- Laboratoire de Microbiologie Signaux et Microenvironnement EA 4312, l’Institut Universitaire de Technologie d’Evreux (IUT), Université de Rouen, Normandy University, Evreux, France
Numerous studies in Gram-negative bacteria have focused on the Type VI Secretion Systems (T6SSs), Quorum Sensing (QS), and social behavior, such as in biofilms. These interconnected mechanisms are important for bacterial survival; T6SSs allow bacteria to battle other cells, QS is devoted to the perception of bacterial cell density, and biofilm formation is essentially controlled by QS. Here, we review data concerning T6SS dynamics and T6SS–QS cross-talk that suggest the existence of inter-bacterial communication via T6SSs.
Introduction
Bacteria are perpetually at war against multiple competitors and thus require weapons to conquer new territory or persist in an ecological niche. Among the mechanisms that aid in the struggle against other bacterial species are the Type VI Secretion Systems (T6SSs) (Hood et al., 2010). The T6SSs of Gram-negative bacteria are effector translocation apparatuses, resembling an inverted bacteriophage-puncturing device, composed of at least 13 proteins called core-components (TssA-M, for Type six secretion) (Boyer et al., 2009; Silverman et al., 2012). Auxiliary components can be associated with these conserved proteins (Tag, for Type six secretion associated genes) (Shalom et al., 2007). T6SSs participate in a broad variety of functions, including virulence and antibacterial activity (Pukatzki et al., 2006, 2007; Ma et al., 2009; Coulthurst, 2013; Sana et al., 2016). T6SSs also participate in metal ion uptake, such as that of iron, manganese, and zinc (Wang et al., 2015; Chen et al., 2016; Lin et al., 2017; Si et al., 2017), conferring an advantage during bacteria–bacteria competition. In this review, we provide an overview of the data on T6SS assembly and emphasize connections between T6SSs and bacterial communication.
T6SS Dynamics
The Global Scenario
Type VI Secretion System contractile nanomachines allow bacteria to inject toxins directly into prey cell membranes or cytoplasm. The machinery of the T6SS is assembled in an orderly manner. It starts with membrane complex formation, allowing baseplate positioning. The baseplate serves as a platform for contractile tail elongation. Contraction of the sheath propels effectors across membranes. Finally, the ATPase, TssH (ClpV), recycles the sheath and probably other T6SS components such as TssA, whereas the membrane-anchoring complex can be used to fire a new salvo (Figure 1).
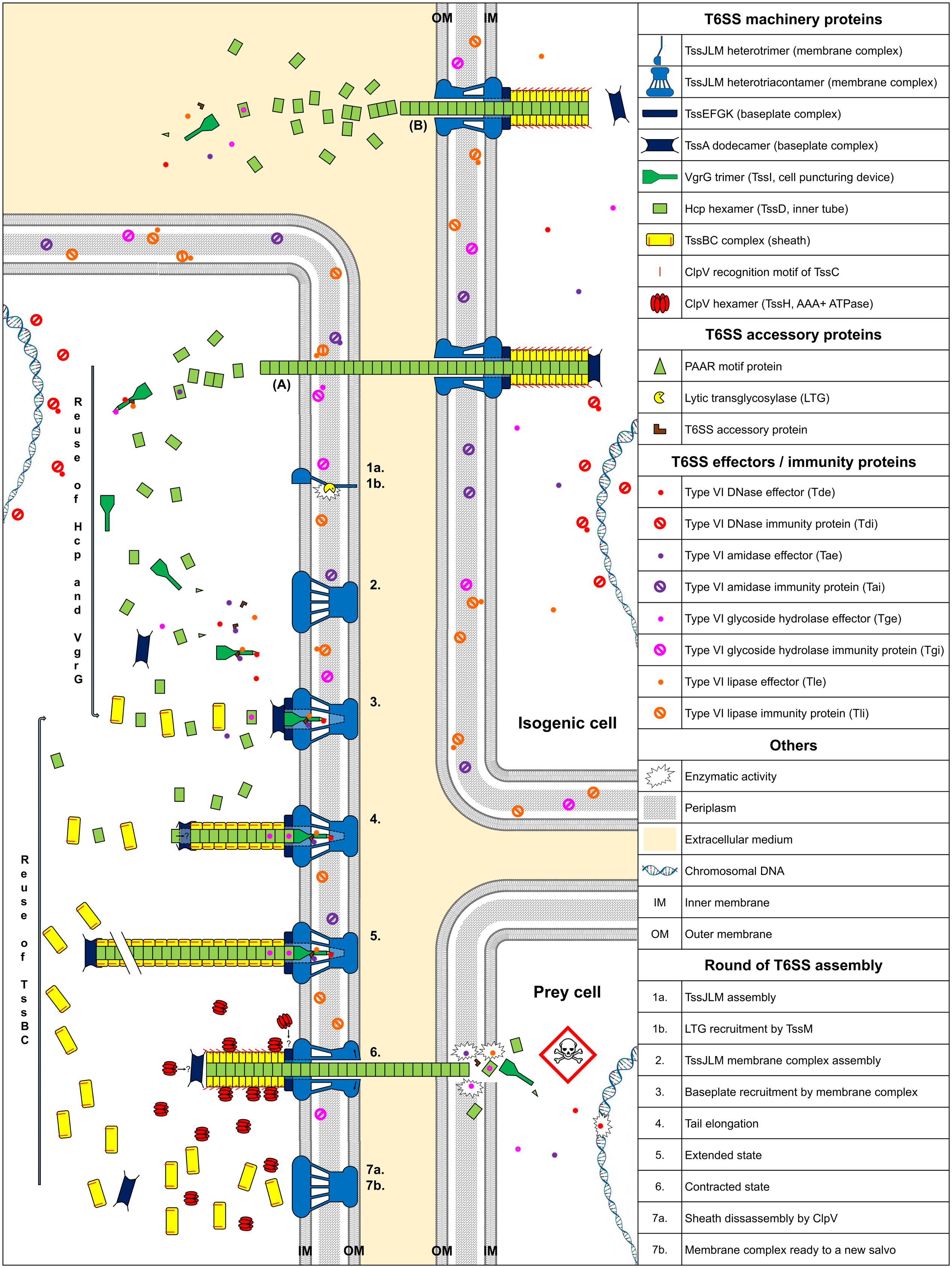
FIGURE 1. Model of T6SS assembly and Hcp/VgrG/effector translocation and recycling. The membrane complex is anchored to the cell envelope with the help of a lytic transglycosylase (1a, 1b, and 2). While effectors are loaded onto their respective Hcp, VgrG, or PAAR proteins, the baseplate platform is positioned (3), aiding inner tube and sheath assembly (4 and 5). TssA interacts with the membrane complex, allowing contractile tail polymerization (3). Hcp or sheath components can be individually added during elongation (4 and 5). Following sheath contraction, Hcp, VgrG/PAAR, with their associated effectors, are pushed out and delivered into prey cells (6). ClpV disassembles the sheath and probably other T6SS components (7a) and the membrane complex may be used for a new assembly cycle using recycled components (7b). (A) Indicates exogenous Hcp and VgrG that can be recycled after isogenic T6SS aggression. (B) Indicates secretion of Hcp and VgrG into the culture medium. The question mark indicates a hypothesis.
