- 1State Key Laboratory of Veterinary Etiological Biology, Key Laboratory of Veterinary Parasitology of Gansu Province, Lanzhou Veterinary Research Institute, Chinese Academy of Agricultural Sciences, Lanzhou, China
- 2Faculty of Medicine and Health Sciences, School of Veterinary Medicine and Science, University of Nottingham, Sutton Bonington Campus, Loughborough, UK
- 3College of Animal Science and Veterinary Medicine, Shandong Agricultural University, Tai’an, China
- 4College of Animal Science and Veterinary Medicine, Heilongjiang Bayi Agricultural University, Daqing, China
Toxoplasma gondii is an obligatory intracellular apicomplexan protozoan which can infect any warm-blooded animal and causes severe diseases in immunocompromised individuals or infants infected in utero. The survival and success of this parasite require that it colonizes the host cell, avoids host immune defenses, replicates within an appropriate niche, and exits the infected host cell to spread to neighboring non-infected cells. All of these processes depend on the parasite ability to synthesis and export secreted proteins. Amongst the secreted proteins, rhoptry organelle proteins (ROPs) are essential for the parasite invasion and host cell manipulation. Even though the functions of most ROPs have been elucidated in the less virulent T. gondii (type II), the roles of ROPs in the highly virulent type I strain remain largely un-characterized. Herein, we investigated the contributions of 15 ROPs (ROP10, ROP11, ROP15, ROP20, ROP23, ROP31, ROP32, ROP33, ROP34, ROP35, ROP36, ROP40, ROP41, ROP46, and ROP47) to the infectivity of the high virulent type I T. gondii (RH strain). Using CRISPR-Cas9, these 15 ROPs genes were successfully disrupted and the effects of gene knockout on the parasite’s ability to infect cells in vitro and BALB/c mice in vivo were investigated. These results showed that deletions of these ROPs did not interfere with the parasite ability to grow in cultured human foreskin fibroblast cells and did not significantly alter parasite pathogenicity for BALB/c mice. Although these ROPs did not seem to be essential for the acute infectious stage of type I T. gondii in the mouse model, they might have different functions in other intermediate hosts or play different roles in other life cycle forms of this parasite due to the different expression patterns; this warrants further investigations.
Introduction
Toxoplasma gondii, an obligate intracellular protozoan pathogen, has the ability to infect a wide range of warm-blooded animals and humans (Pappas et al., 2009; Dubey, 2010; Andiappan et al., 2014; Chemoh et al., 2015; Liu et al., 2015). This parasite has a great medical importance because infection of immunocompromised individuals, including AIDS, cancer, and transplant patients, can lead to serious illness. Also, primary infection during pregnancy can lead to spontaneous abortion, fetal death, and significant congenital complications, such as brain and visual impairment (Pappas et al., 2009; Dubey, 2010; Andiappan et al., 2014; Chemoh et al., 2015; Liu et al., 2015). One of the most remarkable features of T. gondii is its ability to infect another eukaryotic cell and hijack the functions of such a host cell once within. Intracellular survival of this parasite is critically dependent on its ability to actively invade surrogate host cell, establish a replication-permissive vacuole and avoid host cell immune defenses (Melo et al., 2011; Hunter and Sibley, 2012).
Another important feature of T. gondii is the dramatic difference between strains in terms of their virulence in animal models (Su et al., 2003, 2012; Lorenzi et al., 2016) and also in human infections (Niedelman et al., 2012). This explains why disease that T. gondii causes in humans ranges from essentially asymptomatic to debilitating or even life threatening. There are likely many reasons for these differences, but evidence from animal studies and clinical studies in humans indicate that strain differences in the parasite likely play a major role and a number of genes were found to vary between strains and interact with the innate immunity in different ways (Saeij et al., 2006; Taylor et al., 2006; Behnke et al., 2011; Reese et al., 2011; Niedelman et al., 2012). It is intriguing that the major virulence determinants of this parasite are the effector secretory proteins derived from the apical highly specialized secretory organelles known as rhoptries (ROPs). Rhoptries contain many parts of the invasion machinery, located within the rhoptry necks and known as RONs, and a collection of effector proteins known as ROPs that are located within the rhoptry bulbs and intersect several host signaling pathways key to the pathogenesis and immune evasion, such as STAT signaling and immunity-related GTPases (IRGs or p47 GTPases) (Bradley and Sibley, 2007; Boothroyd and Dubremetz, 2008; Melo et al., 2011; Hunter and Sibley, 2012). Many ROPs contain a conserved serine/threonine protein kinase domain and may function as kinases or pseudokinases, which include only part of the catalytic triad, possibly modifying host cell pathways by phosphorylation of specific targets (Melo et al., 2011; Hunter and Sibley, 2012).
Comparative genomic and phylogenetic analyses revealed approximately 50 rhoptry kinases and pseudokinases (ROPKs) in T. gondii genome (Bradley et al., 2005; Peixoto et al., 2010; Talevich and Kannan, 2013). These ROPKs are polymorphic and can account for much of the differences observed in the virulence of different T. gondii strains. Some ROPKs have been shown to play roles in T. gondii virulence, establishing chronic T. gondii infection and manipulating the host innate immune response (Saeij et al., 2006; Taylor et al., 2006; Behnke et al., 2011; Butcher et al., 2011; Reese et al., 2011; Hunter and Sibley, 2012; Camejo et al., 2014; Etheridge et al., 2014; Fox et al., 2016; Jones et al., 2016; Kim et al., 2016). For instance, ROP5, ROP17, and ROP18 forming ROP5/17/18 complex function as virulence factors to prevent immune related GTPases accumulation at the parasitophorous vacuole (PV) (Saeij et al., 2006; Taylor et al., 2006; Behnke et al., 2011; Reese et al., 2011; Etheridge et al., 2014). ROP54, a type II T. gondii virulence factor, modulates the innate immune loading of GTP-binding protein 2, which does not act by interacting with the ROP5/17/18 complex (Kim et al., 2016). ROP16 can inhibit the production of the IL-12 in a parasite strain type dependent manner via activation of the host transcription factors STAT3 and STAT6 (Butcher et al., 2011) and can mediate human neuroblastoma SH-SY5Y apoptosis and cell cycle arrest in G1 phase by affecting the expression of Bax/Bcl-2 and p21/CDKs (Chang et al., 2015). More recently, deletion of 26 ROP gene loci encoding for 31 unique ROPK proteins in the less virulent T. gondii type II strain showed that most ROP proteins are essential for the establishment of chronic infection and that while some ROPs can serve as virulence factors by resisting innate immunity and other ROPs can influence chronic infection without affecting virulence (Fox et al., 2016), indicating the diverse roles that ROPs can play in mediating the parasite interaction with host.
