- 1Department of Energy, Environmental and Chemical Engineering, Washington University, St. Louis, MO, USA
- 2Department of Chemistry and Biochemistry, Clark University, Worcester, MA, USA
- 3The Biodesign Institute, Arizona State University, Tempe, AZ, USA
Chloroflexus aurantiacus is an anoxygenic phototrophic bacterium. Its unique CO2 fixation pathway and primitive light-harvesting antenna complexes have attracted extensive research attentions. In this work, we investigated the photoheterotrophic growth of C. aurantiacus J-10-fl using acetate [at 55°C and without H2(g)]. The results indicate that glycine can promote anaerobic biomass production in a minimal medium by threefold to fivefold. Via 13C-metabolite analysis, we observed that glycine was involved in serine synthesis. Instead of being used as a major carbon source, glycine was degraded to produce C1 units and NAD(P)H. Tracer experiments also suggest that photoheterotrophic cultures growing with a exogenous glycine source exhibited capabilities of assimilating CO2 via multiple routes (including the 3-hydroxypropionate pathway). Finally, glycylglycine, a commonly used culture buffer, also significantly enhanced photoheterotrophic growth of C. aurantiacus, probably due to its thermal or enzymatic breakdown to glycine.
Introduction
Chloroflexus aurantiacus is a filamentous anoxygenic phototrophic bacterium isolated from hot springs (Hanada and Pierson, 2006). It has specialized light-harvesting antenna machines and performs a cyclic photosynthetic electron transport via a type II reaction center (Tang and Blankenship, 2013). Its photosystem does not generate NADPH directly, but can convert light energy into ATP via photosynthetic electron transfer. The genome of C. aurantiacus strain J-10-fl has been sequenced to facilitate our understanding of its physiology and cellular metabolism (Tang et al., 2011). C. aurantiacus shows a versatile carbon metabolism. In aerobic and dark conditions, it can grow on various organic substrates chemoheterotrophically. C. aurantiacus switches to photoheterotrophic growth when supplied with acetate under anaerobic and light conditions. In the presence of H2/CO2, C. aurantiacus can perform a photoautotrophic growth by fixing CO2 via the 3-hydroxypropionate (3HOP) bi-cycle pathway (Eisenreich et al., 1993; Zarzycki et al., 2009).
In its natural habitat, C. aurantiacus consumes organic nutrients (e.g., short-chain fatty acids, acetate, etc.) released from cyanobacteria in the associated microbial mats (Hanada and Pierson, 2006; Lee et al., 2014). Its phototrophic metabolisms for coassimilation of organic substrates have been extensively studied. For photoheterotrophic culture of C. aurantiacus, the common growth medium requires complex nutrients such as yeast extract or casamino acids (Madigan et al., 1974). In some early work on photoheterotrophic cultures, both acetate and mixed gases of H2 and CO2 were provided (Strauss et al., 1992). However, we found that strain J-10-fl was unable to grow well in acetate-based minimal media with phosphate buffer but without amino acids and H2 supplies. To delineate key exogenous nutrients demanded by C. aurantiacus, we performed 13C-tracer experiments. The results enhanced our understanding of C. aurantiacus carbon and energy metabolisms, and shed lights on C. aurantiacus survival in the ecosystem.
Materials and Methods
All chemicals and labeled substrates (13C-sodium acetate and NaH13CO3, purity > 98%) were purchased from Sigma–Aldrich (St. Louis, MO, USA). C. aurantiacus strain J-10-fl was grown in a minimal PE medium at 55°C. One liter medium contained 5 mL phosphate buffer solution, 5 mL basal salt solution, 0.5 g Na2S2O3 and 0.5 g (NH4)2SO4. One liter of phosphate solution contained 75 g KH2PO4 and 78 g K2HPO4. One liter of basal salt solution contained 4.12 g Na3EDTA, 1.11 g FeSO4⋅7H2O, 24.65 g MaSO4⋅7H2O, 2.94 g CaCl2⋅2H2O, 23.4 g NaCl and 10 mL tracer solution. One liter of tracer solution contained 11.2 g MnSO4⋅4H2O, 2.88 g ZnSO4⋅7H2O, 2.92 g Co(NO3)2⋅4H2O, 2.52 g CuSO4⋅5H2O, 2.42 g Na2MoO4⋅2H2O, 3.1 g H3BO3 and 41.2 g Na3EDTA. The medium pH was adjusted to 7.5. Commercial RPMI 1640 vitamins solution (100X, Sigma–Aldrich) was added into the sterile medium. In nutrient studies, yeast extract or amino acids were added into medium to test their effects on cell growth. For photoheterotrophic cultivation, the anaerobic cultures (purged with N2) were grown in sealed serum bottles (containing 30 mL culture) under continuous illumination (20–30 μmol photons m-2 s-1). Neither H2(g) nor CO2(g) was provided in the bottle headspace. Biomass growth was monitored based on optical densities at 600 nm.
