- 1School of Public Health, Wannan Medical College, Wuhu, China
- 2Clinical Laboratory, Institute of Dermatology, Chinese Academy of Medical Sciences – Peking Union Medical College, Nanjing, China
- 3Clinical Laboratory, Zhongda Hospital, School of Medicine, Southeast University, Nanjing, China
- 4Pediatric Research Institute, Nanjing Children’s Hospital, Nanjing Medical University, Nanjing, China
- 5Clinical Laboratory, Ma’anshan Center for Disease Control and Prevention, Ma’anshan, China
Background: Recently, Mycobacterial Interspersed Repetitive Unit (MIRU) was supposed to be associated with drug resistance in Mycobacterium tuberculosis (M. tuberculosis), but whether the association exists actually in local strains in China was still unknown. This research was conducted to explore that association and the predictability of MIRU to drug resistance of Tuberculosis (TB).
Methods: The clinical isolates were collected and the susceptibility test were conducted with Lowenstein–Jensen (LJ) medium for five anti-TB drug. Based on PCR of MIRU-VNTR (Variable Number of Tandem Repeat) genotyping, we tested the number of the repeat unite of MIRU. Then, we used logistic regression to evaluate the association between 15 MIRU and drug resistance. In addition, we explored the most suitable MIRU locus of identified MIRU loci for drug resistance by multivariate logistic regression.
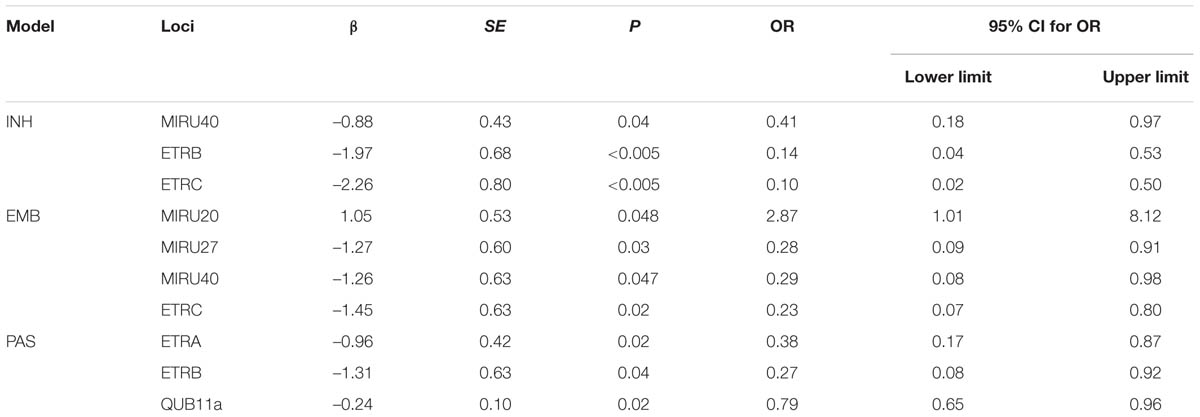
Results: Of the 102 strains, one isolate was resistant to rifampicin and one isolate was resistant to streptomycin. Among these fifteen MIRU, there was a association between MIRU loci polymorphism and anti-tuberculosis drug resistance, ETRB (P = 0.03, OR = 0.19, 95% CI 0.05–0.81) and ETRC (P = 0.01, OR = 0.14, 95% CI 0.03–0.64) were negatively related to isoniazid resistance; MIRU20 (P = 0.05, OR = 2.87, 95% CI 1.01–8.12) was positively associated with ethambutol resistance; and QUB11a (P = 0.02, OR = 0.79, 95% CI 0.65–0.96) was a negative association factor of p-aminosalicylic acid resistance.
Conclusion: Our research showed that MIRU loci may predict drug resistance of tuberculosis in China. However, the mechanism still needs further exploration.
Introduction
Tuberculosis (TB) caused by Mycobacterium tuberculosis (M. tuberculosis) infection has affected human beings for 1000s of years. Due to patients defaulting on TB therapy, there has been an increasing number of drug resistant TB strains, especially multidrug-resistant TB (MDR-TB; Biadglegne et al., 2014). In 2012, 1.3 million people died from TB, of which 3.6% of all new and 20% of all recurring cases involved infection with MDR-TB1. Particularly, the prevalent genotype in East Asia, Beijing genotype M. tuberculosis, has shown drug resistance (Liu et al., 2012). Drug resistance, especially MDR-TB, has made treating TB extremely difficult.
