- Department of the Science of Agriculture, Food and Environment, University of Foggia, Foggia, Italy
Among the innovative trends in the wine sector, the continuous exploration of enological properties associated with wine microbial resources represents a cornerstone driver of quality improvement. Since the advent of starter cultures technology, the attention has been focused on intraspecific biodiversity within the primary species responsible for alcoholic fermentation (Saccharomyces cerevisiae) and, subsequently, for the so-called ‘malolactic fermentation’ (Oenococcus oeni). However, in the last decade, a relevant number of studies proposed the enological exploitation of an increasing number of species (e.g., non-Saccharomyces yeasts) associated with spontaneous fermentation in wine. These new species/strains may provide technological solutions to specific problems and/or improve sensory characteristics, such as complexity, mouth-feel and flavors. This review offers an overview of the available information on the enological/protechnological significance of microbial resources associated with winemaking, summarizing the opportunities and the benefits associated with the enological exploitation of this microbial potential. We discuss proposed solutions to improve quality and safety of wines (e.g., alternative starter cultures, multistrains starter cultures) and future perspectives.
Introduction
Wine has been consumed by humans for thousands of years and produced by crushing grapes and allowing them to ferment using the organisms present on the grapes and in the surrounding environment (Whitener et al., 2016). The microbiology of wines involves two main phases, alcoholic fermentation (AF) and malolactic fermentations (MLF) that rely on a heterogeneous microbiota composed by different indigenous microorganisms (e.g., yeast, bacteria and filamentous fungi). Considerable possible cellular/biochemical interactions can take place among these microbial resources in wine (Liu J. et al., 2017). Although of the entire wine microflora contribute to the wine chemistry, yeasts detain a predominant role (Capozzi et al., 2015). Yeast species are usually classified in two groups: Saccharomyces and non-Saccharomyces (Taillandier et al., 2014). Saccharomyces species play the key role, with Saccharomyces cerevisiae as the dominant species (Masneuf-Pomarède et al., 2010). On the other side, non-Saccharomyces yeast include different genera such as Hanseniaspora, Issatchenkia, Pichia and, Schizosaccharomyces, Brettanomyces, Zygosaccharomyces, Kluyveromyces, Candida, Torulaspora (Taillandier et al., 2014). These yeasts can experience anaerobic or aerobic growth and may persist during the fermentation, competing with Saccharomyces for nutrients, producing secondary compounds or modifying the metabolism of S. cerevisiae (Ciani et al., 2016). Several genera of lactic acid bacteria (LAB) have been reported in association to the wine prokaryotic consortia: Enterococcus, Lactobacillus, Leuconostoc, Oenococcus, Pediococcus, and Weissella (Capozzi et al., 2011; Berbegal et al., 2016; Salvetti et al., 2016; Cappello et al., 2017). In particular, Oenococcus oeni is the main species associated with MLF because of its tolerance to the harsh wine conditions (high ethanol concentration, low pH and nutrient content) (Cappello et al., 2017), even if increasing studies confirmed the relevance of Lactobacillus plantarum strains associated with specific physico-chemical conditions (du Toit et al., 2011).
Inoculation of selected starter cultures in wine must is an established enological practice in order to mitigate product losses or the production of off-flavors (García-Ríos et al., 2014; Whitener et al., 2015). Currently, a debate is still open about the use of commercial starters able to mimic some advantageous enological traits, which are present when the spontaneous fermentation is ruled by indigenous populations (Capozzi et al., 2015). For example, wines produced using single inocula are thought to lack the sensory complexity and rounded palate structure (Bellon et al., 2013). In this respect, mixed fermentation/multi-strains starter cultures including non-Saccharomyces yeasts/malolactic bacteria are generally regarded as having improved characteristics, such as complexity, mouth-feel and flavors (Tronchoni et al., 2017).
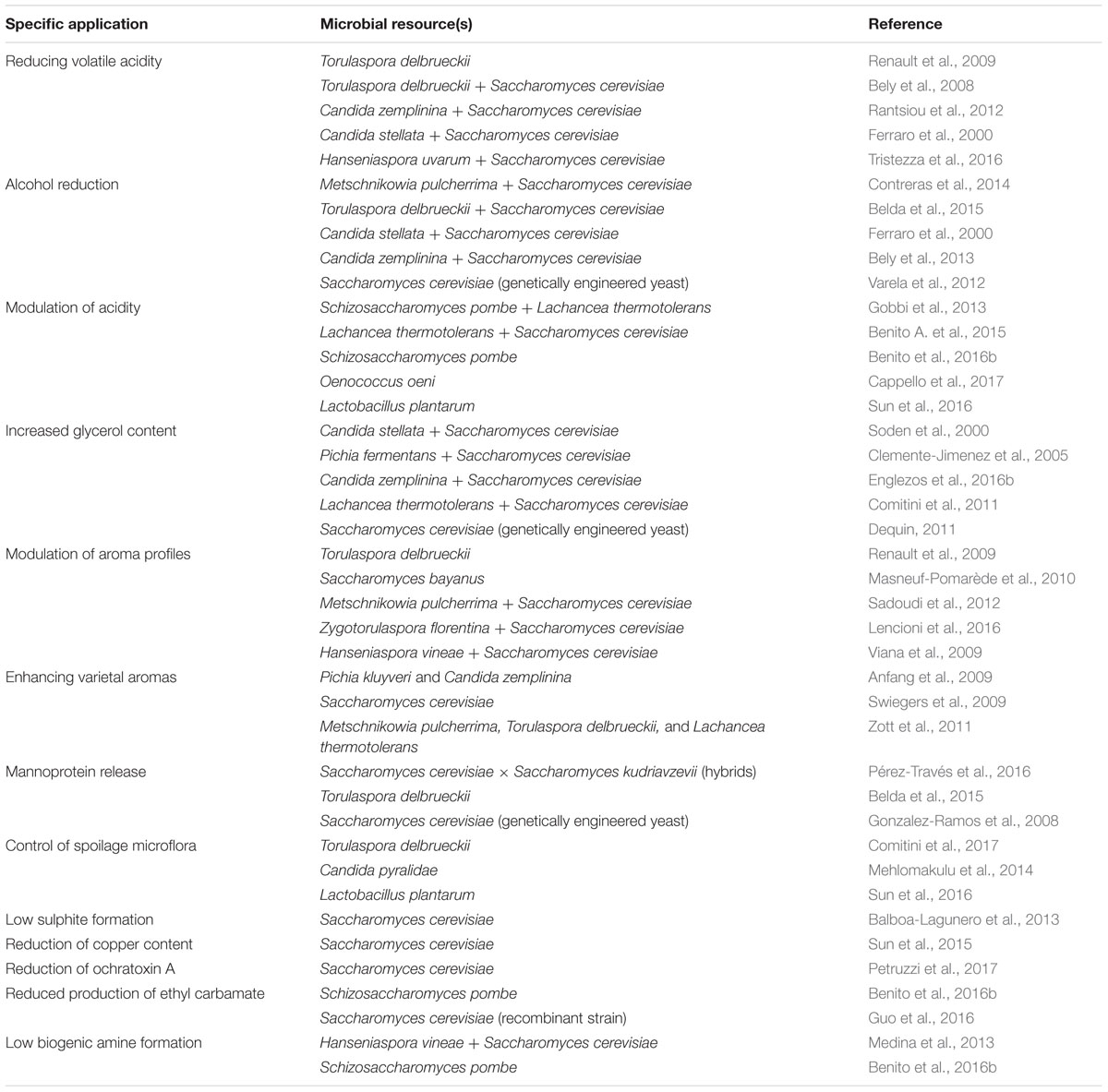
This review offers an overview of the available information on the enological/protechnological significance of microbial resources associated with winemaking, with a special focus on non-Saccharomyces yeast genera/species. A summary of the current state of knowledge about the microbial strategies to solve some technological and safety problems in winemaking is given in Table 1.
