- College of Urban and Environmental Sciences, Peking University, Beijing, China
Syntrophic methanogenesis is an essential link in the global carbon cycle and a key bioprocess for the disposal of organic waste and production of biogas. Recent studies suggest direct interspecies electron transfer (DIET) is involved in electron exchange in methanogenesis occurring in paddy soils, anaerobic digesters, and specific co-cultures with Geobacter. In this study, we evaluate the possible involvement of DIET in the syntrophic oxidation of butyrate in the enrichments from two lake sediments (an urban lake and a natural lake). The results showed that the production of CH4 was significantly accelerated in the presence of conductive nanoscale Fe3O4 or carbon nanotubes in the sediment enrichments. Observations made with fluorescence in situ hybridization and scanning electron microscope indicated that microbial aggregates were formed in the enrichments. It appeared that the average cell-to-cell distance in aggregates in nanomaterial-amended enrichments was larger than that in aggregates in the non-amended control. These results suggested that DIET-mediated syntrophic methanogenesis could occur in the lake sediments in the presence of conductive materials. Microbial community analysis of the enrichments revealed that the genera of Syntrophomonas, Sulfurospirillum, Methanosarcina, and Methanoregula were responsible for syntrophic oxidation of butyrate in lake sediment samples. The mechanism for the conductive-material-facilitated DIET in butyrate syntrophy deserves further investigation.
Introduction
Methane (CH4) is not only an important greenhouse gas, but also a well-known component of biogas, which is used as fuel. Methanogens, which obtain energy for growth by utilizing a few simple substrates (such as H2, formate, and acetate), are widespread in anoxic environments (Thauer et al., 2008). Because methanogens are able to use only a limited number of substrates, they depend on syntrophic cooperation for decomposing complex organic compounds in anoxic environments such as rice paddies, wetlands, fresh water sediments, anaerobic bioreactors, and the intestinal tracts of animals and insects (Stams and Plugge, 2009; Sieber et al., 2012).
It is well recognized that interspecies electron transfer (IET) via H2 or formate diffusion occurs between proton-reducing acetogens and methanogens. IET has been considered a bottleneck step in the methanogenic decomposition of complex organic substances. Recently it was proposed that direct interspecies electron transfer (DIET) is an alternative to the interspecies H2/formate transfer in syntrophic methanogenesis (Morita et al., 2011; Rotaru et al., 2014b). The DIET-mediated syntrophic methanogenesis has been demonstrated in co-cultures of Geobacter metallireducens and Methanosaeta (Rotaru et al., 2014b) or Methanosarcina species (Rotaru et al., 2014a). The outer surface c-type cytochromes and electrically conductive pili are considered to play a role in mediating DIET when Geobacter species are involved (Lovley, 2012; Shrestha and Rotaru, 2014).
In addition to biogenic mediators, naturally occurring conductive or semiconductive materials can also facilitate DIET in syntrophic methanogenesis. Magnetite (Fe3O4), a mineral containing Fe(II) and Fe(III) in a ratio of 1:2 which originates either biologically (microbially) or geologically is ubiquitous in environments like weathered soils and freshwater sediments (Skomurski et al., 2010; Liu et al., 2015). This mineral is electrically conductive. Kato et al. (2012) showed that the supplementation of Fe3O4 in rice paddy soil significantly accelerated CH4 production from ethanol and acetate, and that Geobacter and Methanosarcina species were significantly enriched (Kato et al., 2012). Several other studies suggested that the supplementation with artificial electrically conductive materials (e.g., granular activated carbon, biochar) could also promote DIET-mediated syntrophic methanogenesis in defined co-cultures of Geobacter with Methanosarcina species (Liu et al., 2012; Chen et al., 2014; Rotaru et al., 2014a). A recent study suggested that magnetite could substitute for the pilus-associated cytochrome OmcS of G. sulfurreducens to mediate cell-to-cell electron transfer (Liu et al., 2015). Using of the natural or artificial nanomaterials as electron conductors to accelerate syntrophic CH4 production may help microbes to save energy for the biosynthesis of the extracellular, biological and electrical connections.
Butyrate is one of the most important intermediates in the transformation of complex organic matter to CH4. DIET was not expected to occur in the syntrophic oxidation of butyrate because recognized butyrate syntrophs were not known to possess genetically determined extracellular electron transfer components (Sieber et al., 2012). But a recent study showed that the syntrophic production of CH4 from butyrate oxidation in paddy soil enrichment was significantly accelerated in the presence of nanoFe3O4, which was thought to be due to DIET activity (Li et al., 2015). Anaerobic freshwater lake sediments represent important methanogenic environments (Bastviken et al., 2004). The syntrophic enrichments were prepared from two lake sediments (an urban and a natural lake) with butyrate as the sole substrate in the presence of nanoFe3O4. We determined the effect of nanoFe3O4 on syntrophic methanogenesis and the corresponding community structure in lake sediment samples and attempted the identification of the organisms responsible for DIET. Furthermore, carbon nanotubes (CNTs) were used to determine if artificial electrically conductive nanomaterials could be substituted for nanoFe3O4.
Materials and Methods
Collection of Lake Sediments
Two lake sediment samples were collected at the depth of 0–15 cm from the sediment surface: one from Weiming Lake (WML), a pond on the campus of Peking University (39°59′36″N 116°18′12″E) and the other from Erhai Lake (EHL), which is located on the Yungui plateau within Yunnan Province in the southwestern China (26°01′N 100°03′E). WML has an area of about 3 ha and an average water depth of 1.5 m. The lake freezes temporally in winter. The pond is eutrophic. A sediment sample from it was analyzed. It contained 6.25% total organic carbon, 0.38% total nitrogen, and its pH was 7.78. The EHL is the second largest freshwater lake (256.5 km2) on Yungui plateau. It is situated at an altitude of 1934 m and has average water depth of 10.5 m. The water source of EHL is rainfall and the melt water from ice and snow. The sediment sample from EHL contained the 2.40% total organic carbon and 0.27% total nitrogen. Its pH was 7.20 (Wu et al., 2015). Sediment samples were obtained by scraping the sediment surface (0–15 cm) with a 3-liter sampler.
Preparation of Magnetite Nanoparticles and Multi-Walled CNTs
The nanoFe3O4 was synthesized via a conventional aqueous co-precipitation method (Kang et al., 1996). Briefly, 2 g FeCl2 and 5.2 g FeCl3 were dissolved in 0.4 N HCl solution and 1.5 N NaOH solution was added drop-wisely under vigorous mechanical agitation. Upon the addition of NaOH, precipitation of a black precipitate of magnetite was observed. The precipitate was separated with an external magnet, and then washed repeatedly with deionized water until the supernatant wash water was neutral (pH 7). Commercially available multi-walled CNTs (MWCNTs; 10–20 nm in diameter and 10–30 μm in length) were purchased from DK Nano Technology Co. (Beijing, China). MWCNTs were carboxylic acid functionalized (2% -COOH by weight) and were >98% pure. A stock suspension of 10% (w/v) MWCNTs in the water was used for the experiment.