Genetic factors play important roles in drug resistance specific to TB treatment. For example, isoniazid (INH), rifampicin (RFP), and streptomycin (SM) resistance were found to be related to genes katG315 (Lu et al., 2014), rpoB513 (Jun et al., 2009), and gidB A80P (Perdigão et al., 2014), respectively. In addition, an efflux pump capable of effluxion of INH was found to be related to five genes (Rodrigues et al., 2012). The underlying mechanisms of these resistances, however, are still unclear.
Mycobacterial Interspersed Repetitive Unit is a type of the “minisatellite” VNTR (Variable Number of Tandem Repeat) with the length of MIRU repetitive unit ranging from 50 to 100 base pairs (bp) (Supply et al., 2001). A series of studies suggested that MIRU loci could be involved in the expression regulation of neighboring genes. For example, when the repeat units of Mtub39 locus repeat fourth, the expression level of its downstream gene lpdA was 12-fold as that of the same gene when the frequency of repeat units of Mtub39 locus was 1 (Akhtar et al., 2009). Similar results were showed in another experiment (Pérez-Lago et al., 2013): ∼56% increase in expression level of lpdA when the frequency of repeat units of Mtub39 was from 3 to 4. Tantivitayakul et al. (2010) also found that MIRU10 could promote the expression of downstream gene gfp. In addition, different frequencies of tandem repeats may influence the folding of DNA, thereby affecting the attraction, binding, and interaction of transcription factors (Supply et al., 1997).
Our previous studies have confirmed/verified the hypothesis that some MIRU loci polymorphism were related to anti-tuberculosis drug resistance (Yu-feng et al., 2015). This study was conducted using strains worldwidely, of which 54 were from Germany, 20 were from Ghana, 20 were from Uganda, and 15 were from former Soviet Union. However, whether the association between MIRU loci and drug resistance exists in China or other countries were still unknown. Therefore, in this study, we sampled 102 clinical strains of M. tuberculosis from eastern China to explorethe association between the MIRU loci and drug resistance.
Materials and Methods
Selection of Clinical Isolates
A total of 102 clinical isolates of M. tuberculosis were selected from sputum of patients who were recruited between 2009 and 2012 in the eastern area of Anhui Province, China. The standard M. tuberculosis strain H37Rv was used as the control isolate, which was courteously provided by Dr. Chuanyou Li of Beijing Chest Hospital, China.
Drug Susceptibility Testing
Four first-line anti-tuberculosis drugs including RFP, INH, SM, ethambutol (EMB), and one second-line drug of p-aminosalicylic acid (PAS) were incorporated into LJ medium, at the following concentrations: INH, 0.2 μg/mL; RFP, 40.0 μg/mL; SM, 4.0 μg/mL; EMB, 2.0 μg/mL; PAS, 1.0 μg/mL2. The proportion method was used to detect the drug-resistance of the M. tuberculosis. Strain was scored as resistant to a specific drug, or defined as sensitive if its growth rate was <1% compared to the control. Strain isolation, identification, and drug susceptibility tests (DST) were performed at the laboratory of Ma Anshan Centre for Disease Control and Prevention.
MIRU Loci Selection
According to the Chinese genotyping handbook and protocol referenced literature (Lee et al., 2014), 15 MIRU loci were selected in this study. The primer sequence and length of repeat units of each MIRU locus was shown in Table 1.
DNA Preparation and PCR Amplification
Sputum specimens were treated with 4% NaOH and inoculated on LJ medium for 4–6 weeks. The colony of 1–2 rings was dissolved in 400 μL sterile distilled water on the vortex device. In addition, the liquid supernatant was stored with -20°C cryopreservation after being boiled in a water bath for 30 min and clarified by centrifugation at 4°C at 12,000 rpm for 15 min.
Each PCR mix contained 12.5 μL of 2x PCR buffer (Tiangen Biotech Co., Ltd., Beijing, China), 0.25 μM of each primer and 6 μL of sterile distilled water. Four microliters of template DNA was added to each reaction mixture. Following an initial denaturation at 95°C for 10 min the PCRs were subjected to 40 thermocycles of 94°C for 60 s, 60°C for 60 s, and 72°C for 90 s. This was followed by a final extension at 72°C for 10 min. Positive and negative control reactions, in which PCR mixes were inoculated with 4 μL of boiled cells from the M. tuberculosis H37RV strain and sterile distilled water, respectively, were performed with each set of reactions (Lee et al., 2014).