Membrane Complex Assembly
Type VI Secretion Systems are anchored to the cell envelope by a membrane core complex (Durand et al., 2015), which serves as a T6SS docking station and platform for baseplate assembly and prevents membrane cell damage during effector injection. The membrane core complex is a 1.7 MDa structure with a fivefold symmetry composed of 10 heterotrimeric complexes containing the three proteins TssJ, TssL, and TssM. Hierarchical biogenesis of this complex is initiated by the insertion of the lipoprotein TssJ in the outer membrane (Aschtgen et al., 2008; Zoued et al., 2014). TssJ then interacts with the large periplasmic domain of the inner membrane protein TssM (Felisberto-Rodrigues et al., 2011; Nguyen et al., 2015). The cytoplasmic domain of TssM interacts with the inner membrane protein TssL and the cytoplasmic domain of another TssM subunit, thus enabling oligomerization (Logger et al., 2016). Similarly, the cytoplasmic domain of TssL mediates self-polymerization (Durand et al., 2012; Zoued et al., 2016a). The TssM periplasmic domain recruits a lytic transglycosylase (LTG), which is required for local peptidoglycan layer degradation, necessary for proper assembly of the 1.7 MDa TssJLM complex (Figure 1, assembly steps 1 and 2) (Weber et al., 2016; Santin and Cascales, 2017). Associated proteins with a peptidoglycan-binding domain, such as TagL, which can bind to truncated TssL, can associate with the membrane complex. TagL corresponds in this case to an “ancestral TssL” domain (Aschtgen et al., 2010a,b).
Baseplate Complex Positioning
The T6SS baseplate complex, composed of TssA, TssE, TssF, TssG, TssK, and Valine-glycine repeat protein G (VgrG or TssI) proteins (Brunet et al., 2015), is recruited by the membrane complex (Zoued et al., 2013). This structure serves as a platform for contractile sheath assembly and is essential for the correct assembly of the inner tube, comprised of hexameric rings of Hemolysin-coregulated protein (Hcp or TssD) (Brunet et al., 2015). TssA forms a dodecamer complex, which first binds to the membrane complex (Zoued et al., 2016b) (Figure 1, assembly step 3). Positioning of the baseplate complex may be initiated by recruitment of TssE, TssK, and VgrG via TssA (Planamente et al., 2016; Zoued et al., 2016b). The cytoplasmic domains of TssM and TssL, located at the base of the membrane complex, interact with the TssG/TssK and TssK/TssE baseplate subunits, respectively (Zoued et al., 2013, 2016a; Logger et al., 2016). The baseplate complex thus forms an interface between the membrane complex and the T6SS tail: both the Hcp and TssC sheath subunits interact with baseplate components (Brunet et al., 2015) (Figure 1, assembly step 4). After TssE recruitment, TssA likely properly attaches the sheath onto the baseplate and/or stabilizes the sheath structure (Planamente et al., 2016).
Elongation of the Contractile Tail
The TssE baseplate component may initiate sheath assembly and anchors the sheath to the baseplate (Kudryashev et al., 2015). Hcp proteins assemble into hexameric rings, stacked head-to-tail, under the control of baseplate components (Brunet et al., 2014, 2015). The TssBC sheath component then wraps around the inner Hcp tube (Zoued et al., 2016b). Formation of the inner tube precedes TssBC assembly and is primordial for proper stacking of the subunits (Basler et al., 2012; Kapitein et al., 2013; Brunet et al., 2014). The Hcp tube has an inner diameter of ∼40 Å (Mougous et al., 2006), forming a lumen with a neutral surface, suggesting passive effector translocation into the Hcp tube (Ge et al., 2015). The diameter of the internal sheath, measuring approximately 100–110 Å (Bönemann et al., 2009), coincides with the ∼80–85 Å outer diameter of the Hcp hexamer (Cascales and Cambillau, 2012). The TssA dodecamer is located at the distal end of the tail in Escherichia coli (Zoued et al., 2016b), whereas TssA1 is a component of the baseplate/tail subcomplex in Pseudomonas aeruginosa (Planamente et al., 2016). The TssA complex appears to possess short, flexible arm-like extensions, which may grasp TssBC or Hcp and incorporate them, one by one, at the distal end of the contractile tail (Zoued et al., 2016b). Moreover, the diameter of the central channel of the ring-shaped TssA structure measures approximately 100 Å (Planamente et al., 2016), comparable to the dimension of the Hcp hexamers. Hcp components perhaps pass through the large central lumen of the TssA dodecamer (Figure 1, assembly step 4) and are added to the growing structure. Contrary to bacteriophages, the length of the T6SS tail does not appear to be controlled by a specialized protein (Zoued et al., 2014; Vettiger and Basler, 2016). The length of the T6SS tail can exceed 1 μm (Basler et al., 2012). It is possible that contact with the opposite cell membrane is the physical signal to stop tail elongation (Figure 1, assembly step 5).
Contraction and Sheath Disassembly
Clemens et al. (2015) demonstrated that the sheath of Francisella tularensis has a quaternary structure with handedness opposite to that of the contracted sheath of T4 phage tails. The sheath contracts within a few milliseconds (Basler et al., 2012), propelling the Hcp-VgrG spike and effectors, punching either indiscriminately or in a focused manner into neighboring bacteria. The sheath contracts and becomes shorter and wider than in the extended state (Basler et al., 2012). Once the sheath is contracted, the ClpV recognition motif of TssC, which is buried in the elongated state, becomes accessible (Bönemann et al., 2009; Pietrosiuk et al., 2011; Basler and Mekalanos, 2012; Kapitein et al., 2013; Kube et al., 2014; Douzi et al., 2016), permitting TssBC recycling by the ATPase. Thus, TssB and TssC can be reused for a new round of sheath elongation (Figure 1, assembly steps 6 and 7). An alternative mode of sheath disassembly may involve the TagJ accessory protein (Forster et al., 2014). TagJ is structurally related to particular TssA C-terminal extensions (Planamente et al., 2016). In this case, TagJ interacts with TssB and recruits ClpV to the sheath (Forster et al., 2014). ClpV can also interact with TssA and may be involved in recycling TssA rings (Planamente et al., 2016).
Effector Translocation
The puncturing device, consisting of the VgrG trimer, is located at the top of the inner tube and may be crucial for piercing the prey cell envelope. The VgrG trimer sometimes terminates with a Pro-Ala-Ala-Arg (PAAR)-repeat-containing protein, sharpening the tip (Shneider et al., 2013; Bondage et al., 2016).
Effectors transported by T6SS fall into two groups: “specialized” effectors and “cargo” effectors (Cianfanelli et al., 2016). Specialized effectors are extension domains of a structural component, whereas cargo effectors interact directly with VgrG, PAAR, or Hcp proteins (Silverman et al., 2013), with or without the help of accessory proteins (Alcoforado Diniz and Coulthurst, 2015; Liang et al., 2015; Unterweger et al., 2015). Four main classes of antibacterial effectors have been described, according to the target (Figure 1). Peptidoglycan targeting effectors are comprised of both Type six amidase effectors (Tae) and Type six glycoside hydrolase effectors (Tge) (Whitney et al., 2013). Type six lipase effectors (Tle) hydrolyse membrane phospholipids (Russell et al., 2013; Flaugnatti et al., 2016), whereas Type six DNase effectors (Tde) have nuclease activity (Ma et al., 2014). Some toxins do not belong to any of these four main classes. Pore-forming toxins, such as VasX from Vibrio cholerae, disrupt the inner membrane integrity of target cells (Miyata et al., 2013). Whitney and collaborators identified a NAD(P)+ glycohydrolase toxin in P. aeruginosa (Whitney et al., 2015). This toxin depletes cellular NAD(P)+ levels and induces bacteriostasis. The T6SS is not only an injection mechanism, it also enables the release of a proteinaceous metallophore into the extracellular medium and plays a role in the transport of Mn2+ under conditions of oxidative stress (Si et al., 2017) and iron uptake (Chen et al., 2016; Lin et al., 2017).
Bacteria that secrete antibacterial toxins also produce immunity proteins, which interact with toxic effectors, to allow self-protection and prevent the killing of sibling cells (called Tai, Tgi, Tli, and Tdi, corresponding to their effector family). Immunity proteins and effector targets are located within the same cellular compartment (Russell et al., 2013). Therefore, tli genes encoding outer membrane lipoproteins or periplasmic exposed lipoproteins, the Tle, should act in the periplasm (Figure 1).