Strain J-10-fl was cultivated in four tracer media: (a) 2 g/L [1-13C]acetate and 0.2 g/L unlabeled yeast extract; (b) 2 g/L [1,2-13C2]acetate (fully labeled acetate) and 0.05 g/L unlabeled glycine; (c) 2 g/L [1,2-13C2]acetate, 0.05 g/L glycine, and 0.5 g/L NaH13CO3; and (d) 2 g/L unlabeled acetate, 0.05 g/L unlabeled glycine, and 0.5 g/L NaH13CO3. In each tracer experiment, exponentially growing cells from unlabeled culture were inoculated in the labeled medium at a volume ratio of 1%. The protocol for 13C-metabolite analysis was described elsewhere (Tang et al., 2007; You et al., 2012). In brief, the biomass was hydrolyzed in HCl solution (100°C), and the resulting amino acid mixtures were derivatized by N-(tert-butyldimethylsilyl)-N-methyltrifluoroacetamide (TBDMS). GC-MS equipped with a DB5-MS column (Agilent Technologies, USA) was used to measure amino acid labeling pattern. The mass isotopomer distributions (MIDs) of amino acids were calculated based on five MS fragments (Wahl et al., 2004): [M-159]+ or [M-85]+ (both containing the carbon skeleton of amino acids after loss of their α carboxyl groups), [M-15]+ or [M-57] + (both containing the whole carbon skeleton of amino acids), and f302 (containing the first and the second carbon of amino acids). The final MS results, M0, M1, and M2, represent unlabeled, singly labeled, and doubly labeled amino acids, respectively (Figure 1). We also measured glycine concentrations in the cultures by GC-MS. In brief, culture samples were centrifuged and supernatant was collected. After the supernatant was dried, glycine from the supernatant was derivatized by TBDMS. Meanwhile, we determined the relationship between the glycine standards and corresponding MS abundances, which were used to estimate extracellular glycine concentrations in cultures.
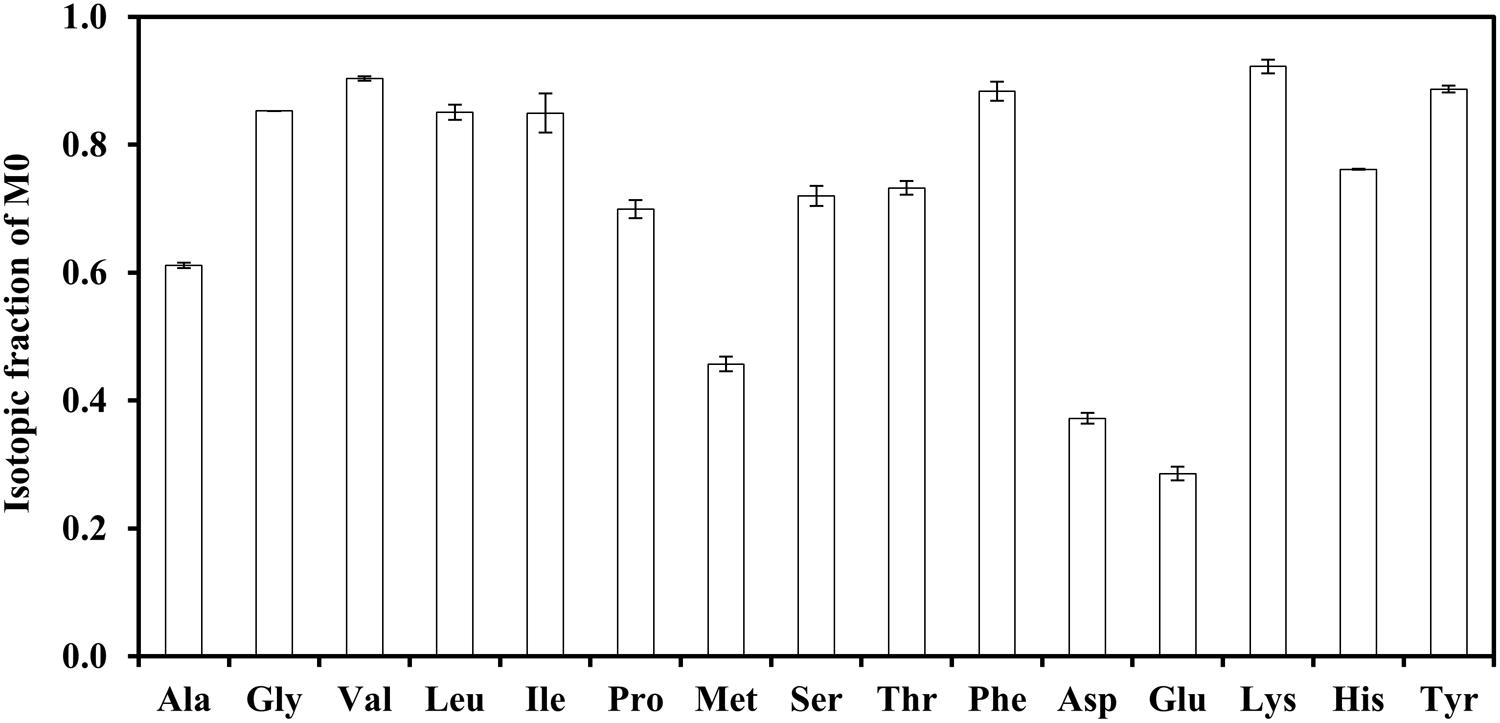
FIGURE 1. Investigation of the key nutrients enhancing the photoheterotrophic growth of strain J-10-fl. The medium contained 2 g/L [1-13C] sodium acetate and 0.2 g/L yeast extract. Unlabeled fractions (M0) of proteinogenic amino acids were mainly derived from yeast extract.