Enrichment Cultivation
Approximately 0.5 g (WML) or 5 g (EHL) fresh sediment (microbial inocula for first enrichments) were put into sterile 120 serum bottles filled with 50 ml of HEPES-buffered (30 mM, pH 7) medium supplemented with sodium butyrate to a final concentration of 10 mM. The basal medium consisted of 0.4 g MgCl2, 0.1 g CaCl2, 0.1 g NH4Cl, 0.2 g KH2PO4, 0.5 g KCl, and 0.0005 g resazurin per liter of distilled water (Lü and Lu, 2012). Supplements of NaHCO3, NaS, vitamin, trace element solutions, and Se/W solution followed the protocol described previously (Lü and Lu, 2012). The cysteine was excluded in the medium to avoid the possible effect of electron shuttle molecules (Kaden et al., 2002). All the bottles were sealed with butyl stoppers and aluminum crimp caps. The enrichments were incubated statically in the dark at 30°C under the atmosphere of N2/CO2 [80:20 (V/V)] in the headspace of each bottle.
The first and second enrichments consisted of two treatments: (1) nanoFe3O4 were supplemented from the stock solution to give the final concentration of 10 mM as Fe atom; and (2) CK (no-amendment control). When the CH4 production from the Fe3O4 treatment approached a plateau, inculum from nanoFe3O4 treatment (4% inoculum) were transferred to medium as described above. In the third and fourth enrichment transfers, three treatments were prepared: control without supplements (CK), Fe3O4 addition at a final concentration of 10 mM as Fe atom and MWCNTs addition at a final concentration of 0.5% (w/v). After the third transfer, each culture was examined microscopically and subjected to molecular phylogenetic analysis. The experiment was carried out in triplicate. The design of experiment is shown in the Supplementary Figure 1.
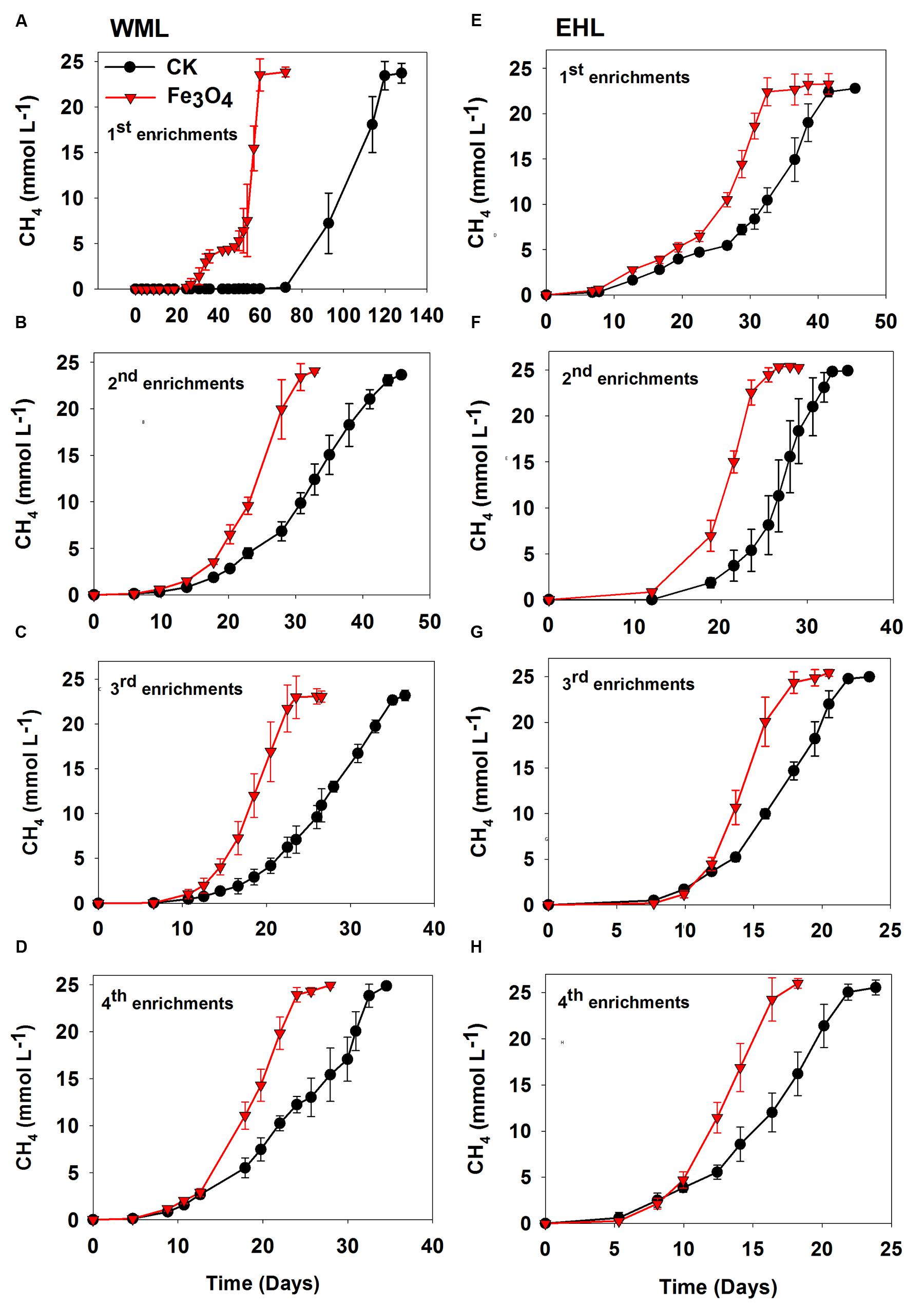
FIGURE 1. Effects of conductive Fe3O4 nanoparticles on the production of CH4 in the enrichments from Weiming Lake sediment, WML (A–D), and Erhai Lake sediment, EHL (E–H). Error bars represent the standard deviation of replicate experiments.
Chemical Analyses
Gas samples (100 μl) were regularly taken from the headspace using a Pressure-Lok precision analytical syringe (Bation Rouge, LA, USA). The CH4 concentration was analyzed using GC-7890 gas chromatograph (Agilent Technologies, USA) equipped with flame ionization detector (FID). The detection limit was 50 ppmv. For the analysis of organic acids including butyrate and acetate, 0.2 ml of culture medium was sampled with sterile syringes. Butyrate and acetate were analyzed by injecting filtered (0.22 μm porosity) liquid sample into a gas chromatograph equipped with FID detector (injection, detection, and column temperatures were 200, 250, and 120°C, respectively). We used a 30 m capillary column (DB-Waxetr, 0.53 mm i.d., 1 μm film thickness) to separate organic acids with a split ratio of 10:1. The HCl-extractable Fe(III) and Fe(II) were determined by the ferrozine technique as described elsewhere (Lovley and Phillips, 1986).
Microscopy
The cells in the mid-log phase were collected by a syringe fixed with 2.5% (wt/vol) glutaraldehyde in phosphate-buffered saline and sequentially dehydrated with ethanol (20, 40, 60, 80, 95, and 100% (v/v) ethanol, 10 min for each treatment). The dried samples were coated with platinum and imaged using scanning electron microscope (FEI NanoSEM 430).
Fluorescence in situ hybridization (FISH) analysis was performed on 4% paraformaldehyde-fixed samples according to a procedure described elsewhere (Moter and Göbel, 2000). Oligonucleotide probes specific for bacteria (Cy3-labeled EUB338mix probes) and archaea (FITC-labeled ARC915 probe) were used in this study. The details of the probes used are available in the probebase (Loy et al., 2003). The labeled samples were visualized using epifluorescence microscopy (Axio imager D2, ZEISS).