Some other proteins secreted by the T6SS are involved in self-recognition, allowing communication between bacteria (Wenren et al., 2013; Cardarelli et al., 2015; Saak and Gibbs, 2016). In bacteria, secreted proteins are involved in many functions and are essential for bacterial fitness (Maffei et al., 2017). In some strains, the T6SS is activated in response of T6SS aggression by neighboring bacteria during cell–cell contact. Thus, the T6SS can modulate the fitness of other bacteria. In addition, T6SSs can be active, even in pure culture, and the presence of Hcp and VgrG in the culture medium is often used as evidence of a functional T6SS (Pukatzki et al., 2006, 2009). What purpose, however, does a functional T6SS serve in the absence of a competitor or prey?
Is the T6SS Involved in Cell-To-Cell Signaling and Communication?
Prelude
Pseudomonas aeruginosa is a widely-used model for T6SS studies. P. aeruginosa possesses three T6SS clusters comprised of TssA-M core component proteins. They are called H1-T6SS, H2-T6SS, and H3-T6SS. Among them, the H1-T6SS machinery is the most widely studied and is involved in antibacterial activity (Hood et al., 2010). The H2-T6SS and H3-T6SS are involved in virulence in eukaryotes (Lesic et al., 2009; Sana et al., 2012, 2015) but also display antibacterial activity by secreting trans-kingdom effectors, such as PldA and TplE via the H2-T6SS or PldB via the H3-T6SS machinery (Russell et al., 2013; Jiang et al., 2014, 2016).
“Dueling” and “Tit-for-Tat”
Two types of T6SS behavior can be distinguished: that of defensive (targeted firing) and offensive cells (arbitrarily firing). P. aeruginosa can discern T6SS-mediated aggression of neighboring sister cells (Basler and Mekalanos, 2012). Similarly, P. aeruginosa can perceive T6SS attacks coming from V. cholerae or Acinetobacter baylyi cells (Basler et al., 2013). In both cases, P. aeruginosa is first assaulted by a nearby bacterium and then attacks, in turn, the aggressive cell. This mechanism is called “T6SS dueling” and is mediated by the H1-T6SS. Thus, P. aeruginosa only counterattacks in response to T6SS firing of V. cholerae or A. baylyi. In general, P. aeruginosa does not target T6SS-defective bacteria, suggesting that it is a defensive bacterium, in contrast to offensive V. cholerae and A. baylyi strains. However, P. aeruginosa strains affected in the hybrid sensor kinase RetS, can attack T6SS-defective bacteria in a H1-T6SS-dependent manner (Hachani et al., 2013, 2014). The perception of a T6SS attack involves the TagQRST threonine phosphorylation pathway, following envelope perturbation after T6SS-mediated perforation (Basler et al., 2013; Casabona et al., 2013; Ho et al., 2013) (Figure 2). Indeed, Polymyxin B, which alters cell membranes of Gram-negative bacteria, mediates activation of the T6SS, confirming that envelope perturbation triggers the T6SS counterattack (Ho et al., 2013). The TagQRST trans-membrane signaling cascade, essential for activation of the H1-T6SS of P. aeruginosa, is composed of four proteins. The ABC transporter complex TagST, anchored to the inner membrane, has ATPase activity (Casabona et al., 2013). In this complex, TagT is required for T6SS activation after cell membrane perturbation (Ho et al., 2013). TagQ, an outer membrane lipoprotein, is necessary for outer membrane localisation of TagR, which is required for protein kinase PpkA phosphorylation (Hsu et al., 2009). PpkA phosphorylates, in turn, Fha1 (which has a forkhead-associated domain), thus activating H1-T6SS assembly (Figure 2). The phosphatase PppA counteracts the role of PpkA by dephosphorylating Fha1 (Mougous et al., 2007). T6SS dueling appears to be an indirect means of communication, in which the occurrence of T6SS attacks may correlate with cell density. As the population increases, the likelihood of targeting sister cells also intensifies. In other words, as the population grows, the incidence of T6SS aggression rises. Thus, the perception of T6SS attacks provides an overall view of bacterial density and a form of social interaction (Figure 2).
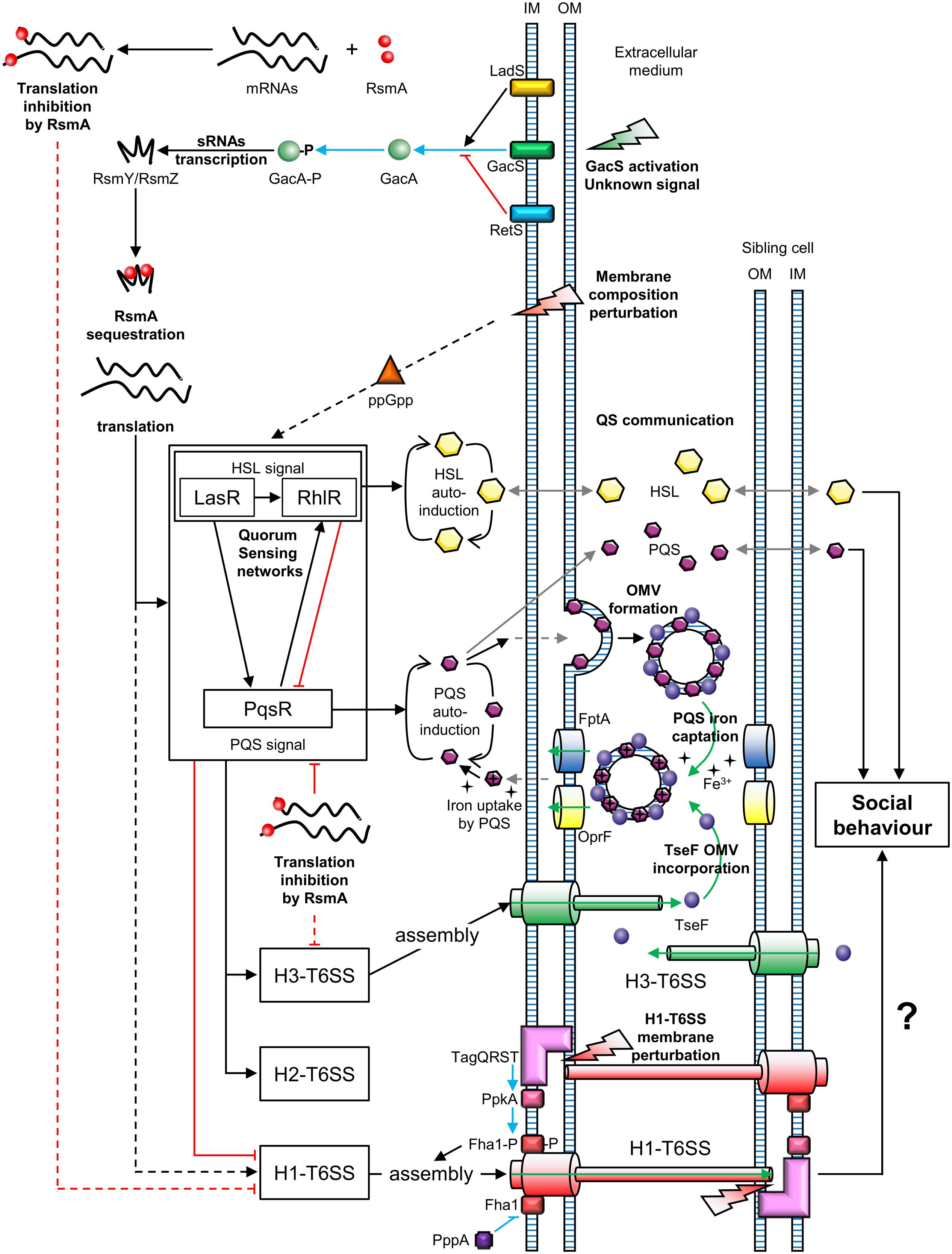
FIGURE 2. Connections between T6SSs and bacterial communication in Pseudomonas aeruginosa. OM, outer membrane; IM, inner membrane; OMV, outer membrane vesicle; P, phosphate; QS, quorum sensing. Green arrows represent molecule transport or uptake; black arrows indicate positive regulation; red blunt lines represent negative regulation; gray arrows represent diffusion; blue arrows indicate phosphorylation and blue blunt lines represent dephosphorylation; uncharacterized regulations are indicated by dotted lines. The question mark indicates a hypothesis.
