- 1Department of Clinical Laboratory, Peking Union Medical College Hospital, Chinese Academy of Medical Sciences, Beijing, China
- 2Department of Biotechnology, National Kaohsiung Normal University, Kaohsiung, Taiwan
- 3Department of Graduate Institute of Biomedical Sciences, Chang Gung University, Tao-Yuan, Taiwan
- 4Departments of Laboratory Medicine and Internal Medicine, National Taiwan University Hospital, National Taiwan University College of Medicine, Taipei, Taiwan
Burkholderia pseudomallei is not represented in the current version of Bruker Biotyper matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) system. A total of 66 isolates of B. pseudomallei, including 30 clinical isolates collected from National Taiwan University Hospital (NTUH, n = 27) and Peking Union Medical College Hospital (PUMCH, n = 3), and 36 isolates of genetically confirmed strains, including 13 from clinical samples and 23 from environmental samples, collected from southern Taiwan were included in this study. All these isolates were identified by partial 16S rDNA gene sequencing analysis and the Bruker Biotyper MALDI-TOF MS system. Among the 30 isolates initially identified as B. pseudomallei by conventional identification methods, one was identified as B. cepacia complex (NTUH) and three were identified as B. putida (PUMCH) by partial 16S rDNA gene sequencing analysis and Bruker Biotyper MALDI-TOF MS system. The Bruker Biotyper MALDI-TOF MS system misidentified 62 genetically confirmed B. pseudomallei isolates as B. thailandensis or Burkholderia species (score values, 1.803–2.063) when the currently available database (DB 5627) was used. However, using a newly created MALDI-TOF MS database (including B. pseudomallei NTUH-3 strain), all isolates were correctly identified as B. pseudomallei (score values >2.000, 100%). An additional 60 isolates of genetically confirmed B. cepacia complex and B. putida were also evaluated by the Bruker Biotyper MALDI-TOF MS system using the newly created database and none of these isolates were identified as B. pseudomallei. MALDI-TOF MS is a versatile and robust tool for the rapid identification of B. pseudomallei using the enhanced database.
Introduction
Melioidosis is a tropical and subtropical infectious disease caused by Burkholderia pseudomallei, a Gram-negative, aerobic, motile rod-shaped bacterium that is widely distributed in rice field soil, and in stagnant water throughout the tropics (Hsueh et al., 2001; Currie et al., 2008; Lau et al., 2014). B. pseudomallei is also a major cause of community-acquired septicemia and pneumonia in adults in the Asia-pacific region, particularly in northeast Thailand (Peto et al., 2014). In Taiwan, the first case of melioidosis was reported in 1985 in a man who acquired the disease after aspirating river water in a near-drowning accident in the Philippines (Lee et al., 1985) Since then several sporadic and epidemic cases have been reported (Hsueh et al., 2001; Ko et al., 2007; Su et al., 2011; Chen et al., 2013, 2014). Previous studies clearly showed that the disease was endemic in Taiwan and demonstrated that all clinical isolates were arabinose non-assimilators (Hsueh et al., 2001; Chen et al., 2013). Other studies have reported high concentrations of ambient B. pseudomallei during typhoon season in regions of Taiwan (Ko et al., 2007; Su et al., 2011; Chen et al., 2014). Several typhoon-related melioidosis epidemics have also been reported (Ko et al., 2007; Su et al., 2011). There is also evidence that melioidosis can be transmitted to humans via environmental aerosols contaminated with B. pseudomallei (Chen P. S. et al., 2015).
In Mainland China, melioidosis was first reported in 1990 (Yang, 2000) and is now known to be endemic to several tropical provinces, including Hainan, Guangdong, and Guangxi (Chen H. et al., 2015; Fang et al., 2015; Zheng et al., 2015). Multilocus sequence typing (MLST) disclosed that B. pseudomallei ST562 is dominant in southern China as well as in Australia and Taiwan and that its wide-ranging presence might be due to recent spread caused by transmission between regions (Chen H. et al., 2015). Whole-genome sequencing of B. pseudomallei has been conducted for isolates obtained from patients with melioidosis in Mainland China (strain BPC006) and Taiwan (strain vgh07; Fang et al., 2012; Chen Y. S. et al., 2015).
MALDI-TOF MS is increasingly being used in clinical microbiology laboratories to identify bacterial isolates to the species level and the technique is expected to further accelerate the routine identification of suspicious isolates (Bizzini and Greub, 2010; Inglis et al., 2012; Lau et al., 2012; Niyompanich et al., 2014; Jang et al., 2015; Lasch et al., 2015). Because diseases due to B. pseudomallei are uncommon in North America and Europe, B. pseudomallei are not included (but B. thailandensis was included) in the reference spectra of the Bruker Biotyper and Vitek MS libraries (SARAMIS database) (Jang et al., 2015).
