- School of Life Sciences, Tokyo University of Pharmacy and Life Sciences, Hachioji, Japan
Shewanella oneidensis MR-1 is a heterotrophic facultative anaerobe that respires using various organic and inorganic compounds. This organism has served as a model to study bacterial metabolic and regulatory systems that facilitate their survival in redox-stratified environments. The expression of many anaerobic respiratory genes in MR-1, including those for the reduction of fumarate, dimethyl sulfoxide, and metal oxides, is regulated by cyclic AMP receptor protein (CRP). However, relatively little is known about how this organism regulates the expression of catabolic enzymes catalyzing the oxidation of organic compounds, including lactate. Here, we investigated transcriptional mechanisms for the lldP (SO_1522) and dld (SO_1521) genes, which encode putative lactate permease and D-lactate dehydrogenase, respectively, and demonstrate that CRP regulates their expression in MR-1. We found that a crp-deletion mutant of MR-1 (Δcrp) showed impaired growth on D-lactate. Complementary expression of dld in Δcrp restored the ability to grow on D-lactate, indicating that the deficient growth of Δcrp on D-lactate is attributable to decreased expression of dld. In vivo transcription and in vitro electrophoretic mobility shift assays reveal that CRP positively regulates the expression of the lldP and dld genes by directly binding to an upstream region of lldP. Taken together, these results indicate that CRP is a global transcriptional regulator that coordinately regulates the expression of catabolic and respiratory pathways in MR-1, including D-lactate dehydrogenase and anaerobic terminal reductases.
Introduction
The genus Shewanella belongs to the class Gammaproteobacteria and is widely distributed in nature, including marine, freshwater, sedimentary, and soil environments (Venkateswaran et al., 1999; Hau and Gralnick, 2007; Fredrickson et al., 2008; Rodionov et al., 2011). Members of this genus are able to utilize a variety of electron acceptors for respiration, such as insoluble solid compounds (e.g., iron and manganese oxides) and soluble organic and inorganic compounds [e.g., oxygen, fumarate, nitrate, nitrite, dimethyl sulfoxide, and trimethylamine N-oxide (TMAO)] (Myers and Nealson, 1988; Fredrickson et al., 2008). The electron-acceptor versatility of this genus may have evolved to allow survival in redox-stratified environments, such as oxic/anoxic interfaces in sediments, where available electron acceptors frequently change.
Shewanella oneidensis MR-1 is the most extensively studied strain in the genus Shewanella. This organism has served as a model to study how bacteria adapt to redox-stratified environments (Fredrickson et al., 2008). Previous studies have shown that MR-1 expresses multiple anaerobic terminal reductase genes, including those for the reduction of metal oxides (omcA and mtrCAB),fumarate (fccA), and DMSO (dmsEFAB), when alternative electron acceptors, e.g., oxygen, are limited (Pirbadian et al., 2014; Barchinger et al., 2016). The simultaneous expression of these genes results in the formation of a dynamic respiratory electron-transfer network consisting of periplasmic and membrane-bound c-type cytochromes, enabling this strain to efficiently discharge electrons to various electron acceptors in response to changes in the environmental redox state (Sturm et al., 2015). Cyclic AMP (cAMP) receptor protein (CRP) plays a central role in the transcriptional regulation of these anaerobic respiratory genes (Saffarini et al., 2003; Fredrickson et al., 2008; Charania et al., 2009; Murphy et al., 2009; Dong et al., 2012; Kasai et al., 2015; Kouzuma et al., 2015). To cite an instance, CRP directly binds to the upstream regions of omcA and mtrC in the presence of cAMP, and activates the transcription of these metal-reduction genes (Kasai et al., 2015). Recent studies have also shown that CRP and cAMP are involved in the regulation of aerobic respiration in MR-1 (Fu et al., 2013; Zhou et al., 2013; Gao et al., 2015; Jin et al., 2016; Yin et al., 2016). These findings are intriguing, since our knowledge of CRP derives mostly from studies on Escherichia coli and other enterobacteria in which CRP is shown to regulate carbon catabolite repression by glucose (Botsford and Harman, 1992; Kolb et al., 1993). Further studies to identify the physiological functions of the cAMP/CRP regulatory system in Shewanella are therefore needed.
In contrast to the relatively well-investigated regulatory mechanisms for the respiratory genes, less is known about how MR-1 regulates catabolic pathways that donate electrons to respiratory pathways. MR-1 preferably utilizes low-molecular-weight organic acids, particularly lactate, as carbon and energy sources under aerobic and anaerobic conditions (Scott and Nealson, 1994; Serres and Riley, 2006). A previous study has identified respiratory L- and D-lactate dehydrogenase (LDH) genes as responsible for the selective oxidation of these isomers to pyruvate in MR-1 (Pinchuk et al., 2009). In this strain, L-LDH is comprised of three subunits encoded by the lldEFG genes (SO_1520 to SO_1518), whereas D-LDH is encoded by the dld gene (SO_1521), a distant homolog of a FAD-dependent LDH gene in yeast (Pinchuk et al., 2009). A previous study has also demonstrated that LlpR (L-lactate-positive regulator, SO_3460) is required for L-lactate utilization by MR-1, suggesting that this regulator is involved in the transcriptional activation of lldEFG (Brutinel and Gralnick, 2012). This work has also uncovered that MR-1 preferentially utilizes D-lactate when both L- and D-lactate isomers are present (Brutinel and Gralnick, 2012). In addition, the expression of dld is up-regulated under oxygen-limited conditions (Barchinger et al., 2016) and high electrode potential-applied conditions in bioelectrochemical systems (Nakagawa et al., 2015), suggesting the possibility that the ability of MR-1 to utilize D-lactate is affected by electron acceptors. These observations suggest that D-lactate is an important catabolic substrate for Shewanella spp., particularly when they grow in anaerobic environments. Nevertheless, the molecular mechanisms underlying the regulation of D-LDH in this genus remain to be elucidated.
Here, we examined the involvement of CRP in the regulation of D-lactate oxidation in S. oneidensis MR-1. We hypothesized that, to thrive in nutrient-limited conditions, bacteria should coordinately regulate electron-donating catabolic pathways (e.g., D-LDH) and electron-consuming respiratory pathways (e.g., metal reductases), and that CRP is involved in this regulation. Findings presented herein provide insights into the coordinated regulation of catabolic and respiratory pathways in bacteria that thrive in the natural environment.
Materials and Methods
Chemicals
Chemicals used in this study were of the highest commercially available purity and purchased from Kanto Chemical (Tokyo, Japan), Wako Pure Chemical (Tokyo, Japan), and Tokyo Kasei Kogyo (Tokyo, Japan). The stock solution of D-lactic acid was neutralized to pH 7.4 with sodium hydroxide before use as a growth substrate for S. oneidensis strains.
Bacterial Strains, Plasmids, and Growth Condition
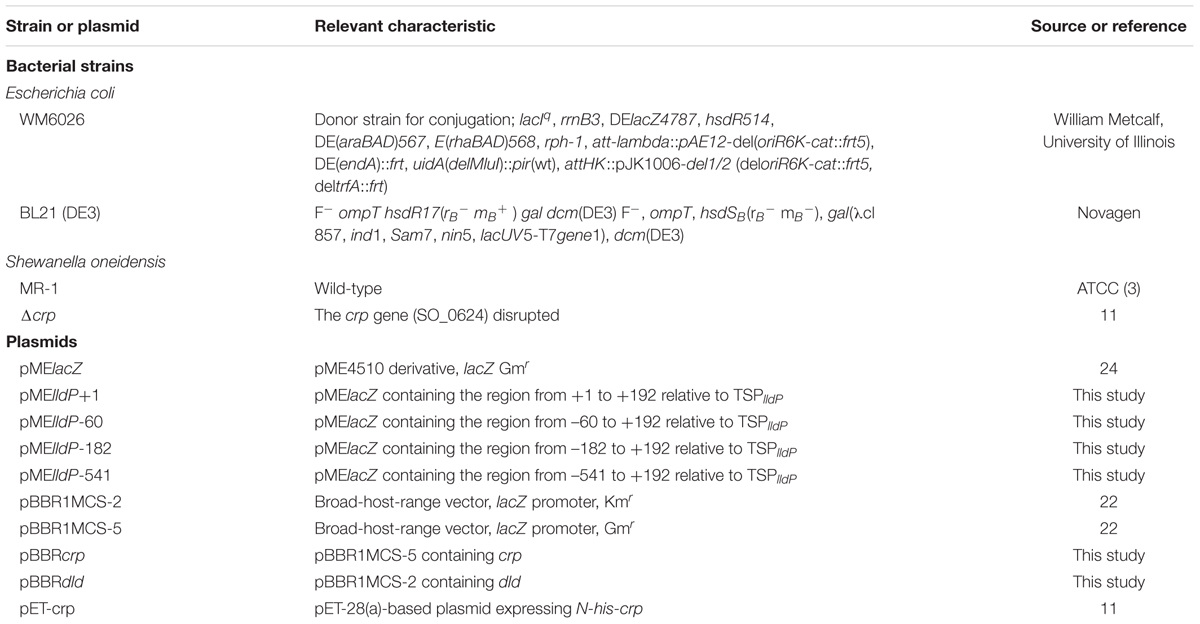
Bacterial strains and plasmids used in the present study are listed in Table 1. Escherichia coli strains were cultivated in Luria-Bertani (LB) or 2 × yeast extract-tryptone (2 × YT) medium at 37°C. The E. coli mating strain (WM6026) required 100 μg/ml 2,6-diaminopimelic acid (DAP) for growth. S. oneidensis strains were cultured at 30°C in LB or minimal medium (MM) (Nakagawa et al., 2015) containing a racemic mixture of DL-lactate, D-lactate, L-lactate, or pyruvate as the carbon and energy source. S. oneidensis strains were grown under aerobic or anaerobic TMAO-reducing conditions, since a CRP-deletion mutant (Δcrp) cannot utilize other electron acceptors (Saffarini et al., 2003). For aerobic cultivation, MM supplemented with each substrate (10 mM) in a test tube was inoculated with an S. oneidensis strain and shaken at 180 rpm. For anaerobic cultivation, MM supplemented with each substrate and TMAO (10 mM or 30 mM) in a test tube was inoculated with an S. oneidensis strain and incubated without shaking. The test tubes containing the anaerobic cultures were capped with butyl rubber septa and polycarbonate screw caps, and purged with pure nitrogen gas. Optical density at 600 nm (OD600) was measured using a mini photo 518R photometer (Taitec, Tokyo, Japan). When necessary, 50 μg/ml kanamycin (Km) or 15 μg/ml gentamicin (Gm) was added to a culture medium. Agar plates contained 1.6% Bacto Agar (Difco).
