- 1Division of Bacteriology, National Institute of Cholera and Enteric Diseases, Kolkata, India
- 2Absolut Data Labs, Absolut Data Research and Analytics, Gurgaon, India
- 3Department of Neonatology, Institute of Postgraduate Medical Education and Research, SSKM Hospital, Kolkata, India
Carbapenem-resistant determinants and their surrounding genetic structure were studied in Acinetobacter spp. from neonatal sepsis cases collected over 7 years at a tertiary care hospital. Acinetobacter spp. (n = 68) were identified by ARDRA followed by susceptibility tests. Oxacillinases, metallo-β-lactamases (MBLs), extended-spectrum β-lactamases and AmpCs, were detected phenotypically and/or by PCR followed by DNA sequencing. Transconjugants possessing the blaNDM−1(New Delhi metallo-β-lactamase) underwent further analysis for plasmids, integrons and associated genes. Genetic environment of the carbapenemases were studied by PCR mapping and DNA sequencing. Multivariate logistic regression was used to identify risk factors for sepsis caused by NDM-1-harboring organisms. A. baumannii (72%) was the predominant species followed by A. calcoaceticus (10%), A. lwoffii (6%), A. nosocomialis (3%), A. junni (3%), A. variabilis (3%), A. haemolyticus (2%), and 14TU (2%). Fifty six percent of the isolates were meropenem-resistant. Oxacillinases present were OXA-23-like, OXA-58-like and OXA-51-like, predominately in A. baumannii. NDM-1 was the dominant MBL (22%) across different Acinetobacter spp. Isolates harboring NDM-1 also possessed bla(VIM−2, PER−1, VEB−2, CTX−M−15), armA, aac(6′)Ib, aac(6′)Ib-cr genes. blaNDM−1was organized in a composite transposon between two copies of ISAba125 in the isolates irrespective of the species. Further, OXA-23-like gene and OXA-58-like genes were linked with ISAba1 and ISAba3 respectively. Isolates were clonally diverse. Integrons were variable in sequence but not associated with carbapenem resistance. Most commonly found genes in the 5′ and 3′conserved segment were aminoglycoside resistance genes (aadB, aadA2, aac4′), non-enzymatic chloramphenicol resistance gene (cmlA1g) and ADP-ribosylation genes (arr2, arr3). Outborn neonates had a significantly higher incidence of sepsis due to NDM-1 harboring isolates than their inborn counterparts. This study demonstrates the significance of both A. baumannii and other species of Acinetobacter in cases of neonatal sepsis over an extended period. Oxacillinases and blaNDM−1 are the major contributors to carbapenem resistance. The dissemination of the blaNDM−1 is likely linked to Tn125 in diverse clones of the isolates.
Introduction
Acinetobacter, a non-fermenting Gram-negative coccobacilli, has been noted as a pathogen in neonatal units in several parts of the globe and outbreaks caused by this organism have also been reported (Huang et al., 2002; Chan et al., 2007; Hammerum et al., 2015). Though A. baumannii has been predominantly implicated in neonatal infections or sepsis, “other species” within the genus have also been reported to have caused sepsis in neonates (Beaufort et al., 1999; Kilic et al., 2008). The ability of the species to acquire antibiotic resistance has now resulted in the acquisition of the potent carbapenem resistant gene, blaNDM−1 (Bonnin et al., 2012; Boulanger et al., 2012; Chuanfu et al., 2013). This enzyme, New Delhi Metallo-β–lactamase-1 (NDM-1), is an addition to the wide array of antimicrobial resistance mechanisms that have been described for Acinetobacter spp.
There have been reports of NDM-1 in Acinetobacter from different countries (Nordmann et al., 2011). Variants of this enzyme from Acinetobacter have also been reported (Espinal et al., 2011). A study has shown that blaNDM−1 is possibly a chimera that was constructed in A. baumannii (Dortet et al., 2014). Though most studies report the presence of this enzyme in A. baumannii, NDM-1 has also been detected in other species such as A. lwoffii and A. pittii (Hu et al., 2012; Pasteran et al., 2014). Apart from NDM-1, carbapenem resistance in Acinetobacter is mediated predominantly by the carbapenem-hydrolyzing-oxacillinases belonging to molecular class D (OXA-type carbapenemases) and also by other metallo-β-lactamases (MBLs) belonging to the class B enzymes such as the IMP-and VIM-types.
In addition to the antibiotic resistance, the identification within this genus remains difficult (Vaneechoutte et al., 1995). Acinetobacter genus consists of more than 30 species (Turton et al., 2011) and identification requires molecular methods which are not available in the clinical laboratories in developing countries. Thus, these organisms are not identified to the species level using appropriate methods. Until this is done, the association of the various species to nosocomial cases can never be achieved.
Most investigations of Acinetobacter in neonatal units have been ad hoc studies that were triggered by an outbreak. There is a dearth of studies that focus on neonatal infections caused by Acinetobacter for an extended period of time. In this study we investigate (i) the prevalence of Acinetobacter in cases of neonatal sepsis for a period of about 7 years, (ii) the carbapenem-resistant determinants and the genetic context of blaNDM−1 in Acinetobacter, (iii) the transmission of blaNDM−1 and (iv) association of clinical factors with episodes of sepsis caused due to NDM-1-harboring isolates. The emergence of NDM-1 in this unit makes it relevant to carry out such a study. The use of molecular methods in this study to discriminate the different species of Acinetobacter also gives an insight about the different species of Acinetobacter as a causative agent of neonatal sepsis.
Materials and Methods
Setting and Patients
The specimens were collected from the neonatal intensive care unit of the IPGMER and SSKM Hospital, Kolkata, India from 2007(January) to 2014(June). Due to some unavoidable circumstances specimens could not be collected from January 2012 to June 2012. This unit has a 16 bed Level III unit, 26 bed Level II unit and 8 bed neonatal surgical units. This unit had 1185 admissions per year (departmental census 2013), including both intramural and extramural births. Blood cultures of the neonates having sepsis were processed by previously described methods (Roy et al., 2013). In brief, 1 ml of blood for culture was drawn with aseptic precautions from a peripheral vein into Peds Plus vials. Blood culture was performed with a BACTEC9050 system (Becton Dickinson, Sparks, MD, USA) from 2008 onwards and manually prior to that. For any culture that flagged positive, Gram-staining was performed and subculture was done on appropriate media based on Gram's stain: MacConkey agar (Difco Laboratories, Detroit, MI, USA) and 5% sheep blood agar (Difco Laboratories) for Gram-negative and Gram-positive organisms, respectively. Clinical data were collected from the hospital registers.
Identification of Acinetobacter
The identity of the Acinetobacter spp. was confirmed by VITEK 2 compact system (BioMérieux, Marcy l'Etoile, France) from 2012 onwards and prior to that by the Mini API system (BioMérieux). Molecular identification of all Acinetobacter spp. was carried out by the ARDRA (amplified ribosomal DNA restriction analysis) method (Dijkshoorn et al., 1998).
Antimicrobial Susceptibility and MIC
Antimicrobial susceptibility testing was done by the VITEK 2 compact system (BioMérieux) and results were interpreted according to the custom set parameters (CLSI, 2014, FDA guidelines and natural resistance interpretation).
The MIC values (mg/L) of meropenem and ceftazidime were also determined using Etest method (AB Biodisk, Solna, Sweden) and were interpreted according to CLSI (CLSI, 2014). The MIC50 and MIC90 for meropenem and ceftazidime were calculated as the MIC at which 50 and 90% of the isolates were inhibited.
