- 1Integrative Microbiology Research Centre, Guangdong Province Key Laboratory of Microbial Signals and Disease Control, State Key Laboratory for Conservation and Utilization of Subtropical Agro-Bioresources, South China Agricultural University, Guangzhou, China
- 2School of Biological and Science Technology, University of Jinan, Jinan, China
- 3Guangdong Innovative and Entrepreneurial Research Team of Sociomicrobiology Basic Science and Frontier Technology, College of Agriculture, South China Agricultural University, Guangzhou, China
- 4Institute of Molecular and Cell Biology, Singapore, Singapore
Ralstonia solanacearum is a ubiquitous soil-borne plant pathogenic bacterium, which frequently encounters and interacts with other soil cohabitants in competition for environmental niches. Ralsolamycin, which is encoded by the rmy genes, has been characterized as a novel inter-kingdom interaction signal that induces chlamydospore development in fungi. In this study, we provide the first genetic evidence that the rmy gene expression is controlled by the PhcBSR quorum sensing (QS) system in strain GMI1000. Mutation of phcB could lead to significant reduction of the expression levels of the genes involved in ralsolamycin biosynthesis. In addition, both the phcB and rmy mutants were attenuated in induction of chlamydospore formation in Fusarium oxysporum f. cubense and diminished in the ability to compete with the sugarcane pathogen Sporisorium scitamineum. Agreeable with the pattern of QS regulation, transcriptional expression analysis showed that the transcripts of the rmy genes were increased along with the increment of the bacterial population density. Taken together, the above findings provide new insights into the regulatory mechanisms that the QS system involves in governing the ralsolamycin inter-kingdom signaling system.
Introduction
Ralstonia solanacearum is a notorious soil-borne pathogen that causes lethal bacterial wilt of many plants around the world (Hayward, 1991). The pathogen encodes numerous virulence determinants, including extracellular polysaccharide (EPS), cell wall-degrading enzymes (CWDE), chemotaxis system, and secretion systems, which are collectively contribute to its virulence (Denny and Ryel, 1991; Arlat et al., 1992; Yao and Allen, 2006; Genin and Denny, 2012). Previous studies have outlined the sophisticated regulatory mechanisms that control the production of virulence factors in R. solanacearum. Among them, PhcA is a LysR family transcriptional regulator (Brumbley et al., 1993), which is located at the center of the complex regulatory network, and can directly or indirectly regulate the genes involved in production of EPS and other virulence factors (Huang et al., 1995). Along with bacterial proliferation, PhcA activity is regulated by accumulated quorum sensing (QS) signal 3-hydroxypalmitic acid methyl ester (3-OH-PAME) or (R)-methyl 3-hydroxymyristate (3-OH-MAME), which is encoded by phcB (Flavier et al., 1997; Kai et al., 2015). Consequently, PhcA directs the production of EPS, CWDE, and other virulence factors in a population density dependent manner. Evidence indicates that a two component system, encoded by phcS and phcR in the same operon as phcB, is involved in detection and response to the QS signal 3-OH-PAME (Clough et al., 1997).
Ralstonia solanacearum species complex is well known not only for their ability to infect a unusually broad range of host plants, but also for their wide geographic distributions and capability to live and compete for versatile and diverse habitats (Salanoubat et al., 2002; Alvarez et al., 2010). Involvement of secondary metabolites in interspecies and inter-kingdom signaling and interference between R. solanacearum and the other organisms in competition for environmental niches has recently attracted much attention. Genome sequencing analysis and genetic studies show that R. solanacearum complex has the potential to produce an array of secondary metabolites. For example, ralfuranone is known to contribute to the virulence of R. solanacearum strain OE1-1 (Kai et al., 2014); staphyloferrin B is a siderophore associated with iron scavenge in strain AW1 (Bhatt and Denny, 2004); the yersiniabactin-like siderophore micacocidin was identified as an anti-mycoplasma agent (Kreutzer et al., 2011). More recently, R. solanacearum was reported to produce ralsolamycin as an inter-kingdom signal to communicate with fungal organisms and consequently induce conserved morphological differentiation, i.e., formation of chlamydospores that are survival structures with thickened cell walls, in 34 species of fungi belonging to three taxa (Spraker et al., 2016). Ralsolamycin is produced by the non-ribosomal peptide synthetase-polyketide synthase hybrid RmyA and RmyB, which also facilitates the bacterial pathogen entry into fungal tissues (Spraker et al., 2016). It is not yet clear how the production of ralsolamycin is regulated in R. solanacearum.
RmyA and RmyB are the homologs of AmbB and AmbE, respectively, which we identified previously are responsible for production of IQS, an integrative QS signal associated with virulence regulation in Pseudomonas aeruginosa (Lee et al., 2013). Production of IQS is controlled not only by the central las QS system, but also influenced by phosphate depletion, which is a host stress signal commonly encountered by invading pathogens (Lee et al., 2013; Lee and Zhang, 2015). In this study, initiated by investigating the role of rmyAB genes in R. solanacearum, we found that the expression of rmyAB was controlled by the PhcBSR QS system. Deletion of phcB resulted in dramatic decreases in transcriptional expression of rmy genes and ralsolamycin transportation related genes, and weakened the bacterial ability to induce formation of chlamydospores in soil-borne phytopathogens Fusarium oxysporum f. cubense (FOC) strain XJZ2 (Li et al., 2014), and lost the antimicrobial activity to inhibit the growth of Sporisorium scitamineum.
Materials and Methods
Bacterial and Fungal Strains, Plasmids, and Media
The plasmids and R. solanacearum strains used in this study are listed in Table 1. Escherichia coli strain DH5α (Invitrogen, Carlsbad, CA, United States) was used as a host in gene cloning and vector construction. R. solanacearum strains were maintained at 30°C in casamino acid-peptone-glucose (CPG) plates (Hendrick and Sequeira, 1984), and cultured in CPG broth for testing CWDE activities (Yap et al., 2005), or on minimal medium (MM) agar plate for screening transformants after tri-parental mating (Hussain et al., 2008). E. coli was grown at 37°C in LB medium. Antibiotics were added at the following final concentrations (μg/L): kanamycin, Km (50), gentamicin, Gm (50), and rifampicin, Rif (50). The fungal strains used in this study were maintained in PDA medium unless otherwise stated.