Saccharomyces cerevisiae Model for Winemaking Workhorse
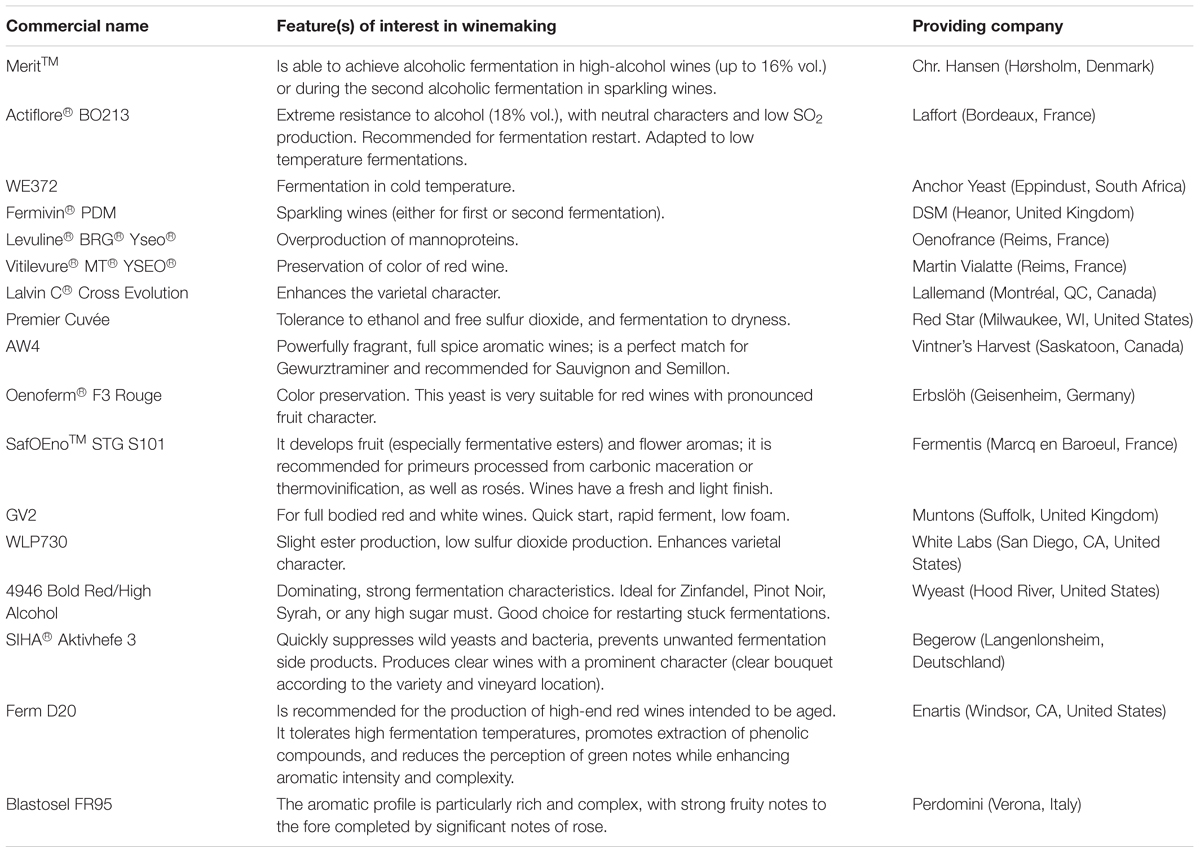
For the last two decades, the natural variation of the yeast S. cerevisiae has been massively exploited with the aim of understanding ecological and evolutionary processes (Cubillos, 2016). It has been used as a model to study aging, regulation of gene expression, signal transduction, cell cycle, metabolism, apoptosis, neurodegenerative disorders, and many other biological processes (Karathia et al., 2011). Genome sequence information is now available for >80 strains of S. cerevisiae in some form (complete, draft, or raw data) (Borneman and Pretorius, 2015). In addition, it is a key player in many industrial fermentation of food matrices such as wine, bread and sake (Cubillos, 2016). The first reported use of a selected yeast starter for wine production dates from 1890, when Müller-Thurgau introduced this technology adapting the techniques developed by Christian Hansen for the Carlsberg Brewery (Padilla et al., 2016). Since then, the role of S. cerevisiae as starter culture in the wine industry has received the most attention. This yeast is not only responsible for the metabolism of grape sugar to alcohol and CO2 but has other important side-roles, including the conversion of grape aroma precursors to sensory active molecules (Jolly et al., 2014; Belda et al., 2017). The metabolism of fermenting S. cerevisiae can be divided into primary and secondary. Primary metabolism is essential for growth, cell division and survival, producing metabolites such as ethanol, glycerol, acetaldehyde, and acetic acid. Secondary metabolites include the fusel alcohols, esters, carbonyls, sulfur compounds, thiols and terpenoids (Romano et al., 2015; Hirst and Richter, 2016). The dominance of S. cerevisiae in the fermentation is expected and desired (Jolly et al., 2014; Capece et al., 2016). One of the main features that allow S. cerevisiae to overcome is its remarkable sugar consumption rate and ethanol production coupled with a high alcohol tolerance. Through this quick proliferation in grape must S. cerevisiae efficiently depletes nitrogen sources and other nutrients required for yeast biomass production from the medium (Tronchoni et al., 2017). In addition, several studies raised evidence that microbial interactions play an important role in the early death of non-Saccharomyces yeasts (Albergaria and Arneborg, 2016). Therefore, S. cerevisiae has been the primary choice for producing wine starters (Albergaria and Arneborg, 2016). In modern winemaking, fermentations are driven largely by single-strain inoculations; pure cultures of selected strains of S. cerevisiae are added to grape must as soon as possible after crushing. This practice ensures the control of vinification, leads to outcomes that are more predictable and decreases the risk of spoilage by other microorganisms (Chambers and Pretorius, 2010). Several studies addressed the genetics underlying these unique properties and tried to unravel the evolutionary path Saccharomyces strains have undergone to become the specialized fermentation organisms they are today. It was shown that duplication of several key genes, such as those encoding alcohol dehydrogenase, hexose transporters, and enzymes linked to glycolysis, as well as global rewiring of the transcriptional network after whole genome duplication, might contribute to the suitability of S. cerevisiae as a driver of industrial fermentations (Steensels and Verstrepen, 2014). There are many-probably hundreds of-different yeast strains available, and the winemaker’s choice could have a significant effect on the quality of the wine (Chambers and Pretorius, 2010). Table 2 shows the main enological properties of some exemplificative commercially available S. cerevisiae wine yeasts. However, while this practice may reduce sources of microbial spoilage, some winemakers feel that the exclusive use of S. cerevisiae has resulted in a lack of organoleptic complexity when compared with successful spontaneous fermentations (Whitener et al., 2015), thus contributing to an increased interest on the role of non-Saccharomyces yeasts in winemaking (Whitener et al., 2016).
Non-Saccharomyces Yeasts
The world wine market is experiencing increasing interest in new yeast strains that can produce unique wines with novel properties (Mylona et al., 2016). Non-Saccharomyces yeasts have garnered interest in winemaking due to their beneficial effects and because consumers are demanding new wine styles (Lleixà et al., 2016). Recently, some commercial yeast manufacturers have already included non-Saccharomyces yeasts as part of their product portfolio (Albergaria and Arneborg, 2016). The main enological properties of some commercially available non-Saccharomyces wine yeasts are outlined in Table 3. The main benefit of using complementary yeast starters in winemaking is the improved aromatic complexity (Comitini et al., 2011; Tronchoni et al., 2017). Non-Saccharomyces yeasts possess enzymatic activities, which can catalyze the release of volatile aroma compounds from non-volatile bound precursors. These yeasts can also affect aroma production directly by their own metabolic activity (production of alcohols and esters) or by the release of extracellular enzymes which transform S. cerevisiae-derived metabolites. Strains of non-Saccharomyces yeasts have also shown potential for producing aroma compounds not associated with fermentation by many strains of S. cerevisiae, such as various monoterpenes and other terpenoid compounds (Rossouw and Bauer, 2016). In addition, the use of non-Saccharomyces yeasts has been proposed to improve glycerol or mannoprotein content, volatile acidity, or color stability (Tronchoni et al., 2017) or to reduce the ethanol levels of wines (Rossouw and Bauer, 2016). On the other hand, there have been a large number of researches on the isolation and characterization of yeast strains degrading malic acid, as alternative to malolactic bacteria-fermentation starters (Kim et al., 2008). Positive features of non-Saccharomyces yeasts have been highlighted regarding to the production of metabolites beneficial for wine quality and stability. This is the case of yeasts producers of active extracellular molecules, able to counteract the development of wild spoilage microorganisms (Comitini et al., 2017).
However, most non-Saccharomyces yeasts cannot ferment to dryness, thus S. cerevisiae should be also inoculated. Two modes of inoculation have been proposed and used (Whitener et al., 2016): the first is known as co-inoculation and some studies have showed that the inoculation of selected non-Saccharomyces yeasts at high cell concentration together with S. cerevisiae might produce wines with distinct characteristics while avoiding stuck fermentations. Others researchers have explored the use of non-Saccharomyces yeasts in sequential inoculations; non-Saccharomyces yeasts are first inoculated at high levels and allowed to ferment on their own for a given amount of time before S. cerevisiae is added to take over the fermentation. This practice gives the non-Saccharomyces yeast more time to express their unique metabolic footprint uninhibited by the stress of Saccharomyces competition (Whitener et al., 2015). Moreover, the effects of the non-Saccharomyces yeasts on fermentation and wine quality were strictly dependent on the Saccharomyces/non-Saccharomyces inoculum ratio (Comitini et al., 2011).
The principal outcomes of fermentations conducted with the aid of non-Saccharomyces yeast genera/species have been documented in the following subsections and summarized in Supplementary Table S1.
Torulaspora delbrueckii
Torulaspora delbrueckii was one of the first commercial non-Saccharomyces yeast to be released (Jolly et al., 2014). This species has been previously suggested for the vinification of low sugar and acidity musts, and it has been used for the production of red and rosé wines in Italy and for Sauvignon Blanc in South Africa (González-Royo et al., 2015). T. delbrueckii is characterized by high purity fermentation, low production of glycerol, acetaldehyde, acetic acid, and ethyl acetate (Loira et al., 2015). When used in sequential or mixed fermentations with S. cerevisiae, it can contribute to correct certain defects such as the volatile acidity (Loira et al., 2015). In Semillon wine, a mixed T. delbrueckii/S. cerevisiae culture at a 20:1 ratio produced 53% less in volatile acidity and 60% less acetaldehyde than a pure culture of S. cerevisiae (Bely et al., 2008). Some authors showed that the strong β-glucosidase activity of this species enhanced wine aroma by modulating the levels of nor-isoprenoids, terpenols, and lactones by hydrolysing their respective precursors (Renault et al., 2015). Maturano et al. (2012) confirmed the high production of extracellular enzymes of enological relevance by this species. Amarone wines produced by co-inoculation/sequential inoculation with T. delbrueckii and S. cerevisiae were judged to have increased aroma intensity, including ‘ripe red fruit’ aroma, increased sweetness and astringency and decreased intensity for vegetal attributes (Azzolini et al., 2012). Gewurztraminer wines produced by sequential inoculation showed increased concentration of terpenes α-terpineol and linalool (Cus and Jenko, 2013). T. delbrueckii/S. cerevisiae multi starters have been proposed to modulate wine flavor in Sauvignon Blanc and Merlot (Renault et al., 2015), as well as Shiraz wines (Loira et al., 2015).
The growth of T. delbrueckii is often negatively affected by the presence of S. cerevisiae. Indirect interaction, i.e., interaction between strains via components of the medium have been demonstrated. Substrate competition or amensalism (production by S. cerevisiae of metabolites that inhibit T. delbrueckii growth) are examples of the indirect interactions already suggested (Taillandier et al., 2014).
Recently, new killer toxins from T. delbrueckii with potential biocontrol activity of Brettanomyces bruxellensis, Pichiaguilliermondii, P. manshurica and P. membranifaciens wine spoilage were identified and characterized (Comitini et al., 2017).