Analysis of Microbial Community Composition
The triplicate cultures growing in the presence of nanoFe3O4 in the fourth transfer were used to extract microbial DNA. Prior to extraction, sonication treatment was conducted to separate the microbial cells from Fe3O4 nanoparticles. Total DNA was extracted using the FastDNA SPIN Kit (MP Biomedicals, USA) according to the manufacturer’s protocol. Prior to PCR amplification, the DNA extracts from each replicate were mixed in equal amount. DNA extracts were stored at -20°C.
Bacterial and archaeal clone libraries were constructed from the WML enrichment. PCR amplification was performed using the primer pairs of Ba27f/907r for bacteria and Ar109f/Ar915r for archaea. PCR products were purified and clone library analyses were conducted as described previously (Peng et al., 2008). One hundred bacterial clones and one hundred archaeal clones were retrieved from the WML enrichment. Sequences of the clone libraries were analyzed by defining operational taxonomic units (OTU), in which representative sequences from each OTU was defined by 97% sequence identity. The closest matching sequences were identified by searching with the BLAST program in the NCBI database. Phylogenetic trees were constructed using MEGA 6.0 with the neighbor-joining method. The 16S rRNA gene sequences obtained in this study were deposited in the GenBank databases under accession numbers from KU577285 to KU577288.
Because relatively diverse cell morphologies were observed in EHL enrichment, the high-throughput sequencing was used for the microbial community analysis. After DNA extraction, the qualified DNA was sent to Sangon Biotech Company (Shanghai, China) for amplicon sequencing using the Illumina Miseq 2 × 300 bp platform (San Diego, CA, USA). The V3–V4 universal primers 314F (CCTACGGGNGGCWGCAG) and 805R (GACTACHVGGGTATCTAATCC) with sample-specific barcodes were used for interrogating bacterial communities. A nested PCR amplification, with the primer sets of 340F (CCCTAYGGGGYGCASCAG) and 1000R (GGCCATGCACYWCYTCTC) for the first round (25 cycles), and 349F (GYGCASCAGKCGMGAAW) and 806R (GGACTACVSGGGTATCTAAT) for the second round (25 cycles) was conducted to amplify the archaeal 16S rRNA genes. More than 30,000 sequences were obtained from each sample. Sequences were clustered into OTUs using UCLUST software1 with a similarity threshold of 97%. The Ribosomal Database Project (RDP) classifier was used to assign the taxonomic data to each representative sequence. Raw sequencing reads obtained in this study have been deposited in the NCBI SRA with the accession number SRP068809 and SRP068811.
Results
CH4 Production
Four enrichment transfers were performed for each sediment sample. In all transfers, we observed consistently increased rates of CH4 production in the presence of nanoFe3O4 (Figure 1). In all incubations, about 25 micromoles of CH4 per liter were produced from 10 micromoles of butyrate added. Acetate showed only intermediate accumulation (Supplementary Figure 2). This result indicated that methanogenesis in the enrichments followed the stoichiometry of complete conversion of butyrate to CH4 and CO2. For the WML sediment, the first enrichment was inoculated with 0.5 g of fresh sample into 50 ml medium. CH4 production displayed a substantial lag (Figure 1A), but this lag was markedly shortened in the presence of nanoFe3O4 (27 days versus 72 days in the control). The conversion of butyrate to CH4 was completed at 61 days in nanoFe3O4 compared with 119 days in the control. The lag phase was reduced significantly in the second and later transfers (Figures 1B–D). For the oligotrophic EHL, the first enrichment was inoculated with 5 g of fresh sediment. The initiation of CH4 production was faster than with the WML sediment. The stoichiometric conversion of butyrate in the first transfer of EHL enrichment was obtained at 32 days in the presence of nanoFe3O4 compared with 42 days in the control (Figure 1E). For both sediments, CH4 production was faster in the third and fourth enrichments compared with the first and second enrichments. The calculated maximum rate of CH4 production during the exponential growth phase was about 60–90% (WML) and 34–56% (EHL) greater in the presence of nanoFe3O4 than in the control (Supplementary Figure 3).
In the third and fourth transfers, MWCNTs were applied in parallel. The transfers involved inocula from previously nanoFe3O4-amended enrichments. The addition of MWCNTs accelerated CH4 production in all of the enrichments (Figure 2). The time needed for the complete conversion of butyrate to CH4 and CO2 was significantly shortened in the presence of MWCNTs compared with the control. The maximum rates of CH4 production in the presence of MWCNTs were approximately 50% greater relative to the control (Supplementary Figure 4).
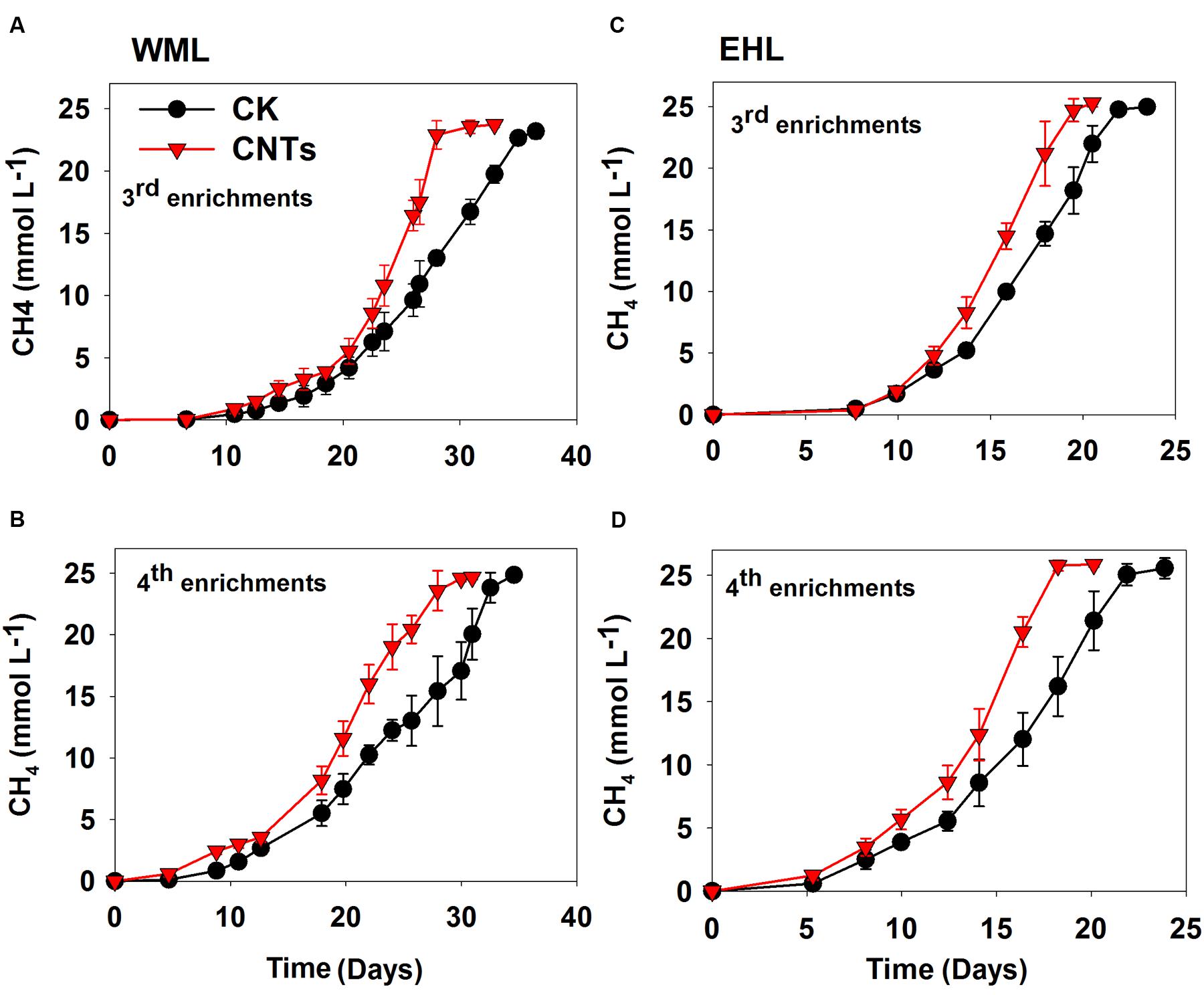
FIGURE 2. Effects of conductive Multi-walled CNTs on the production of CH4 in the third and fourth enrichments from WML (A,B) and EHL (C,D).
