- 1National Risk Assessment Laboratory for Antimicrobial Resistance of Animal Original Bacteria, South China Agricultural University, Guangzhou, China
- 2College of Veterinary Medicine, South China Agricultural University, Guangzhou, China
- 3College of Animal Science and Technology, Guangxi University, Nanning, China
- 4Jiangsu Co-Innovation Centre for Prevention and Control of Important Animal Infectious Diseases and Zoonoses, Yangzhou, China
During a regular monitoring of antimicrobial resistance in a farrowing farm in Southern China, 117 Escherichia coli isolates were obtained from sows and piglets. Compared with the isolates from piglets, the isolates from sows exhibited higher resistance rates to the tested cephalosporins. Correspondingly, the total detection rate of the blaCMY-2/blaCTX-M genes in the sow isolates (34.2%) was also significantly higher than that of the piglet isolates (13.6%; p < 0.05). The blaCMY-2 gene had a relatively high prevalence (11.1%) in the E. coli isolates. MLST and PFGE analysis revealed the clonal spread of ST1121 E. coli in most (7/13) of the blaCMY-2-positive isolates. An indistinguishable IncHI2 plasmid harboring blaCMY-2 was also identified in each of the seven ST1121 E. coli isolates. Complete sequence analysis of this IncHI2 plasmid (pEC5207) revealed that pEC5207 may have originated through recombination of an IncHI2 plasmid with a blaCMY-2-carrying IncA/C plasmid like pCFSAN007427_01. In addition to blaCMY-2, pEC5207 also carried other resistance determinants for aminoglycosides (aacA7), sulfonamides (sul1), as well as heavy metals ions, such as Cu and Ag. The susceptibility testing showed that the pEC5207 can mediate both antibiotic and heavy metal resistance. This highlights the role of pEC5207 in co-selection of blaCMY-2-positive isolates under the selective pressure of heavy metals, cephalosporins, and other antimicrobials. In conclusion, clonal spread of an ST1121 type E. coli strain harboring an IncHI2 plasmid contributed to the dissemination of blaCMY-2 in a farrowing farm in Southern China. We also have determined the first complete sequence analysis of a blaCMY-2-carrying IncHI2 plasmid.
Introduction
Antimicrobial agents are often used as feed and water additives in food animals to treat or prevent disease and to promote general overall health (McEwen and Fedorka-Cray, 2002; Cabello et al., 2013). However, antimicrobial resistance driven by the intensive use of antimicrobial agents in animal husbandry is increasing worldwide (Witte, 1998). In the swine industry, antimicrobial resistance patterns can be traced to particular farms associated with certain management practices (Rasschaert et al., 2012; Brooks et al., 2014). In farrowing farms, antimicrobial resistance in piglets has been shown to be a reflection of antimicrobial use in sows (Mathew et al., 2005; Callens et al., 2015), because sows are an important reservoir of antimicrobial resistant bacteria for their offspring (Callens et al., 2012; Crombe et al., 2013). Additionally, piglet transfer from farrowing to finishing farms increases the likelihood of the transmission of resistant bacteria, thus increasing the risk of antimicrobial resistance transfer between swine farms (Sandvang et al., 2000; van Duijkeren et al., 2008). Therefore, surveillance for antimicrobial resistance in the farrowing farm is important for controlling the dissemination of antimicrobial resistance.
Escherichia coli is an important cause of intestinal and extraintestinal diseases in animals and humans worldwide, and β-lactams are widely used in veterinary medicine to treat colibacillosis. However, the use of extended-spectrum cephalosporins (ESCs) in animals has contributed to β-lactam resistance in E. coli (Greko et al., 2009). Resistance to ESCs in E. coli has been associated with the extended-spectrum β-lactamases (ESBLs) and plasmid-mediated Ambler class C cephamycinases (pAmpC β-lactamases; Seiffert et al., 2013). ESBLs are the major contributors to ESC resistance in E. coli and confer resistance to cephalosporins with an oxyimino side chain (cefotaxime, ceftriaxone, and ceftazidime; Bradford, 2001).
Unlike ESBLs, pAmpC β-lactamases exhibit activity against cephamycins, such as cefoxitin and cannot be inactivated by β-lactamase inhibitor clavulanate (Bradford, 2001; Jacoby, 2009). Among them, CMY-2, the most common pAmpC β-lactamase, has been documented worldwide in bacteria of human and animal origin (Li et al., 2007; Jacoby, 2009). The blaCMY-2 gene which likely originated from the chromosomal AmpC locus of Citrobacter freundii has been horizontally transmitted through plasmids in E. coli from different sources (Martin et al., 2012). Various plasmid types are associated with blaCMY-2, including IncA/C, IncF, IncI1, IncL/M, IncP, IncK, and IncHI2. Of these, IncA/C and IncI1 plasmid are the most common carriers of blaCMY-2 (Verdet et al., 2009; Folster et al., 2011; Martin et al., 2012; Bortolaia et al., 2014; Guo et al., 2014).
The blaCMY-2 gene is prevalent in ESC-resistant E. coli of livestock (Li et al., 2007; Seiffert et al., 2013). In swine, the detection rate of blaCMY-2 in E. coli is quite different, ranging from 0 to up to 80% (Seiffert et al., 2013). Although the detection methods of CMY-2 vary between reports, this may still reflect the different prevalence of blaCMY-2 worldwide. In mainland China, the blaCMY-2 gene was detected for the first time in E. coli of swine origin in Tian et al. (2012), Zheng et al. (2012), and subsequently occured in swine E. coli isolates carrying plasmid-mediated quinolone resistance (PMQR) genes (Liu et al., 2013a). Recently, a surveillance study identified blaCMY-2 mainly in E. coli of pig origin, highlighting the role of CMY-2 in the ESC resistance of swine E. coli (Guo et al., 2014). In the present study, we make an investigation on the prevalence of drug resistance and ESBL/pAmpC genes in E. coli isolates from a swine farm in Southern China. The blaCMY-2-positive isolates were further analyzed to characterize the transmission mechanisms of the blaCMY-2 gene.
Materials and Methods
Bacterial Isolates and Antimicrobial Susceptibility Testing
In August 2011, a regular monitoring of antimicrobial resistance was conducted in a farrowing farm in Southern China. This farm had been in operation about 8 years and consisted of 2,300 sows with production of about 40,000 piglets for market annually. Rectal swab samples were randomly taken from one pig in every batch pen. The rectal swabs were taken by inserting the sterile swab about 2 cm into the rectum, rotated gently and then immersed in sterile PBS. After collection, the swabs were immediately brought to the laboratory in cool conditions. A total of 137 swab samples were collected from sows (1–5 years-old) and piglets (1–60 days-old). The collected samples were plated on MacConkey agar and then incubated at 37°C for 24 h. One suspected colony with typical E. coli morphology was selected from each sample and was identified with API 20E system (BioMerieux, France). Minimal inhibitory concentrations (MICs) of ampicillin (AMP), ceftazidime (CAZ), cefoxitin (FOX), cefotaxime (CTX), ceftiofur (CEF), amikacin (AMK), kanamycin (KAN), florfenicol (FFC), doxycycline (DOX), enrofloxacin (ENR), and trimethoprim-sulfamethoxazole (SXT) were determined by the agar dilution method in accordance with the standard provided by the Clinical and Laboratory Standards Institute (2013a,b). E. coli ATCC 25922 was used as the quality control strain.