Detection of β-Lactamase and Carbapenemase Phenotypes
The production of ESBL, MBL, AmpC was detected by the commercial diagnostic tablets (Rosco Diagnostica A/S, Taastrup, Denmark) following the manufacturer's instructions. The results were interpreted as follows: an increase in ≥5 mm of inhibition zone diameter around boronic acid and dipicoloinic acid, in comparison to the diameter with meropenem alone was considered a positive result for AmpC and MBL respectively, while an increase in ≥5 mm in diameter around clavulanic acid and cloxacillin against cefotaxime alone suggests ESBL and AmpC production respectively (Giske et al., 2010).
Detection of Carbapenemases, AmpCs and ESBLs by Genotypic Method
PCR was carried out for carbapenem-resistant genes (blaVIM, IMP, SPM−1, GIM−1, SIM−1, NDM, and blaOXA−23−like, OXA−24−like, OXA−51−like, OXA−58−like) (Ellington et al., 2007; Roy et al., 2011a; Datta et al., 2014) extended-spectrum-β-lactamase genes (blaCTX−M, VEB, PER) (Cao et al., 2002; Saladin et al., 2002; Colom et al., 2003; Woodford et al., 2006) and blaAmpC genes (Perilli et al., 2007) (Table S1).
Class 1 and class 2 integrons (Corvec et al., 2003; Shibata et al., 2003), plasmid-mediated quinolone resistance genes aac(6′)-Ib-cr and aminoglycoside resistance genes (rmtA, rmtB, rmtC, rmtD, and armA) (Berçot et al., 2011) were additionally investigated in the NDM-1-producing isolates (Table S1).
Conjugation and Electro-Transformation of blaNDM−1
Conjugal transfer of blaNDM−1 to the sodium azide-resistant E. coli J53 recipient (Pasteran et al., 2014) was attempted by a solid mating assay (Walsh et al., 2011). Transconjugants were selected with varying concentrations of meropenem (0.1–2 mg/L) and 100 mg/L of sodium azide. Electro-transformation into commercially available MAX Efficiency® DH10B™ Competent Cells (Life Technologies, Carlsbad, CA, USA) was carried out for non-conjugative strains using 0.1 mg/L meropenem.
Susceptibility test for ceftazidime and meropenem was performed for the blaNDM−1 carrying transconjugants and transformants by Etest method and was followed by PCRs.
Plasmid Characterization
Plasmids harbored by the donor (blaNDM−1-harboring isolates), transconjugants or transformants were purified by Kado and Liu method, 1981 and were subsequently assessed for the number and sizes (approx) using E. coli V517 and S. flexneri 2a YSH6000 (Roy et al., 2011b) as mega-plasmid marker (Kado and Liu, 1981).
Sequencing of the Amplified Product
Amplified products of blaNDM, blaVIM, blaVEB, blaPER, blaAmpC, blaCTX−M were sequenced directly on both strands using Big dye Terminator v3.1 Cycle Sequencing Kit and analyzed with an automated sequencer (ABI 3730 DNA Analyzer, Perkin Elmer, USA). The blaNDM in some isolates could not be directly sequenced and were cloned into a TOPO TA cloning vector (Life technologies) and sequenced with the M13 primers.
Characterization of Integrons
Integrons were amplified and sequenced with primers derived from the 5′ and 3′ conserved segments (Novais et al., 2006) and submitted to the INTEGRALL database (http://integrall.bio.ua.pt) for nomenclature. Some of the large products (>1500 bp) were sequenced by primer walking technique (Sverdlov and Tatyana, 2005).
Genetic Environment of the Carbapenem-Resistant Genes
The genetic structures surrounding the blaNDM−1, were studied by PCR mapping and DNA sequencing using primers based on previously reported structures (Poirel et al., 2011; Bonnin et al., 2012; Table S1). In the NDM-1-possessing transconjugants and transformants, the proximity of the IS elements to genes were determined using combinations of the respective IS primers and the OXA-23-like and OXA-58-like primers (Lin et al., 2010).
Pulsed Field Gel Electrophoresis (PFGE)
PFGE was carried out for all isolates in a CHEF-DR III apparatus (Bio-Rad Laboratories, Hercules and CA) by ApaI enzyme (Abbo et al., 2005) (New England Biolab, Massachusetts).
When indistinguishable isolates of Acinetobacter spp. were cultured from two or more neonates treated in the NICU during an interval of up to 15 days, they were considered as a single episode of cross-transmission (Roy et al., 2010).
Statistical Analysis
The data for the 68 neonates along with all their recorded variables (clinical factors, results of tests, organisms found, antibiotic resistance patterns etc.) were entered in IBM SPSS Statistics Version 20.0 and analyzed systematically using established statistical procedures. Characteristics were described as median and standard deviation (SD) for continuous variables and as frequencies and percentages for categorical variables.
Features like sex, gestational age, birth weight, mode of delivery, use of mechanical ventilator and place of birth (intramural or extramural) that are known to be associated with sepsis were analyzed for association with episodes of sepsis caused due to NDM-1-harboring isolates. Multivariate logistic regression was used to identify the risk factors for infection by NDM-1-harboring organisms. All available clinical factors were entered into the regression simultaneously and a backward selection process was used to identify the risk factors. All relationships were evaluated at a 95% significance level.
Chi-square test of independence was carried out to determine the association of sepsis due to NDM-1 with outcome (death or discharge) and onset of sepsis (early or late). Finally, a simple t-test of proportions was used to test if the incidence of infection via Acinetobacter baumannii or other species was significantly different for each type of antibiotic resistance. All these tests were evaluated at a 95% significance level.
Result
Demographics of the Neonates
Twenty out of 68 (29.4%) neonates from whom Acinetobacter spp. was isolated were female while 46 were male (gender information was missing for 2 neonates). The median gestational age for the neonates was 32.5 weeks (SD 3.4 weeks) and median birth weight was 1363 grams (SD 828 grams). Eighty two percent (56 of 68) of the neonates were pre-term while 77.9% (53 of 68) of the neonates were of low birth weight. Sixty five percent (44 of 68) of the neonates were delivered by cesarean section while the rest had a normal vaginal delivery. Most of the neonates in this unit were born at the facility (inborn)—46 of 68 (67.6%) and the rest were born at other facilities and later transferred to this unit (out born). Sixty two percent of the neonates had early onset sepsis while 31% had late onset of sepsis (time of onset of sepsis could not be recorded with certainty for 5 neonates).
Identification of Acinetobacter spp. by ARDRA
During this period, 68 non-duplicate Acinetobacter spp. were isolated from the blood specimens of 68 septicaemic neonates. All isolates were analyzed in this study. The year-wise breakup of the isolates is depicted in Figure S1 indicating the higher prevalence of Acinetobacter calcoaceticus-baumannii complex (ACB complex) comprising of A. baumannii, A. calcoaceticus, A. nosocomialis, and A. pittii.
Each of the 68 isolates was unambiguously identified to the species level by ARDRA. The predominant species was A. baumannii (72%) followed by A. calcoaceticus (10%), A. lwoffii (6%), A. nosocomialis (3%), A. junni (3%), A. variabilis (3%), A. haemolyticus (2%), and 14TU (2%) (Table 1).