In the present study, we evaluated the ability of the Bruker Biotyper MALDI-TOF MS system to accurately identify genetically confirmed B. pseudomallei that were recovered from patients and environmental sources.
Materials and Methods
Bacterial Isolates
A total of 66 isolates of B. pseudomallei, including 43 clinical isolates and 23 environmental isolates, were collected for study. Among these isolates, 27 were recovered from patients with bloodstream infections who were treated at National Taiwan University Hospital (NTUH, a 2500-bed university-affiliated hospital in northern Taiwan) during the period 1998–2014 (Table 1), and three isolates were recovered from patients treated at Peking Union Medical College Hospital (PUMCH), Beijing, China. The other 36 isolates of B. pseudomallei, which had been identified by sequencing of the 16S rRNA and flagellar genes (Chen et al., 2013), included 13 isolates from various clinical samples obtained from patients treated at several hospitals in southern Taiwan and 23 isolates obtained from different environmental sources during the period 1994–2013 (Table 2; Chen et al., 2013). Cultures and analysis of B. pseudomallei isolates were manipulated in the Mycobacteriology Laboratory at NTUH, a biosafety level 3 laboratory, and followed the biosafety level 3 precaution. Identification of these isolates were initially based on conventional biochemical methods and commercial identification systems, including API (API 20E) and Vitek 2 (ID-GN card) (bioMe'rieux, Marcy l'Etoile, France). Arabinose assimilation testing was performed for all genetically confirmed B. pseudomallei isolates as reported previously (Hsueh et al., 2001).
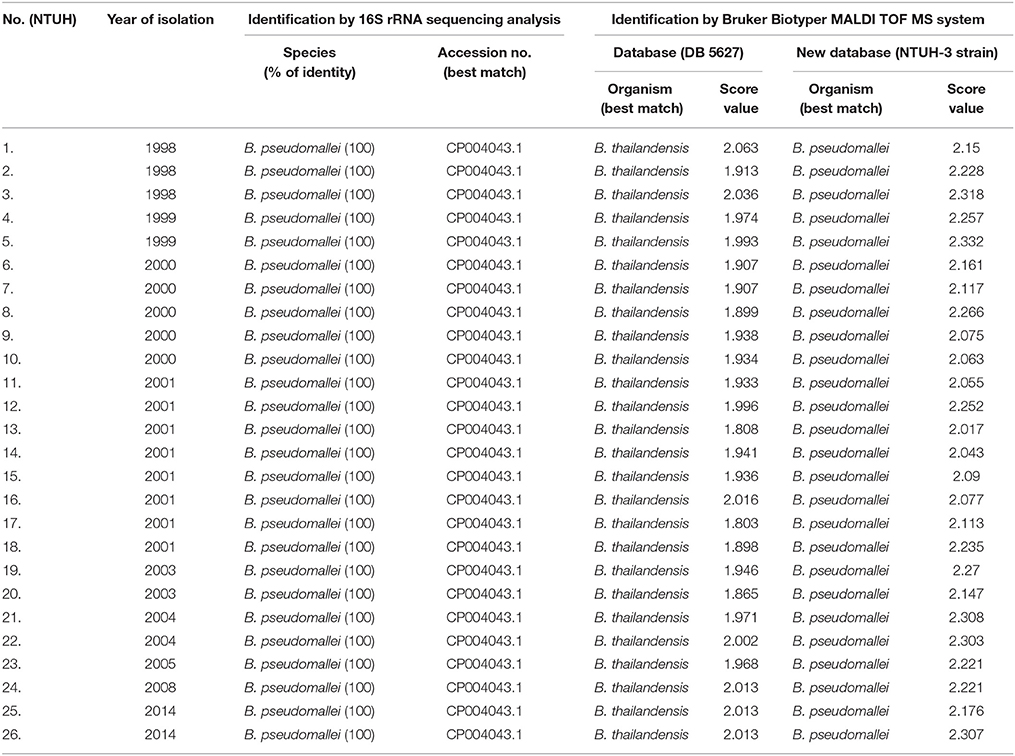
Table 1. Results of 16S rRNA sequencing analysis and Bruker Biotyper MALDI TOF MS for the identification of 26 isolates of B. pseudomallei recovered from patients who were treated at National Taiwan University Hospital (NTUH).