To construct plasmid pBBRcrp, the crp gene was amplified using Phusion High-Fidelity DNA polymerase (New England Biolabs, Beverly, MA, United States) and primers crp-F-EcoRI and crp-R-BamHI (Supplementary Table S1). The PCR product was digested using EcoRI and BamHI, and cloned between the corresponding sites of pBBR1MCS-5 (Kovach et al., 1995). To construct plasmid pBBRdld, the dld gene was amplified using primers dld-F-BamHI and dld-R-XbaI (Supplementary Table S1). The PCR product was digested using BamHI and XbaI, and cloned between the corresponding sites of pBBR1MCS-2 (Kovach et al., 1995). The resulting plasmids, pBBRcrp and pBBRdld, were introduced into Δcrp cells by filter mating with E. coli WM6026 (Kouzuma et al., 2010).
Measurements of D- and L-Lactate
Concentrations of D- and L-lactate in culture supernatants were determined according to a method described previously (Nakagawa et al., 2015). S. oneidensis strains were cultivated in liquid MM until a stationary growth phase, and cells were removed by filtration through a membrane filter unit (0.20 μm pore size, DISMIC-13JP; Advantec, Tokyo, Japan). Concentrations of D- and L-lactate in the filtrate were measured using F-kit reagents (J. K. International, Tokyo, Japan) according to the manufacturer’s instructions.
RNA Extraction
Shewanella oneidensis cells were cultivated in MM containing 10 mM racemic DL-lactate, D-lactate, or L-lactate under aerobic or 10 mM TMAO-reducing conditions and harvested at the logarithmic growth phase (OD600 of 0.2–0.3 or 0.04–0.06, respectively). RNA was extracted from the cells using a Trizol reagent (Invitrogen, Carlsbad, CA, United States) according to the manufacturer’s instructions. The extracted RNA was purified using an RNeasy Mini Kit and RNase-Free DNase Set (Qiagen, Valencia, CA, United States). The quality of purified RNA was evaluated using an Agilent 2100 Bioanalyzer with RNA 6000 Pico reagents and RNA Pico Chips (Agilent Technologies, Santa Clara, CA, USA) according to the manufacturer’s instructions.
Reverse Transcription (RT)-PCR
To synthesize cDNA, 1.0 μg of total RNA extracted from MR-1 cells grown in MM containing 10 mM racemic DL-lactate as described above was reverse-transcribed using Superscript III Reverse Transcriptase (Invitrogen) and Random Primers (Invitrogen) according to the manufacturer’s instructions. The synthesized cDNA was amplified using Ex Taq DNA polymerase (Takara, Tokyo, Japan) and the primer sets listed in Supplementary Table S1. Amplification steps consisted of an initial denaturation step at 96°C for 30 s, followed by 30 amplification cycles of 96°C for 30 s, 52°C for 30 s, and 72°C for 2 min, and a final extension step at 72°C for 7 min. The PCR products were electrophoresed on 2% agarose gels. Negative control reactions without reverse transcriptase were also performed.
Quantitative RT-PCR (qRT-PCR)
Quantitative RT-PCR (qRT-PCR) was performed using a LightCycler 1.5 instrument (Roche, Indianapolis, IN, United States) according to a method described previously (Kouzuma et al., 2014). A PCR reaction mixture contained 15 ng total RNA, 1.3 μL of 50 mM Mn(OAc)2 solution, 7.5 μL of LightCycler RNA Master SYBR Green I (Roche), and 0.15 μM primers listed in Supplementary Table S1. To generate standard curves, DNA fragments of target genes (dld, lldP, lldF, and 16S rRNA genes) were amplified by PCR using Ex Taq DNA polymerase (Takara) and the primer sets listed in Supplementary Table S1, and purified by gel electrophoresis using a QIAEX II Gel Extraction Kit (Qiagen) according to the manufacturer’s instructions. Standard curves were generated by amplifying a dilution series of the purified DNA fragments of each gene. The specificity of quantitative PCR was verified by a dissociation-curve analysis. Expression levels of target genes were normalized based on expression levels of the 16S rRNA gene.
Identification of Transcriptional Start Sites
5′-Rapid amplification of cDNA ends (5′-RACE) PCR reactions were carried out using 1.0 μg total RNA extracted from MR-1 cells grown in 10 mM racemic DL-lactate up to the logarithmic growth phase and a SMATer RACE cDNA Amplification Kit (Takara) in accordance with the manufacturer’s instructions. The first single-strand cDNA was synthesized using the gene-specific primer lldP_race_out (Supplementary Table S1) and subsequently amplified using Universal Primer A Mix (Takara) and the primer lldP_race_in (Supplementary Table S1). The amplified DNA fragments were purified using a QIAquick PCR Purification Kit (Qiagen). The Purified DNA fragments were cloned into T-Vector pMD20 (Takara) and sequenced to determine the 5′-end points.
LacZ Reporter Assay
To construct a series of reporter plasmids containing promoter regions upstream of lldP, these DNA regions were amplified using Phusion High-Fidelity DNA polymerase (New England Biolabs) and the primer sets listed in Supplementary Table S1. After purification using a QIAquick PCR Purification Kit (Qiagen), the DNA fragments were digested with EcoRI and SalI, and cloned between the corresponding sites of pMElacZ (Endoh et al., 2003). The constructed reporter plasmids were introduced into MR-1 and Δcrp by electroporation according to a method reported previously (Choi et al., 2006). The resulting reporter strains were aerobically cultivated in MM containing 10 mM racemic DL-lactate up to an OD600 of 0.3–0.5. β-Galactosidase activity was measured in triplicate according to the method of Miller (1972).
EMSA
The electrophoretic mobility shift assay (EMSA) using purified N-terminal histidine-tagged CRP (N-his-CRP) was performed according to a method described previously (Kasai et al., 2015). Briefly, N-his-CRP was produced by E. coli BL21(DE3) harboring pET-crp (Kasai et al., 2015), and purified using a QuickPick IMAC Metal Affinity Kit for Proteins (Bio-Nobile, Turku, Finland). The purity of the protein samples was assessed by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) and Coomassie blue staining. Cy3-labeled DNA probes PBlldP1 and PBlldP2 were generated by PCR with the 5′-Cy3 labeled primer set listed in Supplementary Table S1. PBlldP3 and PBlldP3m were generated by annealing of the complementary single-strand oligonucleotides listed in Supplementary Table S1. DNA-binding reactions were performed in 20 μl of a previously described reaction mixture (Kasai et al., 2015) containing 2 nM Cy3-labeled DNA probe, 50 ng N-his-CRP, and 50 μM cAMP.
Results
Deficient D-Lactate Utilization by Δcrp
To investigate the involvement of CRP in the catabolism of D-lactate and other organic acids in S. oneidensis MR-1, we compared growth characteristics of wild-type MR-1 (WT) and Δcrp (Kasai et al., 2015) on a 10 mM racemic mixture of D- and L-lactate (DL-lactate; 5 mM each isomer), D-lactate, L-lactate, and pyruvate (Figure 1). When these strains were grown on DL-lactate, the final cell density of Δcrp was approximately half that of WT (Figure 1A). Measurements of residual D- and L-lactate in the culture media (Figure 2) revealed that a large portion of D-lactate remained in the Δcrp culture. These results suggest that Δcrp has a decreased capacity for metabolizing D-lactate. This notion is supported by the growth deficiency of this mutant in the medium containing D-lactate as the sole substrate (Figure 1B). Complementary expression of crp in Δcrp restored the ability to grow on D-lactate (Supplementary Figure S1), demonstrating the involvement of CRP in D-lactate utilization. However, Δcrp was capable of growing on L-lactate and pyruvate, albeit that growth rates and final cell densities were slightly lower than those of WT (Figures 1C,D). These results suggest that CRP is required for the expression of gene(s) involved in D-lactate utilization, such as the dld gene encoding D-LDH in MR-1, although this regulator is not essential for the expression of genes involved in L-lactate and pyruvate utilization.
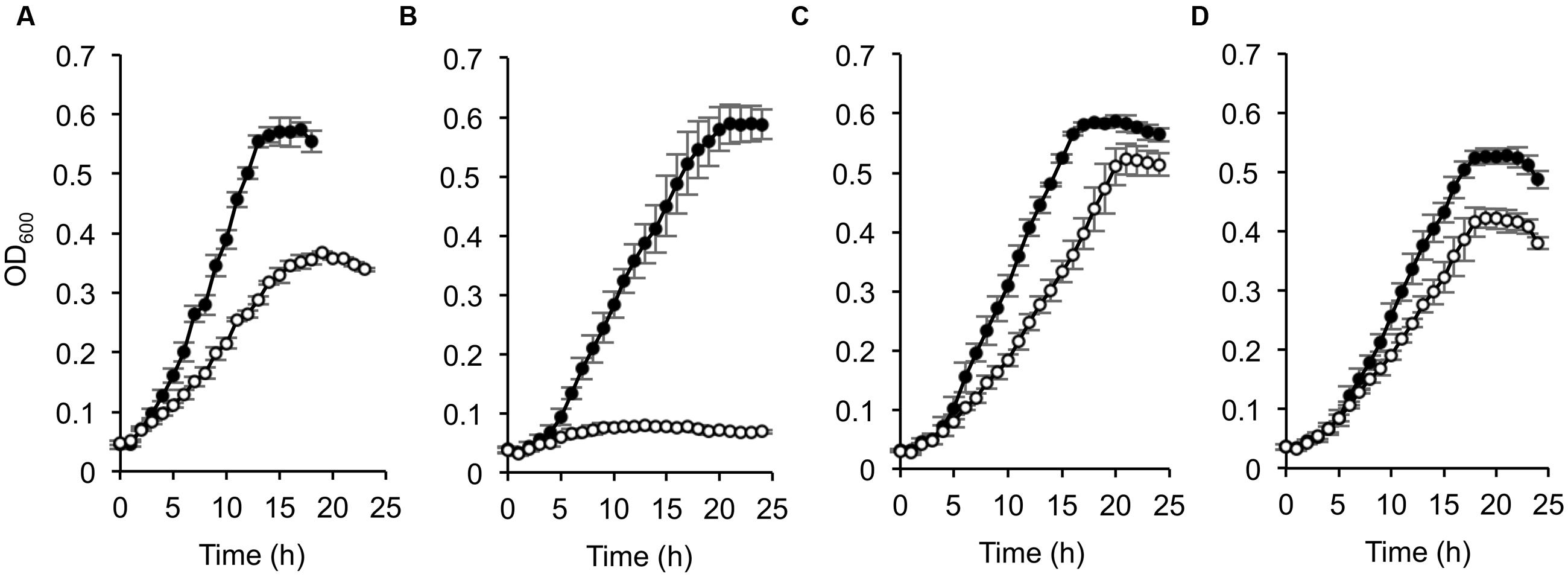
FIGURE 1. Growth of wild-type (WT) and Δcrp on DL-lactate (A), D-lactate (B), L-lactate (C), and pyruvate (D). WT (closed circle) and Δcrp (open circle) were aerobically grown in minimal medium (MM) containing 10 mM of each substrate. Error bars represent standard deviations calculated from at least triplicate independent cultures.
