- 1Department of Zoology, College of Science, King Saud University, Riyadh, Saudi Arabia
- 2Department of Zoology and Entomology, Faculty of Science, Helwan University, Cairo, Egypt
- 3Department of Zoology, Faculty of Science, Beni-Suef University, Beni-Suef, Egypt
- 4Boehringer-Ingelheim Pharma, Biberach, Germany
- 5Department of Biology, Heinrich-Heine-University, Duesseldorf, Germany
Protective vaccination induces self-healing of otherwise fatal blood-stage malaria of Plasmodium chabaudi in female Balb/c mice. To trace processes critically involved in self-healing, the liver, an effector against blood-stage malaria, is analyzed for possible changes of its transcriptome in vaccination-protected in comparison to non-protected mice toward the end of the crisis phase. Gene expression microarray analyses reveal that vaccination does not affect constitutive expression of mRNA and lincRNA. However, malaria induces significant (p < 0.01) differences in hepatic gene and lincRNA expression in vaccination-protected vs. non-vaccinated mice toward the end of crisis phase. In vaccination-protected mice, infections induce up-regulations of 276 genes and 40 lincRNAs and down-regulations of 200 genes and 43 lincRNAs, respectively, by >3-fold as compared to the corresponding constitutive expressions. Massive up-regulations, partly by >100-fold, are found for genes as RhD, Add2, Ank1, Ermap, and Slc4a, which encode proteins of erythrocytic surface membranes, and as Gata1 and Gfi1b, which encode transcription factors involved in erythrocytic development. Also, Cldn13 previously predicted to be expressed on erythroblast surfaces is up-regulated by >200-fold, though claudins are known as main constituents of tight junctions acting as paracellular barriers between epithelial cells. Other genes are up-regulated by <100- and >10-fold, which can be subgrouped in genes encoding proteins known to be involved in mitosis, in cell cycle regulation, and in DNA repair. Our data suggest that protective vaccination enables the liver to respond to P. chabaudi infections with accelerated regeneration and extramedullary erythropoiesis during crisis, which contributes to survival of otherwise lethal blood-stage malaria.
Introduction
Malaria has caused worldwide about 214 million new cases and about 438,000 deaths in 2015 (WHO, 2015). The liver plays an important role in malaria. It is the organ of intra-hepatocyte multiplication of the pre-erythrocytic stages of the malaria-causing agent, parasitic protozoans of the genus Plasmodium (Bertolino and Bowen, 2015; Frevert and Krzych, 2015; Frevert et al., 2015). Moreover, the liver responds to the morbidity- and mortality-causing blood-stages of the parasites. The liver is pathologically changed and even massively injured in patients suffering from blood-stage malaria (Ananad et al., 1992; Kochar et al., 2003; Nautyal et al., 2005; Rupani and Amarapurkar, 2009). Also, the liver with its inherent immune system even functions as an effector against blood-stage malaria (Dockrell et al., 1980; Abo and Sekikawa, 2002; Wunderlich et al., 2005, 2014). The latter view is supported by studies performed with animal models, as e.g., Plasmodium chabaudi in mice. This model shares several characteristics with P. falciparum, the most dangerous malaria species for humans (Longley et al., 2011; Stephens et al., 2012). Mice are either susceptible or resistant to P. chabaudi malaria, i.e., blood-stage infections take either a lethal or a self-healing outcome. This outcome is under complex control involving genes of the mouse-MHC, the H-2 complex, and genes of the non-H-2 background as well as environmental factors (Wunderlich et al., 1988b, 2014). Both lethal and self-healing infections take a similar course of peripheral parasitemia during the pre-crisis phase. For instance, when mice are challenged with 106 P. chabaudi-infected erythrocytes, precrisis culminates at peak parasitemia of about 40–50% on approximately day 8 p.i. The following crisis phase, characterized by dramatically declining peripheral parasitemia to about 2–1% within about 3–4 days, is apparently critical for the outcome of disease. Indeed, all susceptible mice succumb during crisis, whereas resistant mice will survive and generate long-lasting immune mechanisms against homologous re-challenge (Wunderlich and Helwig, 1987).
An effective anti-malaria vaccine for humans is currently not yet available. However, there are promising candidate vaccines in progress (Halbroth and Draper, 2015; Hoffman et al., 2015; Miura, 2016). To date, only RTS,S/AS01 vaccine has completed a phase 3 evaluation (White et al., 2015). Although RTS,S/AS01 received a positive regulatory assessment by the European Medicine Agency for its active immunization of children aged 6 weeks to 17 months against malaria it was not recommended by the WHO (Birkett, 2016; Gosling and von Seidlein, 2016). Moreover, the protective mechanisms of the host defense, which have to be activated by a vaccine, appear to be not yet fully understood despite enormous advances in our knowledge during the past 35 years. At least, however, those protective mechanisms directed against blood-stage malaria can be also conveniently investigated in the P. chabaudi model. Indeed, there has been previously developed a vaccination procedure which raises survival of malaria-susceptible mice from 0 to over 80% (Wunderlich et al., 1988a; Krücken et al., 2009). This vaccination, using non-infectious surface membranes isolated from P. chabaudi-infected erythrocytes, does not induce sterile immunity, but converts lethal blood-stage infections to take a self-healing course. Moreover, there is information available, though still little, that protective vaccination also affects the liver (Krücken et al., 2009; Wunderlich et al., 2014). This becomes evident especially during the crisis phase, when vaccination leads to a dramatic increase in the hepatic capacity to trap injected particles (Krücken et al., 2009).
The processes associated with protective vaccination in the liver during crisis are only poorly investigated to date. To trace these processes, it would be initially meaningful to identify those genes whose expression is changed in the liver of vaccination-induced self-healing infections toward the end of the crisis phase. This would in turn allow deductions for processes possibly involved in the liver to mediate protection against blood-stage malaria. We have therefore decided to analyze the effect of protective vaccination on the transcriptome of the liver toward the end of the crisis phase in vaccination-induced self-healing infections of P. chabaudi in comparison to lethal infections in non-protected Balb/c mice using Agilent's gene expression microarrays.
Materials and Methods
Mice
This study continues our previous work performed with female Balb/c mice (Krücken et al., 2009). It was carried out in strict accordance with the German law on animal protection. The keeping of mice as well as the experimental protocol of the study were officially approved by the State-controlled Committee on the Ethics of Animal Experiments of the State Nordrhein-Westfalen, Germany, and were regularly controlled, without being previously announced, by the local authorities. All efforts were made to minimize suffering. Female Balb/c mice were delivered at an age of 10–12 weeks from our central animal facilities at the University of Düsseldorf, where they were bred under specified pathogen-free conditions. During the experiments, the mice were housed in plastic cages, received a standard diet (Woehrlin, Bad Salzuflen, Germany) and water ad libitum.
Blood-Stage Malaria
Blood-stage infections of P. chabaudi were routinely maintained in outbred mice under sterile conditions by weekly passages of infected blood. Since 1982, a non-clonal line of P. chabaudi is continuously used in our laboratory (Wunderlich et al., 1982) resembling to P. chabaudi as described previously (Wunderlich et al., 2005; Krücken et al., 2009). Balb/c mice were challenged i.p. with 106 P. chabaudi chabaudi AS-infected erythrocytes. Parasitemia was evaluated in Giemsa-stained smears from tail blood. Erythrocytes were counted in a Neubauer chamber.
Protective Vaccination
Mice were vaccinated as detailed previously (Krücken et al., 2009). The vaccine contained host cell plasma membranes isolated in the form of ghosts from P. chabaudi-parasitized red blood cells as detailed previously (Wunderlich et al., 1985, 1987). These membranes were previously characterized to be non-infectious and to be associated with parasite-synthesized proteins (Wunderlich et al., 1988c,d) as it was also reported for surface membranes of P. falciparum-infected erythrocytes (Fontaine et al., 2012). Approximately 106 ghosts were suspended in 100 μl Freund's complete adjuvant (FCA) and subcutaneously injected 3 and 1 week before infecting with P. chabaudi-parasitized erythrocytes. Controls received only FCA.