The GacS/GacA System and the Interplay between the T6SS and Quorum Sensing in P. aeruginosa
Two-Component Signal Transduction systems (TCSTs) are involved in external signal perception via a “sensor” and translate the signal via a “response regulator.” Thus, TCSTs play a key role in adaptive responses during environmental stress. GacS/GacA is a TCST in P. aeruginosa that perceives unknown signals and regulates numerous networks (Reimmann et al., 1997). The GacS/GacA cascade activates the transcription of the small RNAs (sRNAs) RsmZ and RsmY. RsmZ has a high affinity for the RNA-binding protein RsmA and can sequester it, permitting the translation of genes encoding H1-T6SS mRNAs. A rsmZ mutation results in downregulation of the transcription of genes encoding the H1-T6SS and H3-T6SS of P. aeruginosa (Brencic et al., 2009; Moscoso et al., 2011). The GacS/GacA system is under the control of two hybrid sensors, RetS and LadS. The hybrid sensor kinase RetS decreases RsmZ/RsmY transcription via the inhibition of GacS/GacA phospho-relay (Goodman et al., 2004). In contrast, LadS, enhances GacA phosphorylation via GacS (Chen et al., 2015). In summary, H1-T6SS is upregulated by the GacS/GacA/RsmZ regulatory pathway, which depends on the balance between RetS and LadS activation from external signals, unlike the H2-T6SS and H3-T6SS (Figure 2).
Quorum Sensing (QS) is a system that allows social synchronization, based on the perception of population density, according to signal molecule concentration. QS is crucial for collective adaptive responses (similar to a social behavior) and regulates both bacterial virulence and biofilm formation (Deng et al., 2011). P. aeruginosa has four QS networks consisting of three classes of diffusible auto-inducers (Lee and Zhang, 2015). The first class includes two types of Homoserine Lactones (HSLs): N-(3-oxododecanoyl)-Homoserine Lactones (odDHL, 3-oxo-C12-HSL) and N-butyrylhomoserine Lactones (BHL, C4-HSL) controlled by the Las and Rhl cascades, respectively (Schuster and Greenberg, 2007). P. aeruginosa also produces Pseudomonas Quinolone Signal (PQS), 2-heptyl-3-hydroxy-4-quinolone, of which the production is regulated by the PqsR cascade (also known as MvfR) (Cao et al., 2001). The last QS system consists of 2-(2-hydroxyphenyl)-thiazole-4-carbaldehyde, which is involved in the Integrated Quorum Sensing system (IQS) (Lee et al., 2013). The IQS can enhance PQS production depending on the P. aeruginosa strain (Lee et al., 2013; Sun et al., 2016). These QS networks are all interconnected and positively regulated by the Las cascade (Pessi et al., 2001; Lee et al., 2013). At the same time, RsmA is a negative post-transcriptional regulator of both 3-oxo-C12-HSL and C4-HSL production, affecting the Las/Rhl quorum sensing cascades (Pessi et al., 2001). Similarly, PqsR is post-transcriptionally repressed by RsmA (Kulkarni et al., 2014). However, the PQS system and Rhl cascade are upregulated via the Las pathway and PQS positively regulates the Rhl cascade (Rasamiravaka et al., 2015). In summary, the GacS/GacA system is a global activator of QS communication, because the action of the RNA-binding protein RsmA is jointly antagonized by RsmY and RsmZ (Kay et al., 2006) (Figure 2). At the same time, QS positively regulates H2-T6SS and H3-T6SS, whereas it suppresses H1-T6SS associated gene expression (Lesic et al., 2009).
A study published by Lin et al. (2017) revealed a link between the H3-T6SS and cell–cell signaling in P. aeruginosa. The cell–cell signaling compound PQS contributes to the formation of Outer Membrane Vesicles (OMVs) and associates with vesicle membranes. The PQS in OMVs can capture iron from the extracellular medium. In parallel, the protein TseF, secreted by the H3-T6SS, associates with OMVs containing PQS. The TseF from the OMVs then interacts with the Fe(III)-pyochelin receptor FptA and the porin OprF. This enables the delivery of PQS, associated with iron, into bacterial cells. Thus, the effector TseF delivered by the H3-T6SS is involved in the PQS pathway (Figure 2).
T6SS and Social Behavior in Proteus mirabilis
Proteus mirabilis strains are able to recognize isogenic cells, coordinate multicellular swarming motility, and form macroscopic boundaries with non-sister cell swarms (Alteri et al., 2013). Macroscopic demarcation, called Dienes lines, can be observed between swarming P. mirabilis strains. Functional T6SSs are involved in this recognition phenomenon in the region of inter-strain contact. This visible boundary requires physical cell–cell interactions and is the result of T6SS recognition. Indeed, cells at the intersection between two swarming populations of P. mirabilis appear to kill each other using their T6SS effectors (Alteri et al., 2013). The T6SSs appear to assemble and fire deeply beyond the inter-strain boundary into the opposing swarming cells, thus enhancing T6SS effector injection (Alteri et al., 2013). Some Identification of self (Ids) proteins, involved in self-recognition and territorial behavior, are exported by the T6SS (Wenren et al., 2013). For example, IdsD mediates identity recognition between neighboring cells in a T6SS-dependant manner. IdsD interacts specifically with the cognate IdsE protein on the surface of recipient cells. The specific interaction between the two membrane-bound self-recognition proteins IdsD and IdsE drives social behavior (Cardarelli et al., 2015). These binding interactions contribute to the definition of strain identity and discrimination between self and neighboring non-self cells. The lack of binding between IdsD and IdsE correlates with the formation of the visible boundary. The authors speculate that IdsE itself contributes to the repression of swarm colony expansion. Interaction between the two cognate proteins reduces swarm restriction (Saak and Gibbs, 2016). IdsD and IdsE proteins may constitute a lethal effector-immunity (toxin–antitoxin) system. Contrary to QS, which is based on contact-independent recognition, T6SS-associated recognition generally requires cell-to-cell contact (Saak and Gibbs, 2016). P. mirabilis uses the T6SS to discriminate between strains, coordinate multicellular swarming behavior, and direct its collective movement. Thus, the T6SS is essential for boundary formation and mediates cell–cell communication of swarming P. mirabilis via specific self-identity determinants.
The T6SS of the P. fluorescens MFE01 Strain
The P. fluorescens MFE01 strain, like numerous other P. fluorescens strains, does not produce the QS signals of P. aeruginosa (no HSL or PQS) (Gallique et al., 2017). MFE01 is an aggressive T6SS strain which contains a unique T6SS core component cluster and three orphan hcp genes (Decoin et al., 2014, 2015). The MFE01 T6SS is involved in biofilm formation and maturation (Gallique et al., 2017), as shown for other T6SSs (de Pace et al., 2011; Sheng et al., 2013; Lin et al., 2015; Tian et al., 2015). Indeed, P. fluorescens MFE01 is unable to form biofilms once the T6SS machinery is inactive (in a tssC mutant), whereas individual mutations of the three hcp genes affect biofilm maturation, but not formation. Intra-bacterial cooperation in conditions of biofilm formation via T6SS dueling could occur. Indeed, alterations of membrane phospholipid composition increase the ppGpp stress-response signal, which in turn causes the premature production of HSL-QS signals, including in P. aeruginosa (Baysse et al., 2005) (Figure 2). Similarly, communicating pathways could be activated following membrane perturbation due to T6SS perforation in MFE01 strain during “tit-for-tat” interactions.