Lachancea thermotolerans
Lachancea thermotolerans has been investigated for its ability to enhance wine acidity and improve the overall quality (Whitener et al., 2016). L. thermotolerans produces high concentration of L-lactic acid from glucose and fructose as well as low levels of volatile acidity and undesirable flavor compounds (Erten and Tanguler, 2010). This attribute could be of concern to address the problems of increased alcohol content and a reduction in the total acidity of wines associated with global climate change and variations in viticulture and wine-making practices (Balikci et al., 2016). L. thermotolerans was used as starter culture in Airén wine (increase of lactic acid of 3.18 g/L and pH reduction of 0.22) (Benito et al., 2016a), Emir wines (the use of L. thermotolerans in mixed and sequential cultures led to an increase in final total acidity of 5.40–6.28 g/L) (Balikci et al., 2016), Riesling wine (peach/apricot character) (Benito S. et al., 2015), Sangiovese and Cabernet-Sauvignon (spicy attributes) (Gobbi et al., 2013).
Metschnikowia pulcherrima
Metschnikowia pulcherrima is commonly associated with grapes and wine (Whitener, 2016). It is a high producer of β-glucosidase and its presence in mixed cultures can decrease the volatile acidity and increase the production of medium-chain fatty acids, higher alcohol, esters, terpenols and glycerol (González-Royo et al., 2015). Wines of the grape varieties Chardonnay and Shiraz obtained by sequential fermentation with Caesalpinia pulcherrima and S showed ca. 1% v/v lower ethanol concentration (Varela et al., 2016). In addition, M. pulcherrima has been reported to increase: (1) the levels of methyl butyl-, methyl propyl-, and phenethyl esters production in Sauvignon Blanc (Whitener et al., 2016); (2) the ‘overall impression,’ ‘citrus/grape fruit’ and ‘pear’ attributes of Riesling (Benito S. et al., 2015); (3) foam persistence and ‘smoky’ and ‘flowery’ attributes of Macabeo wine (González-Royo et al., 2015). It has also been reported that M. pulcherrima might have an antagonistic effect toward several yeast. cerevisiaes including S. cerevisiae which leads to delays in fermentation. This phenomenon was the result of a pulcherrimin pigment (Jolly et al., 2014).
Candida Species
The genus Candida is large and extremely diverse with over 50 different identified species several of which have been associated with winemaking (Whitener, 2016). Among the non-Saccharomyces wine yeasts involved in grape juice fermentation, C. stellata has been frequently isolated during the course of must fermentations in different countries (fermentation of botrytized wines and other wines produced by overripe grapes, in cooked musts, and in traditional balsamic vinegars) (Tofalo et al., 2012). Ciani and Maccarelli (1998) suggested that it might be used as a starter culture to increase glycerol levels in Trebbiano Toscano wine, where it 11.76 g/L of glycerol, which is higher than the sensory threshold level for glycerol sweetness, i.e., 5.2 g/L (Jolly et al., 2006). Similarly, wines of the grape variety Syrah obtained by mixed and sequential fermentation with C. cantarellii and S. cerevisiae showed 44.3 to 52.8% glycerol content higher than the control wines (Toro and Vazquez, 2002). C. sake increased concentrations of terpenes and higher alcohols in Pedro Gimenez wines (Maturano et al., 2015).
C. zemplinina (synonym Starmerella bacillaris; Duarte et al., 2012) is a non-Saccharomyces yeast, isolated for the first time in Napa Valley (California, United States) in 2002, under the name EJ1 (Mills et al., 2002). This yeast differs from the other common non-Saccharomyces yeasts, since it can survive and resist until the end of the AF due to its ability to tolerate high concentrations of ethanol (Englezos et al., 2016a).
Candida zemplinina showed some interesting characteristics, such as: (1) high glycerol production (Tofalo et al., 2012; Zara et al., 2014; Englezos et al., 2016b); (2) reduced ethanol yield (Bely et al., 2013; Englezos et al., 2016b); (3) increase of aroma complexity (Tofalo et al., 2016; Whitener et al., 2016); (4) ability to metabolize malic acid (Tofalo et al., 2012); (5) reduction of the acetic acid production in combination with S. cerevisiae (Rantsiou et al., 2012). These applications support the use of C. zemplinina, which could be a good choice to achieve various desired results, mainly due to its fructophilic character and the poor ethanol yield from sugar (Masneuf-Pomarede et al., 2015; Englezos et al., 2016a).
Recently, Mehlomakulu et al. (2014) found two novel killer toxins, CpKT1 and CpKT2, from the wine isolated strains of C. pyralidae, able to control the development of spoilage yeast B. bruxellensis. In addition, these killer toxins inhibited neither the Saccharomyces cerevisiae nor the LAB strains tested.
Pichia Species
Pichia fermentans was investigated by Clemente-Jimenez et al. (2005) in lab-scale vinifications of Macabeo wine. Sequential fermentations with P. fermentans and S. cerevisiae produced wines with increased concentrations of acetaldehyde, ethyl acetate, 1-propanol, n-butanol, 1-hexanol, ethyl caprilate, 2,3-butanediol and glycerol.
Sequential inoculum of Riesling must with P. kluyveri and S. cerevisiae increased the ‘overall impression’ and ‘peach/apricot’ character (Benito S. et al., 2015). Co-fermentation of Sauvignon Blanc grape juice with P. kluyveri has been reported to lead to higher levels of 3-mercaptohexyl acetate (Anfang et al., 2009); however, it has also been reported to produce many off odor compounds (Whitener et al., 2016).
Malic acid is sometimes detrimental to the quality of wines when present at high concentrations in some varieties. Several grape varieties contain considerable amounts of malic acid and grapes grown in the cooler regions contain higher amounts of malic acid than those grown in the warmer regions. Excessive amounts of malic acid (15–16 mg/ml) were detected in grapes grown during exceptionally cold summer in cool regions (Kim et al., 2008). In this respect, co-inoculation of P. kudriavzevii enhance the catalysis of malic acid in grape juice fermentation (Kim et al., 2008).
It is reported that P. membranifaciens increased esters production in Muscat wine (Viana et al., 2008).
Schizosaccharomyces pombe
Schizosaccharomyces pombe was initially considered as a spoilage yeast because of the production of undesirable metabolites with a negative sensory impact; on the other hand, it could be successfully used at an industrial level in cane sugar fermentation during rum making, palm wine production and cocoa fermentation (Mylona et al., 2016). However, Schizosaccharomyces pombe species is highly appreciated in colder regions because of its ability to completely transform the malic acid of the must into ethanol, thanks to its particular metabolism of maloalcoholic fermentation (Loira et al., 2015). In this respect, S. pombe perform effective malic acid deacidification and significantly reduces the levels of biogenic amines and ethyl carbamate precursors without the need for any secondary bacterial MLF (Benito et al., 2014, 2016b). Mylona et al. (2016) confirmed the ability of yeast to produce safe wines.
One new application exploits Schizosaccharomyces ability to increase the formation of vitisins and vinylphenolic pyranoanthocyanin (Loira et al., 2015). In addition, the rapid autolytic release of cell wall polysaccharides after death could reduce the time required to complete aging over lees (Benito et al., 2014).
Hanseniaspora Species
The apiculate yeast Hanseniaspora uvarum is the non-Saccharomyces found at highest levels in grape must, in fact it is among the relevant contributors to wine quality (Jolly et al., 2014). In single fermentation, H. uvarum produced low volatile acidity and high levels of glycerol (Tofalo et al., 2016). H. uvarum has also been used in mixed fermentations with S. cerevisiae for wine production. Tristezza et al. (2016) assessed the oenological potential of H. uvarum in co-inoculation and in a sequential inoculation with S. cerevisiae for industrial wine production. The mixed starter was able to successfully dominate the different stages of the fermentation process and the H. uvarum strain ITEM8795 contributed to increase the wine organoleptic quality and reduce the volatile acidity. In Solaris wine, mixed fermentations produced higher levels of glycerol, heptyl acetate, and 2-phenylethyl acetate (Liu et al., 2016).
Hanseniaspora vineae has been demonstrated to increase fruity aromas and produce high amounts of acetate esters, such as 2-phenylethyl acetate and ethyl acetate, in wines produced by a sequential fermentation with S. cerevisiae (Lleixà et al., 2016). Co-fermentation of Bobal grape must with H. vineae and S. cerevisiae produced wines that not only showed an increased concentration of 2-phenylethyl acetate (approximately three–ninefold higher) but also exhibited higher ‘fruity’ sensory scores than wines produced with S. cerevisiae pure culture (Viana et al., 2009). Chardonnay wines produced with H. vineae and S. cerevisiae have shown increased aroma and flavor diversity and reduced biogenic amines content (Medina et al., 2013).
Hanseniaspora guilliermondii can contribute to the overall quality of wines. In single-culture this yeast produced high amounts of 2-phenylethyl acetate (Viana et al., 2008). Tinta Roriz grape must inoculated with H. guilliermondii led to the production of wine with higher concentrations of 1-propanol, 2-phenylethyl acetate and 3-(methylthio)propionic acid, and lower amounts of ethyl hexanoate, pentanoic acid, free fatty acids, 2-methyltetrahydrothiophen-3-one and acetic acid-3-(methylthio)propyl ester (Moreira et al., 2011).
Zygosaccharomyces Species
The genus Zygosaccharomyces is known, together with Brettanomyces species (Capozzi et al., 2016a), for its ability to spoil wine, namely sweet and sparkling wines (Whitener et al., 2016). Zygosaccharomyces bailii and Zygosaccharomyces rouxii are often the source of spoilage in acidic and shelf-stable foods as well as sweet wines due to their ability to tolerate high acid, salt and sugar conditions (Whitener, 2016). However, selected strains of Zygosaccharomyces spp. might be useful (Garavaglia et al., 2015), because they can yield high levels of ethyl esters in Chardonnay wine (Garavaglia et al., 2015). Ethyl esters are highly interesting because they have a pleasant fruity and floral aromatic note, and are responsible for beer and wine aroma (Garavaglia et al., 2015).
Zygosaccharomyces bailii is fructophilic, and metabolizes fructose more easily than glucose (Garavaglia et al., 2015). This triat could be beneficial in grape musts from over-ripened grapes (Jolly et al., 2006). Zuehlke et al. (2015) used inoculation with Z. bailii to remove residual sugar from Cabernet Sauvignon and Syrah sluggish fermentations.
Zygosaccharomyces kombuchaensis is a newly discovered yeast. It can contribute to increase flavor intensity’ and several ‘fruity’ attributes in Ribolla Gialla (Dashko et al., 2015), as well as benzaldehyde in Sauvignon Blanc and Syrah (Whitener et al., 2015) wines.