RNA Isolation
Livers were aseptically removed from sacrificed mice, rapidly frozen in liquid nitrogen and stored at −80°C until use. Frozen livers were grounded in a mortar under liquid nirogen and aliquots of each liver were used to isolate total RNA by the Trizol (Qiagen, Hilden, Germany) standard RNA extraction protocol. Trizol-extracted RNA was additionally cleaned up using the miRNeasy Kit (Qiagen). Integrity and quality of RNA was checked with the Agilent 2100 Bioanalyzer platform (Agilent Technologies). All RNA samples revealed RIN values between 8.7 and 9.1.
Cy3-Labeling of RNA
Equivalents of 100 ng from each RNA sample were used for the linear T7-based amplification step. To produce Cy3-labeled cRNA, the RNA samples were amplified and labeled with the Agilent Low Input Quick Amp Labeling Kit (Agilent Technologies) according to the manufacturer's instructions. Yields of cRNA and dye-incorporation were determined with the ND-1000 Spectrophotometer (NanoDrop Technologies). The incorporations were between 18 and 23 fmol Cy3/ng cRNA.
Hybridization of Agilent Mouse Whole Genome Oligo Microarrays
Hybridization was performed with Agilent's 8 × 60 K oligo microarrays (design 028005), which contain 8 arrays per slide with each array displaying 39,430 Entrez Gene RNAs and 16,251 lincRNAs. Using the Agilent Gene Expression Hybridization Kit, hybridization was carried out as detailed in the Agilent processing protocol (Agilent technologies). In brief, 600 ng Cy3-labeled fragmented cRNA in hybridization buffer was hybridized overnight at 65°C to the microarrays using Agilent's recommended hybridization chamber and oven. Finally, the microarrays were washed with the Agilent Gene Expression Wash Buffer 1 for 1 min at room temperature, which was followed by washing with preheated Agilent Gene Expression Wash Buffer 2 at 37°C for 1 min.
Scanning and Data Analysis
The Agilent's Microarray Scanner System (Agilent Technologies) was used to detect fluorescence signals on the hybridized microarrays. The microarray image files were read out and processed with the Agilent Feature Extraction Software. The latter determines feature intensities (including background substraction), rejects outliers and calculates statistical confidences. A heatmap was generated to visualize the expression levels of each mRNA and lincRNA (Spotfire, TIBCO Software Inc., Palo Alto, USA). The row and column dendrograms were clustered with the unweighted pair group method with arithmetic mean and Euclidean distance measure. Data are publicly available at the EMBL-EBI Array Express repository (Array Express accession number: E-MTAB-4791).
Quantitative Real-Time PCR
Using High Capacity cDNA Reverse Transcription Kit (Life Technologies) and TaqMan mRNA assays (Life Technologies) we performed reverse transcription of mRNAs coding for the following proteins: RHD (assay ID: Mm00456910_m1), ERMAP (Mm_00469273_m1), ADD2 (Mm00478923_m1), ANK1 (Mm00482889_m1), PLK1 (Mm00440924_g1), CCNA2 (Mm00438063_m1), LIFR (Mm00442942_m1), and CD163 (Mm_00474091_m1). PCR reactions were performed with the TaqMan® gene expression master mix (Life Technologies) according manufacturer's protocol on a 7900HT real-time PCR System, as described previously (Al-Quraishy et al., 2014). All samples were run in duplicates and raw ct values were calculated using the SDS software v.2.4 and GAPDH was used for normalization. Fold change of expression was calculated with the comparative Ct method (2−ΔΔct) (Livak and Schmittgen, 2002). Data sets were analyzed for statistical significance using two-tailed unpaired heteroskedastic Student's t-test (*p < 0.01).
Results
Identification of Differentially Expressed Genes in the Liver of Vaccination-Protected vs. Non-protected Mice
To identify differentially expressed genes in the liver of vaccination-protected mice, in comparison with non-vaccinated malaria-susceptible mice, toward the end of the crisis phase of blood-stage infections of P. chabaudi, we have analyzed gene expression microarrays from livers individually prepared from three vaccination-protected mice on day 11 p.i. (Vd11) and from 3 vaccinated mice before infection on day 0 p.i. (Vd0) as well as from 3 non-vaccinated mice on day 0 p.i. (Nd0) and on day 11 p.i. (Nd11). The course of parasitemias of P. chabaudi infections in vaccinated and non-vaccinated mice has been previously determined under identical experimental conditions as used here (Krücken et al., 2009). The parasitemia was 0% at Vd0 and Nd0 and approximately 2% at Vd11 and Nd11, respectively. Then, we have stringently selected for those genes, which displayed more than 3-fold changed mRNA expression levels at a significance level of p < 0.01 at Vd11 and Nd11 relative to the corresponding constitutive mRNA expressions at Vd0 and Nd0, respectively. Figure 1 shows the heat maps of the expression profiles of all self-healing and lethal infections on day 0 p.i. and day 11 p.i. The profiles of Vd0 and Nd0 separately cluster from those of Vd11 and Nd11, respectively.
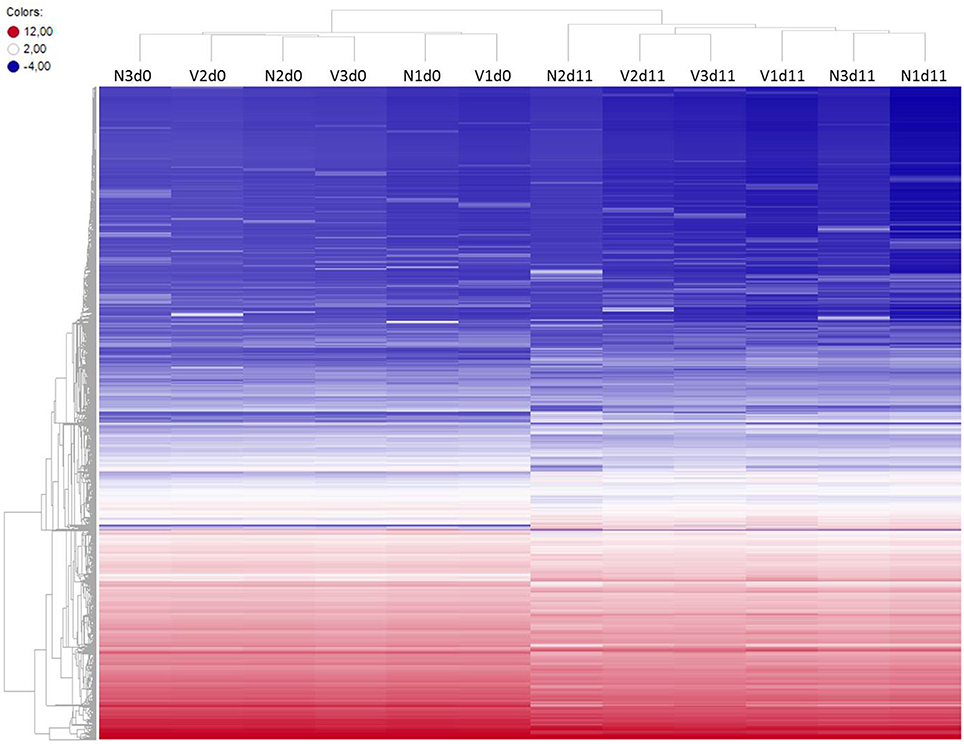
Figure 1. Heatmap of global expression levels of mRNA and lincRNA in the liver of non-vaccinated (N) and vaccinated (V) mice on day 0 p.i. (Nd0, Vd0) and day 11 p.i. (Nd11, Vd11). Expression levels for each sample were hierarchically clustered. Log2 transformed expression levels range from −4 to 12 indicated by blue and red color, respectively.
