the azooxanthellate hexacoral Cirrhipathes lutkeni. ISME J. 1, 654–659. doi: 10.1038/ismej.2007.77
Sarmiento-Vizcaíno, A., González, V., Braña, A. F., Molina, A., Acuña, J. L., García, L. A., et al. (2015). Myceligenerans cantabricum sp. nov., a barotolerant actinobacterium isolated from a deep cold-water coral. Int. J. Syst. Evol. Microbiol. 8, 1328–1334. doi: 10.1099/ijs.0.000107
Schloss, P. D., Westcott, S. L., Ryabin, T., Hall, J. R., Hartmann, M., Hollister, E. B., et al. (2009). Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541. doi: 10.1128/AEM.01541-0
Schneemann, I., Nagel, K., Kajahn, I., Labes, A., Wiese, J., and Imhoff, J. F. (2010). Comprehensive investigation of marine Actinobacteria associated with the sponge Halichondria panacea. Appl. Environ. Microbiol. 76, 3702–3714. doi: 10.1128/AEM.00780-10
Seemann, P., Gernert, C., Schmitt, S., Mebs, D., and Hentschel, U. (2009). Detection of hemolytic bacteria from Palythoa caribaeorum (Cnidaria, Zoantharia) using a novel palytoxin-screening assay. Anton. Leeuw. Int. J. G. 96, 405–411. doi: 10.1007/s10482-009-9353-4
Selvin, J., Gandhimathi, R., Kiran, G. S., Priya, S. S., Ravji, T. R., and Hema, T. A. (2009). Culturable heterotrophic bacteria from the marine sponge Dendrilla nigra: isolation and phylogenetic diversity of actinobacteria. Helgol. Mar. Res. 63, 239–247. doi: 10.1007/s10152-009-0153-z
Sheeja, M. S., Selvakumar, D., and Dhevendaran, K. (2011). Antagonistic potential of Streptomyces associated with the gut of marine ornamental fishes. Middle-East J. Sci. Res. 7, 327–334. doi: 10.1128/AEM.67.1.434-444.2001
Shnit-Orland, M., and Kushmaro, A. (2009). Coral mucus-associated bacteria: a possible first line of defense. FEMS Microbiol. Ecol. 67, 371–380. doi: 10.1111/j.1574-6941.2008.00644.x
Singh, R. P., and Reddy, C. R. K. (2013). Seaweed–microbial interactions: key functions of seaweed-associated bacteria. FEMS Microbiol. Ecol. 88, 2, 213–230. doi: 10.1111/1574-6941.12297
Stach, J. E. M., Maldonado, L. A., Ward, A. C., Goodfellow, M., and Bull, A. T. (2003). New primers for the class Actinobacteria: application to marine and terrestrial environments. Environ. Microbiol. 5, 828–841. doi: 10.1046/j.1462-2920.2003.00483.x
Su, J., Jin, L., Jiang, Q., Sun, W., Zhang, F., and Li, Z. (2013). Phylogenetically diverse ureC genes and their expression suggest the urea utilization by bacterial symbionts in marine sponge Xestospongia testudinaria. PLoS ONE 8:e64848. doi: 10.1371/journal.pone.0064848
Sun, W., Dai, S., Jiang, S., Wang, G., Liu, G., Wu, H., et al. (2010). Culture-dependent and culture-independent diversity of Actinobacteria associated with the marine sponge Hymeniacidon perleve from the South China Sea. Anton. Leeuw. Int. J. G. 98, 65–75. doi: 10.1007/s10482-010-9430-8
Sun, W., Peng, C., Zhao, Y., and Li, Z. (2012). Functional gene-guided discovery of type II polyketides from culturable actinomycetes associated with soft coral Scleronephthya sp. PLoS ONE 7:e42847. doi: 10.1371/journal.pone.0042847
Sun, W., Zhang, F., He, L., and Li, Z. (2014). Pyrosequencing reveals diverse microbial community associated with the Zoanthid Palythoa australiae from the South China Sea. Microb. Ecol. 67, 942–950. doi: 10.1007/s00248-014-0395-4
Sweet, M. J., Croquer, A., and Bythell, J. C. (2011). Bacterial assemblages differ between compartments within the coral holobiont. Coral Reefs 30, 39–52. doi: 10.1007/s00338-010-0695-1
Tabares, P., Pimentel-Elardo, S. M., Schirmeister, T., Hünig, T., and Hentschel, U. (2011). Anti-protease and immunomodulatory activities of bacteria associated with Caribbean sponges. Mar. Biotechnol. 13, 883–892. doi: 10.1007/s10126-010-9349-0
ter Braak, C. J. F., and Šmilauer, P. (2002). CANOCO Reference Manual and CanoDraw for Windows User's Guide: Software for Canonical Community Ordination (version 4.5). Ithaca, NY: Microcomputer Power.