Debaryomyces Species
Debaryomyces may contribute to the maturation, aroma, and flavor of foods, such as cheese and meat products, but it also might spoil fermented food products (Wrent et al., 2014). In winemaking, two species are of interest: Debaryomyces pseudopolymorphus and Debaryomyces vanriji. Co-fermentation of Chardonnay grape juice with D. pseudopolymorphus and S. cerevisiae resulted in an increased concentration of the terpenols (citronellol, nerol, and geraniol) (Cordero Otero et al., 2003). Sequential fermentation of Muscat of Frontignan grape juice with D. vanriji and S. cerevisiae produced wines with increased concentration of geraniol (Garcia et al., 2002). Sequential fermentation of Pedro Gimenez must with D. vanriji and S. cerevisiae increased the concentrations of esters and fatty acids (Maturano et al., 2015).
Kazachstania Species
The Kazachstania genus as a whole is fairly new; it was first mentioned in literature in 2003. Kazachstania aerobia was first identified in 2004 from corn silage while Kazachstania gamospora was discovered as a species in 2007. Being relatively new yeasts and closely related to S. cerevisiae, the genus is worthy of investigation (Whitener, 2016).
Dashko et al. (2015) used K. gamospora in Ribolla Gialla fermentation, resulting in increased acetate and ethyl ester amounts. In Sauvignon blanc and Syrah musts, this yeast caused a 200-fold increase of phenethyl propionate (Whitener et al., 2015). Sauvignon Blanc wines produced with K. aerobia and S. cerevisiae have shown increased ethyl acetate production (Whitener et al., 2016).
Wickerhamomyces anomalus
Wickerhamomyces anomalus is a constituent of the normal grape flora in the early phases of fermentation. It could spoil wine by excessive production of acetic acid and ethyl acetate, but also contributes to wine aroma by the production of volatile compounds. This species has gained considerable biotechnological interest due to its tolerance toward environmental stress factors (e.g., low pH, high osmolarity), metabolic versatility and production of exoenzymes (Sabel et al., 2014). Mazuela wines produced by sequential inoculation of W. anomalus and S. cerevisiae showed increased concentration of acetate- and ethyl- esters, along with a significant and panel preference (Cañas et al., 2014).
Williopsis saturnus
Williopsis saturnus has been reported to increase the levels of main terpenols (linalool, citronellol, and α-terpineol), produce some terpenoid esters (citronellyl and neryl acetate) and retain the concentration of cis-rose oxide in mango wine with ethanol levels of 2–4% (v/v) (Chen et al., 2015). This specie is not generally found from the natural environment of surfaces of grapes and winery equipments: however, it might potentially enhance the fruity flavor in wines obtained from neutral cultivar characteristics (Erten and Tanguler, 2010).
Emir wines produced by co-inoculation with W. saturnus and S. cerevisiae showed higher concentrations of acetic acid, ethyl acetate and isoamyl acetate (Erten and Tanguler, 2010).
Zygotorulaspora florentina
Zygotorulaspora florentina was used in mixed fermentations at different inoculum ratio with S. cerevisiae, and caused an increase of the production of polysaccharides and a modulation of the final concentrations of the various volatile compounds (Domizio et al., 2011). Recently, Sangiovese wines produced by co-inoculation with Zygotorulaspora florentina and S. cerevisiae showed an enhancement of polysaccharides and 2-phenylethanol content, a reduction of volatile acidity, and high concentration of glycerol and esters (Lencioni et al., 2016).
Bacteria
Malolactic fermentations is a secondary microbial-based biochemism that usually takes place in wine during or at the end of AF; MLF is mainly carried out by one or more species of LAB (Lerm et al., 2011; Cappello et al., 2017). During this phase, the biological conversion of interest in enology is the L-malic acid decarboxylation to produce L-lactic acid and carbon dioxide. Aside from impacts on acidity, LAB can also metabolize other precursors present in wine during fermentation and, therefore, affect the chemical composition of the wine resulting in an increased complexity of wine aroma and flavor (Sumby et al., 2014; Campbell-Sills et al., 2016).
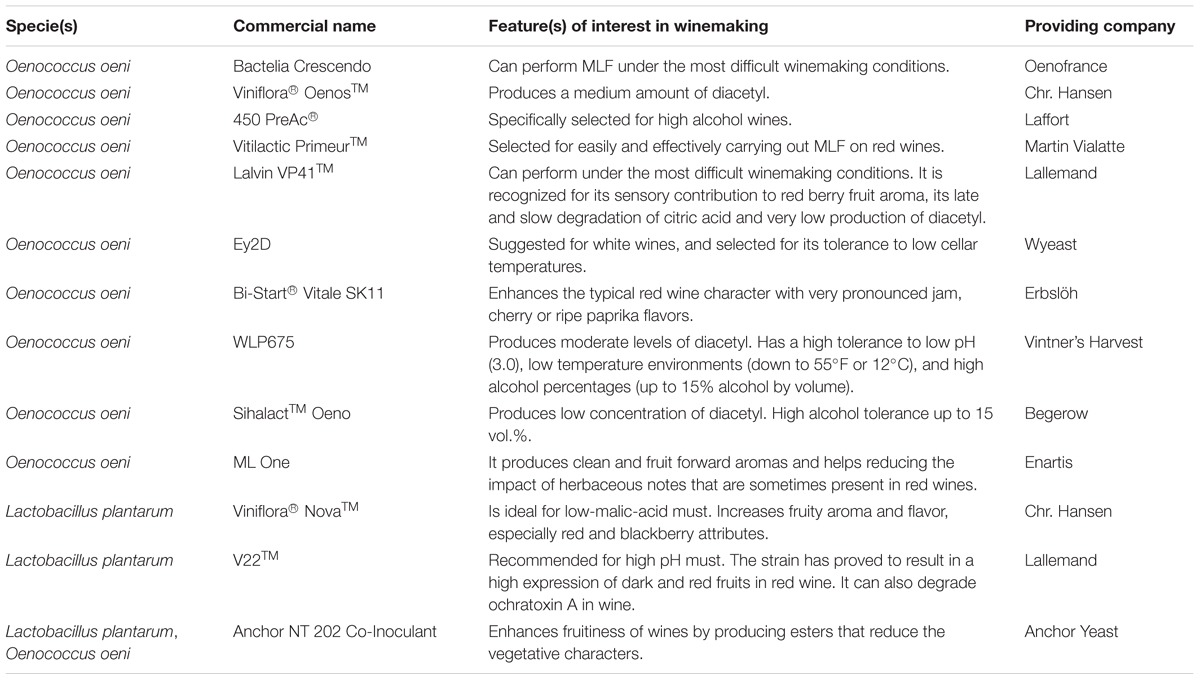
Spontaneous MLF implies several risks, such as a considerable increase in volatile acidity, the consumption of residual sugars, and the formation of undesirable metabolites, such as biogenic amines, that can affect human health and lead to low quality wines (Spano et al., 2010). In recent years, wine industries have moved toward using pure starter cultures of selected LAB to promote a reliable and rapid malic acid bioconversion (Spano et al., 2010). Table 4 shows the main enological properties of exemplificative commercially available wine malolactic bacteria.
Actually, the production of efficient malolactic starter cultures has become one of the main challenges for oenological research (Lerm et al., 2011; Berbegal et al., 2016; Cappello et al., 2017). There are various important criteria to address when selecting LAB for possible use in a starter culture, like the ability to tolerate low pH, high ethanol and SO2 concentrations, good growth characteristics under winemaking conditions, compatibility with the selected yeast strain, the inability to produce biogenic amines and the lack of off-flavor or off-odor production (Capozzi et al., 2010; Lerm et al., 2011). A minor but also important aspect to be considered is the susceptibility of LAB to polyphenols. O. oeni is the major LAB used in commercial starter cultures for MLF (Lerm et al., 2011). O. oeni has been the only species within the Oenococcus genus until the mid-2000s when O. kitaharae was identified in composting distilled shochu residue. Over centuries of selective pressure, O. oeni has honed and perfected various adaptive strategies that enable it to outcompete with other potential MLF bacteria, during the later stages of vinification and thus to dominate in wine (Campbell-Sills et al., 2015). Recently a third Oenococcus species has been identified, O. alcoholitolerans isolated from Brazilian cachaça (Cappello et al., 2017).
In the selection of MLF starters, a challenge is the time of inoculation. Starter cultures can be co-inoculated with yeast (at the beginning or toward the end of AF), or sequentially (after AF) (Bartowsky et al., 2015). Generally, it has been demonstrated that bacteria inoculated in must performed better than those inoculated after AF, especially when cell growth conditions are not favorable (Azzolini et al., 2010). Some Lactobacillus species have also showed the ability to survive the harsh wine conditions; the species Lactobacillus plantarum has shown the most potential as a starter culture (Lerm et al., 2011). This versatile bacterium tolerates ethanol up to 14% v/v and has similar SO2 tolerance of O. oeni (Cappello et al., 2017). The introduction of some L. plantarum strains to the fermenting musts could significantly modify the wine aroma profile due to a different enzymatic profile. In addition, L. plantarum could synthesize antimicrobial peptides which might help preventing the production of undesired compounds, or inhibiting the indigenous LAB microflora (Sun et al., 2016). Due to these characteristics, selected strains of L. plantarum are currently being commercialized to induce MLF in wine (Table 4).
Concluding Remarks
Wine fermentation is generally performed through inoculated or spontaneous fermentation (Martiniuk et al., 2016). One of the main trends in the industry of starter cultures for enology relies on the survey of the microbial resources associated with spontaneous fermentation in order to design products able to maximize wine quality (Bokulich et al., 2016; Corbo et al., 2017; Romano and Capece, 2017; Russo et al., 2017). In this light, our contribute proposes an overview of the opportunities and benefits associated with the exploitation of this microbial potential in winemaking. Considering future perspectives, the increasing number of species/strains used, often associated to new isolations from spontaneous fermentations (e.g., Garofalo et al., 2015; Garofalo et al., 2016), introduces a relevant change in terms of interspecific interactions (Ciani et al., 2016; Liu Y. et al., 2017; Tronchoni et al., 2017). A field of particular interest if we consider that different grape juices and batch volumes could influence the growth and final biomass of yeasts in mixed fermentations (Gobbi et al., 2013) and that most studies have been performed at laboratory scale without an effective validation at industrial or semi-industrial scale, questioning their applicability at cellar (Belda et al., 2016). Moreover, it is relevant to underline how this protechnological potential is often a reservoir of interesting biotechnological applications in the food sector (Capozzi et al., 2011, 2016b; Russo et al., 2016; Petruzzi et al., 2017).
