- 1Department of Clinical Microbiology, Ruijin Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China
- 2Department of Clinical Laboratory, Ruijin Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China
- 3Department of Clinical Laboratory, Xiangya Hospital, Central South University, Changsha, China
- 4Department of Medical Microbiology and Parasitology, Institutes of Medical Sciences, Shanghai Jiao Tong University School of Medicine, Shanghai, China
- 5Department of Pulmonary Medicine, Ruijin Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China
Klebsiella pneumoniae (K.pneumoniae) is a common nosocomial pathogen causing bloodstream infections. Antibiotic susceptibility surveillance and molecular characterization will facilitate prevention and management of K. pneumoniae bloodstream infections. K. pneumoniae isolates causing bloodstream infections were consecutively collected between January 2012 and December 2015 in Shanghai. Eighty isolates (20 per year) were randomly selected and enrolled in this study. Drug susceptibility were determined by the disk diffusion method. Polymerase chain reaction (PCR) was employed to detect extended-spectrum β-lactamases (ESBLs), carbapenemases, and seven housekeeping genes of K. pneumoniae. eBURST was used for multi-locus sequence typing (MLST). More than 50% isolates were resistant to cefuroxime, ampicillin-sulbactam, and piperacillin, while carbapenems had lower resistant rates than other antibiotics. Of the 80 isolates, 22 produced ESBLs, and 14 were carbapenemase producers. In the ESBL-producing K. pneumoniae isolates, the most common ESBL genes were blaSHV and blaCTX−M. Thirteen carbapenemase producers harbored blaKPC−2 and one other carried blaNDM−5. ST11 (14/80) was the most frequent sequence type (ST), followed by ST15 (7/80) and ST29 (4/80). Our data revealed high prevalence of antibiotic resistant K. pneumoniae isolates from bloodstream infections but their genetic diversity suggested no clonal dissemination in the region. Also, one K. pneumoniae isolate harbored blaNDM−5 in this study, which was firstly reported in Shanghai.
Introduction
Klebsiella pneumoniae (K. pneumoniae) can cause ventilator-associated pneumonia, urinary tract infection, sepsis, catheter-related infection, and bacterial meningitis (Tumbarello et al., 2006; Du et al., 2014), and is also the most common causative gram-negative bacterium in nosocomial and community-acquired infections (Hu et al., 2015). Unfortunately, extended and overuse of antibiotics may potentiate antibiotic resistance of K. pneumoniae strains to cephalosporins, aminoglycosides, fluoroquinolones, and even carbapenems (Guana et al., 2014), which will be difficult and costly to control.
Extensive use of invasive procedures and glucocorticoids has increased the incidence of bloodstream infections and caused high mortality in patients (27.2–40.8%; Marra et al., 2006; Yang et al., 2010; Lv et al., 2014; Wang et al., 2016). Of the gram negative bacilli implicated in nosocomial bloodstream infections, K. pneumoniae was one of the most common pathogens, second only to Escherichia coli (Ghadiri et al., 2012; Lv et al., 2014). The prevalence of Klebsiella spp. bloodstream infections was 7.6% in the United States, Canada, South America, and Europe according to SENTRY (Biedenbach et al., 2004). On the basis of a large-scale survey from China, K. pneumoniae caused 11.3% of all bloodstream infections in 2011–2012 (Lv et al., 2014).
The wide dissemination of drug resistant pathogens leads to their increasing prevalence in bloodstream infections. Data from China showed that 53.3% K. pneumoniae isolates were multidrug-resistant (Lv et al., 2014). In Europe, Latin America, and North America, 21.7, 42.7, and 5.8% of Klebsiella spp. had the extended-spectrum β-lactamase (ESBL) phenotype (Biedenbach et al., 2004), while as high as 60.6% K. pneumoniae isolates in Greece were carbapenemase producers (Daikos et al., 2014). And the most prevalent group of carbapenemases was K. pneumonia carbapenemase (KPC). Studies have shown that the drug-resistant organisms bloodstream infection was one important risk factor for mortality and negatively impacted the treatment outcome of patients (Kim et al., 2002; Tumbarello et al., 2006). Limited data on susceptibility and molecular epidemiology of K. pneumoniae causing bloodstream infections were available in Shanghai. In the study, we have monitored resistance phenotype, the prevalent resistant genes and sequence types (STs) of K. pneumoniae isolates from bloodstream infections between 2012 and 2015 in the region.