Results
Chloroflexus aurantiacus exhibits optimal growth in photoheterotrophic conditions when the medium is supplemented with yeast extract. It can also grow well with acetate and H2 in a minimal medium (Strauss et al., 1992). In our experiments, strain J-10-fl supplied with yeast extract reached an OD above 1.0 within 6 days during photoheterotrophic growth. Without H2 and yeast extract in the minimal medium, however, photoheterotrophic cultures grew poorly. To identify the biomass building blocks that were not effectively synthesized from primary substrate acetate, photoheterotrophic cultures were supplied with 13C-labeled sodium acetate and unlabeled yeast extract. Figure 1 shows the contribution of yeast extract to the synthesis of proteinogenic amino acids. Based on tracer experiments, a high fraction of 13C-carbon (40–70%) was incorporated into several amino acids, such as Ala, Glu, Met, and Asp, suggesting effective synthesis of those amino acids from acetate. Since the labeled carbon of [1-13C]acetate (purity > 98%) would be mostly incorporated into amino acids via central metabolic pathways, high unlabeled fractions of amino acids (e.g., Gly, Leu, and Phe) indicated that they were avidly absorbed from exogenous sources.
We then investigated the influences of these highly imported amino acids on the photoheterotrophic growth by replacing yeast extract with different amino acids. We observed the most significant growth enhancement (threefold to fivefold) when glycine was supplied (Figure 2), and a glycine concentration between 0.05 g/L and 0.25 g/L seems to suffice for enhancing the bacterial growth (Supplementary Figure S1). The influence of inorganic carbon was also investigated, which confirmed that addition of NaHCO3 (0.5 g/L) in medium slightly promoted photoheterotrophic growth, but much less than glycine did. Therefore, inorganic carbon source was not the rate limiting nutrients to the strain J-10-fl, and glycine played an important role in supporting the bacterial growth. These observations thus intrigued us to further investigate the role of glycine in C. aurantiacus metabolism.
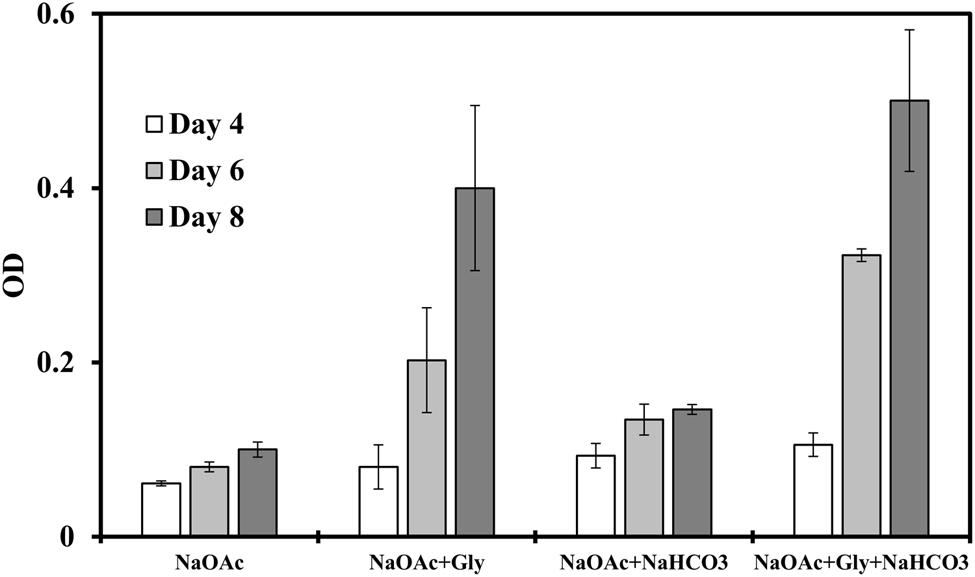
FIGURE 2. Strain J-10-fl growth on sodium acetate (NaOAc, 2 g/L) in a minimal medium with or without glycine (0.05 g/L) addition under photoheterotrophic conditions. NaHCO3 (0.5 g/L) was also supplemented to investigate its influence on the bacterial growth. Error bars represent the standard deviations of two biological replicates.
To this end, we first traced the fate of glycine in the central carbon metabolism via 13C labeling experiments. We cultivated strain J-10-fl photoheterotrophically with [1,2-13C2]acetate and unlabeled glycine, and then measured the isotopomer distributions of proteinogenic amino acids. Figure 3 shows that 13C from labeled acetate was incorporated into many key proteinogenic amino acids (e.g., Ala, Asp, and Glu) at significant levels (>90%), suggesting that glycine was not used as a carbon source for biomass synthesis. The only significant contributions of glycine to proteinogenic amino acids were glycine at 70% and serine (converted via glycine hydroxymethyltransferase, Caur_2543) at 30% (Supplementary Table S1). Substituting glycine with serine in minimal medium led to only a slight increase in cell growth (Supplementary Figure S1). Therefore, the profound growth enhancement by addition of glycine appears to be predominately due to factors other than direct conversion to biomass.