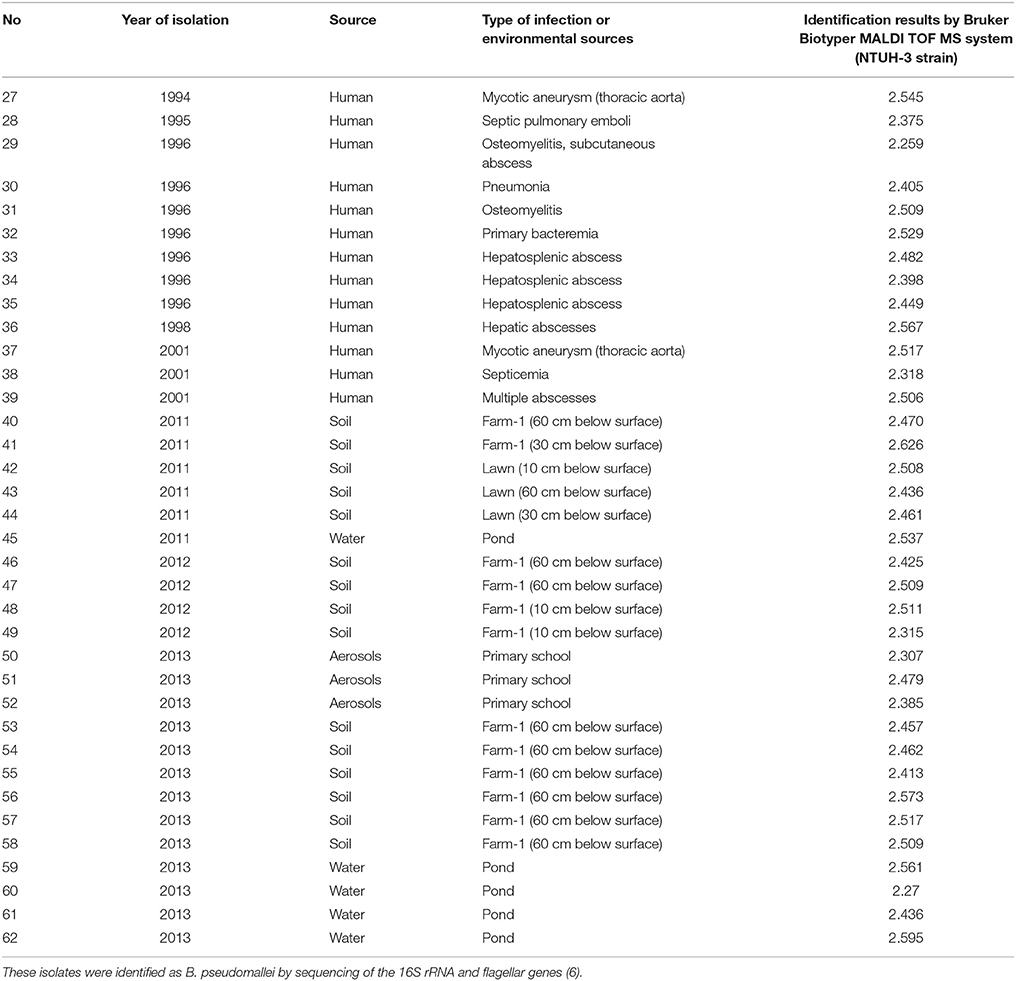
Table 2. Identification of 36 isolates of B. pseudomallei obtained from clinical specimens and environmental sources by Bruker Biotyper MALDI-TOF MS with the NTUH-3 strain.
Identification of Isolates by Gene Sequencing Analysis
Partial 16S rRNA gene sequencing of all 66 isolates was performed using two primers, 8FPL (5′-AGAGTTT GATCCTGGCTCAG-3′) and 1492RPL (5′-GGTTACCTTG TTACGACTT-3′; Cheng et al., 2015). The sequences (1425 bp) obtained were compared with published sequences in the GenBank database using the BLAST algorithm (http://www.ncbi.nlm.nih.gov/blast).
Identification of Isolates by the Bruker Biotyper MALDI-TOF System
Samples of the 66 isolates were prepared for analysis by the Bruker Biotyper MALDI-TOF MS system as previously described (Cheng et al., 2015). B. thailandensis E264 (ATCC700388) was included as the control strain. All isolates were inoculated onto Trypticase soy agar with 5% sheep blood (blood agar plates, BAP; Becton Dickinson Microbiology Systems Sparks, MD, USA) and incubated in 5% CO2 at 37°C for 18 to 24 h. Two to three colonies were transferred to a 1.5-ml screw-cap Eppendorf tube containing 50 μl of 70% formic acid. After incubation for 30 s, 50 μl of acetonitrile (Sigma-Aldrich) was added. The suspension was centrifuged at 13,000 rpm for 2 min, and then 1.0 μl of the supernatant was applied to a 96-spot polished steel target plate (Bruker Daltonik GmbH) and dried. A saturated solution of 1.0 μl of MALDI matrix (alphacyano-4-hydroxycinnamic acid matrix solution; Bruker Daltonik GmbH) was applied to each sample and dried. Measurements were performed with the Bruker Biotyper MALDI-TOF MS system using FlexControl software with Compass Flex Series version 1.3 software and a 60-Hz nitrogen laser (337 nm wavelength). Spectra ranging from 2000 to 20,000 m/z were analyzed using the MALDI Biotyper system's automation control and the current Bruker Biotyper 3.1 software and library (database [DB] 5627 with 5627 entries). Identification scores of ≥2.000 indicated species-level identification, scores of 1.700 to 1.999 indicated genus-level identification, and scores of < 1.700 indicated no reliable identification. B. pseudomallei is not listed in the current Bruker Biotyper MALDI-TOF MS database.