In-Frame Deletion and Complementation
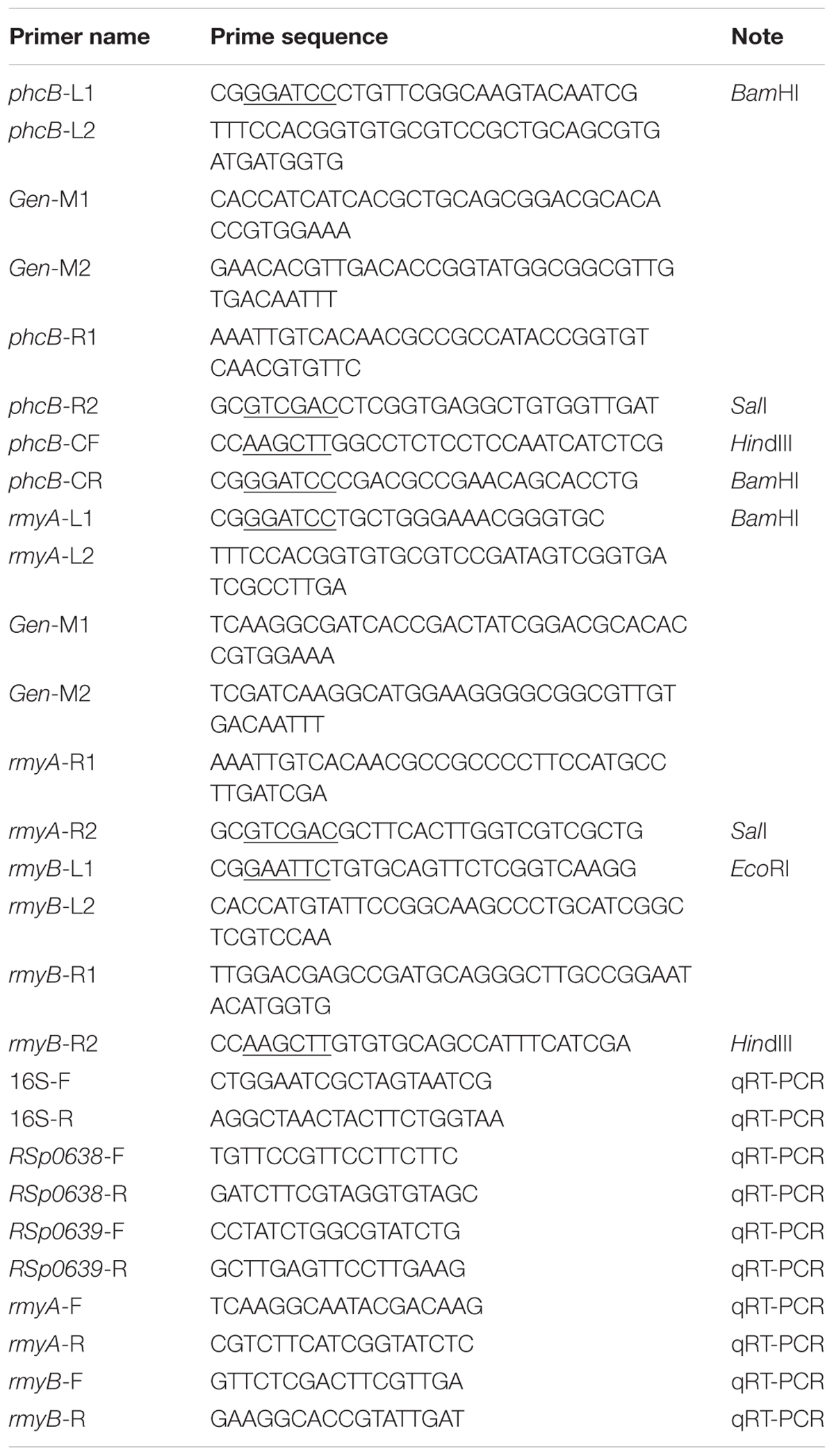
The phcB, rmyA, rmyB, and rmyAB deletion mutants were generated by amplifying two DNA fragments flanking their coding sequences (using primers as described in Table 2), the 859 bp gentamicin resistance gene sequence (Gen) was added between the left and right fragments of phcB or rmyA by using the overlap primers, the left and right DNA fragments of rmyB were ligated without adding the Gen sequence. The two/three fragments were fused using the primer pair L1/R2. All the PCR procedure of this study were amplification using the high fidelity Taq polymerase (PrimeSTAR® HS DNA Polymerase, Takara Bio Inc., Dalian, China), cycling conditions were set as follows: initial denaturation at 98°C for 1 min, followed by 34 cycles of denaturation at 98°C for 15 s, annealing at 60°C for 20 s, extension at 72°C for 30 s, and a final extension at 72°C for 5 min. The fusion fragments and the suicide plasmid pK18mobsacB (ATCC®87097TM) (Schäfer et al., 1994) were digested, respectively, with corresponding restriction enzymes as indicated, followed by purification by using Sangon purification kit (Sangon, Shanghai, China), and then ligated together by using T4 ligase [NEW ENGLAND BioLabs (Beijing) Ltd., China]. The ligation products were transformed into E. coli DH5α competent cells (Life Technologies Corporation, Beijing, China), and the bacterial cells were cultured at 37°C with shaking for 1 h. The transformants were selected in LB medium supplemented with Km and verified by PCR analysis. The plasmid constructs were introduced into R. solanacearum by using tri-parental mating as described previously to generate corresponding in-frame deletion mutants (Ditta et al., 1980). Mutants were selected on CPG plate containing 10% sucrose, antibiotics Gm and Rif, and then confirmed by PCR and DNA sequencing. To construct complementary strains, the DNA fragment containing 235 bp promoter sequence and ORF of phcB was amplified using primers phcB-CF and phcB-CR (Table 2). The purified PCR products were digested by required restriction enzymes, and then purified again prior to ligation with the same enzymes digested expression vector pBBR1MCS2 (Kovach et al., 1995). The ligation products were transformed into E. coli DH5α competent cells, and the transformants were selected on LB plate containing Km. Tri-parental mating was performed as above description, and the complemented strains were selected on MM plates supplemented with Km and Rif, and confirmed by PCR product analysis. The gel electrophoresis results of the mutants and control are provided in Supplementary Information File 1 (SI 1).
Co-culture Experiments and Microscopy Examination of Chlamydospore Formation
Preparation of conidial suspensions of the FOC strain XJZ2 was performed as described (Spraker et al., 2016). For co-culture experiments, R. solanacearum strain GMI1000 and the fungal pathogen were plated as previously described and incubated for 8 days at 28°C. When the colonies met each other, the fungi mycelia of the interaction zone were harvested and examined for FOC chlamydospore formation by using ZEISS Observer Z1 microscopy. For testing bacterial interaction with S. scitamineum, the MAT-1 and MAT-2 (Yan et al., 2016) mating mixture of S. scitamineum were grown on PDA plates for observation of hypha growth. To observe the dose effect of strain GMI1000 on inhibition of S. scitamineum, the GMI1000 (A) was cultured 24 h on the CPG plate before adding GMI1000 (B), other GMI1000 derivatives, and the MAT-1/MAT-2 mating mixture of S. scitamineum on the CPG plate.
RNA Preparation and Quantitative Real-Time PCR (qRT-PCR) Analysis
Cultures of R. solanacearum strains in CPG broth (OD600 = 1.5) were centrifuged and total RNA were isolated by using RNeasy Mini Kit (QIAGEN, Hilden, Germany). The contaminated genomic DNA were removed by treating with DNaseI (Takara, Dalian, China) at 37°C for 1 h, and was confirmed by PCR using the 16S primer pair and visualized on an agarose gel. The qRT-PCR experiments were performed with SuperReal PreMix Color SYBR Green, 2X, (TIANGEN BIOTECH CO. LTD, Beijing, China) on QuantStudio 6 Flex (applied biosystems by life technologies, Carlsbad, CA, United States) following the user’s guide from the manufacturer. The PCR conditions were as follows: 95°C for 15 min, followed by 40 cycles of 95°C for 15 s and 55°C for 31 s, and then the melting curve analysis was carried out to determine the specificity of PCR products. The cDNA samples of each treatment were replicated three times, and the absolute value of –ΔΔCt = –(ΔCt1 – ΔCt2) were calculated as described in the 2-ΔΔCt method (Livak and Schmittgen, 2001).