Conclusion
A recent study showed that bacteria can reuse T6SS components from attacking cells for new T6SS assembly (Vettiger and Basler, 2016) (Figure 1). This suggests that an increase in cell density increases the concentration of T6SS components in bacteria and the ability of the cell to fire again, forming a positive feedback loop. We postulate that T6SS could be a cell-to-cell signal between sibling cells, depending on cell density, similar to the QS pathway, especially in bacteria devoid of QS signals.
Author Contributions
Writing, review, and editing: MG, AM, MB.
Funding
This study was supported by GRR CBS funds from the Region Haute-Normandie, SFR Normandie Végétale (NORVEGE) funds, GEA (grand Evreux agglomeration) funds, and FEDER funds.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Alcoforado Diniz, J., and Coulthurst, S. J. (2015). Intraspecies competition in Serratia marcescens is mediated by type VI-secreted Rhs effectors and a conserved effector-associated accessory protein. J. Bacteriol. 197, 2350–2360. doi: 10.1128/JB.00199-15
Alteri, C. J., Himpsl, S. D., Pickens, S. R., Lindner, J. R., Zora, J. S., Miller, J. E., et al. (2013). Multicellular bacteria deploy the type VI secretion system to preemptively strike neighboring cells. PLoS Pathog. 9:e1003608. doi: 10.1371/journal.ppat.1003608
Aschtgen, M.-S., Bernard, C. S., De Bentzmann, S., Lloubès, R., and Cascales, E. (2008). SciN is an outer membrane lipoprotein required for type VI secretion in enteroaggregative Escherichia coli. J. Bacteriol. 190, 7523–7531. doi: 10.1128/JB.00945-08
Aschtgen, M.-S., Gavioli, M., Dessen, A., Lloubès, R., and Cascales, E. (2010a). The SciZ protein anchors the enteroaggregative Escherichia coli type VI secretion system to the cell wall. Mol. Microbiol. 75, 886–899. doi: 10.1111/j.1365-2958.2009.07028.x
Aschtgen, M.-S., Thomas, M. S., and Cascales, E. (2010b). Anchoring the type VI secretion system to the peptidoglycan: TssL, TagL, TagP...what else? Virulence 1, 535–540.
Basler, M., Ho, B. T., and Mekalanos, J. J. (2013). Tit-for-tat: type VI secretion system counterattack during bacterial cell-cell interactions. Cell 152, 884–894. doi: 10.1016/j.cell.2013.01.042
Basler, M., and Mekalanos, J. J. (2012). Type 6 secretion dynamics within and between bacterial cells. Science 337:815. doi: 10.1126/science.1222901
Basler, M., Pilhofer, M., Henderson, P. G., Jensen, J. G., and Mekalanos, J. (2012). Type VI secretion requires a dynamic contractile phage tail-like structure. Nature 483, 182–186. doi: 10.1038/nature10846
Baysse, C., Cullinane, M., Dénervaud, V., Burrowes, E., Dow, J. M., Morrissey, J. P., et al. (2005). Modulation of quorum sensing in Pseudomonas aeruginosa through alteration of membrane properties. Microbiology 151(Pt 8), 2529–2542. doi: 10.1099/mic.0.28185-0
Bondage, D. D., Lin, J.-S., Ma, L.-S., Kuo, C.-H., and Lai, E.-M. (2016). VgrG C terminus confers the type VI effector transport specificity and is required for binding with PAAR and adaptor-effector complex. Proc. Natl. Acad. Sci. U.S.A. 113, E3931–E3940. doi: 10.1073/pnas.1600428113
Bönemann, G., Pietrosiuk, A., Diemand, A., Zentgraf, H., and Mogk, A. (2009). Remodelling of VipA/VipB tubules by ClpV-mediated threading is crucial for type VI protein secretion. EMBO J. 28, 315–325. doi: 10.1038/emboj.2008.269
Boyer, F., Fichant, G., Berthod, J., Vandenbrouck, Y., and Attree, I. (2009). Dissecting the bacterial type VI secretion system by a genome wide in silico analysis: what can be learned from available microbial genomic resources? BMC Genomics 10:104. doi: 10.1186/1471-2164-10-104
Brencic, A., McFarland, K. A., McManus, H. R., Castang, S., Mogno, I., Dove, S. L., et al. (2009). The GacS/GacA signal transduction system of Pseudomonas aeruginosa acts exclusively through its control over the transcription of the RsmY and RsmZ regulatory small RNAs. Mol. Microbiol. 73, 434–445. doi: 10.1111/j.1365-2958.2009.06782.x
Brunet, Y. R., Hénin, J., Celia, H., and Cascales, E. (2014). Type VI secretion and bacteriophage tail tubes share a common assembly pathway. EMBO Rep. 15, 315–321. doi: 10.1002/embr.201337936
Brunet, Y. R., Zoued, A., Boyer, F., Douzi, B., and Cascales, E. (2015). The type VI secretion TssEFGK-VgrG phage-like baseplate is recruited to the TssJLM membrane complex via multiple contacts and serves as assembly platform for tail tube/sheath polymerization. PLoS Genet. 11:e1005545. doi: 10.1371/journal.pgen.1005545
Cao, H., Krishnan, G., Goumnerov, B., Tsongalis, J., Tompkins, R., and Rahme, L. G. (2001). A quorum sensing-associated virulence gene of Pseudomonas aeruginosa encodes a LysR-like transcription regulator with a unique self-regulatory mechanism. Proc. Natl. Acad. Sci. U.S.A. 98, 14613–14618. doi: 10.1073/pnas.251465298
Cardarelli, L., Saak, C., and Gibbs, K. A. (2015). Two proteins form a heteromeric bacterial self-recognition complex in which variable subdomains determine allele-restricted binding. mBio 6:e00251-15. doi: 10.1128/mBio.00251-15
Casabona, M. G., Silverman, J. M., Sall, K. M., Boyer, F., Couté, Y., Poirel, J., et al. (2013). An ABC-transporter and an outer membrane lipoprotein participate in posttranslational activation of type VI secretion in Pseudomonas aeruginosa. Environ. Microbiol. 15, 471–486. doi: 10.1111/j.1462-2920.2012.02816.x
Cascales, E., and Cambillau, C. (2012). Structural biology of type VI secretion systems. Philos. Trans. R. Soc. Lond. B Biol. Sci. 367, 1102–1111. doi: 10.1098/rstb.2011.0209
Chen, L., Zou, Y., She, P., and Wu, Y. (2015). Composition, function, and regulation of T6SS in Pseudomonas aeruginosa. Microbiol. Res. 172, 19–25. doi: 10.1016/j.micres.2015.01.004
Chen, W.-J., Kuo, T.-Y., Hsieh, F.-C., Chen, P.-Y., Wang, C.-S., Shih,Y.-L., et al. (2016). Involvement of type VI secretion system in secretion of iron chelator pyoverdine in Pseudomonas taiwanensis. Sci. Rep. 6:32950. doi: 10.1038/srep32950
Cianfanelli, F. R., Monlezun, L., and Coulthurst, S. J. (2016). Aim, load, fire: the type VI secretion system, a bacterial nanoweapon. Trends Microbiol. 24, 51–62. doi: 10.1016/j.tim.2015.10.005