Given the polymorphism of most ROPs and distinct biological and pathogenic traits of T. gondii strains, it is likely that these rhoptry paralogs have different functions in different strains of T. gondii (Su et al., 2012; Behnke et al., 2015; Lorenzi et al., 2016). Hence, determining how differences between T. gondii strains result in different pathogenicity in their hosts may help optimize treatment for individual patients. The contributions of most rhoptry kinases to host–pathogen interaction processes have been investigated in the less virulent type II strain, but the functions of most rhoptry kinases in the highly virulent type I strains remain largely unknown. Therefore, in this work, we utilized CRISPR-Cas9 technology to disrupt 15 ROP genes, namely rop10, rop11, rop15, rop20, rop23, rop31, rop32, rop33, rop34, rop35, rop36, rop40, rop41, rop46, and rop47 in the high virulent T. gondii RH strain (type I). Then, we evaluated the capacity of mutant strains of T. gondii deficient in these individual ROPs to induce lesions in vitro growth and to cause illness in experimentally infected BALB/c mice.
Materials and Methods
Parasites and Cell Culture
Tachyzoites of T. gondii (strain RH; type I) were grown in vitro in human foreskin fibroblast (HFF, ATCC® SCRC-1041TM) in a humidified incubator with 5% CO2 at 37°C. HFF cell culture was maintained by biweekly passage in Dulbecco’s modified Eagle medium (DMEM) supplemented with 10% fetal bovine serum (FBS) and 10 μg/mL gentamicin. Infected cells were syringe-lysed using a 27-gauge needle to release tachyzoites from host feeder cells into DMEM medium and then filtered using a 5 μm pore size Millipore filter.
Construction of ROP Knockouts by CRISPR-Cas9
The development of bacterial CRISPR-Cas9 system for use as a genome-editing tool has accelerated the pace of reverse genetic studies including rapid gene knockout (KO) and targeted modifications in T. gondii (Shen et al., 2014; Wang et al., 2016a). Hence, in this study targeted genetic manipulation of T. gondii was performed using CRISPR-Cas9 technology to disrupt 15 previously defined ROP genes, namely rop10, rop11, rop15, rop20, rop23, rop31, rop32, rop33, rop34, rop35, rop36, rop40, rop41, rop46, and rop47. The latter gene was annotated as a putative polo kinase in T. gondii genome and was proposed as rop47 (Talevich and Kannan, 2013). Details of all plasmids, primers and sgRNAs used in this study are provided in the Supplemental Material (Table S1). The CRISPR-Cas9 expression plasmids were constructed using methods previously described (Shen et al., 2014; Wang et al., 2016b). Briefly, sgRNAs were engineered into pSAG1::CAS9-U6::sgUPRT (Addgene #54467) using the Q5 Mutagenesis Kit (NEB). Resistance cassettes DHFR∗-Ts were amplified from the plasmid pUPRT-DHFR-D using Q5 DNA Polymerase (NEB). Freshly harvested tachyzoites were obtained, purified as described above and 40-μg CRISPR-Cas9 plasmids were cotransfected with the 10-μg purified DHFR∗-Ts amplicons. Successfully transgenic parasites were obtained by selection with 3 μM pyrimethamine (Sigma Aldrich, St. Louis, MO, USA) and single clones were screened using methods previously described (Shen et al., 2014; Wang et al., 2016b). Diagnostic PCRs were carried out with genome DNA as template to confirm the insertion of DHFR∗-Ts at the sgRNA-targeted coding region in the 15 endogenous ROP loci.
RT-PCR
First-strand cDNA was generated from total RNA of T. gondii using a PrimeScriptTM 1st Strand cDNA Synthesis Kit (TaKaRa). cDNAs from specific ROP KOs and wild type (WT) RH strain were tested by RT-PCR to amplify a small fragment around the insertion site in each targeted gene using specific primers listed in the Supplemental Material (Table S1).
Plaque Assay
For parasite plaque assays, 200 T. gondii tachyzoites of ROP-deficient or WT strains were inoculated into confluent HFF monolayers grown in six-well cell culture plates. The infected cell cultures were monitored microscopically over 7 days after infection. Then, the cells were fixed with 4% paraformaldehyde and stained for 10 min with 2% crystal violet to visualize and count plaques formed by the growing parasites by using inverted microscope (Wang et al., 2016b; Zhang et al., 2016).
Quantification of Ionophore-Induced Parasite Egress
Briefly, 1 × 105 freshly harvested tachyzoites were seeded into confluent HFF monolayers in 12-well cell culture plates and were incubated for 30–36 h. Then, cell monolayers were washed with sterile phosphate-buffered saline (PBS) followed by incubation with 3 μM calcium ionophore A23187 (Sigma) diluted in DMEM or an equivalent amount of DMSO as a solvent control for 10 min. Live videos were performed to document the progress of parasite egress as previously described (McCoy et al., 2012; Wang et al., 2016b).
Infectivity Studies with Mice
Mice (female inbred BALB/c, 7 weeks old) were purchased from Center of Laboratory Animals, Lanzhou Institute of Biological Products, Lanzhou, China. Mice were housed (five mice/cage) under a 12-h light:dark cycle (8:00–20:00) and under specific-pathogen-free conditions within the animal care facility until the end of the experiment. All mice were handled in strict accordance with the Good Animal Practice Requirements of the Animal Ethics Procedures and Guidelines of the People’s Republic of China. The study was approved by the Animal Ethics Committee of Lanzhou Veterinary Research Institute, Chinese Academy of Agricultural Science. Approximately, 200 freshly harvested tachyzoites were intraperitoneally injected into each BALB/c mice (10 mice/parasite strain) and the animals were monitored daily for signs of illness. Animal experiment began after 1 week of habituation and all efforts were made to minimize animal suffering.