Thomas, S., Burdett, H., Temperton, B., Wick, R., Snelling, D., McGrath, J., et al. (2010). Evidence for phosphonate usage in the coral holobiont. ISME J. 4, 459–461. doi: 10.1038/ismej.2009.129
Tremblay, P., Weinbauer, M. K., Rottier, C., Guérardel, Y., Nozais, C., and Ferrier-pagès, C. (2011). Mucus composition and bacterial communities associated with the tissue and skeleton of three scleractinian corals maintained under culture conditions. J. Mar. Biol. Assoc. UK. 91, 649–654. doi: 10.1017/S002531541000130X
Valliappan, K., Sun, W., and Li, Z. Y. (2014). Marine actinobacteria associated with marine organisms and their potentials in producing pharmaceutical natural products. Appl. Microbiol. Biot. 98, 7365–7377. doi: 10.1007/s00253-014-5954-6
Vicente, J., Stewart, A., Song, B., Hill, R. T., and Wright, J. L. (2013). Biodiversity of Actinomycetes associated with Caribbean sponges and their potential for natural product discovery. Mar. Biotechnol. 15, 413–424. doi: 10.1007/s10126-013-9493-4
Wang, L. R., Yu, K. F., Zhao, H. T., and Zhang, Q. M. (2014). Economic valuation of the coral reefs in South China Sea. Trop. Geogr. 34, 44–49. doi: 10.13284/j.cnki.rddl.000007
Webster, N. S., and Taylor, M. W. (2012). Marine sponges and their microbial symbionts: love and other relationships. Environ. Microbiol. 14, 335–346. doi: 10.1111/j.1462-2920.2011.02460.x
Yang, S., Sun, W., Tang, C., Jin, L., Zhang, F., and Li, Z. (2013). Phylogenetic diversity of Actinobacteria associated with Soft Coral Alcyonium gracllimum and Stony Coral Tubastraea coccinea in the East China Sea. Microb. Ecol. 66, 189–199. doi: 10.1007/s00248-013-0205-4
Zhang, H., Lee, Y. K., Zhang, W., and Lee, H. K. (2006). Culturable actinobacteria from the marine sponge Hymeniacidon perleve: isolation and phylogenetic diversity by 16S rRNA gene-RFLP analysis. Anton. Leeuw. Int. J. G. 90, 159–169. doi: 10.1007/s10482-006-9070-1
Zhang, X. Y., He, F., Wang, G. H., Bao, J., Xu, X. Y., and Qi, S. H. (2013). Diversity and antibacterial activity of culturable actinobacteria isolated from five species of the South China Sea gorgonian corals. World J. Microb. Biot. 29, 1107–1116. doi: 10.1007/s11274-013-1279-3
Zhang, X., Sun, Y., Bao, J., He, F., Xu, X., and Qi, S. (2012). Phylogenetic survey and antimicrobial activity of culturable microorganisms associated with the South China Sea black coral Antipathes dichotoma. FEMS Microbiol. Lett. 336, 122–130. doi: 10.1111/j.1574-6968.2012.02662.x
Keywords: actinobacteria, Porites lutea, diversity, temporal and spatial distribution, 16S rRNA gene
Citation: Kuang WQ, Li J, Zhang S and Long LJ (2015) Diversity and distribution of Actinobacteria associated with reef coral Porites lutea. Front. Microbiol. 6:1094. doi: 10.3389/fmicb.2015.01094
Received: 10 July 2015; Accepted: 22 September 2015;
Published: 21 October 2015.
Edited by:
Sheng Qin, Jiangsu Normal University, ChinaReviewed by:
Syed Gulam Dastager, National Collection of Industrial Microorganisms Resource Center, IndiaWei Sun, Shanghai Jiao Tong University, China
P. Nithyanand, SASTRA University, India
Copyright © 2015 Kuang, Li, Zhang and Long. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Lijuan Long, CAS Key Laboratory of Tropical Marine Bio-Resources and Ecology, RNAM Center for Marine Microbiology, South China Sea Institute of Oceanology, Chinese Academy of Sciences, Xingangxi Road 164, Guangzhou 510301, China,bG9uZ2xqQHNjc2lvLmFjLmNu
†These authors have contributed equally to this work.
 Weiqi Kuang
Weiqi Kuang Jie Li
Jie Li Si Zhang1
Si Zhang1 Lijuan Long
Lijuan Long