LC/MS Analysis of Ralsolamycin
To prepare ralsolamycin extracts, bacteria were inoculated in 1 litre CPG broth and cultured at 28°C for 24 h with shaking at 150 rpm (OD600 = 1.5). Bacterial supernatants were collected by centrifugation at 10,000 rpm for 10 min, mixed with an equal volume of ethyl acetate with shaking. The upper organic phase was collected and air dried, and the residues were dissolved in methanol for further analysis. LC/MS analysis was conducted using a Waters LC-MS system (Waters, MA, United States) with an ACQUITY UPLC system coupled to the Waters Q-Tof Premier high resolution mass spectrometer. An ACQUITY UPLC BEH C18 column (2.1 mm × 50 mm) was used for chromatography analysis. Solution A was composed of 0.01% formic acid in water; solution B was composed of 0.01% formic acid in CH3OH. A linear gradient elution from 10 to 100% B over 10 min at 0.4 ml/min, 100% B over 3 min and re-equilibrated with 10% B for an additional 3 min. The injection volume was 1 μl. The entire column elute was introduced into the Q-Tof mass spectrometer. Ion detection was achieved in ESI mode using a source capillary voltage of 1.5 kV, source temperature of 120°C, desolvation temperature of 350°C, cone gas flow of 50 L/h (N2), and desolvation gas flow of 600 L/h (N2). Presence of the peak in the bacterial extract showing the accurate mass of ralsolamycin (m/z 1291.7142) was identified, and the corresponding peak areas from wild-type and mutant were compared and calculated.
Results
Expression of the rmy Genes and Production of Ralsolamycin Are Controlled by the PhcBSR QS System
To evaluate the role of the PhcBSR QS system in regulation of the rmy genes expression and ralsolamycin production, the QS signal synthase gene phcB deletion mutant was constructed in the genetic background of R. solanacearum strain GMI1000 by deletion of its coding sequence. The resulting strain ΔphcB was non-mucoid and nearly avirulent with weak cellulose activity, but was highly motile and with an increased polygalacturonase activity as previously reported (Flavier et al., 1997). The expression levels of rmyA and rmyB in ΔphcB were determined and compared to its parental strain GMI1000. In the ΔphcB, the rmyA transcription was drastically decreased by about 24-fold (Welch t-test, p-value < 3.64e-4; Figure 1), and the gene rmyB transcription was decreased by around 4.6-fold (Welch t-test, p-value < 3.02e-3; Figure 1). Meanwhile, deletion of phcB also resulted in decreasing the expression level of RSp0638 and RSp0639 by approximately 6- and 7-fold, respectively (Welch t-test, p-value < 2.32e-3 and 2.08e-3, respectively; Figure 1), which are the two neighbor genes of the rmy operon. In trans expression of a wild-type phcB gene in ΔphcB restored their expression to the wild-type levels. Consistent with the results of transcriptional assay, LC-MS analysis of the ethyl acetate extracts from wide-type strain GMI1000 and ΔphcB showed that ralsolamycin production in ΔphcB was decreased by about 81-fold compared with the wide-type strain (Welch t-test, p-value < 6.86e-5; Figure 2).
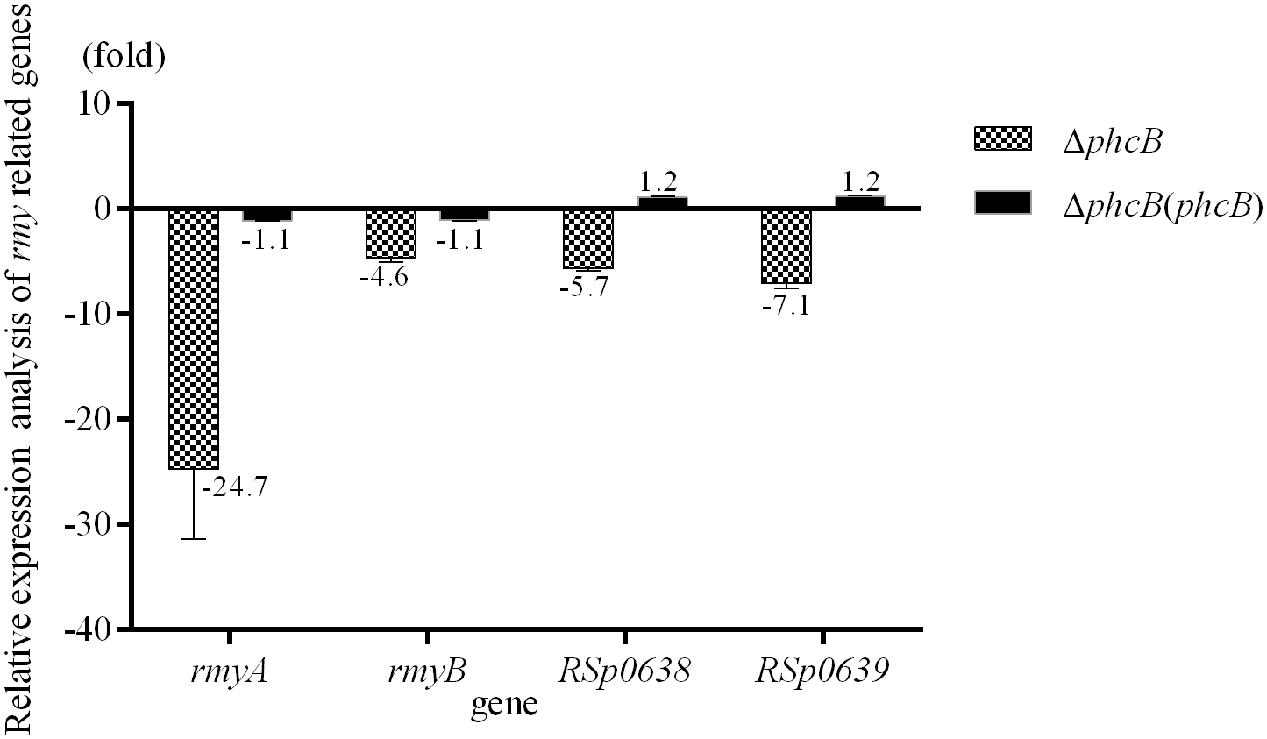
FIGURE 1. Quantitative Real-Time PCR (qRT-PCR) analysis of rmy clusters in phcB deletion mutant of strain GMI1000. The results were calculated as –ΔΔCt = –(ΔCtmutant – ΔCtwt). Three biological repeats (independent cultures) and three technical repeats were done to calculate and compare the values, all statistics were presented as means ± SE.
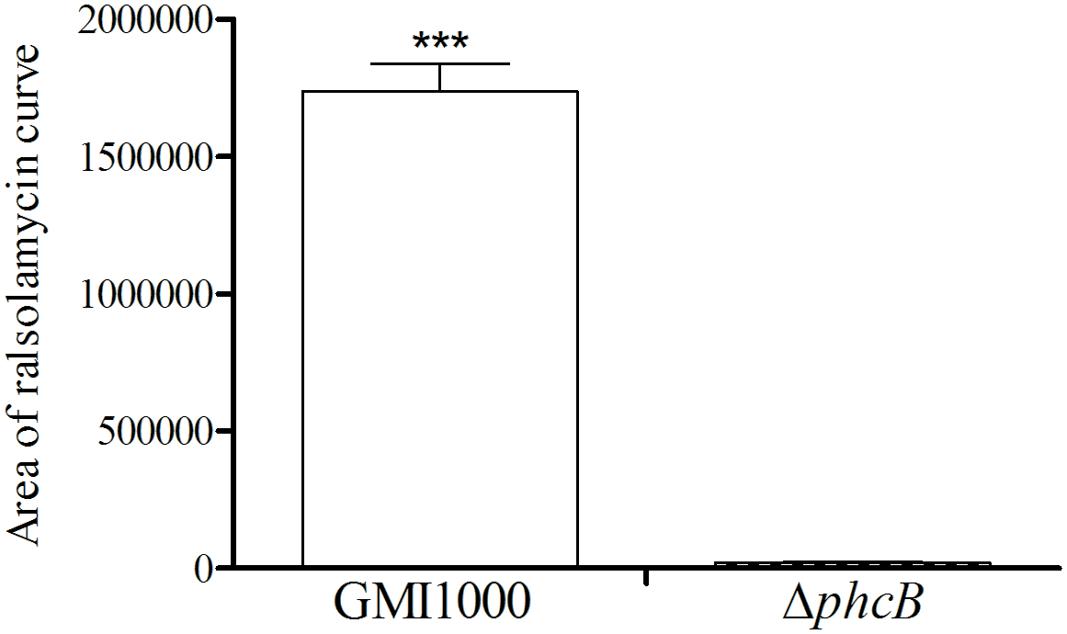
FIGURE 2. LC-MS analysis of ralsolamycin in the ethyl acetate extracts of the wide-type strain GMI1000 and phcB deletion mutant supernatant. The bacterial growth (under shaking) has been followed during 24 h by measurement of OD600 (1.5). The p-values from the Welch t-test comparing the production of wide type strain GMI1000 with the production of the ΔphcB strain (∗∗∗ p-value < 0.001).