Bioinformatic Analyses of T. gondii Rhoptries Proteins (ROPs)
Genomic data (the number of exons, transmembrane domains, and signal peptide) and transcriptomic data (cell cycle expression profiles, oocyst, tachyzoite, and bradyzoite developmental expression profiles) of 15 ROPs were obtained from http://ToxoDB.org (Gajria et al., 2007). Single nucleotide polymorphisms (SNP) from the coding sequence of three Toxoplasma strains (GT1, ME49, and VEG) were analyzed by the Clustal W program in the MEGA 5.0 (Tamura et al., 2011).
Results and Discussion
We have investigated the role of 15 ROP genes (rop10, rop11, rop15, rop20, rop23, rop31, rop32, rop33, rop34, rop35, rop36, rop40, rop41, rop46, and rop47) in the type I T. gondii (RH strain), to determine whether they play critical roles in T. gondii-host interaction, on replication in human cells and virulence in the BALB/c mice.
Generation of ROP Knockouts in Type I RH Strain
To assess the contribution of 15 ROPs to T. gondii infection, CRISPR-Cas9 system was used to disrupt these genes via the insertion of DHFR∗-Ts at the sgRNA-targeted coding region in the 15 endogenous ROP loci (Figure 1A). Diagnostic PCR with expected small KO strains could be yielded in the WT RH strains, while a large fragment containing about 3.3 kb DHFR∗-Ts amplification was not amplified in the KO strains with 30 s extension time (Figures 1B,C), but could be yielded with 4 min extension time (Figures 1D,E). Also, mRNA from the targeted sites was not detected by RT-PCR in the KO strains for each of the ROP genes (Figures 1F,G). Insertion of a large DHFR∗-Ts fragment in the interested coding sequence caused shift mutations followed by gene disruption. The generation of the 15 ROPs KOs was successfully and rapidly achieved.
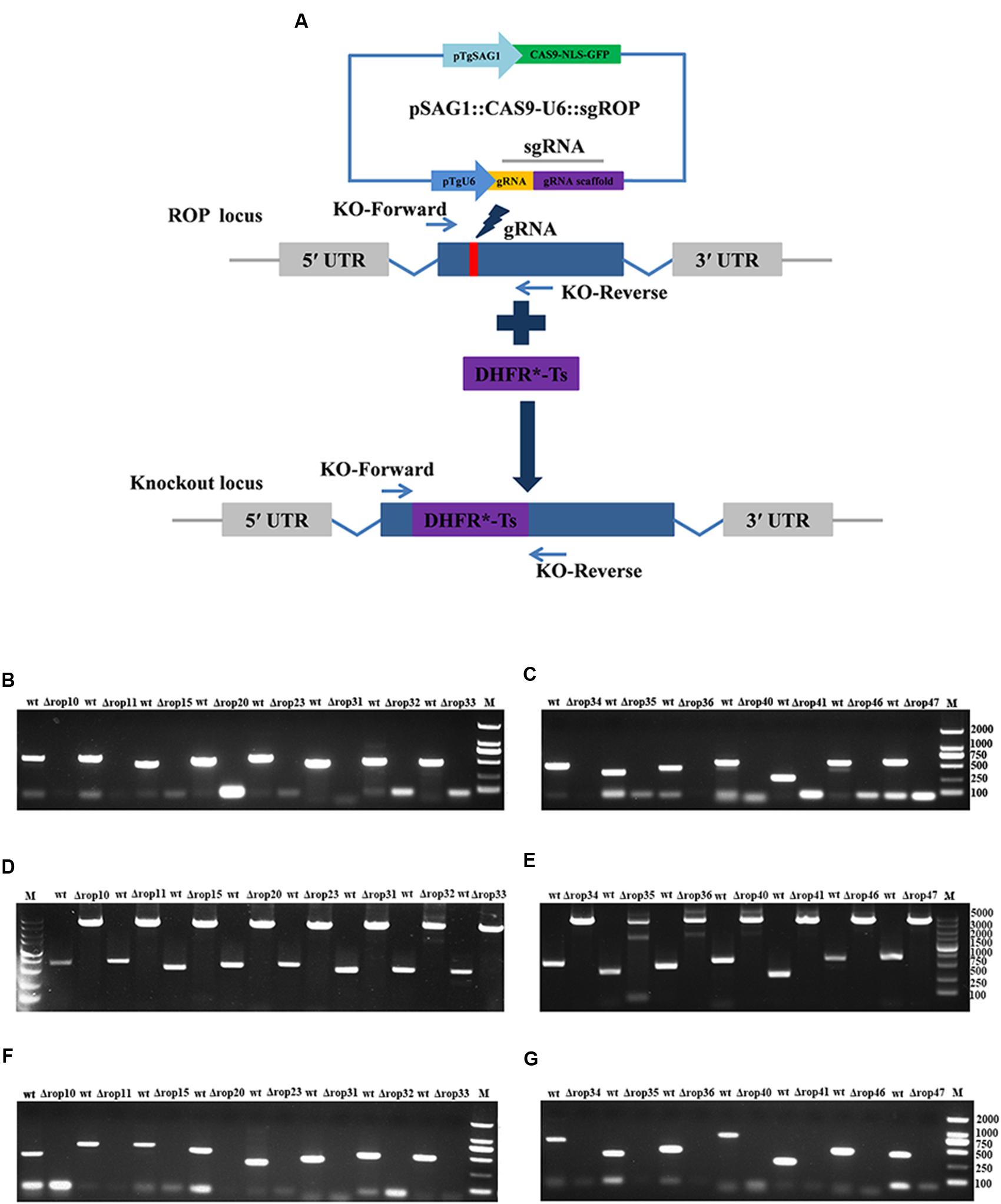
FIGURE 1. Toxoplasma gondii rhoptry organelle proteins (ROP) knockout (KO) mutant construction. (A) Schematic illustration of CRISPR-Cas9-mediated gene KO by insertion of the pyrimethamine-resistant DHFR∗-Ts in the ROPs coding sequence. (B–E) KO forward and KO reverse primers were used to amplify the small fragment with 30 s extension time (B,C) or the large fragment with 4 min extension time (D,E). Diagnostic PCR demonstrates DHFR∗-Ts integration and ROP genes disruption in a representative clone compared with wild-type RH strain. (F,G) RT-PCR screening of ROP gene KOs. Small fragment around the insertion site for each ROP gene was not detected by RT-PCR demonstrating that the coding sequences were disrupted.