Protective vaccination does not affect constitutive mRNA expression in the liver. However, the P. chabaudi infections induce significant changes in the expression of numerous hepatic genes, which differ between vaccination-protected and non-vaccinated Balb/c mice toward the end of the crisis phase of blood-stage infections of P. chabaudi on day 11 p.i. The Venn diagrams in Figure 2 summarize the number of identified genes: malaria induces 302 genes to be up- and 1,147 genes to be down-regulated by more than 3-fold at Nd11 in relation to their constitutive expressions at Nd0. The identity of these genes is provided in the Tables S1–S4. Tables S1, S2 list the genes whose expression is up- and down-regulated by more than 3-fold and less than 10-fold, respectively. In Tables S3, S4, those genes are summarized whose expression is more than 3-fold up and less than 10-fold down-regulated, respectively, in the liver of non-vaccinated mice.
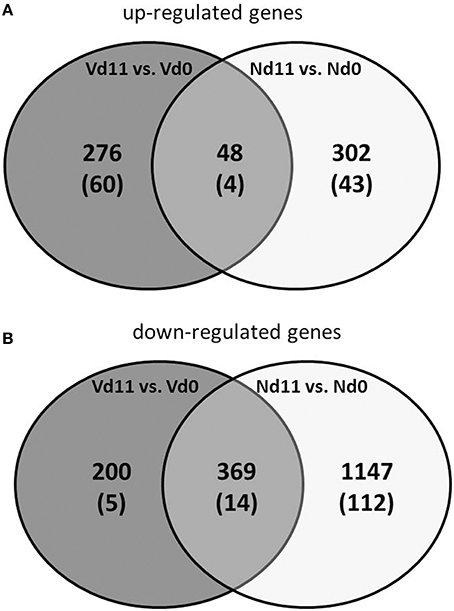
Figure 2. Venn-diagram of the number of up-regulated (A) and down-regulated (B) genes expressed more than 3-fold (p < 0.01) in the liver of vaccinated mice infected with P. chabaudi toward the end of the crisis phase on day 11 p.i. (Vd11) in comparison to constitutive expressions day 0 p.i. (Vd0). Nd11 vs. Nd0 indicates number of those genes identified in the liver of non-vaccinated mice. Numbers in brackets indicate genes more than 10-fold changed.
Moreover, Figure 2 shows 48 and 369 genes exhibiting malaria-induced up- and down-regulated expressions, respectively, by more than 3-fold both at Nd11 and at Vd11 as compared with their constitutive expressions at Nd0 and Vd0. The Tables S5, S6 summarize those genes, whose expression is up- and down-regulated, respectively, by more than 3-fold and less than 10-fold in the liver of both vaccinated and non-vaccinated mice. Table 1 lists those 4 genes and 14 genes, respectively, whose expressions are significantly up- and down-regulated by more than 10-fold in both non-vaccinated susceptible mice and vaccination-protected mice (Figure 2). However, the differences between both groups are not larger than about 2-fold—if at all. The 4 up-regulated genes are Apol10b, Hbq1b, Fcho1, and D1gap5. Among the 14 down-regulated genes, there are Cd209b and the 3 Mups2, 17, and 19, which are known to encode pheromone-binding proteins. Other genes encode proteins which are involved in liver metabolism and these genes are down-regulated to about the same extent in vaccination-protected and non-vaccinated mice.
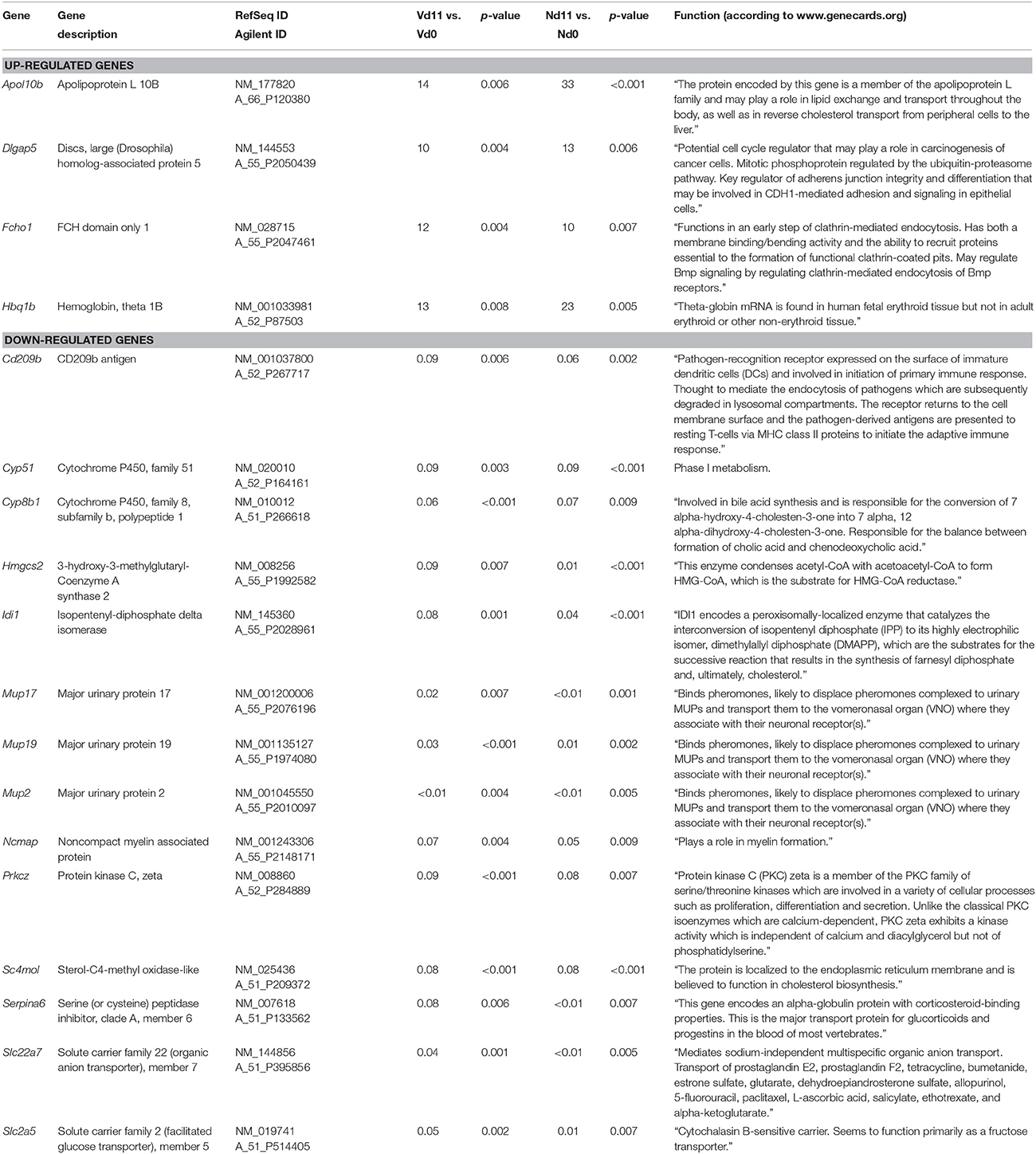
Table 1. Genes, whose expression is up-or down-regulated more than 10-fold (p < 0.01) in the liver of both non-vaccinated (N) and vaccinated mice (V) infected with P. chabaudi on day 11 p.i. (Vd11, Nd11) in comparison to constitutive expression on day 0 p.i. (Vd0, Nd0).
Protective vaccination does not only induce self-healing infections of P. chabaudi, but also the number and identity of the malaria-induced genes in the liver at Vd11 differ from those identified in non-vaccinated mice at Nd11: 276 and 200 genes have been identified to be up- and down-regulated, respectively, more than 3-fold at Vd11 in relation to Vd0 (Figures 2A,B). Tables S7, S8 list the 43 and 112 genes genes whose expression levels are significantly (p < 0.01) up- and down-regulated between 3- and 10-fold at Vd11 in relation to Vd0 (Figures 2A,B). In the following, the focus of this study will be on those genes which are more than 10-fold (p < 0.01) up- or down-regulated by malaria in the liver of vaccination-protected mice at Vd11.