Materials and Methods
Patients and Bacterial Isolates
This retrospective study was conducted in Ruijin Hospital, an 1800-bed general university-affiliated hospital located in Shanghai, with ~1,15,000 patient visits per year. Patients with at least one positive blood culture of K. pneumoniae from January 2012 through December 2015 were enrolled in the study. A total of 254 episodes of K. pneumoniae bloodstream infections (66 in 2012, 62 in 2013, 64 in 2014, and 62 in 2015) were identified during this period. Only the first positive blood culture was reviewed and recorded. Eighty isolates were enrolled: twenty isolates were selected from each year using the random number generation function in Microsoft Office Excel 2010 (Microsoft Corporation, Redmond, WA, USA). Isolates identification was carried out using matrix-assisted laser desorption ionization-time of flight mass spectrometer (bioMérieux, Marcy-l'Étoile, France).
This study was approved by Ruijin Hospital Ethics Committee (Shanghai Jiao Tong University School of Medicine). The Review Board exempted request for informed consent because this retrospective study only focused on the bacteria and did not have impact on the patients.
Antimicrobial Susceptibility Tests
The antimicrobial susceptibility tests were determined by the disk diffusion method according to the Clinical and Laboratory Standards Institute [Clinical and Laboratory Standards Institute (CLSI), 2015]. The antibiotics tested were ceftazidime, ceftriaxone, cefepime, cefotaxime, cefuroxime, amikacin, gentamicin, tobramycin, piperacillin, aztreonam, ciprofloxacin, levofloxacin, ampicillin-sulbactam, piperacillin-tazobactam, imipenem, meropenem, trimethoprim-sulfamethoxazole, and cefoperazone-sulbactam. Pseudomonas aeruginosa ATCC 27853, K. pneumoniae ATCC 700603, and E. coli ATCC 25922 were used for quality control.
ESBL-Producing and Carbapenemase-Producing Isolates Screening and Confirmation
According to the CLSI criteria [Clinical and Laboratory Standards Institute (CLSI), 2015], ceftazidime and cefotaxime were used as screening tests for ESBLs. Ceftazidime, ceftazidime-clavulanate, cefotaxime, and cefotaxime-clavulanate were used as confirmatory test. Imipenem and meropenem were used as screening for carbapenemase production. Detection of carbapenemase genes was performed to confirm production of carbapenemases.
Detection of Resistant Genes
Polymerase chain reaction (PCR) was performed to amplify the resistant genes using previous primers (Du et al., 2014; Zhao et al., 2016), including ESBL genes (blaTEM, blaSHV, blaCTX−M−1, −2, −8, −9, −10, −25 group), blaVEB, blaGES, blaOXA(−1, −2, −10 group), and blaPER) and carbapenemase genes (blaVIM, blaIPM, blaKPC, blaGIM, blaSPM, blaSIM, blaOXA−48 group, and blaNDM). Sample DNA was prepared by boiling the bacteria at 100°C for 15 min, and centrifugation at 3,000 g for 15 min. The PCR conditions used were initial denaturation at 95°C for 5 min, cyclic denaturation at 95°C for 50 s, annealing at 55°C for 30 s, elongation at 72°C for 1 min for 35 cylces and final extension at 72°C for 5 min. PCR products were examined by electrophoresis in 1.5% agarose gel. Positive amplicons were sequenced using ABI3730xl DNAAnalyzer by Sangon Biotech (Shanghai, China) and aligned with subtypes of β-lactamase genes by BLAST (http://blast.ncbi.nlm.nih.gov).