Author Contributions
LP, VC, CB, GS, MC, MS, and AB performed an accurate research in the literature and planned paper. LP wrote the paper. LP, VC, CB, GS, MC, MS, and AB revised the manuscript.
Funding
This research was supported by Apulian Region in the framework of projects: (a) “Innovazioni di processo e di prodotto nel comparto dei vini spumanti da vitigni autoctoni pugliesi” - IPROVISP (Bando “Aiuti a Sostegno Cluster Tecnologici Regionali”; Project code VJBKVF4); (b) “Biotecnologie degli alimenti per l’innovazione e la competitività delle principali filiere regionali: estensione della conservabilità e aspetti funzionali” – BIOTECA (Bando “Aiuti a Sostegno Cluster Tecnologici Regionali”; Project code QCBRAJ6); (c) “Programma regionale a sostegno della specializzazione intelligente e della sostenibilità sociale ed ambientale—FutureInResearch” - ‘Fondo di Sviluppo e Coesione 2007–2013—APQ Ricerca Regione Puglia.’ Pubblicazione realizzata con un contributo sui fondi del 5 × 1000 dell’IRPEF a favore dell’Università di Foggia, in memoria di Gianluca Montel.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2017.00995/full#supplementary-material
References
Albergaria, H., and Arneborg, N. (2016). Dominance of Saccharomyces cerevisiae in alcoholic fermentation processes: role of physiological fitness and microbial interactions. Appl. Microbiol. Biotechnol. 100, 2035–2046. doi: 10.1007/s00253-015-7255-0
Anfang, N., Brajkovich, M., and Goddard, M. R. (2009). Co-fermentation with Pichia kluyveri increases varietal thiol concentrations in Sauvignon Blanc. Aust. J. Grape Wine Res. 15, 1–8. doi: 10.1111/j.1755-0238.2008.00031.x
Azzolini, M., Fedrizzi, B., Tosi, E., Finato, F., Vagnoli, P., Scrinzi, C., et al. (2012). Effects of Torulaspora delbrueckii and Saccharomyces cerevisiae mixed cultures on fermentation and aroma of Amarone wine. Eur. Food Res. Technol. 235, 303–313. doi: 10.1007/s00217-012-1762-3
Azzolini, M., Tosi, E., Vagnoli, P., Krieger, S., and Zapparoli, G. (2010). Evaluation of technological effects of yeast-bacterial co-inoculation in red table wine production. Ital. J. Food Sci. 3, 257–263.
Balboa-Lagunero, T., Arroyo, T., Cabellos, J. M., and Aznar, M. (2013). Yeast selection as a tool for reducing key oxidation notes in organic wines. Food. Res. Int. 53, 252–259. doi: 10.1016/j.foodres.2013.04.006
Balikci, E. K., Tanguler, H., Jolly, N. P., and Erten, H. (2016). Influence of Lachancea thermotolerans on cv. Emir wine fermentation. Yeast 33, 313–321. doi: 10.1002/yea.3166
Bartowsky, E. J., Costello, P. J., and Chambers, P. J. (2015). Emerging trends in the application of malolactic fermentation. Aust. J. Grape Wine Res. 21, 663–669. doi: 10.1111/ajgw.12185
Belda, I., Conchillo, L. B., Ruiz, J., Navascués, E., Marquina, D., and Santos, A. (2016). Selection and use of pectinolytic yeasts for improving clarification and phenolic extraction in winemaking. Int. J. Food Microbiol. 223, 1–8. doi: 10.1016/j.ijfoodmicro.2016.02.003
Belda, I., Navascués, E., Marquina, D., Santos, A., Calderon, F., and Benito, S. (2015). Dynamic analysis of physiological properties of Torulaspora delbrueckii in wine fermentations and its incidence on wine quality. Appl. Microbiol. Biotechnol. 99, 1911–1922. doi: 10.1007/s00253-014-6197-2
Belda, I., Zarraonaindia, I., Perisin, M., Palacios, A., and Acedo, A. (2017). From vineyard soil to wine fermentation: microbiome approximations to explain the “terroir” concept. Front. Microbiol. 8:821. doi: 10.3389/fmicb.2017.00821
Bellon, J. R., Schmid, F., Capone, D. L., Dunn, B. L., and Chambers, P. J. (2013). Introducing a new breed of wine yeast: interspecific hybridisation between a commercial Saccharomyces cerevisiae wine yeast and Saccharomyces mikatae. PLoS ONE 8:e62053. doi: 10.1371/journal.pone.0062053
Bely, M., Renault, P., da Silva, T., Masneuf-Pomarede, I., Albertin, W., Moine, V., et al. (2013). “Non-conventional yeasts and alcohol level reduction,” in Alcohol Level Reduction in Wine, ed. P. L. Teissedre (Bordeaux: Vigne et Vin Publications Internationales), 33–37.
Bely, M., Stoeckle, P., Masneuf-Pomarède, I., and Dubourdieu, D. (2008). Impact of mixed Torulaspora delbrueckii–Saccharomyces cerevisiae culture on high-sugar fermentation. Int. J. Food Microbiol. 122, 312–320. doi: 10.1016/j.ijfoodmicro.2007.12.023
Benito, A., Calderón, F., Palomero, F., and Benito, S. (2015). Combined use of selected Schizosaccharomyces pombe and Lachancea thermotolerans yeast strains as an alternative to the traditional malolactic fermentation in red wine production. Molecules 20, 9510–9523. doi: 10.3390/molecules20069510
Benito,Á., Calderón, F., Palomero, F., and Benito, S. (2016a). Quality and composition of Airén wines fermented by sequential inoculation of Lachancea thermotolerans and Saccharomyces cerevisiae. Food Technol. Biotechnol. 54, 135–144. doi: 10.17113/ftb.54.02.16.4220
Benito,Á., Jeffares, D., Palomero, F., Calderón, F., Bai, F.-Y., Bähler, J., et al. (2016b). Selected Schizosaccharomyces pombe strains have characteristics that are beneficial for winemaking. PLoS ONE 11:e0151102. doi: 10.1371/journal.pone.0151102
Benito, S., Hofmann, T., Laier, M., Lochbühler, B., Schüttler, A., Ebert, K., et al. (2015). Effect on quality and composition of riesling wines fermented by sequential inoculation with non-Saccharomyces and Saccharomyces cerevisiae. Eur. Food Res. Technol. 241, 707–717. doi: 10.1007/s00217-015-2497-8
Benito, S., Palomero, F., Gálvez, L., Morata, A., Calderón, F., Palmero, D., et al. (2014). Quality and composition of red wine fermented with Schizosaccharomyces pombe as sole fermentative yeast, and in mixed and sequential fermentations with Saccharomyces cerevisiae. Food Technol. Biotechnol. 52, 376–382.
Berbegal, C., Peña, N., Russo, P., Grieco, F., Pardo, I., Ferrer, S., et al. (2016). Technological properties of Lactobacillus plantarum strains isolated from grape must fermentation. Food Microbiol. 57, 187–194. doi: 10.1016/j.fm.2016.03.002
Bokulich, N. A., Collins, T. S., Masarweh, C., Allen, G., Heymann, H., Ebeler, S. E., et al. (2016). Associations among wine grape microbiome, metabolome, and fermentation behavior suggest microbial contribution to regional wine characteristics. mBio 7:e00631-16. doi: 10.1128/mBio.00631-16
Borneman, A. R., and Pretorius, I. S. (2015). Genomic insights into the Saccharomyces sensu stricto complex. Genetics 199, 281–291. doi: 10.1534/genetics.114.173633
Campbell-Sills, H., Capozzi, V., Romano, A., Cappellin, L., Spano, G., Breniaux, M., et al. (2016). Advances in wine analysis by PTR-ToF-MS: optimization of the method and discrimination of wines from different geographical origins and fermented with different malolactic starters. Int. J. Mass Spectrom. 39, 42–51. doi: 10.1016/j.ijms.2016.02.001
Campbell-Sills, H., El Khoury, M., Favier, M., Romano, A., Biasioli, F., Spano, G., et al. (2015). Phylogenomic analysis of Oenococcus oeni reveals specific domestication of strains to cider and wines. Genome Biol. Evol. 7, 1506–1518. doi: 10.1093/gbe/evv084
Cañas, P. M. I., García-Romero, E., Manso, J. M. H., and Fernández-González, M. (2014). Influence of sequential inoculation of Wickerhamomyces anomalus and Saccharomyces cerevisiae in the quality of red wines. Eur. Food Res. Technol. 239, 279–286. doi: 10.1007/s00217-014-2220-1
Capece, A., Granchi, L., Guerrini, S., Mangani, S., Romaniello, R., Vincenzini, M., et al. (2016). Diversity of Saccharomyces cerevisiae strains isolated from two Italian wine-producing regions. Front. Microbiol. 7:1018. doi: 10.3389/fmicb.2016.01018
Capozzi, V., Di Toro, M. R., Grieco, F., Michelotti, V., Salma, M., Lamontanara, A., et al. (2016a). Viable but not culturable (VBNC) state of Brettanomyces bruxellensis in wine: new insights on molecular basis of VBNC behaviour using a transcriptomic approach. Food Microbiol. 59, 196–204. doi: 10.1016/j.fm.2016.06.007
Capozzi, V., Garofalo, C., Chiriatti, M. A., Grieco, F., and Spano, G. (2015). Microbial terroir and food innovation: the case of yeast biodiversity in wine. Microbiol. Res. 181, 75–83. doi: 10.1016/j.micres.2015.10.005
Capozzi, V., Ladero, V., Beneduce, L., Fernández, M., Alvarez, M. A., Benoit, B., et al. (2011). Isolation and characterization of tyramine-producing Enterococcus faecium strains from red wine. Food Microbiol. 28, 434–439. doi: 10.1016/j.fm.2010.10.005
Capozzi, V., Makhoul, S., Aprea, E., Romano, A., Cappellin, L., Sanchez Jimena, A., et al. (2016b). PTR-MS characterization of VOCs associated with commercial aromatic bakery yeasts of wine and beer origin. Molecules 21:483. doi: 10.3390/molecules21040483
