- 1Department of Basic Medical Sciences, Biochemistry Section, University of the West Indies, Mona, Jamaica
- 2Veterinary Services Division, Ministry of Agriculture, Hope Gardens, Jamaica
Leptospirosis is a zooanthroponosis aetiologically caused by pathogenic bacteria belonging to the genus, Leptospira. Environmental signals such as increases in temperatures or oxidative stress can trigger response regulatory modes of virulence genes during infection. This study sought to determine the effect of temperature and oxidative stress on virulence associated genes in highly passaged Leptospira borgpeterseneii Jules and L. interrogans Portlandvere. Bacteria were grown in EMJH at 30°C, 37°C, or at 30°C before being transferred to 37°C. A total of 14 virulence-associated genes (fliY, invA, lenA, ligB, lipL32, lipL36, lipL41, lipL45, loa22, lsa21, mce, ompL1, sph2, and tlyC) were assessed using endpoint PCR. Transcriptional analyses of lenA, lipL32, lipL41, loa22, sph2 were assessed by quantitative real-time RT-PCR at the temperature conditions. To assess oxidative stress, bacteria were exposed to H2O2 for 30 and 60 min with or without the temperature stress. All genes except ligB (for Portlandvere) and ligB and mce (for Jules) were detectable in the strains. Quantitatively, temperature stress resulted in significant changes in gene expression within species or between species. Temperature changes were more influential in gene expression for Jules, particularly at 30°C and upshift conditions; at 37°C, expression levels were higher for Portlandvere. However, compared to Jules, where temperature was influential in two of five genes, temperature was an essential element in four of five genes in Portlandvere exposed to oxidative stress. At both low and high oxidative stress levels, the interplay between genetic predisposition (larger genome size) and temperature was biased towards Portlandvere particularly at 30°C and upshift conditions. While it is clear that expression of many virulence genes in highly passaged strains of Leptospira are attenuated or lost, genetic predisposition, changes in growth temperature and/or oxidative intensity and/or duration were factors which acted in isolation or together with other regulatory cues to contribute to the variable gene expression observed in this study. Overall, differential gene expression in serovar Portlandvere was more responsive to temperature and oxidative stress.
Introduction
Leptospirosis is a zooanthroponosis, widely distributed throughout the world and aetiologically caused by pathogenic bacteria belonging to the genus, Leptospira (Bharti et al., 2003; Pappas et al., 2008). Pathogenic Leptospira species are invasive and infection results from their ability to colonize and invade the renal tubes of incidental hosts. While the complete mechanism involved in leptospiral pathogenicity is not fully elucidated, several studies on leptospiral virulence and virulence-associated factors indicate the involvement of haemolysins, adhesins, heat shock proteins, flagellins/motility, lipopolysaccharide (LPS), catalase KatE, heat-inducible ClpB chaperone, and several outer membrane proteins (Artiushin et al., 2004; Cullen et al., 2005; Barbosa et al., 2006; Nally et al., 2007; Dong et al., 2008; Lourdault et al., 2011; Andrade and Brown, 2012; Eshghi et al., 2012).
Pathogenicity is multifactorial, requiring integrated mechanisms and pathways to establish an infection. As diverse pathogenic bacteria share common strategies to cause disease and infection, the extent of damage to host tissue is determined by multiple gene or gene products involved in pathways of signal transduction, invasiveness and toxigenesis. Bacteria rely on the ability to sense and respond to environmental cues, including changes in temperature, pH, osmolarity, oxygen availability, and nutrient conditions (Thomas and Wigneshweraraj, 2014). The changing environment, often influenced by climate change, prompts adaptation, and regulatory responses to enhance the survival of the pathogen, and by extension, the ability to cause infection is largely due to the pathogen’s ability and adaptability (Reis et al., 2008; Lau et al., 2010; Batchelor et al., 2012).
The state of physiological imbalance between the natural or exogenous production of and/or exposure to high levels of oxidants and the organism’s ability to counteract their harmful effects induce oxidative stress (Fulda et al., 2010). In biological systems, oxidation reactions involving organic molecules usually generate unstable free radicals including reactive oxygen species (ROS) and reactive nitrogen species (RNS) (Nathan and Shiloh, 2000; Seis, 2014). Unequivocal interplay between oxidant and anti-oxidant countermeasures trigger a myriad of cascades that are likely to induce oxidative stress. For example, in Escherichia coli, the presence of as little as 0.5 μM H2O2, regardless of the source, can be cytotoxic (Sobota and Imlay, 2011). Unlike enteric bacteria, saprophytic leptospires lack the two main transcriptional regulators of oxidative stress response in enteric bacteria, OxyR and SoxRS (Anjem and Imlay, 2012). However, Leptospira interrogans possesses four predicted FurR homologs and PerR, a negative peroxide regulator with sensitivity to low H2O2 levels (Fillat, 2014), which exhibits similarity to PerR in Bacillus subtilis which controls katA and ahpC expression (Lo et al., 2010). Other leptospiral defense mechanisms against oxidative stress involve but are not limited to peroxiredoxin LinAhpC, catalase KatE in L. interrogans compared to KatG in L. biflexa, heat inducible ClpB chaperone, and glutathione and thiol peroxidases, among others which function in the capacity as metalloproteins (Lourdault et al., 2011; Eshghi et al., 2012).
Adaptation by an organism during serial passage is well established since the generation of live attenuated vaccines (Gamberini et al., 2005). In most instances, gene expression among pathogenic bacteria, including Leptospira is attenuated in highly passaged cultures and associated with loss of or attenuated virulence. Cullen et al. (2002) noted concomitant changes in leptospiral surfaceome, colonial morphology and loss of virulence associated with highly passaged Leptospira. Other studies have reported observations of non-synonymous variant alleles (Lehmann et al., 2015), attenuation of genes involved in invasion (Toma et al., 2014) and plasminogen binding (Vieria et al., 2009). Gene function and virulence may be restored by passage through a host and/ or activation by stimuli.
Temperature and oxidative stress represent two of the main external (host and environmental) stresses which influence virulence and viability of pathogenic Leptospira. Given that L. interrogans serovar Portlandvere and L. borgpetersenii serovar Jules account for more than 60% of cases of human leptospirosis in Jamaica, and with the paucity of information regarding molecular pathogenicity and the drivers of virulence in these species, this study sought to determine the effect of temperature and oxidative stress on virulence associated genes in L. borgpeterseneii Jules and L. interrogans Portlandvere.
Materials and Methods
Bacterial Strains and Culture Conditions
Leptospira interrogans serovar Portlandvere strain MY1039 and L. borgpetersenii serovar Jules strain jules were sub-cultured biweekly into liquid Ellinghausen-McCullough-Johnson-Harris (EMJH) medium at 30°C (Cameron, 2015), to yield highly passaged cultures with >200 serial passages. Leptospires were visualized using an Olympus BX 53 darkfield microscope.
Temperature and Oxidative Stress Conditions
Bacteria were grown in supplemented EMJH liquid medium to a density of 1 × 108 bacteria per mL and subsequently pelleted via centrifugation at 3,200 × g for 15 min. The pelleted bacteria were washed three times with EMJH medium and the bacteria were then re-suspended in EMJH medium and visualized via darkfield microscopy. One milliliter of cultures (3 × 108 cells) was used to seed each 50 mL aliquot of fresh growth medium which was incubated either at 30°C for 14 days, 37°C for 14 days, or 30°C for 7 days before transfer to 37°C for an additional 7 days to simulate upshifted temperature conditions. Biological replicates (rather than technical replicates) were used at each temperature condition. These sets of conditions established the baseline for further comparisons. Following incubation, bacteria were visualized via darkfield microscopy to ensure bacterial viability prior to exogenous oxidative stress. Hydrogen peroxide (H2O2) was added to inoculated media at a final concentration of 1 mM or 10 mM and samples were incubated for 30 or 60 min at the previously incubated temperatures (Eshghi et al., 2012). Four exposure conditions ensued: 1 mM H2O2 for 30 min; 10 mM H2O2 for 30 min; 1 mM H2O2 for 60 min; and 10 mM H2O2 for 60 min. Controls without H2O2 were also analyzed. Following exposures, cells were collected by centrifugation for subsequent analyses.
DNA Isolation and Endpoint PCR Analysis of Virulence Associated Genes
Genomic DNA was isolated from resuspended pellets by using the DNeasy blood and tissue kit (Qiagen, CA, USA) as per the manufacturer’s instructions. DNA quality was assessed by electrophoresis and quantification done using the Qubit 3.0 fluorometer (Invitrogen, USA) and Qubit dsDNA BR Assay Kit (Invitrogen, USA). Each biological replicate was done in duplicate. Endpoint PCR was performed to confirm the presence of open reading frames for the 14 virulence-associated genes investigated, which included genes for outermembrane proteins (lipL36, lipL41, lipL45), genes involved in adherence (lenA, ligB, lipL32, loa22, lsa21, ompL1), invasion (invA, mce), haemolysis (sph2, tlyC), and motility/chemotaxis (fliY). Two nanogrammes of DNA were used for amplification in a total reaction volume of 25 μL containing final concentration of 25 mM MgCl2, 500 μM of each deoxynucleotide triphosphate (dNTP), 5 U Taq polymerase, and 5 μM of each primer listed in Table 1. Primers were synthesized by Integrated DNA Technologies (IDT, IA, USA). Each biological replicate was done in duplicate and amplifications were carried out in a Techne TechGene Peltier thermal cycler and products separated on ethidium bromide-stained agarose gels.