Multilocus Sequence Typing
Multilocus sequence typing (MLST) was carried out as described previously (http://bigsdb.web.pasteur.fr/klebsiella/primers_used.html; Diancourt et al., 2005). Briefly, Seven hosekeeping genes (gapA, infB, mdh, pgi, phoE, rpoB, and tonB) for K. pneumoniae were amplified, sequenced, and analyzed. Alleles and STs were determined according to the database (http://bigsdb.web.pasteur.fr/perl/bigsdb/bigsdb.pl?db=pubmlst_klebsiella_seqdef_public&page=profiles). STs that could not be found in the database were submitted to the curator of the database (klebsiellaMLST@pasteur.fr). eBURST version 3.0 software was used to analyze the clustering of related STs. In this study, isolates were grouped together if six of the seven alleles were homologs.
Statistical Analysis
Data in this study were analyzed by SAS 8.2 (SAS Institute Inc., Cary, NC, USA). Continuous variables were presented as the mean ± SD or median and interquartile range. For categorical variables, the chi-square test was used to compare the disparity between different groups. P < 0.05 was considered to be statistically significant.
Results
Patient Data
From January 2012 to December 2015, there were more male patients (182/254) than females (72/254). The age of patients ranged from 1 to 91 years. Most of the cases (40/254) were derived from the surgery. Fifty-eight males and 22 females were enrolled and their median age was 61 years (range: 12–91 years). Most patients were from the Department of General Surgery (17/80), Transplantation (11/80), and Infectious Disease (9/80).
Antimicrobial Susceptibility Tests
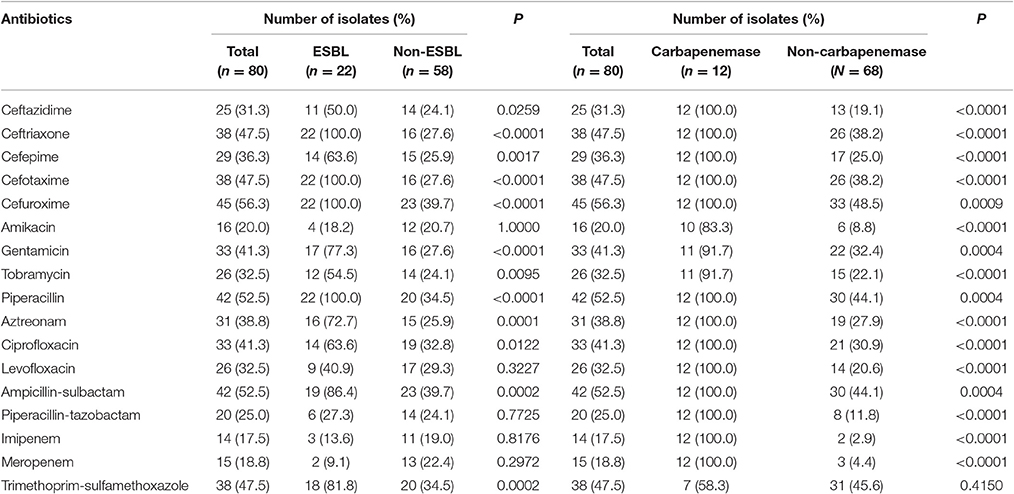
Isolates bear high resistance to cefuroxime (56.3%), ampicillin-sulbactam (52.5%), and piperacillin (52.5%), while there were the least resistance to imipenem (17.5%), meropenem (18.8%), and amikacin (20.0%). Of the 80 isolates, 22 isolates (27.5%) were ESBL producers, and 14 (17.5%) were carbapenemase producers. The resistant rates to most of the antibiotics in ESBL producers were >60%, including cefuroxime (100.0%), piperacillin (100.0%), cefotaxime (100.0%), ceftriaxone (100.0%), ampicillin-sulbactam (86.4%), trimethoprim-sulfamethoxazole (81.8%), gentamicin (77.3%), aztreonam (72.7%), ciprofloxacin (63.6%), and cefepime (63.6%). The resistant rates to antibiotics were significantly higher in ESBL producers than those in non-producers except meropenem, levofloxacin, piperacillin-tazobactam, imipenem, and amikacin. The carbapenemase producers were resistant to all tested drugs except trimethoprim-sulfamethoxazole, tobramycin, amikacin, and gentamicin. Resistant rates to all antibiotics other than trimethoprim-sulfamethoxazole were statistically different between the carbapenemase producers and non-carbapenemase producers (Table 1).