The PCR products were electrophoretically separated on 2% agarose gel (Life Technologies, Grand Island, NY, USA) in 1x Tris-boric acid-EDTA buffer (Tiangen). A 100-bp DNA stepladder (Thermo Fisher Scientific Inc., Waltham, MA, USA) was used as the marker.
Data Analysis
The allelic diversity (h-value) of MIRU loci, individually and in combination, was calculated by the following equation: , where n is the number of isolates and xi is the frequency of its allele at the locus (Kremer et al., 2005).
Univariate logistic regression with the enter method was used to initially evaluate the associations between MIRU loci and anti-tuberculosis drug resistance. Multivariate logistic regression with the stepwise method was used to analyze the major risk and protective factors of MIRU loci initially selected from the univariate logistic regression.
Results
Distribution of Repeat Units, Polymorphism, and Drug Resistance Rate of MIRU Loci
The number of repeat units in the 15 MIRU loci ranged from 0 to 12. The h-value, which measures the allelic diversity of MIRU loci, ranged from 0.03 for ETRC to 0.77 for OUB11a. Of the drug resistance, 22 were resistant to INH, 20 were resistant to RFP, 5 were resistant to EMB, and 26 were resistant to PAS. The anti-TB drug rates of tuberculosis strains with different copies of MIRU locus unit ranged from 0.0 to 100.0% (Supplementary Table 1).
Association between MIRU Loci and Drug-Resistance in M. tuberculosis
According to univariate logistic regression analysis, we found a number of MIRU loci associated with the drug resistance (Table 2). For example, ETRB was associated with INH resistance (P < 0.005). The strain with one repeat of ETRB is the most drug resistant: 6 out of 10 are INH resistant, versus 16 out of 87 being resistant for strains with two repeats of ETRB (Supplementary Table 1). The corresponding odds ratio (OR) was estimated to be 0.14. Overall, 3, 4, and 3 out of the 15 MIRU loci showed resistance to INH, EMB, PAZ, respectively. No MIRU loci were associated with RFP and SM resistance.
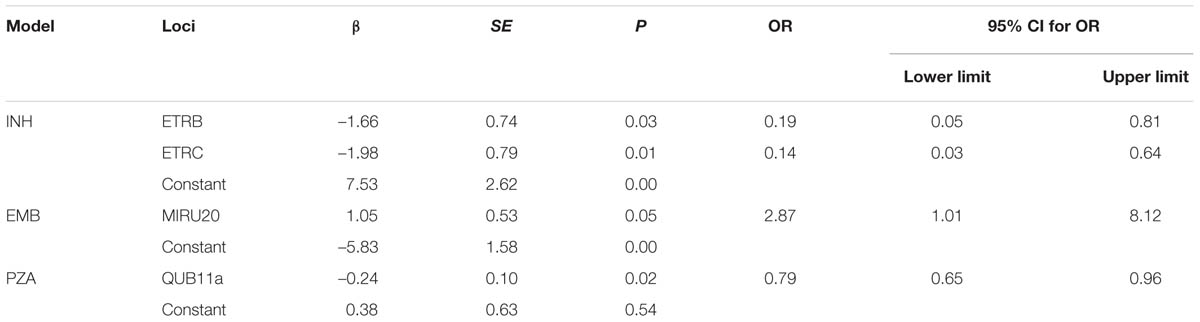
In addition, the ability of the MIRU loci in predicting drug resistance was studied by multivariate logistic regression analysis. Overall, three models were found to provide predictions. For example, with MIRU40, ETRB, and ETRC were the most suitable MIRU loci for the predictive model. We also found other models being able to improve prediction for EMB and PAS resistance (Table 3). The models detected here could potentially aid in future drug resistance prediction.
Discussion
In the present study, we found that ETRB and ETRC were the major predicted factors for INH resistance and QUB11a was the leading predicted factor for PAS resistance. The allelic diversity of QUB11a (0.77) was high but that of ETRB (0.25) and ETRC (0.17) was low.