RNA Isolation, Endpoint RT-PCR, and Quantitative RT-PCR Analyses
Total RNA was isolated from resuspended bacterial pellets using TRIzol LS reagent (Invitrogen) and RNA was purified according to the manufacturer’s instructions. Purified RNA was reconstituted in RNase-free water and any contaminating DNA was removed by treating with Turbo DNase (Ambion, TX, USA) following the manufacturer’s recommendations. RNA was quantified using Qubit 3.0 fluorometer (Invitrogen) and Qubit RNA BR Assay Kit (Invitrogen). cDNA synthesis (reverse transcription at 50°C for 30 min followed by inactivation at 95°C for 15 min) of RNA extracts was performed using the OneStep RT-PCR Kit (Qiagen) in a total volume of 25 μL, with 1 μg total RNA and 0.6 μM of each primer (listed in Table 1), and components of OneStep RT-PCR enzyme mix with Omniscript and Sensiscript reverse transcriptases, based on the manufacturer’s instructions. Endpoint RT-PCR was performed to confirm transcription of the genes being investigated and each biological replicate was done in duplicate.
Quantitative RT-PCR (qRT-PCR) analyses were conducted using custom Taqman Gene Expression assays with fluorescent reporter dye, 6-carboxy-fluorescein (FAM)-labeled primer pairs and probe and Taqman Fast Virus 1-step master mix (Applied Biosystems, CA, USA). Virulence-associated genes analyzed included lenA, lipL32, lipL41, loa22, and sph2. These were selected for further study based on their consistent expression in both species in the previous endpoint RT-PCR analyses. The probes were designed using the software programme Primer ExpressTM (Applied Biosystems), for compatibility with primer sequences used in endpoint RT-PCR: lipL32 (FAM-CCAGGGACAAACGAA-MGBNFQ), lipL41 (FAM-ATCAGATGCCTTCTAAAG-MGBNFQ), loa22 (FAM-CGCAGAAGCAAACA-MGBNGQ), lenA (FAM-AGTTTAACGGGAGCTTAT-MBGNFQ), and sph2 (AM-CACGCTCAACCACC-MGBNFQ). Taqman primer pairs and probes were synthesized by Applied Biosystems in a custom gene expression assay. The labeled MGB probe had the FAM located at the 5′ end of the probe and a non-fluorescent quencher (NFQ) at the 3′ end. For qRT-PCR, cDNA was synthesized in a total reaction volume of 20 μL containing 0.1 μg total RNA with components of the 4x Taqman Fast Virus 1-Step Master mix (Applied Biosystems), 20x Taqman Custom gene expression assay and RT-PCR grade H2O, to provide a final concentration of 5 μM labeled probe and 18 μM of each primer. Amplification was done as singleplex reactions with each biological replicate being amplified in duplicate. Controls with each run included a no-template control (NTC) that contained all the listed reagents except the RNA template and a no-enzyme control to detect the presence of contaminating DNA. Thermal cycling was performed in a 7500 Fast Real-Time PCR System (Applied Biosystems), using the following parameters: reverse transcription at 50°C for 5 min, inactivation at 95°C for 20 s, followed by 40 cycles at 95°C for 3 s and 60°C for 30 s. Gene expression results were reviewed for run validity. Negative reactions were assigned where no amplification occurred at threshold cycle (CT) greater than 38 cycles. The gene expression data (gene abundance) from the qPCR experiments were means of duplicate biological replicates quantified based on a 3-point standard curve; these were relative values of pathogenic leptospiral total RNA. Because of the dispersion of the data, it was necessary to transform them using logarithm base-10. Analysis of Variance (ANOVA) was used to evaluate the differential expression. Expression data for the five genes were normalized using gene expression values the 16S rRNA gene at 30°C for the two strains and analyzed to assess individual contributions of parameters (temperature or oxidative stress – level and duration) to the bacteria (together and individually). Graphs depicting relative fold expression for each treatment were derived using the ΔΔCT formula, with normalization against the 16S rRNA gene expression at 30°C.
Results
Gene Expression of Virulence-Associated Genes in L. borgpetersenii Jules and L. interrogans Portlandvere Exposed to Temperature Stress Conditions
For L. borgpetersenii serovar Jules, 12 (85.7%) of the 14 virulence associated genes were detected at 30°C and included adhesins lenA, lsa21 and loa22; haemolysins sph2 and tlyC; OMP porin ompL1; invasin invA; OMP lipoproteins lipL32, lipL36, lipL41, and lipL45 and the fliY gene involved in chemotaxis. For L. interrogans serovar Portlandvere, 13 (92.9%) of the 14 virulence associated genes were also amplified and included all except ligB.
The expression of five genes, lenA, lipL32, lipL41, loa22, and sph2 in Jules and Portlandvere were selected for quantification determination by quantitative real time RT-PCR. As illustrated in Figure 1 and detailed in Table 2, in most cases, temperature stress resulted in significant changes in gene expression within species or between species (Jules vs. Portlandvere). Firstly, we observed varying degrees of decreased expression of lipL32, loa22, and sph2 in Jules at 30°C compared to 37°C, while the expression of lipL41 was unchanged with the elevation of temperature. In Portlandvere, the increased expression of lenA at 37°C compared to 30°C, contrasts that of loa22 and sph2 where decreased gene expression was observed. Secondly, in Jules exposed to upshifted temperature conditions compared to 30°C, expression levels of lipL41, loa22, and sph2 were similarly higher, while those for lenA and lipL32 decreased in bacteria at 30°C compared to upshifted conditions. Thirdly, expression levels of all genes were increased, except for loa22 in Portlandvere at 30°C compared to upshifted conditions. Fourthly, only lipL32 (in Jules) and lipL41 (in Portlandvere) expression showed increased expression at 37°C compared to upshifted conditions. Overall, expression of the virulence-associated genes observed in Portlandvere was generally lower relative to expression in Jules. When inter-species comparisons were carried out, we noted that at 30°C, there were higher levels of expression of lenA and sph2 (in Jules compared to Portlandvere) and loa22 (in Portlandvere compared to Jules). However, at upshifted temperatures, lenA, sph2, and lipL41 were expressed at a higher level in Jules compared to lipL32 in Portlandvere. Finally, expression of lenA was significantly elevated in Portlandvere at 37°C compared to Jules.
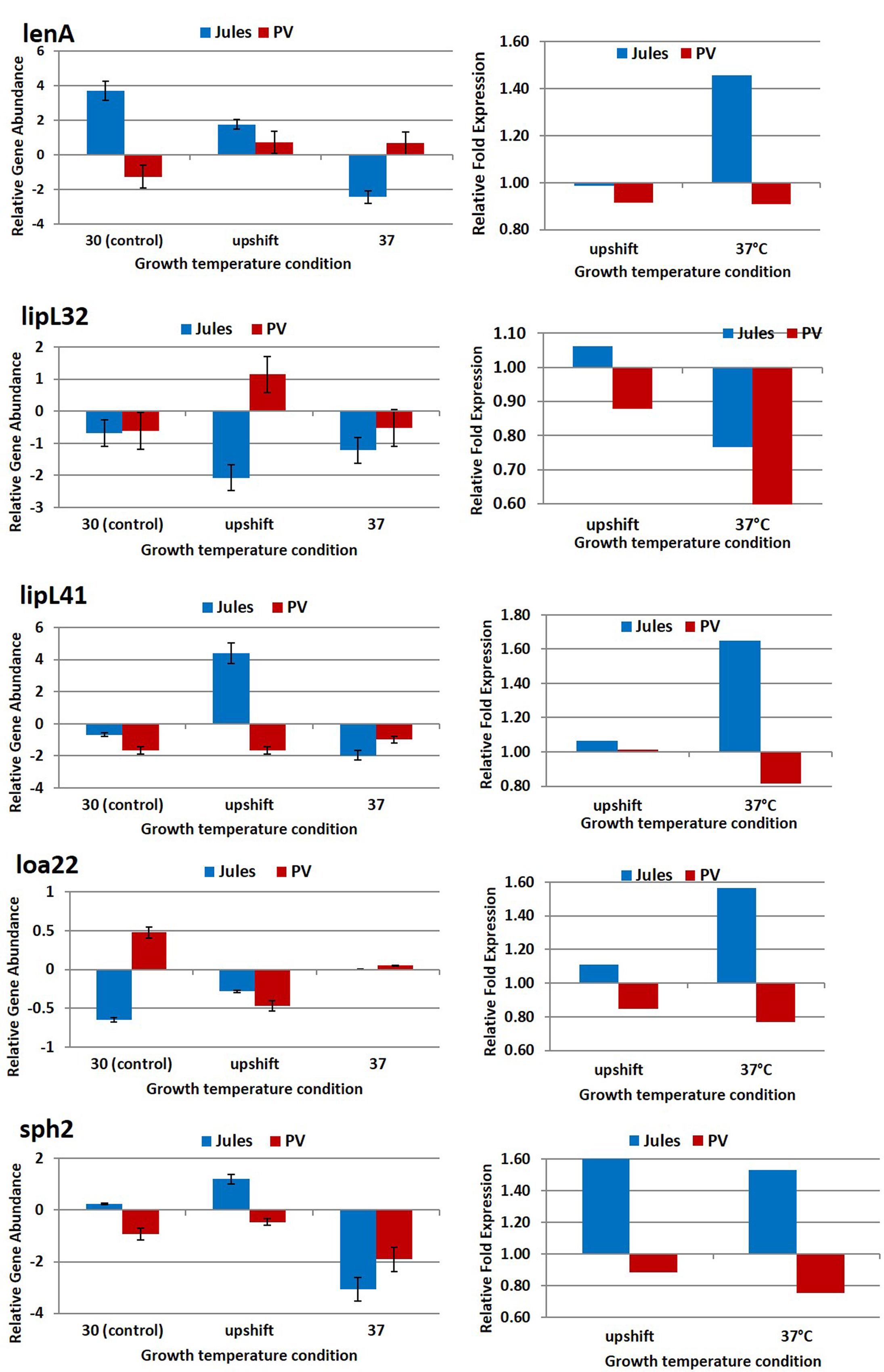
FIGURE 1. Comparative analysis of the effects of temperature on transcription of five virulence associated genes in Leptospira Jules and Leptospira Portlandvere (PV) using qRT-PCR. Each gene is represented as relative gene abundance (based on log10-transformed gene expression data) with error bars (standard error of mean) for bacteria exposed to 30°C, upshift conditions, and 37°C. Gene expression of 16S rRNA at 30°C served as controls for the purpose of normalization of gene expression at upshift and at 37°C conditions, and calculation of fold changes (shown on the right-hand side).