Detection of ESBL/pAmpC Genes
Extended-spectrum β-lactamase genes (blaTEM, blaSHV, blaCTX-M-1G, blaCTX-M-9G, blaCTX-M-2G, and blaCTX-M-25G) among the E. coli isolates were analyzed by PCR amplification using previously published primers and protocols (Liu et al., 2013a). Purified PCR products were sequenced and compared using the β-lactamase classification system1 to confirm the subtypes. Detection of pAmpC genes was performed by a multiplex PCR as previously described (Perez-Perez and Hanson, 2002). For amplification of the entire blaCMY-2 gene, PCR-positive isolates were re-amplified and sequenced with specific primers (Perez-Perez and Hanson, 2002).
Molecular Typing
All blaCMY-2-positive isolates were classified according to XbaI-pulsed-field gel electrophoresis (PFGE) type (Tenover et al., 1995). Comparison of PFGE patterns was performed by BioNumerics®v6.6 (Applied Maths, Ghent, Belgium) with a cut-off at 90% of the similarity values to indicate identical PFGE types. Multilocus sequence typing (MLST) was performed by using the primers and protocol specified at the E. coli MLST web site2.
Transferability of blaCMY-2
Conjugation experiments were performed as previously described (Chen et al., 2007a), using E. coli C600 (streptomycin-resistant; MIC >2000 μg/ml) as a recipient. Putative transconjugants were selected on MacConkey agar plates with streptomycin (2000 μg/ml) and cefoxitin (32 μg/ml), examined for the presence of blaCMY-2 by PCR assay, and finally confirmed by ERIC-PCR (Versalovic et al., 1991). All transconjugants were tested for antimicrobial susceptibility as described above.
The susceptibility of transconjugant EC5207-35T to heavy metals (Cu, Ag) was tested by microdilution in an aerobic atmosphere, as previously described, with some modifications (Mourao et al., 2015). Briefly, the transconjugant was incubated in Mueller–Hinton broth with serial dilutions of CuSO4 (0.25, 0.5, 1, 2, 4, 8, 12, 16, 20, 24, 32, and 36 mM, adjusted to pH 7.2) and AgNO3 (0.0125, 0.025, 0.06, 0.08, 0.125, 0.16, 0.25, 0.32, 0.5, 1.0, 1.5, and 3 mM, adjusted to pH 7.4). E. coli C600 was used as the reference strain.
Plasmid Analysis
Plasmids of the transconjugants were typed with PCR-based replicon typing (PBRT; Carattoli et al., 2005). The size of blaCMY-2-carrying plasmid in the transconjugants was determined using S1 nuclease-digested (TaKaRa Biotechnology, Dalian, China) genomic DNA followed by PFGE and Southern blot hybridization with a blaCMY-2-specific probe (Barton et al., 1995). Plasmid DNA from the transconjugants was extracted using the QIAGEN Plasmid Midi Kit, and was further analyzed by restriction fragment length polymorphism (RFLP) using XbaI (TaKaRa Biotechnology, Dalian, China).
Complete Sequence of blaCMY-2-carrying IncHI2 Plasmid pEC5207
In order to further characterize the blaCMY-2-carrying IncHI2 plasmids of the ST1121 clone in this study, the plasmid pEC5207 from transconjugant EC5207-35T was sequenced using SMRT sequencing approach and assembled by HGAP2.2.0 method (Chin et al., 2013). Open reading frames (ORFs) prediction and annotation were performed with the RAST tools (Aziz et al., 2008). Sequence comparison and map generation were performed using BLAST3 and Easyfig version 2.1 (Sullivan et al., 2011).
Statistical Methods
Statistical significance for the comparison of prevalence data was determined by the χ2 test. Differences were considered statistically significant at p < 0.05.
Nucleotide Sequence Accession Number
The complete DNA sequence of plasmid pEC5207 was assigned GenBank accession number KT347600.
Results
Antimicrobial Susceptibility
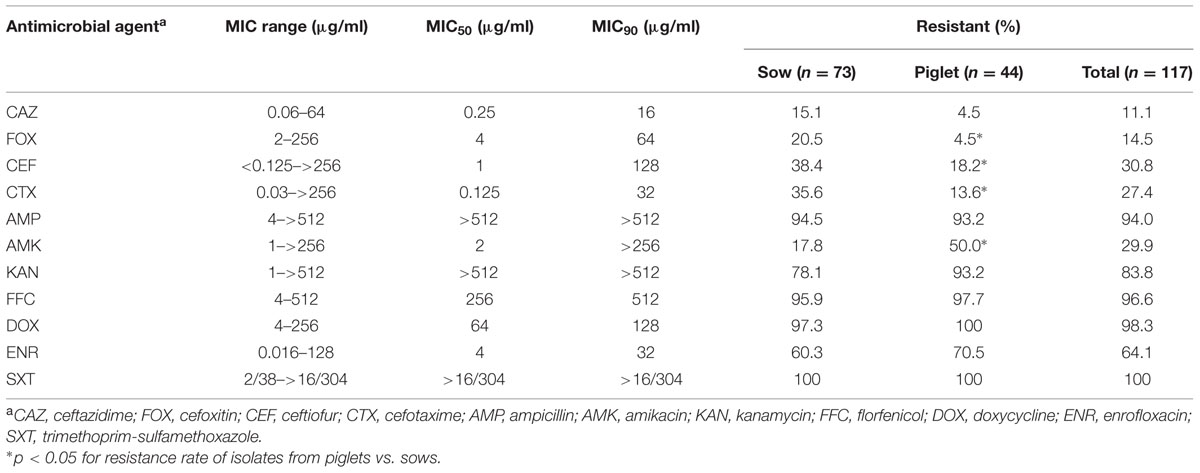
Rectal swab samples were randomly taken from one pig in every batch pen and 117 E. coli samples were isolated from 137 sow and piglet rectal swabs. The incidence for each group was approximately 85% (Table 1). Antimicrobial susceptibility tests showed that part of the isolates were resistant to CEF (30.8%), CTX (27.4%), FOX (14.5%), and CAZ (11.1%). Except for CAZ, cephalosporin resistance rates for sow isolates were significantly higher than those for isolates from piglets (p < 0.05). However, the resistance rate to CAZ for isolates from sows was still three times of that for piglet isolates (Table 2). In addition to cephalosporins, different levels of resistance to other classes of antimicrobials, including SXT (100%), DOX (98.3%), FFC (96.6%), AMP (94.0%), KAN (83.8%), ENR (64.1%) and AMK (29.9%), were also observed in the tested isolates (Table 2). The vast majority of the isolates (95.7%) exhibited a multidrug resistance (MDR) phenotype, and were resistant to four or more tested antimicrobial agents.