Cluster Analysis and Main Spectra Projection by the Bruker Biotyper MALDI TOF System
A clustering analysis of 26 isolates collected from NTUH was performed using ClinProTools 3.0 (Bruker Daltonics GmbH, Bremen, Germany; Cheng et al., 2015). Dendrograms from the MALDI Biotyper data of 26 genetically well-characterized B. pseudomallei isolates were obtained and cluster groups with a default critical distance level of 850 were identified. Isolates from different cluster groups were selected for main spectra projection (MSP; database entrance) creation using MALDI Biotyper software (Bruker Daltonics). The database generated using the isolates selected from different cluster groups was blindly tested against the 26 B. pseudomallei isolates from NTUH.
External validation was performed for another 36 B. pseudomallei isolates collected from southern Taiwan, 30 isolates of B. putida collected from NTUH (n = 10) and PUMCH (n = 20), and 30 isolates of B. cepacia complex (from NTUH). Principal component analysis (PCA) dendrograms were generated from MALDI-TOF MS Biotyper mass spectra of 62 isolates of B. pseudomallei, including 42 clinical isolates and 20 environmental isolates.
Results
Among the 30 clinical isolates collected from clinical microbiology laboratories at the NTUH and PUMCH that were initially identified as B. pseudomallei (all >90% probability) by the Vietk 2 ID-GN card, four (one for NTUH and three from PUMCH) were identified as non-B. pseudomallei by partial 16S rDNA sequencing analysis. Among these four isolates, one was identified as B. cepacia complex (NTUH) and three were identified as B. pudita (PUMCH) by partial 16S rDNA sequencing analysis.
The Bruker Biotyper MALDI-TOF MS system misidentified all the 62 isolates of genetically confirmed B. pseudomallei as B. thailandensis (seven isolates with a score value ≥2.000) or Burkholderia species (19 with score values ranging from 1.803 to 1.996). However, arabinose assimilation testing showed that all 62 isolates were arabinose non-assimilators, indicating that these isolates were not B. thailandensis. Clustering analysis of the 26 genetically confirmed isolates of B. pseudomallei collected from NTUH by the Bruker Biotyper MALDI-TOF MS system identified five cluster groups (Figure 1).
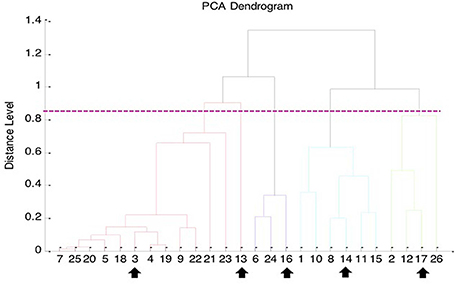
Figure 1. Principal component analysis (PCA) dendrogram generated from Bruker Biotyper MALDI-TOF MS mass spectra of 26 clinical isolates of B. pseudomallei collected from NTUH. The five arrows indicate the MALDI spectra of the five B. pseudomallei isolates selected for MSP (main spectra projection; database entrance) creation using MALDI Biotyper software (Bruker Daltonics).
Because B. pseudomallei was not listed in the current Bruker Biotyper MALDI-TOF MS database (DB5627), spectra of five isolates, namely NTUH-3, NTUH-13, NTUH-14, NTUH-16, and NTUH-17, were selected from each cluster group for MSP (database entrance). The database generated using the five isolates was blindly tested against the spectra of the remaining 57 isolates (21 from NTUH and 36 from southern Taiwan). The best identification scores were found according to the database created using the NTUH-3 strain; all (100%) 57 B. pseudomallei isolates were correctly identified as B. pseudomallei with identification scores of ≥2.000 (2.017 to 2.626). B. thailandensis E264 (ATCC700388) was identified by the Bruker Biotyper MALDI-TOF MS system based on Bruker Biotyper 3.1 software and library (DB5627) plus new database created by NTUH-3 strain (enhanced database) as B. thailandensis (best match, score value of 2.170) or B. pseudomallei (second match, score value of 2.058). Among the four isolates initially mis-identified as B. pseudomallei, one was identified as B. cepacia complex (NTUH; score value of 2.137) and three were identified as B. pudita (PUMCH; 2.012, 2.102, and 2.223, respectively) by standard and enhanced Bruker Biotyper MALDI-TPOF MS database. The characteristic spectra of NTUH-3 strain and B. thailandensis E264 (ATCC700388) are shown in Figure 2.