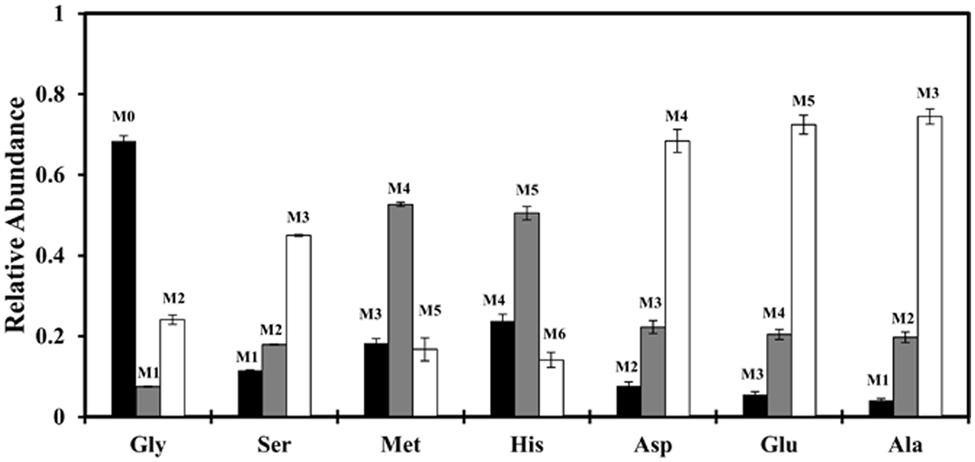
FIGURE 3. Mass isotopomer distributions (MIDs) of selected proteinogenic amino acids ([M-57]+) under photoheterotrophic condition. Strain J-10-fl was grown in a minimal medium supplied with [1,2-13C2] acetate (2 g/L) and unlabeled glycine (0.05 g/L). Error bars represent the standard deviations of two biological replicates.
Glycine metabolism of C. aurantiacus has been previously investigated during photoautotrophic growth (Herter et al., 2001). The researchers have shown that: (1) C. aurantiacus does not generate glycine from glyoxylate (a product of the 3HOP pathway), and (2) it exhibits high enzyme activity to produce C1 units from glycine cleavage. C1 units, including 5, 10-methylene-tetrahydrofolate (5, 10-methylene-THF), 5-methyl-THF and 10-formyl-THF, participate in biosynthesis of amino acids and inosine monophosphate. For example, 5, 10-methylene-THF, derived from glycine degradation, can be converted to 5-methyl-THF and 10-formyl-THF. The former contributes to Met synthesis and the latter His synthesis. As for cultures growing with [1,2-13C2] acetate and unlabeled glycine, the most abundant isotopologs of the resulting proteinogenic Met and His have a single unlabeled carbon (Figure 3), which suggested that C1 units are mainly synthesized from glycine cleavage under photoheterotrophic conditions. Following is the glycine cleavage reaction (Kikuchi et al., 2008):
Glycine cleavage can also be coupled with the serine hydroxymethyltransferase reaction:
In fact, the glycine consumption rate was higher than its requirement as a carbon source for biomass growth (Supplementary Figure S2). Therefore, part of glycine must be cleaved and oxidized by the THF-dependent C1 pathway, which contains successive steps that oxidize 5, 10-methylene-THF to formate or CO2, generating both NADPH and ATP (Fan et al., 2014). The fate of glycine/C1-metabolism and the distribution of C1-metabolic enzymes/genes have been discussed for bacteria and archaea, including Chloroflexi (Braakman and Smith, 2012). These studies suggest that glycine cleavage and C1 degradation can serve as a key energetic route to produce ATP, NADH, and NADPH. Therefore, it is likely that glycine is actively involved in the energy metabolism of C. aurantiacus under photoheterotrophic conditions.
In addition to glycine metabolism, we were also interested in investigating the CO2 fixation activity under photoheterotrophic conditions in the absence of H2, as C. aurantiacus possesses multiple carbon fixation routes (Tang et al., 2011). To this end, we added NaH13CO3 (0.5 g/L) to minimal medium containing unlabeled acetate and glycine (Figures 4A–D). A protein BLAST search against either Escherichia coli or Synechocystis 6803 carbonic anhydrase on NCBI website suggests that the gene encoding carbonic anhydrase that converts H13CO3- to 13CO2 is missing in C. aurantiacus, and an alternative 13CO2 source would be from the HCO3--CO2 equilibrium in the culture medium (pH = 7.5). Although addition of NaHCO3 did not appear to promote strain J-10-fl growth (Figure 2), enzyme activities of CO2 fixation was measurable due to significant 13C incorporation into proteinogenic amino acids (e.g., ∼40% alanine is singly labeled and 5% alanine is doubly labeled, Figure 4A and Supplementary Table S1). As pyruvate is involved in the CO2 fixation pathways and also a precursor to alanine, the labeling patterns of alanine can reflect CO2 fixation routes (Figure 5). MS data show that alanine was mostly labeled at first position, and a small fraction of alanine was labeled at both first and second positions (Supplementary Table S1). Figure 5B shows the origins of different labeling patterns of alanine. Firstly, unlabeled acetyl-CoA and H13CO3- can be condensed to [1-13C]pyruvate by pyruvate:ferredoxin oxidoreductase (PFOR, Caur_2080) or the 3HOP pathway. After a second H13CO3- is incorporated in the 3HOP pathway, doubly labeled malyl-CoA is formed, subsequently resulting in the formation of [1-13C]acetyl-CoA via a cleavage reaction. When the PFOR reaction or the 3HOP pathway converts [1-13C] acetyl-CoA into pyruvate, [1,2-13C2] pyruvate is generated, which explains the origin of [1,2-13C2] alanine. Under anaerobic conditions without H2, the flux of the 3HOP pathway for CO2 fixation is weak, and [1,2-13C2] alanine only accounts for ∼5% of the total alanine (Supplementary Table S1), probably due to the lack of reducing equivalents to power this pathway. As a consequence, most [1-13C] pyruvate would be generated from the PFOR reaction, which also leads to production of [1-13C] serine and [1-13C] glycine (Figures 4B,C) after pyruvate is converted into downstream metabolites. Figure 4D shows similar labeling patterns between Met (derived from oxaloacetate and C1 unit) and Asp (derived from oxaloacetate), further confirming that most C1 units come from glycine degradation (Figure 5A). This phenomenon also indicates that 13CO2 is not preferred to be fixed via the reductive C1 pathway (CO2 ↔ formyl-THF ↔ 5, 10-methylene-THF) under photoheterotrophic conditions, as 5, 10-methylene-THF can be continuously generated from glycine. We conclude that reducing equivalents from glycine degradation are insufficient to drive appreciable CO2 fixation as part of promoting heterotrophic growth.
