- 1N.I. Vavilov Institute of General Genetics, Russian Academy of Science, Moscow, Russia
- 2A.N. Belozersky Institute of Physico-Chemical Biology, Lomonosov Moscow State University, Moscow, Russia
Overlapping genes (OGs) are a universal phenomenon in all kingdoms, and viruses display a high content of OGs combined with a high rate of evolution. It is believed that the mechanism of gene overlap is based on overprinting of an existing gene. OGs help virus genes compress a maximum amount of information into short sequences, conferring viral proteins with novel features and thereby increasing their within-host fitness. Analysis of tobamovirus 3′-terminal genes reveals at least two modes of OG organization and mechanisms of interaction with the host. Originally isolated from Solanaceae species, viruses (referred to as Solanaceae-infecting) such as tobacco mosaic virus do not show 3′-terminal overlap between movement protein (MP) and coat protein (CP) genes but do contain open reading frame 6 (ORF6), which overlaps with both genes. Conversely, tobamoviruses, originally isolated from Brassicaceae species (referred to as Brassicaceae-infecting) and also able to infect Solanaceae plants, have no ORF6 but are characterized by overlapping MP and CP genes. Our analysis showed that the MP/CP overlap of Brassicaceae-infecting tobamoviruses results in the following: (i) genome compression and strengthening of subgenomic promoters; (ii) CP gene early expression directly from genomic and dicistronic MP subgenomic mRNA using an internal ribosome entry site (IRES) and a stable hairpin structure in the overlapping region; (iii) loss of ORF6, which influences the symptomatology of Solanaceae-infecting tobamoviruses; and (iv) acquisition of an IRES polypurine-rich region encoding an MP nuclear localization signal. We believe that MP/CP gene overlap may constitute a mechanism for host range expansion and virus adjustment to Brassicaceae plants.
Introduction
Overlapping genes (OGs) are a universal phenomenon in all kingdoms. OGs originate via various mechanisms, including the use of alternative start codons, ribosomal read through, and frameshift mutations (Krakauer, 2000). Viruses, particularly RNA viruses, possess a high content of OGs in addition to a high mutation rate and a high rate of evolution (Belshaw et al., 2008; Simon-Loriere et al., 2013; Geoghegan et al., 2017).
There are several explanations for the abundance of overlapping genes in RNA viruses. The appearance of OGs (i) helps viruses to compress a maximum amount of information into short sequences (Belshaw et al., 2008) and increase their fitness in various hosts (Krakauer, 2000), (ii) constitutes a mechanism of gene expression regulation via translational coupling of functionally related polypeptides (Belshaw et al., 2007), and (iii) represents an effective mechanism for novel gene generation by introducing a new open reading frame (ORF) over an existing one (Pavesi et al., 2013; Simon-Loriere et al., 2013; Saha et al., 2016).
Tobacco mosaic virus (TMV), the first virus to be discovered, along with other members of the tobamovirus group may provide information about the possible roles of OGs. It is believed that tobamoviruses have coevolved with their hosts (co-divergence) since formation of the asterid, rosid, and caryophyllid plant lineages approximately 112.9 million years ago (Gibbs et al., 2015). A second mechanism of primordial tobamovirus evolution is more rapid, as when there is a sudden appearance of a new host plant, which presumably occurs due to human agricultural activity (host switching) (Gibbs et al., 2010). If we take into account Vavilov's ideas about the centers of origin of cultivated plants (Vavilov and Dorofeev, 1992), it is difficult to imagine a different mechanism because the centers of Solanaceae and Brassicaceae plant origin are on different continents. The current host-virus pair reflects the within-host fitness of tobamovirus to the host plant.
Here, we attempt to resolve the putative role of tobamovirus 3′-terminal gene overlap in within-host fitness.