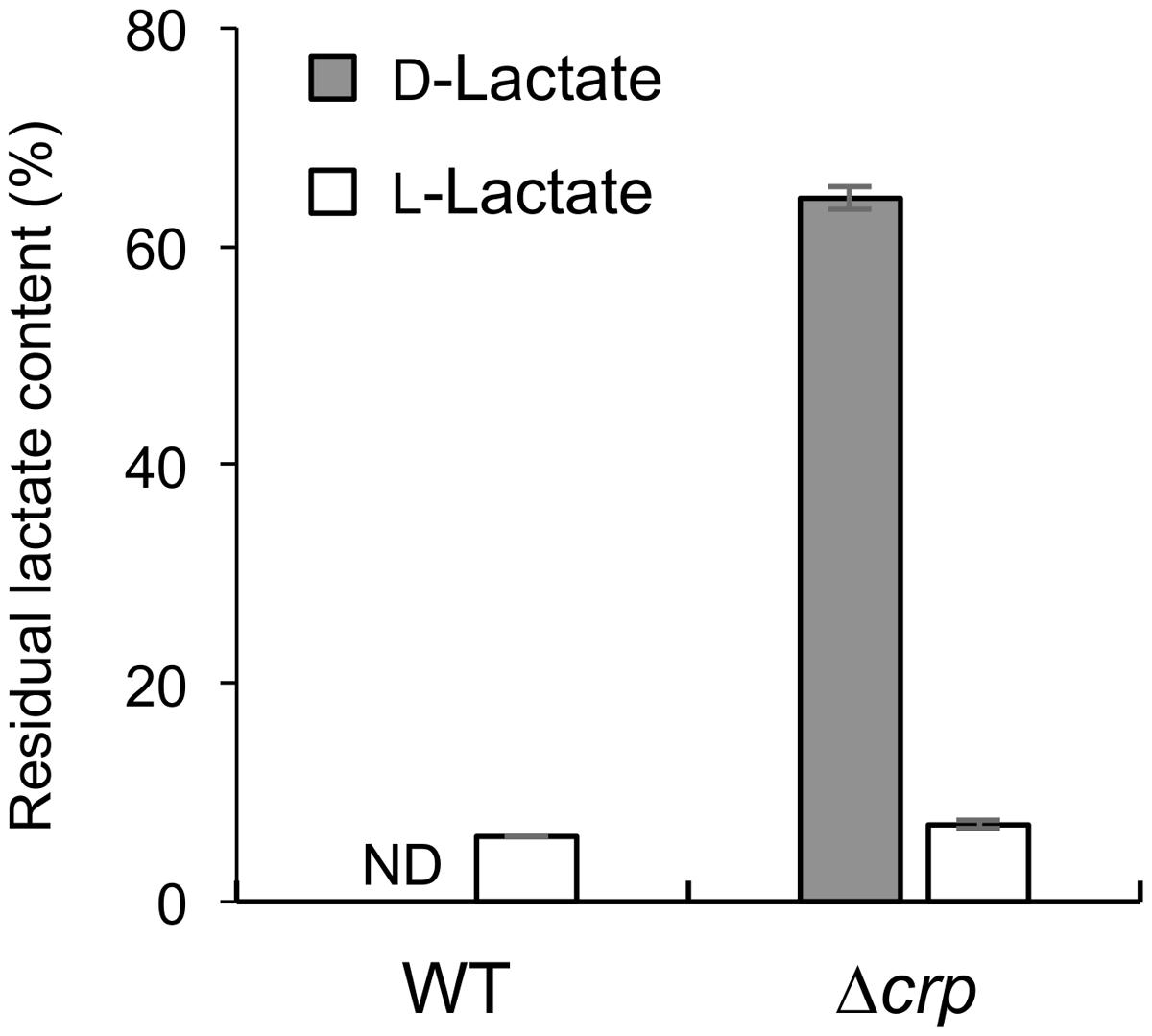
FIGURE 2. Residual D- and L-lactate contents in WT and Δcrp cultures. Error bars represent standard deviations calculated from at least triplicate independent experiments. ND, not detected.
Involvement of CRP in the Expression of dld
In the MR-1 genome, the dld gene is located within a gene cluster consisting of lldP (a putative lactate permease gene), dld, and lldEFG (SO_1522 to SO_1518; Figure 3A) (Pinchuk et al., 2009). Thus, to examine the involvement of CRP in the transcription of genes in this cluster, we first determined transcriptional units by RT-PCR and subsequently compared amounts of transcripts from these genes in WT and Δcrp by qRT-PCR. RT-PCR analysis using total RNA extracted from WT cells grown on DL-lactate detected a transcript containing the intergenic region between lldP and dld (Figure 3B, lane 2), indicating the polycistronic transcription of these two genes. The analysis also amplified the lldE–lldF and lldF–lldG intergenic regions (Figure 3B, lanes 5 and 6). However, no transcripts were detected when the dld–lldE intergenic region was analyzed (Figure 3B, lane 4), demonstrating that the transcription of dld is terminated within this region. These results indicate that the five genes in this gene cluster are transcribed as two operons, the lldP-dld and lldEFG operons.
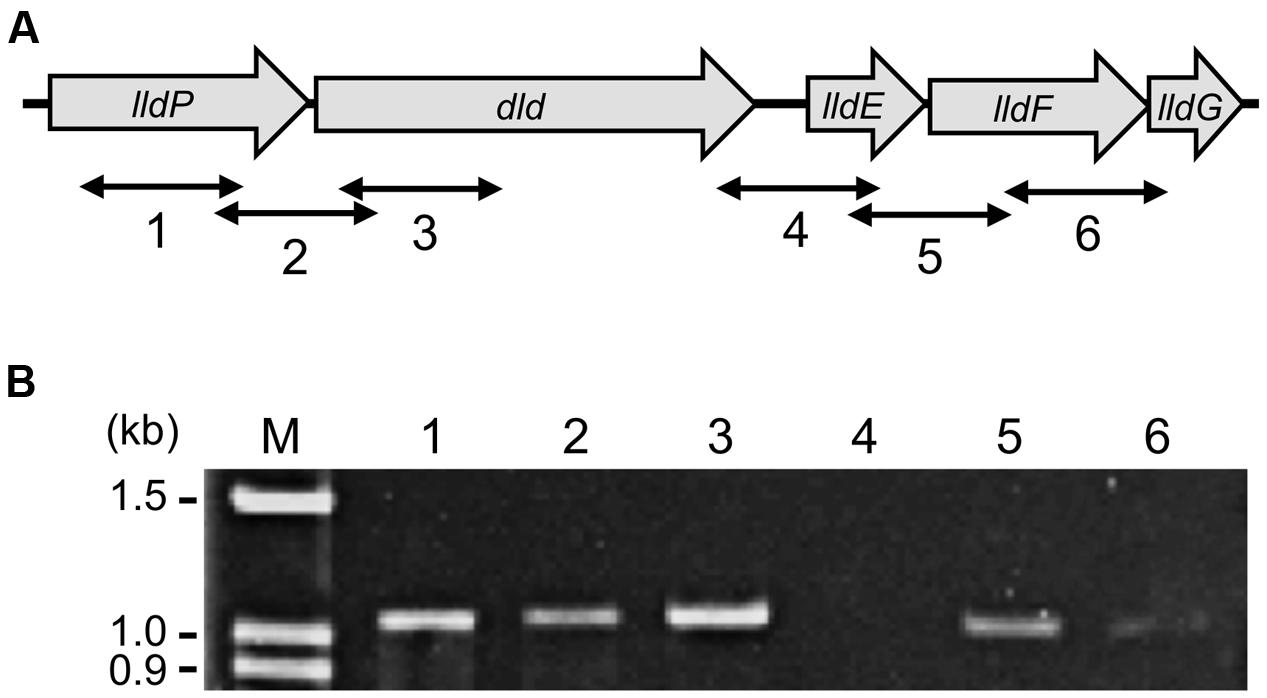
FIGURE 3. Organization and transcriptional units of the dld and lld genes. (A) Schematic illustration of the organization of the dld and lld genes. Bidirectional arrows indicate target regions of the RT-PCR analysis. (B) RT-PCR of the dld and lld genes. RNA was extracted from WT cells grown aerobically in MM containing 10 mM DL-lactate until the logarithmic growth phase. Lane numbers correspond to the target regions shown in A. Left-side numbers indicate molecular sizes (kb) of bands in the marker (lane M).
We performed qRT-PCR analyses to determine expression levels of lldP, dld, and lldF in WT and Δcrp (Figure 4). The results revealed that expression levels of lldP and dld in Δcrp were decreased to approximately 20% of those in WT, although the expression of lldF did not significantly differ between these two strains. This result indicates that CRP is required for the transcriptional activation of the lldP-dld operon, although this regulator is not involved in the transcription of the lldEFG operon. Decreased expression of lldE and dldP in Δcrp was also observed when cells were grown under TMAO-reducing conditions (Supplementary Figure S2), indicating that CRP activates the transcription of these genes under both aerobic and anaerobic conditions.