Although no functions have been found for many MIRU loci, there are evidences that certain MIRU loci can affect nearby gene activities. For example, Yao et al. found that Mtub-39 could affect the promotor activity of the downstream genes, and subsequently regulate the expression of the genes3. Along with our previous results (Wen et al., 2015), the identified resistance-related MIRU in our study provide good hints for generating function-related hypotheses for these MIRU loci, which was briefly described in the following paragraphs. Particularly, MIRU loci might exert functions by regulating the expression of neighboring genes.
For INH resistance, ETRB was immediately downstream of gene qcrB. It has been reported that overexpression of qcrB in Mycobacterium bovis bacillus Calmette–Guérin (BCG) enabled the bacteria to survive at imidazo[1,2-a]pyridines(IP) concentrations that would normally be lethal (Abrahams et al., 2012). Cytochrome b encoded by qcrB was the subunit of bc1 compound which formed a key component in the electron transport chain and included in superoxide generation (Crofts et al., 2008). Previous studies have found that repeated ALU sequences can depress the expression of the reporter gene GFP (Li et al., 2012). We hypothesized that increased copy numbers of ETRB units might inhibit the expression of qcrB, and lead to sensitivity to INH, whose anti-tuberculosis function is associated with the electron transport chain (Herman and Weber, 1980; Lee et al., 2013). ETRC, another protective factor for INH resistance, is located 210 bp upstream of Rv0488.
For EMB resistance, MIRU20 locus is a protective factor. The frequency of MIRU20, located 19 bp upstream of Rv1817, is 2 in H37Rv strain. The function of Rv1817 is still unknown, and is believed to be probably involved in cellular metabolism and metal utilization (Camacho et al., 1999). Interestingly, Zhang et al. (2013) reported that the mutation of Rv1817 may be associated with ofloxacin and kanamycin resistance but not EMB. Thus, we hypothesized that MIRU20 may stimulate the expression of Rv1817, and result in the EMB resistance. However, potential molecular mechanisms about Rv1817 involved in EMB resistance have not been found previously.
QUB11a, a major protective factor for PAS resistance, is included in the gene PPE34. Expression of a recombinant PPE34 protein in Mycobacterium smegmatis showed that PPE34 was likely to be both exposed on the surface of the cell and associated with the cell wall (Lu et al., 2009). PPE34 is polymorphic and related to changes in the numbers of tandem repeat. The high number of copy units of QUB11a may influence the activity of the expression protein of gene PPE34 and then impair the barrier function of cell wall to have M. tuberculosis be resistant to PAS.
In this study, we found that MIRU 40 was related to the increased INH resistance, which was the same as that in our previous results (Wen et al., 2015).
Conclusion
Our research showed that MIRU loci may predict drug resistance of tuberculosis in China. However, the mechanism and exact predictive model still need further exploration.
At last, a number of limitations exist in our study. First, only 15 MIRU loci of M. tuberculosis were selected. This is partially because our work was designed based on the tuberculosis genotyping guideline of Chinese version as the strains were all from China and we thought the initial research method would be better according to the handbook of China version. Second, there is still lack of direct evidence for the mechanisms mentioned above. Further functional study is needed to clarify whether the abnormal expression of neighboring genes can cause resistance. Third, the study may need to include other MIRU loci to investigate the association of MIRU loci with drug resistance in M. tuberculosis.
Author Contributions
X-fC, M. Zhang, M. Zhu, and Y-gJ: acquisition, analysis, and interpretation of data for the work. CJ, DX, and L-lC: drafting the work or revising it critically for important intellectual content. Y-fW: substantial contributions to the conception or design of the work. Final approval of the version to be published.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We owe sincere thanks to Yin Hong, Hui Yang for assistance with drug susceptibility testing. We thank Lingfei Han, Jin Chen for assistance with the PCR testing. This study was reported by the Wannan Medical College University Student Research Fund (WK2014527).