Microscopic Observation
The oligonucleotide probes used for FISH observations targeted bacteria and archaea. The fluorescence images revealed that numerous aggregates were formed in both WML and EHL enrichments (Figure 3). In many cases, it appeared that the bacterial cells formed dense cores with archaeal cells residing peripherally (Figures 3A,B). Most of bacterial cells were curved and rod-shapes and were morphologically similar in the WML and EHL enrichments. Archaeal cells were mostly coccoid in the WML enrichment (Figures 3A,C,E), whereas more of archaeal cells in the EHL enrichment were slender rods (Figures 3B,F). The microbial aggregates displayed distinct architectures between the control and those in nanoFe3O4 and MWCNTs treatments. In the control, the bacterial and archaeal cells in aggregates were in close physical proximity (Figures 3A,B). But in the nanoFe3O4 and MWCNTs treatments, there existed dark areas within aggregates (Figures 3C–F). These dark areas were filled with nanoFe3O4 particles and MWCNTs. The aggregates formed in the presence of nanoFe3O4 or MWCNTs seemed to exhibit greater intercellular distances on average than aggregates in the control. However, most of the cells in the presence of either nanoFe3O4 or MWCNTs were interconnected by the respective particles.
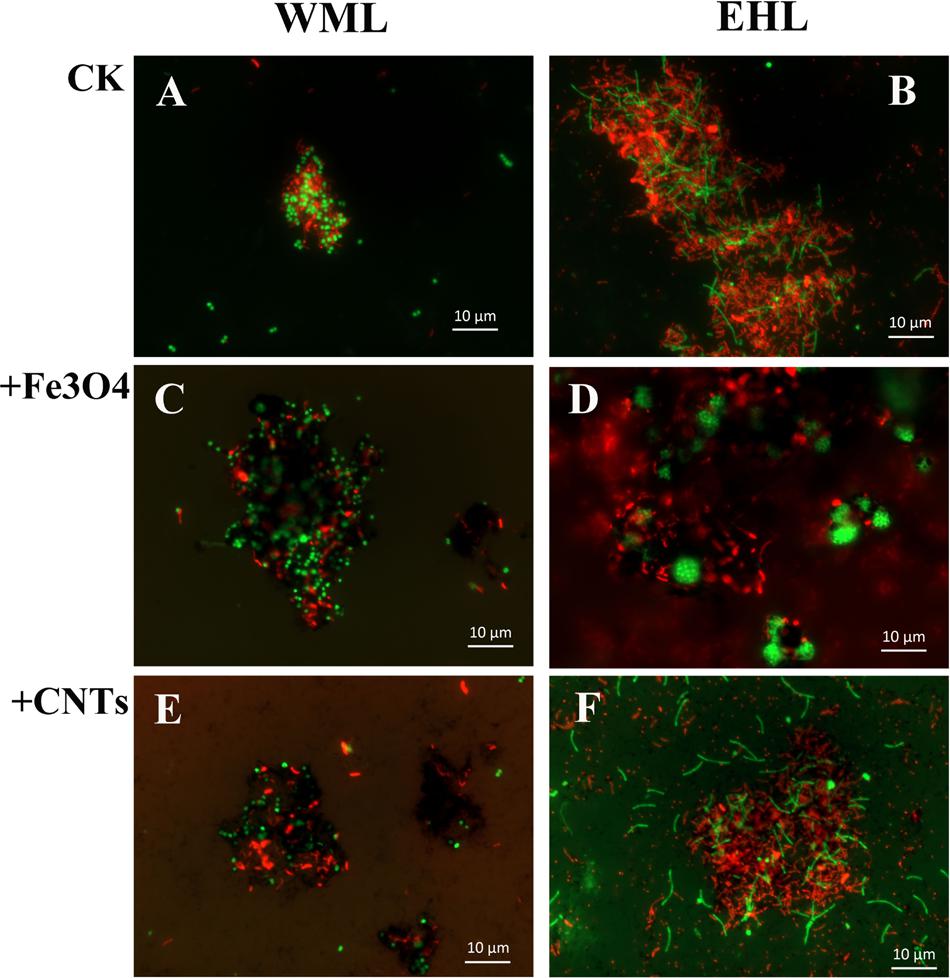
FIGURE 3. Spatial distribution of archaeal (Arc915-FITC, green) and bacterial (EUB338mix-Cy3, red) cells identified by FISH in WML and EHL enrichments with CK, Fe3O4 and MWCNTs treatments. (CK treatment: A,B; Fe3O4 treatment: C,D; WMCNTs treatment: E,F). The Fe3O4 nanoparticles and the MWCNTs appear black in the FISH images from the corresponding treatments.
The scanning electron microscope (SEM) corroborated the FISH observations (Figure 4). Figures 4A,B showed that the cells were in close contact forming dense microbial aggregates in the control. The cells in the WML enrichment were coccus-shaped, rod-shaped and curved (Figure 4A). In the EHL enrichment, Methanosarcina-like cells and long slender rods were observed additionally (Figure 4B). The extracellular polymer-like substances were also detected in the aggregates. The cells from the nanoFe3O4 and MWCNTs treatments (Figures 4C–F) were less densely organized and showed larger intercellular distance on average compared with the control. However, most of these cells were closely associated with nanomaterials and formed cell/nanomaterial mixtures (Figures 4C–F).