Characterization of Genes Up-Regulated in Vaccination-Protected Mice
Among the 276 genes with up-regulated expression in the liver of only vaccination-protected mice toward the end of the crisis phase on day 11 p.i., there are 60 genes, whose expression is more than 10-fold up-regulated at Vd11 in comparison to Vd0, respectively (Figure 2A). These 60 genes are listed in Table 2, which also provides information about the currently annotated functions of their encoded proteins. The 60 up-regulated genes can be subsumed under specific functional groups. The most prominent group contains genes, which can be even up-regulated by far more than 100-fold, and which code for different erythroid-associated proteins, in particular proteins associated with the surface membrane. For instance, the Rhd encodes the RhD blood group, Add 2, Ank1, and Tmod1 encode proteins associated with membrane skeletal proteins, Ermap encodes a cell adhesion mediator, and Slc4a1 an integral membrane protein. Besides these membrane-associated genes, this group of genes also contains Gata1 and Gfi1b which are more than 70-fold up-regulated and which encode transcription factors involved in the development of erythroid cells. Another major group comprises genes which are associated with different phases of mitotis, such as Aspm, Bub1b, Casc5, Ccna2, Cdca5, Cenpe, Diap3, Ercc6l, Hist1h2ab, Hist3h2ba, Kif4, Kif15, Kif18b, Mns1, Nusap1, Prc1, Sgol1, Ska1, Spag5, Ube2c, and Uhrf1. Another group summarizes genes coding for proteins involved in cell cycle regulation, as e.g., Birc5, Bub1, Ccnb1, Ccnb2, Cdc25b, Espl1, Melk, Mybl2, Plk1, Rgcc, and Rrm2. The four genes Bard1, Fanci, Neil3, and Top2a code for proteins which have been described to be involved in DNA repair mechanisms. Moreover, the genes Kcnn4a and Mki67encode proteins involved in signaling and cell proliferation. Furthermore, the genes Cd24a, Lrr1, Pbk, and Trim59 can be assigned to immunity. Two genes can be subgrouped to tissue permeability, among which is Cldn13 that is more than 200-fold up-regulated. The encoded membrane protein claudin13 is known to be associated with tight junctions.
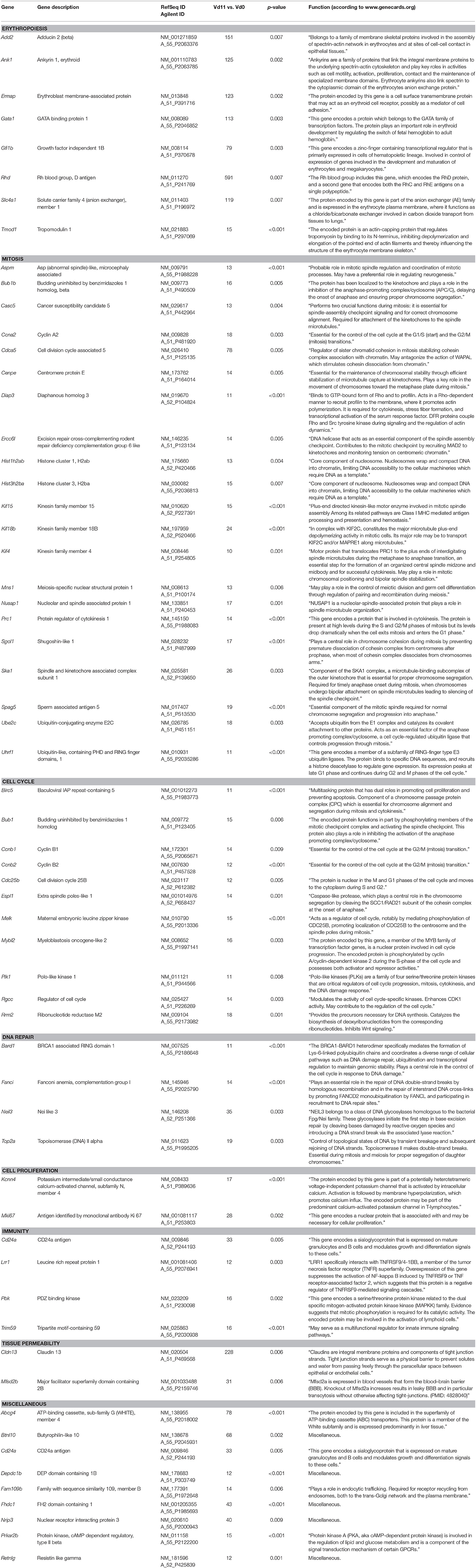
Table 2. Genes expressed more than 10-fold (p < 0.01) in the liver of vaccinated mice infected with P. chabaudi on day 11 p.i. (Vd11) in comparison to constitutive expression on day 0 p.i. (Vd0).
Finally, quantitative PCR of arbitrarily selected up-regulated genes confirms that the expression of the erythroid-associated genes Add2, Ank1, Ermap, and Rhd are indeed up-regulated by more than 100-fold, while the genes Ccna2 associated with mitosis and Plk1 involved in cell cycle regulation are more than 10-fold expressed in the liver of vaccination-protected Balb/c mice infected with P. chabaudi on day 11 p.i. (Vd11) in relation to constitutive expression at Vd0 (Figure 3).
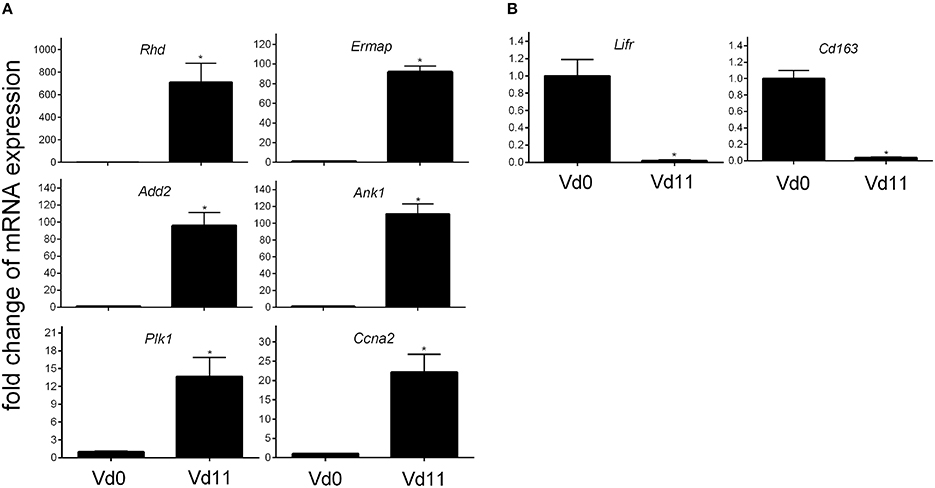
Figure 3. Quantitative PCR of mRNAs of arbitrarily selected up-regulated (A) and down-regulated (B) genes in the liver of vaccinated Balb/c mice infected with P. chabaudi on day 0 p.i. (Vd0) and on day 11 p.i. (Vd11). Values are means of triplicate determinations. Bars indicate half SEM and stars significant differences (p < 0.01) between Vd0 and Vd11.