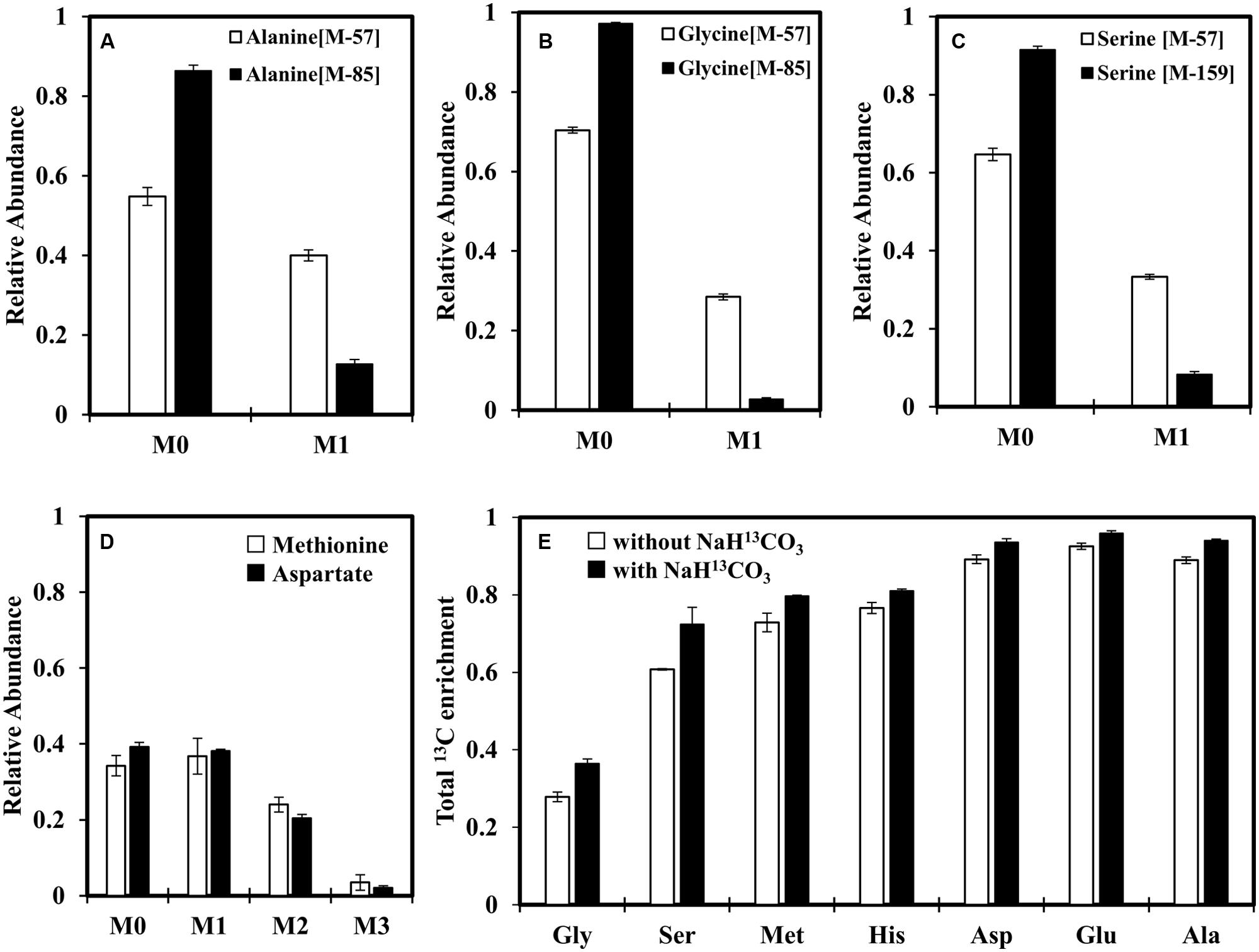
FIGURE 4. Investigation of MID of proteinogenic amino acids to reveal CO2 fixation under photoheterotrophic conditions. (A–C) MIDs of alanine, glycine and serine, respectively. (D) Comparison of MIDs between Met [M-57]+ and Asp [M-57]+. (A–D) Strain J-10-fl grew in a minimal medium supplied with unlabeled acetate (2 g/L), unlabeled glycine (0.05 g/L) and NaH13CO3 (0.5 g/L). (E) The total 13C enrichments of selected amino acids when J-10-fl was grown using [1,2-13C2] acetate (2 g/L) and unlabeled glycine (0.05 g/L) with or without NaH13CO3 (0.5 g/L) addition. Error bars represent the standard deviations of two biological replicates.