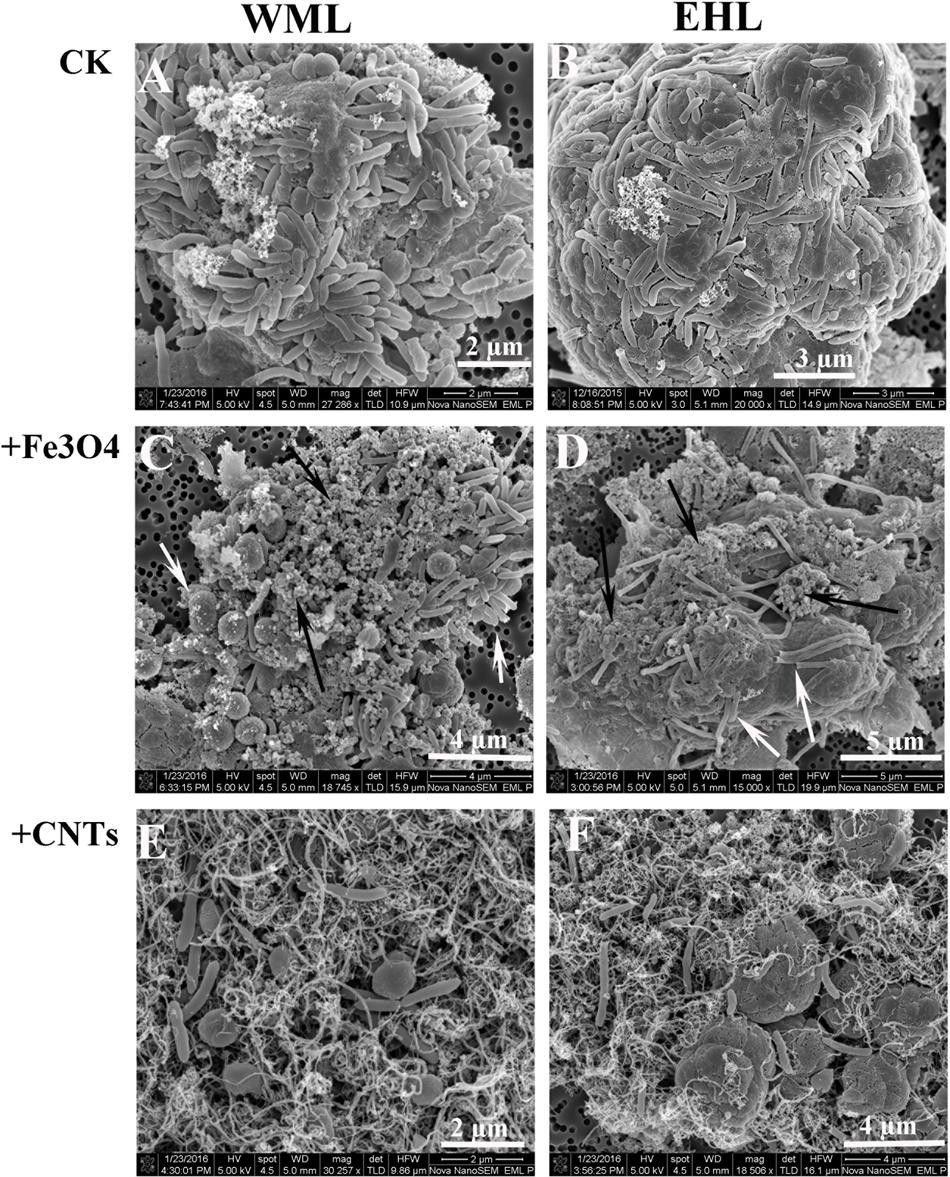
FIGURE 4. Scanning electron micrographs (SEM) of cell aggregates or cell/material mixtures in the enrichment cultures from WML and EHL (WML: A,C,E; EHL: B,D,F). White arrows indicate cells and the black arrows indicate Fe3O4 nanoparticles.
Microbial Communities
The composition of microbial communities in the enrichments was determined by 16S rRNA gene cloning and sequencing. We conducted the conventional cloning and sequencing with the WML enrichment and Miseq sequencing with the EHL enrichment because of the relatively more complex community in the latter. One hundred bacterial clones retrieved from the WML enrichment could be classified into two OTUs (Figure 5A). OTU 1 belonged to Syntrophomonas, accounting for 96% of the total sequences. It was closely related (99% identity) to an uncultured bacterial clone detected in an anode biofilm of a microbial fuel cell (clone BP, JX145977; Ishii et al., 2014). The closest pure culture relative was S. bryantii CuCal (95% identity in 16S rRNA). The remaining four sequences (OTU 2) were related to a known bacterial strain, Desulfovibrio carbinoliphilus D41 (98% identity in 16S rRNA; Allen et al., 2008). Figure 5B shows that the archaeal clone sequences (100 clones) were comprised 84% of the Methanosarcina-like sequences (OTU 2), with the closest pure culture relatives being M. horonobensis HB-1 (99.2% identity), and 16% of Methanomicrobia-like sequences (OTU 1) that were related to Methanosphaerula palustris E1-9c (94% identity).
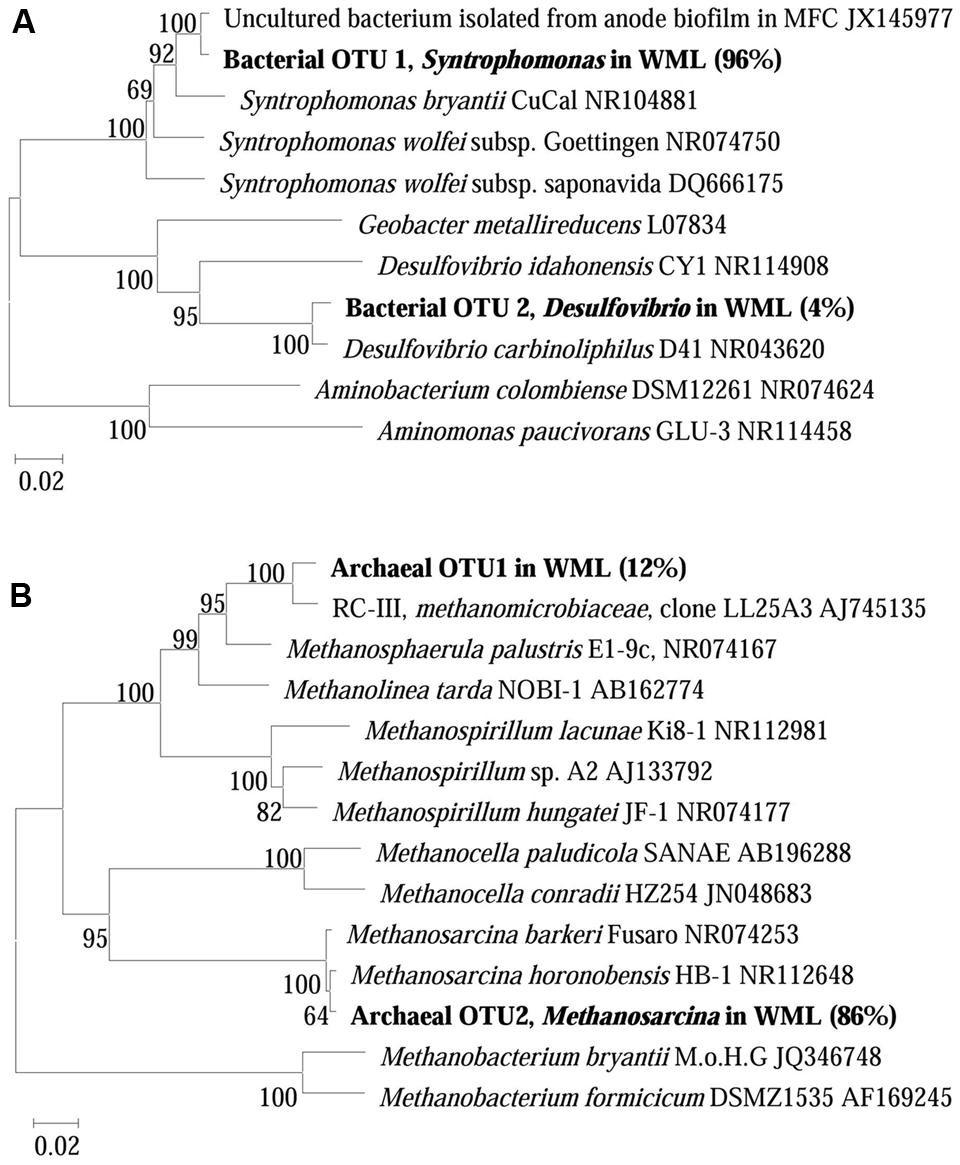
FIGURE 5. Neighbor-joining phylogenetic tree of representative bacterial (A) and archaeal (B) 16S rRNA gene clones obtained from the fourth enrichments of WML with the addition of Fe3O4 nanoparticles. Clones obtained in this study are indicated in boldface and their relative abundances are given in parentheses. GenBank accession numbers of reference sequences are indicated.