3′-Terminal Gene Overlap and Tobamovirus Classification
The genome of viruses originally isolated from Solanaceae- species (referred to as Solanaceae-infecting) (such as the U1 strain) contains six ORFs and encodes six proteins, four of which are a 126-kDa protein (ORF2) and its read-through derivative of 183-kDa (ORF1), a 30-kDa protein (ORF4) or movement protein (MP), and a coat protein (CP) (ORF5) (Cooper, 2014). The fifth, a 54-kDa protein (ORF3), has an undefined role, if any. The sixth, a 4.8-kDa protein encoded by ORF6, which overlaps ORF4 and ORF5, influences symptomatology but is not found in all tobamovirus species (Canto et al., 2004). The 130-kDa and 180-kDa proteins are translated directly from the genomic RNA, whereas MP and CP are synthesized from their respective subgenomic RNAs that are 5′-capped and 3′-coterminal with the genomic RNA. TMV MP mRNA is bicistronic; as in the genomic RNA, the MP and CP genes do not overlap and are separated by a 3-nt spacer.
Currently, the genus Tobamovirus consists of 35 species (ICTV, virus taxonomy: 2015 release, http://www.ictvdb.org/virusTaxonomy.asp), and TMV is the type species. The natural host ranges for different members of the genus Tobamovirus include Solanaceae, Brassicaceae, Cucurbitaceae, Malvaceae, Cactaceae, Passifloraceae, Fabaceae, Apocynaceae, Cannabaceae, and Orchidaceae (Figure 1). Initially, tobamoviruses were classified into two subgroups (Fukuda et al., 1981) with different genomic locations for their origin of virion assembly (OA). The OA is located within the MP gene in subgroup 1 members, including Solanaceae-infecting tobamoviruses; an exception is odontoglossum ringspot virus, which infects orchids. In subgroup 2 tobamoviruses, such as those infecting Cucurbitaceae and Fabaceae, the OA is contained within the CP gene.
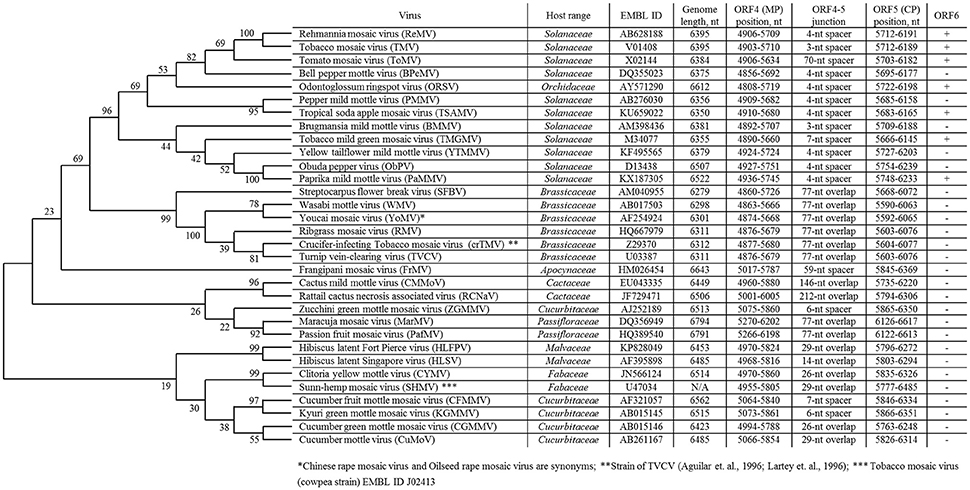
Figure 1. List of overlapping tobamovirus MP/CP genes and tobamovirus molecular phylogenetic analysis based on amino acid sequences of CP proteins using the maximum likelihood method using the JTT matrix-based model (Jones et al., 1992). Evolutionary analyses were conducted in MEGA7 (Kumar et al., 2016).