Phenotypic Analyses of ROPs Knockouts In vitro
To test the phenotypes of ROP KOs during the parasite’s lytic cycle, plaque assay, an assay that captures invasion, egress, motility and replication was performed to compare the intracellular growth of mutants to the WT RH strain. Over a 7-day period of the parasite lytic cycle, the number and the size of individual plaques developed by each strain were recorded. There was no significant difference between all ROP KOs and the RH WT strain (Supplementary Figure S1). We also evaluated whether there was any alteration in ionophore-induced egress (IIE) induced by ROP gene deletion. Calcium ionophore was used to stimulate intracellular tachyzoites to rapidly egress from their PV at any stage during the lytic cycle given that the natural egress normally occurs when there are 64 or more parasites inside the PV (Black and Boothroyd, 2000; Black et al., 2000). Based on time-lapse imaging analysis of infected cultures stimulated with 3 μM calcium ionophore A23187 or equivalent amount of DMSO as control over 10 min revealed that all 15 ROP KO strains and the WT RH strain rapidly egressed from their PVs within 4 min after the stimulation with 3 μM calcium ionophore A23187 (Supplementary Figure S2), while all 15 ROP KO strains and the WT RH strain were still in their PVs after the stimulation with equivalent amount of DMSO. These findings show that infection of HFF cells with these ROP KO strains did not entail a significant reduction in parasite multiplication or any significant alteration in the parasite’s growth pattern. This result was not surprising, as some ROP genes have previously been shown to be not required for the parasite’s ability to invade and colonize HFF cells, such as ROP5 and ROP18 (Saeij et al., 2006; Taylor et al., 2006; Behnke et al., 2011; Reese et al., 2011; Etheridge et al., 2014; Fox et al., 2016).
Characterization of ROP Knockouts In vivo
Even though ROP5, ROP17, and ROP18, as well as other ROPs have been shown as important virulence factors, deletion of these genes did not or slightly affect the parasite infection cycle in vitro. We were also interested in whether deletion of these ROPs in RH strain would affect its virulence as described previously in ROP5-, ROP17- and ROP18-deficient strains (Saeij et al., 2006; Taylor et al., 2006; Behnke et al., 2011; Reese et al., 2011; Etheridge et al., 2014). Approximately, 200 WT RH tachyzoites or an equal number of the generated ROP KOs were used to infect BALB/c mice. Overall, deletion of the 15 ROPs in type I T. gondii RH strain did not cause significant impact on the parasite virulence in mice. Compared to mice infected with WT strain all mice infected with ROP KO strains showed similar mortality or slightly reduced virulence with mice dying between days 7 and 11 post infection (Figure 2). There may be some level of function redundancy in these ROPs, as some ROPs in type II strain both have the ability to increase cyst burdens in order to establish chronic infection in mice (Fox et al., 2016; Jones et al., 2016). Perhaps, the effects of deleting these ROP genes in the T. gondii RH did not result in phenotype changes due to function redundancy, or alternatively these genes may play no significant role in the virulence in mouse model that we have tested. For example, these ROPs genes may serve as virulence factors in other intermediate hosts. However, this question is quite challenging because T. gondii can infect almost all warm-blooded animals.
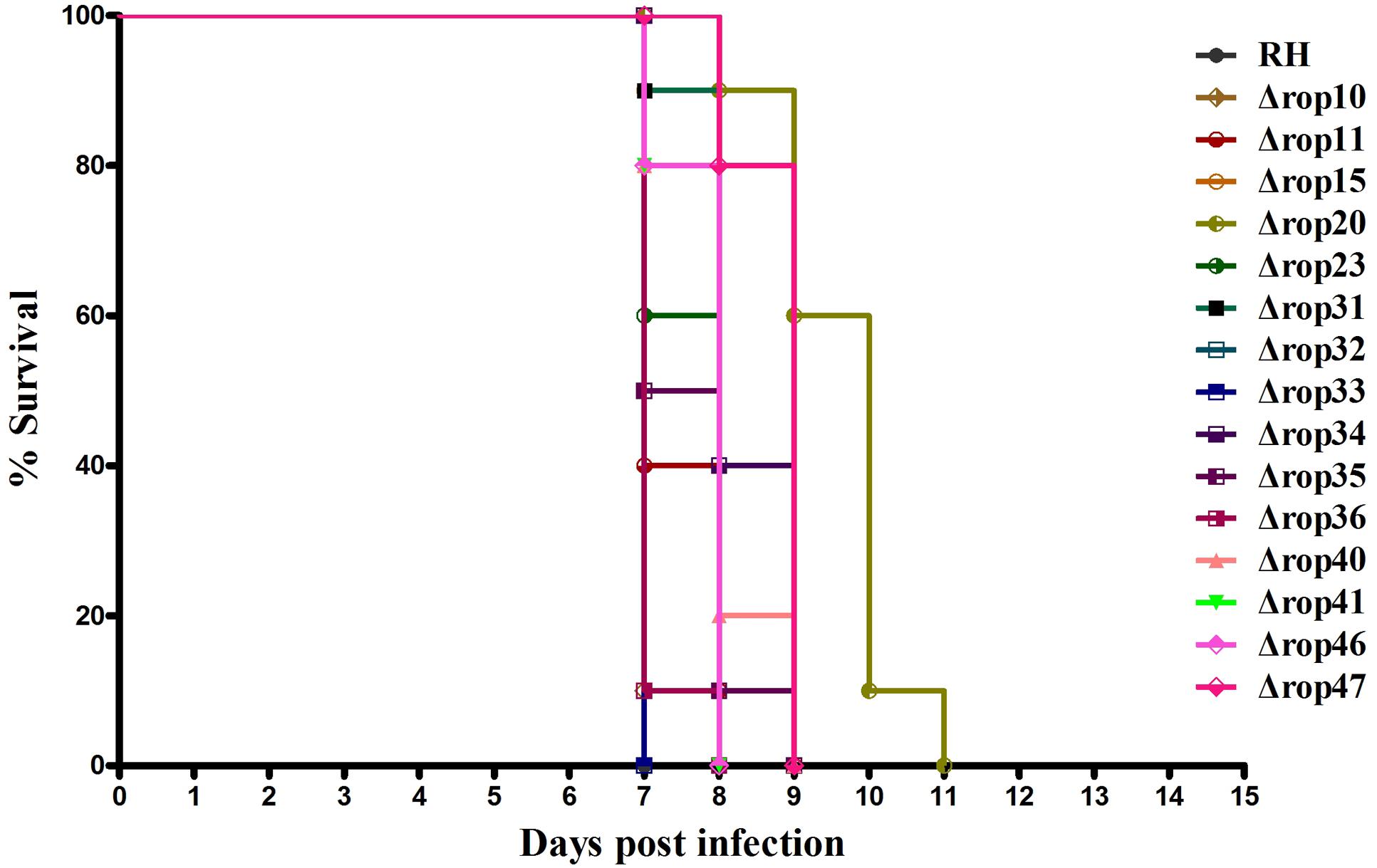
FIGURE 2. Survival curves of BALB/c mice infected with T. gondii (RH) wild-type strain or ROP KO strains. The various groups of BALB/c mice were challenged with 200 tachyzoites of the indicated strains. Each group was composed of 10 mice and survival time was monitored daily for 11 days after challenge.