The Miseq sequencing revealed that the bacterial populations in the EHL enrichment (Figure 6A) were dominated by Syntrophomonas (40%), followed by Sulfurospirillum (26%) and a few other bacterial lineages Paenibacillus, Treponema, and unclassified Rikenellaceae. The archaeal community (Figure 6B) consisted mainly of Methanoregula (57%), Methanosarcina (26%) and Methanospirillum (10%), with a minor proportion of Methanosaeta and Methanobacterium.
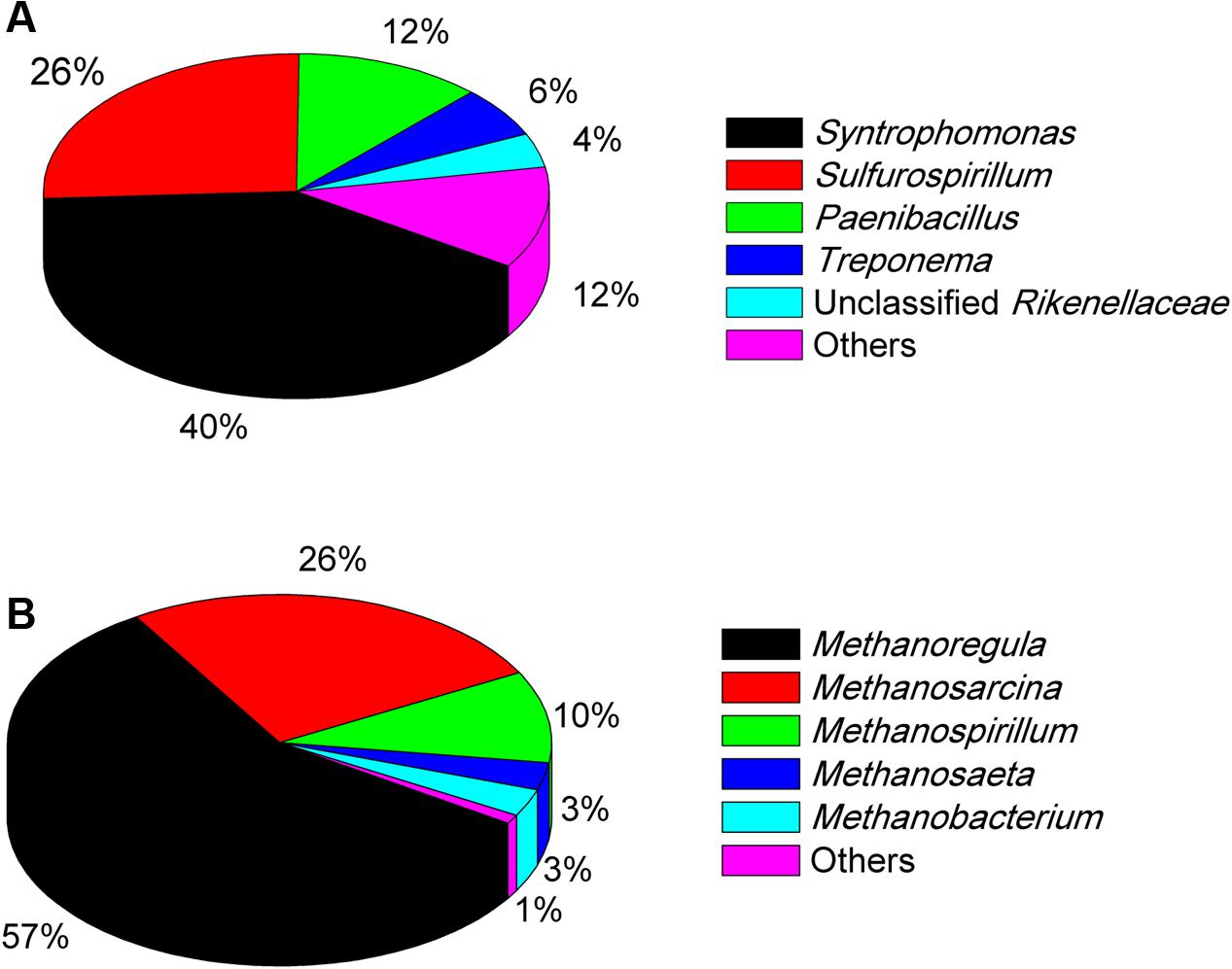
FIGURE 6. The community composition and relative abundance of bacteria (A) and archaea (B) at genus level in the fourth enrichments of EHL as determined by Illumina Miseq sequencing. The genus whose relative abundance was less than 3% was included in the group “Other.” More than 30,000 sequences were obtained for the bacteria and archaea phylogenetic classifications at genus level separately.
Discussion
In all of the enrichments, we found that the addition of nanoFe3O4 substantially facilitated the syntrophic production of CH4 from butyrate oxidation. Furthermore, the addition of MWCNTs, an artificial chemically stable nanomaterial with high electric conductivity (Baughman et al., 2002), displayed a similar stimulatory effect. These results suggest that the electric conductivity of the added nanoparticles played a key role in facilitating the syntrophic oxidation of butyrate.
Microbial aggregates were extensively formed in the sediment enrichments (Figure 3). It has been suggested that the formation of microbial aggregates can enhance the efficiency of interspecies H2 transfer due to the reduction of cell-to-cell distance for the electron-carrier diffusion (Kouzuma et al., 2015). Even though all enrichments showed microbial aggregates formation, the structure of the aggregates formed in the control and in the presence of nanoFe3O4 or MWCNTs differed from each other. The consensus of visual image estimates suggests that the presence of nanoparticles within the aggregates increased rather than shortened the intercellular distance compared with the control. If the intercellular distance were a key factor in the syntrophic interaction based on interspecies H2/formate diffusion, a lower rate of CH4 production would be expected in the presence of nanoparticles. Therefore, our results suggest that the conductive nanoparticles likely facilitated DIET in the butyrate syntrophy by forming the cell-nanomaterial-cell networks. The discrepancy between cell-to-cell distance and the syntrophic activity has also been considered to indicate the involvement of DIET in the anaerobic oxidation of CH4 in ocean sediments (McGlynn et al., 2015). Theoretical calculation suggested that the electron transfer rate among the syntrophic partners via direct electric conduction was substantially higher (106 times) than via interspecies H2 diffusion (Viggi et al., 2014).
The dominant bacterial clones in both WML enrichment (96%) and EHL enrichment (40%) were affiliated with Syntrophomonas. The clone sequence from the WML enrichment was closely (99% identity) related to an uncultured bacterial clone retrieved from an anode biofilm of a microbial fuel cell fed with butyrate and propionate (Ishii et al., 2014). It has to be noted that the clone analysis and phylogenetic information reflected only the bacterial composition in the enrichment rather than the community in the original sediment sample. Syntrophomonas have been well recognized as an obligate syntroph in anaerobic oxidation of butyrate. Genomic analyses have so far not been able to identify the mechanism of extracellular electron transfer in Syntrophomonas (Sieber et al., 2010, 2012). In other studies, involving anaerobic membrane bioreactor (Smith et al., 2015), microbial fuel cells (Ishii et al., 2014) and microbial electrolysis cells (Zhao et al., 2016), Syntrophomonas were detected in the biofilms or anodic biofilms, and indicated that they was able to contribute to butyrate degradation and electricity generation.