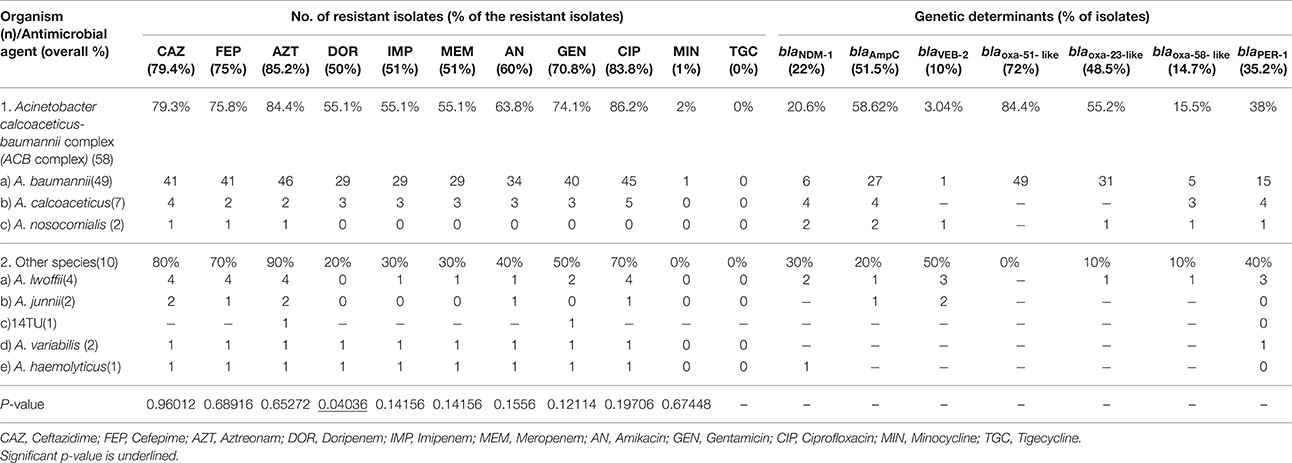
Table 1. Antibiotic susceptibility pattern (by VITEK 2 compact) and genetic determinants of Acinetobacter spp.
The identity of the isolates using the ARDRA method was compared with Mini API or VITEK 2 results. Twenty six of the 38 strains were identified as the same species when results of the mini API and ARDRA (Table S2) were compared. Similarly, 19 of the 30 isolates were assigned as the same species by VITEK 2 and ARDRA (Table S2).
Antimicrobial Susceptibility Tested by VITEK 2
Table 1 summarizes the antibiotic resistance pattern found in Acinetobacter spp. during this period. More than 75% isolates were resistant to extended spectrum cephalosporins (79% isolates were resistant to ceftazidime, 75% isolates were resistant to cefepime), 85% isolates were resistant to aztreonam, 50% were resistant to doripenem, 51% were resistant to both imipenem and meropenem. More than 60% isolates were resistant to at least one aminoglycosides (60% resistant to amikacin, 71% resistant to gentamicin). Eighty four percent isolates were resistant to ciprofloxacin. But all isolates showed high susceptibility to minocycline (99%) and tigecycline (100%).
All the Acinetobacter isolates were categorized into two groups, the ACB complex (A. baumannii, A. calcoaceticus, A. nosocomialis, and A. pittii) and other species (A. lwoffii, A. junni, A. haemolyticus, A. variabilis, and 14TU). Several analyses in this study were done where comparison between these two groups were made.
While analyzing resistance to different groups of antibiotics against these two categories, differences were noted for doripenem (Table 1). The percentage of doripenem-resistant isolates was significantly higher in ACB complex (55%) than in other species (20%) (p-value 0.0404). However, since the number of strains in the category “other species” is small (n = 10), these differences should not be overemphasized.
Distribution of MIC Values of Different Groups of Antibiotics
There was a marked increase in resistance to carbapenems among Acinetobacter spp. from 25% in 2007 to 87% in 2013 (Figure S1). The range of MIC values for meropenem and ceftazidime over this period were 0.025− ≥32 mg/L and 1− ≥256 mg/L respectively. For individual years, the range of MIC values is depicted in Table S3.
Over this period the MIC50 and MIC90 values for meropenem were 32 mg/L and for ceftazidime were 256 mg/L. Detailed year-wise distribution of MIC50 and MIC90 values (mg/L) for meropenem and ceftazidime is shown in Table S3. This result depicted a significant increase in MIC value of meropenem since 2011 whereas for ceftazidime high resistance persisted throughout this period (except for 2010) (Table S3).
Detection of Different β-Lactamases by Phenotypic Test
Of the 68 isolates, 19, 24, and 28% showed phenotypic evidence of MBLs, ESBLs and AmpCs respectively. Among MBL-producers (n = 13), 50% isolates showed ESBL and AmpC phenotype both and 64% showed only ESBL phenotype. Whereas, among non-MBL (n = 55) phenotype, 18% were ESBL producers, 27% were AmpC producers and 11% produced both of the enzymes. Presence of MBL sometimes complicates the detection of ESBLs and AmpCs. Therefore, the presence of ESBLs and AmpCs in these isolates was confirmed by PCR subsequently.
Genotypic Distribution of Various β-Lactamases
Isolates possessed different β-lactamases such as ESBLs (50%), AmpC (52%), and carbapenemases (56%, includes both oxacillinases and MBLs). Sequencing revealed the predominant presence of blaPER−1(35%), followed by blaVEB−2 and blaCTXM−15. Among the carbapenem-resistant determinants the blaOXA−23−like gene (49%) was the most common, followed by blaOXA−58−like gene (15%). Sequence analysis revealed that NDM-1 was the predominant (22%) MBL detected (Table 1). No other variant of NDM was found.
Detailed distribution of these genes in the different species has been presented in Table 1. blaNDM−1 was present in 21% of the ACB complex (A. calcoaceticus, A. baumannii, A. nosocomialis) and 30% of the other species (A. haemolyticus, A. lwoffii). In contrast, most of the oxacillinases were confined within ACB complex except 2 isolates of A. lwoffii carried either blaOXA−23−like or blaOXA−58−like gene.
While correlating the phenotypic and genotypic test results of all the strains, it was observed that one strain that could not be detected as MBL producers by phenotypic tests harbored blaNDM−1 (A_134) as detected by PCR. Similarly, there were 8 strains which did not show the presence of ESBL by the phenotypic tests but harbored blaPER−1 or blaCTXM−15. Of these 8 strains, 3 showed the presence of NDM-1. Similarly, 17 of the AmpC non-producers (as tested by the phenotypic tests) harbored blaAmpC gene and 5 of these strains also possessed NDM-1. This showed the failure to detect ESBLs and AmpCs by phenotypic tests particularly in MBL–producing strains.
Diversity of the Acinetobacter Isolates
The analyses of the pulsotypes were done separately for A. baumannii, A. calcoaceticus, and A. lwoffii. Strains of A. baumannii were primarily diverse and dissemination of any particular clone was not evident (Figure 1). One sporadic case of cross-transmission was seen: A_151 (DOB 10/01/2013) and A_153 (DOB 24/01/2013). A few pairs of strains were found to be indistinguishable but could not be considered as episodes of cross-transmission as they were isolated from neonates who were not treated in the NICU during an interval of 15 days (Figure 1). In A. calcoaceticus and A. lwoffii group no cross-transmission was noted and all the isolates were distinct.