Characterization of Genes Down-Regulated in Vaccination–Protected Mice
Among the 200 genes down-regulated in the liver of vaccination-protected mice at Vd11, only 5 genes in the liver respond to malaria with a significant down-regulation by more than 10-fold (p < 0.01) (Figure 2B). One of these 5 genes is Cd163 which is down-regulated by approximately 50-fold (Table 3). The encoded CD163 is a transmembrane protein of 130 KDa expressed on Kupffer cells in the liver serving as a scavenger receptor cysteine-rich superfamily in the clearence and endocytosis of hemoglobin/haptoglobin complexes (Gronbaek et al., 2012; Etzerodt et al., 2013). Another gene is Lifr, whose expression is down-regulated by approximately 17-fold, encodes the transmembrane leukemia inhibitory factor (LIF) receptor (LIFR). Ectodomain shedding creates soluble receptor forms of both LIFR and CD163, which have different functions as the membrane-bound receptors (Layton et al., 1994; Owczarek et al., 1996; Moller, 2012; Ingels et al., 2013; Onishi and Zandstra, 2015). Quantitative PCR confirms that the expression of both Lifr and Cd163 is strongly down-regulated at Vd11 as compared with Vd0 (Figure 3).
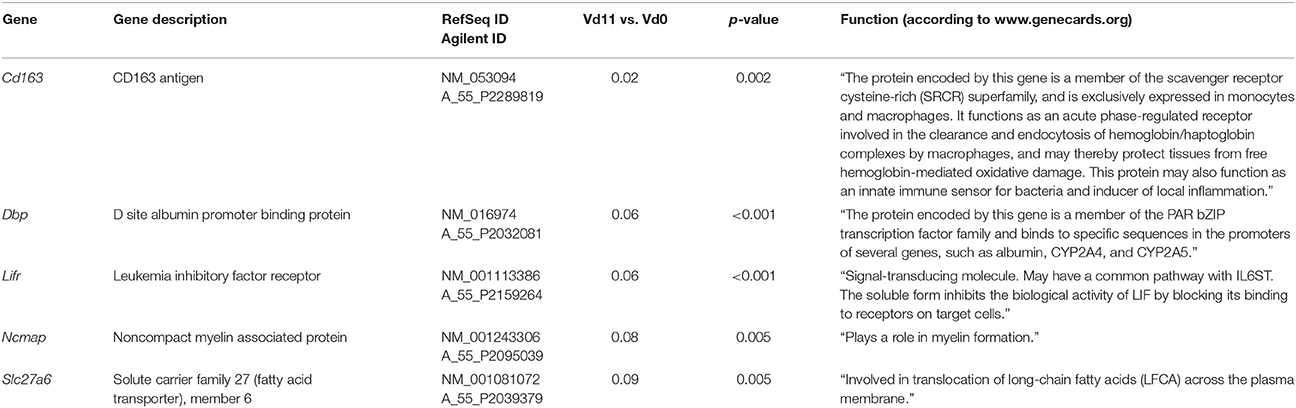
Table 3. Genes down-regulated more than 10-fold (p < 0.01) in the liver of vaccinated mice infected with P. chabaudi on day 11 p.i. (Vd11) in comparison to constitutive expression on day 0 p.i. (Vd0).
Identification of Differentially Expressed lincRNAs
Besides the 39,430 Entrez Gene RNAs, the used microarrays also contain 16,251 probes for detecting lincRNAs. In general, lincRNAs are known to range in length between 200 nts and about 100 kb, but lincRNAs are never transcribed into proteins. However, they are increasingly recognized as to play critical roles in different diseases, including diverse liver diseases (Takashi et al., 2014). We therefore also screened the microarrays for lincRNAs using the same stringent conditions as above for the mRNAs. Protective vaccinaton does not change the constitutive expression of lincRNA. However, malaria has induced changes in the expression of lincRNAs toward the end of the crisis phase, which differ between vaccination-protected mice and non-protected mice. The Venn-diagrams in Figure 4 show that, vaccination-protected mice, malaria induces 40 lincRNAs to be up-regulated more than 3-fold (Table S9), among which are 6 lincRNAs up-regulated by more than 10-fold at Vd11 in relation to Vd0. About the same number, namely 43 lincRNAs, are down-regulated by more than 3-fold in non-vaccinated mice at Vd11, but none of them is more than 10-fold down-regulated (Table S10). However, annotated functions are not yet available for any of the differentially regulated lincRNAs.
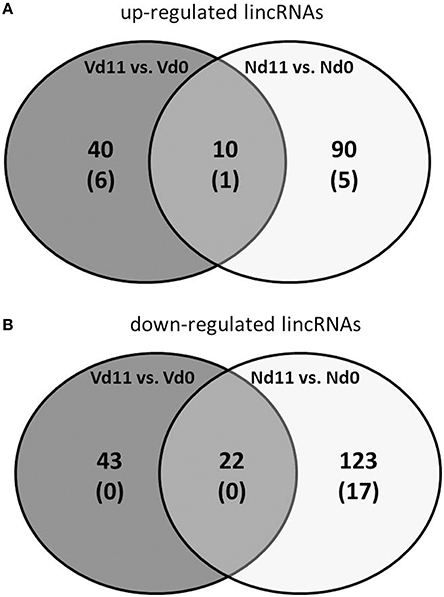
Figure 4. Number of up-regulated (A) and down-regulated (B) lincRNAs expressed more than 3-fold (p < 0.01) in the liver of non-vaccinated (N) and vaccinated mice (V) infected with P. chabaudi on day 11 p.i. as compared to constitutive expressions on day 0 p.i. (Nd11 vs. Nd0 and Vd11 vs. Vd0). Numbers in brackets indicate lincRNAs expressed more than 10-fold.
Discussion
The vaccination procedure used here has been previously shown to protect female Balb/c mice from fatal P. chabaudi blood-stage malaria (Krücken et al., 2009). This becomes evident toward the end of the crisis phase on day 11 p.i., when all non-vaccinated mice will succumb to infections, but the majority of the vaccinated mice will survive. This survival is associated with malaria-induced differential expression of genes and lincRNAs in the liver, as identified in the present study. These changes in gene expression are presumably associated with processes in the liver, which contribute to survival of blood-stage malaria of P. chabaudi.
Extramedullary erythropoiesis obviously is one of these malaria-activated processes in the liver of vaccination-protected mice. Indeed, we have detected a massive up-regulation of hepatic genes by far more than 100-fold, which encode diverse proteins of erythroid cells. In accordance, previous findings have provided evidence for the onset of hepatic erythropoiesis during blood-stage malaria in mice (Abo and Sekikawa, 2002; Halder et al., 2003; Wunderlich et al., 2014). It is therefore rather likely that reticulocytes produced in the liver contribute to the increasing number of peripheral reticulocytes in vaccination-induced self-healing infections of P. chabaudi. This increase begins at approximately peak parasitemia on day 8 p.i. and reaches its maximum of approximately 80% at the end of the crisis phase on approximately days 11 p.i., when the number of peripheral erythrocytes has concomitantly declined to a minimum (Krücken et al., 2009). This dramatic decline of erythrocytes is presumably achieved both by mechanical destruction of P. chabaudi-infected erythrocytes, when the merozoite stages are released from erythrocytes, and by erythrophagocytosis mediated through natural auto-antibodies directed, for example, against clustered band 3 membrane proteins of senescent and aberrant erythrocytes including even P. falciparum-infected erythrocytes (Arese et al., 2005; Winograd et al., 2005; Lutz, 2012; Lutz and Bogdanova, 2013). In this context, it is worth mentioning that the spleen is rather likely not that organ which massively removes P. chabaudi-infected erythrocytes from circulation during the crisis phase of infection. Indeed, the spleen during crisis dramatically decreases the uptake of injected particulate material including fluorescently labeled P. chabaudi-infected erythrocytes (Krücken et al., 2005, 2009), while, concomitantly, the liver dramatically increases its capacity to trap injected particles. Thus, the liver, for example the Kupffer cells, rather than the “closed” spleen may massively remove P. chabaudi-infected erythrocytes from circulation during crisis, and this may even trigger hepatic erythropoiesis.
