- 1Department of Clinical Laboratory, Peking Union Medical College Hospital, Chinese Academy of Medical Sciences, Beijing, China
- 2Graduate School, Peking Union Medical College, Chinese Academy of Medical Sciences, Beijing, China
- 3Centre for Infectious Diseases and Microbiology Laboratory Services, Institute of Clinical Pathology and Medical Research–Pathology West, Westmead Hospital, University of Sydney, Sydney, NSW, Australia
Candida nivariensis and C. bracarensis are two emerging cryptic species within the C. glabrata complex. Thirteen of these isolates from 10 hospitals in China were studied for their species identification and antifungal susceptibilities. Phenotypic and molecular [rDNA ITS sequencing, D1/D2 sequencing and ITS sequencer-based capillary gel electrophoresis (SCGE)] and matrix-assisted laser desorption ionization-time of flight (MALDI-TOF) MS identification methods were compared for their performance in species identification. Twelve of 13 (92.3%) isolates were identified as C. nivariensis and one as C. bracarensis using ITS sequencing as the reference method. Results obtained by D1/D2 sequencing and ITS SCGE were concordant with ITS sequencing results for all (100%) isolates. SCGE was able to subtype 12 C. nivariensis into four ITS SCGE length types. All isolates failed to be identified by the Vitek MALDI-TOF MS system (bioMérieux), whilst the Bruker MS system (Bruker Daltoniks) correctly identified all C. nivariensis isolates but using a lowered (≥1.700) cut-off score for species assignment; the C. bracarensis isolate was identified but with score <1.700. The Vitek 2 Compact system could not identify 11 C. nivariensis and one C. bracarensis isolate and misidentified the remaining C. nivarensis strain as “C. glabrata.” All isolates were susceptible-dose dependent to fluconazole [minimum inhibitory concentration (MIC) range 0.5–4 μg/mL] and were classed as susceptible to echinocandins (MICs ≤ 0.06 μg/mL). All 13 isolates had low MICs for other azoles (MICs ≤ 0.5 μg/mL), amphotericin B (MICs ≤ 2 μg/mL) and 5-flucytosine (MICs ≤ 0.25 μg/mL). Our results reinforce the need for molecular differentiation of species of C. nivarensis and C. bracarensis. The performance of MALDI-TOF may be improved by adding mass spectral profiles (MSPs) into the current databases. The antifungal susceptibility profile of isolates should be monitored.
Introduction
Candida remains the most important cause of opportunistic mycoses worldwide (Pfaller and Diekema, 2007). Although overall, C. albicans remains the most common Candida species responsible for invasive candidiasis, non-albicans Candida species increasingly represent a significant burden (Pfaller et al., 2010). With evolving and changing fungal taxonomy due to advances in DNA sequencing techniques, previously unrecognized species, as well as “cryptic” species within species complexes have been identified. Examples include the distinction between closely related species within the C. parapsilosis species complex (C. parapsilosis sensu stricto, C. orthopsilosis and C. metapsilosis), C. haemulonii complex (C. haemulonii, C. duobushaemulonii and C. haemulonii var. vulnera) and C. glabrata complex (C. glabrata sensu stricto, C. nivariensis and C. bracarensis; Alcoba-Florez et al., 2005; Tavanti et al., 2005; Correia et al., 2006; Cendejas-Bueno et al., 2012).
Although cryptic species of the C. glabrata complex likely cause similar disease manifestations as C. glabrata sensu stricto, their distinction is important due to differences in antifungal susceptibility where both C. nivariensis and C. bracarensis have been reported to be more drug resistant including to the azoles (Fujita et al., 2007; Bishop et al., 2008; Borman et al., 2008; Figueiredo-Carvalho et al., 2016). However, phenotypic identification methods cannot identify these two species where they are “misidentified” as “C. glabrata” by identification systems such as Vitek 2 Compact system and API ID32C (bioMérieux, Marcy l’Etoile, France; Alcoba-Florez et al., 2005; Correia et al., 2006; Lockhart et al., 2009). Matrix-assisted laser desorption ionization–time of flight mass spectrometry (MALDI-TOF MS) is able to rapidly and accurately identify Candida species (Pinto et al., 2011). The mass spectra (MS) of C. glabrata sensu stricto, C. nivariensis and C. bracarensis are included in most commercial MS databases (Pinto et al., 2011; Gorton et al., 2013). Nonetheless, confirmation of species identification remains reliant on DNA sequencing of the fungal rDNA ITS region (considered the reference method) or the D1/D2 region of the 28S rRNA gene in many instances (Alcoba-Florez et al., 2005; Correia et al., 2006). Other molecular methods may also be employed to identify and distinguish within C. glabrata species complex including an ITS-targeted SCGE method. We had previously used SCGE to distinguish between subtypes of both C. glabrata (cgl-1 and cgl-2) and C. nivariensis (cni-1 and cni-2; Hou et al., 2016).
Despite growing awareness of their clinical significance, data on the occurrence and distribution of C. nivariensis and C. bracarensis in clinical specimens in China has not been described. Therefore, here we have studied the epidemiology and antifungal susceptibility of C. nivariensis and C. bracarensis clinical isolates collected from a multi-center surveillance in China over 5 years.
Materials and Methods
Ethics Statement
The study was approved by the Human Research Ethics Committee of Peking Union Medical College Hospital (No. S-263). Written informed consent was obtained from patients for the use of the samples in research.