Characterization of Resistance Genes
In this study, 40.9% (9/22) blaTEM, 100.0% (22/22) blaSHV, and 77.3% (17/22) blaCTX−M were identified in the 22 ESBL producers. The dominan ESBL gene was blaCTX−M. The subtypes of blaTEM were blaTEM−1(n = 8) and blaTEM−103(n = 1). The main subtype of blaSHV was blaSHV−1 (n = 11), and the most common subtype of blaCTX−M was blaCTX−M−14 (n = 7). Other subtypes including blaSHV−33, blaSHV−36, blaSHV−11, blaSHV−12, blaSHV−28, blaCTX−M−15, and blaCTX−M−65 were also found. Five of the 22 isolates (22.7%) only harbored blaSHV. While the other 17 isolates additionally harbored one or two ESBL genes, including blaSHV along with blaCTX−M(8/22), and blaSHV along with blaCTX−M and blaTEM (9/22) (Table 2). No blaCTX−M(−2, −8, −10, −25 group), blaVEB, blaGES, blaOXA(−1, −2, −10 group), or blaPERgenes were found.
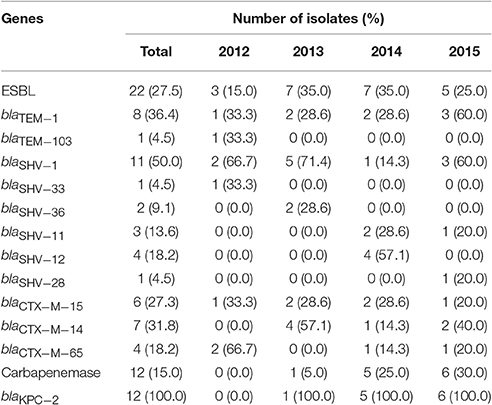
Table 2. Resistant genes in ESBL-producing or carbapenemase-producing isolates from K. pneumoniae bloodstream isolates during 2012–2015.
The blaKPC−2 gene was detected in 92.9% (13/14) carbapenemase producers and blaNDM−5was found in another isolate. No other carbapenemase genes were detected.
ST and Clonal Complexes
Forty-seven STs among 80 K. pneumoniae isolates, including three new STs (ST2247, ST2248, and ST2249) were identified. The most prevalent ST in K. pneumoniae isolates was ST11 (n = 14, 17.5%), followed by ST15 (n = 7, 8.7%), and ST29 (n = 4, 5.0%) (Table 3). eBURST indicated that these 47 STs could be clustered into one clonal complex, four groups, and 35 singletons (Figure 1). Notably, ESBL producers were mostly ST15 (5/22, 22.7%), ST11 (3/22, 13.6%), and ST628 (3/22, 13.6%), while all carbapenemase producers belonged to ST11 except the blaNDM−5-positive one (ST1).

Table 3. Antibiotic resistance profiles and genotypes in MLST of eighty K. pneumoniae isolates from bloodstream infections.

Figure 1. Clonal groupings among K. pneumoniae bloodstream isolates. There were 35 singletons, four groups (group 1: ST37, ST1779; group 2: ST11, ST340; group 3: ST347, ST1023; group 4: ST485, ST45) and one clonal complexes (ST218, ST23, ST2248) with ST23 as the putative founder. One blob represented one ST and the size reflected the number of isolates belonging to it.
