- 1Division of Neonatology, Department of Pediatrics, College of Medicine, University of Florida, Gainesville, FL, United States
- 2Department of Microbiology and Cell Science, Genetics Institute, Institute of Food and Agricultural Sciences, University of Florida, Gainesville, FL, United States
- 3College of Nursing, University of Florida, Gainesville, FL, United States
- 4Department of Health Outcomes and Policy, College of Medicine, University of Florida, Gainesville, FL, United States
The American Academy of Pediatrics recommends that extremely preterm infants receive mother’s own milk (MOM) when available or pasteurized donor breast milk (DBM) when MOM is unavailable. The goal of this study was to determine whether DBM could be inoculated with MOM from mothers of preterm infants to restore the live microbiota (RM). Culture dependent and culture independent methods were used to analyze the fluctuations in the overall population and microbiome, respectively, of DBM, MOM, and RM samples over time. Using MOM at time = 0 (T0) as the target for the restoration process, this level was reached in the 10% (RM-10) and 30% (RM-30) mixtures after 4 h of incubation at 37∘C, whereas, the larger dilutions of 1% (RM-1) and 5% (RM-5) after 8 h. The diversity indexes were similar between MOM and DBM samples, however, different genera were prevalent in each group. Interestingly, 40% of the bacterial families were able to expand in DBM after 4 h of incubation indicating that a large percentage of the bacterial load present in MOM can grow when transferred to DBM, however, no core microbiome was identified. In summary, the microbiome analyses indicated that each mother has a unique microbiota and that live microbial reestablishment of DBM may provide these microbes to individual mothers’ infants. The agreement between the results obtained from the viable bacterial counts and the microbiome analyses indicate that DBM incubated with 10–30% v/v of the MOM for 4 h is a reasonable restoration strategy.
Introduction
The benefits of human milk for preterm infants include immune and nutritional protection against infection, decreased necrotizing enterocolitis (NEC), and other morbidities (Wight, 2001). In the absence of mother’s own milk (MOM), the American Academy of Pediatrics recommends using DBM over formula in very preterm infants (Moretti, 2012). Thus, the current practice at the University of Florida, UF Health Neonatal Intensive Care Unit (NICU), is to provide premature infants of gestational age less than 30 weeks with DBM if the MOM supply is low or not available.
Numerous components of human milk are thought to be beneficial for the infant (Newburg et al., 2005). Accumulating data suggests that microbes that are indigenous to human milk may not be contaminants and play a beneficial role for the infant (Jeurink et al., 2013). Up to 200 different bacterial species have been found in human milk. Hunt et al. (2011) studied milk samples from 16 healthy women collected at three different time points. A common group of nine bacterial genera was present in all samples, but in different concentrations among the subjects. Each individual demonstrated a unique milk microbiome that was stable over time (Hunt et al., 2011). Collectively, these data highlight a personalized collection of milk microbes from each mother that is optimized for the health of her own infant.
Donor breast milk used in most NICUs is pasteurized due to safety concerns (Landers and Updegrove, 2010). Pasteurization of DBM kills 99% of bacteria and may also inactivate a large proportion of the bioactive components. Since DBM is pooled and pasteurized, it lacks the unique live maternal milk microbiome, which may be of benefit to the infant (for a review see Jost et al., 2015). The majority of mothers of very preterm infants are able to express small amounts of their own milk and although it may be of insufficient volume to meet the daily nutritrional requirements of their infant, it can still provide lasting health benefits. Our objective was to encourage the NICU mothers to continue pumping and use a small amount of MOM to inoculate the pasteurized DBM to add back the potentially beneficial naturally occurring microbes. We hypothesized that fresh MOM can be mixed with DBM to improve the quality of DBM by cultivating the beneficial milk microbiome to reflect MOM. Studies were performed to determine optimal dilutions and incubation times to obtain a microbial content most similar to that of the MOM.