Detection of ESBL/pAmpC Genes
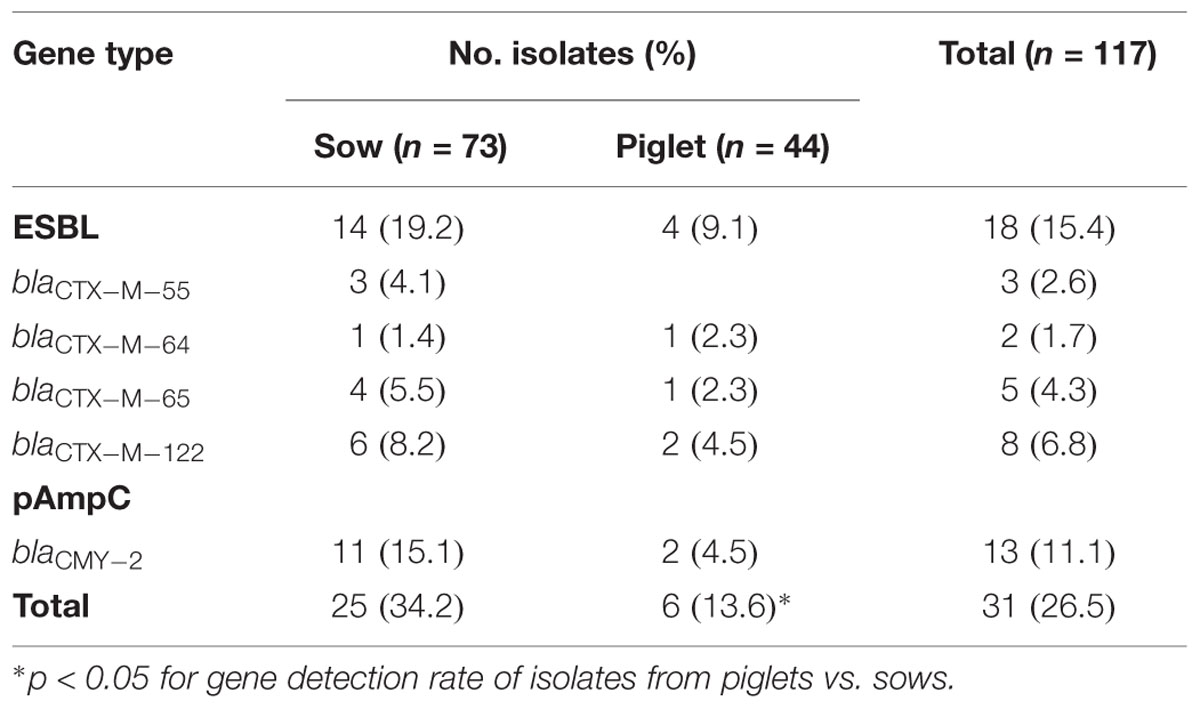
Among the 117 E. coli isolates, CTX-M-type ESBL genes were detected in 18 isolates (15.4%). The most common blaCTX-M-type were blaCTX-M-122 (n = 8), followed by blaCTX-M-65 (n = 5), blaCTX-M-55 (n = 3) and blaCTX-M-64 (n = 2). TEM- and SHV-type ESBL genes were not detected in any of the isolates. Compared with the diverse blaCTX-M genes detected in this study, blaCMY-2 was the only identified pAmpC gene, and was found in 13 isolates (11.1%). All the blaCMY-2-positive isolates were resistant to FOX with MICs ranging from 64 to 256 μg/ml, and also exhibited multi-resistance to AMP, CTX, CEF, KAN, FFC, DOX, ENR, and SXT (Figure 1).
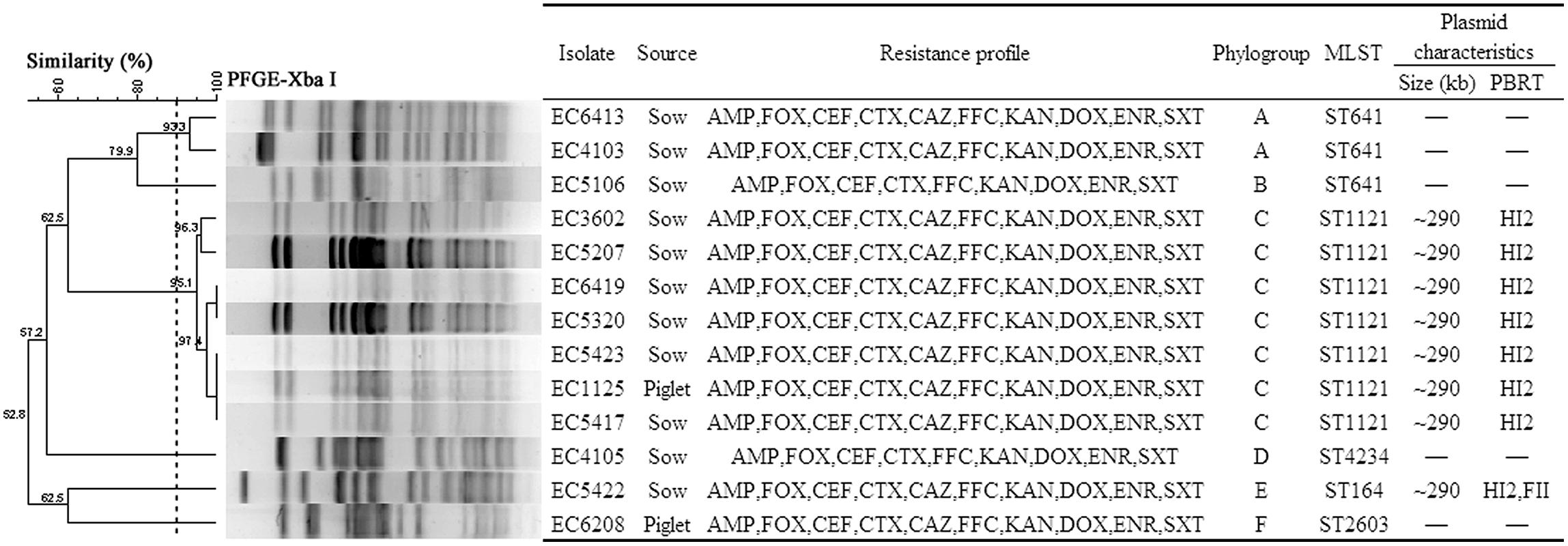
FIGURE 1. Clonal relationship and plasmid characteristics of the 13 blaCMY-2-positive Escherichia coli isolates. -, not detected. AMP, CAZ, FOX, CEF, CTX, KAN, FFC, DOX, ENR, and SXT are represented as in Table 2.