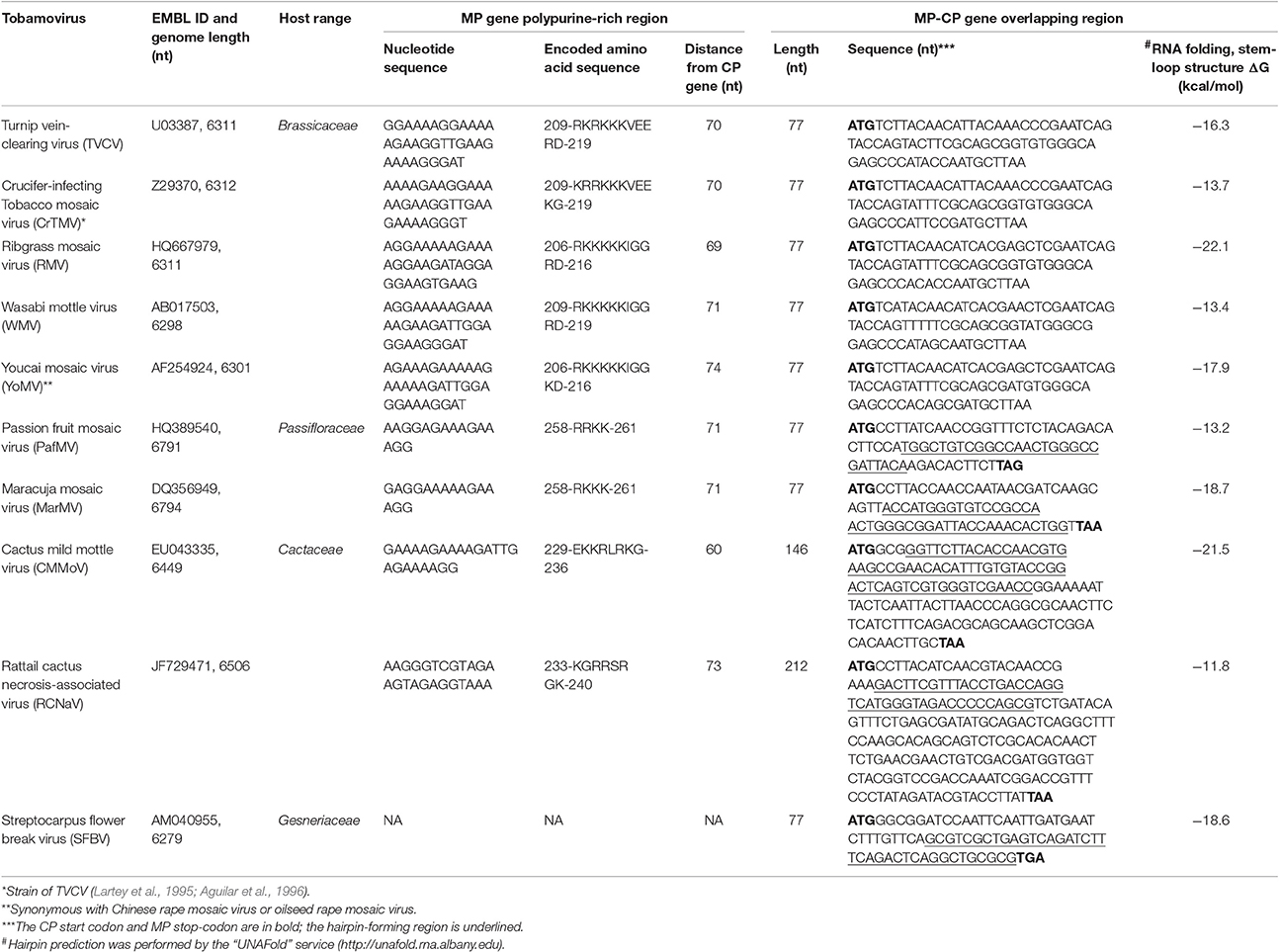
In 1994, we first obtained the complete nucleotide sequence of the crucifer-infecting tobamovirus (crTMV) isolated from a Brassicaceae plant, garlic mustard (Alliaria petiolata) (Dorokhov et al., 1994), which later came to be regarded by ICTV as a strain of turnip vein-clearing virus (TVCV) (Aguilar et al., 1996; Lartey et al., 1996). The genome organization of crTMV is different from that of other tobamoviruses, as the MP and CP ORFs overlap by 77 nt (Table 1). Once the complete nucleotide sequences of the other two viruses originally isolated from Brassicaceae species (referred to as Brassicaceae-infecting) became available (Aguilar et al., 1996; Lartey et al., 1996), a third subgroup of the genus was proposed (Aguilar et al., 1996; Lartey et al., 1996). The OAs of viruses from subgroup 3 are also located in the MP ORF; however, the MP and CP ORFs overlap by 77 nt, and members of this subgroup are able to infect both Solanaceae and Brassicaceae species. Isolation of new tobamoviruses, such as frangipani mosaic virus (Lim et al., 2010), maracuja mosaic virus (Song et al., 2006), passion fruit mosaic virus (Song and Ryu, 2011), cactus mild mottle virus (Min et al., 2006) and rattail cactus necrosis-associated virus (Kim et al., 2012), along with the complete sequencing of their genomes, showed that MP/CP overlap is not unique to Brassicaceae-infecting tobamoviruses but is also present in Apocynaceae-, Passifloraceae-, and Cactaceae-infecting viruses (Figure 1).