Yeast Isolates
All isolates were collected as part of the nationwide surveillance program for IFDs in China -CHIF-NET from August 2009 to July 2014 (Table 1). The CHIF-NET was a nationwide, prospective, laboratory-based, surveillance network established to provide updated information on the epidemiology of IFDs in China and the study inclusion criteria are previously described (Wang et al., 2012). Isolates were collected from different study centres and then forwarded to a central reference laboratory (Department of Clinical Laboratory, Peking Union Medical College Hospital) for species identification. Species were identified at study sites but confirmed in all cases at the reference laboratory. Specifically, species identification was performed by MALDI-TOF MS and by sequencing of the ITS region according to the algorithm of Zhang et al. (2014). All isolates identified as “C. glabrata” and all uncommon or interesting Candida species were also identified by sequencing of the ITS regions. Over 8000 Candida isolates were collected during 5 years and a total of 12 C. nivariensis and one C. bracarensis were identified. Then phenotypic and molecular (rDNA ITS sequencing, D1/D2 sequencing and ITS SCGE) and MALDI-TOF MS identification methods were compared and evaluated for their performance in species identification.
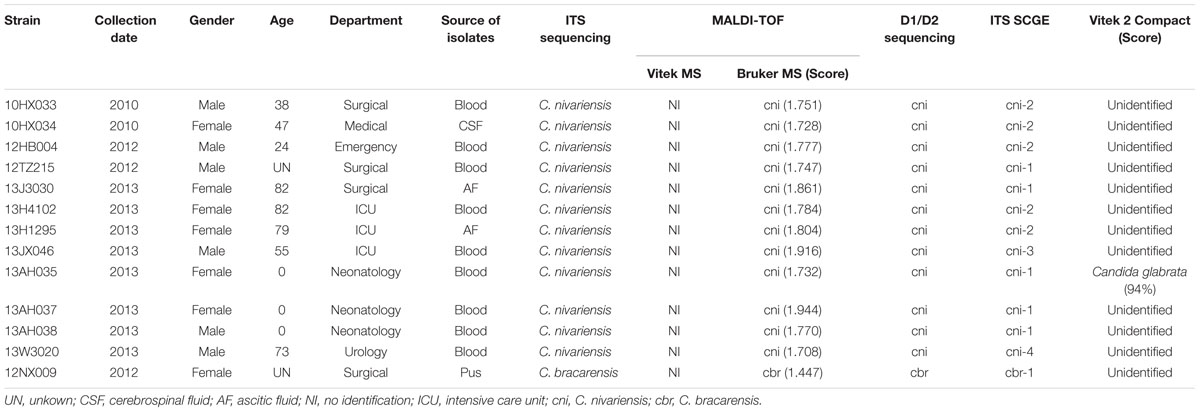
TABLE 1. Demographic data and identification of the Candida nivariensis and Candida bracarensis isolates included in this study.
Vitek 2 Compact System Identification
All 13 isolates were also re-identified using the Vitek-2 yeast identification card (Vitek-2 Yeast ID; bioMérieux, Marcy l’Etoile, France) following the manufacturer’s instructions. The inoculum suspensions for the Vitek 2 were prepared in sterile saline at turbidity equal to a 2.0 McFarland standard, as measured using a DensiChek instrument (bioMérieux). The individual test cards were automatically filled with the prepared culture suspension, sealed, and incubated by the Vitek 2 instrument. Cards were incubated for 18 h at 35°C and read every 15 min. The final profile results were compared with the database, and the identification of the unknown organism was obtained.
Sequencer-Based Identification
(i) For DNA sequencing, all isolates were identified by sequencing the ITS region gene and the D1/D2 domain of the 28S rRNA gene was performed as previously described (Amberg et al., 2005; Zhang et al., 2014). The ITS region and D1/D2 sequences of strains used in this study have been deposited in GenBank (Supplementary Table S1).
(ii) For ITS SCGE, both the ITS1 and full-length ITS regions of each isolate were amplified by a duplex PCR as previously described (Hou et al., 2016).
Phylogenetic Analysis
ITS and D1/D2 nucleotide sequences of C. nivariensis and C. bracarensis were obtained from GenBank (Supplementary Table S1). C. glabrata ATCC 2001 was also included. The ITS and D1/D2 sequences were then used for phylogenetic analysis by the maximum-likelihood algorithm with 1000 bootstrap replication to ensure robustness using MEGA software (version 6.0, MEGA Inc., Englewood, NJ, USA).
Matrix-Assisted Laser Desorption Ionization-Time of Flight Mass Spectrometry (MALDI-TOF MS) Analysis
All isolates were identified by both Vitek MS system (IVD Knowledgebase version 2.0, bioMérieux, Marcy l’Etoile, France) and Bruker Autoflex Speed TOF/TOF MS system using Biotyper version 3.1 software (Bruker Daltoniks, Billerica, MA, USA) according to the manufacturer’s instructions. For the Vitek MS system, the results were scored in one of three ways, (i) a single identification (confidence value of 60.0 to 99.9%), (ii) a split identification for which a set of possible organisms is displayed, or (iii) NI when no match is found (Zhang et al., 2014). C. nivariensis and C. bracarensis reference spectra are not included in the Vitek MS v2.0 database. With the Bruker system, identification was provided according to manufacturer-determined criteria: a spectral score of <1.700 was considered not to provide reliable identification. A score of 1.700 but <2.000 indicated identification at the genus level, and a score of ≥2.000, identification at the species level (Deak et al., 2015). The current Bruker Daltonik v3.1 database contains 5,989 MSPs, which include reference spectra of C. nivariensis and C. bracarensis.