Sequence and Expression Analyses of ROPs in T. gondii
Previous studies have identified ∼50 ROPs in T. gondii genome and some of these, such as ROP5, ROP16, ROP17, and ROP18, are typically expressed in a characteristic cell cycle-dependent profile where their mRNA expression levels were found to increase to maximum 2–3 h post-thymidine release (S to M phase) and then dramatically decrease in early G1 (minimum 6 h) followed by a rise again in the next S phase (peak 10 h) (Behnke et al., 2010; Camejo et al., 2014). However, some ROPs did not exhibit any obvious cyclical expression profile, such as ROP21, ROP27, and ROP28 (Jones et al., 2016). To examine the expression of the 15 ROP genes tested in the present study during cell cycle expression, we analyzed microarray data obtained from ToxoDB1 (Behnke et al., 2010). Profiling the cell cycle expression profiles of the 15 ROPs revealed that several ROPs (ROP10, ROP11, ROP15, ROP20, and ROP40) are expressed in a characteristic cell cycle profile while most are not (ROP23, ROP31, ROP32, ROP33, ROP34, ROP35, ROP36, ROP41, and ROP46) (Figure 3A). Other typical bioinformatic features for ROPs are expressed from single exon genes and have a predicted signal peptide that mediates their trafficking to the parasitophorous vacuole membrane (PVM) or the host cell (Camejo et al., 2014; Lorenzi et al., 2016). However, we found that some of these ROPs are encoded by multi-exons and some do not have a predicted signal peptide (Table 1). Our analysis also suggests that some annotated ROPs are not consistent with and may have different functions from the canonical ROPs. We also analyzed the transcriptomic levels of 15 ROP genes during various life cycle forms of the parasite2 based on a previous published transcriptomic data (Fritz et al., 2012). This analysis revealed the transcriptomic expression level of some ROPs (ROP32, ROP33, ROP34, ROP35, and ROP36) to be constitutively expressed while others (ROP10, ROP11, ROP15, ROP20, ROP23, ROP31, ROP40, ROP41, and ROP46) are differentially expressed during Toxoplasma development (Figure 3B). The presence of some ROPs that are differentially expressed during the parasite developmental cycle and other ROPs that are expressed throughout the parasite life cycle indicate that these ROPs may play different roles during different life cycle stages, such as the sexual phase in the cat intestine or during oocyst development. However, the RH strain lost its capacity to complete the life cycle in cat (Frenkel et al., 1976). Using CRISPR-Cas9 in other cat-competent type I T. gondii strains, such as the GT1, would allow for testing the functions of ROPs during sexual development or during oocyst development.
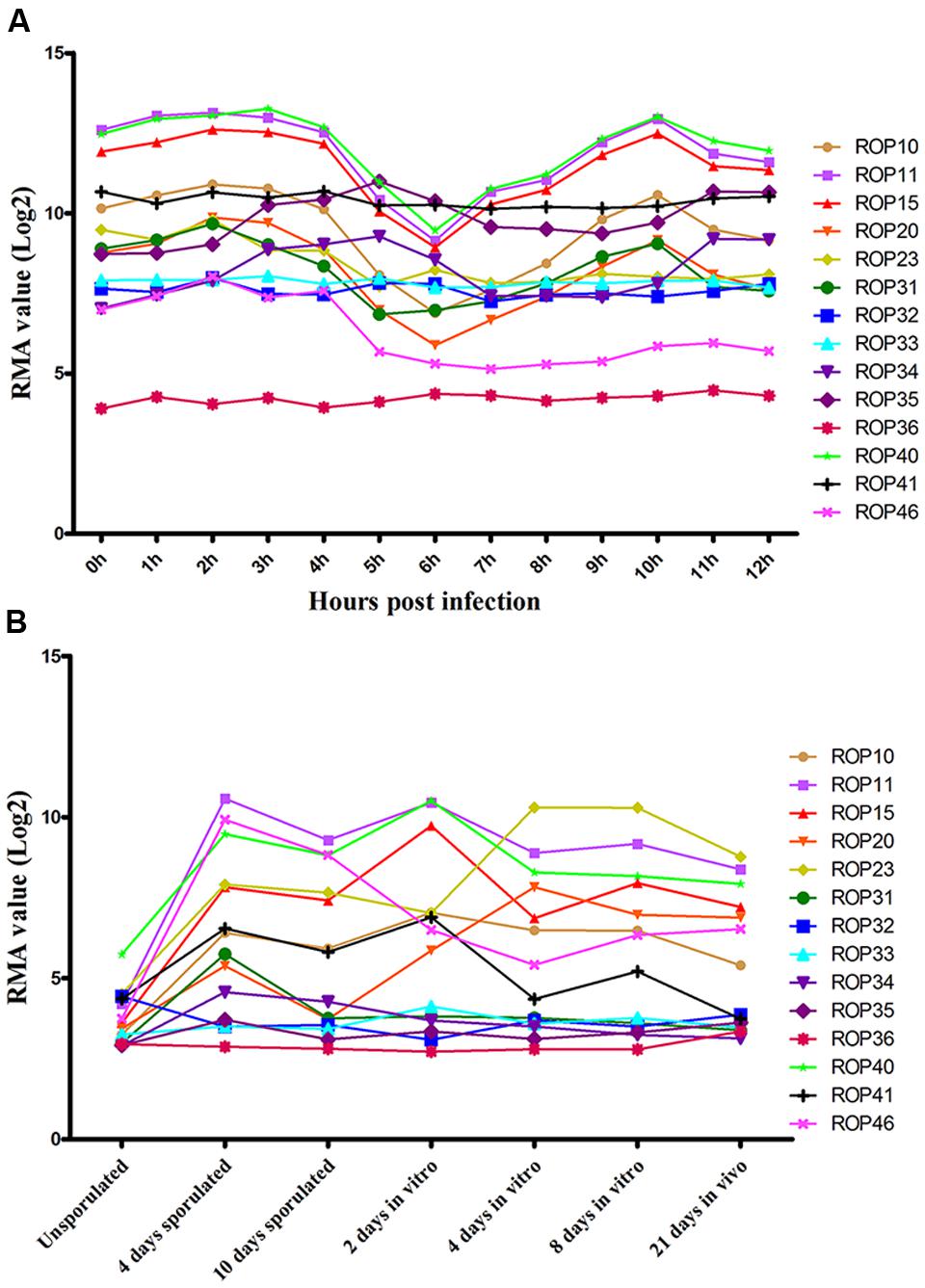
FIGURE 3. Expression analysis of ROPs in T. gondii (Note: data were not available for ROP47). (A) Toxoplasma RH cell cycle microarray expression data for ROP genes were obtained from http://ToxoDB.org. h, the time post-thymidine release; S, synthesis; M, mitosis; C, cytokinesis; G1, G1 phase of cell cycle; RMA, robust multiarray average algorithm. (B) Transcriptomic expression data of Toxoplasma ROP genes of the parasite developmental stages, oocyst, tachyzoite, and bradyzoite, of type II strain M4 were obtained from http://ToxoDB.org. Sources of the RNAs: oocysts recovered from cat feces included day 0 (unsporulated) oocysts, day 4 post-induction of sporulation (4 days sporulated) and day 10 post-induction of sporulation (10 days sporulated). In vitro, 2 dpi tachyzoite, 4 and 8 dpi bradyzoite samples (2 days in vitro, 4 days in vitro, 8 days in vitro) are from separately infected cultures of HFF cell. In vivo, 21 dpi bradyzoite cysts (21 days in vivo) harvested from the brains of mice infected with oocysts.