Discussion
Local epidemiologic data on prevalence of specific clones of K. pneumoniae bloodstream infection were indispensible to develop clinical treatment regimen and evaluate outcomes of different therapeutic strategy (Pai et al., 2004; Marra et al., 2006; Neuner et al., 2011; Harris et al., 2015). However, there were limited data on antibiotic resistance, resistant genes and STs of K. pneumoniae bloodstream isolates in Shanghai. In our pilot study conducted in 2012, we found that K. pneumoniae was the second frequent gram negative bacillus from blood cultures in our hospital, representing 14.3% of all the isolates (Zhao et al., 2014). In current study, we have extended our research through 2015 to acquire more comprehensive molecular epidemiologic data.
K. pneumoniae bloodstream isolates in the study showed threatened resistance to the most of routine antibiotics and only meropenem, imipenem, amikacin, and piperacillin-tazobactam had relative low resistant rates (<30%) which made them better candidates for empiric therapy of K. pneumoniae bloodstream infections. However, prudent and rational uses of antibiotics based on the results of antimicrobial susceptibility tests are still essential to the success of treatment. Notably, the resistant rates to meropenem, imipenem, amikacin, cefepime, and ciprofloxacin determined in this study were much higher than what were found in the surveillance of 2011–2012 in China (Li et al., 2014). This dynamic change of the pattern of drug resistance warrant active ongoing surveillance on antibiotic resistance and consistent prevention and control of K. pneumoniae.
The proportion of ESBL-producing K. pneumoniae (27.5%) in this study was lower than that in Italy (32.6%), America (51.8%), Korea (52.9%), and Russia (60.8%) (Kim et al., 2002; Edelstein et al., 2003; Marra et al., 2006; Tumbarello et al., 2006), but the multidrug-resistant phenotype, and the dominate ESBL enzyme (CTX-M) of ESBL producers were similar (Edelstein et al., 2003). Typing of ESBL producers revealed a high level of genetic diversity, with ST15 (22.7%, 5/22) and ST11 (13.6%, 3/22) as the most common STs. ST15 K. pneumoniae has been identified in both animals and humans in several countries. Although it was not yet a dominant clone in our study, ST15 K. pneumoniae had achieved a highly successful clonal spread in some countries, such as Bulgaria, Portugal, and Thailand (Netikul et al., 2014; Rodrigues et al., 2014; Markovska et al., 2015), and we should be cautious about its high potential of becoming a major clone associated with ESBL producers in Shanghai in future. Isolates belonged to ST11 were isolated from different departments, in different years with different antibiotic resistance profiles. This suggested no clonal dissemination in the region.
Different from previous studies where no carbapenemase-producing E. coli bloodstream isolate was found (Zhao et al., 2015; Wang et al., 2016), the carbapenemase-producing K. pneumoniae accounted for 17.5% in our hospital, and they, as reported elsewhere (David et al., 2013; Du et al., 2014), also showed extremely high resistant rates to major antibiotics except aminoglycosides and trimethoprim-sulfamethoxazole. Molecular analysis suggested 92.9% of the carbapenemase producers were harbored with blaKPC−2, and belonged to ST11, confirming that ST11 was associated with KPC. Unlike the widespread of KPC-producing ST258 K. pneumoniae in Europe, the dominant ST of KPC-producing K. pneumoniae was ST11 in China (Dautzenberg et al., 2016). Also, One ST1 carbapenemase producers carried blaNDM−5 was found in this study. NDM-5, which mostly found in E. coli, was firstly reported in K. pneumoniae in Shanghai. Taken together, our data indicated the KPC-producing ST11 K. pneumoniae isolate was a high-risk clone in our hospital, and should be taken as the major consideration when developing strategy to control resistant isolates dissemination. NDM, along with KPC, VIM, and OXA-48, were four major carbapenemases detected in K. pneumoniae. The most common type of NDM found was NDM-1. NDM-5, an emerging carbapenemase in K. pneumoniae, should attract our attention.