Antifungal Susceptibility Testing
The in vitro susceptibility to nine antifungal drugs – FLC, VRC, ITC, POS, CAS, MCF, ANF, AMB and 5FC – was determined using Sensititre YeastOneTM YO10 methodology (Thermo Scientific, Cleveland, OH, USA) following the manufacturer’s instructions. Briefly, isolates were sub-cultured onto Sabouraud dextrose agar (Oxoid Ltd., Hampshire, UK) and incubated at 35°C for 48 h. After this, 20 μL of 0.5 McFarland yeast suspension was transferred into 11 mL of inoculum broth and then 100 μL of the inoculated broth was transferred to each well of the manufacturer’s plate. Plates were incubated at 35°C and the MIC endpoints were read at 24 h. C. krusei ATCC 6258 and C. parapsilosis ATCC22019 were used as quality control strains in every test run. For C. glabrata complex, MIC values were interpreted according to the CBPs for FLC and echinocandins according to the CLSI M27-S4 standard, and to ECVs for the other agents (Clinical and Laboratory Standards Institute, 2012; Pfaller and Diekema, 2012).
Review of C. nivariensis and C. bracarensis Infections Reported in PubMed
For comprehensive understanding of the current epidemiology and antifungal susceptibility profiles of C. nivariensis and C. bracarensis, we reviewed and summarized all published literature in PubMed1 database as of July 12, 2016.
Results
Detailed information relating to the study isolates is summarized in Table 1. The 13 isolates were collected from 13 patients at 10 hospitals situated in eight provinces across China. For C. nivariensis, 25% (3/12) of the isolates were from patients admitted in the surgery department, 25% (3/12) from the ICU, 25% (3/12) from neonatology department, and the remaining 25% (3/12) were from other departments. The C. bracarensis isolate was grown from pus from a patient admitted to the surgery department in 2012 (Table 1). C. nivariensis recovered from blood cultures comprised 75% (9/12) of the strains with two isolates (2/12, 16.7%) recovered from CSF and one (8.3%) from AF.
Species Identification of C. nivariensis and C. bracarensis by Sequencer-Based Identification
The ITS sequences of the 13 study isolates exhibited 98.6 to 99.7% and 98.5% sequence identity to ITS sequences of reference isolates archived in the GenBank database (C. nivariensis CBS 10161 and C. bracarensis 153MT; Supplementary Table S1). Twelve isolates were identified as C. nivariensis and one as C. bracarensis (Table 1).
Analysis of the D1/D2 gene region sequences identified all 13 clinical isolates with 100% concordance with ITS sequencing results (Table 1). The D1/D2 region sequence of the 12 C. nivariensis isolate (KX499363 to KX499375) showed 99.6% sequence identity to those of C. nivariensis isolates in GenBank (C. nivariensis CBS 10161, accession no. EF056323.1) and the C. bracarensis sequence showed 99.7% sequence identity to that of C. bracarensis 153MT (accession no. AY589572.1). Results of ITS SCGE identified lengths of ITS1 and full-length ITS regions for C. nivariensis and C. bracarensis, which were identified as “cni-1,” “cni-2,” “cni-3,” “cni-4” and “cbr-1” as previously described (Hou et al., 2016).
Phylogenetic Analysis
The nucleotide sequence alignments within C. nivariensis and C. bracarensis, with reference to sequences of C. nivariensis type strain CBS 10161 and C. bracarensis type strain 153MT as the reference points, showed these species had high inter-species genetic similarity within ITS region (98.4 to 100%) and D1/D2 region (98.1 to 99.8%; Supplementary Table S1). The maximum-likelihood analysis of the ITS and D1/D2 region yielded similar results (Figures 1 and 2). For ITS, three groups of C. nivariensis were distinguished and our 12 isolates were in Groups I and II (Figure 1). Intra-species ITS diversity in Candida species has been described before (Merseguel et al., 2015). There was no intra-species D1/D2 diversity in 12 C. nivariensis isolates (Figure 2). Both ITS and D1/D2 sequencing can be used for the differentiation of C. glabrata, C. nivariensis and C. bracarensis.
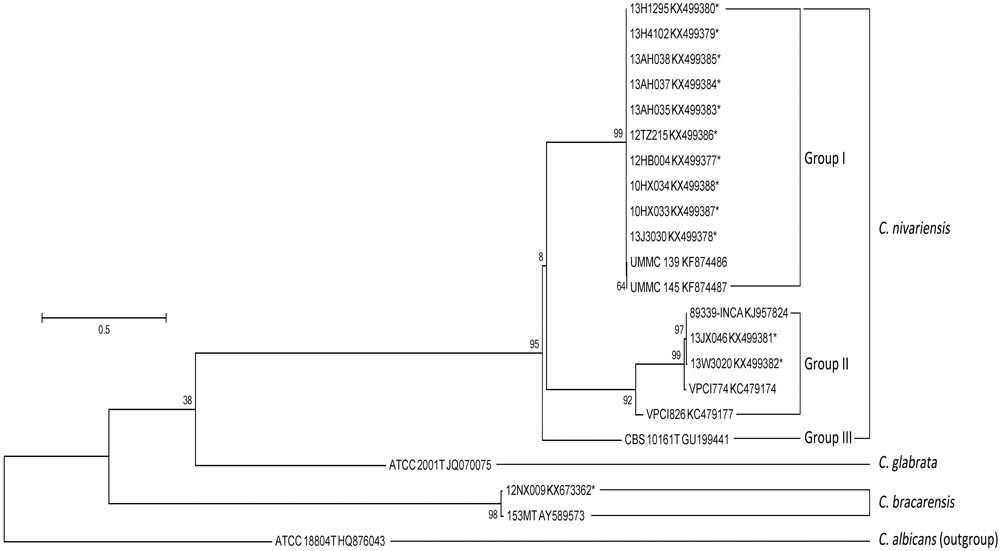
FIGURE 1. The maximum-likelihood tree of C. nivariensis and C. bracarensis with ITS sequences available in GenBank, using Candida albicans ATCC 18804 as an outgroup.