Cell Density Affects the Expression Level of the rmy Genes
Given that bacterial QS systems modulate target gene expression in a population density dependent manner, the transcriptional levels of rmyA and rmyB in strain GMI1000 was then determined at different growth stages of bacterial cells. The results showed that the transcripts level of rmyA and rmyB in strain GMI1000 at a high cell density (∼108 CFU/ml) was increased by about 2.3- and 4.8-fold, respectively (Welch t-test, p-value < 3.22e-3 and 2.18e-3, respectively; Figure 3), compared with those at a low cell density (∼106 CFU/ml).
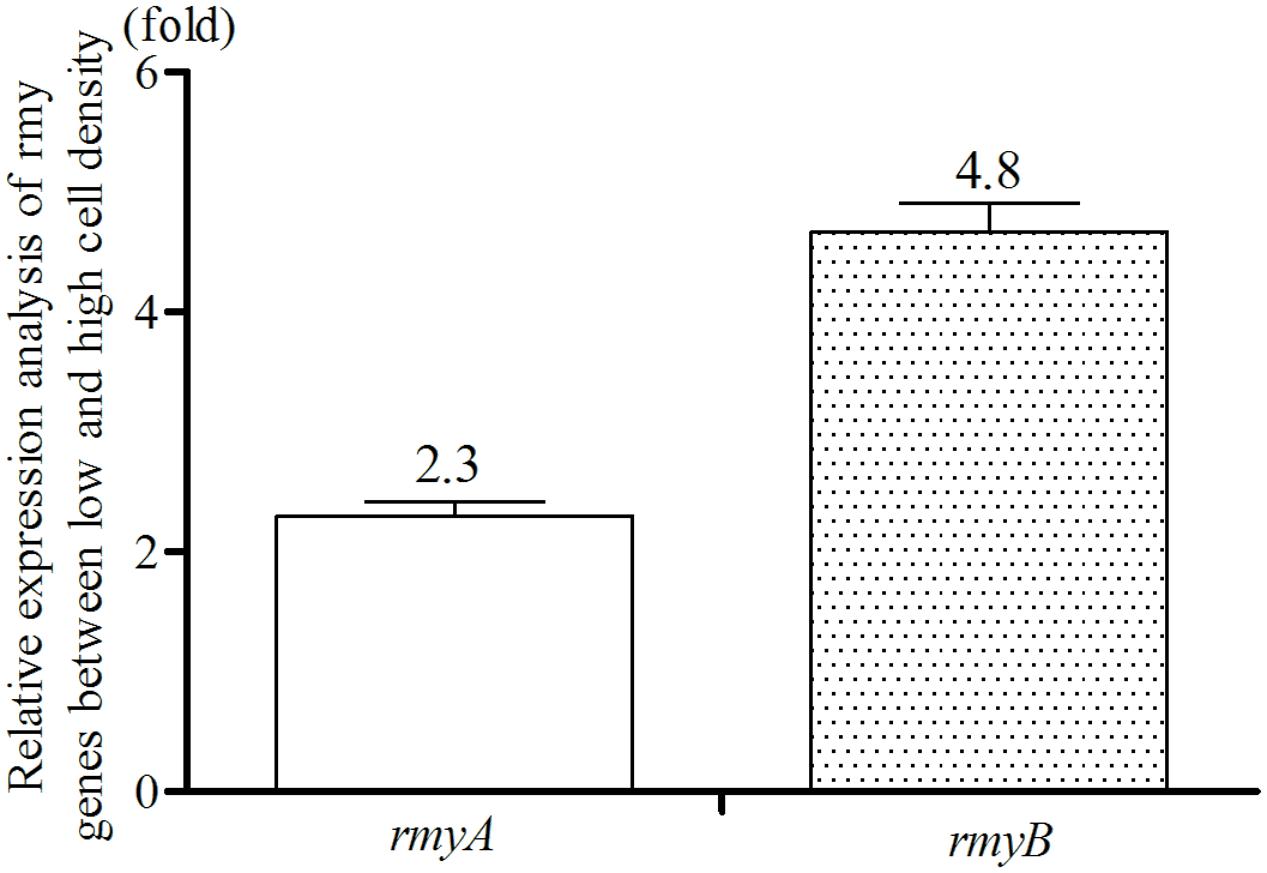
FIGURE 3. Quantitative Real-Time PCR analyses of rmyA and rmyB at high and low cell density. The results were calculated as –ΔΔCt = –(ΔCthighcelldensity – ΔCtlowcelldensity). Three biological repeats (indedendent cultures) and three technical repeats were done to calculate and compare the values, all statistics were presented as means ± SE.