Our study described the phenotypic and molecular properties of K. pneumoniae bloodstream isolates in Shanghai for the first time. This study also suggested ST11 K. pneumoniae harbored blaKPC−2 had absolute predominance in carbapenemase producers, and NDM-5 was an emerging carbapenemase. Although our conclusion based on a single hospital cannot be directly extrapolated to the whole area, it provides the stepstone for the future expanded research associated with multicenter and further resistant mechanism surveillance to prevent further possible dissemination in this region.
Author Contributions
Conceived and designed the experiments: LH, SX, SW, and SZ. Performed the experiments: SW and SX. Analyzed the data: SX and SW. Contributed reagents/materials/analysis tools: LH, YN, and XG. Wrote the paper: SX, SW, LH, WW, FG, and JQ.
Funding
This work was supported by Special Fund for Health-scientific Research in the Public Interest: Research & application for the prevention & control of nosocomial infections caused by multi-drug resistant bacteria (201002021) and The Shanghai 3-Year Plan of the Key Subjects Construction in Public Health-Infectious Diseases and Pathogenic Microorganism (15GWZK0102). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Biedenbach, D. J., Moet, G. J., and Jones, R. N. (2004). Occurrence and antimicrobial resistance pattern comparisons among bloodstream infection isolates from the SENTRY Antimicrobial Surveillance Program (1997–2002). Diagn. Microbiol. Infect. Dis. 50, 59–69. doi: 10.1016/j.diagmicrobio.2004.05.003
Clinical Laboratory Standards Institute (CLSI) (2015). Performance Standards for Antimicrobial Susceptibility Testing[S]: Twenty-fourth Informational Supplement. Wayne, PA: Clinical and Laboratory Standards Institute.
Daikos, G. L., Tsaousi, S., Tzouvelekis, L. S., Anyfantis, I., Psichogiou, M., Argyropoulou, A., et al. (2014). Carbapenemase-producing Klebsiella pneumoniae bloodstream infections: lowering mortality by antibiotic combination schemes and the role of Carbapenems. Antimicrob. Agents Chemother. 58, 2322–2328. doi: 10.1128/AAC.02166-13
Dautzenberg, M. J., Haverkate, M. R., Bonten, M. J., and Bootsma, M. C. (2016). Epidemic potential of Escherichia coli ST131 and Klebsiella pneumoniae ST258: a systematic review and meta-analysis. BMJ Open 6:e009971. doi: 10.1136/bmjopen-2015-009971
David, V. D., Keith, S. K., Elizabeth, A. N., and Robert, A. B. (2013). Carbapenem-resistant Enterobacteriaceae: a review of treatment and outcomes. Diagn. Microbiol. Infect. Dis. 75, 6. doi: 10.1016/j.diagmicrobio.2012.11.009
Diancourt, L., Passet, V., Verhoef, J., Grimont, P. A., and Brisse, S. (2005). Multilocus sequence typing of Klebsiella pneumoniae nosocomial isolates. J. Clin. Microbiol. 43, 4178–4182. doi: 10.1128/JCM.43.8.4178-4182.2005
Du, J., Li, P., Liu, H., Lü, D., Liang, H., and Dou, Y. (2014). Phenotypic and molecular characterization of multidrug resistant Klebsiella pneumoniae isolated from a University Teaching Hospital, China. PLoS ONE 9:e95181. doi: 10.1371/journal.pone.0095181
Edelstein, M., Pimkin, M., Palagin, I., Edelstein, I., and Stratchounski, L. (2003). Prevalence and molecular epidemiology of CTX-M extended-spectrum -lactamase-producing Escherichia coli and Klebsiella pneumoniae in Russian Hospitals. Antimicrob. Agents Chemother. 47, 3724–3732. doi: 10.1128/AAC.47.12.3724-3732.2003