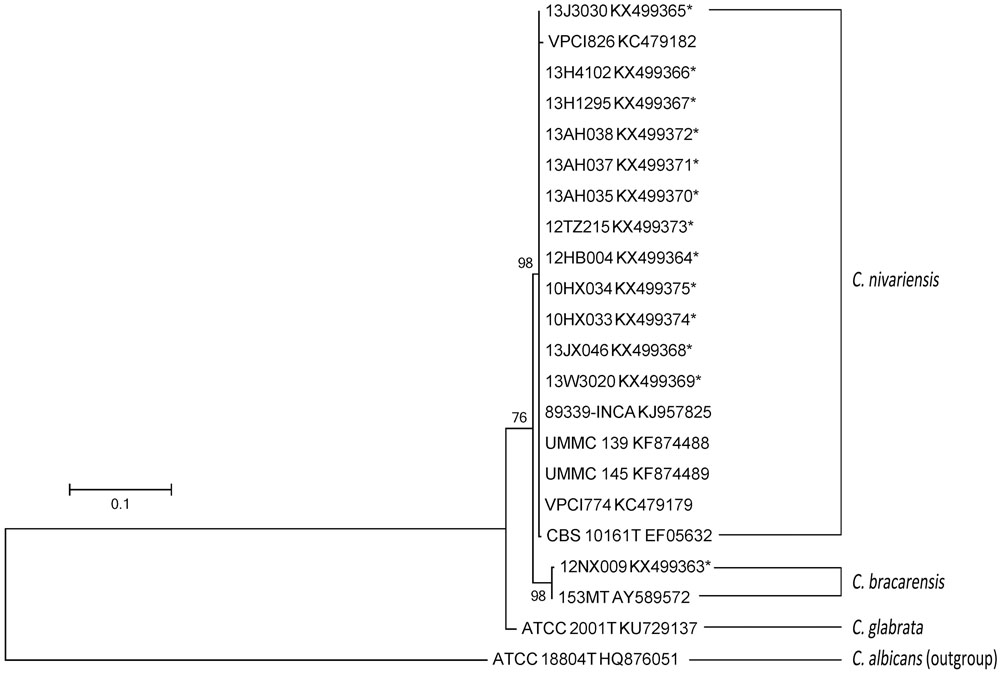
FIGURE 2. The maximum-likelihood tree of C. nivariensis and C. bracarensis with D1/D2 sequences available in GenBank, using Candida albicans ATCC 18804 as an outgroup.
Performance of Vitek-2 Compact and Two MALDI-TOF MS Systems
As compared with ITS sequencing, 11 C. nivariensis and the C. bracarensis isolates could not be identified by the Vitek 2 Compact system (bioMérieux). The remaining C. nivariensis was misidentified as C. glabrata (score = 94%; Table 1). All isolates were not identified by the Vitek MS system because C. nivariensis and C. bracarensis reference spectra were not included in the Vitek MS v2.0 database (Table 1). However, the Bruker system correctly identified all C. nivariensis isolates to the genus level with a MS score of <2.000 but ≥1.700. The C. bracarensis isolate was identified as such but with a MS score of 1.447 (Table 1).
Antifungal Susceptibility
The antifungal susceptibilities of the study isolates to nine antifungal agents are shown in Table 2. All isolates were S-DD to FLC (MIC range 0.5–4 μg/mL) and were classed as susceptible to echinocandins (MICs ≤ 0.06 μg/mL). All 13 isolates were also wild type to VRC (MICs ≤ 0.12 μg/mL), ITC (MICs ≤ 0.25 μg/mL), POS (MICs ≤ 0.5 μg/mL), AMB (MICs ≤ 2 μg/mL) and 5FC (MICs ≤ 0.25 μg/mL).