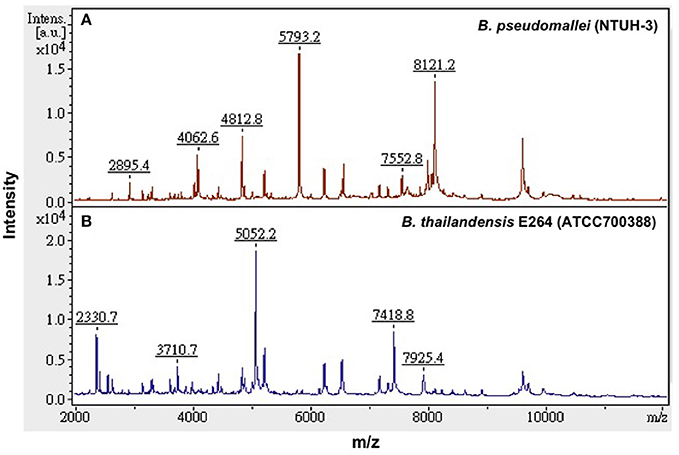
Figure 2. Characteristic MALDI-TOF MS spectra. Characteristic spectra of (A) B. pseudomallei (NTUH-3 strain) and (B) B. thailandensis E264 (ATCC 700388) generated by the Bruker Biotyper MALDI-TOF MS system. The absolute intensities of the ions are shown on the y axis, and the masses (m/z) of the ions are shown on the x axis. The m/z values represent the mass-to-charge ratios.
The PCA dendrogram generated from MALDI Biotyper mass spectra of 62 isolates of B. pseudomallei, including 42 clinical isolates and 20 environmental isolates collected from northern and southern Taiwan is illustrated in Figure 3. There were five cluster groups with dividing branches linked at a distance level of 850.
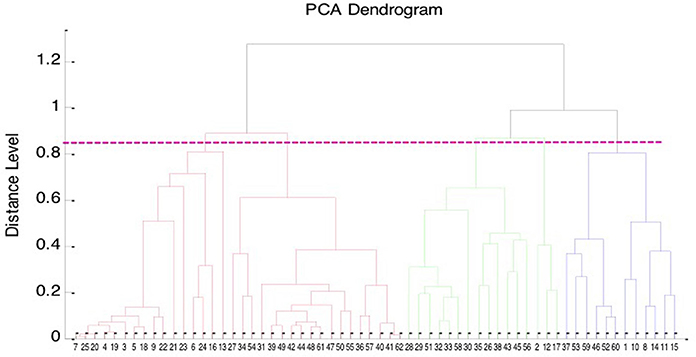
Figure 3. Principal component analysis (PCA) dendrogram of B. pseudomallei. PCA dendrogram generated from Bruker Biotyper MALDI TOF MS mass spectra of the 62 isolates of B. pseudomallei, including 42 clinical isolates, and 20 environmental isolates collected from northern and southern Taiwan.
Among the 30 isolates of genotypically characterized B. putida and 30 isolates of B. cepacia complex, all were identified as B. putida (score values of >2.000) and B. cepacia complex (26 B. cenocepacia and four B. cepacia; score values of >2.000), respectively, using the pre-existing database and that created by NTUH-3 strain.
Discussion
This study revealed several important findings. First, the Bruker Biotyper MALDI-TOF MS system failed to correctly identify clinical and environmental isolates of B. pseudomallei because B. pseudomallei is not listed in the current version of FDA-cleared Bruker library. Although there is a “security-relevant library” which contains B. pseudomallei, this library is not available in most clinical microbiological laboratories. Second, when we included the B. pseudomallei NTUH-3 strain only into the current library, all isolates of B. pseudomallei could be identified B. pseudomallei by the Bruker Biotyper MALDI-TOF MS system with a correct identification rate of 100%. Third, the control strain of B. thailandensis was still identified as B. thailandensis as the best match organism when the enhanced database was used. Finally, all isolates obtained from environmental sources were confirmed to be B. pseudomallei not B. thailandensis.