Materials and Methods
Subjects
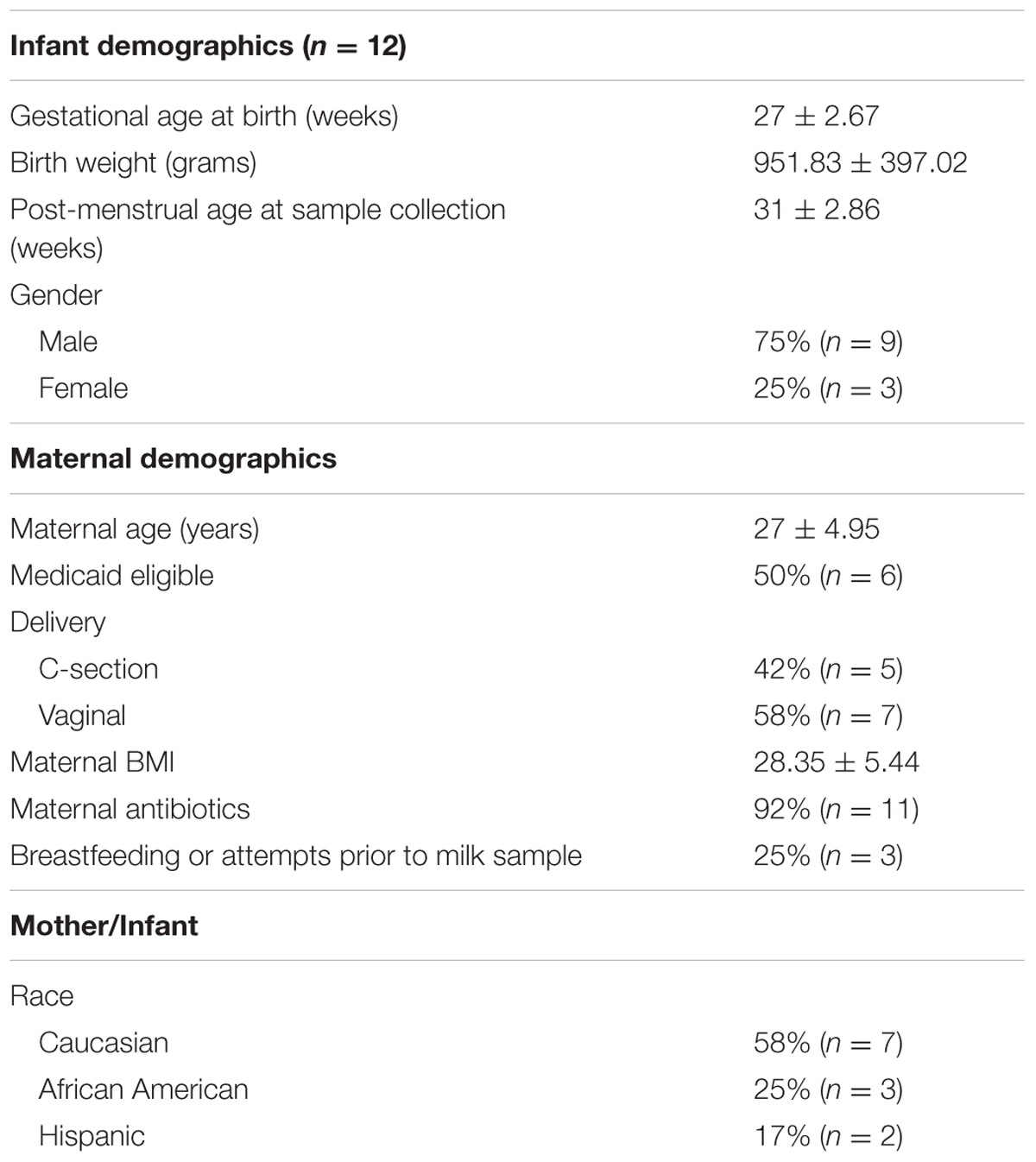
The study cohort consisted of twelve mothers who provided a breast milk sample between December 2014 and February 2016. Mothers had delivered an infant at less than 32 weeks gestation, weighing less than 1500 g at birth, and who were expressing over 100 mL of breast milk per day and producing at least 45 mL with each expression session. This pilot study was approved by the Institutional Review Board (IRB201400527) at the University of Florida and mothers provided written informed consent. Exclusion criteria included mothers who had delivered an infant with a chromosomal abnormality or who was severely ill and mothers who were currently taking antibiotics. Data collected included gender, gestational age at birth, gestational age at breast milk collection, birth weight, race, maternal age, parity, mode of delivery, maternal BMI, maternal infections, maternal medications, maternal antibiotics, and Medicaid eligibility. Baseline demographics are summarized in Table 1.