In the EHL enrichment, we also found that Sulfurospirillum was enriched (26%). Sulfurospirillum is a sulfur-reducing bacterium able to use Fe(III), arsenate, elemental sulfur, thiosulfate, and nitrate as electron acceptors (Kodama et al., 2007), and acetate, lactate, and butyrate as electron donors (Héry et al., 2008). The present experiment was performed under methanogenic conditions. Therefore, Sulfurospirillum probably also directly participated in the syntrophic oxidation of butyrate in the EHL enrichment. The mechanisms deserve further investigations.
Methanosarcina were specifically enriched (86%) in the WML enrichment after continuous transfers in the presence of nanoFe3O4. In the EHL enrichment the hydrogenotrophic Methanoregula dominated (accounting for 57%) with Methanosarcina being the second major methanogens (26%). Methanogens serve as the syntrophic partner utilizing the electrons released from syntrophic bacteria. The molecular mechanisms of the involvement of methanogens in DIET remain unknown (Kouzuma et al., 2015). However, pure cultures of Methanosaeta harundinacea and Methanosarcina barkeri have been used in the construction of syntrophic co-cultures with Geobacter, which performed DIET in CH4 production from ethanol (Liu et al., 2012; Rotaru et al., 2014a,b). We showed earlier that Methanosarcinaceae, Methanocellales, and Methanobacteriales were possibly involved in DIET in paddy soil (Li et al., 2015). Other studies showed that Methanococcus maropaludis, Methanobacterium palustre, and Methanothermobacter spp. were capable of accepting electrons in a more direct way from cathode electrodes than from H2 in the electrochemical systems (Cheng et al., 2009; Lohner et al., 2014; Fu et al., 2015). In an electrochemical study, Methanoregula were found to be specifically enriched when the applied voltage was set lower than the threshold for H2 production, indicating that Methanoregula produced CH4 via cathode reaction: CO2 + 8H+ + 8e- → CH4 + 2H2O (Chen et al., 2016). Thus, the extracellular electron uptake may occur with various methanogens. More studies are needed to elucidate the mechanisms of direct extracellular electron uptake by methanogens.
Based on the origin of mediators, DIET can be categorized into two forms, DIET of biological origin, which employs outer-membrane c-type cytochromes and pili or exudation of molecular redox shuttles, and DIET of environmental origin, which utilizes electron-conductive material of natural origin and/or the artificial supplements (Shrestha and Rotaru, 2014). The DIET of biological origin has been intensively studied using the defined co-cultures with Geobacter spp. (Lovley, 2012; Kouzuma et al., 2015). However, DIET of environment origin has been less understood. Utilization of DIET of environment origin shall confer an ecological advantage to microbes using DIET because biosynthesis of molecular conduits is not needed (Kato et al., 2012; Li et al., 2015). Furthermore, biological mediators are probably not ubiquitous among microorganisms, while the conductive-semiconductive minerals have widespread occurrence in nature. Additional studies shall clarify the significance of DIET of environment origin in methanogenesis and carbon cycling in various environments.
Author Contributions
JZ and YL devised the study, JZ conducted the experiments and analyses, all the authors contributed to data interpretation and writing of the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgment
This work was partly supported by the National Natural Science Foundation of China (41130527).
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2016.01316
Footnotes
References
Allen, T. D., Kraus, P. F., Lawson, P. A., Drake, G. R., Balkwill, D. L., and Tanner, R. S. (2008). Desulfovibrio carbinoliphilus sp. nov., a benzyl alcohol-oxidizing, sulfate-reducing bacterium isolated from a gas condensate-contaminated aquifer. Int. J. Syst. Evol. Microbiol. 58, 1313–1317. doi: 10.1099/ijs.0.65524-0
Bastviken, D., Cole, J., Pace, M., and Tranvik, L. (2004). Methane emissions from lakes: dependence of lake characteristics, two regional assessments, and a global estimate. Glob. Biogeochem. Cycle 18, GB4009. doi: 10.1029/2004GB002238
Baughman, R. H., Zakhidov, A. A., and de Heer, W. A. (2002). Carbon nanotubes–the route toward applications. Science 297, 787–792. doi: 10.1126/science.1060928
Chen, S., Rotaru, A.-E., Shrestha, P. M., Malvankar, N. S., Liu, F., Fan, W., et al. (2014). Promoting interspecies electron transfer with biochar. Sci. Rep. 4:5019. doi: 10.1038/srep05019
Chen, Y., Yu, B., Yin, C., Zhang, C., Dai, X., Yuan, H., et al. (2016). Biostimulation by direct voltage to enhance anaerobic digestion of waste activated sludge. RSC Adv. 6, 1581–1588. doi: 10.1039/c5ra24134k
Cheng, S., Xing, D., Call, D. F., and Logan, B. E. (2009). Direct biological conversion of electrical current into methane by electromethanogenesis. Environ. Sci. Technol. 43, 3953–3958. doi: 10.1021/es803531g
Fu, Q., Kuramochi, Y., Fukushima, N., Maeda, H., Sato, K., and Kobayashi, H. (2015). Bioelectrochemical analyses of the development of a thermophilic biocathode catalyzing electromethanogenesis. Environ. Sci. Technol. 49, 1225–1232. doi: 10.1021/es5052233
Héry, M., Gault, A. G., Rowland, H. A. L., Lear, G., Polya, D. A., and Lloyd, J. R. (2008). Molecular and cultivation-dependent analysis of metal-reducing bacteria implicated in arsenic mobilisation in south-east asian aquifers. Appl. Geochem. 23, 3215–3223. doi: 10.1016/j.apgeochem.2008.07.003
Ishii, S. I., Suzuki, S., Norden-Krichmar, T. M., Phan, T., Wanger, G., Nealson, K. H., et al. (2014). Microbial population and functional dynamics associated with surface potential and carbon metabolism. ISME J. 8, 963–978. doi: 10.1038/ismej.2013.217
Kaden, J., Galushko, A. S., and Schink, B. (2002). Cysteine-mediated electron transfer in syntrophic acetate oxidation by cocultures of Geobacter sulfurreducens and Wolinella succinogenes. Arch. Microbiol. 178, 53–58. doi: 10.1007/s00203-002-0425-3
Kang, Y. S., Risbud, S., Rabolt, J. F., and Stroeve, P. (1996). Synthesis and characterization of nanometer-size Fe3O4 and γ-Fe2O3 Particles. Chem. Mat. 8, 2209–2211. doi: 10.1021/cm960157j
Kato, S., Hashimoto, K., and Watanabe, K. (2012). Methanogenesis facilitated by electric syntrophy via (semi) conductive iron-oxide minerals. Environ. Microbiol. 14, 1646–1654. doi: 10.1111/j.1462-2920.2011.02611.x
Kodama, Y., Ha, L. T., and Watanabe, K. (2007). Sulfurospirillum cavolei sp. nov., a facultatively anaerobic sulfur-reducing bacterium isolated from an underground crude oil storage cavity. Int. J. Syst. Evol. Microbiol. 57, 827–831. doi: 10.1099/ijs.0.64823-0
Kouzuma, A., Kato, S., and Watanabe, K. (2015). Microbial interspecies interactions: recent findings in syntrophic consortia. Front. Microbiol. 6:477. doi: 10.3389/fmicb.2015.00477
Li, H., Chang, J., Liu, P., Fu, L., Ding, D., and Lu, Y. (2015). Direct interspecies electron transfer accelerates syntrophic oxidation of butyrate in paddy soil enrichments. Environ. Microbiol. 17, 1533–1547. doi: 10.1111/1462-2920.12576
Liu, F., Rotaru, A.-E., Shrestha, P. M., Malvankar, N. S., Nevin, K. P., and Lovley, D. R. (2012). Promoting direct interspecies electron transfer with activated carbon. Energy Environ. Sci. 5, 8982–8989. doi: 10.1039/c2ee22459c
Liu, F., Rotaru, A.-E., Shrestha, P. M., Malvankar, N. S., Nevin, K. P., and Lovley, D. R. (2015). Magnetite compensates for the lack of a pilin-associated c-type cytochrome in extracellular electron exchange. Environ. Microbiol. 17, 648–655. doi: 10.1111/1462-2920.12485
Lohner, S. T., Deutzmann, J. S., Logan, B. E., Leigh, J., and Spormann, A. M. (2014). Hydrogenase-independent uptake and metabolism of electrons by the archaeon Methanococcus maripaludis. ISME J. 8, 1673–1681. doi: 10.1038/ismej.2014.82
Lovley, D. R. (2012). Electromicrobiology. Annu. Rev. Microbiol. 66, 391–409. doi: 10.1146/annurev-micro-092611-150104
Lovley, D. R., and Phillips, E. J. P. (1986). Availability of ferric iron for microbial Reduction in bottom sediments of the freshwater tidal Potomac River. Appl. Environ. Microbiol. 52, 751–757.