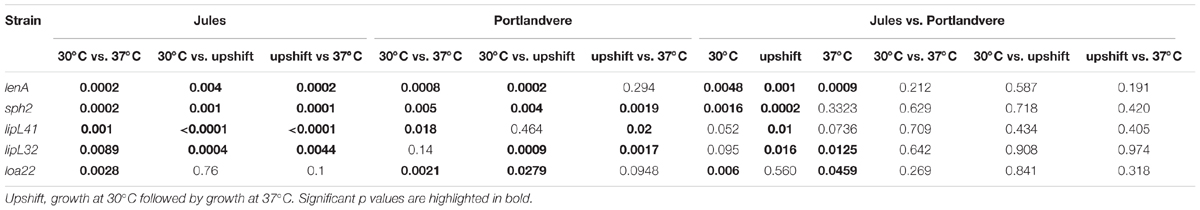
TABLE 2. P-values associated with comparative analysis of the effects of temperature on transcription of five virulence associated genes in L. Jules and L. Portlandvere using qPCR.
The greatest fold increases in gene expression was observed for lenA, lipL41, and loa22 (1.4–1.6-fold) in Jules at 37°C and sph2 (1.5–1.6-fold) in Jules at upshift and 37°C.
Qualitative Gene Expression of Virulence-Associated Genes in L. borgpetersenii Jules and L. interrogans Portlandvere Exposed to Oxidative Stress Conditions
Hydrogen peroxide, H2O2, a ROS, causes alterations in cellular redox potential, where small perturbations stimulate the cell’s anti-oxidant system and larger changes result in apoptosis and necrosis. H2O2-induced cytotoxicity derived from destabilization of cellular components such as DNA, proteins, and lipids, is enhanced by its lipid solubility which facilitates diffusion across cellular membranes. However, much of the damage caused by ROS occurs within the DNA structure to effect base damage and DNA nicking, leading to mutations.
Generally, growth at 37°C coupled with hydrogen peroxide-induced oxidative stress resulted in gene expression significantly higher in L. interrogans Portlandvere compared to L. borgpetersenii Jules. This increase in gene expression was more noticeable in upshifted Portlandvere compared to bacteria at 30 and 37°C.
Specifically, differential gene expression under oxidative conditions yielded expression of loa22, lipL32, and lipL41 in both Jules and Portlandvere with fliY and lsa21 transcripts observed solely in Portlandvere. Collective gene expression among upshifted Jules and Portlandvere yielded more transcripts compared to growth at 30 and 37°C, with growth at 37°C yielding the least gene transcripts. This was particularly noticeable under conditions of low oxidative intensity, 1 mM H2O2 and varied durations of 30 and 60 min oxidative exposure. Of note, lsa21, transcribed in Portlandvere, appeared to be preferentially expressed at low oxidative intensity, regardless of oxidative duration in bacteria grown at 30°C and upshifted temperature conditions. Thus, it is possible that lsa21 transcription may be both temperature-sensitive (negligible detection at long term growth at 37°C) and oxidative stress-sensitive (detection at ≤1 mM H2O2). Similar to lsa21, fliY expression may be temperature sensitive as long term growth at 37°C yielded reduced fliY expression in Portlandvere compared to transcription at all oxidative conditions in bacteria grown at 30°C and upshifted temperatures. Further, neither gene was visibly expressed at 30°C in Portlandevere, suggestive of reduced responsiveness to oxidative stress.
Oxidative stress conditions resulted in differential gene expression: in Portlandvere, loa22 and lipL32 were expressed in all 12 combinations of growth temperature, oxidative intensity and duration compared to Jules, however, at 30°C loa22 was preferentially expressed at low oxidative intensity regardless of duration. On the other hand, the expression of loa22 was sensitive to oxidative duration at upshifted temperature conditions in Jules, as it was expressed at 60 min duration regardless of intensity.
Quantitative Gene Expression of Virulence-Associated Genes in L. borgpetersenii Jules and L. interrogans Portlandvere Exposed to Oxidative Stress Conditions
The quantitative effect of oxidative stress on the expression of the five selected virulence-associated genes were analyzed by trend analysis for bacteria exposed to three temperature conditions, 30, 37, and upshifted conditions, each with subsequent independent exposure to the four hydrogen peroxide-induced oxidative stress states of 1 mM peroxide for 30 min (low oxidative intensity/short durational exposure); 10 mM peroxide for 30 min (high oxidative intensity/short durational exposure); 1 mM peroxide for 60 min (low oxidative intensity/long durational exposure) and 10 mM peroxide for 60 min (high oxidative intensity/long durational exposure. Gene expression in unexposed cultures at the respective growth temperatures were used as comparative baseline control values.
Selected genes were differentially expressed by Jules and Portlandvere as temperature changes had a measureable impact on the transcriptional response observed for lipL41, lipL32, and loa22 (Figures 2–6). For each gene analyzed, the fold change in expression was illustrated below the relative gene abundance plots. Specifically, lenA expression was significantly reduced in Jules at low oxidative intensity (p = 0.007) and between exposure times (p = 0.02). The other gene that had significant change was sph2 in Jules for short (p = 0.004) and long exposure (p = 0.02), and between low and high intensities (p = 0.003). When expression in Jules was compared with that in Portlandvere, we noted that all genes except lipL41 were significantly different.
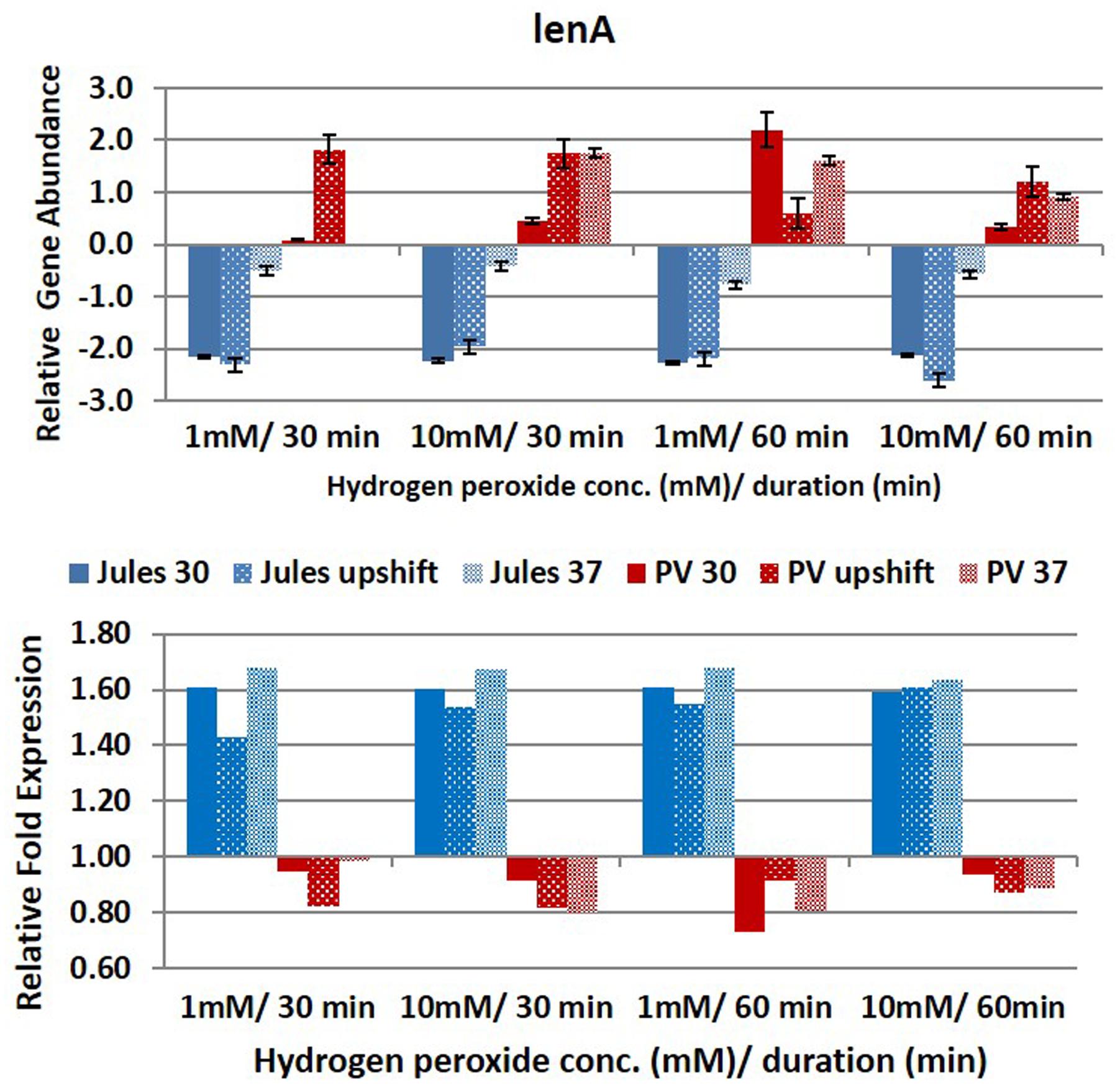
FIGURE 2. Comparative analysis of the effects of peroxide-induced oxidative stress on transcription of gene lenA in L. Jules and L. Portlandvere (PV) using qRT-PCR. The gene is represented as relative gene abundance (based on log10-transformed gene expression data) with error bars (standard error of mean) for bacteria exposed to hydrogen peroxide at 1 mM/30 min, 10 mM/30 min, 1 mM/60 min, 10 mM/60 min, at the three temperature conditions. Gene expression of 16S rRNA at 30°C (without oxidative stress) served as controls for the purpose of normalization of gene expression in the presence of oxidative stress at the various combinations of exposures, and calculation of fold changes (shown below).