Mutation of phcB Attenuates the Bacterial Induction Activity on FOC Chlamydospore Formation and Inhibitory Ability on S. scitamineum Growth
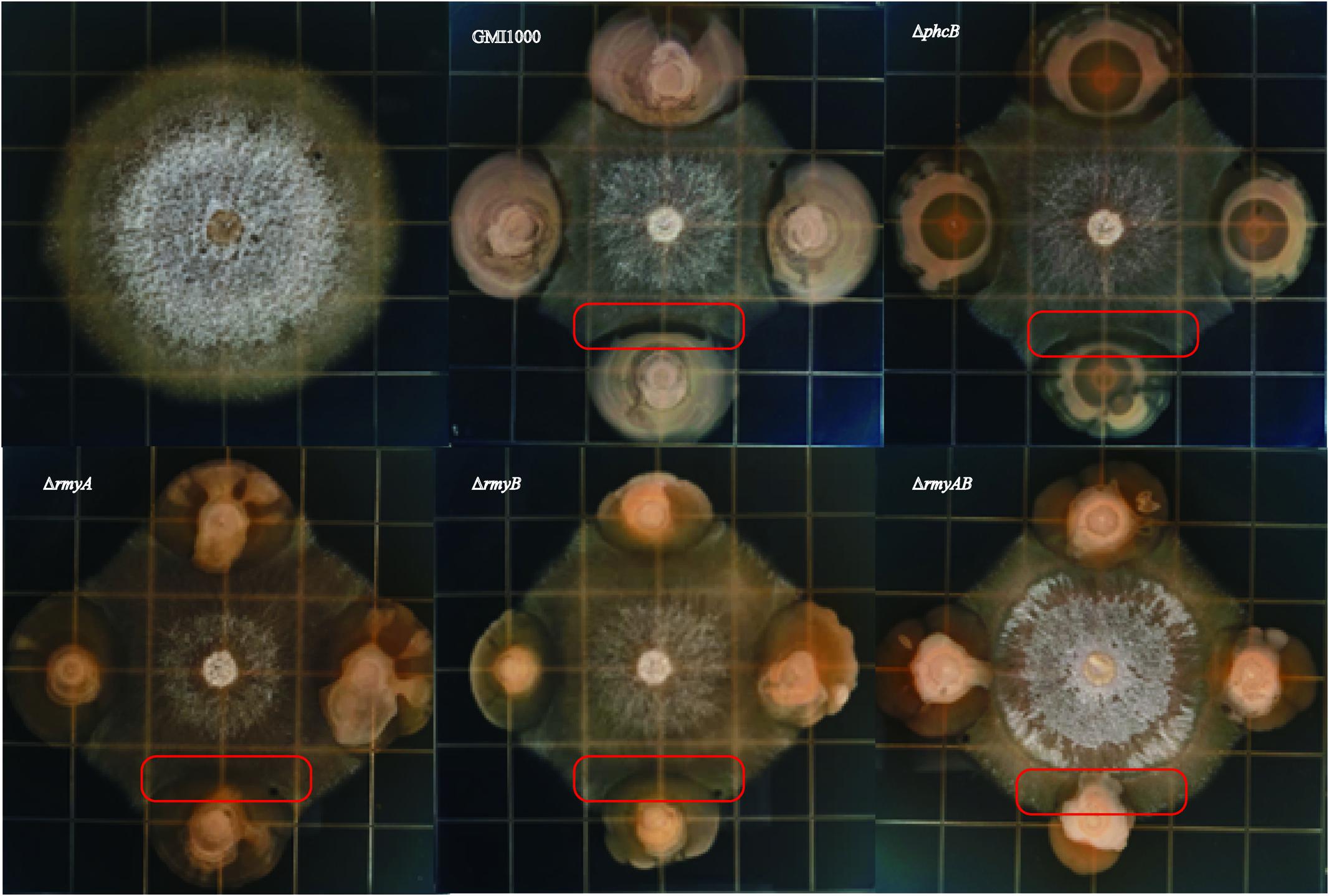
Interaction of R. solanacearum strain GMI1000 and fungal pathogen FOC strain XJZ2 were examined by spotting them side by side in the same plate as shown in Figure 4. After 8 days of co-culture, distinct zones of fungal inhibition were formed in the area containing R. solanacearum wide-type strain GMI1000. We also detected the diameter of XJZ2 colony co-cultured with GMI1000 and its derivatives to quantify the inhibition effect, result demonstrated that the XJZ2 colony co-cultured with wide-type strain GMI1000 with the smallest diameter. In comparison to GMI1000, the reduction of phcB deletion mutant inhibition effect reached significant level (Welch t-test, p-value = 0.02; Figure 5). In addition, the chlamydospores of strain XJZ2 were routinely found in the interaction zones with wide type strain GMI1000 Supplementary Information File 2 (SI 2), whereas in the area containing phcB deletion mutant ΔphcB, chlamydospores were hardly found. Complementation of the mutant ΔphcB with a wild-type phcB gene restored the bacterial induction ability on chlamydospore formation Supplementary Information File 2 (SI 2). When the mutant ΔrmyA, or ΔrmyB, or the double deletion mutant ΔrmyAB was co-cultured with the fungal strain XJZ2, the chlamydospore was also hardly found within their interaction zones Supplementary Information File 2 (SI 2).
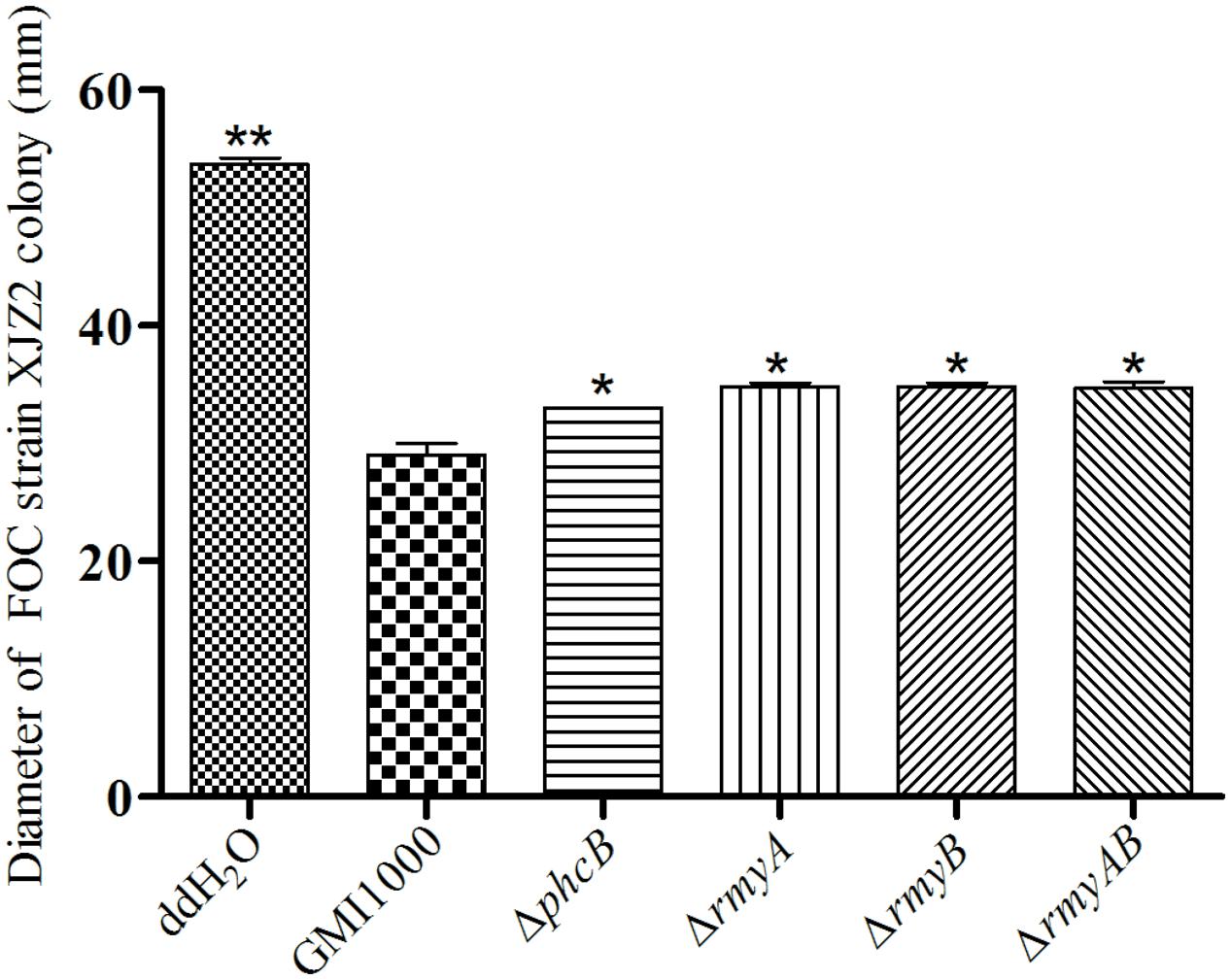
FIGURE 5. Diameter of XJZ2 colony co-culture with GMI1000 and its derivatives. Three biological repeats were conducted per strain. Bars indicate standard deviation (Welch t-test, comparison with the wide type strain GMI1000, ∗p-value < 0.05, ∗∗ p-value < 0.01).