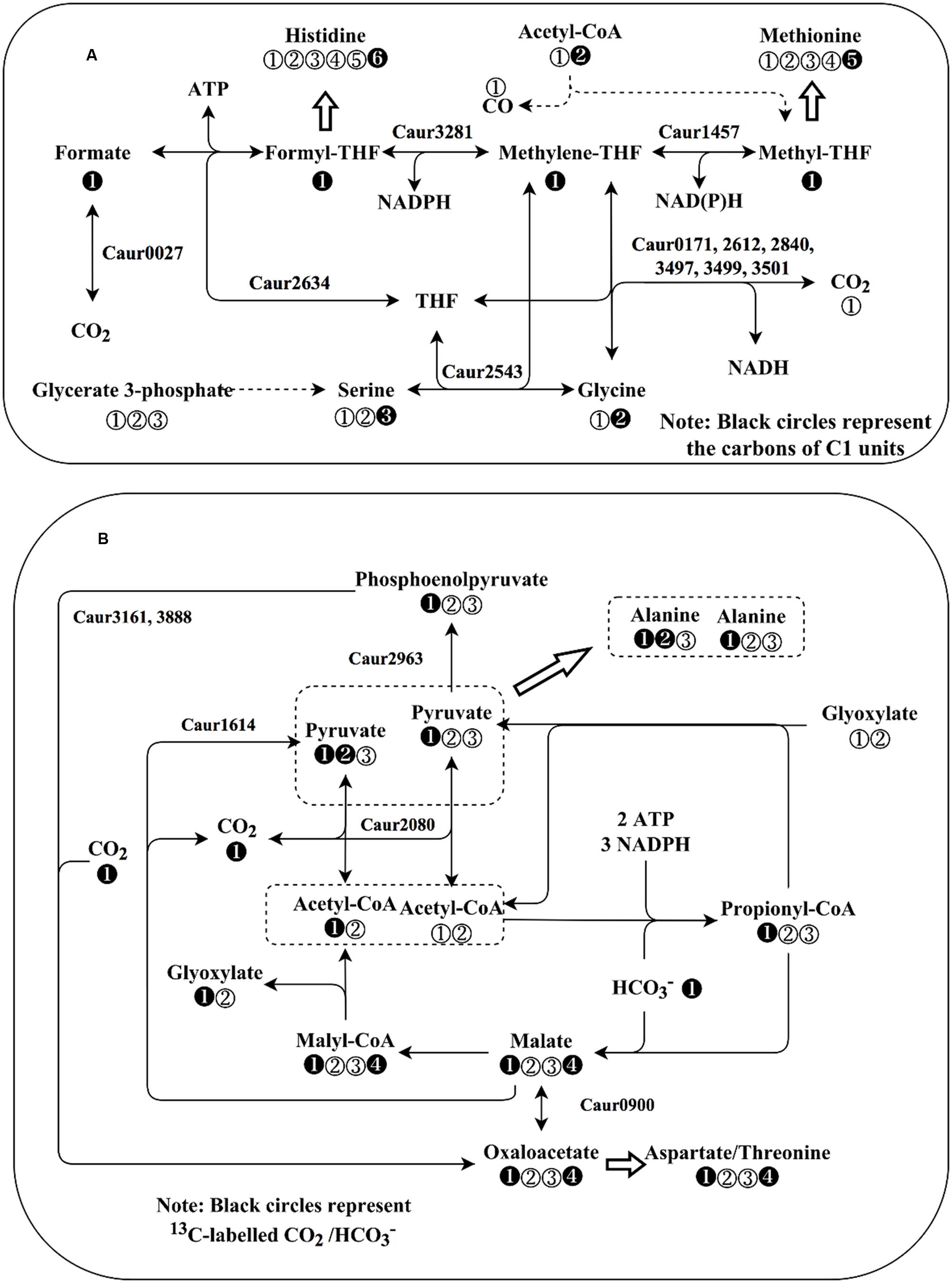
FIGURE 5. Schematic representation of C1 metabolism and CO2 fixation pathways in Chloroflexus aurantiacus. (A) C1 metabolism of C. aurantiacus. Black circles in (A) represent carbon atoms that will be donated to or originate from C1 unit carbon atoms. Dashed lines represent the genes that are not annotated in strain J-10-fl. Note: C. aurantiacus lacks the annotation of formate dehydrogenase gene, but it may use alternative enzyme (formylmethanofuran dehydrogenase, Caur_0027) for formate oxidation (Bertram et al., 1994). (B) Annotated CO2 fixation pathways in C. aurantiacus. The secondary CO2 fixation route (3HOP pathway) can generate acetyl-CoA of different labeling patterns. Black circles in (B) represent 13CO2 or H13CO3-. In both (A,B), the numbers in the circles represent the positions of carbon atoms in corresponding intracellular metabolites.
Figure 4E shows that 13C enrichment shifts of various amino acids after NaH13CO3 addition into minimal medium containing [1,2-13C2]acetate and unlabeled glycine. When strain J-10-fl was grown without NaH13CO3, 70% proteinogenic glycine and 40% proteinogenic serine were unlabeled. Other key proteinogenic amino acids (i.e., Ala, Asp and Glu) were labeled significantly (13C enrichments ∼90%). The small fraction of 12C in these amino acids was possibly from the metabolic assimilation of 12CO2 released from glycine degradation. When strain J-10-fl was cultivated with NaH13CO3, unlabeled glycine and fully labeled acetate, 13C enrichments in all proteinogenic amino acids were further raised (e.g., ∼95% of Glu carbons were 13C-carbons). This observation confirms photoheterotrophic CO2 co-utilizations by strain J-10-fl.
Discussion
In this study, we attempted to understand why glycine can promote C. aurantiacus growth on acetate. Herter et al. (2001) have previously reported the contribution of glycine to photoautotrophic growth of C. aurantiacus strain OK-70-fl with H2/CO2. The study indicated that glycine could participate in serine synthesis and contribute approximately half of the C1 units to biomass synthesis. As a comparison, we tested the growth effect of glycine on strain OK-70-fl strain and revealed a similar growth enhancement (by threefold to fourfold) on strain J-10-fl during photoheterotrophic growth (Supplementary Figure S3 and related contents). Whether glycine can promote photoheterotrophic growth in the presence of H2 remains to be tested.