The co-existence of ESBL and pAmpC genes was not observed in any of the isolates. The total detection rate of the blaCMY-2/blaCTX-M genes in the sow isolates (34.2%) was significantly higher than that in the piglet isolates (13.6%; p < 0.05). The distribution of ESBL/pAmpC genes among the isolates are listed in Table 3.
Molecular Typing
We used cluster analysis of the blaCMY-2-positive isolates to generate dendrograms from PFGE profiles (Figure 1). Six phylogenetic groups (designated A–F) each with more than 90% similarity were represented in these 13 isolates. Group C contained seven isolates, Group A two, and the other four groups one each.
Multilocus sequence typing analysis of the 13 blaCMY-2-positive isolates identified five different STs including a novel one (ST4234). The most prevalent STs were ST1121 (n = 7) and ST641 (n = 3). The remaining isolates were each of a single ST type (Figure 1). Interestingly, the seven ST1121 isolates from six sows and one piglet were all contained in Group C. The three ST641 isolates from sows were divided between two groups (Figure 1).
Conjugation Assays and Plasmid Analysis
Eight blaCMY-2-positive transconjugants were successfully obtained from the seven ST1121 and one ST164 E. coli isolates. S1 nuclease-PFGE analysis identified a single plasmid in each of the seven transconjugants from ST1121 isolates. Two plasmids were identified in the transconjugant EC5422-25T derived from the ST164 isolate EC5422 (Supplementary Figure S1). Subsequently, Southern blot hybridization identified the blaCMY-2 gene located on a ∼290 kb plasmid in each of the transconjugants (Supplementary Figure S1). Replicon typing revealed the presence of the IncHI2 replicon in each of the eight transconjugants, but one of them (EC5422-25T) carried two replicons (IncHI2 and IncFII; Figure 1).
The IncHI2 plasmids of the ST1121 isolates shared indistinguishable RFLP profiles that were generated using XbaI digestion (data not shown). All of the transconjugants were resistant to FOX, CEF,CTX, AMP, KAN, and AMK. In addition, transfer of resistance to FFC, DOX, and SXT was also observed in the transconjugant EC5422-25T.
The metal susceptibility testing showed that the MICs of CuSO4 and AgNO3 for transconjugant EC5207-35T were higher than that of E. coli C600 (MICCuSO4 = 12 vs. 8 mM; MICAgNO3 = 1 vs. 0.0125 mM).
Complete Sequence of Plasmid pEC5207
We determined the DNA sequence of the plasmid derived from the blaCMY-2-positive transconjugants that were obtained from the seven ST1121 isolates. This plasmid pEC5207 is 272,865 bp in length with a GC content of 46.16% and harbors 253 predicted ORFs. The plasmid backbone is organized similarly to that of pSH111_227 (GenBank JN983042) from Salmonella sp. and encodes typical IncHI2 plasmid replication, partition, maintenance and transfer functions (Figure 2). The replication region of pEC5207 included a repHI2 gene. The parA and parB genes were involved in the plasmid partition. Two tra transfer regions and a hipAB toxin-antitoxin gene cluster were associated with the transfer and maintenance function of pEC5207. Interestingly, pEC5207 also contained a large number of genes encoding resistance to heavy metals including tellurium (terABCDEFWXYZ), silver (silABCEPRS) and copper (copABCE; Figure 2).
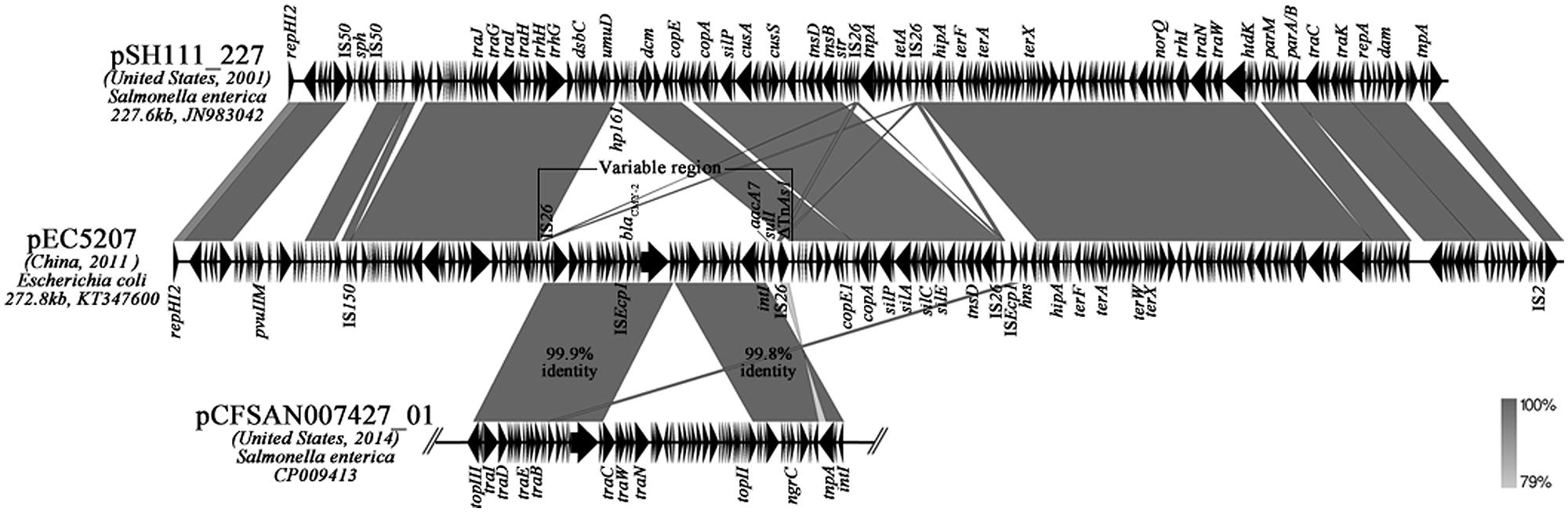
FIGURE 2. Linear comparison of IncHI2 plasmids pEC5207, pSH111_227 and blaCMY-2-carrying IncA/C plasmid pCFSAN007427_01. The arrows represent the positions and transcriptional direction of the ORFs. Regions of homology are shaded in gray.
The blaCMY-2 gene in pEC5207 is contained within a 48,888 bp variable region (Figure 2). The variable portion is located in a region between umuD and dcm in pEC5207, where is the hp161 locus in pSH111_227. The variable region showed high homology (>99%) with two parts of the blaCMY-2-carrying IncA/C plasmid pCFSAN007427_01 (GenBank CP009413) from Salmonella and contained an IS26 and a truncated TnAs1 at the ends. In addition to blaCMY-2, a new class I integron harboring aminoglycoside resistance gene aacA7 and sulfonamide resistance gene sulI was also found in this variable region.