Identification of B. pseudomallei poses difficulties in clinical microbiology laboratories. Although, B. pseudomallei is included in the API 20NE, the Vitek 1, and the Vitek 2 databases, the accuracy of identification by these systems varies (Lau et al., 2015). Misidentification of B. pseudomallei as other Burkholderia species such as B. cepacia complex and B. putida as well as Pseudomonas aeruginosa is common using these commercial systems (around 20%; Lowe et al., 2006; Deepak et al., 2008; Zong et al., 2012; Lau et al., 2015). Furthermore, commercial bacterial identification kits might fail to differentiate between B. pseudomallei and a closely related species such as B. thailandensis, although >99% of cases of melioidosis are caused by B. pseudomallei (Lau et al., 2015). The issue of misidentification of B. pseudomallei is of importance for patient care as well as laboratory safety.
Traditionally, B. pseudomallei can be distinguished from B. thailandensis by arabinose assimilation (Lau et al., 2015). Genotypic differentiation between B. pseudomallei and B. thailandensis can be achieved by specific PCR-based identification using B. pseudomallei-specific gene targets, such as the Type III secretion system and Tat-domain protein and sequencing of gene targets of 16S rRNA and groEL (Lau et al., 2015).
MALDI-TOF MS, a revolutionary technique for pathogen identification, has been shown to be a potentially useful tool for rapid identification of B. pseudomallei, although existing databases require optimization by adding reference spectra for B. pseudomallei (Inglis et al., 2012; Lau et al., 2012; Niyompanich et al., 2014; Lasch et al., 2015). Jang et al. reported a case of multifocal aneurysms in the aortic arch of the thoracic aorta and pseudoaneurysm in the abdominal aorta and the inferior area of the superior mesenteric artery caused by B. pseudomallei (Jang et al., 2015). Colonies from positive blood and tissue were compatible with the presumptive identification of B. pseudomallei. However, using the Vitek 2 system, the blood isolate was identified as B. pseudomallei and the tissue isolate was identified as B. cepacia complex. The blood isolate was identified as B. thailandensis with a score value of 1.901 by the Bruker Biotyper MALDI-TOF MS system (Jang et al., 2015). Lau et al. used the Bruker database extended with B. pseudomallei reference strains, three B. thailandensis isolates were misidentified as B. pseudomallei. In this study, the reference strain of B. thailandensis was correctly identified with enhanced Bruker database. More isolates of B. thailandensis isolates are needed for verify the accuracy of this new database.
Recently, bioMerieux recognized misidentification of B. pseudomallei as an important issue and addressed the problem by altering the algorithm parameters. The new parameters are included in the most recent software release (version 4.03 for Vitek 2 60/XL and version 2.01 for Vitek 2 Compact; Lowe et al., 2006).
Interestingly, in this study all four isolates initially identified as B. pseudomallei by the Vitek 2 ID-GN card were identified as B. cepacia complex or B. pudita by 16S rRNA sequencing analysis. In areas with high endemicity of melioidoisis like Taiwan, laboratory staff tends to report the results of B. pseudomallei isolation without further clarification. In contrast, most clinical microbiologists in Beijing often recheck the identification of B. pseudomallei because of the low incidence of melioidoisis in northern China (Yang, 2000; Currie et al., 2008; Fang et al., 2015; Zheng et al., 2015).
Summary
In areas where this organism is endemic, such as in South Asia and Northern Australia, identification of B. pseudomallei is not usually problematic (Currie et al., 2008; Lau et al., 2015). With increased international travel and threats of bioterrorism, recognition, and accurate identification of these organisms is important (Lau et al., 2015). The use of automated identification systems, including MALDI-TOF MS, in the clinical microbiology laboratory is becoming common as the pressure of cost containment impacts staff resources. In this study, using our newly created database, all B. pseudomallei isolates were correctly identified to the species level using the Bruker Biotyper MALDI-TOF MS system. These findings suggest that MALDI-TOF MS is a versatile and robust tool for the rapid identification of B. pseudomallei isolates. Expansion of commercially available databases with pathogens endemic in different regions is crucial to improve the usefulness of MALDI-TOF MS. However, this successful application of MALDI-TOF can only be regarded as pilot study, due to the small sample size, which needs independent validation before it can be offered as routine technique in the clinic.
Author Contributions
HW, PH, and YX conceived and designed the experiments, performed the experiments, analyzed the data, and wrote the paper. YC, ST, ZX performed the experiments and analyzed the data. HW, YC, ST, ZX, PH, YX read and approved the final version of the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This study was conducted at Taoyuan General Hospital and four privately run respiratory care wards in Northern Taiwan.