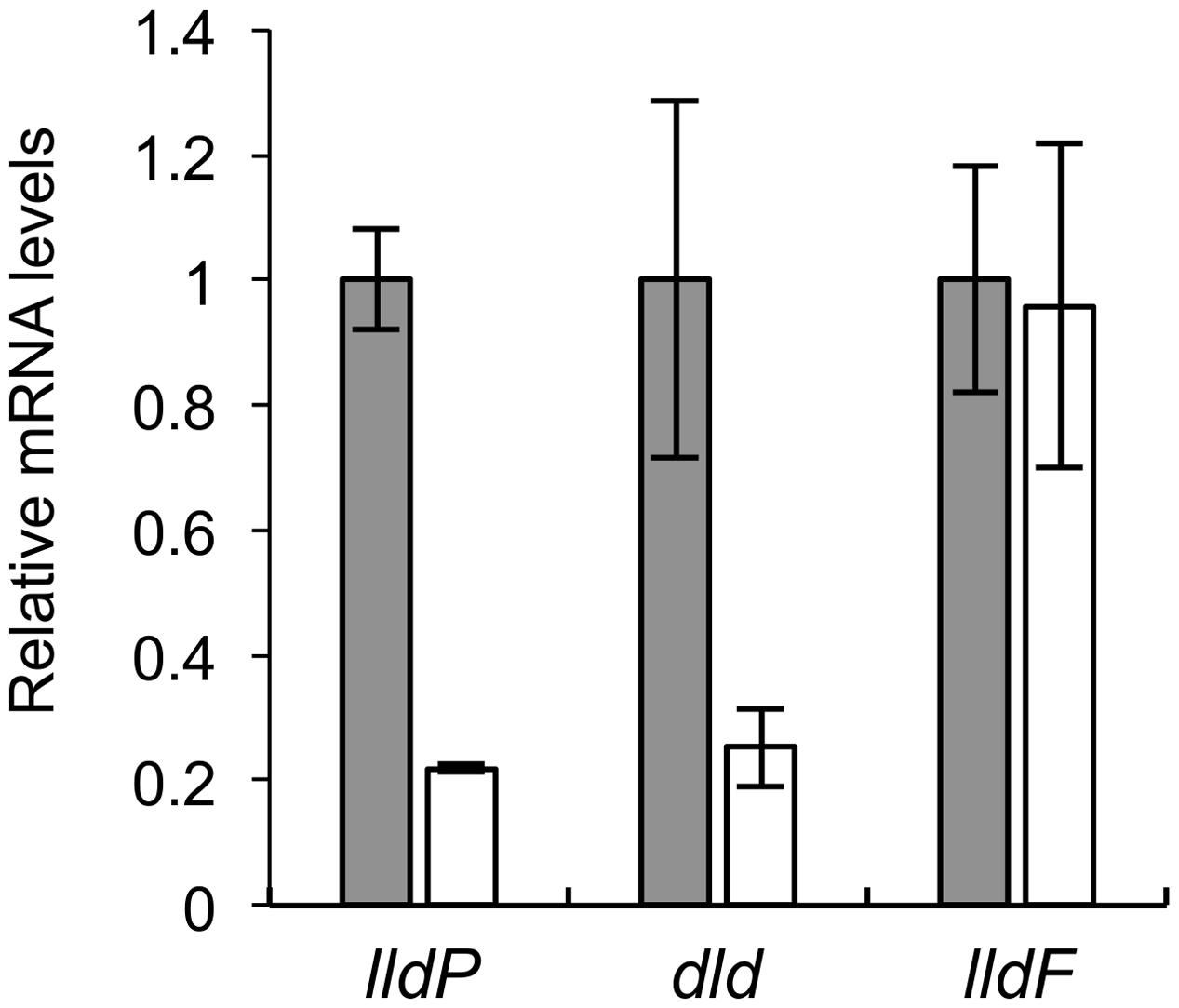
FIGURE 4. Quantitative RT-PCR (qRT-PCR) analyses of lldP, dld, and lldF in WT (gray bars) and Δcrp (white bars). Cells were aerobically grown in MM containing 10 mM DL-lactate and harvested in the logarithmic growth phase. Results are expressed as relative values to mRNA levels in the WT cells. Error bars represent standard deviations calculated from at least three independent experiments.
Restored D-Lactate Utilization by dld-Complemented Δcrp
The above-mentioned results suggest that the decreased expression of the lldP-dld operon in Δcrp is related to the deficient growth of this mutant on D-lactate. However, it remained possible that transcriptional activation of lldP and/or other CRP-regulated genes might also be required for D-lactate utilization. To test this possibility, we transformed Δcrp with a plasmid constitutively expressing the dld gene, pBBRdld (Table 1), and examined the growth of the resultant strain, Δcrp(pBBRdld), on D-lactate. When cells were grown under aerobic conditions (Figure 5), the growth of Δcrp(pBBRdld) on D-lactate was comparable to that of the control strain, MR-1(pBBR1MCS-2). The restored growth of Δcrp(pBBRdld) on D-lactate was also observed when cells were grown under TMAO-reducing conditions (Supplementary Figure S3), although its growth was slower than that of MR-1(pBBR1MCS-2). We consider that this growth retardation is due to the decreased ability of Δcrp to grow under anaerobic conditions because similar growth retardation was observed when Δcrp(pBBR1MCS-2) was cultivated using L-lactate and TMAO (Supplementary Figure S4). We therefore concluded that the deficient growth of Δcrp on D-lactate is attributable to the decreased expression of the dld gene.
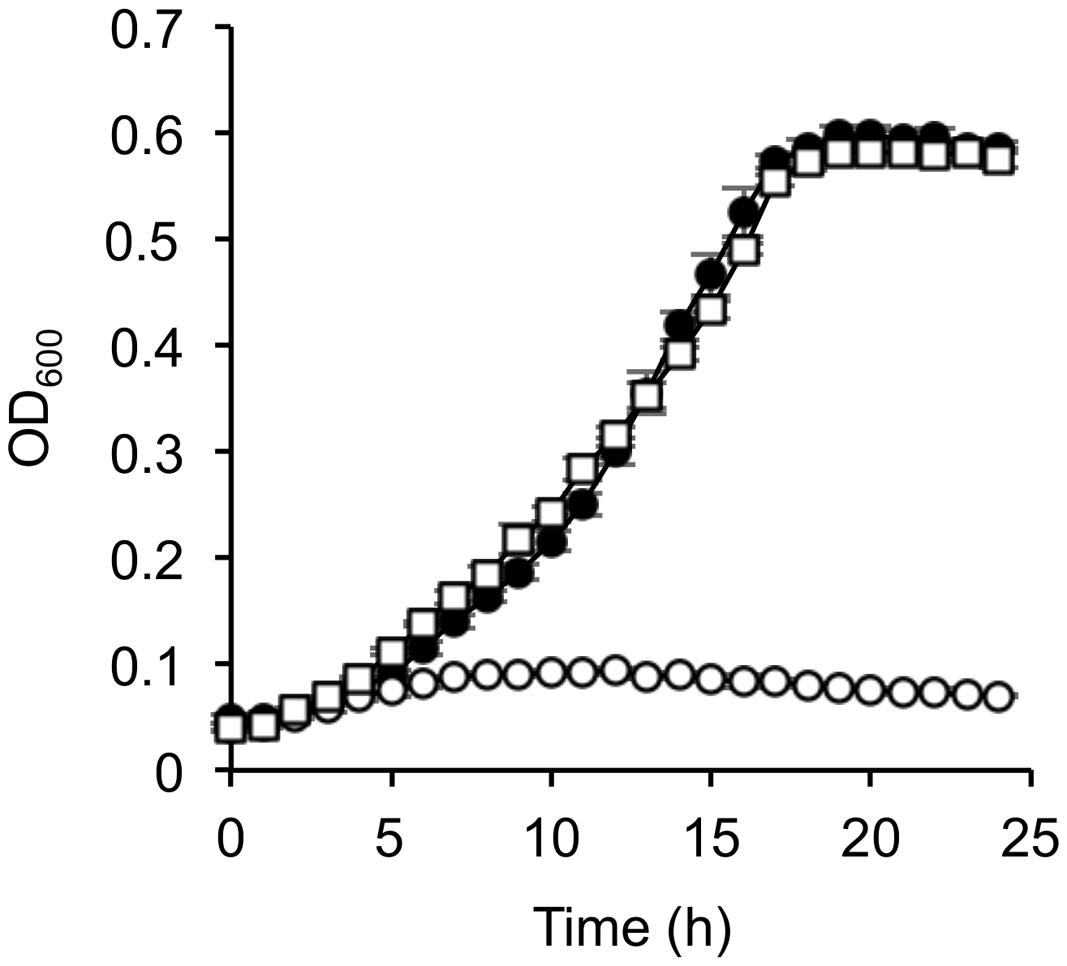
FIGURE 5. Growth of dld-complemented Δcrp on D-lactate. WT harboring the control vector pBBR1MCS-2 (closed circle), Δcrp harboring pBBR1MCS-2 (open circle), and Δcrp harboring pBBRdld (open squire) were aerobically grown in MM containing 10 mM D-lactate. Error bars represent standard deviations calculated from at least three independent experiments.
D-Lactate-Independent Expression of the lldP-dld Operon
A previous study reported that D-LDH activity in MR-1 was independent of growth substrates, whereas L-LDH activity was increased when it was grown on L-lactate (Brutinel and Gralnick, 2012). However, it was unclear how the presence of D- and L-lactate affects transcription of the lldP-dld and lldEFG operons. To address this, we cultivated MR-1 cells in the presence of D- or L-lactate, and investigated the expression levels of lldP and lldF by qRT-PCR analyses (Figure 6). We found that, when cells were grown on L-lactate, the expression of lldF was markedly increased, suggesting that the lldEFG operon is regulated by an L-lactate-dependent regulatory mechanism (i.e., the LlpR transcriptional regulator) (Brutinel and Gralnick, 2012). In contrast, the expression of lldP did not significantly differ between the two growth conditions. These results support the idea that the lldP-dld and lldEFG operons are regulated by different mechanisms, and that expression of the lldP-dld operon is not affected by the presence or absence of D-lactate.