Milk Collection
Each mother pumped one sample of 45 mL of breast milk (MOM) into a sterile container using a new breast pump kit (Symphony Breast Pump Kit, Medela LLC, McHenry, IL, United States) and hospital grade electric breast pump (Symphony Breast Pump, Medela LLC, McHenry, IL, United States). Prior to sample collection, mothers were provided the verbal instructions regarding hand hygiene during milk expression and techniques for breast pump cleaning per NICU protocol. Immediately following collection, the samples were placed on ice and delivered to the laboratory for processing. The pasteurized DBM was obtained frozen from the Human Milk Banking Association of North America (HMBANA) milk bank (see http://rmchildren.org/mothers-milk-bank/donate-milk/collection-and-storage/ for details on the collection guidelines for donors) and were thawed immediatly prior to each restoration process.
Restoration of the Live Microbiome of DBM
The restoration strategy was to add increasing amounts of MOM (1, 5, 10, and 30% v/v) into pasteurized DBM. As controls, pure DBM and MOM were included. Once blended, the milk mixtures were incubated at 37°C. Samples were taken at time 0, 4, and 8 h. For viable bacterial counts, samples were analyzed immediately. For microbiome analyses, 2 mL were centrifuged at 12,000 rpm for 5 min at 4°C, the supernatant removed, and pellets stored at -80°C until the DNA was extracted. The pH and physical appearance of the samples at each time point was also recorded.
Culture Dependent Bacterial Analysis
Viable cell counts were determined by plating serial dilutions of each sample on selective and non-selective agar plates at time 0, 4, and 8 h (T0, T4, and T8, respectively). Based on the most common groups of bacteria cultivated from human milk (Wight, 2001; Martín et al., 2003; Reviriego et al., 2005; Jost et al., 2013; Menon and Williams, 2013), the following media were used: acidified Man, Rogosa and Sharp (MRS) agar for lactic acid bacteria (Fisher Scientific Company), Berens agar (BSM agar) for Bifidobacterium (Sigma–Aldrich) amended with BSM supplement Mupirocin Lithium Antibiotic (Fisher Scientific Company), Mannitol salt agar (MSA) for Staphylococcus (Fisher Scientific Company), Nutrient rich agar for facultative aerobes including Streptococcus (Sigma–Aldrich), and MacConkey agar for enterobacteria (Fisher Scientific Company). All plates were incubated at 37°C for 48–72 h. MRS and BSM agar plates were incubated in jars enclosing a burning candle to create a reduced oxygen environment while MacConkey and Nutrient agar plates remained incubated aerobically.
DNA Isolation, Library Construction and Sequencing
DNA was extracted from milk samples and preserved at -80°C using the PowerFecal® DNA isolation kit (MoBio Lab, Inc. United States) with the following modification: the pellet was homogenized in 750 μL of bead solution, then 100 μL of Protease from Streptomyces griseus 20 mg/mL (Sigma–Aldrich, Steinheim, Germany) was added (Marcial et al., 2017) The mixture was incubated at 37°C for 15 min, then the samples were processed according to the manufacturer’s protocol. In the elution step, the DNA was collected in 70 μL of water and quantified. The DNA concentration was standardized to 1 ng/μL before the amplification of the V4 region using primers 515F/806R barcoded for Illumina HiSeq platform (Caporaso et al., 2012). To reduce variability and potential bias from potential sources of DNA contamination, all samples were processed with the same batch of DNA extraction kits as well as PCR reagents.
Bioinformatics and Statistical Analysis
Clustering of Operational Taxonomic Units (OTUs) at 97% similarity was performed with the subsampled open-reference OTU picking method (Rideout et al., 2014) with no removal of singletons. The Greengenes reference dataset version 13.8 (DeSantis et al., 2006) was used as the reference for OTU picking and for taxonomy assignment with uclust (Edgar, 2010). OTUs identified as mitochondrial DNA or as chloroplasts were removed from further analyses using R studio.
Community structure was analyzed in R with phyloseq (McMurdie and Holmes, 2013) and plotted with ggplot2 (Wickham, 2009). Differences in taxonomic profiles were analyzed by Welch’s t-test (for two groups) or by ANOVA (for multiple groups) with Tukey-Kramer post hoc tests with STAMP (Parks et al., 2014) and PAST (Hammer et al., 2001).