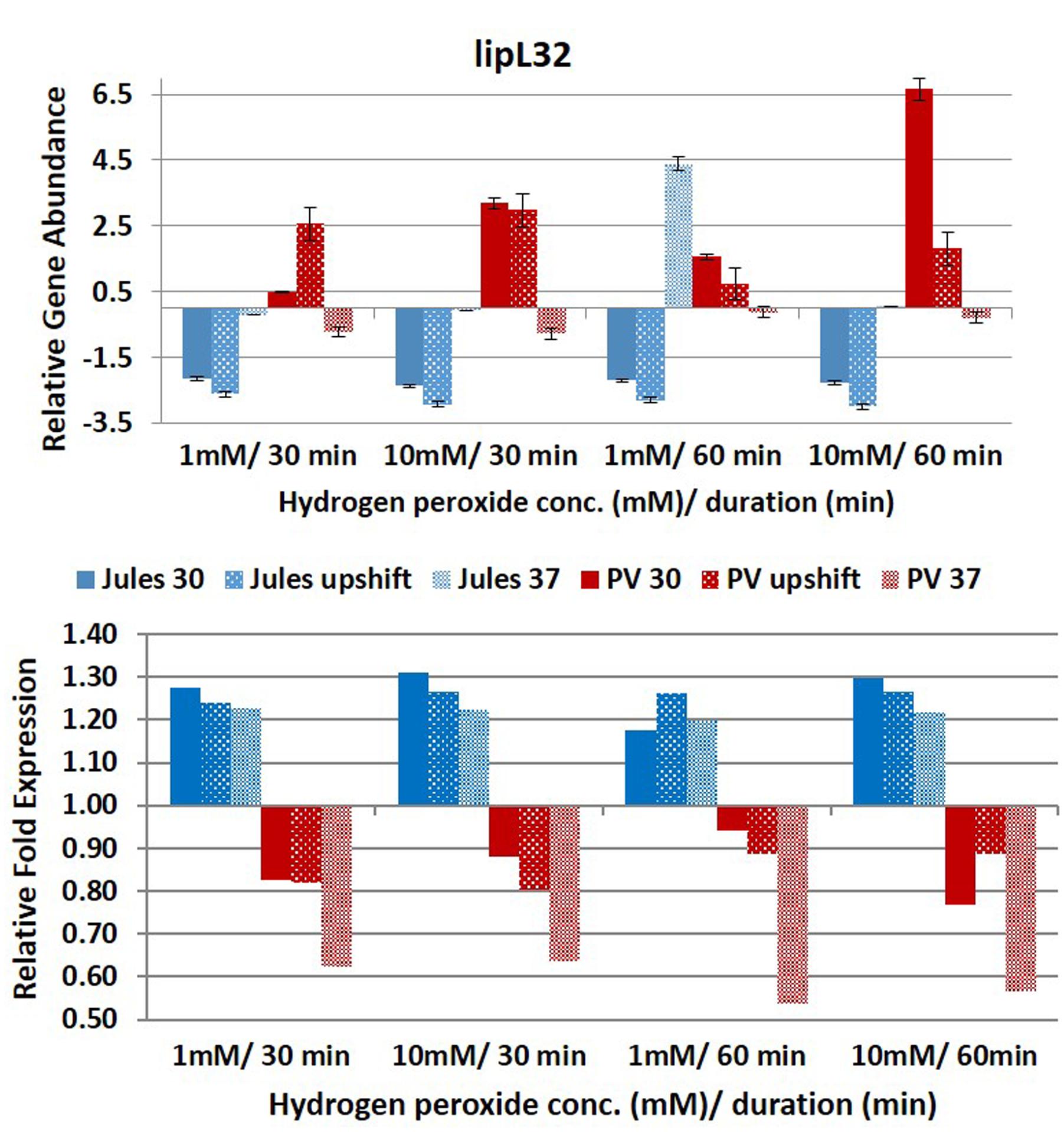
FIGURE 3. Comparative analysis of the effects of peroxide-induced oxidative stress on transcription of gene lipL32 in L. Jules and L. Portlandvere (PV) using qRT-PCR. The gene is represented as relative gene abundance (based on log10-transformed gene expression data) with error bars (standard error of mean) for bacteria exposed to hydrogen peroxide at 1 mM/30 min, 10 mM/30 min, 1 mM/60 min, 10 mM/60 min, at the three temperature conditions. Gene expression of 16S rRNA at 30°C (without oxidative stress) served as controls for the purpose of normalization of gene expression in the presence of oxidative stress at the various combinations of exposures, and calculation of fold changes (shown below).
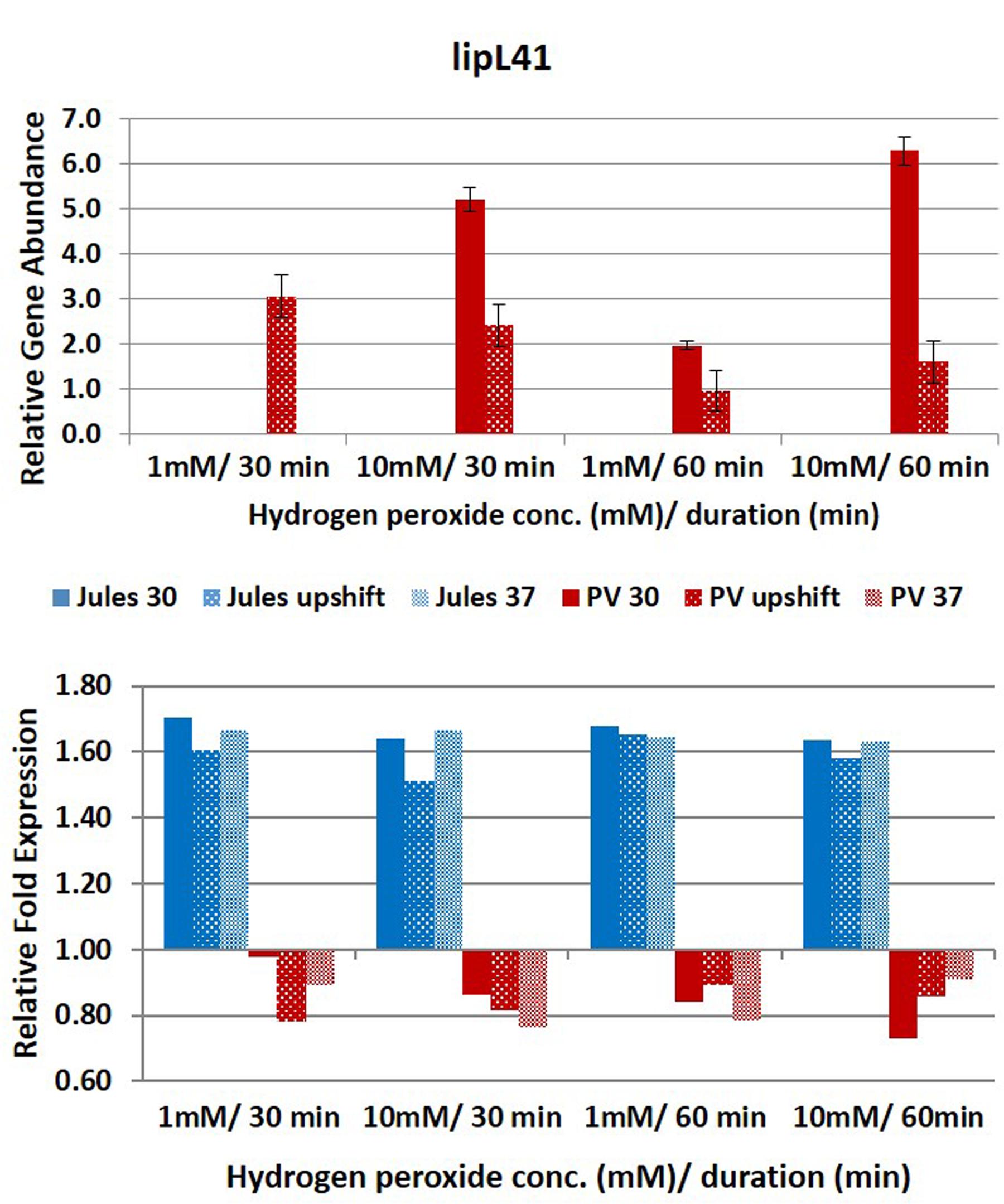
FIGURE 4. Comparative analysis of the effects of peroxide-induced oxidative stress on transcription of gene lipL41 in L. Jules and L. Portlandvere (PV) using qRT-PCR. The gene is represented as relative gene abundance (based on log10-transformed gene expression data) with error bars (standard error of mean) for bacteria exposed to hydrogen peroxide at 1 mM/30 min, 10 mM/30 min, 1 mM/60 min, 10 mM/60 min, at the three temperature conditions. Gene expression of 16S rRNA at 30°C (without oxidative stress) served as controls for the purpose of normalization of gene expression in the presence of oxidative stress at the various combinations of exposures, and calculation of fold changes (shown below).