Ghadiri, H., Vaez, H., Khosravi, S., and Soleymani, E. (2012). The antibiotic resistance profiles of bacterial strains isolated from patients with hospital-acquired bloodstream and urinary tract infections. Crit. Care Res. Pract. 2012:890797. doi: 10.1155/2012/890797
Guana, J., Zhuo, C., and DH, S. (2014). CHINET 2012 surveillance of antibiotic resistance in Klebsiella spp in China. Chin. J. Infect. Chemother. 14, 398–404. doi: 10.3969/j.issn.1009-7708.2014.05.006
Harris, P. N. A., Yin, M., Jureen, R., Chew, J., Ali, J., Paynter, S., et al. (2015). Comparable outcomes for β-lactam/β-lactamase inhibitor combinations and carbapenems in definitive treatment of bloodstream infections caused by cefotaxime-resistant Escherichia coli or Klebsiella pneumoniae. Antimicrob. Resist. Infect. Control 4:14. doi: 10.1186/s13756-015-0055-6
Hu, F., Zhu, D., and Wang, F. (2015). CHINET 2014 surveillance of bacterial resistance in China. Chin. J. Infect. Chemother. 15, 401–410. doi: 10.3969/j.issn.1009-7708.2015.05.001
Kim, Y. K., Pai, H., Lee, H. J., Park, S. E., Choi, E. H., Kim, J., et al. (2002). Bloodstream infections by extended-spectrum β-lactamase-producing Escherichia coli and Klebsiella pneumoniae in children: epidemiology and clinical outcome. Antimicrob. Agents Chemother. 46, 1481–1491. doi: 10.1128/AAC.46.5.1481-1491.2002
Li, Y., Lv, Y., and Xue, F. (2014). Antimicrobial susceptibility surveillance of gram–negative bacterial from Mohnarin 2011–2012. Chin. J. Clin. Pharmacol. 30, 260–277.
Lv, Y., Li, Y., and Xue, F. (2014). Mohnarin report of 2011–2012: surveillance for resistance of bacteria causing bloodstream infections. Chin. J. Clin. Pharmacol. 30, 278–288.
Markovska, R., Stoeva, T., Schneider, I., Boyanova, L., Popova, V., Dacheva, D., et al. (2015). Clonal dissemination of multilocus sequence type ST15 KPC-2-producing Klebsiella pneumoniae in Bulgaria. APMIS 123, 887–894. doi: 10.1111/apm.12433
Marra, A. R., Wey, S. B., Castelo, A., Gales, A. C., Cal, R. G. R., Filho, J. C., et al. (2006). Nosocomial bloodstream infections caused by Klebsiella pneumoniae: impact of extended-spectrum β-lactamase (ESBL) production on clinical outcome in a hospital with high ESBL prevalence. BMC Infect. Dis. 6:24. doi: 10.1186/1471-2334-6-24
Netikul, T., Sidjabat, H. E., Paterson, D. L., Kamolvit, W., Tantisiriwat, W., Steen, J. A., et al. (2014). Characterization of an IncN2-type blaNDM-(1)-carrying plasmid in Escherichia coli ST131 and Klebsiella pneumoniae ST11 and ST15 isolates in Thailand. J. Antimicrob. Chemother. 69, 3161–3163. doi: 10.1093/jac/dku275
Neuner, E. A., Yeh, J. Y., Hall, G. S., Sekeres, J., Endimiani, A., Bonomo, R. A., et al. (2011). Treatment and outcomes in Carbapenem-resistant Klebsiella pneumoniae bloodstream infections. Diagn. Microbiol. Infect. Dis. 69, 357–362. doi: 10.1016/j.diagmicrobio.2010.10.013
Pai, H., Kang, C. I., Byeon, J. H., Lee, K. D., Park, W. B., Kim, H. B., et al. (2004). Epidemiology and clinical features of bloodstream infections caused by AmpC-Type-β-lactamase-producing Klebsiella pneumoniae. Antimicrob. Agents Chemother. 48, 3720–3728. doi: 10.1128/AAC.48.10.3720-3728.2004