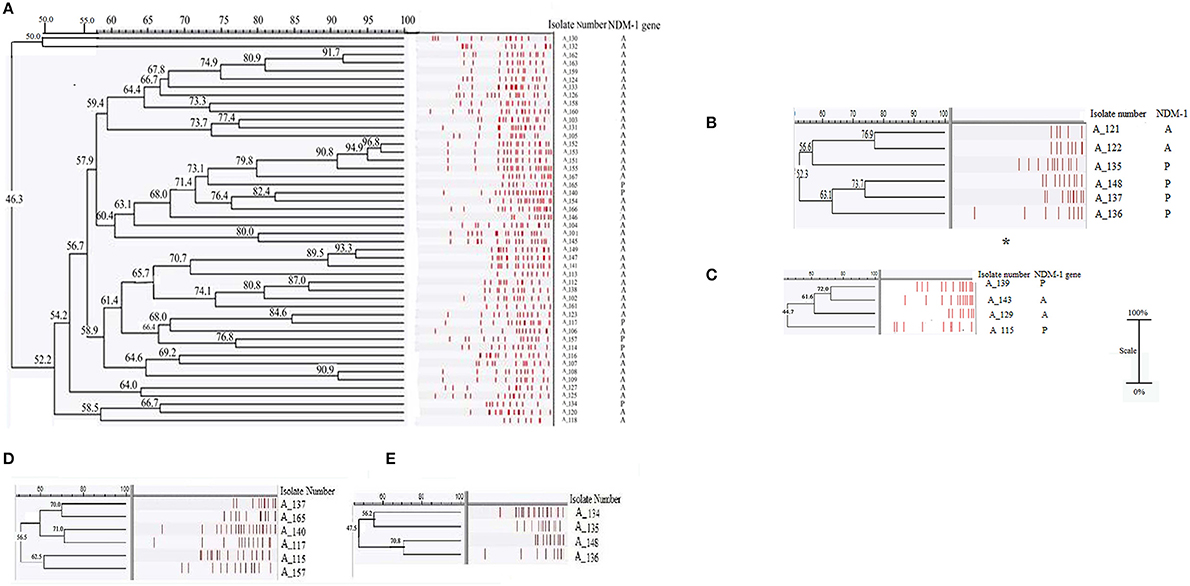
Figure 1. Analysis of genetic relationship according to Dice's similarity coefficient and the unweighted pair group method with arithmetic mean (UPGMA) (the position tolerance and optimization were set at 1.5 and 1% respectively) of the ApaI patterns of (A) A. baumannii (n = 49), (B) A. calcoaceticus (n = 6), (C) A. lwoffii (n = 4) isolates, (D) A. baumannii (n = 6), and (E) A. calcoaceticus (n = 4) harboring NDM-1. More than 90% similarity in PFGE band pattern interpreted as indistinguishable. A few pairs of strains were found to be indistinguishable but could not be considered as episodes of cross-transmission as they were isolated from neonates who were not treated in the NICU during an interval of 15 days: (i) A_152 (DOB 06/01/2013) and A_153 (DOB 24/01/2013), (ii) A_162 (DOB 5/09/2013) and A_163 (DOB 21/10/2013), (iii) A_151 (DOB 10/01/2013) and A_155 (DOB 08/03/2013), (iv) A_108 (DOB 24/08/2008) and A_109 (DOB 24/11/2008), (v) A_147 (DOB 25/01/2012) and A_149 (DOB 01/04/2012). *One A. calcoaceticus isolate was untypable. A represents absence of NDM-1, P represents presence of NDM-1.
The number of isolates in other groups (A. junni, A. nosocomialis, 14TU, A. variabilis, A. haemolyticus) was small for cluster analysis and were visually analyzed. All were diverse.
Characterization of the blaNDM−1-Harboring Isolates
Twenty two percent (n = 15) of the isolates harbored NDM-1 across all species of Acinetobacter. The presence of NDM-1 was not significantly different between the ACB complex and the other species (p-value: 0.7640). Most of the isolates (n = 11) harboring NDM-1 had an MIC value ≥32 mg/L for meropenem except for a few that had low MIC values (0.125, 0.25, 0.5, 1 mg/L) (Table 2). One of the isolates, A_117 (0.5 mg/L) had at least three carbapenem resistant determinants blaNDM−1, blaOXA−23−like and blaVIM−2gene. All NDM-1-harboring isolates were resistant to a series of antibiotics except tigecycline and colistin.
Most isolates that possessed blaNDM−1 also possessed blaOXA−23−like or blaOXA−58−like. NDM-1 possessing isolates also harbored blaVIM−2 (6%)′ blaVEB−2 (20%), blaPER−1 (60%), blaAmpC (60%), armA (47%), aac(6′)-Ib (27%). Presence of class 1 integron was observed in 8 of the NDM-1-possessing isolates; IntI2 was present in one isolate (Table 2).
PFGE of the A. baumannii and A. calcoaceticus isolates possessing NDM-1 revealed that diverse clones of these species harbored NDM-1(Figure 1). No cases of cross-transmission were noted in the NDM-1 isolates.,
Characterization of the blaNDM−1-Harboring Transconjugants
Fourteen of the 15 NDM-1 harboring isolates were able to transfer blaNDM−1 by conjugation (13/15) or transformation (1/15) to the recipients. Fifty percent of the transconjugants showed MIC values of ≥3 mg/L for meropenem and 79% of the isolates were also resistant to ceftazidime (MICs ≥ 8 mg/L). One transformant (A_139-DH10B/EC) was sensitive to both meropenem (0.023 mg/L) and ceftazidime (0.5 mg/L) (Table 3). A single plasmid was detected in all of the blaNDM−1-harboring transconjugants and the sizes of these plasmids ranged between 86 and 213 kb (Table 3).
Investigation of the immediate genetic environment of blaNDM−1 gene revealed the presence of a conserved structure that always associates with the complete transposon, Tn125. This structure starts with one copy of the ISAba125 at the 5′ end of blaNDM−1 gene and bleMBL gene followed by trpF, tat, cutA1, groES, groEL, insE, and ISAba125 at the 3′ end. All strains showed similar genetic structure surrounding the blaNDM−1. As the primer walking and PCR results were consistent for all transconjugants, sequences were thoroughly analyzed for 4 isolates (Figure 2).
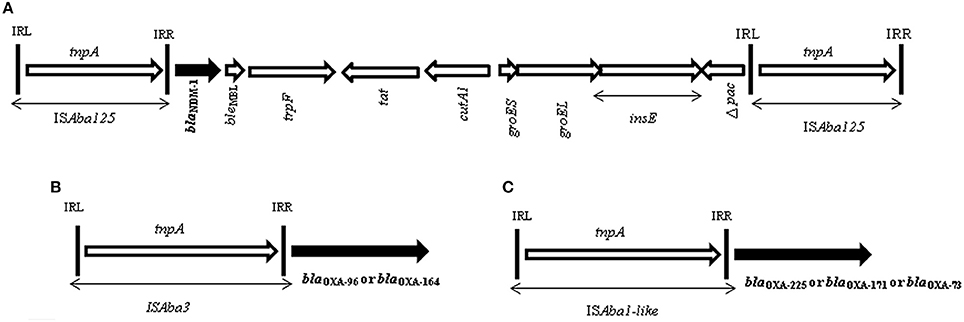
Figure 2. (A) Schematic representation of Tn125 carrying blaNDM−1 gene in a representative transconjugant. Genes and their transcription orientation are indicated by arrows. The lengths of the target genes and their exact location of the target genes are not to scale. Gene names are abbreviated according to the corresponding proteins (Bonnin et al., 2012): cutA1 for divalent cation tolerence protein; groES, groEL for heat-chaperonin protein; insE for ISCR21 of tnpA family. Δ pac for truncated phospholipid acetyltransferase. IRL and IRR are for inverted repeat left and right, respectively. This structure was found in all of the transcojugants (except one transformant). (B) Diagram showing the linkage between blaOXA−58−like with ISAba3 (A_114-J53, A_136-J53, and A_156-J53). (C) Diagram showing the linkage between blaoxa−23−like with ISAba1 (A_134-J53, A_140-J53, and A_148-J53).
PCRs were also carried out to map the position of the respective IS elements with the blaOXA−23−like and blaOXA−58−like genes in the transconjugants harboring NDM-1. Among the NDM-1-harboring transconjugants, three possessed blaOXA−23−like and three others harbored blaOXA−58−like gene (Table 3). All transconjugants with blaOXA−23−likewere linked to ISAba1 and all three isolates with blaOXA−58−likewere linked to ISAba3 (Figure 2). Sequence analysis revealed the presence of different variants of OXA-23-like enzymes, i.e., OXA-73 (A_134), OXA-225 (A_140), and OXA-171(A_148). Similarly, blaOXA−58−like genes were identified as OXA-96 (A_114, A_156) and OXA-164 (A_136) (Table 3).