Capozzi, V., Russo, P., Beneduce, L., Weidmann, S., Grieco, F., Guzzo, J., et al. (2010). Technological properties of Oenococcus oeni strains isolated from typical southern Italian wines. Lett. Appl. Microbiol. 50, 327–334. doi: 10.1111/j.1472-765X.2010.02795.x
Cappello, M. S., Zapparoli, G., Logrieco, A., and Bartowsky, E. J. (2017). Linking wine lactic acid bacteria diversity with wine aroma and flavour. Int. J. Food Microbiol. 243, 16–27. doi: 10.1016/j.ijfoodmicro.2016.11.025
Chambers, P. J., and Pretorius, I. S. (2010). Fermenting knowledge: the history of winemaking, science and yeast research. EMBO Rep. 11, 914–920. doi: 10.1038/embor.2010.179
Chen, D., Yap, Z. Y., and Liu, S.-Q. (2015). Evaluation of the performance of Torulaspora delbrueckii, Williopsis saturnus, and Kluyveromyces lactis in lychee wine fermentation. Int. J. Food Microbiol. 206, 45–50. doi: 10.1016/j.ijfoodmicro.2015.04.020
Ciani, M., and Maccarelli, F. (1998). Oenological properties of non-Saccharomyces yeasts associated with wine-making. World J. Microbiol. Biotechnol. 14, 199–203. doi: 10.1023/A:1008825928354
Ciani, M., Morales, P., Comitini, F., Tronchoni, J., Canonico, L., Curiel, J. A., et al. (2016). Non-conventional yeast species for lowering ethanol content of wines. Front. Microbiol. 7:642. doi: 10.3389/fmicb.2016.00642
Clemente-Jimenez, J. M., Mingorance-Cazorla, L., Martínez-Rodríguez, S., Las Heras-Vázquez, F. J., and Rodríguez-Vico, F. (2005). Influence of sequential yeast mixtures on wine fermentation. Int. J. Food Microbiol. 98, 301–308. doi: 10.1016/j.ijfoodmicro.2004.06.007
Comitini, F., Capece, A., Ciani, M., and Romano, P. (2017). New insights on the use of wine yeasts. Curr. Opin. Food Sci. 13, 44–49. doi: 10.1016/j.cofs.2017.02.005
Comitini, F., Gobbi, M., Domizio, P., Romani, C., Lencioni, L., Mannazzu, I., et al. (2011). Selected non-Saccharomyces wine yeasts in controlled multistarter fermentations with Saccharomyces cerevisiae. Food Microbiol. 28, 873–882. doi: 10.1016/j.fm.2010.12.001
Contreras, A., Hidalgo, C., Henschke, P. A., Chambers, P. J., Curtin, C., and Varela, C. (2014). Evaluation of non-Saccharomyces yeasts for the reduction of alcohol content in wine. Appl. Environ. Microbiol. 80, 1670–1678. doi: 10.1128/AEM.03780-13
Corbo, M. R., Racioppo, A., Monacis, N., and Speranza, B. (2017). “Commercial starters or autochtonous strains? That is the question,” in Starter Cultures in Food Production, eds B. Speranza, A. Bevilacqua, M. R. Corbo, and M. Sinigaglia (Hoboken, NJ: John Wiley & Sons), 174–198. doi: 10.1002/9781118933794.ch10
Cordero Otero, R., Ubeda Iranzo, J. F., Briones-Perez, A. I., Potgieter, N., Villena, M. A., Pretorius, I. S., et al. (2003). Characterization of the β-glucosidase activity produced by enological strains of non-Saccharomyces yeast. J. Food Sci. 68, 2564–2569. doi: 10.1111/j.1365-2621.2003.tb07062.x
Cubillos, F. A. (2016). Exploiting budding yeast natural variation for industrial processes. Curr. Genet. 62, 745–751. doi: 10.1007/s00294-016-0602-6
Cus, F., and Jenko, M. (2013). The Influence of Yeast Strains on the Composition and Sensory Quality of Gewürztraminer Wine. Available at: http://www.ftb.com.hr/index.php/info-for-authors/136-volume-51-issue-no-4/1171-the-influence-of-yeast-strains-on-the-composition-and-sensory-quality-of-gewuerztraminer-wine [accessed February 21, 2017].
Dashko, S., Zhou, N., Tinta, T., Sivilotti, P., Lemut, M. S., Trost, K., et al. (2015). Use of non-conventional yeast improves the wine aroma profile of Ribolla Gialla. J. Ind. Microbiol. Biotechnol. 42, 997–1010. doi: 10.1007/s10295-015-1620-y
Dequin, S. (2011). The potential of genetic engineering for improving brewing, wine-making and baking yeasts. Appl. Microbiol. Biotechnol. 56, 577–588. doi: 10.1007/s002530100700
Domizio, P., Romani, C., Lencioni, L., Comitini, F., Gobbi, M., Mannazzu, I., et al. (2011). Outlining a future for non-Saccharomyces yeasts: selection of putative spoilage wine strains to be used in association with Saccharomyces cerevisiae for grape juice fermentation. Int. J. Food Microbiol. 147, 170–180. doi: 10.1016/j.ijfoodmicro.2011.03.020
du Toit, M., Engelbrecht, L., Lerm, E., and Krieger-Weber, S. (2011). Lactobacillus: the next generation of malolactic fermentation starter cultures—an overview. Food Bioprocess Technol. 4, 876–906. doi: 10.1007/s11947-010-0448-8
Duarte, F. L., Pimentel, N. H., Teixeira, A., and Fonseca, A. (2012). Saccharomyces bacillaris is not a synonym of Candida stellata: reinstatement as Starmerella bacillaris comb. nov. Antonie Van Leeuwenhoek 102, 653–658. doi: 10.1007/s10482-012-9762-7
Englezos, V., Rantsiou, K., Cravero, F., Torchio, F., Ortiz-Julien, A., Gerbi, V., et al. (2016a). Starmerella bacillaris and Saccharomyces cerevisiae mixed fermentations to reduce ethanol content in wine. Appl. Microbiol. Biotechnol. 100, 5515–5526. doi: 10.1007/s00253-016-7413-z
Englezos, V., Torchio, F., Cravero, F., Marengo, F., Giacosa, S., Gerbi, V., et al. (2016b). Aroma profile and composition of Barbera wines obtained by mixed fermentations of Starmerella bacillaris (synonym Candida zemplinina) and Saccharomyces cerevisiae. LWT Food Sci. Technol. 73, 567–575. doi: 10.1016/j.lwt.2016.06.063
Erten, H., and Tanguler, H. (2010). Influence of Williopsis saturnus yeasts in combination with Saccharomyces cerevisiae on wine fermentation. Lett. Appl. Microbiol. 50, 474–479. doi: 10.1111/j.1472-765X.2010.02822.x
Ferraro, L., Fatichenti, F., and Ciani, M. (2000). Pilot scale vinification process using immobilized Candida stellata cells and Saccharomyces cerevisiae. Process Biochem. 35, 1125–1129. doi: 10.1016/S0032-9592(00)00148-5
Garavaglia, J., Schneider, R., de, C., de, S., Camargo Mendes, S. D., Welke, J. E., et al. (2015). Evaluation of Zygosaccharomyces bailii BCV 08 as a co-starter in wine fermentation for the improvement of ethyl esters production. Microbiol. Res. 173, 59–65. doi: 10.1016/j.micres.2015.02.002
Garcia, A., Carcel, C., Dulau, L., Samson, A., Aguera, E., Agosin, E., et al. (2002). Influence of a mixed culture with Debaryomyces vanriji and Saccharomyces cerevisiae on the volatiles of a muscat wine. J. Food Sci. 67, 1138–1143. doi: 10.1111/j.1365-2621.2002.tb09466.x
García-Ríos, E., López-Malo, M., and Guillamón, J. M. (2014). Global phenotypic and genomic comparison of two Saccharomyces cerevisiae wine strains reveals a novel role of the sulfur assimilation pathway in adaptation at low temperature fermentations. BMC Genomics 15:1059. doi: 10.1186/1471-2164-15-1059
Garofalo, C., El Khoury, M., Lucas, P., Bely, M., Russo, P., Spano, G., et al. (2015). Autochthonous starter cultures and indigenous grape variety for regional wine production. J. Appl. Microbiol. 118, 1395–1408. doi: 10.1111/jam.12789
Garofalo, C., Tristezza, M., Grieco, F., Spano, G., and Capozzi, V. (2016). From grape berries to wine: population dynamics of cultivable yeasts associated to “Nero di Troia” autochthonous grape cultivar. World J. Microbiol. Biotechnol. 32:59. doi: 10.1007/s11274-016-2017-4
Gobbi, M., Comitini, F., Domizio, P., Romani, C., Lencioni, L., Mannazzu, I., et al. (2013). Lachancea thermotolerans and Saccharomyces cerevisiae in simultaneous and sequential co-fermentation: a strategy to enhance acidity and improve the overall quality of wine. Food Microbiol. 33, 271–281. doi: 10.1016/j.fm.2012.10.004
Gonzalez-Ramos, D., Cebollero, E., and Gonzalez, R. (2008). A recombinant Saccharomyces cerevisiae strain overproducing mannoproteins stabilizes wine against protein haze. Appl. Environ. Microbiol. 74, 5533–5540. doi: 10.1128/AEM.00302-08
González-Royo, E., Pascual, O., Kontoudakis, N., Esteruelas, M., Esteve-Zarzoso, B., Mas, A., et al. (2015). Oenological consequences of sequential inoculation with non-Saccharomyces yeasts (Torulaspora delbrueckii or Metschnikowia pulcherrima) and Saccharomyces cerevisiae in base wine for sparkling wine production. Eur Food Res. Technol. 240, 999–1012. doi: 10.1007/s00217-014-2404-8
Guo, X. W., Li, Y. Z., Guo, J., Wang, Q., Huang, S. Y., Chen, Y. F., et al. (2016). Reduced production of ethyl carbamate for wine fermentation by deleting CAR1 in Saccharomyces cerevisiae. J. Ind. Microbiol. Biotechnol. 43, 671–679. doi: 10.1007/s10295-016-1737-7
Hirst, M. B., and Richter, C. L. (2016). Review of aroma formation through metabolic pathways of Saccharomyces cerevisiae in beverage fermentations. Am. J. Enol. Vitic. 67, 361–370. doi: 10.5344/ajev.2016.15098
Jolly, N. P., Augustyn, O. P. H., and Pretorius, L. S. (2006). The role and use of non-Saccharomyces yeasts in wine production. S. Afr. J. Enol. Vitic. 27, 15–39.