The MP/CP overlap of Brassicaceae-infecting tobamovirus is characterized by a +1 frameshift, which is the most frequent for viral gene overlap (Belshaw et al., 2007), yet the mechanisms supporting the evolutionary stability of MP/CP OGs remains unclear. It is known that for a random missense mutation in an overlapping region to remain in the population, this mutation must be beneficial for one of the genes and at least neutral for the other (Brandes and Linial, 2016). The origin and function of the 126/183-kDa OGs confirm a general idea of the origin of eukaryotic and viral OGs (Belshaw et al., 2007) regarding their association with regulation of gene expression via the translational coupling of functionally related polypeptides. Nonetheless, the role of MP/CP OGs functional coupling is unknown.
Unlike the 126- and 183- kDa proteins involved in RNA viral replication functions, MP and CP perform diverse functions. MP is a nonstructural protein, the properties of which are revealed by its interaction with cellular factors associated with the cell-to-cell transport of macromolecules. In contrast, CP is a structural protein, and its amino acid sequence is determined by the mechanism of virion assembly and formation. It can be assumed that joint evolution of primordial virus MP and CP genes led to their overlap, which allowed the virus to utilize Brassicaceae as a host in the evolutionary host-switching process.
Below we define a possible benefit for acquisition of MP/CP overlap by Brassicaceae-infecting tobamoviruses.
Influence of MP/CP Overlap on CP mRNA Synthesis
Confirmation of the dominant role of gene overlap in genome compression can be found in the size of the genome of tobamoviruses that infect both Brassicaceae and Solanaceae and have a 77-nt MP/CP overlap, making their genomic RNA shorter (6298-6312 nt) in comparison with those infecting only Solanaceae (6350-6522 nt) (Figure 1) (Belshaw et al., 2008). However, the size of genomes containing overlapping MP/CP genes in Passifloraceae-infecting tobamoviruses (PafMV and MarMV), with genome lengths close to 6800 nt, does not agree with this assumption (Figure 1). One possible explanation for the appearance of MP/CP gene overlap in tobamovirus evolution is the necessity of strengthening the activity of subgenomic promoters directing the synthesis of a dicistronic and monocistronic mRNA encoding MP and CP, respectively. This view is primarily supported by the results of Dawson et al. (Culver et al., 1993), who observed the earlier and higher expression of MP when the gene was closer to the 3′-nontranslated region (NTR), which contains three pseudoknots followed by a tRNA-like structure. CP gene proximity to pseudoknots also determines gene expression (Dawson, 2014).
The 3′-NTR of Brassicaceae-infecting tobamoviruses is ~30 nt longer than the 3′-NTR of those infecting only Solanaceae; however, as we have shown, crTMV genomic RNA contains six pseudoknots and one additional pseudoknot at the 3′ end of the CP gene (Dorokhov et al., 1994). We can assume that the maximum proximity of the CP gene to pseudoknots promotes CP RNA synthesis from a subgenomic promoter.
Thus, the MP/CP gene overlap of Brassicaceae-infecting tobamoviruses should be evaluated in terms of not only genome compression but also the contribution of strengthening subgenomic promoters as the distance to 3′-NTR pseudoknots is decreased.
Influence of MP/CP Overlap on CP mRNA Translation
TMV CP, in addition to its primary functions of virion assembly and systemic virus movement, is involved in the formation of virus replication complexes (VRCs) in the early stage of infection (Asurmendi et al., 2004). Moreover, CP supports early synthesis of MP, which facilitates the rapid movement of viral genetic material (Bendahmane et al., 2002).