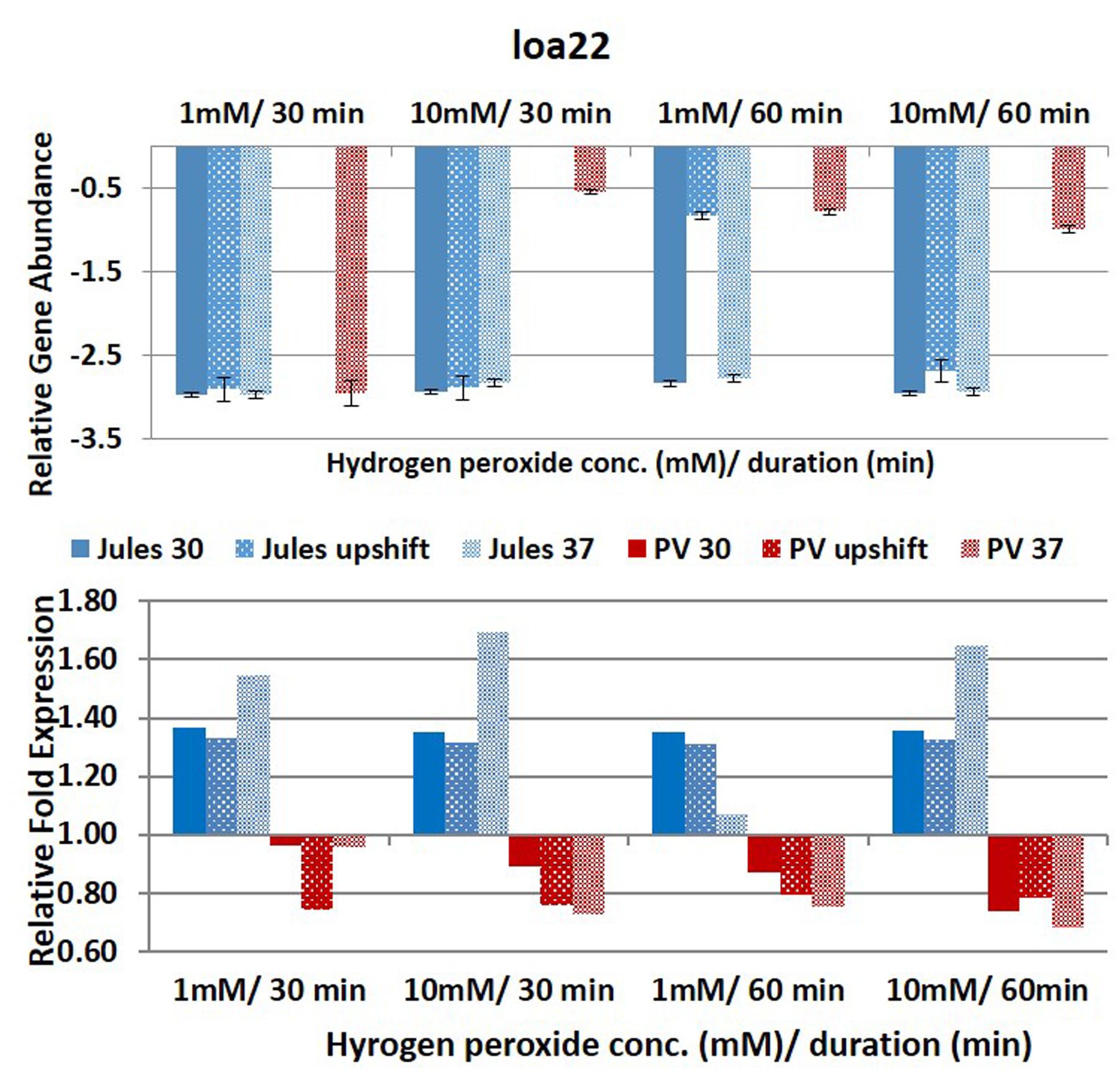
FIGURE 5. Comparative analysis of the effects of peroxide-induced oxidative stress on transcription of gene loa22 in L. Jules and L. Portlandvere (PV) using qRT-PCR. The gene is represented as relative gene abundance (based on log10-transformed gene expression data) with error bars (standard error of mean) for bacteria exposed to hydrogen peroxide at 1 mM/30 min, 10 mM/30 min, 1 mM/60 min, 10 mM/60 min, at the three temperature conditions. Gene expression of 16S rRNA at 30°C (without oxidative stress) served as controls for the purpose of normalization of gene expression in the presence of oxidative stress at the various combinations of exposures, and calculation of fold changes (shown below).
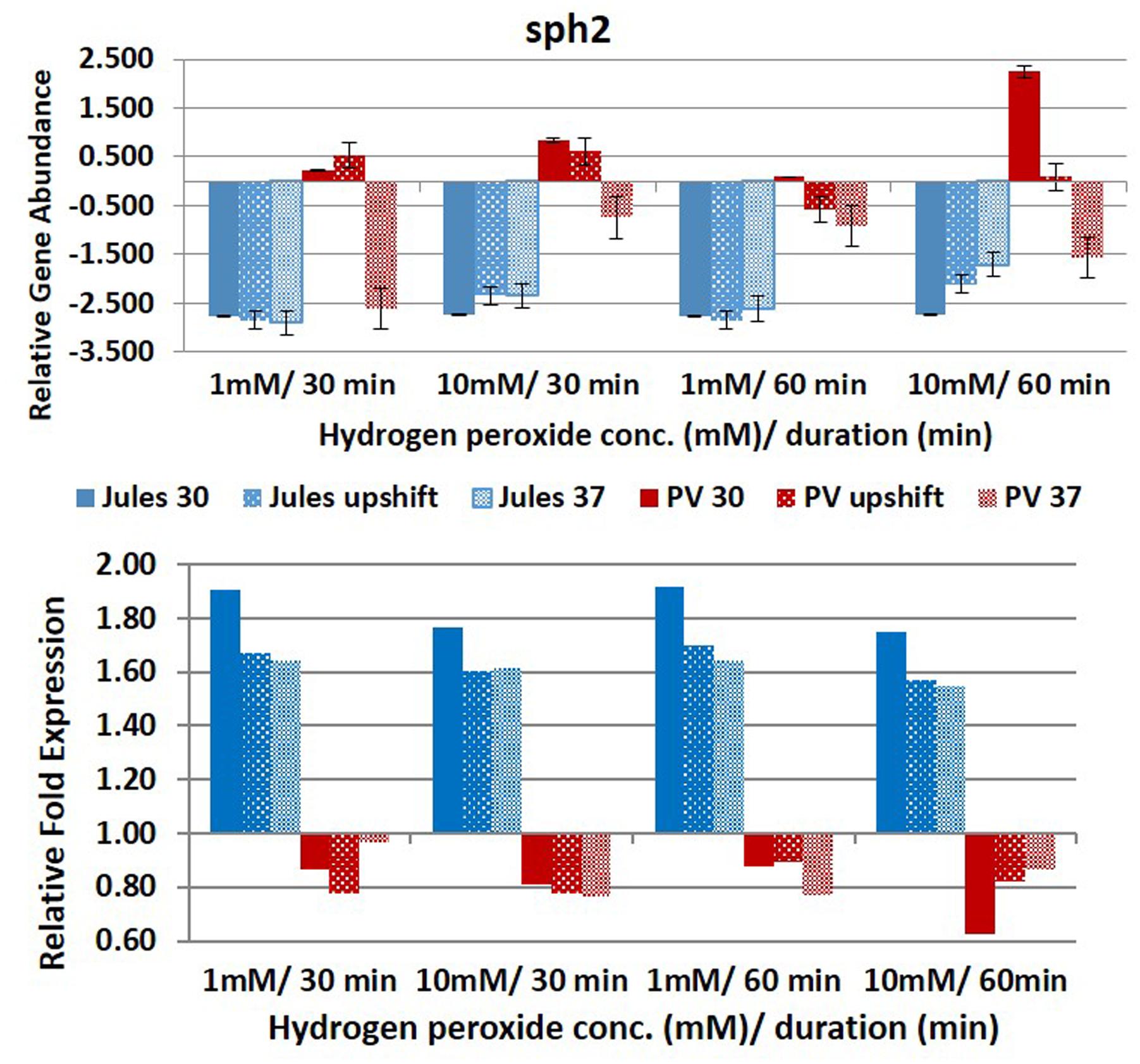
FIGURE 6. Comparative analysis of the effects of peroxide-induced oxidative stress on transcription of gene sph2 in L. Jules and L. Portlandvere (PV) using qRT-PCR. The gene is represented as relative gene abundance (based on log10-transformed gene expression data) with error bars (standard error of mean) for bacteria exposed to hydrogen peroxide at 1 mM/30 min, 10 mM/30 min, 1 mM/60 min, 10 mM/60 min, at the three temperature conditions. Gene expression of 16S rRNA at 30°C (without oxidative stress) served as controls for the purpose of normalization of gene expression in the presence of oxidative stress at the various combinations of exposures, and calculation of fold changes (shown below).
With the complexity of the oxidative stress response so intricately linked with heat stress response, it was not surprising to note that oxidative intensity and duration were influential at upshifted and elevated growth temperatures. The intensity of H2O2-induced oxidative stress was an essential element in both lenA and sph2 expressions. An increase in oxidative stress intensity from 1 to 10 mM H2O2 in Jules at 37°C and upshifted conditions yielded increased sph2 and lenA expression, respectively. Further, increased duration of exposure from 30 to 60 min resulted in decreased lenA transcription among upshifted cultures of Jules.