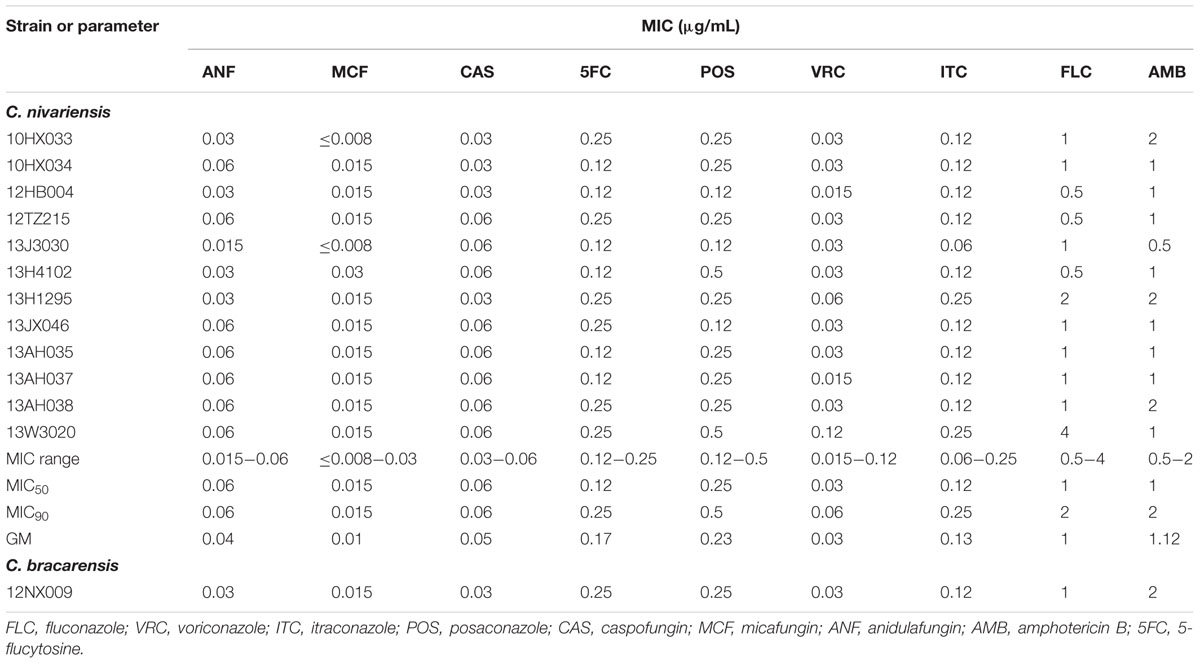
TABLE 2. In vitro susceptibility of 12 Candida nivariensis and one Candida bracarensis to nine antifungal agents.
C. nivariensis and C. bracarensis Reported in PubMed
By searching the literature in PubMed1 using “C. nivariensis or C. bracarensis” as the keywords, a total of 46 articles were found as of July 12, 2016. Among these, 15 and seven, full text articles reporting on the detection of C. nivariensis and C. bracarensis isolates were found, respectively; with 65 cases identified in 12 regions and 16 cases identified in seven regions, respectively (Supplementary Table S1). C. nivariensis caused mucosal superficial disease as well as IFDs and have been isolated from body sites including the mouth, toenails, urine, sputum, vagina, CSF, blood, BAL and other sterile body fluids. Two C. nivariensis isolates have been recovered from soil and bark. C. bracarensis has been isolated from throat, stool, abscess, sputum, vagina, blood and other sterile body fluids.
Data on C. nivariensis (12 articles were found) and C. bracarensis (six articles) antifungal susceptibility profiles are relatively limited. An overview of the published data is presented in Supplementary Table S2. Some strains exhibited high MICs to FLC alone or to all azoles, suggesting that the same mechanisms of resistance as found in C. glabrata may be involved (Fujita et al., 2007; Bishop et al., 2008; Borman et al., 2008; Gorton et al., 2013; Figueiredo-Carvalho et al., 2016). Further studies, using the C. nivariensis and C. bracarensis genome data, that are now available, might be useful to confirm this hypothesis (Angoulvant et al., 2016). MICs to echinocandins of C. nivariensis and C. bracarensis are overall low (MIC90 < 0.5 μg/ml for CAS; Supplementary Table S2). To date there are no reports on emergence of echinocandin-resistant C. nivariensis or C. bracarensis strains.
Discussion
Candida nivariensis and C. bracarensis are reportedly uncommon amongst clinical Candida isolates. Although the three species are closely related phylogenetically, DNA sequencing has shown that C. glabrata, C. nivariensis and C. bracarensis are sufficiently genetically distinct to justify their assignment as separate species (Figures 1 and 2) (Alcoba-Florez et al., 2005; Correia et al., 2006; Mirhendi et al., 2011).
In the present study, we have estimated the proportion of C. nivariensis and C. bracarensis amongst Candida strains cultured from patient with IFDs in China. Overall, during the 5-year CHIF-NET study period, C. nivariensis and C. bracarensis collectively represented only 0.11% (13/9673) of all yeast isolates, while 947 (9.79%, 947/9673) C. glabrata were collected. Among the 13 isolates, C. nivariensis (12/9673) was more frequently isolated compared with C. bracarensis (1/9673), which is in accordance with a retrospective study from four French university hospitals (2010–2014) where C. nivariensis and C. bracarensis were, respectively, identified in 0.12 and 0.01% of the 55,464 yeast strains (Angoulvant et al., 2016). Conversely, in the ARTEMIS DISK Global Antifungal Surveillance Study (2001–2006), where a total of 1,598 isolates phenotypically identified as C. glabrata were re-tested by peptide nucleic acid fluorescence in situ hybridization, only one C. nivariensis and two C. bracarensis were identified (Lockhart et al., 2009). In a Spanish study over a period of 2 years (2008–2009), a total of three (2%) of 143 C. glabrata clinical strains were identified as C. bracarensis and none as C. nivariensis (Cuenca-Estrella et al., 2011). Data for prevalence of C. nivariensis and C. bracarensis are directly influenced by the identification method used and comparisons between studies need to take into account the different identification methodologies used. In this study we have reinforced that molecular methods are the preferred approach for definitive identification of members within the species complex. Currently, ITS and D1/D2 regions were considered as universal DNA barcode markers for Candida spp. identification (Taverna et al., 2013). In this study, ITS sequencing was able to identify all 13 isolates and of note, concordance of results with ITS sequencing was 100% with those obtained by D1/D2 sequencing and ITS SCGE. An ITS SCGE assay has also shown promise for identification of these cryptic species and indeed to identify subtypes within these species. Twelve C. nivariensis were subtyped into four different ITS LTs, i.e., five cni-1, five cni-2, one cni-3 and one cni-4. The technique has the advantage of being simple to use (Hou et al., 2016).