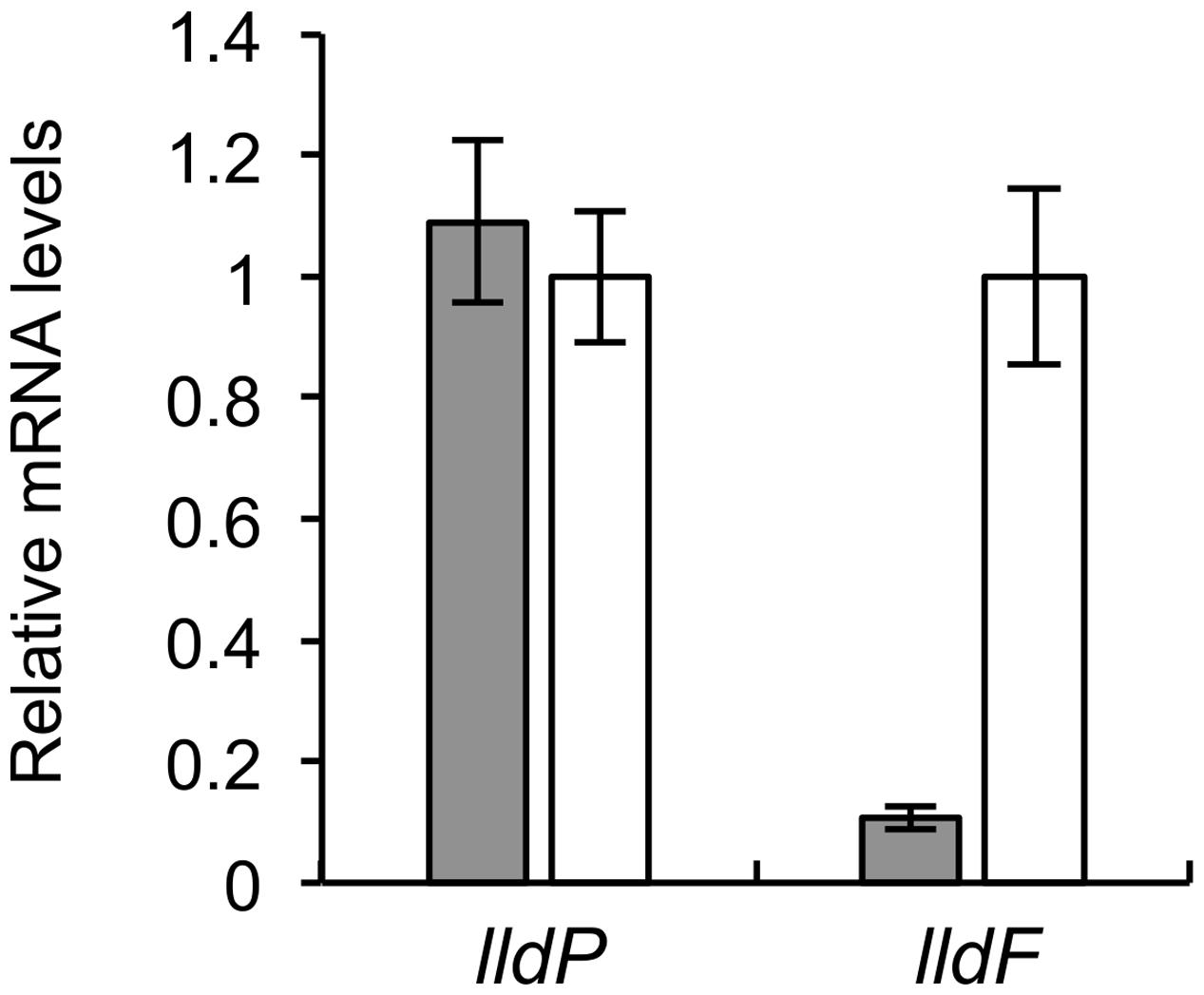
FIGURE 6. Quantitative RT-PCR analyses of lldP and lldF in WT grown on D-lactate (gray bars) or L-lactate (white bars). Cells were grown in MM containing 10 mM D- or L-lactate and harvested in the logarithmic growth phase. Data are expressed as relative values to mRNA levels in cells grown on L-lactate. Error bars represent standard deviations calculated from at least three independent experiments.
Identification of the lldP Promoter Region
To investigate the transcriptional mechanism of the lldP-dld operon in more detail, we determined a transcription start site (TSS) and upstream promoter region for the lldP-dld operon. 5′-RACE PCR analysis detected a TSS 192 bp upstream of the ATG start codon of lldP (TSSlldP; Figure 7). A sequence similar to the consensus sequence of the E. coli σ70-dependent promoter (–10 and –35 regions) (deHaseth et al., 1998) was found in the upstream region of TSSlldP (Figure 7). Although a previous study reported that the RpoE-binding motif is detected upstream of many anaerobic respiratory genes in MR-1 (Barchinger et al., 2016), a sequence similar to that motif was not found in the upstream of lldP. However, a candidate CRP-binding sequence (5′-TTAAGTGACACCGATCACAGTT-3′) predicted by Gao et al. (2010) was located in position –70 to –49 relative to TSSlldP. We therefore hypothesized that CRP binds to this sequence and activates the downstream lldP promoter (PlldP).
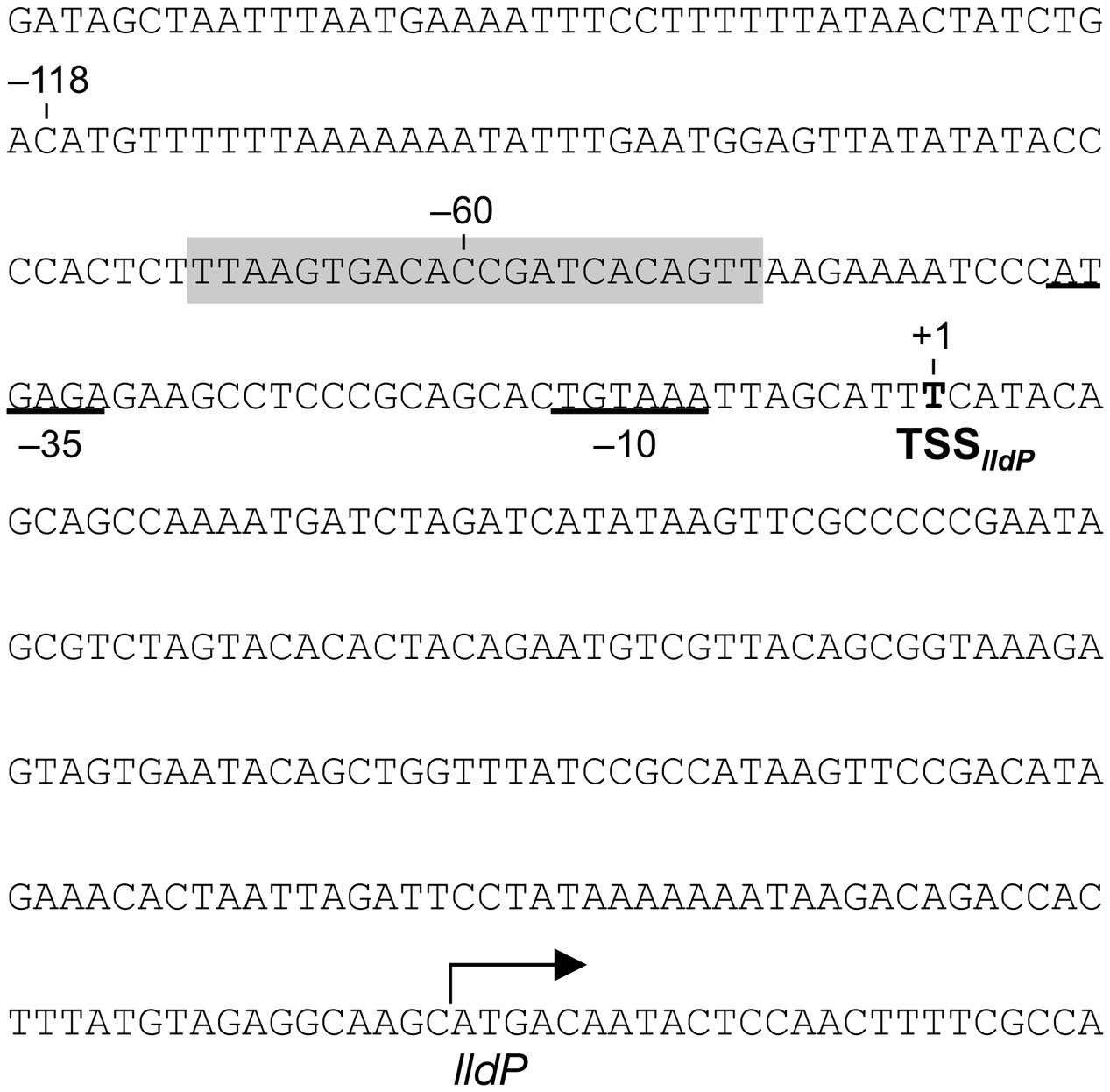
FIGURE 7. Nucleotide sequence upstream of lldP. The position of TSSlldP is indicated by boldface. Nucleotides are numbered relative to TSSlldP (+1). Putative –10 and –35 promoter sequences are underlined. A shaded sequence indicates a putative CRP-binding motif.
To determine the DNA regions involved in the activation of PlldP by CRP, we performed 5′-deletion analysis of the sequence upstream of TSSlldP. For this purpose, WT and Δcrp strains were transformed with the lacZ reporter plasmids with promoter region deletions (pMElldP-541 to pMElldP+1; Table 1), and resultant transformants were grown on DL-lactate until the logarithmic growth phase. Measurement of the LacZ activities of these strains (Figure 8) revealed that, while Δcrp cells transformed with these reporter plasmids consistently exhibited very low levels of LacZ activities, WT cells transformed with pMElldP-541 and pMElld-182 exhibited much higher LacZ activities than these Δcrp cells. This result supports the idea that CRP plays a critical role in the activation of PlldP. The analysis also revealed that LacZ activity in WT cells was decreased to the same level as that in Δcrp cells when they were transformed with pMElld-60. This result demonstrates that the activation of PlldP by CRP requires the DNA region from –182 to –60 relative to TSSlldP, which partially contains the putative CRP-binding site (Figure 7).
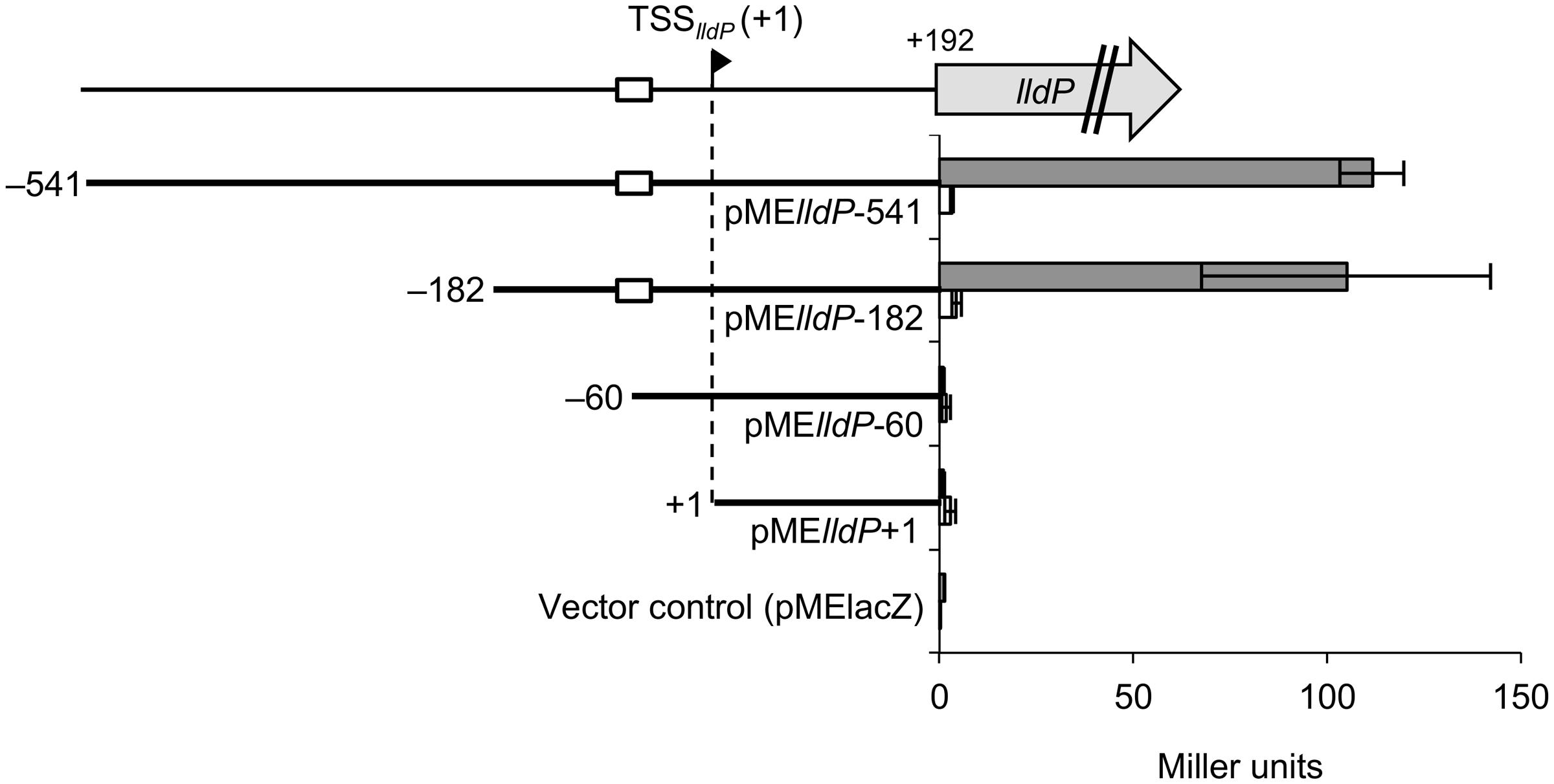
FIGURE 8. 5′-deletion analyses of the upstream region of lldP. WT (gray bars) or Δcrp (white bars) cells harboring the pME-series plasmids were grown on DL-lactate, and their LacZ activities were measured in the logarithmic growth phase. White boxes represent the putative CRP-binding motif. Error bars show standard deviations calculated from at least three independent experiments.