Discussion
Antimicrobial susceptibility testing showed different resistance levels to cephalosporins and other classes of antimicrobials in E. coli isolates from a farrowing farm in Southern China. The antimicrobial usage records for this farm showed that doxycycline and the first generation cephalosporin cefradine were frequently added to drinking water or compound feeds as prophylactics. Other antimicrobials such as ceftiofur, enrofloxacin, florfenicol, and sulfamethoxazole were also commonly used for treatment during production. This background may be favorable for developing resistance to cephalosporins and other antimicrobials in this farm. The ESC-resistant E. coli usually carry additional genes conferring resistance to other veterinary antimicrobial agents like quinolone, aminoglycoside and florfenicol. In our previous study, the ESBL and pAmpC genes were detected in 67.5% of the PMQR-positive E. coli, indicating a strong association between ESC and quinolone resistance (Liu et al., 2013a). The 16S rRNA methylase genes, especially rmtB, and the florfenicol resistance gene floR are also commonly identified in ESBLs and pAmpC-producing E. coli isolates (Kang et al., 2009; Wu et al., 2009; Yu et al., 2010; Deng et al., 2011; Liu et al., 2013b; Guo et al., 2014). The use of different antimicrobial agents may increase the potential risk for selection of multidrug resistant isloates, and contribute to the MDR phenotypes of E. coli isolates in this farm.
Compared with the isolates from piglets, the isolates from sows showed significantly higher resistance rates to FOX, CEF and CTX, as well as a relatively higher resistance rate to CAZ. This coincided with the higher occurrence of the blaCMY-2 and blaCTX-M genes in the isolates from sows than in the isolates from piglets. Previous studies have shown that cephalosporin treatment can result in the selection of ESBL and AmpC producing E. coli in animals and aggravate the problem of cephalosporin resistance (Tragesser et al., 2006; Cavaco et al., 2008; Kanwar et al., 2013; Barton, 2014). The longer antibiotic exposure times of the sows may promote the persistence of ESBL/AmpC-producing E. coli in the gastrointestinal tract, thus resulting in the serious cephalosporin resistance and relatively high prevalence of ESBL/AmpC genes observed in sow isolates.
In this study the prevalence of the blaCMY-2 gene (11.1%) was lower than that of the blaCTX-M genes (15.4%). However, this level was much higher than those in our previous study (2.9%; Fang et al., 2015) and others’ (1.0–3.0%) in China (Liu et al., 2007; Guo et al., 2014; Rao et al., 2014). Previous studies have shown that cefoxitin/ceftiofur-resistant isolates from E. coli and Salmonella had a high degree of association with the production of CMY enzymes (Barton, 2014). Therefore, the use of ceftiofur in this farm probably contributed to the selection of blaCMY-2-producing E. coli isolates. MLST typing revealed a major ST type, ST1121, in seven of the thirteen blaCMY-2-positive isolates. Since they shared a high similarity (>95%) in PFGE profiles, this suggested a clonal spread of these ST1121 isolates. Furthermore, plasmid analysis of the transconjugants from the seven ST1121 isolates revealed an indistinguishable IncHI2 plasmid harboring blaCMY-2 in these isolates. Thus, the clonal spread of ST1121 isolates harboring IncHI2 plasmid may play an important role in the dissemination of blaCMY-2 in this farm. Notably, among the 13 blaCMY-2-positive isolates, one of the ST1121 isolates and the ST2603 isolate were isolated from piglets for market, which would indicate that the blaCMY-2-positive isolates may have been introduced into other swine farms by piglet trading and therefore would accelerate the spread of blaCMY-2. Further surveillance is necessary to determine the prevalence of blaCMY-2-positive isolates in these swine farms.
Since its first identification on plasmid from Klebsiella pneumoniae (Bauernfeind et al., 1990), the blaCMY-2 gene has been associated with various plasmid types. In our previous surveillance study, the blaCMY-2 gene was identified on IncA/C, IncHI2, and IncX plasmids (Fang et al., 2015). In the present study, blaCMY-2-carrying IncHI2 plasmids were not only detected in the clonal ST1121 isolates but also in a ST164 isolate. Additionally, the strong association of blaCMY-2 with IncHI2 plasmid was also observed in Salmonella (Shi, 2015). This highlighted the role of the IncHI2 plasmid in the transfer of blaCMY-2. Besides blaCMY-2, IncHI2 plasmid is also associated with another pAmpC gene blaCMY -8 (Chen et al., 2007b). In addition, IncHI2 plasmids have been implicated in the spread of blaCTX-M genes and are also frequently linked with the other antimicrobial resistance genes such as blaSHV, blaIMP, blaVIM, armA, qnrA1, qnrS1, and qnrB2 (Garcia Fernandez et al., 2007; Veldman et al., 2010; Coelho et al., 2012). Together, our data indicate that the IncHI2 plasmid played a significant role in the dissemination of antimicrobial resistance. However, to the best of our knowledge, there was no complete sequence analysis of a blaCMY-2-carrying IncHI2 plasmid to date. Therefore, in order to further characterize the blaCMY-2-carrying IncHI2 plasmids of ST1121 isolates, plasmid pEC5207 was sequenced in our study.
This plasmid possesses a typical IncHI2 plasmid backbone organized similarly to that of pSH111_227. A portion of hp161 in pSH111_227 was replaced with a blaCMY-2-harboring variable region in pEC5207. A sequence comparison revealed that this variable region might originate from IncA/C plasmid pCFSAN007427_01, and is flanked with IS26 and a truncated TnAs1. Considering the potential for genetic rearrangements by insertion sequence and transposon (Hallet and Sherratt, 1997; Bennett, 2004), we speculate that IS26 and TnAs1 transposition into the hp161 locus of a pSH111_227-like IncHI2 plasmid was followed by recombination with a pCFSAN007427_01-like blaCMY-2-carrying IncA/C plasmid. This could set the stage for the integration of blaCMY-2 into the IncHI2 plasmid.
Apart from the blaCMY-2 gene, pEC5207 also carried other resistance determinants for aminoglycosides (aacA7), sulfonamides (sul1), as well as heavy metals ions, such as Cu and Ag. And the result of susceptibility tests also confirm the functionality of pEC5207 in the transfer of copper and silver resistance. Copper is often present as trace element feed additives for animal production in China (Wang et al., 2013). Silver is used as disinfectants in water or surfaces in the animal production setting (Mourao et al., 2015). Under the selective pressure of heavy metals, cephalosporins, and other antimicrobials, plasmid pEC5207 could play a critical role in the persistence of host bacteria in the intestine of pigs, thereby increasing the risk for co-selection of isolates carrying blaCMY-2 gene. This may further explain the high prevalence of blaCMY-2 in this farm.