References
Bizzini, A., and Greub, G. (2010). Matrix-assisted laser desorption ionization time-of-flight mass spectrometry, a revolution in clinical microbial identification. Clin. Microbiol. Infect. 16, 1614–1619. doi: 10.1111/j.1469-0691.2010.03311.x
Chen, H., Xia, L., Zhu, X., Li, W., Du, X., Wu, D., et al. (2015). Burkholderia pseudomallei sequence type 562 in China and Australia. Emerg. Infect. Dis. 21, 166–168. doi: 10.3201/eid2101.140156
Chen, P. S., Chen, Y. S., Lin, H. H., Liu, P. J., Ni, W. F., Hsueh, P. T., et al. (2015). Airborne transmission of melioidosis to humans from environmental aerosols contaminated with B. pseudomallei. PLoS. Negl. Trop. Dis. 9:e0003834. doi: 10.1371/journal.pntd.0003834
Chen, Y. L., Lin, Y. C., Chen, Y. S., Chen, S. C., Liu, Y. M., and Tseng, I. L. (2013). Characterisation of predominant molecular patterns of Burkholderia pseudomallei in Taiwan. Trans. R. Soc. Trop. Med. Hyg. 107, 165–169. doi: 10.1093/trstmh/trs093
Chen, Y. L., Yen, Y. C., Yang, C. Y., Lee, M. S., Ho, C. K., Mena, K. D., et al. (2014). The concentrations of ambient Burkholderia pseudomallei during typhoon season in endemic area of melioidosis in Taiwan. PLoS Negl. Trop. Dis. 8:e2877. doi: 10.1371/journal.pntd.0002877
Chen, Y. S., Lin, H. H., Hsueh, P. T., Liu, P. J., Ni, W. F., Chung, W. C., et al. (2015). Whole-genome sequence of an epidemic strain of Burkholderia pseudomallei vgh07 in Taiwan. Genome Announc. 3, e00345–e00315. doi: 10.1128/genomeA.00345-15
Cheng, W. C., Jan, I. S., Chen, J. M., Teng, S. H., Teng, L. J., Sheng, W. H., et al. (2015). Evaluation of the Bruker Biotyper matrix-assisted laser desorption ionization-time of flight mass spectrometry system for identification of blood isolates of Vibrio species. J. Clin. Microbiol. 53, 1741–1744. doi: 10.1128/JCM.00105-15
Currie, B. J., Dance, D. A., and Cheng, A. C. (2008). The global distribution of Burkholderia pseudomallei and melioidosis: an update. Trans. R. Soc. Trop. Med. Hyg. 102, S1–S4. doi: 10.1016/s0035-9203(08)70002-6
Deepak, R. N., Crawley, B., and Phang, E. (2008). Burkholderia pseudomallei identification: a comparison between the API 20NE and VITEK2 GN systems. Trans. R. Soc. Trop. Med. Hyg. 102, S42–S44. doi: 10.1016/s0035-9203(08)70012-9
Fang, Y., Chen, H., Li, Y. L., Li, Q., Ye, Z. J., and Mao, X. H. (2015). Melioidosis in Hainan, China: a restrospective study. Trans. R. Soc. Trop. Med. Hyg. 109, 636–642. doi: 10.1093/trstmh/trv065
Fang, Y., Huang, Y., Li, Q., Chen, H., Yao, Z., Pan, J., et al. (2012). First genome sequence of a Burkholderia pseudomallei Isolate in China, strain BPC006, obtained from a melioidosis patient in Hainan. J. Bacteriol. 194, 6604–6605. doi: 10.1128/JB.01577-12
Hsueh, P. R., Teng, L. J., Lee, L. N., Yu, C. J., Yang, P. C., Ho, S. W., et al. (2001). Melioidosis: an emerging infection in Taiwan? Emerg. Infect. Dis. 7, 428–433. doi: 10.3201/eid0703.017310
Inglis, T. J., Healy, P. E., Fremlin, L. J., and Golledge, C. L. (2012). Use of matrix-assisted laser desorption/ionization time-of-flight mass spectrometry analysis for rapid confirmation of Burkholderia pseudomallei in septicemic melioidosis. Am. J. Trop. Med. Hyg. 86, 1039–1042. doi: 10.4269/ajtmh.2012.11-0454
Jang, H. R., Lee, C. W., Ok, S. J., Kim, M. J., Bae, M. J., Song, S., et al. (2015). Melioidosis presenting as a mycotic aneurysm in a Korean patient, diagnosed by 16S rRNA sequencing and matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Int. J. Infect. Dis. 38, 62–64. doi: 10.1016/j.ijid.2015.07.012
Ko, W. C., Cheung, B. M., Tang, H. J., Shih, H. I., Lau, Y. J., Wang, L. R., et al. (2007). Melioidosis outbreak after typhoon, southern Taiwan. Emerg. Infect. Dis. 13, 896–898. doi: 10.3201/eid1306.060646