We can assume that early synthesis of CP creates favorable conditions for the virus effective cell-to-cell movement and long-distance spreading. In general, tobamoviruses are characterized by the capacity for rapid and early synthesis of viral proteins, ensuring rapid movement of viral genetic material through the plant and accumulation of viral particles in quantities exceeding the yields of other plant viruses. Apparently, rapid synthesis of proteins and rapid movement from cell to cell allows tobamoviruses to overcome the development of host protective reactions. A benefit with regard to expanding the plant host range is conferred on the virus, which is able to synthesize CP not only from subgenomic mRNA but also from genomic or MP directing dicistronic I2-RNA. Brassicaceae-infecting tobamoviruses appear to have such an advantage, as they are capable of early synthesis of CP through internal translation initiation.
Indeed, internal initiation of translation has been shown for MP gene expression in both TMV U1 and crTMV (Zvereva et al., 2004). In addition, we also revealed that the sequence of crTMV genomic RNA upstream of the CP gene contains a 148-nt internal ribosome entry site (IRES148,CPCR) (Ivanov et al., 1997), which can mediate translation initiation in yeast, plant and animal systems in vitro and in vivo from both dicistronic constructs and constructs with blocked 5′-end-dependent translation (Dorokhov et al., 2002). Sequence analysis of IRESCP,148CR revealed a stem-loop hairpin structure surrounded on both sides by a polypurine-rich region (PPR) with a long 33-nt PPR33 before and a short 11-nt PPR11 after the hairpin. Moreover, PPR33 is sufficient for IRESCP,148CR activity, and a simplified version of such a sequence, consisting of only 16 (GAAA) repeats, also enables high cross-kingdom efficiency of cap-independent translation initiation in plant and mammalian cells (Dorokhov et al., 2002). It should be noted that the corresponding 148-nt region in the TMV U1 RNA exhibits no homology with IRESCP,148CR and is not able to direct internal initiation of translation (Ivanov et al., 1997). Thus, unlike TMV U1, crTMV CP appears to be translated both from subgenomic mRNA by a cap-dependent mechanism and from dicistronic I2-RNA or genomic RNA via internal translation initiation.
To examine the possibility of CP synthesis in a plant cell directly from crTMV genomic RNA, we created a binary vector containing the crTMV infectious copy; in this system, the promoter controlling the synthesis of CP subgenomic mRNA was inactivated via introduction of nucleotide substitutions while IRESCP,148CR remained intact (Dorokhov et al., 2006). Agroinjection experiments allowed us to establish the possibility of CP synthesis from genomic RNA. Moreover, we estimated that the contribution of IRESCP,148CR to the total synthesis of CP during viral infection was at least 3%.
Although this experimental setup leaves some doubts about the IRES's ability to be active under natural conditions when the subgenomic promoter is functional, we hypothesized that internal translation initiation is important for the synthesis of CP, providing early formation of the VRC and enhanced systemic movement of the virus (Dorokhov et al., 2006).
Here, we have analyzed Brassicaceae-infecting tobamoviruses and other tobamoviruses with overlapping MP/CP genes to identify a common crTMV nucleotide sequence or signature for CP gene expression via an internal initiation translation mechanism. Table 1 shows that all examined tobamoviruses with overlapping MP/CP genes are characterized by two features that can contribute to internal translation initiation of the CP ORF. The first feature, previously noted for crTMV, is the PPR of the MP gene, located at a distance of 60–70 nt from the CP start codon. PPR lengths varies among viruses but is completely absent only in streptocarpus flower break virus. The second feature is the existence of a stable stem-loop hairpin structure at the end of the overlapping region, directly upstream of the MP gene termination codon (Table 1). It can be assumed that such a hairpin structure assists in ribosome recognition of the CP initiation codon during translation of the genomic RNA.
An important consequence of the presence of the PPR in the MP gene of Brassicaceae-infecting tobamoviruses is the emergence of an arginine and lysine-rich amino acid stretch (Table 1), which functions as a nuclear localization signal and enables entry of TVCV MP into the nucleus (Levy et al., 2013). This observation led the authors to reasonably presume a nuclear phase of TVCV MP biogenesis, one function of which is blocking host defense. It should be noted that the appearance of overlapping MP/CP genes in Brassicaceae-infecting tobamovirus evolution is accompanied by loss of nested and overlapping ORF6, which is typical of viruses infecting only Solanaceae (Figure 1). ORF6 functions as a pathogenicity factor (Canto et al., 2004), affecting the nuclear apparatus of infected cells (Gushchin et al., 2013; Erokhina et al., 2017). Thus, tobamoviruses may invoke at least two different strategies for influencing the host cell nucleus: (i) via ORF6 overlap with MP and CP genes, in tobamoviruses originally isolated from Solanaceae species, and (ii) by acquiring a new property of MP to infect Brassicaceae.