Results
Mother’s Own Milk Shows a High Variability in the Number of Culturable Bacteria
At baseline t = 0, the amount of four bacterial populations were quantified in MOM and used as the target goal for each individual mother. It was found that the amount of bacteria in MRS plates (lactic acid bacteria including Lactobacillus) were between 103 and 105 CFU/mL with most of them being at 104 CFU/mL (Figure 1B). The bacterial load in MSA and nutrient broth were between 103 and 106 CFU/mL with an equal distribution in concentrations among the MOMs (Figures 1A,C). Only four MOM samples (30% of samples) grew on MacConkey agar at concentrations between 101 and 102 CFU/mL (data not shown). Bifidobacterium colonies were not recovered on Berens agar under our experimental conditions. The viable counts were also determined in DBM. Around 102 CFU/mL were counted in nutrient broth for half of the DBM samples (Figure 1C).
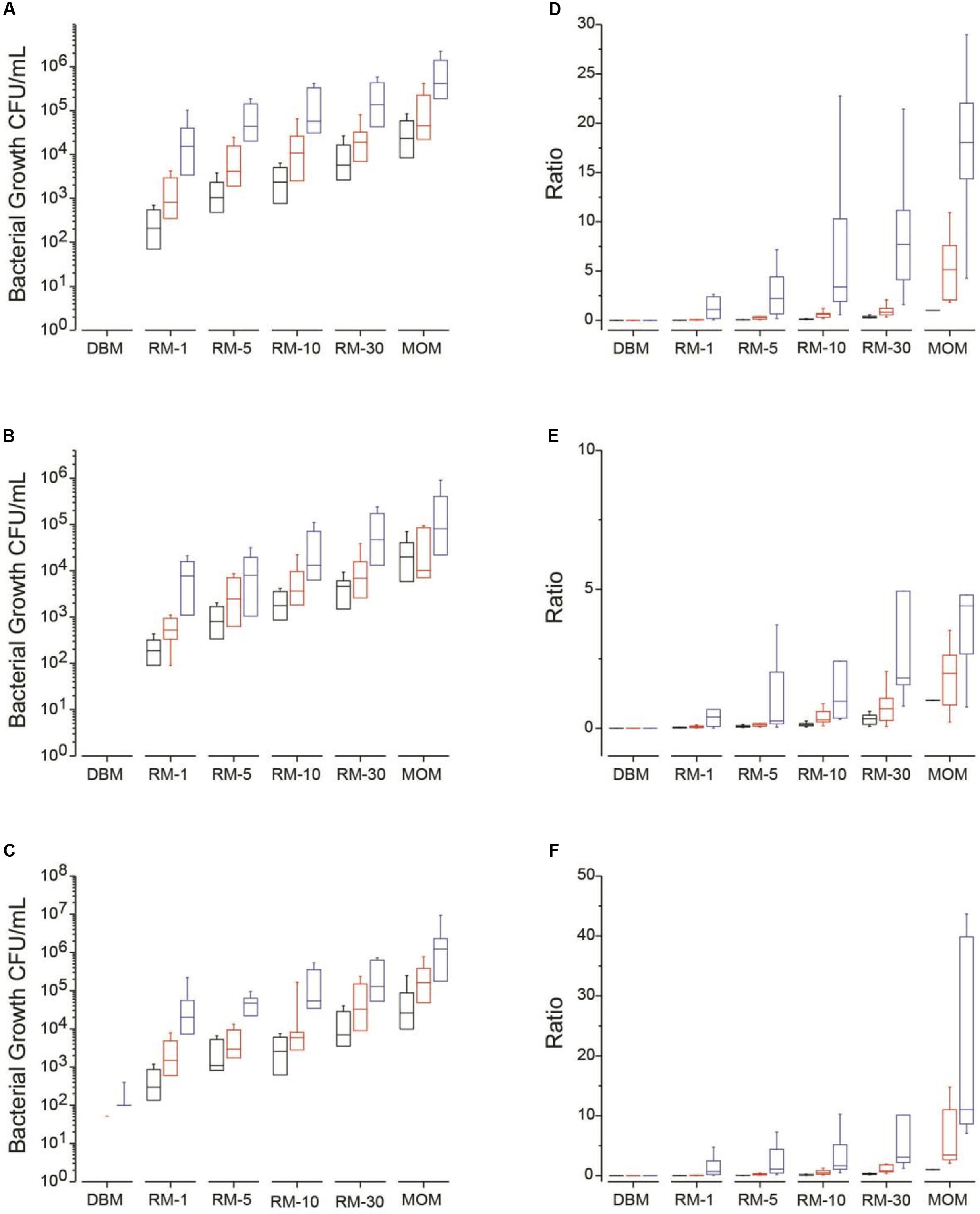
FIGURE 1. Median bacterial growth (CFU/mL) for each milk sample (DBM, RM-1, RM-5, RM-10, RM-30, and MOM) on different media types at time = 0 (T0, black boxes) and incubated over 4 h (T4, red boxes) and 8 h (T8, blue boxes) at 37 °C. Bacterial growth was determined on (A) MSA, (B) MRS and (C) on Nutrient media. Mean ratios of bacterial growth of RM samples to MOM (D–F). The ratio was calculated for each RM sample by dividing the CFU/mL in each RM sample to its corresponding MOM sample at T0 (MOM = 1). The ratios were calculated at each time point in (D) MSA, (E) MRS, and (F) Nutrient media.