Clemens, D. L., Ge, P., Lee, B.-Y., Horwitz, M. A., and Zhou, Z. H. (2015). Atomic structure of T6SS reveals interlaced array essential to function. Cell 160, 940–951. doi: 10.1016/j.cell.2015.02.005
Coulthurst, S. J. (2013). The type VI secretion system - a widespread and versatile cell targeting system. Res. Microbiol. 164, 640–654. doi: 10.1016/j.resmic.2013.03.017
de Pace, F., Boldrin de Paiva, J., Nakazato, G., Lancellotti, M., Sircili, M. P., Guedes Stehling, E., et al. (2011). Characterization of IcmF of the type VI secretion system in an avian pathogenic Escherichia coli (APEC) strain. Microbiology 157(Pt 10), 2954–2962. doi: 10.1099/mic.0.050005-0
Decoin, V., Barbey, C., Bergeau, D., Latour, X., Feuilloley, M. G. J., Orange, N., et al. (2014). A type VI secretion system is involved in Pseudomonas fluorescens bacterial competition. PLoS ONE 9:e89411. doi: 10.1371/journal.pone.0089411
Decoin, V., Gallique, M., Barbey, C., Le Mauff, F., Poc, C. D., Feuilloley, M. G. J., et al. (2015). A Pseudomonas fluorescens type 6 secretion system is related to mucoidy, motility and bacterial competition. BMC Microbiol. 15:72. doi: 10.1186/s12866-015-0405-9
Deng, Y., Wu, J., Tao, F., and Zhang, L.-H. (2011). Listening to a new language: DSF-based quorum sensing in Gram-negative bacteria. Chem. Rev. 111, 160–173. doi: 10.1021/cr100354f
Douzi, B., Brunet, Y. R., Spinelli, S., Lensi, V., Legrand, P., Blangy, S., et al. (2016). Structure and specificity of the type VI secretion system ClpV-TssC interaction in enteroaggregative Escherichia coli. Sci. Rep. 6:34405. doi: 10.1038/srep34405
Durand, E., Nguyen, V. S., Zoued, A., Logger, L., Péhau-Arnaudet, G., Aschtgen, M.-S., et al. (2015). Biogenesis and structure of a type VI secretion membrane core complex. Nature 523, 555–560. doi: 10.1038/nature14667
Durand, E., Zoued, A., Spinelli, S., Watson, P. J. H., Aschtgen, M.-S., Journet, L., et al. (2012). Structural characterization and oligomerization of the TssL protein, a component shared by bacterial type VI and type IVb secretion systems. J. Biol. Chem. 287, 14157–14168. doi: 10.1074/jbc.M111.338731
Felisberto-Rodrigues, C., Durand, E., Aschtgen, M.-S., Blangy, S., Ortiz-Lombardia, M., Douzi, B., et al. (2011). Towards a structural comprehension of bacterial type VI secretion systems: characterization of the TssJ-TssM complex of an Escherichia coli pathovar. PLoS Pathog. 7:e1002386. doi: 10.1371/journal.ppat.1002386
Flaugnatti, N., Le, T. T. H., Canaan, S., Aschtgen, M.-S., Nguyen, V. S., Blangy, S., et al. (2016). A phospholipase A1 antibacterial type VI secretion effector interacts directly with the C-terminal domain of the VgrG spike protein for delivery. Mol. Microbiol. 99, 1099–1118. doi: 10.1111/mmi.13292
Forster, A., Planamente, S., Manoli, E., Lossi, N. S., Freemont, P. S., and Filloux, A. (2014). Coevolution of the ATPase ClpV, the sheath proteins TssB and TssC, and the accessory protein TagJ/HsiE1 distinguishes type VI secretion classes. J. Biol. Chem. 289, 33032–33043. doi: 10.1074/jbc.M114.600510
Gallique, M., Decoin, V., Barbey, C., Rosay, T., Feuilloley, M. G. J., Orange, N., et al. (2017). Contribution of the Pseudomonas fluorescens MFE01 type VI secretion system to biofilm formation. PLoS ONE 12:e0170770. doi: 10.1371/journal.pone.0170770
Ge, P., Scholl, D., Leiman, P. G., Yu, X., Miller, J. F., and Zhou, Z. H. (2015). Atomic structures of a bactericidal contractile nanotube in its pre- and postcontraction states. Nat. Struct. Mol. Biol. 22, 377–382. doi: 10.1038/nsmb.2995
Goodman, A. L., Kulasekara, B., Rietsch, A., Boyd, D., Smith, R. S., and Lory, S. (2004). A signaling network reciprocally regulates genes associated with acute infection and chronic persistence in Pseudomonas aeruginosa. Dev. Cell 7, 745–754. doi: 10.1016/j.devcel.2004.08.020
Hachani, A., Allsopp, L. P., Oduko, Y., and Filloux, A. (2014). The VgrG proteins are “a la carte” delivery systems for bacterial type VI effectors. J. Biol. Chem. 289, 17872–17884. doi: 10.1074/jbc.M114.563429
Hachani, A., Lossi, N. S., and Filloux, A. (2013). A visual assay to monitor T6SS-mediated bacterial competition. J. Vis. Exp. 73:e50103. doi: 10.3791/50103
Ho, B. T., Basler, M., and Mekalanos, J. J. (2013). Type 6 secretion system-mediated immunity to type 4 secretion system-mediated gene transfer. Science 342, 250–253. doi: 10.1126/science.1243745
Hood, R. D., Singh, P., Hsu, F., Güvener, T., Carl, M. A., Trinidad, R. R. S., et al. (2010). A type VI secretion system of Pseudomonas aeruginosa targets a toxin to bacteria. Cell Host Microbe 7, 25–37. doi: 10.1016/j.chom.2009.12.007
Hsu, F., Schwarz, S., and Mougous, J. D. (2009). TagR promotes PpkA-catalyzed type VI secretion activation in Pseudomonas aeruginosa. Mol. Microbiol. 72, 1111–1125. doi: 10.1111/j.1365-2958.2009.06701.x
Jiang, F., Wang, X., Wang, B., Chen, L., Zhao, Z., Waterfield, N. R., et al. (2016). The Pseudomonas aeruginosa type VI secretion PGAP1-like effector induces host autophagy by activating endoplasmic reticulum stress. Cell Rep. 16, 1502–1509. doi: 10.1016/j.celrep.2016.07.012
Jiang, F., Waterfield, N. R., Yang, J., Yang, G., and Jin, Q. (2014). A Pseudomonas aeruginosa type VI secretion phospholipase D effector targets both prokaryotic and eukaryotic cells. Cell Host Microbe 15, 600–610. doi: 10.1016/j.chom.2014.04.010
Kapitein, N., Bönemann, G., Pietrosiuk, A., Seyffer, F., Hausser, I., Locker, J. K., et al. (2013). ClpV recycles VipA/VipB tubules and prevents non-productive tubule formation to ensure efficient type VI protein secretion. Mol. Microbiol. 87, 1013–1028. doi: 10.1111/mmi.12147
Kay, E., Humair, B., Dénervaud, V., Riedel, K., Spahr, S., Eberl, L., et al. (2006). Two GacA-dependent small RNAs modulate the quorum-sensing response in Pseudomonas aeruginosa. J. Bacteriol. 188, 6026–6033. doi: 10.1128/JB.00409-06
Kube, S., Kapitein, N., Zimniak, T., Herzog, F., Mogk, A., and Wendler, P. (2014). Structure of the VipA/B type VI secretion complex suggests a contraction-state-specific recycling mechanism. Cell Rep. 8, 20–30. doi: 10.1016/j.celrep.2014.05.034