Concluding Remarks
Current host-virus pairs reflect the tobamoviruses within-host fitness together with optimal combination of tobamoviruses and their host plants. Therefore, the typical tobamovirus U1 strain is adapted to Solanaceae plants, and Brassicaceae-infecting viruses are adapted to the Brassicaceae plants from which they are isolated. An evolutionarily acquired ability to infect Brassicaceae plants is a manifestation of a common-to-all-tobamovirus strategy of genome expression characterized by rapid and early synthesis of viral proteins encoding 3′-terminal genes. Acquisition of MP/CP gene overlap leads to early and enhanced synthesis of CP, and loss of ORF6 by Brassicaceae-infecting tobamoviruses is somehow compensated for by acquisition of the ability to affect nuclear function by MP. MP/CP gene coupling and the evolutionary stability of the overlap could be explained by the benefit to both ORFs and is accompanied by overprinting of the C-terminal sequence of MP, which is not essential for the movement function.
Although our analysis is limited to Solanaceae- and Brassicaceae-infecting tobamoviruses, which are relatively well studied, 3′-terminal OGs are also observed in other viruses infecting other plant families. It can be assumed that their study may in the future reveal other mechanisms of tobamovirus within-host fitness.
Author Contributions
YD designed the sequence analysis; ES performed the bioinformatics analysis; YD and TK wrote the manuscript with input from all authors who reviewed the final paper.
Funding
This work was supported by the Russian Science Foundation (project No. 16-14-00002).
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank reviewers for their constructive remarks and suggestions.
References
Aguilar, I., Sánchez, F., Martin, A., Martinez-Herrera, D., and Ponz, F. (1996). Nucleotide sequence of Chinese rape mosaic virus (oilseed rape mosaic virus), a crucifer tobamovirus infectious on Arabidopsis thaliana. Plant Mol. Biol. 30, 191–197. doi: 10.1007/BF00017814
Asurmendi, S., Berg, R. H., Koo, J. C., and Beachy, R. N. (2004). Coat protein regulates formation of replication complexes during tobacco mosaic virus infection. Proc. Natl. Acad. Sci. U.S.A. 101, 1415–1420. doi: 10.1073/pnas.0307778101
Belshaw, R., Gardner, A., Rambaut, A., and Pybus, O. G. (2008). Pacing a small cage: mutation and RNA viruses. Trends Ecol. Evol. 23, 188–193. doi: 10.1016/j.tree.2007.11.010
Belshaw, R., Pybus, O. G., and Rambaut, A. (2007). The evolution of genome compression and genomic novelty in RNA viruses. Genome Res. 17, 1496–1504. doi: 10.1101/gr.6305707
Bendahmane, M., Szecsi, J., Chen, I., Berg, R. H., and Beachy, R. N. (2002). Characterization of mutant tobacco mosaic virus coat protein that interferes with virus cell-to-cell movement. Proc. Natl. Acad. Sci. U.S.A. 99, 3645–3650. doi: 10.1073/pnas.062041499
Brandes, N., and Linial, M. (2016). Gene overlapping and size constraints in the viral world. Biol. Direct 11, 26. doi: 10.1186/s13062-016-0128-3
Canto, T., MacFarlane, S. A., and Palukaitis, P. (2004). ORF6 of Tobacco mosaic virus is a determinant of viral pathogenicity in Nicotiana benthamiana. J. Gen. Virol. 85, 3123–3133. doi: 10.1099/vir.0.80270-0
Cooper, B. (2014). Proof by synthesis of Tobacco mosaic virus. Genome Biol. 15:R67. doi: 10.1186/gb-2014-15-5-r67