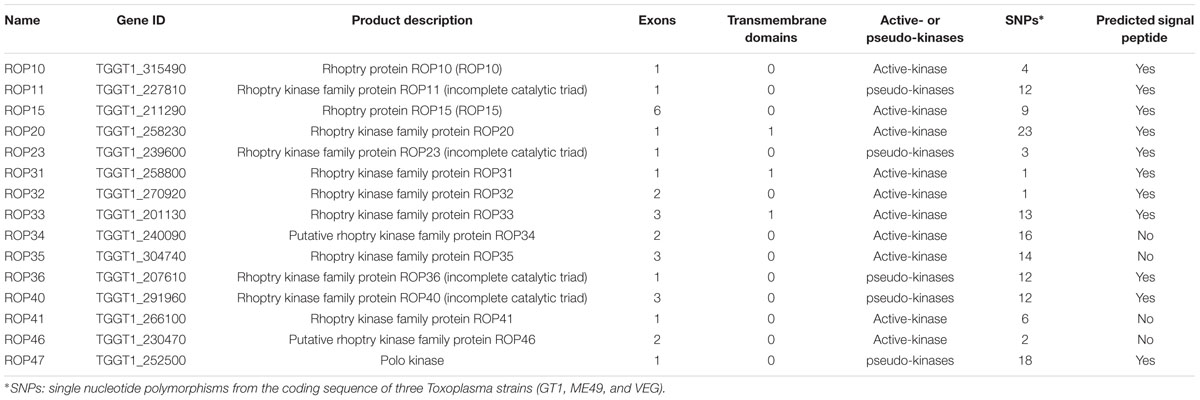
TABLE 1. Characterization sequence features of rhoptry organelle proteins (ROPs) of Toxoplasma gondii.
Conclusion
Toxoplasma synthesizes and exports several proteins, such as rhoptry kinases (ROPs), to the host cells to ensure parasite’s invasion and survival within the host. The present study investigated the contribution of 15 ROPs to T. gondii infection process in vitro and in vivo by disrupting individual ROP genes using the CRISPR-Cas9 approach. Our results showed that deletion of these ROPs does not seem to interfere with the parasite ability growth in vitro or to induce illness in experimentally infected BALB/c mice. Infection of BALB/c mice with the KO mutants demonstrated that the deletion of single ROP genes was insufficient to induce a high level of attenuation compared with the parental parasite strain. The different expression patterns of some ROPs suggest that they may have different functions during various stages of the parasite’s life cycle. Elucidation of the roles of these ROPs in the various life cycle stages of T. gondii and in other strains is needed.
Ethics Statement
The study was approved by the Animal Ethics Committee of Lanzhou Veterinary Research Institute, Chinese Academy of Agricultural Science.
Author Contributions
X-QZ and S-YH designed this study and critically revised the manuscript. J-LW, T-TL, and KC performed the experiments, analyzed data and drafted the manuscript. HE, W-NZ, and D-MY participated in manuscript revision. All the authors read and approved the final manuscript.
Funding
Project support was provided, in part, by the Natural Science Foundation of Gansu Province for Distinguished Young Scholars (Grant No. 1506RJDA133), the National Natural Science Foundation of China (Grant Nos. 31472184 and 31230073) and the Fundamental Research Funds of Chinese Academy of Agricultural Sciences (Grant No. Y2016JC05).
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgment
We are grateful to Professor Bang Shen (Huazhong Agricultural University) for providing pSAG1::CAS9-U6::sgUPRT and pUPRT-DHFR-D vectors.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2017.00084/full#supplementary-material
FIGURE S1 | Phenotype analyses of wild type RH and ROP-deficient parasites. Two hundred freshly harvested Toxoplasma tachyzoites of wild type RH and ROP-deficient parasites were seeded in six-well plates. After 7 days of growth, the plaque numbers were recorded. All parasite strains were performed three times independently, each with triplicates.
FIGURE S2 | Egress phenotype of the parental strain RH and one of ROP mutants (ROP10). Live time-lapse microscopy of parental parasite (A) and one of ROP mutants (ROP10) (B) show both parental parasite and ROP10 mutant can activate egress within 2 min after addition of 3 μM calcium ionophore A23187. The egress phenotype of other ROP mutants is not shown due to the similar results.