Kudryashev, M., Wang, R. Y.-R., Brackmann, M., Scherer, S., Maier, T., Baker, D., et al. (2015). Structure of the type VI secretion system contractile sheath. Cell 160, 952–962. doi: 10.1016/j.cell.2015.01.037
Kulkarni, P. R., Jia, T., Kuehne, S. A., Kerkering, T. M., Morris, E. R., Searle, M. S., et al. (2014). A sequence-based approach for prediction of CsrA/RsmA targets in bacteria with experimental validation in Pseudomonas aeruginosa. Nucleic Acids Res. 42, 6811–6825. doi: 10.1093/nar/gku309
Lee, J., Wu, J., Deng, Y., Wang, J., Wang, C., Wang, J., et al. (2013). A cell-cell communication signal integrates quorum sensing and stress response. Nat. Chem. Biol. 9, 339–343. doi: 10.1038/nchembio.1225
Lee, J., and Zhang, L. (2015). The hierarchy quorum sensing network in Pseudomonas aeruginosa. Protein Cell 6, 26–41. doi: 10.1007/s13238-014-0100-x
Lesic, B., Starkey, M., He, J., Hazan, R., and Rahme, L. G. (2009). Quorum sensing differentially regulates Pseudomonas aeruginosa type VI secretion locus I and homologous loci II and III, which are required for pathogenesis. Microbiology 155, 2845–2855. doi: 10.1099/mic.0.029082-0
Liang, X., Moore, R., Wilton, M., Wong, M. J. Q., Lam, L., and Dong, T. G. (2015). Identification of divergent type VI secretion effectors using a conserved chaperone domain. Proc. Natl. Acad. Sci. U.S.A. 112, 9106–9111. doi: 10.1073/pnas.1505317112
Lin, J., Cheng, J., Chen, K., Guo, C., Zhang, W., Yang, X., et al. (2015). The icmF3 locus is involved in multiple adaptation- and virulence-related characteristics in Pseudomonas aeruginosa PAO1. Front. Cell. Infect. Microbiol. 5:70. doi: 10.3389/fcimb.2015.00070
Lin, J., Zhang, W., Cheng, J., Yang, X., Zhu, K., Wang, Y., et al. (2017). A Pseudomonas T6SS effector recruits PQS-containing outer membrane vesicles for iron acquisition. Nat. Commun. 8:14888. doi: 10.1038/ncomms14888
Logger, L., Aschtgen, M.-S., Guérin, M., Cascales, E., and Durand, E. (2016). Molecular dissection of the interface between the type VI secretion TssM cytoplasmic domain and the TssG baseplate component. J. Mol. Biol. 428, 4424–4437. doi: 10.1016/j.jmb.2016.08.032
Ma, A. T., McAuley, S., Pukatzki, S., and Mekalanos, J. J. (2009). Translocation of a Vibrio cholerae type VI secretion effector requires bacterial endocytosis by host cells. Cell Host Microbe 5, 234–243. doi: 10.1016/j.chom.2009.02.005
Ma, L.-S., Hachani, A., Lin, J.-S., Filloux, A., and Lai, E.-M. (2014). Agrobacterium tumefaciens deploys a superfamily of type VI secretion DNase effectors as weapons for interbacterial competition in planta. Cell Host Microbe 16, 94–104. doi: 10.1016/j.chom.2014.06.002
Maffei, B., Francetic, O., and Subtil, A. (2017). Tracking proteins secreted by bacteria: what’s in the toolbox? Front. Cell. Infect. Microbiol. 7:221. doi: 10.3389/fcimb.2017.00221
Miyata, S. T., Unterweger, D., Rudko, S. P., and Pukatzki, S. (2013). Dual expression profile of type VI secretion system immunity genes protects pandemic Vibrio cholerae. PLoS Pathog. 9:e1003752. doi: 10.1371/journal.ppat.1003752
Moscoso, J. A., Mikkelsen, H., Heeb, S., Williams, P., and Filloux, A. (2011). The Pseudomonas aeruginosa sensor RetS switches type III and type VI secretion via c-di-GMP signalling. Environ. Microbiol. 13, 3128–3138. doi: 10.1111/j.1462-2920.2011.02595.x
Mougous, J. D., Cuff, M. E., Raunser, S., Shen, A., Zhou, M., Gifford, C. A., et al. (2006). A virulence locus of Pseudomonas aeruginosa encodes a protein secretion apparatus. Science 312, 1526–1530. doi: 10.1126/science.1128393
Mougous, J. D., Gifford, C. A., Ramsdell, T. L., and Mekalanos, J. J. (2007). Threonine phosphorylation post-translationally regulates protein secretion in Pseudomonas aeruginosa. Nat. Cell Biol. 9, 797–803. doi: 10.1038/ncb1605
Nguyen, V. S., Logger, L., Spinelli, S., Desmyter, A., Le, T. T. H., Kellenberger, C., et al. (2015). Inhibition of type VI secretion by an anti-TssM Llama nanobody. PLoS ONE 10:e0122187. doi: 10.1371/journal.pone.0122187
Pessi, G., Williams, F., Hindle, Z., Heurlier, K., Holden, M. T. G., Cámara, M., et al. (2001). The global posttranscriptional regulator RsmA modulates production of virulence determinants and N-acylhomoserine lactones in Pseudomonas aeruginosa. J. Bacteriol. 183, 6676–6683. doi: 10.1128/JB.183.22.6676-6683.2001
Pietrosiuk, A., Lenherr, E. D., Falk, S., Bonemann, G., Kopp, J., Zentgraf, H., et al. (2011). Molecular basis for the unique role of the AAA+ chaperone ClpV in type VI protein secretion. J. Biol. Chem. 286, 30010–30021. doi: 10.1074/jbc.M111.253377
Planamente, S., Salih, O., Manoli, E., Albesa-Jové, D., Freemont, P. S., and Filloux, A. (2016). TssA forms a gp6-like ring attached to the type VI secretion sheath. EMBO J. 35, 1613–1627. doi: 10.15252/embj.201694024
Pukatzki, S., Ma, A. T., Revel, A. T., Sturtevant, D., and Mekalanos, J. J. (2007). Type VI secretion system translocates a phage tail spike-like protein into target cells where it cross-links actin. Proc. Natl. Acad. Sci. U.S.A. 104, 15508–15513. doi: 10.1073/pnas.0706532104
Pukatzki, S., Ma, A. T., Sturtevant, D., Krastins, B., Sarracino, D., Nelson, W. C., et al. (2006). Identification of a conserved bacterial protein secretion system in Vibrio cholerae using the Dictyostelium host model system. Proc. Natl. Acad. Sci. U.S.A. 103, 1528–1533. doi: 10.1073/pnas.0510322103
Pukatzki, S., McAuley, S. B., and Miyata, S. T. (2009). The type VI secretion system: translocation of effectors and effector-domains. Curr. Opin. Microbiol. 12, 11–17. doi: 10.1016/j.mib.2008.11.010
Rasamiravaka, T., Labtani, Q., Duez, P., and El Jaziri, M. (2015). The formation of biofilms by Pseudomonas aeruginosa: a review of the natural and synthetic compounds interfering with control mechanisms. BioMed Res. Int. 2015:759348. doi: 10.1155/2015/759348
Reimmann, C., Beyeler, M., Latifi, A., Winteler, H., Foglino, M., Lazdunski, A., et al. (1997). The global activator GacA of Pseudomonas aeruginosa PAO positively controls the production of the autoinducer N-butyryl-homoserine lactone and the formation of the virulence factors pyocyanin, cyanide, and lipase. Mol. Microbiol. 24, 309–319.