Jolly, N. P., Varela, C., and Pretorius, I. S. (2014). Not your ordinary yeast: non-Saccharomyces yeasts in wine production uncovered. FEMS Yeast Res. 14, 215–237. doi: 10.1111/1567-1364.12111
Karathia, H., Vilaprinyo, E., Sorribas, A., and Alves, R. (2011). Saccharomyces cerevisiae as a model organism: a comparative study. PLoS ONE 6:e16015. doi: 10.1371/journal.pone.0016015
Kim, D.-H., Hong, Y.-A., and Park, H.-D. (2008). Co-fermentation of grape must by Issatchenkia orientalis and Saccharomyces cerevisiae reduces the malic acid content in wine. Biotechnol. Lett. 30, 1633–1638. doi: 10.1007/s10529-008-9726-1
Lencioni, L., Romani, C., Gobbi, M., Comitini, F., Ciani, M., and Domizio, P. (2016). Controlled mixed fermentation at winery scale using Zygotorulaspora florentina and Saccharomyces cerevisiae. Int. J. Food Microbiol. 234, 36–44. doi: 10.1016/j.ijfoodmicro.2016.06.004
Lerm, E., Engelbrecht, L., and Du Toit, M. (2011). Selection and Characterisation of Oenococcus oeni and Lactobacillus plantarum South African Wine Isolates for Use as Malolactic Fermentation Starter Cultures. Available at: http://scholar.sun.ac.za/handle/10019.1/20651 [accessed February 21, 2017].
Liu, J., Arneborg, N., Toldam-Andersen, T. B., Zhang, S., Petersen, M. A., and Bredie, W. L. P. (2017). Impact of sequential co-culture fermentations on flavour characters of Solaris wines. Eur. Food Res. Technol. 243, 437–445. doi: 10.1007/s00217-016-2757-2
Liu, Y., Forcisi, S., Harir, M., Deleris-Bou, M., Krieger-Weber, S., Lucio, M., et al. (2016). New molecular evidence of wine yeast-bacteria interaction unraveled by non-targeted exometabolomic profiling. Metabolomics 12:69. doi: 10.1007/s11306-016-1001-1
Liu, Y., Rousseaux, S., Tourdot-Maréchal, R., Sadoudi, M., Gougeon, R., Schmitt-Kopplin, P., et al. (2017). Wine microbiome: a dynamic world of microbial interactions. Crit. Rev. Food Sci. Nutr. 57, 856–873. doi: 10.1080/10408398.2014.983591
Lleixà, J., Martín, V., Portillo, M. D. C., Carrau, F., Beltran, G., and Mas, A. (2016). Comparison of fermentation and wines produced by inoculation of Hanseniaspora vineae and Saccharomyces cerevisiae. Front. Microbiol. 7:338. doi: 10.3389/fmicb.2016.00338
Loira, I., Morata, A., Comuzzo, P., Callejo, M. J., González, C., Calderón, F., et al. (2015). Use of Schizosaccharomyces pombe and Torulaspora delbrueckii strains in mixed and sequential fermentations to improve red wine sensory quality. Food Res. Int. 76, 325–333. doi: 10.1016/j.foodres.2015.06.030
Martiniuk, J. T., Pacheco, B., Russell, G., Tong, S., Backstrom, I., and Measday, V. (2016). Impact of commercial strain use on Saccharomyces cerevisiae population structure and dynamics in pinot noir vineyards and spontaneous fermentations of a Canadian winery. PLoS ONE 11:e0160259. doi: 10.1371/journal.pone.0160259
Masneuf-Pomarède, I., Bely, M., Marullo, P., Lonvaud-Funel, A., and Dubourdieu, D. (2010). Reassessment of phenotypic traits for Saccharomyces bayanus var. uvarum wine yeast strains. Int. J. Food Microbiol. 139, 79–86. doi: 10.1016/j.ijfoodmicro.2010.01.038
Masneuf-Pomarede, I., Juquin, E., Miot-Sertier, C., Renault, P., Laizet, Y., Salin, F., et al. (2015). The yeast Starmerella bacillaris (synonym Candida zemplinina) shows high genetic diversity in winemaking environments. FEMS Yeast Res. 15:fov045. doi: 10.1093/femsyr/fov045
Maturano, Y. P., Assof, M., Fabani, M. P., Nally, M. C., Jofré, V., Rodríguez Assaf, L. A., et al. (2015). Enzymatic activities produced by mixed Saccharomyces and non-Saccharomyces cultures: relationship with wine volatile composition. Antonie Van Leeuwenhoek 108, 1239–1256. doi: 10.1007/s10482-015-0578-0
Maturano, Y. P., Rodríguez Assaf, L. A., Toro, M. E., Nally, M. C., Vallejo, M., Castellanos de Figueroa, L. I., et al. (2012). Multi-enzyme production by pure and mixed cultures of Saccharomyces and non-Saccharomyces yeasts during wine fermentation. Int. J. Food Microbiol. 155, 43–50. doi: 10.1016/j.ijfoodmicro.2012.01.015
Medina, K., Boido, E., Fariña, L., Gioia, O., Gomez, M. E., Barquet, M., et al. (2013). Increased flavour diversity of Chardonnay wines by spontaneous fermentation and co-fermentation with Hanseniaspora vineae. Food Chem. 141, 2513–2521. doi: 10.1016/j.foodchem.2013.04.056
Mehlomakulu, N. N., Setati, M. E., and Divol, B. (2014). Characterization of novel killer toxins secreted by wine- related non-Saccharomyces yeasts and their action on Brettanomyces spp. Int. J. Food Microbiol. 188, 83–91. doi: 10.1016/j.ijfoodmicro.2014.07.015
Mills, D. A., Johannsen, E. A., and Cocolin, L. (2002). Yeast diversity and persistence in botrytis-affected wine fermentations. Appl. Environ. Microbiol. 68, 4884–4893. doi: 10.1128/AEM.68.10.4884-4893.2002
Moreira, N., Pina, C., Mendes, F., Couto, J. A., Hogg, T., and Vasconcelos, I. (2011). Volatile compounds contribution of Hanseniaspora guilliermondii and Hanseniaspora uvarum during red wine vinifications. Food Control 22, 662–667. doi: 10.1016/j.foodcont.2010.07.025
Mylona, A. E., Del Fresno, J. M., Palomero, F., Loira, I., Bañuelos, M. A., Morata, A., et al. (2016). Use of Schizosaccharomyces strains for wine fermentation-Effect on the wine composition and food safety. Int. J. Food Microbiol. 232, 63–72. doi: 10.1016/j.ijfoodmicro.2016.05.023
Padilla, B., Gil, J. V., and Manzanares, P. (2016). Past and future of non-Saccharomyces yeasts: from spoilage microorganisms to biotechnological tools for improving wine aroma complexity. Front. Microbiol. 7:411. doi: 10.3389/fmicb.2016.00411
Pérez-Través, L., Querol, A., and Pérez-Torrado, R. (2016). Increased mannoprotein content in wines produced by Saccharomyces kudriavzevii × Saccharomyces cerevisiae hybrids. Int. J. Food Microbiol. 237, 35–38. doi: 10.1016/j.ijfoodmicro.2016.08.014
Petruzzi, L., Bevilacqua, A., Corbo, M. R., Speranza, B., Capozzi, V., and Sinigaglia, M. (2017). A Focus on quality and safety traits of Saccharomyces cerevisiae isolated from uva di troia grape variety. J. Food Sci. 82, 124–133. doi: 10.1111/1750-3841.13549
Rantsiou, K., Dolci, P., Giacosa, S., Torchio, F., Tofalo, R., Torriani, S., et al. (2012). Candida zemplinina can reduce acetic acid produced by Saccharomyces cerevisiae in sweet wine fermentations. Appl. Enviro. Microbiol. 78, 1987–1994. doi: 10.1128/AEM.06768-11
Renault, P., Coulon, J., de Revel, G., Barbe, J.-C., and Bely, M. (2015). Increase of fruity aroma during mixed T. delbrueckii/S. cerevisiae wine fermentation is linked to specific esters enhancement. Int. J. Food Microbiol. 207, 40–48. doi: 10.1016/j.ijfoodmicro.2015.04.037
Renault, P., Miot-Sertier, C., Marullo, P., Hernández-Orte, P., Lagarrigue, L., Lonvaud-Funel, A., et al. (2009). Genetic characterization and phenotypic variability in Torulaspora delbrueckii species: potential applications in the wine industry. Int. J. Food Microbiol. 134, 201–210. doi: 10.1016/j.ijfoodmicro.2009.06.008
Romano, P., and Capece, A. (2017). “Wine microbiology,” in Starter Cultures in Food Production, eds B. Speranza, A. Bevilacqua, M. R. Corbo, and M. Sinigaglia (Hoboken, NY: John Wiley & Sons), 255–282. doi: 10.1002/9781118933794.ch13
Romano, P., Pietrafesa, R., Romaniello, R., Zambuto, M., Calabretti, A., and Capece, A. (2015). Impact of yeast starter formulations on the production of volatile compounds during wine fermentation. Yeast 32, 245–256. doi: 10.1002/yea.3034
Rossouw, D., and Bauer, F. F. (2016). Exploring the phenotypic space of non-Saccharomyces wine yeast biodiversity. Food Microbiol. 55, 32–46. doi: 10.1016/j.fm.2015.11.017
Russo, P., Capozzi, V., Spano, G., Corbo, M. R., Sinigaglia, M., and Bevilacqua, A. (2016). Metabolites of microbial origin with an impact on health: ochratoxin a and biogenic amines. Front. Microbiol. 7:482. doi: 10.3389/fmicb.2016.00482
Russo, P., Spano, G., and Capozzi, V. (2017). “Safety evaluation of starter cultures,” in Starter Cultures in Food Production, eds B. Speranza, A. Bevilacqua, M. R. Corbo, and M. Sinigaglia (Hoboken, NY: John Wiley & Sons), 101–128. doi: 10.1002/9781118933794.ch6