Footnotes
References
Andiappan, H., Nissapatorn, V., Sawangjaroen, N., Khaing, S. L., Salibay, C. C., Cheung, M. M., et al. (2014). Knowledge and practice on Toxoplasma infection in pregnant women from Malaysia, Philippines, and Thailand. Front. Microbiol. 5:291. doi: 10.3389/fmicb.2014.00291
Behnke, M. S., Khan, A., Lauron, E. J., Jimah, J. R., Wang, Q., Tolia, N. H., et al. (2015). Rhoptry proteins ROP5 and ROP18 are major murine virulence factors in genetically divergent South American strains of Toxoplasma gondii. PLoS Genet. 11:e1005434. doi: 10.1371/journal.pgen.1005434
Behnke, M. S., Khan, A., Wootton, J. C., Dubey, J. P., Tang, K., and Sibley, L. D. (2011). Virulence differences in Toxoplasma mediated by amplification of a family of polymorphic pseudokinases. Proc. Natl. Acad. Sci. U.S.A. 108, 9631–9636. doi: 10.1073/pnas.1015338108
Behnke, M. S., Wootton, J. C., Lehmann, M. M., Radke, J. B., Lucas, O., Nawas, J., et al. (2010). Coordinated progression through two subtranscriptomes underlies the tachyzoite cycle of Toxoplasma gondii. PLoS ONE 5:e12354. doi: 10.1371/journal.pone.0012354
Black, M. W., Arrizabalaga, G., and Boothroyd, J. C. (2000). Ionophore-resistant mutants of Toxoplasma gondii reveal host cell permeabilization as an early event in egress. Mol. Cell. Biol. 20, 9399–9408. doi: 10.1111/j.1462-5822.2009.01357.x
Black, M. W., and Boothroyd, J. C. (2000). Lytic cycle of Toxoplasma gondii. Microbiol. Mol. Biol. Rev. 64, 607–623. doi: 10.1128/MMBR.64.3.607-623.2000
Boothroyd, J. C., and Dubremetz, J. F. (2008). Kiss and spit: the dual roles of Toxoplasma rhoptries. Nat. Rev. Microbiol. 6, 79–88. doi: 10.1038/nrmicro1800
Bradley, P. J., and Sibley, L. D. (2007). Rhoptries: an arsenal of secreted virulence factors. Curr. Opin. Microbiol. 10, 582–587. doi: 10.1016/j.mib.2007.09.013
Bradley, P. J., Ward, C., Cheng, S. J., Alexander, D. L., Coller, S., Coombs, G. H., et al. (2005). Proteomic analysis of rhoptry organelles reveals many novel constituents for host-parasite interactions in Toxoplasma gondii. J. Biol. Chem. 280, 34245–34258. doi: 10.1074/jbc.M504158200
Butcher, B. A., Fox, B. A., Rommereim, L. M., Kim, S. G., Maurer, K. J., Yarovinsky, F., et al. (2011). Toxoplasma gondii rhoptry kinase ROP16 activates STAT3 and STAT6 resulting in cytokine inhibition and arginase-1-dependent growth control. PLoS Pathog. 7:e1002236. doi: 10.1371/journal.ppat.1002236
Camejo, A., Gold, D. A., Lu, D., McFetridge, K., Julien, L., Yang, N., et al. (2014). Identification of three novel Toxoplasma gondii rhoptry proteins. Int. J. Parasitol. 44, 147–160. doi: 10.1016/j.ijpara.2013.08.002
Chang, S., Shan, X., Li, X., Fan, W., Zhang, S. Q., Zhang, J., et al. (2015). Toxoplasma gondii rhoptry protein ROP16 mediates partially SH-SY5Y cells apoptosis and cell cycle arrest by directing Ser15/37 phosphorylation of p53. Int. J. Biol. Sci. 11, 1215–1225. doi: 10.7150/ijbs.10516
Chemoh, W., Sawangjaroen, N., Siripaitoon, P., Andiappan, H., Hortiwakul, T., Sermwittayawong, N., et al. (2015). Toxoplasma gondii - prevalence and risk factors in HIV-infected patients from Songklanagarind Hospital, Southern Thailand. Front. Microbiol. 6:1304. doi: 10.3389/fmicb.2015.01304
Etheridge, R. D., Alaganan, A., Tang, K., Lou, H. J., Turk, B. E., and Sibley, L. D. (2014). The Toxoplasma pseudokinase ROP5 forms complexes with ROP18 and ROP17 kinases that synergize to control acute virulence in mice. Cell Host Microbe. 15, 537–550. doi: 10.1016/j.chom.2014.04.002
Fox, B. A., Rommereim, L. M., Guevara, R. B., Fall, A., Hortua Triana, M. A., Sun, Y., et al. (2016). The Toxoplasma gondii rhoptry kinome is essential for chronic infection. MBio. 7:e00193–16. doi: 10.1128/mBio.00193-116
Frenkel, J. K., Dubey, J. P., and Hoff, R. L. (1976). Loss of stages after continuous passage of Toxoplasma gondii and Besnoitia jellisoni. J. Protozool. 23, 421–424.
Fritz, H. M., Buchholz, K. R., Chen, X., Durbin-Johnson, B., Rocke, D. M., Conrad, P. A., et al. (2012). Transcriptomic analysis of Toxoplasma development reveals many novel functions and structures specific to sporozoites and oocysts. PLoS ONE 7:e29998. doi: 10.1371/journal.pone.0029998
Gajria, B., Bahl, A., Brestelli, J., Dommer, J., Fischer, S., Gao, X., et al. (2007). ToxoDB: an integrated Toxoplasma gondii database resource. Nucl. Acids Res. 36, D553–556. doi: 10.1093/nar/gkm981
Hunter, C. A., and Sibley, L. D. (2012). Modulation of innate immunity by Toxoplasma gondii virulence effectors. Nat. Rev. Microbiol. 10, 766–778. doi: 10.1038/nrmicro2858
Jones, N. G., Wang, Q., and Sibley, L. D. (2016). Secreted protein kinases regulate cyst burden during chronic Toxoplasmosis. Cell. Microbiol. doi: 10.1111/cmi.12651 [Epub ahead of print].