Conclusion
The present study revealed a relatively high prevalence of blaCMY-2 in a farrowing farm in Southern China. The clonal spread of ST1121 type E. coli harboring an IncHI2 plasmid mediated the dissemination of blaCMY-2 in this farm. Nucleotide sequence analysis and comparisons indicated that the blaCMY-2-carrying IncHI2 plasmid pEC5207 may have been generated by recombination with an IncA/C plasmid. pEC5207 may play an important role in the persistence of host bacteria under the selective pressure of heavy metal and antimicrobials. Although limited to a farrowing farm, our study indicates that there is a serious risk of dissemination of the blaCMY-2 gene by clonal spread and piglet training. Broad and longitudinal studies to determine the prevalence of blaCMY-2-positive E. coli in swine farms are required in the future.
Ethics Statement
This study protocol was reviewed and approved by the South China Agriculture University Animal Ethics Committee. The owner of the farm from which rectal swabs were taken gave permission for their animals to be used in this study.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by the Program for Changjiang Scholars and Innovative Research Team at University of Ministry of Education of China (Grant No. IRT13063), the Special Fund for Agro-scientific Research in the Public Interest (Grant No. 201203040), the National Natural Science Foundation (Grant No.31272609), and the Science and Technology Program of Guangzhou, China(Grant No. 2011J2200054).
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2015.01210
Footnotes
- ^ http://www.lahey.org/studies/webt.asp
- ^ http://mlst.ucc.ie/mlst/dbs/Ecoli
- ^ http://blast.ncbi.nlm.nih.gov
References
Aziz, R. K., Bartels, D., Best, A. A., Dejongh, M., Disz, T., Edwards, R. A., et al. (2008). The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9:75. doi: 10.1186/1471-2164-9-75
Barton, B. M., Harding, G. P., and Zuccarelli, A. J. (1995). A general method for detecting and sizing large plasmids. Anal. Biochem. 226, 235–240. doi: 10.1006/abio.1995.1220
Barton, M. D. (2014). Impact of antibiotic use in the swine industry. Curr. Opin. Microbiol. 19, 9–15. doi: 10.1016/j.mib.2014.05.017
Bauernfeind, A., Schweighart, S., Dornbusch, K., and Giamarellou, H. (1990). “A transferable cephamycinase in Klebsiella pneumoniae, abstr. A190,” in Program and Abstracts of the 30th Interscience Conference on Antimicrobial Agents and Chemotherapy. Washington, DC: American Society for Microbiology.
Bennett, P. M. (2004). Genome plasticity: insertion sequence elements, transposons and integrons, and DNA rearrangement. Methods Mol. Biol. 266, 71–113.
Bortolaia, V., Hansen, K. H., Nielsen, C. A., Fritsche, T. R., and Guardabassi, L. (2014). High diversity of plasmids harbouring blaCMY-2 among clinical Escherichia coli isolates from humans and companion animals in the upper Midwestern USA. J. Antimicrob. Chemother. 69, 1492–1496. doi: 10.1093/jac/dku011
Bradford, P. A. (2001). Extended-spectrum β-lactamases in the 21st century: characterization, epidemiology, and detection of this important resistance threat. Clin. Microbiol. Rev. 14, 933–951. doi: 10.1128/CMR.14.4.933-951.2001
Brooks, J. P., Adeli, A., and Mclaughlin, M. R. (2014). Microbial ecology, bacterial pathogens, and antibiotic resistant genes in swine manure wastewater as influenced by three swine management systems. Water Res. 57, 96–103. doi: 10.1016/j.watres.2014.03.017
Cabello, F. C., Godfrey, H. P., Tomova, A., Ivanova, L., Dolz, H., Millanao, A., et al. (2013). Antimicrobial use in aquaculture re-examined: its relevance to antimicrobial resistance and to animal and human health. Environ. Microbiol. 15, 1917–1942. doi: 10.1111/1462-2920.12134
Callens, B., Faes, C., Maes, D., Catry, B., Boyen, F., Francoys, D., et al. (2015). Presence of antimicrobial resistance and antimicrobial use in sows are risk factors for antimicrobial resistance in their offspring. Microb. Drug Resist. 21, 50–58. doi: 10.1089/mdr.2014.0037
Callens, B., Maes, D., Boyen, F., Haesebrouck, F., and Dewulf, J. (2012). “Antimicrobial resistance profile in faecal Escherichia coli of sows and their offspring in the farrowing unit and their interaction,” in Proceeding of the 13th International Symposium on Veterinary Epidemiology and Economics (ISVEE XIII), Wageningen: Wageningen Academic Publishers.
Carattoli, A., Bertini, A., Villa, L., Falbo, V., Hopkins, K. L., and Threlfall, E. J. (2005). Identification of plasmids by PCR-based replicon typing. J. Microbiol. Methods 63, 219–228. doi: 10.1016/j.mimet.2005.03.018
Cavaco, L. M., Abatih, E., Aarestrup, F. M., and Guardabassi, L. (2008). Selection and persistence of CTX-M-producing Escherichia coli in the intestinal flora of pigs treated with amoxicillin, ceftiofur, or cefquinome. Antimicrob. Agents Chemother. 52, 3612–3616. doi: 10.1128/AAC.00354-08
Chen, L., Chen, Z. L., Liu, J. H., Zeng, Z. L., Ma, J. Y., and Jiang, H. X. (2007a). Emergence of RmtB methylase-producing Escherichia coli and Enterobacter cloacae isolates from pigs in China. J. Antimicrob. Chemother. 59, 880–885. doi: 10.1093/jac/dkm065
Chen, Y. T., Lauderdale, T. L., Liao, T. L., Shiau, Y. R., Shu, H. Y., Wu, K. M., et al. (2007b). Sequencing and comparative genomic analysis of pK29, a 269-kilobase conjugative plasmid encoding CMY-8 and CTX-M-3 beta-lactamases in Klebsiella pneumoniae. Antimicrob. Agents Chemother. 51, 3004–3007. doi: 10.1128/AAC.00167-07
Chin, C. S., Alexander, D. H., Marks, P., Klammer, A. A., Drake, J., Heiner, C., et al. (2013). Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat. Methods 10, 563–569. doi: 10.1038/nmeth.2474
Clinical and Laboratory Standards Institute (2013a). Performance Standards for Antimicrobial Susceptibility Testing; 23rd Informational Supplement M100–S23. Wayne, PA: Clinical and Laboratory Standards Institute.
Clinical and Laboratory Standards Institute (2013b). Performance Standards for Antimicrobial Disk and Dilution Susceptibility Tests for Bacteria Isolated From Animals; Approved Standard-Fourth Edition. CLSI document VET01-A4. Wayne, PA: Clinical and Laboratory Standards Institute.