Compared to Jules where temperature was instrumental in only two of five genes, temperature was an essential element in four of five genes (lipL32, lipL41, loa22, and sph2) in Portlandvere subjected to oxidative stress. A temperature change from 30°C to 37°C was important for the decreased expressions of lipL32 (p = 0.0280) and sph2 (p = 0.0342) but extremely important in the reduced transcription of loa22 (p < 0.0001), much lower in Portlandvere at 37°C and upshifted temperatures. Sph2 transcripts were significantly lower in Portlandvere at 30°C (p = 0.0342), particularly following 60 rather than 30 min of oxidative exposure and 10 mM H2O2 rather than low oxidative intensity. Temperature changes from 30°C to upshift were more important in decreased loa22 expression (p = 0.0007) whereas changes from upshift to 37°C were impactful on decreased lipL41 (p = 0.0119) and lipL32 (p = 0.0304) transcripts in Portlandvere. No other combinations, when analyzed, were significantly different.
At low oxidative intensity, the interplay between genetic predisposition and temperature was biased towards Portlandvere particularly when grown at 30°C and upshifted temperatures (Figures 2–6). Under these conditions, higher lenA, lipL32, lipL41, and sph2 transcripts were observed in Portlandvere compared to Jules grown at the corresponding temperatures. Further, increased oxidative intensity (10 mM peroxide) was considered an essential element in intra-comparative differential expressions, as all five genes were expressed several fold higher in Portlandvere when compared to the respective cultures of Jules. This was particularly evident at high oxidative intensity/long duration for lipL32, lipL41, loa22, and sph2 gene expression.
In all cases, fold increases in expression of genes was positive (20–70%) for Jules subsequent to normalization with expression data for the 16S rRNA gene.
Discussion
The transmission cycle of pathogenic Leptospira species necessitates the ability to respond to environmental changes and is supported by altered gene expression of leptospiral exoproteins predominantly involved in motility, signal transduction and energy-generating functions during infection (Matsui et al., 2012; Eshghi et al., 2015). This study examined the effects of temperature and oxidative stress on virulence associated genes in L. interrogans serovar Portlandvere and L. borgpetersenii serovar Jules. Several studies have demonstrated differential gene transcription in leptospires transitioned from environmental to host simulated conditions involving changes in temperature (Lo et al., 2006, 2009), osmolarity (Matsunaga et al., 2007); serum exposure (Patarakul et al., 2010), and macrophage interactions (Xue et al., 2010).
In this study, the variability in gene expression may possibly be attributed to any individual and/or combination of growth temperature, strain diversity, oxidative conditions or other unknown factors. In most cases, temperature by itself resulted in significant changes in gene expression within the individual strains and between the strains for the five genes analyzed in this study. Not surprisingly, but noteworthy, there was generally higher transcription in Portlandvere compared to Jules (notwithstanding the higher fold in Jules increases relative to 16S rRNA expression). This supports reports of loss of gene function involved in environmental sensing and metabolic transport and utilization in L. borgpetersenii (Picardeau, 2015), possibly due to the presence of more pseudogenes (around 12%) in L. borgpetersenii compared to <4% in L. interrogans, and gene reduction in L. borgpetersenii (Bulach et al., 2006). The approximately 700 kb reduced genome of L. borgpetersenii, being about 16% smaller than L. interrogans, exemplifies a restrictive lifecycle of a host to host mode of transmission compared to the adaptability to either aqueous or mammalian host environs for the larger sized genome of L. interrogans. The lack of expression of ligB or mce using both genomic and expression analysis may be due in part to loss or attenuation of these genes in highly passaged cultures, lack of ORFs and/or regulatory dysfunction. In virulent leptospiral strains, ligB has been shown to be upregulated upon exposure to temperature and osmolarity, with its expression lost in high passaged cultures (Bulach et al., 2006). The eleven and nine genes that were negligibly detected using endpoint RT-PCR in Jules and Portlandvere, respectively, may have possibly been expressed below a detectable threshold; subject to amplification failure, pre- or post-transcriptional modifications in the highly passaged cultures and/or responsive to factors other than temperature, oxidative intensity and duration. Differential gene expression in Jules was observed across conditions of low and high oxidative stress intensities for both short and long durations of oxidative stress. We propose that under oxidative stress, the expressions of loa22, lipL32, and lipL41 in Jules are co-regulated by temperature and oxidative stress, as their expressions were reduced at 37°C, and oxidative conditions of long duration yielded the most abundant transcription when the bacteria were grown at 30°C and upshifted temperatures. For Portlandvere, temperature played a greater role in differential gene expression compared to oxidative stress and duration.
Notwithstanding lower lenA expression in Jules, it was clear that there was co-regulation by temperature and oxidative stress in Portlandvere, evidenced by better yields at upshifted temperature vs 30°C, at 10 mM (high oxidative intensity) vs. 1 mM H2O2 (low oxidative intensity), and at 30 min vs 60 min. Interestingly, in the absence of oxidative stress, differential lenA expression in Portlandvere was particularly enhanced at upshifted temperatures and at 37°C compared to 30°C and increased following 30 min oxidative duration, particularly at high oxidation intensity. While the specific function of lenA remains unclear, it is reported to facilitate adherence to host ECM and plasma components to result in degradation of fibrinogen, connective tissue and immunoglobulin (Verma et al., 2010), suggestive of a putative role in leptospiral dissemination and/or evasive strategies. The finding of higher lenA transcripts among upshifted cultures of Portlandvere alludes to the possible involvement at early onset of infection possibly in host recognition to facilitate adhesion, bind plasminogen, overcome host derived ROS, and/or other unknown functions.
In the present study, the expression of the calcium-mediated LipL32, one of several plasminogen binding leptospiral OMPs (Picardeau, 2015), was influenced by temperature and the duration of oxidative stress. Temperature was an essential factor in lipL32 expression in both Jules and Portlandvere where changes from 30°C to upshift conditions resulted in increased and decreased lipL32 transcription in Portlandvere and Jules, respectively. Synergistic changes in temperature and the duration of oxidative stress may also possibly co-regulate lipL32 in Portlandvere, as long rather than short duration was important in the increased lipL32 expression in Portlandvere at 30°C and upshift conditions. This suggests that lipL32 is responsive to oxidative stress for Portlandvere, which contrasts down-regulation of LipL32 upon macrophage interaction with L. interrogans serovar Lai (Xue et al., 2010) and in in vivo studies using animal models of infection (Matsui et al., 2012). With some 38,000 copies per cell, the immunodominant subsurface lipoprotein LipL32 (Pinne et al., 2012), conserved among pathogenic Leptospira, is both highly antigenic and immunogenic, and has been shown to induce a robust inflammatory response via the NF-κB signaling pathway in cultured human and murine renal cells within 2 h via TLR2 activation (Yang et al., 2006; Fitzgerald et al., 2007). This is suggestive of an early inflammatory response likely leading to detrimental effects observed in tubule-interstitial nephritis observed during leptospirosis.
This observation contrasted with decreased lipL32 expression in Jules at upshift conditions and 37°C, although of lower abundance when compared to Portlandvere and warrants further investigation of possible strain specificity and a putative role for LipL32 in the oxidative stress mediated responses of L. borgpetersenii in mammalian hosts. Decreased lipL32 expression in Jules at 37°C and upshift compared to 30°C, decreased lenA and sph2 among Jules grown at 37°C and upshifted temperatures suggest no putative role during early infection. Because L. borgpetersenii is usually transmitted host-to-host transmission where the normal body temperature is ∼37°C, it is likely that the genes are expressed constitutively in Jules with a non-significant regulation by oxidative stress. In light of this, we postulate that lenA might be involved in an anticipatory adaptive response to ‘low dosage pre-exposure’ which facilitate resistance to the damaging effects of host derived H2O2 (Cabiscol et al., 2000). With more genes involved in signal transduction, regulatory and metabolic processes in the larger sized genome of L. interrogans, the role of lipL32 may be relegated to known functions such as outer membrane stabilization, and adhesion of host cells rather than in oxidative stress response.
Further, the results suggest that possible interplay between oxidative stress duration, temperature and strain diversity may have roles in lipL41 expression, as the gene was expressed following both short and long durations of oxidative stress of varying oxidative intensity in Portlandvere at 30°C and preferentially following high oxidative intensity. Conversely, differentially expressed lipL41 displayed temperature sensitivity in Jules, temperature changes from 30°C to upshift were important to yield increased lipL41 transcripts. However, neither changes in oxidative stress intensity nor duration significantly influenced lipL41 transcription in Jules. This decline in lipL41 in Jules as temperature increased from upshift to 37°C, while concurring with similar reports of downregulation of LipL41 upon interaction with macrophage derived cells of L. interrogans at 37°C (Xue et al., 2010), does contrast with other reports which chronicle expression of LipL41 in infection and the urine of rats. On the other hand, the minimal lipL41 expression at upshifted temperatures and the undetectable lipL41 transcripts among L. borgpetersenii Jules grown at 30°C and 37°C concurred with findings of Lo et al. (2009), indicating unaltered lipL41 expression at 30°C and 37°C. This may be suggestive of regulatory factors other than oxidative stress as indicated by Cullen et al. (2004) and Matsui et al. (2012), or lack of co-transcription of the lep chaperone (King et al., 2013).