Culver, J. N., Lehto, K., Close, S. M., Hilf, M. E., and Dawson, W. O. (1993). Genomic position affects the expression of tobacco mosaic virus movement and coat protein genes. Proc. Natl. Acad. Sci. U.S.A. 90, 2055–2059. doi: 10.1073/pnas.90.5.2055
Dawson, W. O. (2014). A personal history of virus-based vector construction. Curr. Top. Microbiol. Immunol. 375, 1–18. doi: 10.1007/82_2011_182
Dorokhov, Y. L., Ivanov, P. A., Komarova, T. V., Skulachev, M. V., and Atabekov, J. G. (2006). An internal ribosome entry site located upstream of the crucifer-infecting tobamovirus coat protein (CP) gene can be used for CP synthesis in vivo. J. Gen. Virol. 87, 2693–2697. doi: 10.1099/vir.0.82095-0
Dorokhov, Y. L., Ivanov, P. A., Novikov, V. K., Agranovsky, A. A., Morozov, S. Y., Efimov, V. A., et al. (1994). Complete nucleotide sequence and genome organization of a tobamovirus infecting cruciferae plants. FEBS Lett. 350, 5–8. doi: 10.1016/0014-5793(94)00721-7
Dorokhov, Y. L., Skulachev, M. V., Ivanov, P. A., Zvereva, S. D., Tjulkina, L. G., Merits, A., et al. (2002). Polypurine (A)-rich sequences promote cross-kingdom conservation of internal ribosome entry. Proc. Natl. Acad. Sci. U.S.A. 99, 5301–5306. doi: 10.1073/pnas.082107599
Erokhina, T. N., Lazareva, E. A., Richert-Pöggeler, K. R., Sheval, E. V., Solovyev, A. G., and Morozov, S. Y. (2017). Subcellular localization and detection of tobacco mosaic virus ORF6 protein by Immunoelectron Microscopy. Biochem. Biokhimiia 82, 60–66. doi: 10.1134/S0006297917010060
Fukuda, M., Meshi, T., Okada, Y., Otsuki, Y., and Takebe, I. (1981). Correlation between particle multiplicity and location on virion RNA of the assembly initiation site for viruses of the tobacco mosaic virus group. Proc. Natl. Acad. Sci. U.S.A. 78, 4231–4235. doi: 10.1073/pnas.78.7.4231
Geoghegan, J. L., Duchêne, S., and Holmes, E. C. (2017). Comparative analysis estimates the relative frequencies of co-divergence and cross-species transmission within viral families. PLoS Pathog. 13:e1006215. doi: 10.1371/journal.ppat.1006215
Gibbs, A. J., Fargette, D., García-Arenal, F., and Gibbs, M. J. (2010). Time–the emerging dimension of plant virus studies. J. Gen. Virol. 91, 13–22. doi: 10.1099/vir.0.015925-0
Gibbs, A. J., Wood, J., Garcia-Arenal, F., Ohshima, K., and Armstrong, J. S. (2015). Tobamoviruses have probably co-diverged with their eudicotyledonous hosts for at least 110 million years. Virus Evol. 1:vev019. doi: 10.1093/ve/vev019
Gushchin, V. A., Lukhovitskaya, N. I., Andreev, D. E., Wright, K. M., Taliansky, M. E., Solovyev, A. G., et al. (2013). Dynamic localization of two tobamovirus ORF6 proteins involves distinct organellar compartments. J. Gen. Virol. 94, 230–240. doi: 10.1099/vir.0.045278-0
Ivanov, P. A., Karpova, O. V., Skulachev, M. V., Tomashevskaya, O. L., Rodionova, N. P., Dorokhov, Y. L., et al. (1997). A tobamovirus genome that contains an internal ribosome entry site functional in vitro. Virology 232, 32–43. doi: 10.1006/viro.1997.8525
Jones, D. T., Taylor, W. R., and Thornton, J. M. (1992). The rapid generation of mutation data matrices from protein sequences. Comput. Appl. Biosci. 8, 275–282. doi: 10.1093/bioinformatics/8.3.275
Kim, N. R., Hong, J. S., Song, Y. S., Chung, B. N., Park, J. W., and Ryu, K. H. (2012). The complete genome sequence of a member of a new species of tobamovirus (rattail cactus necrosis-associated virus) isolated from Aporcactus flagelliformis. Arch. Virol. 157, 185–187. doi: 10.1007/s00705-011-1142-5