Distinct inhibition zones were also found between wide-type strain GMI1000 and the sugarcane fungal pathogen S. scitamineum (Figure 6). The colony of strain GMI1000 inoculated 1 day earlier generated a more obvious inhibition zones than the bacterial colony inoculated at the same time with the fungal strain, suggesting a clear dosage effect of the inhibitory compound(s) produced by strain GMI1000. The mutant ΔphcB could also inhibit the growth of S. scitamineum, but was weaker than the wide-type. In contrast, deletion of rmyA, rmyB, or rmyAB completely abolished the growth inhibition ability on S. scitamineum, as the hyphe of S. scitamineum could grow around or cover the colonies of these rmy gene deletion mutants, and the E. coli as a control showed no inhibition effect on the fungi.
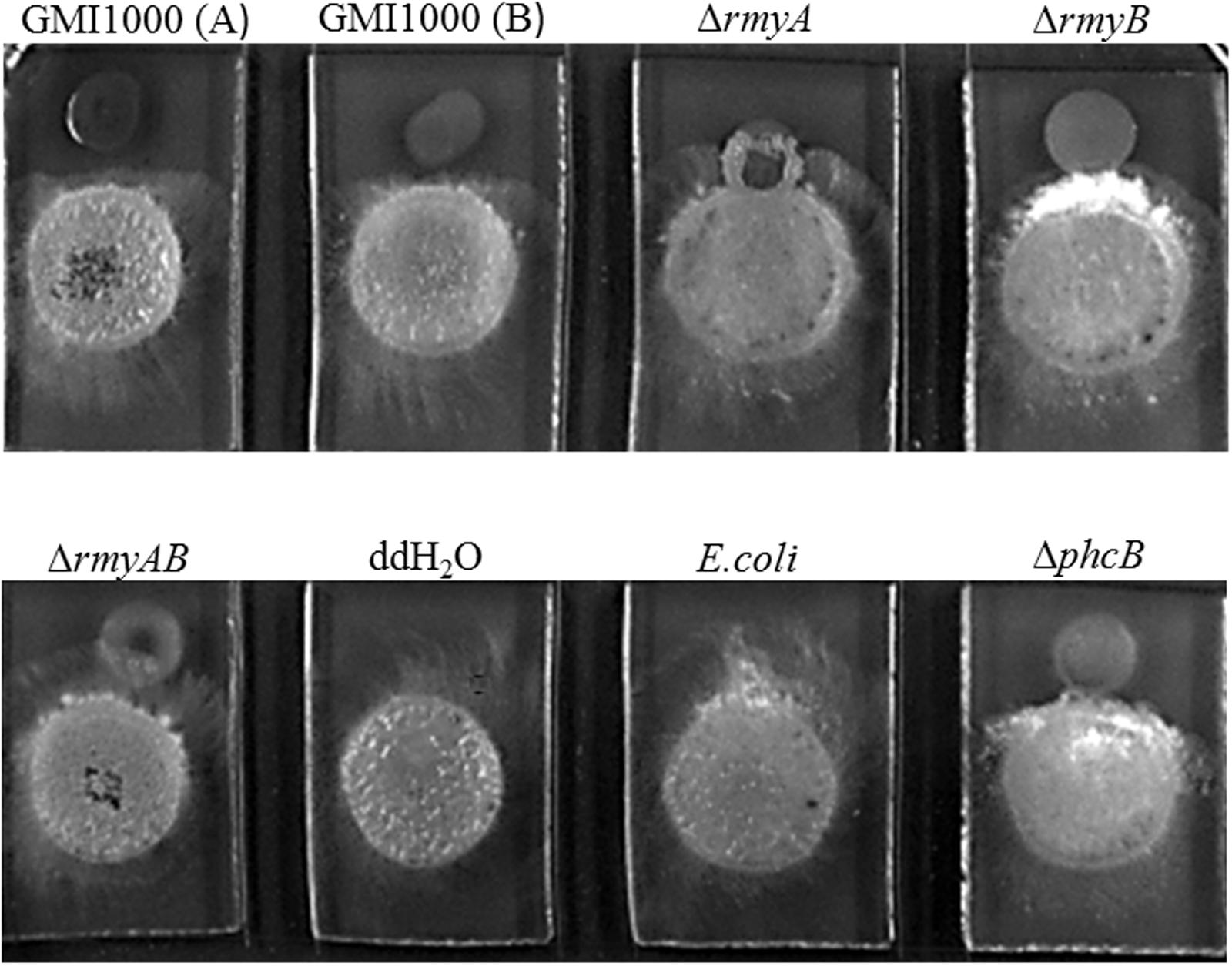

FIGURE 6. Co-culture experiments between R. solanacearum and Sporisorium scitamineum. GMI1000 (A) was cultured 24 h before adding the GMI1000 (B), other GMI1000 derivatives, and MAT-1 and MAT-2 mating mixture of S. scitamineum.
Discussion
Quorum sensing system is a widely conserved mechanism of bacterial cell-cell communications, which acts in a coordinated manner, and the individual bacteria can benefit from group behavior in competition for survive and persistence in nature (Fuqua et al., 1994; Von-Bodman et al., 2003). In R. solanacearum, the PhcBSR QS system serves as a master regulator to regulate most of the traits needed for infection and virulence (Denny, 1995, 2000; Schell, 2000). As an inter-species communication signal associated with competition for environmental niches, we are curious whether ralsolamycin biosynthesis is regulated by QS mechanisms. In this study, we provide sufficient genetic and chemical evidences that the two rmy genes associated with ralsolamycin biosynthesis and production in R. solanacearum are positively regulated by the PhcBSR QS system. During the course of bacteria–fungi interaction, we also showed that deletion of phcB attenuated the bacterial induction activity on chlamydospore development in fungal organisms, and reduced the inhibition activity to fungi.
It was predicted that up to five genes are involved in biosynthesis of ralsolamycin biosynthesis and transport, including rmyA, rmyB, RSp0638, RSp0639, and RSp0640 (Spraker et al., 2016). Although the later three genes are not in the same operon as rmyA and rmyB, their protein characteristics, and the similar QS-dependent expression pattern as the rmy genes unveiled in this study seem to support that these genes are functionally related probably, for deleting phcB also resulted in significant reduction in the transcripts levels of the two genes (RSp0638 and RSp0639) next to the rmy cluster.
Comparing with the NRPs that have been intensively investigated (Stachelhaus and Marahiel, 1995; Marahiel et al., 1997), the rmy genes in R. solanacearum GMI1000 are highly related to the syringomycin synthetase gene, which is required for the production of syringomycin in P. syringae (Bender et al., 1999). Another two NRPs product nunamycin and nunapeptin isolated from P. fluorescens In5 are the key biocontrol components against Rhizoctonia solani (Michelsen et al., 2015). A BLAST search found that RmyA and RmyB are also similar to AmbB and AmbE associated with IQS signal biosynthesis in human pathogen P. aeruginosa (Lee et al., 2013), with over 74% coverage and 35% identity at amino acids level. For the phosphate depletion is a host stress signal commonly encountered by invading pathogen P. aeruginosa (Lee et al., 2013), which actives the IQS system. Moreover, evidence reveals that EfpR is a novel key component of the complex regulatory network of the R. solanacearum cell, tightly linking the bacterial metabolism to virulence in response to multiple environmental signals (Perrier et al., 2016). Accordingly, whether there is any stress or environmental signal will involve in modulating the ralsolamycin production is still worth for further research.