Binding of CRP to the lldP Promoter Region
To investigate whether CRP binds directly to the upstream activation region of PlldP, we performed an EMSA using purified CRP protein (Figure 9A) and DNA probes containing the upstream regions of PlldP (Figure 9B). When a DNA probe containing the region from –182 to –61 relative to TSSlldP (PBlldP1) was mixed with CRP, no shifted bands were observed (Figure 9C). However, when a probe containing the region from –182 to –1 relative to TSSlldP (PBlldP2) was used, a shifted band was detected in a cAMP-dependent manner (Figure 9C), indicating that CRP directly binds to this region. Together with the results of the 5′-deletion analysis, these results suggested that CRP binds to a region around position –60, which is consistent with the position of the putative CRP-binding motif (Figure 7). To examine the binding of CRP to this motif, we performed an EMSA using a 50-bp probe containing the region from –84 to –35 (PBlldP3) and a mutated probe in which the core motif sequence was replaced (PBlldP3m). The results revealed that CRP specifically bound to PBlldP3, but not to PBlld3m (Figure 9C). We therefore conclude that this motif is involved in the binding of CRP and the transcriptional activation of the lldP-dld operon.
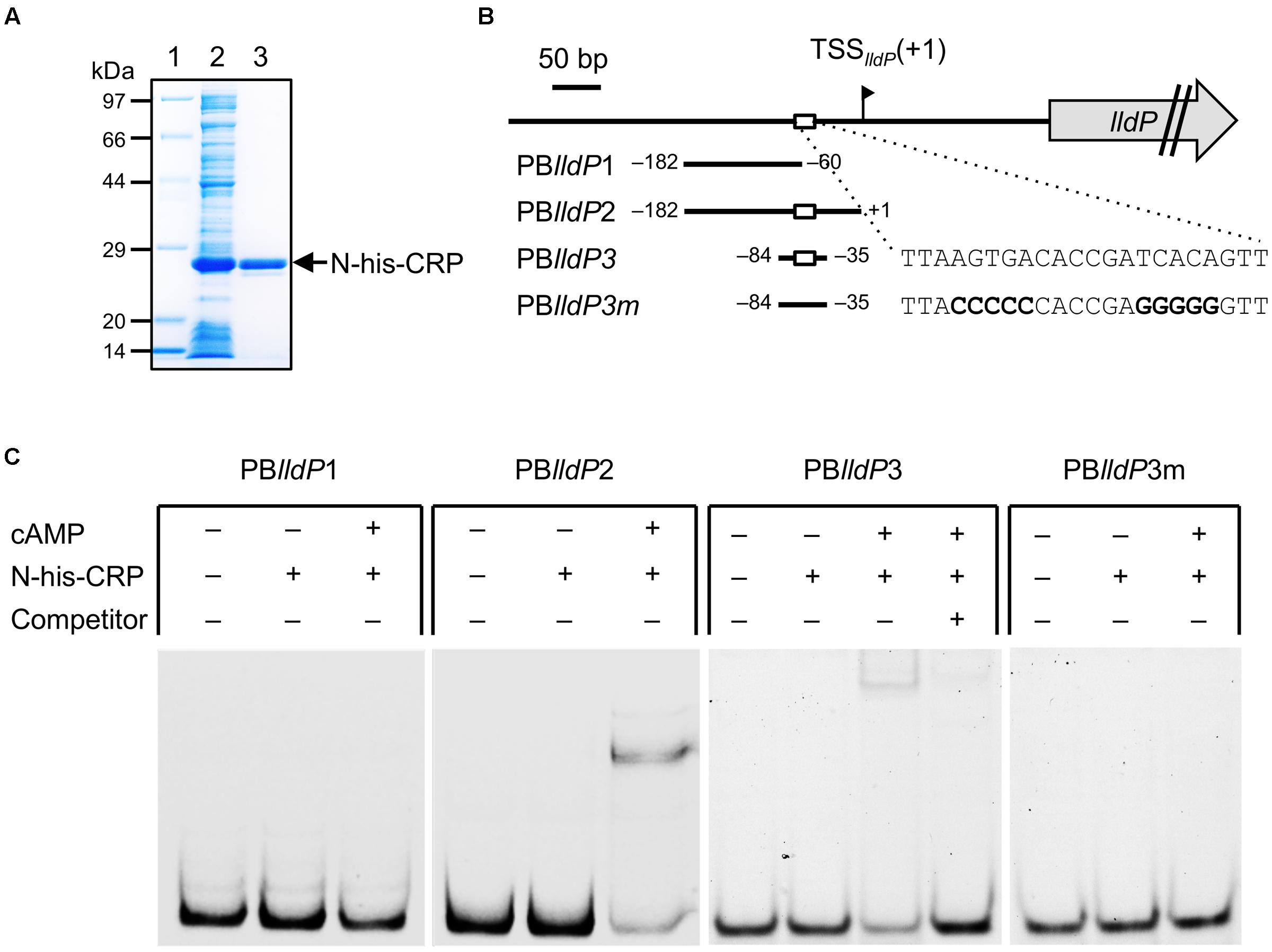
FIGURE 9. Electrophoretic mobility shift assay (EMSA) using CRP and upstream regions of lldP. (A) Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS–PAGE) of N-his-CRP protein samples. Protein samples (5 μg) were analyzed on 12.5% SDS-polyacrylamide gels. Lane 1, Escherichia coli BL21 (DE3) (pET-crp) crude extract; lane 2, purified N-his-CRP; lane 3, molecular weight marker. (B) DNA fragments used as probes. Positions of 5′ and 3′ ends of the fragments relative to TSSlldP (+1) are shown. White boxes represent the putative CRP-binding motif. The mutated sequences in PBlldP3m are shown in bold. (C) Binding of CRP to each probe. DNA-binding reactions were performed in the presence (+) or absence (–) of CRP, cAMP, and specific competitor (2 μM unlabeled PBlldP3 probe).
Discussion
Like many other anaerobic respiratory bacteria, Shewanella spp. preferentially catabolize low-molecular-weight organic acids (Serres and Riley, 2006; Fredrickson et al., 2008), and studies have used lactate (mostly racemic DL-lactate) as a substrate for cultivating S. oneidensis MR-1. Despite the importance of lactate as a catabolic substrate for Shewanella spp., however, little is known about how these bacteria regulate catabolic pathways for this compound. Here, we demonstrated that CRP plays a critical role in the transcriptional regulation of the dld gene encoding D-LDH in MR-1. As CRP is also involved in the expression of many anaerobic respiratory genes (Saffarini et al., 2003; Kouzuma et al., 2015), we suggest that MR-1 uses this transcriptional regulator to coordinately regulate D-lactate metabolism and anaerobic respiration. A previous study reported that, in Corynebacterium glutamicum, a CRP/Fnr-type global transcriptional regulator, GlxR, binds to promoter regions of the ldhA gene encoding a fermentative L-LDH and the narKGHJI operon encoding nitrate respiratory enzymes in a cAMP-dependent manner (Kohl et al., 2008). It is therefore likely that a broad range of bacteria utilizes cAMP-dependent regulatory mechanisms for the coordinated expression of catabolic and respiratory pathways.
In many bacteria, lactate is oxidized to pyruvate by NADH-independent LDHs (iLDHs) (Garvie, 1980; Jiang et al., 2014). For example, in E. coli and C. glutamicum, D-lactate is catabolized through membrane-bound D-iLDHs that utilize membrane-associated quinones as electron-accepting cofactors (Dym et al., 2000; Kato et al., 2010). S. oneidensis MR-1 also oxidizes D-lactate by D-iLDH (Dld) (Pinchuk et al., 2009), and is thereby likely to reduce membrane quinones. In this strain, electrons accumulated in the membrane quinone pool are transferred to anaerobic electron acceptors, such as fumarate, DMSO, and metal oxides, via an inner membrane-anchored cytochrome, CymA, and periplasm- and outer membrane-localized proteins, such as FccA, DmsABEF, MtrCAB, and OmcA (Beliaev et al., 2001; Myers and Myers, 2002; Gralnick et al., 2006; Shi et al., 2006; Hartshorne et al., 2009; Schuetz et al., 2009; Fonseca et al., 2012). It is therefore suggested that D-lactate oxidation in MR-1 is metabolically linked to anaerobic respiratory pathways via membrane-associated quinones.