Krakauer, D. C. (2000). Stability and evolution of overlapping genes. Evol. Int. J. Org. Evol. 54, 731–739. doi: 10.1111/j.0014-3820.2000.tb00075.x
Kumar, S., Stecher, G., and Tamura, K. (2016). MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874. doi: 10.1093/molbev/msw054
Lartey, R. T., Voss, T. C., and Melcher, U. (1995). Completion of a cDNA sequence from a tobamovirus pathogenic to crucifers. Gene 166, 331–332. doi: 10.1016/0378-1119(95)00674-5
Lartey, R. T., Voss, T. C., and Melcher, U. (1996). Tobamovirus evolution: gene overlaps, recombination, and taxonomic implications. Mol. Biol. Evol. 13, 1327–1338. doi: 10.1093/oxfordjournals.molbev.a025579
Levy, A., Zheng, J. Y., and Lazarowitz, S. G. (2013). The tobamovirus turnip vein clearing virus 30-kilodalton movement protein localizes to novel nuclear filaments to enhance virus infection. J. Virol. 87, 6428–6440. doi: 10.1128/JVI.03390-12
Lim, M. A., Hong, J. S., Song, Y. S., and Ryu, K. H. (2010). The complete genome sequence and genome structure of frangipani mosaic virus. Arch. Virol. 155, 1543–1546. doi: 10.1007/s00705-010-0766-1
Min, B. E., Chung, B. N., Kim, M. J., Ha, J. H., Lee, B. Y., and Ryu, K. H. (2006). Cactus mild mottle virus is a new cactus-infecting tobamovirus. Arch. Virol. 151, 13–21. doi: 10.1007/s00705-005-0617-7
Pavesi, A., Magiorkinis, G., and Karlin, D. G. (2013). Viral proteins originated de novo by overprinting can be identified by codon usage: application to the gene nursery of Deltaretroviruses. PLoS Comput. Biol. 9:e1003162. doi: 10.1371/journal.pcbi.1003162
Saha, D., Podder, S., and Ghosh, T. C. (2016). Overlapping regions in HIV-1 genome act as potential sites for host-virus interaction. Front. Microbiol. 7:1735. doi: 10.3389/fmicb.2016.01735
Simon-Loriere, E., Holmes, E. C., and Pagán, I. (2013). The effect of gene overlapping on the rate of RNA virus evolution. Mol. Biol. Evol. 30, 1916–1928. doi: 10.1093/molbev/mst094
Song, Y. S., Min, B. E., Hong, J. S., Rhie, M. J., Kim, M. J., and Ryu, K. H. (2006). Molecular evidence supporting the confirmation of maracuja mosaic virus as a species of the genus Tobamovirus and production of an infectious cDNA transcript. Arch. Virol. 151, 2337–2348. doi: 10.1007/s00705-006-0823-y
Song, Y. S., and Ryu, K. H. (2011). The complete genome sequence and genome structure of passion fruit mosaic virus. Arch. Virol. 156, 1093–1095. doi: 10.1007/s00705-011-0961-8
Vavilov, N. I., and Dorofeev, V. F. (1992). Origin and Geography of Cultivated Plants. Cambridge: Cambridge University Press.
Keywords: overlapping genes, plant virus, virus genome, evolution, tobamovirus, movement protein, coat protein
Citation: Dorokhov YL, Sheshukova EV and Komarova TV (2017) Tobamovirus 3′-Terminal Gene Overlap May be a Mechanism for within-Host Fitness Improvement. Front. Microbiol. 8:851. doi: 10.3389/fmicb.2017.00851
Received: 27 January 2017; Accepted: 25 April 2017;
Published: 11 May 2017.
Edited by:
Ricardo Flores, Universitat Politècnica de València, SpainReviewed by:
Massimo Turina, Consiglio Nazionale Delle Ricerche (CNR), ItalyIsrael Pagan, Universidad Politécnica de Madrid, Spain
Copyright © 2017 Dorokhov, Sheshukova and Komarova. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yuri L. Dorokhov, ZG9yb2tob3ZAZ2VuZWJlZS5tc3Uuc3U=
 Yuri L. Dorokhov
Yuri L. Dorokhov Ekaterina V. Sheshukova
Ekaterina V. Sheshukova Tatiana V. Komarova
Tatiana V. Komarova