Ralstonia solanacearum is known to wide distributed and persisted in soils for remarkably long periods (Alvarez et al., 2010), and the soil is a heterogeneous and complex microcosm replete with inter-organismal interactions, thus, the R. solanacearum will encounter a diversity of other soil microbes. The FOC and S. scitamineum are two spread worldwide and causes considerable yield losses and reduction in their host plants (Singh et al., 2004; O’Donnell et al., 2009). Based on our results, the ralsolamycin functioned in the crosstalk between R. solanacearum and the two tested fungi. On one side, the R. solanacearum can use the ralsolamycin to antagonize the fungi; on the other hand, which can also induce the fungi around to formate chlamydospores significantly, together with the fact that R. solanacearum can enter the endofungal lifestyle with chlamydospores (Spraker et al., 2016), which is a novel persistence mechanism for bacterial survival in the harsh environments. Moreover, we also want to determine whether the ralsolamycin can affect the sexual matting of S. scitamineum, result showed that except for the antagonization on S. scitamineum, the ralsolamycin does not inhibit the sexual matting. It was noteworthy that there was still weak inhibition effects on the fungi after deleting the rmy genes, thus, maybe some other metabolisms are also with the antagonization effect on the fungi. Obviously, the ralsolamycin plays a major role. It has been demonstrated that the “trade-off” existed between virulence factor production and bacterial proliferation is controlled by PhcBSR system dependent regulatory protein PhcA, a phcA mutant has an expanded metabolic versatility to metabolize up to 17 substrates and proliferation ability more than the wild-type (Peyraud et al., 2016). Rather paradoxically, our results indicate that the production of ralsolamycin is decreased dramatically when phcB is inactive. Hence, we can hypothese that the R. solanacearum can compete for a regnant niche and enhance survival ability for the partnering fungus under the modulation of PhcBSR QS system, and it shall be one kind of active regulatory mechanism to adapt to the harsh environment when the phcB is active, which may also contribute to understand why the R. solanacearum species complex could share such a broad ecological range with various host plants and soil cohabitants.
Conclusion
We have demonstrated that the biosynthesis of inter-kingdom communication signal ralsolamycin is regulated by the phcB dependent QS system. Significantly, the findings from this study unveiled a link between QS and inter-kingdom communication. Further studies are required to understand the molecular mechanisms and signaling pathways that govern the QS-dependent expression of the rmy genes and ralsolamycin.
Author Contributions
PL, YD, and L-HZ designed the experiments and wrote the paper; WY, JY, YC, JZ, and ML helped to obtain the mutants; SF and SS helped to perform the LC-MS analysis; YD and L-HZ revised the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by National Key Project for Basic Research of China (973 Program, No. 2015CB150600), Pearl River Nova Program of Guangzhou (No. 201506010067), and National Natural Science Foundation of China (No. 31571969).
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2017.01172/full#supplementary-material
References
Alvarez, B., Biosca, E. G., and Lopez, M. M. (2010). On the life of Ralstonia solanacearum, a destructive bacterial plant pathogen. Technol. Educ. Top. Appl. Microbiol. Microb. Biotechnol. 1, 267–279.
Arlat, M., Gough, C. L., Zischek, C., Barberis, P. A., Trigalet, A., and Boucher, C. A. (1992). Transcriptional organization and expression of the large hrp gene cluster of Pseudomonas solanacearum. Mol. Plant Microbe Interact. 5, 187–193. doi: 10.1094/MPMI-5-187
Bender, C. L., Alarcon-Chaidez, F., and Gross, D. C. (1999). Pseudomonas syringae phytotoxins: mode of action, regulation, and biosynthesis by peptide and polyketide synthetases. Microbiol. Mol. Biol. Rev. 63, 266–292.
Bhatt, G., and Denny, T. P. (2004).Ralstonia solanacearum iron scavenging by the siderophore staphyloferrin B is controlled by PhcA, the global virulence regulator. J. Bacteriol. 186, 7896–7904. doi: 10.1128/JB.186.23.7896-7904.20042004
Brumbley, S. M., Carney, B. F., and Denny, T. P. (1993). Phenotype conversion in Pseudomonas solanacearum due to spontaneous inactivation of PhcA, a putative LysR transcriptional regulator. J. Bacteriol. 175, 5477–5487. doi: 10.1128/jb.175.17.5477-5487.1993
Clough, S. J., Flavier, A. B., Schell, M. A., and Denny, T. P. (1997). Differential expression of virulence genes and motility in Ralstonia (Pseudomonas) solanacearum during exponential growth. Appl. Environ. Microbiol. 63, 844–850.
Denny, T. P. (1995). Involvement of bacterial polysaccharides in plant pathogenesis. Annu. Rev. Phytopathol. 33, 173–197. doi: 10.1146/annurev.py.33.090195.001133
Denny, T. P. (2000). Ralstonia solanacearum: a plant pathogen in touch with its host. Trends Microbiol. 8, 486–489. doi: 10.1016/S0966-842X(00)01860-6
Denny, T. P., and Ryel, B. S. (1991). Genetic evidence that extracellular polysaccharide is a virulence factor of Pseudomonas solanacearum. Mol. Plant Microbe Interact. 4, 198–206. doi: 10.1094/MPMI-4-198
Ditta, G., Stanfield, S., Corbin, D., and Helinski, D. R. (1980). Broad host range DNA cloning system for gram-negative bacteria: construction of a gene bank of Rhizobium meliloti. Proc. Natl. Acad. Sci. U.S.A. 77, 7347–7351. doi: 10.1073/pnas.77.12.7347
Flavier, A. B., Clough, S. J., Schell, M. A., and Denny, T. P. (1997). Identification of 3-hydroxypalmitic acid methyl ester as a novel autoregulator controlling virulence in Ralstonia solanacearum. Mol. Microbiol. 26, 251–259. doi: 10.1046/j.1365-2958.1997.5661945.x
Fuqua, W. C., Winans, S. C., and Greenberg, E. P. (1994). Quorum sensing in bacteria: the LuxR-LuxI family of cell density-responsive transcriptional regulators. J. Bacteriol. 176, 269–275. doi: 10.1128/jb.176.2.269-275.1994
Genin, S., and Denny, T. P. (2012). Pathogenomics of the Ralstonia solanacearum species complex. Annu. Rev. Phytopathol. 50, 67–89. doi: 10.1146/annurev-phyto-081211-173000
Hayward, A. C. (1991). Biology and epidemiology of bacterial wilt caused by Pseudomonas solanacearum. Annu. Rev. Phytopathol. 29, 65–87. doi: 10.1146/annurev.py.29.090191.000433
Hendrick, C. A., and Sequeira, L. (1984). Lipopolysaccharide-defective mutants of the wilt pathogen Pseudomonas solanacearum. Appl. Environ. Microbiol. 48, 94–101.