D- and L-Lactate are major fermentation end products from glucose and other carbohydrates. Many fermentative microbes, including lactic acid bacteria, produce these organic acids and release them into the environment (Garvie, 1980). It is therefore possible that Shewanella spp. utilize these fermentation products as catabolic substrates in their habitats. Here, we demonstrate that the catabolic pathways of D- and L-lactate in MR-1 are regulated by different mechanisms. Expression analysis in the presence of D- or L-lactate (Figure 6) revealed that expression of the lldEFG operon (encoding L-LDH) is induced by L-lactate, while expression of the lldP-dld operon (encoding D-LDH) is independent of the presence of D- or L-lactate. Considering that the lldP-dld operon is regulated by CRP, this operon is likely regulated in an intracellular energy status-dependent manner. It is therefore interesting to consider why only D-lactate catabolism is regulated in a substrate-independent manner. We speculate that this difference reflects how these lactate isomers are supplied to MR-1 in its habitats. The substrate-dependent regulation of the L-LDH genes suggests that the supply of L-lactate may be a relatively rare event in such environments, and thus MR-1 expresses these genes only when L-lactate is available. In contrast, the CRP-dependent regulation of the dld gene implies that MR-1 is frequently exposed to D-lactate in its habitats (mostly anaerobic environments). This notion is likely, given that MR-1 itself produces D-lactate as an electron sink when it catabolizes carbohydrates under electron acceptor-limited conditions. In addition to the respiratory D- and L-iLDH genes, MR-1 possesses the ldhA gene, which encodes a putative NADH-dependent D-LDH that catalyzes the fermentative production of D-lactate from pyruvate (Heidelberg et al., 2002), suggesting that this strain has the ability to produce D-lactate under electron acceptor-limited conditions. It is likely that MR-1 produces D-lactate from carbohydrates by LdhA as a temporal electron sink when electron acceptors are limited, and subsequently catabolizes D-lactate using Dld and anaerobic respiratory chains. Supporting this speculation, we recently found that an engineered S. oneidensis strain capable of utilizing glucose produces D-lactate from this sugar when grown under electron acceptor (fumarate)-limited conditions (Nakagawa et al., 2015). Gene-knockout experiments also demonstrate that LdhA and Dld catalyze the production and degradation of D-lactate, respectively, under these conditions (Nakagawa et al., 2015). The wild-type MR-1 cannot utilize glucose but has the ability to utilize other carbohydrates, such as N-acetylglucosamine (Yang et al., 2006; Rodionov et al., 2010). We therefore speculate that these carbohydrates are important catabolic substrates for Shewanella spp., and that D-lactate metabolism plays a role in maintaining the intracellular redox balance. Interestingly, a previous study also reported that a putative CRP-binding site is present upstream of ldhA, and that expression of the ldhA gene is up-regulated under oxygen-limited conditions, as is found for dld and anaerobic respiratory genes (Barchinger et al., 2016). We therefore hypothesize that the cAMP/CRP system contributes more widely to the regulation of catabolic pathways in Shewanella than previously thought.
The present study also demonstrates that the dld gene is co-transcribed with the lldP gene encoding a putative lactate permease (Figures 3A,B). In E. coli, the L-lactate permease LldP and glycolate permease GlcA are both involved in the uptake of D-lactate, although no specific D-lactate permeases are found (Núñez et al., 2002). LldP in MR-1 shows low but significant homologies to LldP and GlcA in E. coli (22.8 and 24.8% identities, respectively, in amino acid sequences), suggesting the involvement of this protein in the uptake of D- and/or L-lactate. In addition to lldP, the MR-1 genome encodes another putative lactate permease, LctP (SO_0827), which shows high homologies to LldP and GlcA in E. coli (66.0 and 65.7% identities, respectively). It is therefore likely that MR-1 takes up D- and/or L-lactate using one or both of these permeases. Notably, a putative CRP-binding site is also found in the upstream region of the lctP gene (Barchinger et al., 2016), suggesting that expression of the lldP and lctP genes is controlled by a common transcriptional regulator. It would therefore be interesting to investigate the presence of functional differences between lldP and lctP. In this regard, we showed that the growth deficiency of Δcrp on D-lactate was restored by introducing the plasmid from which only Dld was expressed (Figure 5), indicating that transcriptional activation of lldP by CRP is not essential for growth on D-lactate. This result suggests two possibilities for D-lactate uptake in MR-1: (i) LldP expressed at a low level is sufficient for this strain to grow on D-lactate, and (ii) uptake is mainly due to the alternative D-lactate permease, LctP. Confirming these hypotheses requires additional investigation, such as knockout analyses of the permease genes.
While the present study clearly demonstrates that CRP is involved in transcriptional regulation of the lldP-dld operon, further studies are needed to fully understand the regulatory systems for this operon. For instance, although the intracellular cAMP concentration is a key determinant for the expression of CRP regulons, including the lldP-dld operon, it is currently unclear how the concentration of this signaling molecule is controlled in Shewanella. Since previous studies have shown that many CRP-regulated genes are up-regulated under electron acceptor-limited conditions (Wang et al., 2012; Kasai et al., 2015; Barchinger et al., 2016), cAMP concentration may be affected by cellular redox and/or energy states. In addition, it has been previously found that expression of the dld gene is up-regulated in the presence of a high-potential electrode in a bioelectrochemical reactor, while that of the ldhA gene is not affected by the electrode potential (Nakagawa et al., 2015). This result implies that an additional regulatory factor(s) is involved in the expression of the lldP-dld operon and/or the ldhA gene. We expect that further investigation of this operon will reveal as yet unexplored regulatory mechanisms in bacteria.
Author Contributions
TK carried out the majority of the experimental work and drafted the manuscript. AK conceived of the study, participated in its design and coordination, and drafted the manuscript. KW supervised the study and performed manuscript editing. All authors read and approved the final manuscript.
Funding
This work was supported by a Grant-in-Aid for JSPS Fellows (to TK; Grant Number 16J08653) and Grant-in-Aid for Young Scientists (B) (to AK; Grant Number 26850056).
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2017.00869/full#supplementary-material
References
Barchinger, S. E., Pirbadian, S., Sambles, C., Baker, C. S., Leung, K. M., Burroughs, N. J., et al. (2016). Regulation of gene expression in Shewanella oneidensis MR-1 during electron acceptor limitation and bacterial nanowire formation. Appl. Environ. Microbiol. 82, 5428–5443. doi: 10.1128/AEM.01615-16
Beliaev, A. S., Saffarini, D. A., McLaughlin, J. L., and Hunnicutt, D. (2001). MtrC, an outer membrane decahaem c cytochrome required for metal reduction in Shewanella putrefaciens MR-1. Mol. Microbiol. 39, 722–730. doi: 10.1046/j.1365-2958.2001.02257.x
Brutinel, E. D., and Gralnick, J. A. (2012). Preferential utilization of D-lactate by Shewanella oneidensis. Appl. Environ. Microbiol. 78, 8474–8476. doi: 10.1128/AEM.02183-12
Charania, M. A., Brockman, K. L., Zhang, Y., Banerjee, A., Pinchuk, G. E., Fredrickson, J. K., et al. (2009). Involvement of a membrane-bound class III adenylate cyclase in regulation of anaerobic respiration in Shewanella oneidensis MR-1. J. Bacteriol. 191, 4298–4306. doi: 10.1128/JB.01829-08
Choi, K. H., Kumar, A., and Schweizer, H. P. (2006). A 10-min method for preparation of highly electrocompetent Pseudomonas aeruginosa cells: application for DNA fragment transfer between chromosomes and plasmid transformation. J. Microbiol. Methods 64, 391–397. doi: 10.1016/j.mimet.2005.06.001
deHaseth, P. L., Zupancic, M. L., and Record, M. T. (1998). RNA polymerase-promoter interactions: the comings and goings of RNA polymerase. J. Bacteriol. 180, 3019–3025.