Russell, A. B., LeRoux, M., Hathazi, K., Agnello, D. M., Ishikawa, T., Wiggins, P. A., et al. (2013). Diverse type VI secretion phospholipases are functionally plastic antibacterial effectors. Nature 496, 508–512. doi: 10.1038/nature12074
Saak, C. C., and Gibbs, K. A. (2016). The self-identity protein IdsD is communicated between cells in swarming Proteus mirabilis colonies. J. Bacteriol. 198, 3278–3286. doi: 10.1128/JB.00402-16
Sana, T. G., Baumann, C., Merdes, A., Soscia, C., Rattei, T., Hachani, A., et al. (2015). Internalization of Pseudomonas aeruginosa strain PAO1 into epithelial cells is promoted by interaction of a T6SS effector with the microtubule network. mBio 6:e00712-15. doi: 10.1128/mBio.00712-15
Sana, T. G., Flaugnatti, N., Lugo, K. A., Lam, L. H., Jacobson, A., Baylot, V., et al. (2016). Salmonella Typhimurium utilizes a T6SS-mediated antibacterial weapon to establish in the host gut. Proc. Natl. Acad. Sci. U.S.A. 113, E5044–E5051. doi: 10.1073/pnas.1608858113
Sana, T. G., Hachani, A., Bucior, I., Soscia, C., Garvis, S., Termine, E., et al. (2012). The second type VI secretion system of Pseudomonas aeruginosa strain PAO1 is regulated by quorum sensing and fur and modulates internalization in epithelial cells. J. Biol. Chem. 287, 27095–27105. doi: 10.1074/jbc.M112.376368
Santin, Y. G., and Cascales, E. (2017). Domestication of a housekeeping transglycosylase for assembly of a type VI secretion system. EMBO Rep. 18, 138–149. doi: 10.15252/embr.201643206
Schuster, M., and Greenberg, E. P. (2007). Early activation of quorum sensing in Pseudomonas aeruginosa reveals the architecture of a complex regulon. BMC Genomics 8:287. doi: 10.1186/1471-2164-8-287
Shalom, G., Shaw, J. G., and Thomas, M. S. (2007). In vivo expression technology identifies a type VI secretion system locus in Burkholderia pseudomallei that is induced upon invasion of macrophages. Microbiology 153(Pt 8), 2689–2699. doi: 10.1099/mic.0.2007/006585-0
Sheng, L., Lv, Y., Liu, Q., Wang, Q., and Zhang, Y. (2013). Connecting type VI secretion, quorum sensing, and c-di-GMP production in fish pathogen Vibrio alginolyticus through phosphatase PppA. Vet. Microbiol. 162, 652–662. doi: 10.1016/j.vetmic.2012.09.009
Shneider, M. M., Buth, S. A., Ho, B. T., Basler, M., Mekalanos, J. J., and Leiman, P. G. (2013). PAAR-repeat proteins sharpen and diversify the type VI secretion system spike. Nature 500, 350–353. doi: 10.1038/nature12453
Si, M., Zhao, C., Burkinshaw, B., Zhang, B., Wei, D., Wang, Y., et al. (2017). Manganese scavenging and oxidative stress response mediated by type VI secretion system in Burkholderia thailandensis. Proc. Natl. Acad. Sci. U.S.A. 114, E2233–E2242. doi: 10.1073/pnas.1614902114
Silverman, J. M., Agnello, D. M., Zheng, H., Andrews, B. T., Li, M., Catalano, C. E., et al. (2013). Haemolysin co-regulated protein is an exported receptor and chaperone of type VI secretion substrates. Mol. Cell 51, 584–593. doi: 10.1016/j.molcel.2013.07.025
Silverman, J. M., Brunet, Y. R., Cascales, E., and Mougous, J. D. (2012). Structure and regulation of the type VI secretion system. Annu. Rev. Microbiol. 66, 453–472. doi: 10.1146/annurev-micro-121809-151619
Sun, S., Zhou, L., Jin, K., Jiang, H., and He, Y.-W. (2016). Quorum sensing systems differentially regulate the production of phenazine-1-carboxylic acid in the rhizobacterium Pseudomonas aeruginosa PA1201. Sci. Rep. 6:30352. doi: 10.1038/srep30352
Tian, Y., Zhao, Y., Wu, X., Liu, F., Hu, B., and Walcott, R. R. (2015). The type VI protein secretion system contributes to biofilm formation and seed-to-seedling transmission of Acidovorax citrulli on melon. Mol. Plant Pathol. 16, 38–47. doi: 10.1111/mpp.12159
Unterweger, D., Kostiuk, B., Otjengerdes, R., Wilton, A., Diaz-Satizabal, L., and Pukatzki, S. (2015). Chimeric adaptor proteins translocate diverse type VI secretion system effectors in Vibrio cholerae. EMBO J. 34, 2198–2210. doi: 10.15252/embj.201591163
Vettiger, A., and Basler, M. (2016). Type VI secretion system substrates are transferred and reused among sister cells. Cell 167, 99.e–110.e. doi: 10.1016/j.cell.2016.08.023
Wang, T., Si, M., Song, Y., Zhu, W., Gao, F., Wang, Y., et al. (2015). Type VI secretion system transports Zn2+ to combat multiple stresses and host immunity. PLoS Pathog. 11:e1005020. doi: 10.1371/journal.ppat.1005020
Weber, B. S., Hennon, S. W., Wright, M. S., Scott, N. E., de Berardinis, V., Foster, L. J., et al. (2016). Genetic dissection of the type VI secretion system in Acinetobacter and identification of a novel peptidoglycan hydrolase, TagX, required for its biogenesis. mBio 7:e01253-16. doi: 10.1128/mBio.01253-16
Wenren, L. M., Sullivan, N. L., Cardarelli, L., Septer, A. N., and Gibbs, K. A. (2013). Two independent pathways for self-recognition in Proteus mirabilis are linked by type VI-dependent export. mBio 4:e00374-13. doi: 10.1128/mBio.00374-13
Whitney, J. C., Chou, S., Russell, A. B., Biboy, J., Gardiner, T. E., Ferrin, M. A., et al. (2013). Identification, structure, and function of a novel type VI secretion peptidoglycan glycoside hydrolase effector-immunity pair. J. Biol. Chem. 288, 26616–26624. doi: 10.1074/jbc.M113.488320
Whitney, J. C., Quentin, D., Sawai, S., LeRoux, M., Harding, B. N., Ledvina, H. E., et al. (2015). An interbacterial NAD(P)+ glycohydrolase toxin requires elongation factor Tu for delivery to target cells. Cell 163, 607–619. doi: 10.1016/j.cell.2015.09.027
Zoued, A., Brunet, Y. R., Durand, E., Aschtgen, M.-S., Logger, L., Douzi, B., et al. (2014). Architecture and assembly of the type VI secretion system. Biochim. Biophys. Acta 1843, 1664–1673. doi: 10.1016/j.bbamcr.2014.03.018
Zoued, A., Cassaro, C. J., Durand, E., Douzi, B., España, A. P., Cambillau, C., et al. (2016a). Structure-function analysis of the TssL cytoplasmic domain reveals a new interaction between the type VI secretion baseplate and membrane complexes. J. Mol. Biol. 428, 4413–4423. doi: 10.1016/j.jmb.2016.08.030
Zoued, A., Durand, E., Brunet, Y. R., Spinelli, S., Douzi, B., Guzzo, M., et al. (2016b). Priming and polymerization of a bacterial contractile tail structure. Nature 531, 59–63. doi: 10.1038/nature17182
Zoued, A., Durand, E., Bebeacua, C., Brunet, Y. R., Douzi, B., Cambillau, C., et al. (2013). TssK is a trimeric cytoplasmic protein interacting with components of both phage-like and membrane anchoring complexes of the type VI secretion system. J. Biol. Chem. 288, 27031–27041. doi: 10.1074/jbc.M113.499772
Keywords: type six secretion system, quorum sensing, communication, social behavior, membrane perturbation
Citation: Gallique M, Bouteiller M and Merieau A (2017) The Type VI Secretion System: A Dynamic System for Bacterial Communication? Front. Microbiol. 8:1454. doi: 10.3389/fmicb.2017.01454
Received: 12 May 2017; Accepted: 18 July 2017;
Published: 28 July 2017.
Edited by:
Mickael Desvaux, Institut National de la Recherche Agronomique (INRA), FranceReviewed by:
Efstathios D. Giaouris, University of the Aegean, GreeceSophie Bleves, Aix-Marseille University, France
Copyright © 2017 Gallique, Bouteiller and Merieau. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Annabelle Merieau, YW5uYWJlbGxlLm1lcmllYXVAdW5pdi1yb3Vlbi5mcg==
 Mathias Gallique
Mathias Gallique Annabelle Merieau
Annabelle Merieau