Expression of loa22, the second most abundant leptospiral OMP (Ristow et al., 2007; Zhang et al., 2010), was noteworthy in Portlandvere compared to Jules, particularly when grown at 30°C and even more so following high oxidative intensity. Temperature was an essential factor in loa22 expression in Portlandvere, where changes from 30°C through to upshift to 37°C resulted in decreased loa22 transcription in Portlandvere. Loa22 sensitivity to intensity of oxidative stress may allude to possible co-regulation as higher yields were observed following treatment with 10mM H2O2 in Portlandvere. One study reported modest downregulation upon macrophage interaction with serovar Lai (Xue et al., 2010) while another attributed differences between leptospiral strains and macrophages leading to strain-specific interactions (Toma et al., 2011). Similarities between leptospiral OmpA-like, Loa22 and OmpA in E. coli suggest that Loa22 may be osmoregulated, growth rate/phase dependent, and with reduced expression at lower than optimal temperatures (Smith et al., 2007).
Increased lipL32 and loa22 transcription in Portlandvere under conditions of oxidative stress observed during this study were converse to down-regulation in L. interrogans Lai reported by Xue et al. (2010). In fact, in that study, alterations in the outer membrane of Lai upon interaction with macrophages resulted in a highly downregulated clade 1 consisting of major OMPs, lipL41, ompL1, lipL32, lipL48, and ompL47 and a moderately downregulated clade 2 comprising lipL45 and loa22, among other genes.
Differential sph2 transcription was observed in Jules vs Portlandvere with possible co-regulation by temperature in Jules (growth at upshift gave better yields vs growth at 30°C or 37°C), or temperature and oxidative intensity in Portlandvere (high oxidative intensity gave better yields than low intensity). For this gene, the duration of oxidative stress was not considered a significant regulatory factor for Jules. Further, temperature by itself was not an important driver and contrasted with the findings of other thermo-regulatory studies by Qin et al. (2006) and Eshghi et al. (2015) where lowered sph2 expression was observed at 37°C. Other studies have shown upregulation of magnesium-sensitive sph2 in the presence of physiological osmolarity (Matsunaga et al., 2007) and during infection (Narayanavari et al., 2015). SphH in L. borgpetersenii and Sph2 in L. interrogans share >50% structural similarity to Smase C of pathogenic Listeria ivanovii and the beta toxin of Staphylococcus aureus (Narayanavari et al., 2012), which mediate escape from phagocytic vacuoles to release the bacterial cells in the cytosol (Gonzálvez-Zorn et al., 1999). Notwithstanding the significant differential expressions observed in this study, we cannot rule out the possibility of observations due to the differences in experimental temperature, harvesting time, source and complexity of oxidative stress (i.e., simplified H2O2 induction versus macrophage interaction).
Conclusion
Differential gene expressions corresponding with temperature changes from 30°C to upshift; 30°C to 37°C and upshift to 37°C and responsiveness to increased intensity and duration of oxidative stress were of keen interest and prompt further investigation of possible role during infection. With an appreciation of the complexity of integrated mechanisms, and genes and gene products involved in the oxidative response of a cell, the in vitro conditions of this study were not meant to simulate physiological conditions of the complex system of oxidative stress, however, the results serve as an important snapshot of selected gene expression in response to temperature and oxidative stress. While it is clear that expression of many virulence genes in highly passaged strains of Leptospira are attenuated or lost, genetic predisposition, changes in growth temperature and/or oxidative intensity and/or duration were factors which acted in isolation or together with other regulatory cues to contribute to the variable gene expression observed in this study. Overall, differential gene expression in serovar Portlandvere was more responsive to temperature and oxidative stress, although relative to 16S rRNA, fold increases in gene expression were associated with Jules.
Author Contributions
TF assisted in the design of the study, carried out the expression assays, performed statistical analyses and drafted the manuscript. PB conceived of the study, its design and coordination and edited the manuscript. Both authors read and approved the final manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgment
Financial support was obtained from an intramural grant from the Office of Graduate Studies and Research, UWI, Mona Campus, Jamaica.
References
Andrade, G. I., and Brown, P. D. (2012). A comparative analysis of the attachment of Leptospira interrogans and L. borgpetersenii to mammalian cells. FEMS Immunol. Med. Microbiol. 65, 105–115. doi: 10.1111/j.1574-695X.2012.00953.x
Anjem, A., and Imlay, A. (2012). Mononuclear iron enzymes are primary targets for peroxide stress. J. Biol. Chem. 287, 15544–15546. doi: 10.1074/jbc.M111.330365
Artiushin, S., Timoney, J. F., Nally, J., and Verma, A. (2004). Host inducible immunogenic sphingomyelinase-like protein Lk 73.5, of Leptospira interrogans. Infect. Immun. 72, 742–749. doi: 10.1128/IAI.72.2.742-749.2004
Barbosa, A., Abreu, P. A. E., Neves, F. O., Atzingen, M. V., Watanabe, M. M., Viria, M. L., et al. (2006). A newly identified leptospiral adhesion mediates attachment to laminin. Infect. Immun. 74, 6356–6364. doi: 10.1128/IAI.00460-06
Batchelor, T., Stephenson, T., Brown, P. D., Amarakoon, D., and Taylor, M. A. (2012). Impact of climate variability on the incidence of leptospirosis in Jamaica. Clim. Res. 55, 79–90. doi: 10.3354/cr01120
Bharti, A. R., Nally, J. E., Ricaldi, J. N., Matthias, M. A., Diaz, M. M., Lovett, M. A., et al. (2003). Leptospirosis: a zoonotic disease of global importance. Lancet Infect. Dis. 3, 757–771. doi: 10.1016/S1473-3099(03)00830-2
Bulach, D., Zuerner, R. L., Wilson, P., Seeman, T., McGrath, A., Cullen, P. A., et al. (2006). Genome reduction in Leptospira borgpetersenii reflects limited transmission potential. Proc. Natl. Acad. Sci. U.S.A. 103, 14560–14565. doi: 10.1073/pnas.0603979103
Cabiscol, E., Tamarit, J., and Ros, J. (2000). Oxidative stress in bacteria and protein damage by ROS. Int. Microbiol. 3, 3–8.
Cameron, C. E. (2015). “Leptospiral structure, physiology, and metabolism,” in Leptospira and Leptospirosis, Vol. 387, ed. B. Adler (Berlin: Springer), 21–41. doi: 10.1007/978-3-662-45059-8_3
Cullen, P. A., Cordwell, S. J., Bulach, D. M., Haake, D. A., and Adler, B. (2002). Global analysis of outer membrane proteins from Leptospira interrogans serovar Lai. Infect. Immun. 70, 2311–2318. doi: 10.1128/IAI.70.5.2311-2318.2002
Cullen, P. A., Haake, D. A., and Adler, B. (2004). Outer membrane proteins of pathogenic spirochetes. FEMS Microbiol. Rev. 28, 291–318. doi: 10.1016/j.femsre.2003.10.004
Cullen, P. A., Xu, X., Matsunaga, J., Sanchez, J., Ko, A. I., Haake, D. A., et al. (2005). Surfaceome of Leptospira species. Infect. Immun. 73, 4853–4863. doi: 10.1128/IAI.73.8.4853-4863.2005
Dong, H., Hu, Y., Xue, F., Sun, D., Ojcius, D. M., Mao, Y., et al. (2008). Characterization of the ompL1 gene of pathogenic Leptospira species in China and cross-immunogenicity of the OmpL1 protein. BMC Microbiol. 17:223. doi: 10.1186/1471-2180-8-223
Eshghi, A., Lourdault, K., Murray, G. L., Bartpho, T., Sermswan, R. W., Picardeau, M., et al. (2012). Leptospira interrogans catalase is required for resistance to H2O2 and for virulence. Infect. Immun. 80, 3892–3899. doi: 10.1128/IAI.00466-12
Eshghi, A., Pappalardo, E., Hester, S., Thomas, B., Pretre, G., and Picardeau, M. (2015). Pathogenic Leptospira interrogans exoproteins are primarily involved in heterotrophic processes. Infect. Immun. 83, 3061–3073. doi: 10.1128/IAI.00427-15
Fillat, M. F. (2014). The FUR (ferric uptake regulator) superfamily: diversity and versatility of key transcriptional regulators. Arch. Biochem. Biophys. 546, 41–52. doi: 10.1016/j.abb.2014.01.029
Fitzgerald, D. C., Meade, K. G., McEvoy, A. N., Lillis, L., Murphy, E. P., MacHugh, D. E., et al. (2007). Tumor necrosis factor- alpha (TNF-alpha) increases nuclear factor kappa B (NF-κB) activity in and interleukin-8 (IL-8) release from bovine mammary epithelial cells. Vet. Immunol. Immunopathol. 116, 59–68. doi: 10.1016/j.vetimm.2006.12.008
Fulda, S., Gormann, A. M., Hori, O., and Samali, A. (2010). Cellular stress responses: cell death and survival. Int. J. Cell Biol. 2010:214074. doi: 10.1155/2010/214074