Coelho, A., Piedra-Carrasco, N., Bartolome, R., Quintero-Zarate, J. N., Larrosa, N., Cornejo-Sanchez, T., et al. (2012). Role of IncHI2 plasmids harbouring blaVIM-1, blaCTX-M-9, aac(6’)-Ib and qnrA genes in the spread of multiresistant Enterobacter cloacae and Klebsiella pneumoniae strains in different units at Hospital Vall d’Hebron. Barcelona, Spain. Int. J. Antimicrob. Agents 39, 514–517. doi: 10.1016/j.ijantimicag.2012.01.006
Crombe, F., Argudin, M. A., Vanderhaeghen, W., Hermans, K., Haesebrouck, F., and Butaye, P. (2013). Transmission dynamics of methicillin-resistant staphylococcus aureus in pigs. Front. Microbiol. 4:57. doi: 10.3389/fmicb.2013.00057
Deng, Y., He, L., Chen, S., Zheng, H., Zeng, Z., Liu, Y., et al. (2011). F33:A-:B- and F2:A-:B- plasmids mediate dissemination of rmtB-blaCTX-M-9 group genes and rmtB-qepA in Enterobacteriaceae isolates from pets in China. Antimicrob. Agents Chemother. 55, 4926–4929. doi: 10.1128/AAC.00133-11
Fang, L. X., Sun, J., Li, L., Deng, H., Huang, T., Yang, Q. E., et al. (2015). Dissemination of the chromosomally encoded CMY-2 cephalosporinase gene in Escherichia coli isolated from animals. Int. J. Antimicrob. Agents 46, 209–213. doi: 10.1016/j.ijantimicag.2015.04.003
Folster, J. P., Pecic, G., Mccullough, A., Rickert, R., and Whichard, J. M. (2011). Characterization of blaCMY-encoding plasmids among Salmonella isolated in the United States in 2007. Foodborne Pathog. Dis. 8, 1289–1294. doi: 10.1089/fpd.2011.0944
Garcia Fernandez, A., Cloeckaert, A., Bertini, A., Praud, K., Doublet, B., Weill, F. X., et al. (2007). Comparative analysis of IncHI2 plasmids carrying blaCTX-M-2 or blaCTX-M-9 from Escherichia coli and Salmonella enterica strains isolated from poultry and humans. Antimicrob. Agents Chemother. 51, 4177–4180. doi: 10.1128/AAC.00603-07
Greko, C., Badiola, J. I., Catry, B., Moreno, M. A., Pyörälä, S., Ruzauskas, M., et al. (2009). Reflection paper on the use of third and fourth generation cephalosporins in food producing animals in the European Union: development of resistance and impact on human and animal health. J. Vet. Pharmacol. Ther. 32, 515–533. doi: 10.1111/j.1365-2885.2009.01075.x
Guo, Y. F., Zhang, W. H., Ren, S. Q., Yang, L., Lu, D. H., Zeng, Z. L., et al. (2014). IncA/C plasmid-mediated spread of CMY-2 in multidrug-resistant Escherichia coli from food animals in China. PLoS ONE 9:e96738. doi: 10.1371/journal.pone.0096738
Hallet, B., and Sherratt, D. J. (1997). Transposition and site-specific recombination: adapting DNA cut-and-paste mechanisms to a variety of genetic rearrangements. FEMS Microbiol. Rev. 21, 157–178. doi: 10.1111/j.1574-6976.1997.tb00349.x
Jacoby, G. A. (2009). AmpC β-lactamases. Clin. Microbiol. Rev. 22, 161–182. doi: 10.1128/CMR.00036-08
Kang, H. Y., Kim, J., Seol, S. Y., Lee, Y. C., Lee, J. C., and Cho, D. T. (2009). Characterization of conjugative plasmids carrying antibiotic resistance genes encoding 16S rRNA methylase, extended-spectrum beta-lactamase, and/or plasmid-mediated AmpC beta-lactamase. J. Microbiol. 47, 68–75. doi: 10.1007/s12275-008-0158-3
Kanwar, N., Scott, H. M., Norby, B., Loneragan, G. H., Vinasco, J., Mcgowan, M., et al. (2013). Effects of ceftiofur and chlortetracycline treatment strategies on antimicrobial susceptibility and on tet(A), tet(B), and blaCMY-2 resistance genes among E. coli isolated from the feces of feedlot cattle. PLoS ONE 8:e80575. doi: 10.1371/journal.pone.0080575
Li, X. Z., Mehrotra, M., Ghimire, S., and Adewoye, L. (2007). Beta-Lactam resistance and beta-lactamases in bacteria of animal origin. Vet. Microbiol. 121, 197–214. doi: 10.1016/j.vetmic.2007.01.015
Liu, B. T., Liao, X. P., Yue, L., Chen, X. Y., Li, L., Yang, S. S., et al. (2013a). Prevalence of beta-lactamase and 16S rRNA methylase genes among clinical Escherichia coli isolates carrying plasmid-mediated quinolone resistance genes from animals. Microb. Drug Resist. 19, 237–245. doi: 10.1089/mdr.2012.0179
Liu, B. T., Yang, Q. E., Li, L., Sun, J., Liao, X. P., Fang, L. X., et al. (2013b). Dissemination and Characterization of Plasmids Carrying oqxAB-blaCTX-M Genes in Escherichia coli Isolates from Food-Producing Animals. PLoS ONE 8:e73947. doi: 10.1371/journal.pone.0073947
Liu, J. H., Wei, S. Y., Ma, J. Y., Zeng, Z. L., Lu, D. H., Yang, G. X., et al. (2007). Detection and characterisation of CTX-M and CMY-2 β-lactamases among Escherichia coli isolates from farm animals in Guangdong Province of China. Int. J. Antimicrob. Agents 29, 576–581. doi: 10.1016/j.ijantimicag.2006.12.015
Martin, L. C., Weir, E. K., Poppe, C., Reid-Smith, R. J., and Boerlin, P. (2012). Characterization of blaCMY-2 plasmids in Salmonella and Escherichia coli isolates from food animals in Canada. Appl. Environ. Microbiol. 78, 1285–1287. doi: 10.1128/AEM.06498-11
Mathew, A. G., Garner, K. N., Ebner, P. D., Saxton, A. M., Clift, R. E., and Liamthong, S. (2005). Effects of antibiotic use in sows on resistance of E. coli and Salmonella enterica Typhimurium in their offspring. Foodborne Pathog. Dis. 2, 212–220. doi: 10.1089/fpd.2005.2.212
McEwen, S. A., and Fedorka-Cray, P. J. (2002). Antimicrobial use and resistance in animals. Clin. Infect. Dis. 34 (Suppl. 3), S93–S106. doi: 10.1086/340246
Mourao, J., Novais, C., Machado, J., Peixe, L., and Antunes, P. (2015). Metal tolerance in emerging clinically relevant multidrug-resistant Salmonella enterica serotype 4,[5],12:i:- clones circulating in Europe. Int. J. Antimicrob. Agents 45, 610–616. doi: 10.1016/j.ijantimicag.2015.01.013