Lasch, P., Wahab, T., Weil, S., Pályi, B., Tomaso, H., Zange, S., et al. (2015). Identification of highly pathogenic microorganisms by matrix-assisted laser desorption ionization-time of flight mass spectrometry: results of an interlaboratory ring trial. J. Clin. Microbiol. 53, 2632–2640. doi: 10.1128/JCM.00813-15
Lau, S. K., Chan, S. Y., Curreem, S. O., Hui, S. W., Lau, C. C., Lee, P., et al. (2014). Burkholderia pseudomallei in soil samples from an oceanarium in Hong Kong detected using a sensitive PCR assay. Emerg. Microbes Infect. 3:e69. doi: 10.1038/emi.2014.69
Lau, S. K., Sridhar, S., Ho, C. C., Chow, W. N., Lee, K. C., Lam, C. W., et al. (2015). Laboratory diagnosis of melioidosis: past, present and future. Exp. Biol. Med. (Maywood) 240, 742–751. doi: 10.1177/1535370215583801
Lau, S. K., Tang, B. S., Curreem, S. O., Chan, T. M., Martelli, P., Tse, C. W., et al. (2012). Matrix-assisted laser desorption ionization-time of flight mass spectrometry for rapid identification of Burkholderia pseudomallei: importance of expanding databases with pathogens endemic to different localities. J. Clin. Microbiol. 50, 3142–3143. doi: 10.1128/JCM.01349-12
Lee, N., Wu, J. L., Lee, C. H., and Tsai, W. C. (1985). Pseudomonas pseudomallei infection from drowning: the first reported case in Taiwan. J. Clin. Microbiol. 22, 352–354.
Lowe, P., Haswel, H., and Lewis, K. (2006). Use of various common isolation media to evaluate the new VITEK 2 colorimetric GN Card for identification of Burkholderia pseudomallei. J. Clin. Microbiol. 44, 854–856. doi: 10.1128/JCM.44.3.854-856.2006
Niyompanich, S., Jaresitthikunchai, J., Srisanga, K., Roytrakul, S., and Tungpradabkul, S. (2014). Source-identifying biomarker ions between environmental and clinical Burkholderia pseudomallei using whole-cell matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS). PLoS ONE. 9:e99160. doi: 10.1371/journal.pone.0099160
Peto, L., Nadjm, B., Horby, P., Ngan, T. T., van Doorn, R., Van Kinh, N., et al. (2014). The bacterial aetiology of adult community-acquired pneumonia in Asia: a systematic review. Trans. R. Soc. Trop. Med. Hyg. 108, 326–337. doi: 10.1093/trstmh/tru058
Su, H. P., Chan, T. C., and Chang, C. C. (2011). Typhoon-related leptospirosis and melioidosis, Taiwan, 2009. Emerg. Infect. Dis. 17, 1322–1324. doi: 10.3201/eid1707.101050
Yang, S. (2000). Melioidosis research in China. Acta Trop. 77, 157–165. doi: 10.1016/S0001-706X(00)00139-X
Zheng, X., Chen, H., Zhu, X., Cai, H., Li, L., Li, H., et al. (2015). Discovery of Burkholderia thailandensis isolated from melioidosis endemic areas of Hainan China. Zhonghua. Liu. Xing. Bing. Xue. Za. Zhi. 36, 97–98. doi: 10.3760/cma.j.issn.0254-6450.2015.01.023
Keywords: Burkholderia pseudomallei, B. thailandensis, matrix-assisted, laser desorption/ionization time-of-flight mass spectrometry, 16S rDNA gene sequencing analysis, enhanced database
Citation: Wang H, Chen Y-L, Teng S-H, Xu Z-P, Xu Y-C and Hsueh P-R (2016) Evaluation of the Bruker Biotyper Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry System for Identification of Clinical and Environmental Isolates of Burkholderia pseudomallei. Front. Microbiol. 7:415. doi: 10.3389/fmicb.2016.00415
Received: 06 December 2015; Accepted: 14 March 2016;
Published: 08 April 2016.
Edited by:
Fabrice Merien, Auckland University of Technology, New ZealandReviewed by:
Tamas Szakmany, Cardiff University, UKOnya Opota, Lausanne University Hospital, Switzerland
Copyright © 2016 Wang, Chen, Teng, Xu, Xu and Hsueh. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ying-Chun Xu, eHljcHVtY2hAMTM5LmNvbQ==;
Po-Ren Hsueh, aHNwb3JlbkBudHUuZWR1LnR3
 He Wang1
He Wang1 Po-Ren Hsueh
Po-Ren Hsueh