Chloroflexus aurantiacus does not contain a type I reaction center, and its cyclic photosynthetic electron transport system within the type-II reaction center generates ATP. We searched the genome database of strain J-10-fl and found the gene encoding a nickel-dependent hydrogenase (Caur_1188), which may generate NADPH in the presence of H2. However, when C. aurantiacus grows on acetate without H2, its NADPH generation could be less efficient. The major routes in central carbon metabolism for producing reducing power are the TCA cycle and oxidative pentose phosphate pathway (OPPP). However, acetate-based metabolism usually does not show strong fluxes through the OPPP, and both PFOR reaction and gluconeogenesis consume reducing equivalents. Additionally, genes encoding transhydrogenase have not yet been reported to exist in the genome of C. aurantiacus. Therefore, NADH and NADPH production from glycine cleavage and C1 degradation may increase energy flexibility and thus promote C. aurantiacus anaerobic growth. On the other hand, the THF-dependent C1 degradation pathway in C. aurantiacus appears to be influenced by H2. In our study, glycine cleavage contributes to C1 units for both Met and His synthesis under photoheterotrophic conditions. In comparisons, Herter et al. (2001) have investigated C. aurantiacus phototrophic growth with fully labeled glycine and H2/unlabeled CO2. Their labeling data of Met and His revealed that 5,10-methylene-THF (the C1 unit for Met synthesis) derived from fully labeled glycine was not converted to formyl-THF (the C1 unit for His synthesis). This phenomenon implied that H2 may inhibit the C1 unit oxidation, and that formyl-THF synthesis could be formed reductively from CO2 in the presence of H2. Lastly, it is possible that glycine may be also involved in other unknown mechanisms promoting C. aurantiacus photoheterotrophic growth. For example, a previous report has discovered the marine bacterium Pelagibacter ubique possesses all the genes for amino acid biosynthesis, but is still effectively auxotrophic for glycine (Tripp et al., 2009).
An interesting fact is that glycylglycine, a dipeptide of glycine, has been employed in the earliest studies of C. aurantiacus (Madigan et al., 1974), which gave the best growth of C. aurantiacus compared to other buffers (e.g., Tris, phosphate, and MOPS). Notably, glycylglycine is known to be a good buffer for biological systems since it is relatively non-toxic (Smith and Smith, 1949). In the absence of H2, C. aurantiacus grew much better in glycylglycine than in MOPS or other buffers (Supplementary Figure S3). Glycylglycine could be hydrolyzed at high temperature during medium autoclave and cell incubations (Radzicka and Wolfenden, 1996), or degraded by a membrane dipeptidase (Caur_2632). Our isotopic analysis further confirmed that C. aurantiacus growing on 100% [1-13C] acetate in the presence of unlabeled glycylglycine buffer possessed significantly unlabeled proteinogenic glycine (i.e., ∼90% of proteinogenic glycine was completely unlabeled, while only 3% of proteinogenic glutamate was completely unlabeled). Since the culture was grown in minimal medium containing 13C-acetate as the sole carbon source, the unlabeled glycine in the biomass must come from unlabeled glycylglycine. All these evidences imply that glycylglycine could be considered as an exogenous source of glycine contributing to growth enhancement for C. aurantiacus.
Conclusion
In this study, we traced glycine in the central carbon metabolism to answer how it can enhance photoheterotrophic growth of C. aurantiacus in the absence of H2. Our results, together with previous studies and genome annotations, indicate that glycine can be used for producing biomass (mainly glycine and serine), but more importantly glycine molecules are degraded via cleavage reactions, serving as an important route for NAD(P)H production for acetate-grown C. aurantiacus cultures. As it is widely known, glycine is the simplest and also the most abundant amino acid that can be synthesized abiotically on the primitive Earth (Miller, 1953). Glycine cleavage and C1 metabolism might be the ancestral energy generation pathways (Braakman and Smith, 2012).
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank the funding support from the NASA astrobiology program (NNX12AD85G) and NSF (DBI 1356669). We also would like to thank Professor Madigan at Southern Illinois University for providing Chloroflexus aurantiacus strain OK-70-fl.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2015.01467
References
Bertram, P. A., Karrasch, M., Schmitz, R. A., Böcher, R., Albracht, S. P., and Thauer, R. K. (1994). Formylmethanofuran dehydrogenases from methanogenic Archaea Substrate specificity, EPR properties and reversible inactivation by cyanide of the molybdenum or tungsten iron-sulfur proteins. Eur. J. Biochem. 220, 477–484. doi: 10.1111/j.1432-1033.1994.tb18646.x
Braakman, R., and Smith, E. (2012). The emergence and early evolution of biological carbon-fixation. PLoS Comput. Biol. 8:e1002455. doi: 10.1371/journal.pcbi.1002455
Eisenreich, W., Strauss, G., Werz, U., Fuchs, G., and Bacher, A. (1993). Retrobiosynthetic analysis of carbon fixation in the phototrophic eubacterium Chloroflexus aurantiacus. Eur. J. Biochem. 215, 619–632. doi: 10.1111/j.1432-1033.1993.tb18073.x
Fan, J., Ye, J., Kamphorst, J. J., Shlomi, T., Thompson, C. B., and Rabinowitz, J. D. (2014). Quantitative flux analysis reveals folate-dependent NADPH production. Nature 510, 298–302. doi: 10.1038/nature13236
Hanada, S., and Pierson, B. (2006). “The family Chloroflexaceae,” in The Prokaryotes, eds M. Dworkin, S. Falkow, E. Rosenberg, K.-H. Schleifer, and E. Stackebrandt (New York, NY: Springer), 815–842.