Analysis of the integrons revealed different combinations of resistance genes bracketed between the 5′ and 3′ conserved region and each of these different resistance gene cassettes are presented in Figure 3. Most commonly found genes in this conserved segment were aminoglycoside resistance (aadB, aadA2, aac4′), non-enzymatic chloramphenicol resistance gene (cmlA1g) and ADP-ribosylation gene (arr2, arr3).
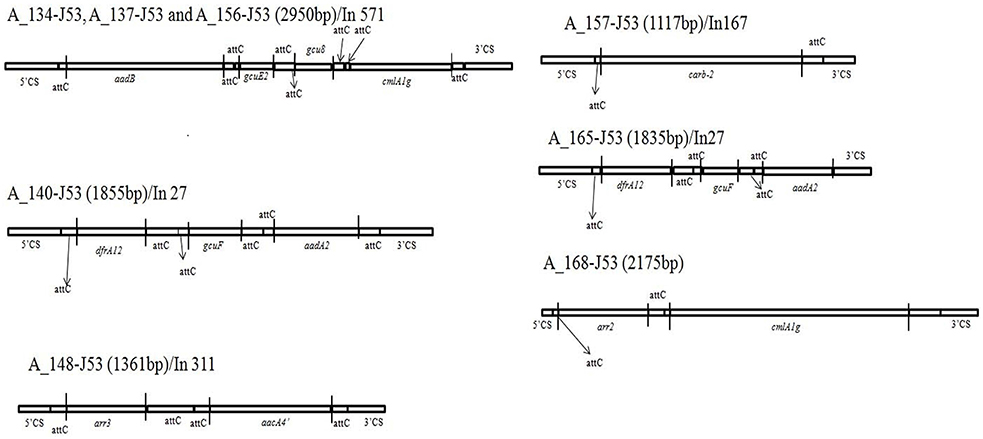
Figure 3. Schematic diagram of genes in between 5′CS and 3′CS of class 1 integron of 8 transconjugants of Acinetobacter spp. Three of the transconjugants (A_134-J53, A_137-J53, and A_156-J53) share similar structure. The genes are not to scale and vertical lines represent the boundaries of each gene. Gene names are abbreviated according to their corresponding proteins: aadB, aadA2, aacA4′− resistance to trimethoprim, gcuF, gcuE2, gcu8- unknown ORF, arr3, and arr2- ADP-ribosylation gene, carb-2- resistance to β-lactamases. attC site- attachment sites of each gene.
Association of Clinical Factors and Mortality of the Neonates with the Presence of NDM-1-Harboring Acinetobacter
The results of a comparison of neonates with sepsis caused due to NDM-1-harboring Acinetobacter with those without NDM-1 is shown in Table 4. Multivariate logistic regression established that neonates who were outborn had a significantly higher incidence of NDM-1 harboring isolates than their inborn counterparts [Odds Ratio (OR) 2.39; p-value <0.001]. None of the other factors were found to be statistically significant.
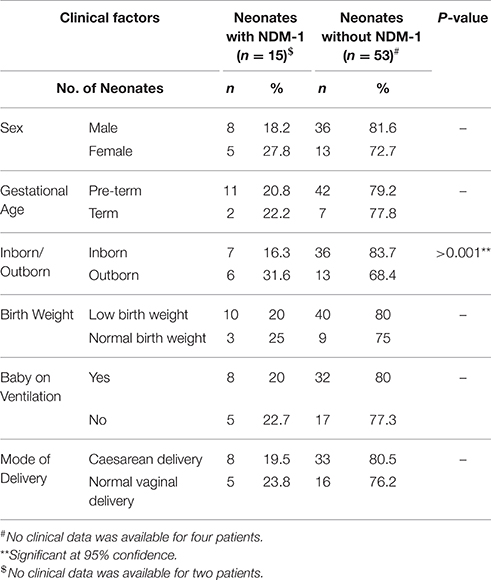
Table 4. Association of clinical factors with sepsis caused by NDM-1-harboring Acinetobacter spp. in neonates.
Incidence of NDM-1 in late onset sepsis cases (38.1%) was significantly higher than those in early onset sepsis (12.2%) [OR 4.43; p < 0.0177].Finally, the association analysis between NDM-1-harboring Acinetobacter isolates and mortality (18.8% in death cases vs. 21.7% in discharge cases), showed no statistically significant difference [p-value 0.8003].
Discussion
The data show that the potential impact of Acinetobacter sepsis on neonates in a tertiary care center is significant. There has been an increase of sepsis due to lactose non-fermenting Gram-negative bacteria in recent years and Acinetobacter is an important contributor to this burden (Viswanathan et al., 2014). This study also shows that though A. baumannii is still by far the most common of the Acinetobacter species causing sepsis in this unit, other species are also emerging in the clinical settings. Though infections due to other species of Acinetobacter has been reported (Turton et al., 2011; Tien et al., 2012), infection in the neonatal population caused by species of Acinetobacter other than A. baumannii are rare. The use of the ARDRA method in this study has helped in the elucidation of the species providing an insight about the presence of the other species of Acinetobacter in cases of neonatal septicaemia.
Another issue that emerges from the study is drug resistance. Carbapenem resistance is high and the clinician is left with very little choice of antibiotics. The only antibiotic that still retains activity is tigecycline, an observation that has also been made by other authors (Golanbar et al., 2011; Hsieh et al., 2014). In this unit, piperacillin/tazobactum and amikacin are being used as a pre-emptive (first line) therapy for clinically suspected cases of sepsis. As a second line of therapy ofloxacin and amikacin are used in the unit. In rare cases where the above therapies are ineffective and the antibiogram results are still awaited, meropenem is used.
Further, this study also shows that the phenotypic detection of ESBLs and AmpCs in presence of MBLs is challenging. The failure to detect the ESBLs in the MBL–producing clinical isolates may lead to the hidden spread of such β-lactamases complicating the situation even further. The authors have recently devised a method to circumvent this problem (Datta et al., 2015).
Resistance to carbapenems and cephalosporins could be correlated to the diverse repertoire of genetic determinants that were identified. The most prevalent were the oxacillinases. Among the transmissible oxacillinases, OXA-23-like was the most common and this result is confirmatory of other studies (Woodford et al., 2006; Golanbar et al., 2011). The presence of the oxacillinases was significantly higher in the ACB complex compared to the other Acinetobacter species. The presence of ESBLs like PER-1, VEB-2 and CTXM-15 were also noted along with the carbapenemases in some isolates but compared to the carbapenemases their numbers were lower (Ben et al., 2011). Of the class B β-lactamases, only NDM-1 and VIM-2 were identified. Most earlier reports of NDM-1 or a variant of the gene in clinical settings have been in A. baumannii (Wang et al., 2007; Boulanger et al., 2012; Revathi et al., 2013) but the presence of NDM-1 in this study was noted across other species of Acinetobacter along with A. baumannii.
PFGE revealed that NDM-1 harboring isolates were diverse. Acinetobacter spp. possessing NDM-1 was isolated predominantly from neonates delivered at other (extramural) centers reflecting the dissemination of this gene but not any clone in particular. A study with A. baumannii isolates collected across Europe also reported diversity of clones in Acinetobacter infection (Bonnin et al., 2012). Studies that have reported the presence of NDM-1 in other genuses have also described similar diversity of clones (Pasteran et al., 2014).