Conventional phenotypic-based methods of classification of species within the C. glabrata complex have proven ineffective in accurately identifying these species as was the case in the present study (Alcoba-Florez et al., 2005; Correia et al., 2006), where all 13 isolates could not be identified by the Vitek 2 system (bioMérieux), except for one isolate “misidentified” as C. glabrata. Hence any unusual identification result should be confirmed by an alternative more discriminatory method. One simple method is to visualize the color of colonies on CHROMagar (Becton Dickinson, Heidelberg, Germany). C. nivariensis and C. bracarensis isolates yield white colonies on CHROMagar in contrast to the purple colonies of C. glabrata (Lockhart et al., 2009), and may be useful as a screen for members of the C. glabrata clade.
MALDI-TOF MS has become a routine identification tool for the identification of yeast and yeast-like organisms in many laboratories (Pinto et al., 2011; Gorton et al., 2013). In one study, four isolates collected from vagina were identified as C. nivariensis by the Bruker MS system (Bruker Daltoniks), with a log score between 1.802 and 2.086 (Aznar-Marin et al., 2015). MALDI-TOF MS has been reported to have the ability to distinguish within the C. glabrata clade, i.e., between C. glabrata, C. nivariensis and C. bracarensis, with scores of <1.700 as C. nivariensis (3/10 strains) and C. bracarensis (1/1 strain; Pinto et al., 2011). We found the database of the Vitek MS system (bioMérieux; IVD Knowledgebase version 2.0) to be limited by the absence of reference spectra for C. nivariensis and C. bracarensis. In comparison, the Bruker system (Bruker Daltoniks) was able to distinguish between all three members of the C. glabrata complex; the log scores of all C. nivariensis isolates were >1.700 but <2.000, while the log score of the C. bracarensis isolate was <1.700. The current Bruker Daltonik v3.1 database contains six MSPs for C. nivariensis and two MSPs for C. bracarensis. The addition of further reference spectra to the database may improve the utility of MALDI-TOF MS-based identification (Pinto et al., 2011). To improve the identification capacity of MALDI-TOF MS, it is important for MS databases to contain more reference mass spectra from type strains of different species, and also spectra representing different strains of the same species (Singhal et al., 2015).
Accurate identification of C. nivariensis and C. bracarensis in clinical samples is important, because the antifungal susceptibility patterns of C. nivariensis and C. bracarensis may differ from that of C. glabrata. In vitro susceptibility to azoles and 5FC appears to be lower for C. nivariensis than for C. glabrata (Fujita et al., 2007; Bishop et al., 2008; Borman et al., 2008). Although our isolates had low MICs to the azoles, echinocandins, AMB and 5FC, several studies have reported azoles resistance in these two species (Fujita et al., 2007; Bishop et al., 2008; Borman et al., 2008; Gorton et al., 2013; Figueiredo-Carvalho et al., 2016). For C. nivariensis, the MICs range of FLC has been 0.5 – >256 μg/ml, ITC MICs range 0.03 – >16 μg/ml, while the MICs range of FLC of C. bracarensis was 2–256 μg/ml and for ITC, 0.06 – >16 μg/ml (Supplementary Table S2). All isolates reported thus far have been susceptible to the echinocandins. Additional strains of C. nivariensis and C. bracarensis need to be studied to better understand their susceptibility profile and to determine whether these species have clinically significant differences in responses to antifungal therapy compared to C. glabrata. Moreover, studies from multiple geographical locations are needed to elucidate the epidemiology of infection, colonization and antifungal resistance of C. nivariensis and C. bracarensis.
Summary
This is the first systemic study regarding the epidemiology, identification and antifungal susceptibility profiles of C. nivariensis and C. bracarensis isolates in China. Our study reinforces the need for molecular identification of these two new and rarely species. Further improvements in the MALDI-TOF databases are needed to increase the identification accuracy. Studies from multiple geographical locations and additional data are required to better characterize their frequency, geographical distribution, susceptibility profiles and clinical features of infections due to C. nivariensis and C. bracarensis.
Author Contributions
XH, MX, and Y-CX conceived and designed the experiments, performed the experiments, analyzed the data, and wrote the paper. HW, S-YY, and XF performed the experiments and analyzed the data. SC and FK revised the paper critically for important intellectual content. XH, MX, SC, HW, S-YY, XF, FK, and Y-CX read and approved the final version of the manuscript.
Funding
This work was supported by Research Special Fund for Public Welfare Industry of Health (No. 201402001) and Innovation Fund of Peking Union Medical College (No. 2016-1001-15). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgment
We thank all the laboratories that participated in the CHIF-NET program in 2010–2014.
Abbreviations
5FC, 5-flucytosine; AF, ascitic fluid; AMB, amphotericin B; ANF, anidulafungin; BAL, bronchoalveolar lavage fluid; CAS, caspofungin; CBPs, clinical breakpoints; cbr, C. bracarensis; CHIF-NET, China Hospital Invasive Fungal Surveillance Net; CLSI, Clinical and Laboratory Standards Institute; cni, C. nivariensis; CSF, cerebrospinal fluid; ECVs, epidemiological cut-off values; FLC, fluconazole; ICU, intensive care unit; IFDs, invasive fungal diseases; ITC, itraconazole; ITS LTs, ITS length types; ITS, internal transcribed spacer; MALDI-TOF, matrix-assisted laser desorption ionization-time of flight; MCF, micafungin; MIC, minimum inhibitory concentration; MSPs, mass spectral profiles; NI, no identification; POS, posaconazole; SCGE, sequencer-based capillary gel electrophoresis; S-DD, susceptible-dose dependent; STs, sequence types; UN, unkown; VRC, voriconazole.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2017.00005/full#supplementary-material
Footnotes
References
Alcoba-Florez, J., Mendez-Alvarez, S., Cano, J., Guarro, J., Perez-Roth, E., and del Pilar Arevalo, M. (2005). Phenotypic and molecular characterization of Candida nivariensis sp. nov., a possible new opportunistic fungus. J. Clin. Microbiol. 43, 4107–4111. doi: 10.1128/JCM.43.8.4107-4111.2005
Amberg, D. C., Burke, D. J., and Strathern, J. N. (2005). Methods in Yeast Genetics: A Cold Spring Harbor Laboratory Course Manual. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press.