Loy, A., Horn, M., and Wagner, M. (2003). probeBase: an online resource for rRNA-targeted oligonucleotide probes. Nucleic Acids Res. 31, 514–516. doi: 10.1093/nar/gkg016
Lü, Z., and Lu, Y. (2012). Methanocella conradii sp. nov., a thermophilic, obligate hydrogenotrophic methanogen, isolated from Chinese rice field soil. PLoS ONE 7:e35279. doi: 10.1371/journal.pone.0035279
McGlynn, S. E., Chadwick, G. L., Kempes, C. P., and Orphan, V. J. (2015). Single cell activity reveals direct electron transfer in methanotrophic consortia. Nature 526, 531–535. doi: 10.1038/nature15512
Morita, M., Malvankar, N. S., Franks, A. E., Summers, Z. M., Giloteaux, L., Rotaru, A. E., et al. (2011). Potential for direct interspecies electron transfer in methanogenic wastewater digester aggregates. MBio 2, e00159-11. doi: 10.1128/mBio.00159-11
Moter, A., and Göbel, U. B. (2000). Fluorescence in situ hybridization (FISH) for direct visualization of microorganisms. J. Microbiol. Methods 41, 85–112. doi: 10.1016/S0167-7012(00)00152-4
Peng, J., Lü, Z., Rui, J., and Lu, Y. (2008). Dynamics of the methanogenic archaeal community during plant residue decomposition in an anoxic Rice field soil. Appl. Environ. Microbiol. 74, 2894–2901. doi: 10.1128/aem.00070-08
Rotaru, A.-E., Shrestha, P. M., Liu, F., Markovaite, B., Chen, S., Nevin, K. P., et al. (2014a). Direct Interspecies electron transfer between Geobacter metallireducens and Methanosarcina barkeri. Appl. Environ. Microbiol. 80, 4599–4605. doi: 10.1128/aem.00895-14
Rotaru, A.-E., Shrestha, P. M., Liu, F., Shrestha, M., Shrestha, D., Embree, M., et al. (2014b). A new model for electron flow during anaerobic digestion: direct interspecies electron transfer to Methanosaeta for the reduction of carbon dioxide to methane. Energy Environ. Sci. 7, 408–415. doi: 10.1039/c3ee42189a
Shrestha, P. M., and Rotaru, A.-E. (2014). Plugging in or going wireless: strategies for interspecies electron transfer. Front. Microbiol. 5:237. doi: 10.3389/fmicb.2014.00237
Sieber, J. R., McInerney, M. J., and Gunsalus, R. P. (2012). Genomic insights into syntrophy: the paradigm for anaerobic metabolic cooperation. Annu. Rev. Microbiol. 66, 429–452. doi: 10.1146/annurev-micro-090110-102844
Sieber, J. R., Sims, D. R., Han, C., Kim, E., Lykidis, A., Lapidus, A. L., et al. (2010). The genome of Syntrophomonas wolfei: new insights into syntrophic metabolism and biohydrogen production. Environ. Microbiol. 12, 2289–2301. doi: 10.1111/j.1462-2920.2010.02237.x
Skomurski, F. N., Kerisit, S., and Rosso, K. M. (2010). Structure, charge distribution, and electron hopping dynamics in magnetite (Fe3O4) (100) surfaces from first principles. Geochim. Cosmochim. Acta 74, 4234–4248. doi: 10.1016/j.gca.2010.04.063
Smith, A. L., Skerlos, S. J., and Raskin, L. (2015). Membrane biofilm development improves COD removal in anaerobic membrane bioreactor wastewater treatment. Microb. Biotechnol. 8, 883–894. doi: 10.1111/1751-7915.12311
Stams, A. J. M., and Plugge, C. M. (2009). Electron transfer in syntrophic communities of anaerobic bacteria and archaea. Nat. Rev. Microbiol. 7, 568–577. doi: 10.1038/nrmicro2166
Thauer, R. K., Kaster, A.-K., Seedorf, H., Buckel, W., and Hedderich, R. (2008). Methanogenic archaea: ecologically relevant differences in energy conservation. Nat. Rev. Microbiol. 6, 579–591. doi: 10.1038/nrmicro1931
Viggi, C. C., Rossetti, S., Fazi, S., Paiano, P., Majone, M., and Aulenta, F. (2014). Magnetite particles triggering a faster and more robust syntrophic pathway of methanogenic propionate degradation. Environ. Sci. Technol. 48, 7536–7543. doi: 10.1021/es5016789
Wu, X., Wang, C., Tian, C., Xiao, B., and Song, L. (2015). Evaluation of the potential of anoxic biodegradation of intracellular and dissolved microcystins in lake sediments. J. Hazard. Mater. 286, 395–401. doi: 10.1016/j.jhazmat.2015.01.015
Keywords: syntrophy, methane, methanogenesis, direct interspecies electron transfer, lake sediments, Fe3O4, butyrate
Citation: Zhang J and Lu Y (2016) Conductive Fe3O4 Nanoparticles Accelerate Syntrophic Methane Production from Butyrate Oxidation in Two Different Lake Sediments. Front. Microbiol. 7:1316. doi: 10.3389/fmicb.2016.01316
Received: 16 June 2016; Accepted: 09 August 2016;
Published: 22 August 2016.
Edited by:
Dennis A. Bazylinski, University of Nevada, Las Vegas, USAReviewed by:
Zhe-Xue Quan, Fudan University, ChinaMarja A. Tiirola, University of Jyväskylä, Finland
Copyright © 2016 Zhang and Lu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yahai Lu, bHV5aEBwa3UuZWR1LmNu
 Jianchao Zhang
Jianchao Zhang Yahai Lu
Yahai Lu