Dong, Y., Wang, J., Fu, H., Zhou, G., Shi, M., and Gao, H. (2012). A Crp-dependent two-component system regulates nitrate and nitrite respiration in Shewanella oneidensis. PLoS ONE 7:e51643. doi: 10.1371/journal.pone.0051643
Dym, O., Pratt, E. A., Ho, C., and Eisenberg, D. (2000). The crystal structure of D-lactate dehydrogenase, a peripheral membrane respiratory enzyme. Proc. Natl. Acad. Sci. U.S.A. 97, 9413–9418. doi: 10.1073/pnas.97.17.9413
Endoh, T., Habe, H., Yoshida, T., Nojiri, H., and Omori, T. (2003). A CysB-regulated and σ54-dependent regulator, SfnR, is essential for dimethyl sulfone metabolism of Pseudomonas putida strain DS1. Microbiology 149,991–1000. doi: 10.1099/mic.0.26031-0
Fonseca, B., Paquete, C., Neto, S., Pacheco, I., Soares, C., and Louro, R. (2012). Mind the gap: cytochrome interactions reveal electron pathways across the periplasm of Shewanella oneidensis MR-1. Biochem. J. 108, 101–108. doi: 10.1042/BJ20121467
Fredrickson, J. K., Romine, M. F., Beliaev, A. S., Auchtung, J. M., Driscoll, M. E., Gardner, T. S., et al. (2008). Towards environmental systems biology of Shewanella. Nat. Rev. Microbiol. 6, 592–603. doi: 10.1038/nrmicro1947
Fu, H., Chen, H., Wang, J., Zhou, G., Zhang, H., Zhang, L., et al. (2013). Crp-dependent cytochrome bd oxidase confers nitrite resistance to Shewanella oneidensis. Environ. Microbiol. 15, 2198–2212. doi: 10.1111/1462-2920.12091
Gao, H., Wang, X., Yang, Z. K., Chen, J., Liang, Y., Chen, H., et al. (2010). Physiological roles of ArcA, Crp, and EtrA and their interactive control on aerobic and anaerobic respiration in Shewanella oneidensis. PLoS ONE 5:e15295. doi: 10.1371/journal.pone.0015295
Gao, T., Ju, L., Yin, J., and Gao, H. (2015). Positive regulation of the Shewanella oneidensis OmpS38, a major porin facilitating anaerobic respiration, by Crp and Fur. Sci. Rep. 5:14263. doi: 10.1038/srep14263
Gralnick, J. A., Vali, H., Lies, D. P., and Newman, D. K. (2006). Extracellular respiration of dimethyl sulfoxide by Shewanella oneidensis strain MR-1. Proc. Natl. Acad. Sci. U.S.A. 103, 4669–4674. doi: 10.1073/pnas.0505959103
Hartshorne, R. S., Reardon, C. L., Ross, D., Nuester, J., Clarke, T. A., Gates, A. J., et al. (2009). Characterization of an electron conduit between bacteria and the extracellular environment. Proc. Natl. Acad. Sci. U.S.A. 106, 22169–22174. doi: 10.1073/pnas.0900086106
Hau, H. H., and Gralnick, J. A. (2007). Ecology and biotechnology of the genus Shewanella. Annu. Rev. Microbiol. 61, 237–258. doi: 10.1146/annurev.micro.61.080706.093257
Heidelberg, J. F., Paulsen, I. T., Nelson, K. E., Gaidos, E. J., Nelson, W. C., Read, T. D., et al. (2002). Genome sequence of the dissimilatory metal ion-reducing bacterium Shewanella oneidensis. Nat. Biotechnol. 20, 1118–1123. doi: 10.1038/nbt749
Jiang, T., Gao, C., Ma, C., and Xu, P. (2014). Microbial lactate utilization: enzymes, pathogenesis, and regulation. Trends Microbiol. 22, 589–599. doi: 10.1016/j.tim.2014.05.008
Jin, M., Fu, H., Yin, J., Yuan, J., and Gao, H. (2016). Molecular underpinnings of nitrite effect on CymA-dependent respiration in Shewanella oneidensis. Front. Microbiol. 7:1154. doi: 10.3389/fmicb.2016.01154
Kasai, T., Kouzuma, A., Nojiri, H., and Watanabe, K. (2015). Transcriptional mechanisms for differential expression of outer membrane cytochrome genes omcA and mtrC in Shewanella oneidensis MR-1. BMC Microbiol. 15:68. doi: 10.1186/s12866-015-0406-8
Kato, O., Youn, J. W., Stansen, K. C., Matsui, D., Oikawa, T., and Wendisch, V. F. (2010). Quinone-dependent D-lactate dehydrogenase Dld (Cg1027) is essential for growth of Corynebacterium glutamicum on D-lactate. BMC Microbiol. 10:321. doi: 10.1186/1471-2180-10-321
Kohl, T. A., Baumbach, J., Jungwirth, B., Pühler, A., and Tauch, A. (2008). The GlxR regulon of the amino acid producer Corynebacterium glutamicum: in silico and in vitro detection of DNA binding sites of a global transcription regulator. J. Biotechnol. 135, 340–350. doi: 10.1016/j.jbiotec.2008.05.011
Kolb, A., Busby, S., Buc, H., Garges, S., and Adhya, S. (1993). Transcriptional regulation by cAMP and its receptor protein. Annu. Rev. Biochem. 62, 749–795. doi: 10.1146/annurev.bi.62.070193.003533
Kouzuma, A., Kasai, T., Hirose, A., and Watanabe, K. (2015). Catabolic and regulatory systems in Shewanella oneidensis MR-1 involved in electricity generation in microbial fuel cells. Front. Microbiol. 6:609. doi: 10.3389/fmicb.2015.00609
Kouzuma, A., Meng, X. Y., Kimura, N., Hashimoto, K., and Watanabe, K. (2010). Disruption of the putative cell surface polysaccharide biosynthesis gene SO3177 in Shewanella oneidensis MR-1 enhances adhesion to electrodes and current generation in microbial fuel cells. Appl. Environ. Microbiol. 76, 4151–4157. doi: 10.1128/AEM.00117-10
Kouzuma, A., Oba, H., Tajima, N., Hashimoto, K., and Watanabe, K. (2014). Electrochemical selection and characterization of a high current-generating Shewanella oneidensis mutant with altered cell-surface morphology and biofilm-related gene expression. BMC Microbiol. 14:190. doi: 10.1186/1471-2180-14-190
Kovach, M. E., Elzer, P. H., Hill, D. S., Robertson, G. T., Farris, M. A., Roop, R. M., et al. (1995). Four new derivatives of the broad-host-range cloning vector pBBR1MCS, carrying different antibiotic-resistance cassettes. Gene 166, 175–176. doi: 10.1016/0378-1119(95)00584-1
Miller, J. H. (1972). Experiments in Molecular Genetics. New York, NY: Cold Spring Harbor Laboratory Press.
Murphy, J. N., Durbin, K. J., and Saltikov, C. W. (2009). Functional roles of arcA, etrA, cyclic AMP (cAMP)-cAMP receptor protein, and cya in the arsenate respiration pathway in Shewanella sp. strain ANA-3. J. Bacteriol. 191,1035–1043. doi: 10.1128/JB.01293-08
Myers, C. R., and Myers, J. M. (2002). MtrB is required for proper incorporation of the cytochromes OmcA and OmcB into the outer membrane of Shewanella putrefaciens MR-1. Appl. Environ. Microbiol. 68, 5585–5594. doi: 10.1128/AEM.68.11.5585
Myers, C. R., and Nealson, K. H. (1988). Bacterial manganese reduction and growth with manganese oxide as the sole electron acceptor. Science 240, 1319–1321. doi: 10.1126/science.240.4857.1319
Nakagawa, G., Kouzuma, A., Hirose, A., Kasai, T., Yoshida, G., and Watanabe, K. (2015). Metabolic characteristics of a glucose-utilizing Shewanella oneidensis strain grown under electrode-respiring conditions. PLoS ONE 10:e0138813. doi: 10.1371/journal.pone.0138813
Núñez, M. F., Kwon, O., Wilson, T. H., Aguilar, J., Baldoma, L., and Lin, E. C. C. (2002). Transport of L-Lactate, D-Lactate, and glycolate by the LldP and GlcA membrane carriers of Escherichia coli. Biochem. Biophys. Res. Commun. 290, 824–829. doi: 10.1006/bbrc.2001.6255
Pinchuk, G. E., Rodionov, D. A., Yang, C., Li, X., Osterman, A. L., Dervyn, E., et al. (2009). Genomic reconstruction of Shewanella oneidensis MR-1 metabolism reveals a previously uncharacterized machinery for lactate utilization. Proc. Natl. Acad. Sci. U.S.A. 106, 2874–2879. doi: 10.1073/pnas.0806798106
Pirbadian, S., Barchinger, S. E., Leung, K. M., Byun, H. S., Jangir, Y., Bouhenni, R. A., et al. (2014). Shewanella oneidensis MR-1 nanowires are outer membrane and periplasmic extensions of the extracellular electron transport components. Proc. Natl. Acad. Sci. U.S.A. 111, 12883–12888. doi: 10.1073/pnas.1410551111
Rodionov, D. A., Novichkov, P. S., Stavrovskaya, E. D., Rodionova, I. A., Li, X., Kazanov, M. D., et al. (2011). Comparative genomic reconstruction of transcriptional networks controlling central metabolism in the Shewanella genus. BMC Genomics 12(Suppl. 1):S3. doi: 10.1186/1471-2164-12-S1-S3
Rodionov, D. A., Yang, C., Li, X., Rodionova, I. A., Wang, Y., Obraztsova, A. Y., et al. (2010). Genomic encyclopedia of sugar utilization pathways in the Shewanella genus. BMC Genomics 11:494. doi: 10.1186/1471-2164-11-494
Saffarini, D. A., Schultz, R., and Beliaev, A. (2003). Involvement of cyclic AMP (cAMP) and cAMP receptor protein in anaerobic respiration of Shewanella oneidensis. J. Bacteriol. 185, 3668–3671. doi: 10.1128/JB.185.12.3668-3671.2003
Schuetz, B., Schicklberger, M., Kuermann, J., Spormann, A. M., and Gescher, J. (2009). Periplasmic electron transfer via the c-type cytochromes MtrA and FccA of Shewanella oneidensis MR-1. Appl. Environ. Microbiol. 75, 7789–7796. doi: 10.1128/AEM.01834-09
Scott, J. H., and Nealson, K. H. (1994). A biochemical study of the intermediary carbon metabolism of Shewanella putrefaciens. J. Bacteriol. 176, 3408–3411. doi: 10.1128/jb.176.11.3408-3411.1994
Serres, M. H., and Riley, M. (2006). Genomic analysis of carbon source metabolism of Shewanella oneidensis MR-1: predictions versus experiments. J. Bacteriol. 188, 4601–4609. doi: 10.1128/JB.01787-05
Shi, L., Chen, B., Wang, Z., Elias, D. A., Mayer, M. U., Gorby, Y. A., et al. (2006). Isolation of a high-affinity functional protein complex between OmcA and MtrC: two outer membrane decaheme c-type cytochromes of Shewanella oneidensis MR-1. J. Bacteriol. 188, 4705–4714. doi: 10.1128/JB.01966-05
Sturm, G., Richter, K., Doetsch, A., Heide, H., Louro, R. O., and Gescher, J. (2015). A dynamic periplasmic electron transfer network enables respiratory flexibility beyond a thermodynamic regulatory regime. ISME J. 9, 1802–1811. doi: 10.1038/ismej.2014.264
Venkateswaran, K., Moser, D. P., Dollhopf, M. E., Lies, D. P., Saffarini, D. A., MacGregor, B. J., et al. (1999). Polyphasic taxonomy of the genus Shewanella and description of Shewanella oneidensis sp. nov. Int. J. Syst. Bacteriol. 49, 705–724. doi: 10.1099/00207713-49-2-705
Wang, H., Correa, E., Dunn, W. B., Winder, C. L., Goodacre, R., and Lloyd, J. R. (2012). Metabolomic analyses show that electron donor and acceptor ratios control anaerobic electron transfer pathways in Shewanella oneidensis. Metabolomics 9, 642–656. doi: 10.1007/s11306-012-0488-3
Yang, C., Rodionov, D. A., Li, X., Laikova, O. N., Gelfand, M. S., Zagnitko, O. P., et al. (2006). Comparative genomics and experimental characterization of N-acetylglucosamine utilization pathway of Shewanella oneidensis. J. Biol. Chem. 281, 29872–29885. doi: 10.1074/jbc.M605052200
Yin, J., Meng, Q., Fu, H., and Gao, H. (2016). Reduced expression of cytochrome oxidases largely explains cAMP inhibition of aerobic growth in Shewanella oneidensis. Sci. Rep. 6:24449. doi: 10.1038/srep24449
Keywords: CRP, transcriptional regulation, lactate, Shewanella, catabolism
Citation: Kasai T, Kouzuma A and Watanabe K (2017) CRP Regulates D-Lactate Oxidation in Shewanella oneidensis MR-1. Front. Microbiol. 8:869. doi: 10.3389/fmicb.2017.00869
Received: 28 February 2017; Accepted: 28 April 2017;
Published: 16 May 2017.
Edited by:
Dongsheng Zhou, Beijing Institute of Microbiology and Epidemiology, ChinaReviewed by:
Thomas Hindré, UMR5525 TIMC-IMAG, Université Grenoble-Alpes, CNRS, FranceHaichun Gao, Zhejiang University, China
Copyright © 2017 Kasai, Kouzuma and Watanabe. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Atsushi Kouzuma, YWtvdXp1bWFAdG95YWt1LmFjLmpw
 Takuya Kasai
Takuya Kasai Atsushi Kouzuma
Atsushi Kouzuma Kazuya Watanabe
Kazuya Watanabe