Huang, J., Carney, B. F., Denny, T. P., Weissinger, A. K., and Schell, M. A. (1995). A complex network regulates expression of eps and other virulence genes of Pseudomonas solanacearum. J. Bacteriol. 177, 1259–1267. doi: 10.1128/jb.177.5.1259-1267.1995
Hussain, M. B. B. M., Zhang, H. B., Xu, J. L., Liu, Q. G., Jiang, Z. D., and Zhang, L. H. (2008). The acyl-homoserine lactone-type quorum-sensing system modulates cell motility and virulence of Erwinia chrysanthemi pv. zeae. J Bacteriol 190, 1045–1053. doi: 10.1128/JB.01472-07
Kai, K., Ohnishi, H., Mori, Y., Kib, A., Ohnishi, K., and Hikichi, Y. (2014). Involvement of ralfuranone production in the virulence of Ralstonia solanacearum OE1-1. ChemBioChem 15, 2590–2597. doi: 10.1002/cbic.201402404
Kai, K., Ohnishi, H., Shimatani, M., Ishikawa, S., Mori, Y., Kiba, A., et al. (2015). Methyl 3-Hydroxymyristate, a diffusible signal mediating phc quorum sensing in Ralstonia solanacearum. ChemBioChem 16, 2309–2318. doi: 10.1002/cbic.201500456
Kovach, M. E., Elzer, P. H., Steven, H. D., Robertson, G. T., Farris, M. A., Roop, I. R. M., et al. (1995). Four new derivatives of the broad-host-range cloning vector pBBR1MCS, carrying different antibiotic-resistance cassettes. Gene 166, 175–176. doi: 10.1016/0378-1119(95)00584-1
Kreutzer, M. F., Kage, H., Gebhardt, P., Wackler, B., Saluz, H. P., Hoffmeister, D., et al. (2011). Biosynthesis of a complex yersiniabactin-like natural product via the mic locus in phytopathogen Ralstonia solanacearum. Appl. Environ. Microbiol. 77, 6117–6124. doi: 10.1128/AEM.05198-11
Lee, J., Wu, J. E., Deng, Y. Y., Wang, J., Wang, C., Wang, J. H., et al. (2013). A cell-cell communication signal integrates quorum sensing and stress response. Nat. Chem. Biol. 9, 339–343. doi: 10.1038/nchembio.12251225
Lee, J., and Zhang, L. H. (2015). The hierarchy quorum sensing network in Pseudomonas aeruginosa. Protein Cell 6, 26–41. doi: 10.1007/s13238-014-0100-x0100-x
Li, M. H., Xie, X. L., Lin, X. F., Shi, J. X., Ding, Z. J., Ling, J. F., et al. (2014). Functional characterization of the gene FoOCH1 encoding a putative alpha-1,6-mannosyltransferase in Fusarium oxysporum f. sp. cubense. Fungal Genet. Biol. 65, 1–13. doi: 10.1016/j.fgb.2014.01.005
Livak, K. J., and Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25, 402–408. doi: 10.1006/meth.2001.1262
Marahiel, M. A., Stachelhaus, T., and Mootz, H. D. (1997). Modular peptide synthetases involved in nonribosomal peptide synthesis. Chem. Rev. 97, 2651–2674. doi: 10.1021/cr960029e
Michelsen, C. F., Watrous, J., Glaring, M. A., Kersten, R., Koyama, N., Dorrestein, P. C., et al. (2015). Nonribosomal peptides, key biocontrol components for Pseudomonas fluorescens In5, isolated from a greenlandic suppressive soil. mBio 6:e00079-15. doi: 10.1128/mBio.00079-1579-15
O’Donnell, K., Gueidan, C., Sink, S., Johnston, P. R., Crous, P. W., Glenn, A., et al. (2009). A two-locus DNA sequence database for typing plant and human pathogens within the Fusarium oxysporum species complex. Fungal Genet. Biol. 46, 936–948. doi: 10.1016/j.fgb.2009.08.006
Perrier, A., Peyraud, R., Rengel, D., Barlet, X., Lucasson, E., Gouzy, J., et al. (2016). Enhanced in planta fitness through adaptive mutations in EfpR, a dual regulator of virulence and metabolic functions in the plant pathogen Ralstonia solanacearum. PLoS Pathog. 12:e1006044. doi: 10.1371/journal.ppat.10060446044
Peyraud, R., Cottret, L., Marmiesse, L., Gouzy, J., and Genin, S. (2016). A resource allocation trade-off between virulence and proliferation drives metabolic versatility in the plant pathogen Ralstonia solanacearum. PLoS Pathog. 12:e1005939. doi: 10.1371/journal.ppat.10059395939
Salanoubat, M., Genin, S., Artiguenave, F., Gouzy, J., Mangenot, S., Arlat, M., et al. (2002). Genome sequence of the plant pathogen Ralstonia solanacearum. Nature 415, 497–502. doi: 10.1038/415497a
Schäfer, A., Tauch, A., Jäger, W., Kalinowski, J., Thierbach, G., and Pühler, A. (1994). Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene 145, 69–73. doi: 10.1016/0378-1119(94)90324-7
Schell, M. A. (2000). Control of virulence and pathogenicity genes of Ralstonia solanacearum by an elaborate sensory network. Annu. Rev. Phytopathol. 38, 263–292. doi: 10.1146/annurev.phyto.38.1.263
Singh, N., Somai, B. M., and Pillay, D. (2004). Smut disease assessment by PCR and microscopy in inoculated tissue cultured sugarcane cultivars. Plant Sci. 167, 987–994. doi: 10.1016/j.plantsci.2004.05.00605.006
Spraker, J. E., Sanchez, L. M., Lowe, T. M., Dorrestein, P. C., and Keller, N. P. (2016). Ralstonia solanacearum lipopeptide induces chlamydospore development in fungi and facilitates bacterial entry into fungal tissues. ISME J. 10, 2317–2330. doi: 10.1038/ismej.2016.32
Stachelhaus, T., and Marahiel, M. A. (1995). Modular structure of genes encoding multifunctional peptide synthetases required for non-ribosomal peptide synthesis. FEMS Microbiol. Lett. 125, 3–14. doi: 10.1111/j.1574-6968.1995.tb07328.x
Von-Bodman, S. B., Bauer, W. D., and Coplin, D. L. (2003). Quorum sensing in plant-pathogenic bacteria. Annu. Rev. Phytopathol. 41, 455–482. doi: 10.1146/annurev.phyto.41.052002.095652
Yan, M. X., Zhu, G. N., Lin, S. Y., Xian, X. Y., Chang, C. Q., Xi, P. G., et al. (2016). The mating-type locus b of the sugarcane smut Sporisorium scitamineum is essential for mating, filamentous growth and pathogenicity. Fungal Genet. Biol. 86, 1–8. doi: 10.1016/j.fgb.2015.11.005
Yao, J., and Allen, C. (2006). Chemotaxis is required for virulence and competitive fitness of the bacterial wilt pathogen Ralstonia solanacearum. J. Bacteriol. 188, 3697–3708. doi: 10.1128/JB.188.10.3697-3708.2006
Keywords: bacterial wilt, non-ribosomal peptide, interaction, soil microbes, regulatory mechanism
Citation: Li P, Yin W, Yan J, Chen Y, Fu S, Song S, Zhou J, Lyu M, Deng Y and Zhang L-H (2017) Modulation of Inter-kingdom Communication by PhcBSR Quorum Sensing System in Ralstonia solanacearum Phylotype I Strain GMI1000. Front. Microbiol. 8:1172. doi: 10.3389/fmicb.2017.01172
Received: 23 March 2017; Accepted: 08 June 2017;
Published: 23 June 2017.
Edited by:
Philippe Prior, Institut National de la Recherche Agronomique, FranceReviewed by:
Yasufumi Hikichi, Kōchi University, JapanPeyraud Rémi, INRA Centre Occitanie-Toulouse, France
Copyright © 2017 Li, Yin, Yan, Chen, Fu, Song, Zhou, Lyu, Deng and Zhang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Lian-Hui Zhang, bGh6aGFuZzAxQHNjYXUuZWR1LmNu;, cmFsc3RvbmlhQDE2My5jb20= Yinyue Deng, eWRlbmdAc2NhdS5lZHUuY24=
 Peng Li
Peng Li Wenfang Yin
Wenfang Yin Jinli Yan
Jinli Yan Yufan Chen1
Yufan Chen1 Shihao Song
Shihao Song Jianuan Zhou
Jianuan Zhou Yinyue Deng
Yinyue Deng