The blaNDM−1 gene was organized in a composite transposon bracketed between two copies of ISAba125 in all the isolates irrespective of the species. This arrangement of the gene in Acinetobacter has been reported earlier (Pfeifer et al., 2011; Bonnin et al., 2012; Fu et al., 2012). The similar organization of the blaNDM−1 in the diverse clones of A. baumannii and also the other species lends credence to the hypothesis that the current dissemination of the gene in Acinetobacter is likely linked to Tn125 and not plasmids. The linkage of blaOXA−23−like and blaOXA−58−like to ISAba1 and ISAba3 respectively as observed here, has also been reported in several earlier studies. This shows that the genetic context of these genes in Acinetobacter remain similar even in geographically distant places (Chen et al., 2010; Lin et al., 2010).
Analysis of the class I integron showed that though there was variability in the genetic determinants present between the 5′ and 3′ CS segments, most of the genes were not associated with carbapenem or cephalosporin resistance (except carb-2/A_157).
A few isolates harboring NDM-1 and oxacillinases had low MIC values (even as low as 0.125 mg/L in one isolate). Similar observation was also reported from earlier studies (Datta et al., 2014; Zmarlicka et al., 2015). Further work is needed to understand the reason for such differences which can occur due to alterations in the expression of the enzymes or other changes in the mechanism of resistance toward β-lactamases.
Statistical analysis indicated that a neonate born in other facilities (outborn) had a significantly higher incidence of NDM-1 harboring isolates than neonates born at this center. This indicates the possibility of the neonates having acquired NDM-1-harboring isolates from other hospitals where they were born. This is supported by the fact that most had a late onset of sepsis, the initial infection probably having acquired from the other hospitals. The clonal diversity of the isolates also lends support to this possibility.
This study documents the significance of both Acinetobacter baumannii and the other species, in cases of neonatal sepsis in a unit over a period of 7 years. We have been able to gain a unique assessment of the diverse genetic determinants responsible for carbapenem resistance in these isolates. The genetic context of the carbapenem-resistant genes also provides insight about the importance of the ISAba125 in the spread of NDM-1 in Acinetobacter. The emergence of NDM-1 among an already existing repertoire of oxacillinases is a challenge for clinicians and microbiologists alike. As new resistance mechanisms constantly evolve, both laboratory detection systems and infection control measures need to be enhanced.
Ethical Approval
The study protocol was carefully reviewed and approved by the Institutional Ethics Committee of the National Institute of Cholera and Enteric Diseases (Indian Council of Medical Research) (No. A-1/2013-IEC, dated 10th January 2014). Individual informed consent was waived because this study used currently existing sample collected during the course of routine diagnosis of sepsis and did not pose any additional risks to the patients. The patient records/information was anonymized and de-identified prior to analysis.
Author Contributions
Conceived and designed the experiments: SB. Performed the experiments: SC, SD, SR. Analyzed the data: SC, SD, SR, LR, SB. Coordinated collection of specimens, maintenance of clinical data: AS, RV, TS. Contributed reagents/materials/analysis tools: AS, RV, TS, LR, SB. Contributed to the writing of the manuscript: SC, LR, SB.
Funding
The study was supported by Indian Council of Medical Research (ICMR) intramural fund. SC and SD were supported by senior research fellowships from ICMR and SR received a postdoctoral fellowship from ICMR.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We extend our thanks to Dr. Arun K. Singh and the staff of the Department of Neonatology for their support during the study. We also thank the technical staff of central facility of Bose Institute, Kolkata for sequencing data. Our sincere thanks to Dr. Simona Pollini, Dr. George A. Jacoby, Dr. Anne Marie Queenan, Dr. Olivier Moquet, and Dr. Kyungwon Lee for providing us the control strains for PCRs.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2016.01126
Figure S1. Distribution of Acinetobacter spp. from 2007 to 2014. Isolates were collected for 6 months in 2012 and till June in 2014.
Table S1. Primers used in this study.
Table S2. Comparison of identity of species by Mini API (a)/VITEK 2 compact (b) and ARDRA.
Table S3. Year wise distribution of MIC50, MIC90, and range of MIC values for meropenem and ceftazidime.
References
Abbo, A., Venezia, S. N., Muntz, H. O., Krichali, T., Siegman, I. Y., Carmeli, Y., et al. (2005). Multidrug-Resistant Acinetobacter baumannii. Emerg. Infect. Dis. 11, 22–29. doi: 10.3201/eid1101.040001
Beaufort, A. J. D., Bernards, A. T., Dijkshoorn, L., and Boven, C. P. A. (1999). Acinetobacter junii causes life-threatening sepsis in preterm infants. ActaPñdiat 88, 772–775. doi: 10.1111/j.1651-2227.1999.tb00041.x
Ben, R., Yang, M., Hsueh, J., Shiang, J. C., and Chien, S. T. (2011). Molecular characterization of multiple drug-resistant Acinetobacter baumanni isolates in southern Taiwan. Int. J. Antimicrob. Agents 38, 403–408. doi: 10.1016/j.ijantimicag.2011.05.014
Berçot, B., Poirel, L., and Nordmann, P. (2011). Updated multiplex polymerase chain reaction for detection of 16S rRNA methylases: high prevalence among NDM-1 producers. Diagn. Microbiol. Infect. Dis. 71, 442–445. doi: 10.1016/j.diagmicrobio.2011.08.016
Bonnin, R. A., Poirel, L., Naas, T., Pirs, M., Seme, K., Schrenzel, J., et al. (2012). Dissemination of New Delhi metallo-β-lactamase-1-producing Acinetobacter baumannii in Europe. Clin. Microbiol. Infect. 18, E362–E365. doi: 10.1111/j.1469-0691.2012.03928.x
Boulanger, A., Naas, T., Fortineau, N., Figueiredo, S., and Nordmann, P. (2012). NDM-1-producing Acinetobacter baumannii from Algeria. Antimicrob. Agents Chemother. 56, 2214–2215. doi: 10.1128/AAC.05653-11
Cao, V., Lambert, T., Nhu, D. Q., Loan, H. K., Hoang, N. K., Arlet, G., et al. (2002). Distribution of extended-spectrum-β-lactamases in clinical isolates of Enterobacteriaceae in Vietnam. Antimicrob. Agents Chemother. 46, 3739–3743. doi: 10.1128/AAC.46.12.3739-3743.2002
Chan, P. C., Huang, L. M., Lin, H. C., Chang, L. Y., Chen, M. L., Liu, C. Y., et al. (2007). Control of an outbreak of pandrug-resistant Acinetobacter baumannii colonization and infection in a neonatal intensive care unit. Infect. Control Hosp. Epidemiol. 28, 423–429. doi: 10.1086/513120
Chen, T. L., Chang, W. C., Kuo, S. C., Lee, Y. T., Chen, C. P., Kei, S. L., et al. (2010). Contribution of a plasmid-borne blaoxa−58 gene with its hybrid promoter provided by IS1006 and an ISAba3-like element to β-lactam resistance in Acinetobacter genomic species 13TU. Antimicrob. Agents Chemother. 54, 3107–3112. doi: 10.1128/AAC.00128-10
Chuanfu, Z., Shaofu, Q., Yong, W., Qi, L., Hao, R., Liu, X., et al. (2013). Higher Isolation of NDM-1 Producing Acinetobacter baumannii from the sewage of the hospitals in Beijing. PLoS ONE 8:e64857. doi: 10.1371/journal.pone.0064857
Clinical Laboratory Standards Institute (2014). Performance Standards for Antimicrobial Susceptibility Testing: Eighteenth Informational Supplement M100-S18. Wayne, PA: CLSI.