Angoulvant, A., Guitard, J., and Hennequin, C. (2016). Old and new pathogenic Nakaseomyces species: epidemiology, biology, identification, pathogenicity and antifungal resistance. FEMS Yeast Res. 16, fov114. doi: 10.1093/femsyr/fov114
Aznar-Marin, P., Galan-Sanchez, F., Marin-Casanova, P., Garcia-Martos, P., and Rodriguez-Iglesias, M. (2015). Candida nivariensis as a new emergent agent of vulvovaginal candidiasis: description of cases and review of published studies. Mycopathologia 181, 445–449. doi: 10.1007/s11046-015-9978-y
Bishop, J. A., Chase, N., Magill, S. S., Kurtzman, C. P., Fiandaca, M. J., and Merz, W. G. (2008). Candida bracarensis detected among isolates of Candida glabrata by peptide nucleic acid fluorescence in situ hybridization: susceptibility data and documentation of presumed infection. J. Clin. Microbiol. 46, 443–446. doi: 10.1128/JCM.01986-07
Borman, A. M., Petch, R., Linton, C. J., Palmer, M. D., Bridge, P. D., and Johnson, E. M. (2008). Candida nivariensis, an emerging pathogenic fungus with multidrug resistance to antifungal agents. J. Clin. Microbiol. 46, 933–938. doi: 10.1128/JCM.02116-07
Cendejas-Bueno, E., Kolecka, A., Alastruey-Izquierdo, A., Theelen, B., Groenewald, M., Kostrzewa, M., et al. (2012). Reclassification of the Candida haemulonii complex as Candida haemulonii (C. haemulonii group I), C. duobushaemulonii sp. nov. (C. haemulonii group II), and C. haemulonii var. vulnera var. nov.: three multiresistant human pathogenic yeasts. J. Clin. Microbiol. 50, 3641–3651. doi: 10.1128/JCM.02248-12
Clinical and Laboratory Standards Institute (2012). Reference Method for Broth Dilution Antifungal Susceptibility Testing of Yeasts: Fourth Informational Supplement M27-S4. Wayne, PA: CLSI.
Correia, A., Sampaio, P., James, S., and Pais, C. (2006). Candida bracarensis sp. nov., a novel anamorphic yeast species phenotypically similar to Candida glabrata. Int. J. Syst. Evol. Microbiol. 56(Pt 1), 313–317. doi: 10.1099/ijs.0.64076-0
Cuenca-Estrella, M., Gomez-Lopez, A., Isla, G., Rodriguez, D., Almirante, B., Pahissa, A., et al. (2011). Prevalence of Candida bracarensis and Candida nivariensis in a Spanish collection of yeasts: comparison of results from a reference centre and from a population-based surveillance study of candidemia. Med. Mycol. 49, 525–529. doi: 10.3109/13693786.2010.546373
Deak, E., Charlton, C. L., Bobenchik, A. M., Miller, S. A., Pollett, S., McHardy, I. H., et al. (2015). Comparison of the Vitek MS and Bruker Microflex LT MALDI-TOF MS platforms for routine identification of commonly isolated bacteria and yeast in the clinical microbiology laboratory. Diagn. Microbiol. Infect. Dis. 81, 27–33. doi: 10.1016/j.diagmicrobio.2014.09.018
Figueiredo-Carvalho, M. H., Ramos Lde, S., Barbedo, L. S., Chaves, A. L., Muramoto, I. A., Santos, A. L., et al. (2016). First description of Candida nivariensis in Brazil: antifungal susceptibility profile and potential virulence attributes. Mem. Inst. Oswaldo Cruz 111, 51–58. doi: 10.1590/0074-02760150376
Fujita, S., Senda, Y., Okusi, T., Ota, Y., Takada, H., Yamada, K., et al. (2007). Catheter-related fungemia due to fluconazole-resistant Candida nivariensis. J. Clin. Microbiol. 45, 3459–3461. doi: 10.1128/JCM.00727-07
Gorton, R. L., Jones, G. L., Kibbler, C. C., and Collier, S. (2013). Candida nivariensis isolated from a renal transplant patient with persistent candiduria-molecular identification using ITS PCR and MALDI-TOF. Med. Mycol. Case Rep. 2, 156–158. doi: 10.1016/j.mmcr.2013.10.001
Hou, X., Xiao, M., Chen, S. C., Wang, H., Zhang, L., Fan, X., et al. (2016). Sequencer-based capillary gel electrophoresis (SCGE) targeting the rDNA internal transcribed spacer (ITS) regions for accurate identification of clinically important yeast species. PLoS ONE 11:e0154385. doi: 10.1371/journal.pone.0154385
Lockhart, S. R., Messer, S. A., Gherna, M., Bishop, J. A., Merz, W. G., Pfaller, M. A., et al. (2009). Identification of Candida nivariensis and Candida bracarensis in a large global collection of Candida glabrata isolates: comparison to the literature. J. Clin. Microbiol. 47, 1216–1217. doi: 10.1128/JCM.02315-08