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2016.00378
Footnotes
- ^ http://www.who.int/tb/publications/global_report/en/
- ^ http://www.cdc.gov/tb/topic/laboratory/Drug_testing.htm
- ^ http://www.stmopen.net/research-on-relationship-between-polymorphisms-of-mtub-39-and-expression-of-downstream-genes-of-mycobacterium-tuberculosis/
References
Abrahams, K. A., Cox, J., Spivey, V. L., Loman, N. J., Pallen, M. J., Constantinidou, C., et al. (2012). Identification of novel imidazo [1, 2-a] pyridine inhibitors targeting M. tuberculosis QcrB. PLoS ONE 7:e52951. doi: 10.1371/journal.pone.0052951
Akhtar, P., Singh, S., Bifani, P., Kaur, S., Srivastava, B. S., and Srivastava, R. (2009). Variable-number tandem repeat 3690 polymorphism in Indian clinical isolates of Mycobacterium tuberculosis and its influence on transcription. J. Med. Microbiol. 58, 798–805. doi: 10.1099/jmm.0.002550-0
Biadglegne, F., Tessema, B., Sack, U., and Rodloff, A. C. (2014). Drug resistance of Mycobacterium tuberculosis isolates from tuberculosis lymphadenitis patients in Ethiopia. Indian J. Med. Res. 140, 116–122.
Camacho, L. R., Ensergueix, D., Perez, E., Gicquel, B., and Guilhot, C. (1999). Identification of a virulence gene cluster of Mycobacterium tuberculosis by signature-tagged transposon mutagenesis. Mol. Microbiol. 34, 257–267. doi: 10.1046/j.1365-2958.1999.01593.x
Crofts, A. R., Holland, J. T., Victoria, D., Kolling, D. R., Dikanov, S. A., Gilbreth, R., et al. (2008). The Q-cycle reviewed: how well does a monomeric mechanism of the bc 1 complex account for the function of a dimeric complex? Biochim. Biophys. Acta 1777, 1001–1019. doi: 10.1016/j.bbabio.2008.04.037
Herman, R. P., and Weber, M. M. (1980). Site of action of isoniazid on the electron transport chain and its relationship to nicotinamide adenine dinucleotide regulation in Mycobacterium phlei. Antimicrob. Agents Chemother. 17, 450–454. doi: 10.1128/AAC.17.3.450
Jun, L., Shan, J., Song, Y., and Zongchang, H. (2009). Sequence analysis on drug-resistant rpoB gene of Mycobacterium tuberculosis L-form of isolated from pneumoconiosis workers. J. Med. Colleges PLA 24, 223–227. doi: 10.3892/mmr.2014.1948
Kremer, K., Au, B. K. Y., Yip, P. C. W., Skuce, R., Supply, P., Kam, K. M., et al. (2005). Use of variable-number tandem-repeat typing to differentiate Mycobacterium tuberculosis Beijing family isolates from Hong Kong and comparison with IS6110 restriction fragment length polymorphism typing and spoligotyping. J. Clin. Microbiol. 43, 314–320. doi: 10.1128/JCM.43.1.314-320.2005
Lee, J., Kang, H., Kim, S., Yoo, H., Kim, H. J., and Park, Y. K. (2014). Optimal combination of VNTR typing for discrimination of isolated Mycobacterium tuberculosis in Korea. Tuberc. Respir. Dis. 76, 59–65. doi: 10.4046/trd.2014.76.2.59
Lee, K. K., Fujimoto, K., Zhang, C., Schwall, C. T., Alder, N. N., Pinkert, C. A., et al. (2013). Isoniazid-induced cell death is precipitated by underlying mitochondrial complex I dysfunction in mouse hepatocytes. Free Radic. Biol. Med. 65, 584–594. doi: 10.1016/j.freeradbiomed.2013.07.038
Li, S., Feng, J., Wang, H., Wang, X., and Lv, Z. (2012). [The effects of SV40 PolyA sequence and its AATAAA signal on upstream GFP gene expression and transcription termination]. Yi Chuan 34, 113–119. doi: 10.3724/SP.J.1005.2012.00113
Liu, B., Lu, L., Lü, B., Wan, K., and Yan, Y. (2012). [Meta analysis on the correlation between Mycobacterium tuberculosis Beijing family strains and drug resistance]. Zhonghua Yu Fang Yi Xue Za Zhi 46, 158–164.
Lu, B., Li, Z., Liu, M., Liu, Z., Zhao, X., and Wan, K. (2009). [Discrimination power evaluation for 45 loci of variable number tandem repeats in Mycobacterium tuberculosis strains isolated from China]. Zhonghua Liu Xing Bing Xue Za Zhi 30, 58–62.