Sabel, A., Martens, S., Petri, A., König, H., and Claus, H. (2014). Wickerhamomyces anomalus AS1: a new strain with potential to improve wine aroma. Ann. Microbiol. 64, 483–491. doi: 10.1007/s13213-013-0678-x
Sadoudi, M., Tourdot-Maréchal, R., Rousseaux, S., Steyer, D., Gallardo-Chacón, J.-J., Ballester, J., et al. (2012). Yeast–yeast interactions revealed by aromatic profile analysis of Sauvignon Blanc wine fermented by single or co-culture of non-Saccharomyces and Saccharomyces yeasts. Food Microbiol. 32, 243–253. doi: 10.1016/j.fm.2012.06.006
Salvetti, E., Campanaro, S., Campedelli, I., Fracchetti, F., Gobbi, A., Tornielli, G. B., et al. (2016). Whole-metagenome-sequencing-based community profiles of Vitis vinifera L. cv. Corvina berries withered in two post-harvest conditions. Front. Microbiol. 7:937. doi: 10.3389/fmicb.2016.00937
Soden, A., Francis, I. L., Oakey, H., and Henschke, P. A. (2000). Effect of co-fermentation with Candida stellata and Saccharomyces cerevisiae on the aroma and composition of Chardonnay wine. Aust. J. Grape Wine Res. 6, 21–30. doi: 10.1111/j.1755-0238.2000.tb00158.x
Spano, G., Russo, P., Lonvaud-Funel, A., Lucas, P., Alexandre, H., Grandvalet, C., et al. (2010). Biogenic amines in fermented foods. Eur. J. Clin. Nutr. 64(Suppl. 3), S95–S100. doi: 10.1038/ejcn.2010.218
Steensels, J., and Verstrepen, K. J. (2014). Taming wild yeast: potential of conventional and nonconventional yeasts in industrial fermentations. Annu. Rev. Microbiol. 68, 61–80. doi: 10.1146/annurev-micro-091213-113025
Sumby, K. M., Grbin, P. R., and Jiranek, V. (2014). Implications of new research and technologies for malolactic fermentation in wine. Appl. Microbiol. Biotechnol. 98, 8111–8132. doi: 10.1007/s00253-014-5976-0
Sun, S. Y., Gong, H. S., Liu, W. L., and Jin, C. W. (2016). Application and validation of autochthonous Lactobacillus plantarum starter cultures for controlled malolactic fermentation and its influence on the aromatic profile of cherry wines. Food Microbiol. 55, 16–24. doi: 10.1016/j.fm.2015.11.016
Sun, X. Y., Zhao, Y., Liu, L. L., Jia, B., Zhao, F., Huang, W. D., et al. (2015). Copper tolerance and biosorption of Saccharomyces cerevisiae during alcoholic fermentation. PLoS ONE 10:e0128611. doi: 10.1371/journal.pone.0128611
Swiegers, J. H., Kievit, R. L., Siebert, T., Lattey, K. A., Bramley, B. R., Francis, I. L., et al. (2009). The influence of yeast on the aroma of Sauvignon Blanc wine. Food Microbiol. 26, 204–211. doi: 10.1016/j.fm.2008.08.004
Taillandier, P., Lai, Q. P., Julien-Ortiz, A., and Brandam, C. (2014). Interactions between Torulaspora delbrueckii and Saccharomyces cerevisiae in wine fermentation: influence of inoculation and nitrogen content. World J. Microbiol. Biotechnol. 30, 1959–1967. doi: 10.1007/s11274-014-1618-z
Tofalo, R., Patrignani, F., Lanciotti, R., Perpetuini, G., Schirone, M., Di Gianvito, P., et al. (2016). Aroma profile of Montepulciano d’Abruzzo wine fermented by single and co-culture starters of autochthonous Saccharomyces and non-saccharomyces yeasts. Front. Microbiol. 7:610. doi: 10.3389/fmicb.2016.00610
Tofalo, R., Schirone, M., Torriani, S., Rantsiou, K., Cocolin, L., Perpetuini, G., et al. (2012). Diversity of Candida zemplinina strains from grapes and Italian wines. Food Microbiol. 29, 18–26. doi: 10.1016/j.fm.2011.08.014
Toro, M. E., and Vazquez, F. (2002). Fermentation behaviour of controlled mixed and sequential cultures of Candida cantarellii and Saccharomyces cerevisiae wine yeasts. World J. Microbiol. Biotechnol. 18, 351–358. doi: 10.1023/A:1015242818473
Tristezza, M., Tufariello, M., Capozzi, V., Spano, G., Mita, G., and Grieco, F. (2016). The oenological potential of Hanseniaspora uvarum in simultaneous and sequential co-fermentation with Saccharomyces cerevisiae for industrial wine production. Front. Microbiol. 7:670. doi: 10.3389/fmicb.2016.00670
Tronchoni, J., Curiel, J. A., Morales, P., Torres-Pérez, R., and Gonzalez, R. (2017). Early transcriptional response to biotic stress in mixed starter fermentations involving Saccharomyces cerevisiae and Torulaspora delbrueckii. Int. J. Food Microbiol. 241, 60–68. doi: 10.1016/j.ijfoodmicro.2016.10.017
Varela, C., Kutyna, D. R., Solomon, M. R., Black, C. A., Borneman, A., and Henschke, P. A. (2012). Evaluation of gene modification strategies for the development of low-alcohol-wine yeasts. Appl. Environ. Microbiol. 78, 6068–6077. doi: 10.1128/AEM.01279-12
Varela, C., Sengler, F., Solomon, M., and Curtin, C. (2016). Volatile flavour profile of reduced alcohol wines fermented with the non-conventional yeast species Metschnikowia pulcherrima and Saccharomyces uvarum. Food Chem. 209, 57–64. doi: 10.1016/j.foodchem.2016.04.024
Viana, F., Gil, J. V., Genovés, S., Vallés, S., and Manzanares, P. (2008). Rational selection of non-Saccharomyces wine yeasts for mixed starters based on ester formation and enological traits. Food Microbiol. 25, 778–785. doi: 10.1016/j.fm.2008.04.015
Viana, F., Gil, J. V., Vallés, S., and Manzanares, P. (2009). Increasing the levels of 2-phenylethyl acetate in wine through the use of a mixed culture of Hanseniaspora osmophila and Saccharomyces cerevisiae. Int. J. Food Microbiol. 135, 68–74. doi: 10.1016/j.ijfoodmicro.2009.07.025
Whitener, M. E. B. (2016). Metabolomic Profiling of Non-Saccharomyces Yeasts in Wine. Ph.D, thesis, Stellenbosch University, Stellenbosch.
Whitener, M. E. B., Carlin, S., Jacobson, D., Weighill, D., Divol, B., Conterno, L., et al. (2015). Early fermentation volatile metabolite profile of non-Saccharomyces yeasts in red and white grape must: a targeted approach. LWT Food Sci. Technol. 64, 412–422. doi: 10.1016/j.lwt.2015.05.018
Whitener, M. E. B., Stanstrup, J., Panzeri, V., Carlin, S., Divol, B., Toit, M. D., et al. (2016). Untangling the wine metabolome by combining untargeted SPME–GCxGC-TOF-MS and sensory analysis to profile Sauvignon blanc co-fermented with seven different yeasts. Metabolomics 12:53. doi: 10.1007/s11306-016-0962-4
Wrent, P., Rivas, E. M., Gil de Prado, E., Peinado, J. M., and de Silóniz, M. I. (2014). “Debaryomyces,” in Encyclopedia of Food Microbiology, 2nd Edn, eds C. Batt and C. A. Batt, (Amsterdam: Elsevier), 563–570. doi: 10.1016/B978-0-12-384730-0.00081-1
Zara, G., Mannazzu, I., Del Caro, A., Budroni, M., Pinna, M. B., Murru, M., et al. (2014). Wine quality improvement through the combined utilisation of yeast hulls and Candida zemplinina/Saccharomyces cerevisiae mixed starter cultures. Aust. J. Grape Wine Res. 20, 199–207. doi: 10.1111/ajgw.12078
Zott, K., Thibon, C., Bely, M., Lonvaud-Funel, A., Dubourdieu, D., and Masneuf-Pomarede, I. (2011). The grape must non-Saccharomyces microbial community: impact on volatile thiol release. Int. J. Food Microbiol. 151, 210–215. doi: 10.1016/j.ijfoodmicro.2011.08.026
Keywords: wine, yeasts, non-Saccharomyces, lactic acid bacteria, microbial resources, starter cultures, alcoholic fermentation, malolactic fermentation
Citation: Petruzzi L, Capozzi V, Berbegal C, Corbo MR, Bevilacqua A, Spano G and Sinigaglia M (2017) Microbial Resources and Enological Significance: Opportunities and Benefits. Front. Microbiol. 8:995. doi: 10.3389/fmicb.2017.00995
Received: 22 February 2017; Accepted: 17 May 2017;
Published: 08 June 2017.
Edited by:
Rosalba Lanciotti, Università di Bologna, ItalyReviewed by:
Sofía M. Arvizu-Medrano, Universidad Autónoma de Querétaro, MexicoChrysoula C. Tassou, Hellenic Agricultural Organisation DEMETER, Greece
Copyright © 2017 Petruzzi, Capozzi, Berbegal, Corbo, Bevilacqua, Spano and Sinigaglia. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Giuseppe Spano, Z2l1c2VwcGUuc3Bhbm9AdW5pZmcuaXQ=
 Leonardo Petruzzi
Leonardo Petruzzi Vittorio Capozzi
Vittorio Capozzi Carmen Berbegal
Carmen Berbegal Maria R. Corbo
Maria R. Corbo Antonio Bevilacqua
Antonio Bevilacqua Giuseppe Spano
Giuseppe Spano Milena Sinigaglia
Milena Sinigaglia