Kim, E. W., Nadipuram, S. M., Tetlow, A. L., Barshop, W. D., Liu, P. T., Wohlschlegel, J. A., et al. (2016). The rhoptry pseudokinase ROP54 modulates Toxoplasma gondii virulence and host GBP2 Loading. mSphere. 1:e00045–16. doi: 10.1128/mSphere.00045-16
Liu, Q., Wang, Z. D., Huang, S. Y., and Zhu, X. Q. (2015). Diagnosis of toxoplasmosis and typing of Toxoplasma gondii. Parasit. Vectors. 8:292. doi: 10.1186/s13071-015-0902-906
Lorenzi, H., Khan, A., Behnke, M. S., Namasivayam, S., Swapna, L. S., Hadjithomas, M., et al. (2016). Local admixture of amplified and diversified secreted pathogenesis determinants shapes mosaic Toxoplasma gondii genomes. Nat. Commun. 7:10147. doi: 10.1038/ncomms10147
McCoy, J. M., Whitehead, L., van Dooren, G. G., and Tonkin, C. J. (2012). TgCDPK3 regulates calcium-dependent egress of Toxoplasma gondii from host cells. PLoS Pathog. 8:e1003066. doi: 10.1371/journal.ppat.1003066
Melo, M. B., Jensen, K. D., and Saeij, J. P. (2011). Toxoplasma gondii effectors are master regulators of the inflammatory response. Trends Parasitol. 27, 487–495. doi: 10.1016/j.pt.2011.08.001
Niedelman, W., Gold, D. A., Rosowski, E. E., Sprokholt, J. K., Lim, D., Farid Arenas, A., et al. (2012). The rhoptry proteins ROP18 and ROP5 mediate Toxoplasma gondii evasion of the murine, but not the human, interferon-gamma response. PLoS Pathog. 8:e1002784. doi: 10.1371/journal.ppat.1002784
Pappas, G., Roussos, N., and Falagas, M. E. (2009). Toxoplasmosis snapshots: global status of Toxoplasma gondii seroprevalence and implications for pregnancy and congenital toxoplasmosis. Int. J. Parasitol. 39, 1385–1394. doi: 10.1016/j.ijpara.2009.04.003
Peixoto, L., Chen, F., Harb, O. S., Davis, P. H., Beiting, D. P., Brownback, C. S., et al. (2010). Integrative genomic approaches highlight a family of parasite-specific kinases that regulate host responses. Cell Host Microbe. 8, 208–218. doi: 10.1016/j.chom.2010.07.004
Reese, M. L., Zeiner, G. M., Saeij, J. P., Boothroyd, J. C., and Boyle, J. P. (2011). Polymorphic family of injected pseudokinases is paramount in Toxoplasma virulence. Proc. Natl. Acad. Sci. U.S.A. 108, 9625–9630. doi: 10.1073/pnas.1015980108
Saeij, J. P., Boyle, J. P., Coller, S., Taylor, S., Sibley, L. D., Brooke-Powell, E. T., et al. (2006). Polymorphic secreted kinases are key virulence factors in toxoplasmosis. Science 314, 1780–1783. doi: 10.1126/science.1133690
Shen, B., Brown, K. M., Lee, T. D., and Sibley, L. D. (2014). Efficient gene disruption in diverse strains of Toxoplasma gondii using CRISPR/CAS9. mBio. 5:e01114–14. doi: 10.1128/mBio.01114-1114
Su, C., Evans, D., Cole, R. H., Kissinger, J. C., Ajioka, J. W., and Sibley, L. D. (2003). Recent expansion of Toxoplasma through enhanced oral transmission. Science 299, 414–416. doi: 10.1126/science.1078035
Su, C., Khan, A., Zhou, P., Majumdar, D., Ajzenberg, D., Dardé, M. L., et al. (2012). Globally diverse Toxoplasma gondii isolates comprise six major clades originating from a small number of distinct ancestral lineages. Proc. Natl. Acad. Sci. U.S.A. 109, 5844–5849. doi: 10.1073/pnas.1203190109
Talevich, E., and Kannan, N. (2013). Structural and evolutionary adaptation of rhoptry kinases and pseudokinases, a family of coccidian virulence factors. BMC Evol. Biol. 13:117. doi: 10.1186/1471-2148-13-117
Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M., and Kumar, S. (2011). MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739. doi: 10.1093/molbev/msr121
Taylor, S., Barragan, A., Su, C., Fux, B., Fentress, S. J., Tang, K., et al. (2006). A secreted serine-threonine kinase determines virulence in the eukaryotic pathogen Toxoplasma gondii. Science 314, 1776–1780. doi: 10.1126/science.1133643
Wang, J. L., Huang, S. Y., Behnke, M. S., Chen, K., Shen, B., and Zhu, X. Q. (2016a). The past, present, and future of genetic manipulation in Toxoplasma gondii. Trends Parasitol. 32, 542–553. doi: 10.1016/j.pt.2016.04.013
Wang, J. L., Huang, S. Y., Li, T. T., Chen, K., Ning, H. R., and Zhu, X. Q. (2016b). Evaluation of the basic functions of six calcium-dependent protein kinases in Toxoplasma gondii using CRISPR-Cas9 system. Parasitol. Res. 115, 697–702. doi: 10.1007/s00436-015-4791-4796
Keywords: Toxoplasma gondii, CRISPR-Cas9 system, rhoptry proteins (ROPs), virulence factors, gene knockout, host–pathogen interaction
Citation: Wang J-L, Li T-T, Elsheikha HM, Chen K, Zhu W-N, Yue D-M, Zhu X-Q and Huang S-Y (2017) Functional Characterization of Rhoptry Kinome in the Virulent Toxoplasma gondii RH Strain. Front. Microbiol. 8:84. doi: 10.3389/fmicb.2017.00084
Received: 11 September 2016; Accepted: 12 January 2017;
Published: 24 January 2017.
Edited by:
Guan Zhu, Texas A&M University, USAReviewed by:
Xun Suo, China Agricultural University, ChinaHaider Abdul-Lateef Mousa, University of Basrah, Iraq
Yutaka Handa, National Center for Global Health and Medicine, Japan
Copyright © 2017 Wang, Li, Elsheikha, Chen, Zhu, Yue, Zhu and Huang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xing-Quan Zhu, eGluZ3F1YW56aHUxQGhvdG1haWwuY29t Si-Yang Huang, c2l5YW5nLmh1YW5nQGhvdG1haWwuY29t
 Jin-Lei Wang
Jin-Lei Wang Ting-Ting Li1
Ting-Ting Li1 Hany M. Elsheikha
Hany M. Elsheikha Xing-Quan Zhu
Xing-Quan Zhu