Colom, K., Pérez, J., Alonso, R., Fernández, A. A., Lariño, E., and Cisterna, R. (2003). Simple and reliable multiplex PCR assay for detection of blaTEM, blaSHV and blaOXA−1 genes in Enterobacteriaceae. FEMS Microbiol. Lett. 223, 147–151. doi: 10.1016/S0378-1097(03)00306-9
Corvec, S., Caroff, N., Espaze, E., Giraudeau, C., Drugeon, H., and Reynaud, A. (2003). AmpC cephalosporinase hyperproduction in Acinetobacter baumannii clinical strains. J. Antimicrob. Chemother. 52, 629–635. doi: 10.1093/jac/dkg407
Datta, S., Chatterjee, S., Mitra, S., and Basu, S. (2015). A reliable phenotypic assay for detection of ESBLs and AmpCs in MBL-producing gram-negative bacteria with the use of amino-phenyleboronic acid, dipicolonic acid and cloxacillin. J. Microb. Methods. 115, 100–103. doi: 10.1016/j.mimet.2015.05.028
Datta, S., Roy, S., Chatterjee, S., Saha, A., Sen, B., Pal, T., et al. (2014). A five-year experience of carbapenem resistance in Enterobacteriaceae causing neonatal septicaemia: predominance of NDM-1. PLoS ONE 9:e112101. doi: 10.1371/journal.pone.0112101
Dijkshoorn, L., Harsselaar, B. V., and Tjernberg, I. (1998). Evaluation of Amplified Ribosomal DNA restriction analysis for identification of Acinetobacter genomic species. System. Appl. Microbiol. 21, 33–39. doi: 10.1016/S0723-2020(98)80006-4
Dortet, L., Poirel, L., and Nordmann, P. (2014). Worldwide Dissemination of the NDM-Type carbapenemases in gram-negative bacteria. Biomed. Res. Int. 2014:249856. doi: 10.1155/2014/249856
Ellington, M. J., Kistler, J., Livermore, D. M., and Woodford, N. (2007). Multiplex PCR for rapid detection of genes encoding acquired metallo-β-lactamases. J. Antimicrob. Chemother. 59, 321–322. doi: 10.1093/jac/dkl481
Espinal, P., Fugazza, G., López, M. Y., Kasma, M., Lerman, Y., Malhotra, K. S., et al. (2011). Dissemination of an NDM-2-producing Acinetobacter baumanni clone in an Israeli Rehabilitation Centre. Antimicrob. Agents Chemother. 55, 5396–5398. doi: 10.1128/AAC.00679-11
Fu, Y., Du, X., Ji, J., Jiang, Y., and Yu, Y. (2012). Epidemiological characteristics and genetic structure of blaNDM−1 in non-baumannii Acinetobacter spp. in China. J. Antimicrob. Chemother. 67, 2114–2122. doi: 10.1093/jac/dks192
Giske, C. G., Gezelius, L., Samuelsen, Ø., Warner, M., Sundsfjord, A., and Woodford, N. (2010). A sensitive and specific phenotypic assay for detection of metallo-β-lactamases and KPC in Klebsiella pneumoniae using meropenem discs supplied with boronic acid, dipicolinic acid and cloxacillin. Clin. Microbiol. Infect. 17, 552–556. doi: 10.1111/j.1469-0691.2010.03294.x
Golanbar, G. D., Lam, C. K., Chu, Y., Cueva, C., Tan, S. W., Silva, I., et al. (2011). Phenotypic and molecular characterization of Acinetobacter clinical isolates obtained from inmates of California correctional facilities. J. Clin. Microbiol. 49, 2121–2131. doi: 10.1128/JCM.02373-10
Hammerum, A. M., Hansen, F., Skov, M. N., Stegger, M., Andersen, P. S., Holm, A., et al. (2015). Investigation of a possible outbreak of carbapenem-resistant Acinetobacter baumannii in Odense, Denmark using PFGE, MLST and whole-genome-based SNPs. J. Antimicrob. Chemother. 70, 1965–1968. doi: 10.1093/jac/dkv072
Hsieh, W. S., Wang, N. Y., Feng, J. A., Weng, L. C., and Wu, H. H. (2014). Types and Prevalence of carbapenem-resistant Acinetobacter calcoaceticus-Acinetobacter baumanni complex in Northern Taiwan. Antimicrob. Agents Chemother. 58, 201–204. doi: 10.1128/AAC.00779-13
Hu, H., Hu, Y., Pan, B. Y., Liang, H., Wang, H., Wang, X., et al. (2012). Novel plasmid and its variant harbouring both a blaNDM−1 gene and type IV secretion system in clinical isolates of Acinetobacter lwoffii. Antimicrob. Agents Chemother. 56, 1698–1702. doi: 10.1128/AAC.06199-11
Huang, Y. C., Su, L. H., Wu, T. L., Hsieh, W. S., Chang, T. M., and Lin, T. Y. (2002). Outbreak of Acinetobacter baumanni bacteraemia in a neonatal intensive care unit: clinical implications and genotyping analysis. Pediatr. Infect. Dis. J. 21, 1105–1109. doi: 10.1097/00006454-200212000-00004
Jiang, X., Ni, Y., Jiang, Y., Yuan, F., Han, L., Li, M., et al. (2005). Outbreak of infection caused by Enterobacter cloacae producing the novel VEB-3 ß-lactamase in China. J. Clin. Microbiol. 43, 826–831. doi: 10.1128/JCM.43.2.826-831.2005
Kado, C. I., and Liu, S. T. (1981). Rapid procedure for detection and isolation of large and small plasmids. J. Bacteriol. 145, 1365–1373.