Rodrigues, C., Machado, E., Ramos, H., Peixe, L., and Novais, A. (2014). Expansion of ESBL-producing Klebsiella pneumoniae in hospitalized patients: a successful story of international clones (ST15, ST147, ST336) and epidemic plasmids (IncR, IncFIIK). Int. J. Med. Microbiol. 304, 1100–1108. doi: 10.1016/j.ijmm.2014.08.003
Tumbarello, M., Spanu, T., Sanguinetti, M., Citton, R., Montuori, E., Leone, F., et al. (2006). Bloodstream infections caused by Extended-spectrum-β-lactamase-producing Klebsiella pneumoniae: risk factors, molecular epidemiology, and clinical outcome. Antimicrob. Agents Chemother. 50, 498–504. doi: 10.1128/AAC.50.2.498-504.2006
Wang, S., Zhao, S. Y., Xiao, S. Z., Gu, F. F., Liu, Q. Z., Tang, J., et al. (2016). Antimicrobial resistance and molecular epidemiology of Escherichia coli causing bloodstream infections in three hospitals in Shanghai, China. PLoS ONE 11:e0147740. doi: 10.1371/journal.pone.0147740
Yang, Z., Zhan, S., and Wang, B. (2010). Fatality anf secular trend of bloodstream infections during hospitalization in China: a systematie review and meta-analysis. J. Peking Univ. (Health Sciences) 42, 304–307.
Zhao, S. Y., Wang, Y. C., Xiao, S. Z., Jiang, X. F., Guo, X. K., Ni, Y. X., et al. (2015). Drug susceptibility and molecular epidemiology of Escherichia coli in bloodstream infections in Shanghai, China, 2011–2013. Infect. Dis. 47, 310–318. doi: 10.3109/00365548.2014.990509
Zhao, S. Y., Xiao, S. Z., Han, L. Z., Mi, C. R., and Ni, Y. X. (2014). Distribution and antimicrobial resistance of pathogens isolated from hospitalized patients with bloodstream infections Chin. J. Infect. Control 13, 266–270. doi: 10.3969/j.issn.1671-9638.2014.05.003
Zhao, S. Y., Zhang, J., Zhang, Y. L., Wang, Y. C., Xiao, S. Z., Gu, F. F., et al. (2016). Epidemiology and risk factors for faecal extended-spectrum β-lactamase-producing Enterobacteriaceae (ESBL-E) carriage derived from residents of seven nursing homes in western Shanghai, China. Epidemiol. Infect. 144, 695–702. doi: 10.1017/s0950268815001879
Keywords: Klebsiella pneumoniae, bloodstream infections, resistance phenotype, molecular epidemiology, extended-spectrum β-lactamase
Citation: Xiao S-z, Wang S, Wu W-m, Zhao S-y, Gu F-f, Ni Y-x, Guo X-k, Qu J-m and Han L-z (2017) The Resistance Phenotype and Molecular Epidemiology of Klebsiella pneumoniae in Bloodstream Infections in Shanghai, China, 2012–2015. Front. Microbiol. 8:250. doi: 10.3389/fmicb.2017.00250
Received: 27 August 2016; Accepted: 06 February 2017;
Published: 23 February 2017.
Edited by:
Dongsheng Zhou, Beijing Institute of Microbiology and Epidemiology, ChinaReviewed by:
Marquita Vernescia Gittens-St.Hilaire, The University of the West Indies at Cave Hill, BarbadosHaijian Zhou, Chinese Center for Disease Control and Prevention, China
Copyright © 2017 Xiao, Wang, Wu, Zhao, Gu, Ni, Guo, Qu and Han. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jie-ming Qu, am1xdTA5MDZAMTYzLmNvbQ==
Li-zhong Han, aGFubGl6aG9uZzExMDdAMTYzLmNvbQ==
†These authors have contributed equally to this work and should be considered as co-first author.
 Shu-zhen Xiao1
Shu-zhen Xiao1 Li-zhong Han
Li-zhong Han