Herter, S., Farfsing, J., Gad’On, N., Rieder, C., Eisenreich, W., Bacher, A., et al. (2001). Autotrophic CO2 fixation by Chloroflexus aurantiacus: study of glyoxylate formation and assimilation via the 3-hydroxypropionate cycle. J. Bacteriol. 183, 4305–4316. doi: 10.1128/jb.183.14.4305-4316.2001
Kikuchi, G., Motokawa, Y., Yoshida, T., and Hiraga, K. (2008). Glycine cleavage system: reaction mechanism, physiological significance, and hyperglycinemia. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 84, 246–263. doi: 10.2183/pjab.84.246
Lee, J. Z., Burow, L. C., Woebken, D., Everroad, R. C., Kubo, M. D., Spormann, A. M., et al. (2014). Fermentation couples Chloroflexi and sulfate-reducing bacteria to Cyanobacteria in hypersaline microbial mats. Front. Microbiol. 5:61. doi: 10.3389/fmicb.2014.00061
Madigan, M., Petersen, S., and Brock, T. (1974). Nutritional studies on Chloroflexus, a filamentous photosynthetic, gliding bacterium. Arch. Microbiol. 100, 97–103. doi: 10.1007/BF00446309
Miller, S. L. (1953). A production of amino acids under possible primitive earth conditions. Science 117, 528–529. doi: 10.2307/1680569
Radzicka, A., and Wolfenden, R. (1996). Rates of uncatalyzed peptide bond hydrolysis in neutral solution and the transition state affinities of proteases. J. Am. Chem. Soc. 118, 6105–6109. doi: 10.1021/ja954077c
Smith, M. E., and Smith, L. B. (1949). Piperazine dihydrochloride and glycylglycine as non-toxic buffers in distilled water and in sea water. Biol. Bull. 96, 233–237. doi: 10.2307/1538357
Strauss, G., Eisenreich, W., Bacher, A., and Fuchs, G. (1992). 13C-NMR study of autotrophic CO2 fixation pathways in the sulfur-reducing Archaebacterium Thermoproteus neutrophilus and in the phototrophic Eubacterium Chloroflexus aurantiacus. Eur. J. Biochem. 205, 853–866. doi: 10.1111/j.1432-1033.1992.tb16850.x
Tang, K.-H., Barry, K., Chertkov, O., Dalin, E., Han, C., Hauser, L., et al. (2011). Complete genome sequence of the filamentous anoxygenic phototrophic bacterium Chloroflexus aurantiacus. BMC Genomics 12:334. doi: 10.1186/1471-2164-12-334
Tang, K. H., and Blankenship, R. E. (2013). “Photosynthetic electron transport,” in Encyclopedia of Biophysics, ed. G. C. K. Roberts (Berlin: Springer-Verlag).
Tang, Y. J., Meadows, A. L., Kirby, J., and Keasling, J. D. (2007). Anaerobic central metabolic pathways in Shewanella oneidensis MR-1 reinterpreted in the light of isotopic metabolite labeling. J. Bacteriol. 189, 894–901. doi: 10.1128/JB.00926-06
Tripp, H. J., Schwalbach, M. S., Meyer, M. M., Kitner, J. B., Breaker, R. R., and Giovannoni, S. J. (2009). Unique glycine-activated riboswitch linked to glycine-serine auxotrophy in SAR11. Environ. Microbiol. 11, 230–238. doi: 10.1111/j.1462-2920.2008.01758.x
Wahl, S. A., Dauner, M., and Wiechert, W. (2004). New tools for mass isotopomer data evaluation in 13C flux analysis: mass isotope correction, data consistency checking, and precursor relationships. Biotechnol. Bioeng. 85, 259–268. doi: 10.1002/bit.10909
You, L., Page, L., Feng, X., Berla, B., Pakrasi, H. B., and Tang, Y. J. (2012). Metabolic pathway confirmation and discovery through 13C-labeling of proteinogenic amino acids. J. Vis. Exp. 26:e3583. doi: 10.3791/3583
Keywords: 3-hydroxypropionate, 13C, C1 metabolism, CO2 fixation, formyltetrahydrofolate
Citation: He L, Wang Y, You L, Khin Y, Tang JK-H and Tang YJ (2015) Glycine Cleavage Powers Photoheterotrophic Growth of Chloroflexus aurantiacus in the Absence of H2. Front. Microbiol. 6:1467. doi: 10.3389/fmicb.2015.01467
Received: 18 March 2015; Accepted: 07 December 2015;
Published: 22 December 2015.
Edited by:
John R. Battista, Louisiana State University and A & M College, USAReviewed by:
Aharon Oren, The Hebrew University of Jerusalem, IsraelRogier Braakman, Massachusetts Institute of Technology, USA
Copyright © 2015 He, Wang, You, Khin, Tang and Tang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yinjie J. Tang, eWluamllLnRhbmdAc2Vhcy53dXN0bC5lZHU=; Joseph K.-H. Tang, amt0YW5nQGFzdS5lZHU=
 Lian He
Lian He Yaya Wang
Yaya Wang Le You1
Le You1 Joseph K.-H. Tang
Joseph K.-H. Tang Yinjie J. Tang
Yinjie J. Tang