Lu, J., Jiang, S., Liu, Q. Y., Ma, S., Li, Y., and Li, C. P. (2014). Analysis of mutational characteristics of the drug-resistant gene katG in multi-drug resistant Mycobacterium tuberculosis L-form among patients with pneumoconiosis complicated with tuberculosis. Mol. Med. Rep. 9, 2031–2035. doi: 10.3892/mmr.2014.2045
Perdigão, J., Macedo, R., Machado, D., Silva, C., Jordão, L., Couto, I., et al. (2014). GidB mutation as a phylogenetic marker for Q1 cluster Mycobacterium tuberculosis isolates and intermediate-level streptomycin resistance determinant in Lisbon, Portugal. Clin. Microbiol. Infect. 20, O278–O284. doi: 10.1111/1469-0691.12392
Pérez-Lago, L., Navarro, Y., Herranz, M., Bouza, E., and García-de-Viedma, D. (2013). Differences in gene expression between clonal variants of Mycobacterium tuberculosis emerging as a result of microevolution. Int. J. Med. Microbiol. 303, 674–677. doi: 10.1016/j.ijmm.2013.09.010
Rodrigues, L., Machado, D., Couto, I., Amaral, L., and Viveiros, M. (2012). Contribution of efflux activity to isoniazid resistance in the Mycobacterium tuberculosis complex. Infect. Genet. Evol. 12, 695–700. doi: 10.1016/j.meegid.2011.08.009
Supply, P., Lesjean, S., Savine, E., Kremer, K., Van Soolingen, D., and Locht, C. (2001). Automated high-throughput genotyping for study of global epidemiology of Mycobacterium tuberculosis based on mycobacterial interspersed repetitive units. J. Clin. Microbiol. 39, 3563–3571. doi: 10.1128/JCM.39.10.3563-3571.2001
Supply, P., Magdalena, J., Himpens, S., and Locht, C. (1997). Identification of novel intergenic repetitive units in a mycobacterial two-component system operon. Mol. Microbiol. 26, 991–1003. doi: 10.1046/j.1365-2958.1997.6361999.x
Tantivitayakul, P., Panapruksachat, S., Billamas, P., and Palittapongarnpim, P. (2010). Variable number of tandem repeat sequences act as regulatory elements in Mycobacterium tuberculosis. Tuberculosis (Edinb.) 90, 311–318. doi: 10.1016/j.tube.2010.08.003
Wen, Y.-F., Jiang, C., Cheng, X.-F., Zhang, Z.-P., Chen, B.-F., and Zhu, Y. (2015). Predictive power of ETRE polymorphism and Katg463 mutation to INH-resistance of M. tuberculosis. Iran. J. Public Health 44,k263–268.
Yu-feng, W., Chao, J., and Xian-feng, C. (2015). Drug-resistant tuberculosis can be predicted by Mycobacterial interspersed repetitive unit locus. Front. Microbiol. 6:147. doi: 10.3389/fmicb.2015.00147
Keywords: Mycobacterium tuberculosis, Mycobacterial Interspersed Repetitive Unit, drug resistance, repeat units, genotyping
Citation: Cheng X-f, Jiang C, Zhang M, Xia D, Chu L-l, Wen Y-f, Zhu M and Jiang Y-g (2016) Mycobacterial Interspersed Repetitive Unit Can Predict Drug Resistance of Mycobacterium tuberculosis in China. Front. Microbiol. 7:378. doi: 10.3389/fmicb.2016.00378
Received: 24 September 2015; Accepted: 08 March 2016;
Published: 23 March 2016.
Edited by:
Tzi Bun Ng, The Chinese University of Hong Kong, ChinaReviewed by:
Noton Kumar Dutta, Johns Hopkins University, USAArnold H. Zea, Louisiana State University Health Sciences Center, USA
Paras Jain, Albert Einstein College of Medicine, USA
Leila Vali, Kuwait University, Kuwait
Copyright © 2016 Cheng, Jiang, Zhang, Xia, Chu, Wen, Zhu and Jiang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yu-feng Wen, d3lmQHdubWMuZWR1LmNu
†These authors have contributed equally to this work and are considered as co-first authors.
 Xian-feng Cheng
Xian-feng Cheng Chao Jiang
Chao Jiang Min Zhang3
Min Zhang3