Anti-erythrocyte autoimmune mechanisms are obviously not directed against reticulocytes, since their number is increasingly replacing the declining number of erythrocytes during crisis. Moreover, the massive increase in reticulocytes is—at least transiently—advantageous for host survival, since reticulocytes, in contrast to erythrocytes, are not the preferred host cells of P. chabaudi. This non-preference may be due to the different composition and arrangement of plasma membrane-associated proteins of reticulocytes as compared with erythrocyte (Liu et al., 2010), which would make it more difficult for P. chabaudi to invade and/or survive in reticulocytes than in erythrocytes (Ott, 1968). This view is also supported by our finding showing an massive up-regulation of genes encoding erythroid skeletal membrane proteins such as Add2 and Ank1. In this context, it is also noteworthy that C57BL/10 mice which have survived blood-stage malaria of P. chabaudi and subsequently acquired immunity to homologous re-infection have been previously shown to possess erythrocytes with a deficiency in the band 4.1a membrane protein (Wunderlich and Helwig, 1987). Erythrocytes deficient in 4.1a are known to be prone to ovalocytosis (Alloisio et al., 1983) and to be less penetrable by Plasmodium parasites (Hadley et al., 1983).
Furthermore, we have identified quite a number of up-regulated genes in the liver, which are known to control progress of cell cycle, mitosis and DNA repair. This may be, at least in part, also associated with increased hepatic erythropoiesis including enucleation of erythroblasts which may be regarded as an asymmetric form of mitosis (Thompson et al., 2010). However, the most straightforward interpretation of these data is that the liver of vaccination-protected mice, in comparison with non-vaccinated mice, undergoes accelerated regeneration toward the end of the crisis phase. This presumably enhances recovery of the liver from the pathological damages and injuries induced by the P. chabaudi infections. This view of accelerated liver regeneration may be also supported by our finding that the claudin13 encoding gene Cldn13 is one of the most massively up-regulated genes in the liver of vaccination-protected mice toward the end of the crisis phase. In the mouse genome, Cldn13 is one of 26 genes encoding claudins, which are 4-pass integral membrane proteins of 20–27 KDa and which are the main constituents of tight junctions (Morita et al., 1999; Lal-Nag and Morin, 2009). These restrict as a paracellular barrier the uncontrolled transfer of water and solutes in the intercellular space between cells of epithelia (Van Itallie and Anderson, 2004). The massive upregulation of Cldn13 may therefore contribute to osmotic stability of liver tissue of the regenerating liver. However, there is also information available that the mouse-specific Cldn13 encodes the most abundant claudin in the spleen of mice, where it has been predicted to be localized on the surface of erythroblasts and to be coordinately regulated as a part of stress-induced erythropoiesis by Trypanosoma congolense (Thompson et al., 2010). On the other hand, it is known that the liver is the only erythropoietic organ in the fetus, and the adult liver of mice is induced to resume erythropoiesis upon infection with the self-healing blood-stage malaria of P. yoelii 17X (Abo and Sekikawa, 2002; Halder et al., 2003). De novo clusters of erythroid cells have been shown in the parenchymal space of the liver. Thus, our finding, that Cldn13 is massively upregulated in the liver of vaccinated Balb/c mice, may reflect a local expression of Cldn13 in/around the microenvironment of erythroid clusters, thus protecting erythroid clusters and erythropoiesis, respectively, from osmotic perturbations. Local expressions of Cldn13 have been also described for Sertoli cell tight junctions (Chakraborty et al., 2014) as well as for neonatal rabbit and rat proximal renal tubules (Abuazza et al., 2006).
Moreover, even down-regulated genes, for example Lifr, in the liver may contribute to enhanced liver regeneration in vaccination-protected mice. The LIF/LIFR system has been hypothesized to be involved in the expansion and differentiation of the liver stem cell compartment, which is activated during liver regeneration (Omori et al., 1996). Moreover, the LIF/LIFR system shares a common pathway through the common signal transducer GP130 with the polyfunctional interleukin 6 class cytokine family, consisting of IL-6, IL-11, OSM (oncostatin M), CNTF (cilary neurotrophic factor), CT-1 (cardiotrophin-1), and CLC (cardiotrophin-like cytokine) (Heinrich et al., 2003). These mutually interacting polyfunctional signaling networks are rather complex and involve both classic and trans-signaling mechanisms. For instance, IL-6 classic signaling takes place only in immune cells and hepatocytes expressing membrane-bound IL-6 receptor alpha. However, when bound to the soluble receptor created by ectodomain shedding from the membrane-bound receptor, IL-6 is then able to communicate by trans-signaling with all other cells through the ubiquitously expressed signal transducer GP130. IL-6 trans-signaling has been shown to play a decisive role in the regulation of liver regeneration (Scheller et al., 2011; Schmidt-Arras and Rose-John, 2016). Inhibition of IL-6 trans-signaling protects from P. chabaudi malaria (Wunderlich et al., 2012).
Moreover, even the differential expression of distinct lincRNA species with no annotated functions in the liver indicate that liver regeneration and hepatic erythropoiesis toward the end of the crisis phase require changes in the expression of diverse lincRNA species (Quagliata and Terraciano, 2014). Erythropoiesis has been recently shown to be associated with synthesis of diverse lncRNAs (Xu et al., 2013), targeted in part by the transcription factor GATA1 (Alvarez-Dominguez et al., 2014). The gene coding for GATA1 has been found here to be massively up-regulated by more than 100-fold in vaccination-protected mice toward the end of the crisis phase.
Collectively, our data indicate that protective vaccination against P. chabaudi malaria leads to changes in the liver, evidenced as altered gene expression in response to P. chabaudi malaria toward the end of the crisis phase. These changes in gene expression support the view that the dramatic decline of P. chabaudi-infected erythrocytes during crisis is counter-regulated by hepatic erythropoiesis leading to massive up-regulation of peripheral reticulocytes, which are not favored as host cells by P. chabaudi. Concomitantly, there is an accelerated regeneration of the liver. Both hepatic regeneration and erythropoiesis contribute to vaccination-induced survival of otherwise lethal blood-stage malaria of P. chabaudi.
Author Contributions
MD, SA, AA, and FW designed the study, MA and DD carried out the experiments and analyzed the data. All authors wrote and revised the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This project was funded by the national plan for science, technology and innovation (MAARIFAH), King Abdulaziz city for science and technology, Kingdom of Saudi Arabia, awarded number (13-B101206-02).
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2016.01087
References
Abo, T., and Sekikawa, H. (2002). Extrathymic T cells in malaria protection, including evidence for the onset of erythropoiesis in the liver during infection. Arch. Histol. Cytol. 65, 127–132. doi: 10.1679/aohc.65.127
Abuazza, G., Becker, A., Williams, S. S., Chakravarty, S., Truong, H.-T., Lin, F., et al. (2006). Claudins 6, 9, and 13 are developmentally expressed renal tight junction proteins. Am. J. Physiol. Renal. Physiol. 291, F1132–F1141. doi: 10.1152/ajprenal.00063.2006
Alloisio, N., Dorleac, E., Morle, L., Girot, R., Galant, C., Boivin, P., et al. (1983). The genetic abnormalities involving red cell membrane protein 4.1 with or without elliptocytosis. Biomed. Biochim. Acta 42, S38–S42.
Al-Quraishy, S., Dkhil, M. A., Abdel-Baki, A. A., Araúzo-Bravo, M. J., Delic, D., and Wunderlich, F. (2014). Testosterone persistently dysregulates hepatic expression of Tlr6 and Tlr8 induced by Plasmodium chabaudi malaria. Parasitol. Res. 113, 3609–3620. doi: 10.1007/s00436-014-4026-2
Alvarez-Dominguez, J. R., Hu, W., Yuan, B., Shi, J., Park, S. S., Gromatzky, A. A., et al. (2014). Global discovery of erythroid long noncoding RNAs reveals novel regulators of red cell maturation. Blood 123, 570–581. doi: 10.1182/blood-2013-10-530683
Ananad, A. C., Ramji, C., Narula, A. S., and Singh, W. (1992). Malarial hepatitis: a heterogenous syndrome. Natl. Med. J. India 5, 59–62.