Merseguel, K. B., Nishikaku, A. S., Rodrigues, A. M., Padovan, A. C., e Ferreira, R. C., de Azevedo Melo, A. S., et al. (2015). Genetic diversity of medically important and emerging Candida species causing invasive infection. BMC Infect. Dis. 15:57. doi: 10.1186/s12879-015-0793-3
Mirhendi, H., Bruun, B., Schonheyder, H. C., Christensen, J. J., Fuursted, K., Gahrn-Hansen, B., et al. (2011). Differentiation of Candida glabrata, C. nivariensis and C. bracarensis based on fragment length polymorphism of ITS1 and ITS2 and restriction fragment length polymorphism of ITS and D1/D2 regions in rDNA. Eur. J. Clin. Microbiol. Infect. Dis. 30, 1409–1416. doi: 10.1007/s10096-011-1235-9
Pfaller, M. A., and Diekema, D. J. (2007). Epidemiology of invasive candidiasis: a persistent public health problem. Clin. Microbiol. Rev. 20, 133–163. doi: 10.1128/CMR.00029-06
Pfaller, M. A., and Diekema, D. J. (2012). Progress in antifungal susceptibility testing of Candida spp. by use of Clinical and Laboratory Standards Institute broth microdilution methods, 2010 to 2012. J. Clin. Microbiol. 50, 2846–2856. doi: 10.1128/JCM.00937-12
Pfaller, M. A., Diekema, D. J., Gibbs, D. L., Newell, V. A., Ellis, D., Tullio, V., et al. (2010). Results from the ARTEMIS DISK global antifungal surveillance study, 1997 to 2007: a 10.5-year analysis of susceptibilities of Candida species to fluconazole and voriconazole as determined by CLSI standardized disk diffusion. J. Clin. Microbiol. 48, 1366–1377. doi: 10.1128/JCM.02117-09
Pinto, A., Halliday, C., Zahra, M., van Hal, S., Olma, T., Maszewska, K., et al. (2011). Matrix-assisted laser desorption ionization-time of flight mass spectrometry identification of yeasts is contingent on robust reference spectra. PLoS ONE 6:e25712. doi: 10.1371/journal.pone.0025712
Singhal, N., Kumar, M., Kanaujia, P. K., and Virdi, J. S. (2015). MALDI-TOF mass spectrometry: an emerging technology for microbial identification and diagnosis. Front. Microbiol. 6:791. doi: 10.3389/fmicb.2015.00791
Tavanti, A., Davidson, A. D., Gow, N. A., Maiden, M. C., and Odds, F. C. (2005). Candida orthopsilosis and Candida metapsilosis spp. nov. to replace Candida parapsilosis groups II and III. J. Clin. Microbiol. 43, 284–292. doi: 10.1128/JCM.43.1.284-292.2005
Taverna, C. G., Bosco-Borgeat, M. E., Murisengo, O. A., Davel, G., Boite, M. C., Cupolillo, E., et al. (2013). Comparative analyses of classical phenotypic method and ribosomal RNA gene sequencing for identification of medically relevant Candida species. Mem. Inst. Oswaldo Cruz 108, 178–185. doi: 10.1590/0074-0276108022013009
Wang, H., Xiao, M., Chen, S. C., Kong, F., Sun, Z. Y., Liao, K., et al. (2012). In vitro susceptibilities of yeast species to fluconazole and voriconazole as determined by the 2010 National China Hospital Invasive Fungal Surveillance Net (CHIF-NET) study. J. Clin. Microbiol. 50, 3952–3959. doi: 10.1128/JCM.01130-12
Zhang, L., Xiao, M., Wang, H., Gao, R., Fan, X., Brown, M., et al. (2014). Yeast identification algorithm based on use of the Vitek MS system selectively supplemented with ribosomal DNA sequencing: proposal of a reference assay for invasive fungal surveillance programs in China. J. Clin. Microbiol. 52, 572–577. doi: 10.1128/JCM.02543-13
Keywords: Candida nivariensis, Candida bracarensis, ITS sequencing, D1/D2 sequencing, ITS sequencer-based capillary gel electrophoresis (SCGE), matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF), antifungal susceptibility
Citation: Hou X, Xiao M, Chen SC -A, Wang H, Yu S-Y, Fan X, Kong F and Xu Y-C (2017) Identification and Antifungal Susceptibility Profiles of Candida nivariensis and Candida bracarensis in a Multi-Center Chinese Collection of Yeasts. Front. Microbiol. 8:5. doi: 10.3389/fmicb.2017.00005
Received: 10 September 2016; Accepted: 03 January 2017;
Published: 19 January 2017.
Edited by:
Miguel Cacho Teixeira, Universidade de Lisboa, PortugalReviewed by:
Lucilla Iacumin, University of Udine, ItalyMehdi Razzaghi-Abyaneh, Pasteur Institute of Iran, Iran
Po-Ren Hsueh, National Taiwan University, Taiwan
Copyright © 2017 Hou, Xiao, Chen, Wang, Yu, Fan, Kong and Xu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ying-Chun Xu, eHljcHVtY2hAMTM5LmNvbQ==
 Xin Hou
Xin Hou Meng Xiao
Meng Xiao Sharon C.-A. Chen3
Sharon C.-A. Chen3 He Wang
He Wang Xin Fan
Xin Fan Fanrong Kong
Fanrong Kong