Perez-Perez, F. J., and Hanson, N. D. (2002). Detection of plasmid-mediated AmpC β-lactamase genes in clinical isolates by using multiplex PCR. J. Clin. Microbiol. 40, 2153–2162. doi: 10.1128/JCM.40.6.2153-2162.2002
Rao, L., Lv, L., Zeng, Z., Chen, S., He, D., Chen, X., et al. (2014). Increasing prevalence of extended-spectrum cephalosporin-resistant Escherichia coli in food animals and the diversity of CTX-M genotypes during 2003-2012. Vet. Microbiol. 172, 534–541. doi: 10.1016/j.vetmic.2014.06.013
Rasschaert, G., Michiels, J., Arijs, D., Wildemauwe, C., De Smet, S., and Heyndrickx, M. (2012). Effect of farm type on within-herd Salmonella prevalence, serovar distribution, and antimicrobial resistance. J. Food Prot. 75, 859–866. doi: 10.4315/0362-028X.JFP-11-469
Sandvang, D., Jensen, L. B., Baggesen, D. L., and Baloda, S. B. (2000). Persistence of a Salmonella enterica serotype typhimurium clone in Danish pig production units and farmhouse environment studied by pulsed field gel electrophoresis (PFGE). FEMS Microbiol. Lett. 187, 21–25. doi: 10.1111/j.1574-6968.2000.tb09130.x
Seiffert, S. N., Hilty, M., Perreten, V., and Endimiani, A. (2013). Extended-spectrum cephalosporin-resistant Gram-negative organisms in livestock: an emerging problem for human health? Drug Resist. Updat. 16, 22–45. doi: 10.1016/j.drup.2012.12.001
Shi, C. (2015). “Screening and characterization of antibiotic-resistant plasmids in Salmonella,” in Proceeding of the 2015 Annual Meeting July 25–28, 2015 (Portland: International Association for Food Protection).
Sullivan, M. J., Petty, N. K., and Beatson, S. A. (2011). Easyfig: a genome comparison visualizer. Bioinformatics 27, 1009–1010. doi: 10.1093/bioinformatics/btr039
Tenover, F. C., Arbeit, R. D., Goering, R. V., Mickelsen, P. A., Murray, B. E., Persing, D. H., et al. (1995). Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J. Clin. Microbiol. 33, 2233–2239.
Tian, G. B., Wang, H. N., Zhang, A. Y., Zhang, Y., Fan, W. Q., Xu, C. W., et al. (2012). Detection of clinically important beta-lactamases in commensal Escherichia coli of human and swine origin in western China. J. Med. Microbiol. 61, 233–238. doi: 10.1099/jmm.0.036806-0
Tragesser, L. A., Wittum, T. E., Funk, J. A., Winokur, P. L., and Rajala-Schultz, P. J. (2006). Association between ceftiofur use and isolation of Escherichia coli with reduced susceptibility to ceftriaxone from fecal samples of dairy cows. Am. J. Vet. Res. 67, 1696–1700. doi: 10.2460/ajvr.67.10.1696
van Duijkeren, E., Ikawaty, R., Broekhuizen-Stins, M. J., Jansen, M. D., Spalburg, E. C., De Neeling, A. J., et al. (2008). Transmission of methicillin-resistant Staphylococcus aureus strains between different kinds of pig farms. Vet. Microbiol. 126, 383–389. doi: 10.1016/j.vetmic.2007.07.021
Veldman, K., Dierikx, C., Van Essen-Zandbergen, A., Van Pelt, W., and Mevius, D. (2010). Characterization of multidrug-resistant, qnrB2-positive and extended-spectrum-β-lactamase-producing Salmonella Concord and Salmonella Senftenberg isolates. J. Antimicrob. Chemother. 65, 872–875. doi: 10.1093/jac/dkq049
Verdet, C., Gautier, V., Chachaty, E., Ronco, E., Hidri, N., Decre, D., et al. (2009). Genetic context of plasmid-carried blaCMY-2-like genes in Enterobacteriaceae. Antimicrob. Agents Chemother. 53, 4002–4006. doi: 10.1128/AAC.00753-08
Versalovic, J., Koeuth, T., and Lupski, J. R. (1991). Distribution of repetitive DNA sequences in eubacteria and application to fingerprinting of bacterial genomes. Nucleic Acids Res. 19, 6823–6831. doi: 10.1093/nar/19.24.6823
Wang, H., Dong, Y., Yang, Y., Toor, G. S., and Zhang, X. (2013). Changes in heavy metal contents in animal feeds and manures in an intensive animal production region of China. J. Environ. Sci. (China) 25, 2435–2442. doi: 10.1016/S1001-0742(13)60473-8
Witte, W. (1998). Medical consequences of antibiotic use in agriculture. Science 279, 996–997. doi: 10.1126/science.279.5353.996
Wu, Q., Zhang, Y., Han, L., Sun, J., and Ni, Y. (2009). Plasmid-mediated 16S rRNA methylases in aminoglycoside-resistant Enterobacteriaceae isolates in Shanghai. China. Antimicrob. Agents Chemother. 53, 271–272. doi: 10.1128/AAC.00748-08
Yu, F. Y., Yao, D., Pan, J. Y., Chen, C., Qin, Z. Q., Parsons, C., et al. (2010). High prevalence of plasmid-mediated 16S rRNA methylase gene rmtB among Escherichia coli clinical isolates from a Chinese teaching hospital. BMC Infect. Dis. 10:184. doi: 10.1186/1471-2334-10-184
Keywords: Escherichia coli, clonal spread, blaCMY-2, IncHI2 plasmid, farrowing farm
Citation: Deng H, Si H-B, Zeng S-Y, Sun J, Fang L-X, Yang R-S, Liu Y-H and Liao X-P (2015) Prevalence of extended-spectrum cephalosporin-resistant Escherichia coli in a farrowing farm: ST1121 clone harboring IncHI2 plasmid contributes to the dissemination of blaCMY-2. Front. Microbiol. 6:1210. doi: 10.3389/fmicb.2015.01210
Received: 06 August 2015; Accepted: 19 October 2015;
Published: 03 November 2015.
Edited by:
Alexandre Gonçalves, University of Trás-os-Montes e Alto Douro, PortugalCopyright © 2015 Deng, Si, Zeng, Sun, Fang, Yang, Liu and Liao. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xiao-Ping Liao, eHBsaWFvQHNjYXUuZWR1LmNu
†These authors have contributed equally to this work.
 Hui Deng1,2†
Hui Deng1,2† Jian Sun
Jian Sun Xiao-Ping Liao
Xiao-Ping Liao