Arese, P., Turrini, F., and Schwarzer, E. (2005). Band 3/complement-mediated recognition and removal of normally senescent and pathological human erythrocytes. Cell. Physiol. Biochem. 16, 133–146. doi: 10.1159/000089839
Bertolino, P., and Bowen, D. G. (2015). Malaria and the liver: immunological hide-and-seek or subversion of immunity from within? Front. Microbiol. 18:41. doi: 10.3390/pathogens5010002
Birkett, A. J. (2016). Status of vaccine research and development of vaccines for malaria. Vaccine 34, 2915–2920. doi: 10.1016/j.vaccine.2015.12.074
Chakraborty, P., Buass, F. W., Sharma, M., Smith, B. E., Greenlee, A. R., Eacker, S. M., et al. (2014). Androgen-dependent Sertoli cell tight junction remodeling is mediated by multiple tight junctions components. Mol. Endocrinol. 28, 1055–1072. doi: 10.1210/me.2013-1134
Dockrell, H. M., de Souza, J. B., and Playfair, J. H. (1980). The role of the liver in immunity to blood-stage murine malaria. Immunology 41, 421–430.
Etzerodt, A., Kjolby, M., Nielsen, M. J., Maniecki, M., Svendsen, P., and Moestrup, S. K. (2013). Plasma clearence of hemoglobin and haptoglobin in mice and effect of CD163 gene targeting disruption. Antooxid Redox Signal 18, 2254–2263. doi: 10.1089/ars.2012.4605
Fontaine, A., Bourdon, S., Belghazi, M., Pophillat, M., Fourquet, P., Granjeaud, S., et al. (2012). Plasmodium falciparum infection-induced changes in erythrocyte proteins. Parasitol. Res. 110, 545–556. doi: 10.1007/s00436-011-2521-2
Frevert, U., and Krzych, U. (2015). Plasmodium cellular effector mechanisms and the hepatic microenvironment. Front. Microbiol. 6:482. doi: 10.3389/fmicb.2015.00482
Frevert, U., Krzych, U., and Richie, T. L. (2015). Editorial: breaking the cycle: attacking the malaria parasite in the liver. Front. Microbiol. 6:810. doi: 10.3389/fmicb.2015.00810
Gosling, R., and von Seidlein, L. (2016). The Future of the RTS,S/AS01 Malaria vaccine: an alternative development plan. PLoS Med. 13:e1001994. doi: 10.1371/journal.pmed.1001994
Gronbaek, H., Sandahl, T. D., Mortensen, C., Vilstrup, H., Moller, H. J., and Moller, S. (2012). Soluble CD163, a marker of Kupffer cell activation, is related to portal hypertension in patients with liver cirrhosis. Aliment. Pharmacol. Ther. 36, 173–180. doi: 10.1111/j.1365-2036.2012.05134.x
Hadley, T., Saul, A., Lamont, G., Hudson, D. E., Miller, L. H., and Kidson, C. (1983). Resistance of Melanesian elliptocytes (ovalocytes) to invasion by Plasmodium knowlesi and Plasmodium falciparum malaria parasites in vitro. J. Clin. Inves. 71, 780–782. doi: 10.1172/JCI110827
Halbroth, B. R., and Draper, S. J. (2015). Recent developments in malaria vaccinology. Adv. Parasitol. 88, 1–49. doi: 10.1016/bs.apar.2015.03.001
Halder, R. C., Abe, T., Mannoor, M. K., Morshed, S. R., Ariyasinghe, A., Watanabe, H., et al. (2003). Onset of hepatic erythropoiesis after malarial infection in mice. Parasitol. Int. 52, 259–268. doi: 10.1016/S1383-5769(03)00029-1
Heinrich, P. C., Behrmann, I., Haan, S., Hermanns, H. M., Müller-Neven, G., and Schaper, F. (2003). Principles of interleukin (IL)-6-type cytokine signaling and its regulation. Biochem. J. 374, 1–20. doi: 10.1042/bj20030407
Hoffman, S. L., Vekemans, J., Richie, T. L., and Duffy, P. E. (2015). The march toward malaria vaccines. Am. J. Prev. Med. 49(6 Suppl. 4), S319–S333. doi: 10.1016/j.amepre.2015.09.011
Ingels, C., Moller, H. J., Hansen, T. K., Wouters, P. J., Vanhorebeek, I., and van den Berghe, G. (2013). Circulating levels of the scavenger receptor sCD163 and association with outcome of critically ill patients. J. Clin. Immunol. 33, 619–629. doi: 10.1007/s10875-012-9830-9
Kochar, D. K., Singh, P., Agarwal, P., Kochar, S. K., and Sareen, P. K. (2003). Malarial hepatitis. J. Assoc. Physicians India 51, 1069–1072.
Krücken, J., Delić, D., Pauen, H., Wojtalla, A., El-Khadragy, M., Dkhil, M. A., et al. (2009). Augmented particle trapping and attenuated inflammation in the liver by protective vaccination against Plasmodium chabaudi malaria. Malaria J. 8–54. doi: 10.1186/1475-2875-8-54
Krücken, J., Mehnert, L. I., Dkhil, M., El-Khadragy, M., Benten, W. P. M., Mossmann, H., et al. (2005). Massive destruction of malaria-parasitized red blood cells despite spleen closure. Infect. Immun. 73, 6390–6398. doi: 10.1128/IAI.73.10.6390-6398.2005
Lal-Nag, M., and Morin, P. J. (2009). The claudins. Genome Biol. 10:235. doi: 10.1186/gb-2009-10-8-235
Layton, M. J., Owczarek, C. M., Metcalf, D., Lock, P., Wilson, T. A., Gough, N. M., et al. (1994). Complex biding of leukemia inhibitory factor to its membrane expressed and soluble receptors. Proc. Soc. Exp. Biol. Med. 206, 295–298. doi: 10.3181/00379727-206-43762
Liu, J., Guo, X., Mohandas, N., Chasis, J. A., and An, X. (2010). Membrane remodeling during reticulocyte maturation. Blood 115, 2021–2027. doi: 10.1182/blood-2009-08-241182
Livak, K. J., and Schmittgen, T. D. (2002). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408. doi: 10.1006/meth.2001.1262
Longley, R., Smith, C., Fortin, A., Berghout, J., McMorran, B., Burgio, G., et al. (2011). Host resistance to malaria: using mouse models to explore the host response. Mamm. Genome 22, 32–42. doi: 10.1007/s00335-010-9302-6
Lutz, H. U. (2012). Naturally occurring anti-band 3 antibodies in clearence of senescent and oxidatively stressed human red blood cells. Transfus. Med. Hemother. 39, 321–327. doi: 10.1159/000342171
Lutz, H. U., and Bogdanova, A. (2013). Mechanisms tagging senescent red blood cells for clearence in healthy humans. Front. Physiol. 4:387. doi: 10.3389/fphys.2013.00387
Miura, K. (2016). Progress and prospects for blood-stage malaria vaccines. Expert Rev. Vaccines 3, 1–17. doi: 10.1586/14760584.2016.1141680
Moller, H. J. (2012). Soluble CD163. Scand. J. Clin. Lab. Invest. 72, 1–13. doi: 10.3109/00365513.2011.626868
Morita, K., Furuse, M., Fujimoto, K., and Tsikita, S. (1999). Claudin multigene family encoding four-transmembrane domain protein components of tight junction strands. Proc. Natl. Acad. Sci. U.S.A. 96, 511–516. doi: 10.1073/pnas.96.2.511
Nautyal, A., Singh, S., Parameswaran, G., and DiSalle, M. (2005). Hepatic dysfunction in a patient with Plasmodium vivax infection. MedGenMed. 7, 8–9.
Omori, N., Evarts, R. P., Omori, M., Hu, Z., Marsden, E. R., and Thorgeirsson, S. S. (1996). Expression of leukemia inhibitory factor and its receptor during liver regeneration in the adult rat. Lab. Invest. 75, 15–24.