Kilic, A., Li, H., Mellmann, A., Basustaoglu, A. C., Kul, M., Senses, Z., et al. (2008). Acinetobacter septicus sp. nov. association with a nosocomial outbreak of bacteremia in a neonatal intensive care unit. J. Clin. Microbiol. 46, 902–908. doi: 10.1128/JCM.01876-07
Kim, E. S., Jeong, J. Y., Jun, J. B., Choi, S. H., Lee, S. O., Kim, M. N., et al. (2009). Prevalence of aac(6')-Ib-cr encoding a ciprofloxacin modifying enzyme among Enterobacteriaceae blood isolates in Korea. Antimicrob. Agents Chemother. 53, 2643–2645. doi: 10.1128/AAC.01534-08
Lin, Y. C., Hsia, K. C., Chen, Y. C., Sheng, W. H., Chang, S. C., Liao, M. H., et al. (2010). Genetic basis of multidrug resistance in Acinetobacter clinical isolates in Taiwan. Antimicrob. Agents Chemother. 54, 2078–2084. doi: 10.1128/AAC.01398-09
Nordmann, P., Poirel, L., and Walsh, T. R. (2011). The emerging NDM carbapenemases. Trends Microbiol. 19, 588–595. doi: 10.1016/j.tim.2011.09.005
Novais, A., Cantón, R., Valverde, A., Machado, E., Galán, J. C., Peixe, L., et al. (2006). Dissemination and persistence of blaCTXM−9 are linked to class 1integrons containing CR1 associated with defective transposon derivatives from Tn402 located in early antibiotic resistance plasmids of IncHI2, IncP1-α, and IncFI groups. Antimicrob. Agents Chemother. 50, 2741–2750. doi: 10.1128/AAC.00274-06
Pasteran, F., Mora, M. M., Albornoz, E., Faccone, D., Franco, R., Ortellado, J., et al. (2014). Emergence of genetically unrelated NDM-1-producing Acinetobacter pitti strains in Paraguay. J. Antimicrob. Chemother. 69, 2575–2578. doi: 10.1093/jac/dku139
Perilli, M., Santis, F. D., Mugnaioli, C., Rossolini, G. M., Luzzaro, F., Stefani, S., et al. (2007). Spread of Enterobacteriaceae carrying the PER-1 extended-spectrum-β-lactamase gene as a chromosomal insert: a report from Italy. J. Antimicrob. Chemother. 59, 323–324. doi: 10.1093/jac/dkl487
Pfeifer, Y., Matten, J., and Rabsch, W. (2009). Salmonella enterica serovar Typhi with CTX-M ß-lactamase, Germany. Emerg. Infect. Dis. 15, 1533–1535. doi: 10.3201/eid1509.090567
Pfeifer, Y., Wilharm, G., Zander, E., Wichelhaus, T. A., Göttig, S., Hunfeld, K. P., et al. (2011). Molecular characterization of blaNDM−1 in an Acinetobacter baumannii strain isolated in Germany in 2007. J. Antimicrob. Chemother. 66, 1998–2001. doi: 10.1093/jac/dkr256
Poirel, L., Bonnin, R. A., Boulanger, A., Schrenzel, J., Kaase, M., and Nordmann, P. (2011). Tn125-related acquisition of blaNDM-like genes in Acinetobacter baumannii. Antimicrob. Agents Chemother. 56, 1087–1089. doi: 10.1128/AAC.05620-11
Revathi, G., Siu, L. K., Lu, P. L., and Huang, L. Y. (2013). First report of NDM-1-producing Acinetobacter baumannii in East Africa. Int. J. Infect. Dis. 17, E1255–E1258. doi: 10.1016/j.ijid.2013.07.016
Roy, S., Datta, S., Viswanathan, R., Singh, A. K., and Basu, S. (2013). Tigecycline susceptibility in Klebsiella pneumoniae and Escherichia coli causing neonatal septicaemia (2007-10) and role of an efflux pump in tigecycline non-susceptibility. J. Antimicrob. Chemother. 68, 1036–1042. doi: 10.1093/jac/dks535
Roy, S., Singh, A. K., Viswanathan, R., Nandy, R. K., and Basu, S. (2011b). Transmission of imipenem resistance determinants during the course of an outbreak of NDM-1 Escherichia coli in a sick newborn care unit. J. Antimicrob. Chemother. 66, 2273–2280. doi: 10.1093/jac/dkr376
Roy, S., Viswanathan, R., Singh, A., Das, P., and Basu, S. (2010). Gut colonization by multidrug-resistant and carbapenem-resistant Acinetobacter baumannii in neonates. Eur. J. Clin. Microbiol. Infect. Dis. 29, 1495–1500. doi: 10.1007/s10096-010-1030-z
Roy, S., Viswanathan, R., Singh, A. K., Das, P., and Basu, S. (2011a). Sepsis in neonates due to imipenem resistant Klebsiella pneumoniae producing NDM-1 in India. J. Antimicrob. Chemother. 66, 1411–1413. doi: 10.1093/jac/dkr068
Saladin, M., Cao, V. T., Lambert, T., Donay, J. L., Herrmann, J. L., Ould-Hocine, Z., et al. (2002). Diversity of CTX-M β-lactamases and their promoter regions from Enterobacteriaceae isolated in three Parisian hospitals. FEMS Microbiol. Lett. 209, 161–168. doi: 10.1016/s0378-1097(02)00484-6
Shibata, N., Doi, Y., Yamane, K., Yagi, T., Kurokawa, H., Shibayama, K., et al. (2003). PCR Typing of genetic determinants for metallo-β-lactamases and integrases carried by gram-negative bacteria isolated in japan, with focus on the Class 3 Integron. J. Clin. Microbiol. 41, 5407–5413. doi: 10.1128/JCM.41.12.5407-5413.2003
Sverdlov, F., and Tatyana, A. (2005). “Primer walking,” in Encyclopaedia of Life Sciences (John Wiley & Sons, Ltd., Press). doi: 10.1038/npg.els.0005382
Tien, N., You, B., Chang, H., Lin, H. S., Chin, Y. L., Chung, T. C., et al. (2012). Comparison of genospecies and antimicrobial resistance profiles of isolates in the Acinetobacter calcoaceticus- Acinetobacter baumannii complex from various clinical specimens. Antimicrob. Agents Chemother. 56, 6267–6271. doi: 10.1128/AAC.01304-12
Turton, J. F., Shah, J., Ozongwu, C., and Pik, R. (2011). Incidence of Acinetobacter species other than A. baumannii among clinical isolates of Acinetobacter: evidence for emerging species. J. Clin. Microbiol. 48, 1445–1449. doi: 10.1128/JCM.02467-09
Vaneechoutte, M., Dijkshoorn, L., Tjernberg, I., Elaichouni, A., Vos, P. D., Claeys, G., et al. (1995). Identification of Acinetobacter genomic species, by amplified ribosomal DNA restriction analysis. J. Clin. Microbiol. 33, 11–15.
Viswanathan, R., Singh, A. K., Basu, S., Chatterjee, S., Roy, S., and Isaacs, D. (2014). Multi-drug-resistant, non-fermenting, gram negative bacilli in neonatal sepsis in Kolkata, India: a 4-year study. Paediatr. Int. Child Health. 34, 56–59. doi: 10.1179/2046905513Y.0000000072
Walsh, T. R., Weeks, J., Livermore, D. M., and Toleman, M. A. (2011). Dissemination of NDM-1 positive bacteria in the New Delhi environment and its implications for human health: an environmental point prevalence study. Lancet Infect. Dis. 11, 355–362. doi: 10.1016/S1473-3099(11)70059-7
Wang, H., Guo, P., Sun, H., Wang, H., Yang, Q., Chen, M., et al. (2007). Molecular epidemiology of clinical isolates of carbapenem-resistant Acinetobacter spp. from Chinese hospitals. Antimicrob. Agents Chemother. 51, 4022–4028. doi: 10.1128/AAC.01259-06
Woodford, N., Ellington, M. J., Coelho, J. M., Turton, J. F., Ward, M. E., Brown, S., et al. (2006). Multiplex PCR for genes encoding prevalent OXA carbapenemases in Acinetobacter spp. Int. J. Antimicrob. Agents. 27, 351–353. doi: 10.1016/j.ijantimicag.2006.01.004
Keywords: Acinetobacter spp., ARDRA, NDM-1, OXA-23, Tn125, 5′–3′ CS, neonatal sepsis, India
Citation: Chatterjee S, Datta S, Roy S, Ramanan L, Saha A, Viswanathan R, Som T and Basu S (2016) Carbapenem Resistance in Acinetobacter baumannii and Other Acinetobacter spp. Causing Neonatal Sepsis: Focus on NDM-1 and Its Linkage to ISAba125. Front. Microbiol. 7:1126. doi: 10.3389/fmicb.2016.01126
Received: 23 March 2016; Accepted: 06 July 2016;
Published: 08 August 2016.
Edited by:
Yuji Morita, Aichi Gakuin University, JapanReviewed by:
Giovanni Gherardi, Università Campus Bio-Medico, ItalyAyush Kumar, University of Manitoba, Canada
Paul G. Higgins, University of Cologne, Germany
Jean-Philippe Lavigne, University Hospital, France
Copyright © 2016 Chatterjee, Datta, Roy, Ramanan, Saha, Viswanathan, Som and Basu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Sulagna Basu, c3VwYWJhc3VAeWFob28uY28uaW4=
†Present Address: Rajlakshmi Viswanathan, Diagnostic Virology, National Institute of Virology, Pune, India
 Somdatta Chatterjee
Somdatta Chatterjee Saswati Datta
Saswati Datta Subhasree Roy
Subhasree Roy Lavanya Ramanan2
Lavanya Ramanan2 Sulagna Basu
Sulagna Basu