Gamberini, M., Gómez, R. M., Atzingen, M. V., Martins, E. A. L., Vasconcellos, S. A., Romero, E. C., et al. (2005). Whole-genome analysis of Leptospira interrogans to identify potential vaccine candidates against leptospirosis. FEMS Microbiol. Lett. 244, 305–313. doi: 10.1016/j.femsle.2005.02.004
Gonzálvez-Zorn, B., Domínguez-Bernal, G., Suarez, M., Ripio, M.-T., Vega, Y., Novella, S., et al. (1999). The smcL gene of Listeria ivanovii encodes a sphingomyelinase C that mediate bacterial escape from the phagocytic vacuole. Mol. Microbiol. 33, 510–523. doi: 10.1046/j.1365-2958.1999.01486.x
King, A., Bartpho, T., Sermswan, R. W., Bulach, D. M., Eshghi, A., Picardeau, M., et al. (2013). Leptospiral outer membrane protein LipL41 is not essential for acute leptospirosis but requires a small chaperone protein, lep, for stable expression. Infect. Immun. 81, 2768–2776. doi: 10.1128/IAI.00531-13
Lau, C. L., Smythe, L. D., Craig, S. B., and Weintein, P. (2010). Climate change, flooding, urbanisation and leptospirosis: fuelling the fire? Trans. R. Soc. Trop. Med. Hyg. 104, 631–638. doi: 10.1016/j.trstmh.2010.07.002
Lehmann, J., Corey, V. C., Ricaldi, J. N., Vinetz, J. M., Winzeter, E. A., and Matthias, M. A. (2015). Whole genome shotgun sequencing shows selection on Leptospira regulatory proteins during in vitro culture attenuation. Am. J. Trop. Med. Hyg. 94, 302–313. doi: 10.4269/ajtmh.15-0401
Lo, M., Bulach, D. M., Powell, D. R., Haake, D. A., Matsunaga, J., Paustian, M. L., et al. (2006). Effects of temperature on gene expression patterns in Leptospira interrogans serovar Lai as assessed by whole genome microarrays. Infect. Immun. 74, 5848–5859. doi: 10.1128/IAI.00755-06
Lo, M., Cordwell, S. J., Bulach, D. M., and Adler, B. (2009). Comparative transcriptional and translational analysis of leptospiral outer membrane protein expression in response to temperature. PLoS Negl. Trop. Dis. 3:e560. doi: 10.1371/journal.pntd.0000560
Lo, M., Murray, G. L., Khoo, C. A., Haake, D. A., Zuerner, R. L., and Adler, B. (2010). Transcriptional response of Leptospira interrogans to iron limitation and characterization of PerR homolog. Infect. Immun. 78, 4850–4859. doi: 10.1128/IAI.00435-10
Lourdault, K., Cerqueira, G. M., Wunder, E. A. Jr., and Picardeau, M. (2011). Inactivation of clpB in the pathogen Leptospira interrogans reduces virulence and resistance to stress conditions. Infect. Immun. 79, 3711–3717. doi: 10.1128/IAI.05168-11
Matsui, M., Soupé, M.-E., Becam, J., and Goarant, C. (2012). Differential in vivo gene expression of major leptospiral proteins in resistant or susceptible animal models. Appl. Environ. Microbiol. 78, 6372–6376. doi: 10.1128/AEM.00911-12
Matsunaga, J., Medejros, M. M., Sanchez, Y., and Ko, A. I. (2007). Osmotic regulation of expression of two extracellular matrix-binding proteins and a hemolysin of Leptospira interrogans: differential effects on LigA and Sph2 extracellular release. Microbiology 153, 3390–3398. doi: 10.1099/mic.0.2007/007948-0
Nally, J. E., Whitelegge, J. P., Bassilians, S., Blanco, D. R., and Lovett, M. A. (2007). Characterization of the outer membrane proteome of Leptospira interrogans expressed during acute lethal infection. Infect. Immun. 75, 766–773. doi: 10.1128/IAI.00741-06
Narayanavari, S. A., Lourdault, K., Sritharan, M., Haake, D. A., and Matsunaga, J. (2015). Role of sph2 gene regulation in hemolytic and sphingomyelinase activities produced by Leptospira interrogans. PLoS Negl. Trop. Dis. 9:e0003952. doi: 10.1371/journal.pntd.0003952
Nathan, C., and Shiloh, M. U. (2000). Reactive oxygen and nitrogen intermediates in the relationship between mammalian hosts and microbial pathogens. Proc. Natl. Acad. Sci. U.S.A. 97, 8847–8876. doi: 10.1073/pnas.97.16.8841
Narayanavari, S. A., Kishore, N. M., and Sritharan, M. (2012). Structural analysis of the leptospiral sphingomyelinases: in silico and experimental evaluation of Sph2 as an Mg++-dependent sphingomyelinase. J. Mol. Microbiol. Biotechnol. 22, 24–34. doi: 10.1159/000337013
Pappas, G., Papadimitriou, P., Siozopoulou, V., Christou, L., and Kritidis, N. (2008). The globalization of leptospirosis : worldwide incidence trends. Int. J. Infect. Dis. 12, 351–357. doi: 10.1016/j.ijid.2007.09.011
Patarakul, K., Lo, M., and Adler, B. (2010). Global transcriptomic response of Leptospira interrogans serovar Copenhageni upon exposure to serum. BMC Microbiol. 10:31. doi: 10.1186/1471-2180-10-31
Picardeau, M. (2015). “Genomics, proteomics, and genetics of Leptospira, physiology, and metabolism,” in Leptospira and Leptospirosis, Vol. 387, ed. B. Adler (Berlin: Springer), 43–63.
Pinne, M., Matsunaga, J., and Haake, D. A. (2012). A novel approach to identification of host ligand binding proteins: leptospiral outer membrane protein microarray. J. Bacteriol. 194, 6074–6087. doi: 10.1128/JB.01119-12
Qin, J.-H., Sheng, Y.-Y., Zhang, Z.-M., Shi, Y.-Z., Hu, B.-Y., Yang, Y., et al. (2006). Genome wide transcriptional analysis of temperature shifts in Leptospira interrogans serovar Lai strain 5660. BMC Microbiol. 6:51. doi: 10.1186/1471-2180-6-51
Reis, R. B., Riberio, G. S., Felzemburgh, R. D. M., Santana, F. S., Mohr, S., Melendez, A. X. T. O., et al. (2008). Impact of environment and social gradient on Leptospira infection in urban slums. PLoS Negl. Trop. Dis. 2:e228. doi: 10.1371/journal.pntd.0000228
Ristow, P., Bourhy, P., McBride, F. W. C., and Picardeau, M. (2007). The OmpA-like protein Loa22 is essential for leptospiral virulence. PLoS Pathog. 3:e97. doi: 10.1371/journal.ppat.0030097
Seis, H. (2014). Role of metabolic H2O2 generation Redox signaling and oxidative stress. J. Biol. Chem. 289, 8735–8741. doi: 10.1074/jbc.R113.544635
Smith, S. G., Mahon, V., Lambert, M. A., and Fagan, R. P. (2007). A molecular Swiss army knife: OmpA structure, function and expression. FEMS Microbiol. Lett. 273, 1–11. doi: 10.1111/j.1574-6968.2007.00778.x
Sobota, J., and Imlay, J. (2011). Iron enzyme ribulose-5-phosphate 3 epimerase in Escherichia coli is rapidly damaged by H2O2 but can be protected by manganese. Proc. Natl. Acad. Sci. U.S.A. 108, 5402–5407. doi: 10.1073/pnas.1100410108
Thomas, M. S., and Wigneshweraraj, S. (2014). Regulation of virulence gene expression. Virulence 5, 832–834. doi: 10.1080/21505594.2014.995573
Toma, C., Nohora, T., Mizuyama, M., Koizumi, N., Adler, B., and Suzuki, T. (2014). Leptospiral outer membrane protein LMB216 is involved in enhancement of phagocytic uptake of macrophages. Cell. Microbiol. 16, 1366–1377. doi: 10.1111/cmi.12296
Toma, C., Okura, N., Takayama, C., and Suzuki, T. (2011). Characteristic features of intracellular pathogenic Leptospira in infected murine macrophages. Cell. Microbiol. 13, 1783–1792. doi: 10.1111/j.1462-5822.2011.01660.x
Verma, A., Brissette, C. A., Bowman, A. A., Shah, S. T., Zipfel, P. F., and Stevenson, B. (2010). Leptospiral endostatin-like protein A is a bacterial cell surface receptor for human plasminogen. Infect. Immun. 78, 2053–2059. doi: 10.1128/IAI.01282-09
Vieria, M. L., Vasconcellos, S. A., Gonçales, A. P., de Morais, Z. M., and Nascimento, A. L. T. O. (2009). Plasminogen acquisition and activation at the surface of Leptospira species lead to fibronectin degradation. Infect. Immun. 77, 4092–4101. doi: 10.1128/IAI.00353-09
Xue, F., Dong, H., Wu, J., Wu, Z., Hu, W., Sun, A., et al. (2010). Transcriptional responses of Leptospira interrogans to host innate immunity: significant changes in metabolism, oxygen tolerance and outer membrane. PLoS Negl. Trop. Dis. 4:e857. doi: 10.1371/journal.pntd.0000857
Yang, C. W., Hung, C.-C., Wu, M. S., Tian, Y.-C., Chang, C.-T., Pan, M.-J., et al. (2006). Toll-like receptor 2 mediates early inflammation by leptospiral outer membrane proteins in proximal tubule cells. Kidney Int. 69, 815–822. doi: 10.1038/sj.ki.5000119
Keywords: Leptospira, virulence, temperature, oxidative stress, regulation
Citation: Fraser T and Brown PD (2017) Temperature and Oxidative Stress as Triggers for Virulence Gene Expression in Pathogenic Leptospira spp. Front. Microbiol. 8:783. doi: 10.3389/fmicb.2017.00783
Received: 05 January 2017; Accepted: 18 April 2017;
Published: 09 May 2017.
Edited by:
Fabrice Merien, Auckland University of Technology, New ZealandReviewed by:
Miguel A. De la Cruz, Instituto Mexicano del Seguro Social, MexicoJi Wang, Harvard Medical School, USA
Mansel William Griffiths, University of Guelph, Canada
Copyright © 2017 Fraser and Brown. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Paul D. Brown, cGF1bC5icm93bkB1d2ltb25hLmVkdS5qbQ==
 Tricia Fraser1,2
Tricia Fraser1,2 Paul D. Brown
Paul D. Brown