Onishi, K., and Zandstra, P. W. (2015). LIF signaling in stem cells and development. Development 142, 2230–2236. doi: 10.1242/dev.117598
Ott, K. J. (1968). Influence of reticulocytosis on the course of infection of Plasmodium chabaudi and P. berghei. J. Eukaryot. Microbiol. 15, 365–369.
Owczarek, C. M., Layton, M. J., Robb, L. G., and Nicola, N. A. (1996). Molecular basis of the soluble and membrane-bound forms of the murine leukemia inhibitory factor receptor alpha-chain. J. Biol. Chem. 271, 5495–5504. doi: 10.1074/jbc.271.10.5495
Quagliata, L., and Terraciano, L. M. (2014). Liver diseases and long non-coding RNAs: new insight and perspective. Front. Med. 1:35. doi: 10.3389/fmed.2014.00035
Rupani, A. B., and Amarapurkar, A. D. (2009). Hepatic changes in fatal malaria: an emerging problem. Ann. Trop. Med. Parasitol. 103, 119–127. doi: 10.1179/136485909X385054
Scheller, J., Chalaris, A., Schmidt-Arras, D., and Rose-John, S. (2011). The pro- and anti-inflammatory properties of the cytokine interleukin 6. Biochim. Biophys. Acta 1813, 878–888. doi: 10.1016/j.bbamcr.2011.01.034
Schmidt-Arras, D., and Rose-John, S. (2016). Il-6 pathway in the liver: from pathophysiology to therapy. J. Hepatol. 64, 1403–1415. doi: 10.1016/j.jhep.2016.02.004
Stephens, R., Culleton, R. L., and Lamb, T. J. (2012). The contribution of Plasmodium chabaudi to our understanding of malaria. Trends Parasitol. 28, 73–82. doi: 10.1016/j.pt.2011.10.006
Takashi, K., Yan, I., Haga, H., and Patel, T. (2014). Long noncoding RNA in liver diseases. Hepatology 60, 744–753. doi: 10.1002/hep.27043
Thompson, P. D., Tipney, H., Brass, A., Noyes, H., Kemp, S., Naessens, J., et al. (2010). Claudin 13, a member of the claudin family regulated in mouse stress induced erythropoiesis. PLoS ONE 9:e12667. doi: 10.1371/journal.pone.0012667
Van Itallie, C. M., and Anderson, J. M. (2004). The molecular physiology of tight junctions pores. Physiology 19, 331–338. doi: 10.1152/physiol.00027.2004
White, M. T., Verity, R., Griffin, J. T., Asante, K. P., Owusu-Agyei, S., Greenwood, B., et al. (2015). Immunogenicity of the RTS,S/AS01 malaria vaccine and implications for duration of vaccine efficacy: secondary analysis of data from a phase 3 randomised controlled trial. Lancet Infect. Dis.15, 1450–1458. doi: 10.1016/S1473-3099(15)00239-X
WHO (2015). World Malaria Report 2015, Released in December 2015 by World Health Organization 2015, WHO Document Production Services. Geneva, Switzerland. Available online at: http://www.who.int/malaria/publications/world-malaria-report-2015/report/en
Winograd, E., Prudhomme, J. G., and Sherman, I. W. (2005). Band 3 clustering promotes the exposure of neoantigens in Plasmodium falciparum-infected erythrocytes. Mol. Biochem. Parasitol. 142, 98–105. doi: 10.1016/j.molbiopara.2005.03.013
Wunderlich, C. M., Delic, D., Behnke, K., Meryk, A., Ströhle, P., Chaurasia, B., et al. (2012). Cutting edge: inhibition of IL-6 trans-signaling protects from malaria-induced lethality in mice. J. Immunol. 188, 4141–4144. doi: 10.4049/jimmunol.1102137
Wunderlich, F., Al-Quraishy, S., and Dkhil, M. (2014). Liver-inherent immune system: its role in blood-stage malaria. Front. Microbiol. 5:559. doi: 10.3389/fmicb.2014.00559
Wunderlich, F., Brenner, H., and Helwig, M. (1988a). Plasmodium chabaudi malaria: protective immunization with surface membranes of erythrocytes infected with Plassmodium chabaudi. Infect. Immun. 56, 3326–3328.
Wunderlich, F., Dkhil, M., Mehnert, L., Braun, J., El-Khadragy, M., Borsch, E., et al. (2005). Testosterone responsiveness of spleen and liver in female lymphotoxin beta receptor-deficient mice resistant to blood-stage malaria. Microbes Infect. 7, 399–409. doi: 10.1016/j.micinf.2004.11.016
Wunderlich, F., and Helwig, M. (1987). Plasmodium chabaudi malaria: red blood cells with altered membrane proteins in immune mice. Eur. J. Cell Biol. 43, 499–500.
Wunderlich, F., Helwig, M., Schillinger, G., and Speth, V. (1988c). Cryptic disposition of antigenic parasite proteins in plasma membranes of erythrocytes infected with Plasmodium chabaudi. Mol. Biochem. Parasitol. 30, 55–65. doi: 10.1016/0166-6851(88)90132-6
Wunderlich, F., Helwig, M., Schillinger, G., Speth, V., and Wiser, M. F. (1988d). Expression of the parasite protein Pc90 in plasma membranes of erythrocytes infected with Plasmodium chabaudi. Eur. J. Cell Biol. 47, 157–164.
Wunderlich, F., Helwig, M., Schillinger, G., Vial, H., Philippot, J., and Speth, V. (1987). Isolation and characterization of parasites and host cell ghosts from erythrocytes infected with Plasmodium chabaudi. Mol. Biochem. Parasitol. 23, 103–115. doi: 10.1016/0166-6851(87)90145-9
Wunderlich, F., Mossmann, H., Helwig, M., and Schillinger, G. (1988b). Resistance to Plasmodium chabaudi in B10 mice: influence of the H-2 complex and testosterone. Infect. Immun. 56, 2400–2406.
Wunderlich, F., Schillinger, G., and Helwig, M. (1985). Fractionation of Plasmodium chabaudi-infected erythrocytes into parasites and ghosts. Z. Parasitenkd. 71, 545–551. doi: 10.1007/BF00928358
Wunderlich, F., Stuebig, H., and Koenigk, E. (1982). Development of Plasmodium chabaudi in mouse red blood cells: structural properties of the host and parasite membranes. J. Protozool. 29, 60–66. doi: 10.1111/j.1550-7408.1982.tb02880.x
Keywords: blood-stage malaria, Plasmodium chabaudi, protective vaccination, gene expression, lincRNA, hepatic erythropoiesis, liver regeneration
Citation: Al-Quraishy SA, Dkhil MA, Abdel-Baki A-AA, Delic D and Wunderlich F (2016) Protective Vaccination against Blood-Stage Malaria of Plasmodium chabaudi: Differential Gene Expression in the Liver of Balb/c Mice toward the End of Crisis Phase. Front. Microbiol. 7:1087. doi: 10.3389/fmicb.2016.01087
Received: 22 April 2016; Accepted: 29 June 2016;
Published: 14 July 2016.
Edited by:
Heinrich Korner, Menzies Research Institute Tasmania, AustraliaReviewed by:
Ann-Kristin Mueller, University Hospital Heidelberg, GermanyWai-Hong Tham, The Walter and Eliza Hall Institute, Australia
Copyright © 2016 Al-Quraishy, Dkhil, Abdel-Baki, Delic and Wunderlich. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Mohamed A. Dkhil, bW9oYW1lZGRraGlsQHlhaG9vLmNvbQ==
 Saleh A. Al-Quraishy
Saleh A. Al-Quraishy Mohamed A. Dkhil
Mohamed A. Dkhil Abdel-Azeem A. Abdel-Baki
Abdel-Azeem A. Abdel-Baki Denis Delic
Denis Delic Frank Wunderlich
Frank Wunderlich