Restoration of the Live Microbiome of DBM with Mother’s Own Milk
As described in the methods section, each of the twelve samples of MOM were inoculated in DBM at 1% (RM-1), 5% (RM-5), 10% (RM-10) and 30% (RM-30). Samples, including incubated DBM and MOM, were taken across three time points. The goal of this experiment was to determine the cultivable bacterial load of each sample to establish the minimum time and dilution required to reach the initial bacterial concentration found in MOM.
For each growth media, the concentration of bacteria was determined and ratios were calculated using MOM concentration at time 0 as the target concentration (represented as 1) (Figures 1D–F). On average, all RM samples increased in bacterial concentration over time (Figures 1A–C). A good correlation was found between the size of the inoculum and the amount of bacterial growth while the initial concentration of bacteria did not affect the outcome.
For MSA media, which targets mostly Staphylococcus, after 4 h of incubation at 37°C, 75% of the RM-10 reached a ratio of 0.6 compared to the MOM original bacterial load while all RM-30 reached a ratio of 1. For MRS media, which targets mostly lactic acid bacteria, 33% of the RM-10 reached a ratio of 1 compared to MOM while 58% of RM-30 reached the same ratio after 4 h. In Nutrient agar, which is a general purpose media targeting non-fastidious organisms, after 4 h of incubation 42% of samples in RM-10 reached a ratio of 1 compared to MOM while 83% reached a ratio of 1 in the RM-30 samples. After 8 h of incubation the bacterial load in all growth media tested (MSA, MRS, and Nutrient agar) for RM-10 and RM-30 went over the initial concentration of MOM (Figure 1). In contrast, the two highest milk dilution ratios (RM-1 and RM-5) were less than 0.5 at 4 h of incubation and then at 8 h of incubation reached a ratio > 1, exceeding the MOM bacterial load (Figure 1).
Since microbial growth may result in changes in pH as well as physical changes to the milk, the overall appearance (i.e., phase separation and curdling) and pH was monitored throughout the incubation period. Visual inspection of the milk samples did not reveal changes during the incubation period. Analysis of pH indicated that MOM samples were more alkaline (pH 7.5 ± 0.11) than pasteurized DBM (pH 6.5 ± 0.15) at T0. The incubation of MOM milk for 8 h resulted in a significant decrease (p = 0.047) in pH to 7 ± 0.5 while no significant changes were observed in DBM samples or RM samples over time.
Donor Milk versus Mother’s Own Milk Have a Similar Diversity Index
Illumina sequencing of the V4 region of the bacterial 16S rRNA was performed on all milk samples. After quality control, a total of 15,575,142 sequences were obtained with a mean of 75,976 sequences per sample. The rationale of this microbiome analyses was to have an unbiased view of the changes in the microbial community during the restoration process.
First, we compared the community structure of DBM and MOM at T0. The alpha diversity expressed as Chao1 and Shannon index was similar between the two sets of samples (Figure 2A). It cannot be concluded, however, that due to the similarity in alpha diversity between DBM and MOM that the bacterial load is the same since DBM is pasteurized. The relative abundance of genera was compared between the two samples (Figure 2B). Acinetobacter, Staphylococcus, Halomonas, Bacillus, Stenotrophomonas, unclassified Enterobacteriaceae genus, Streptococcus, Shewanella, Pseudomonas, Serratia, Enterococcus, unclassified Enterobacteriaceae genus, unclassified Methylobacteriaceae genus, unclassified Pseudomonadaceae genus, unclassified Xanthomonadaceae genus and Bacteroides constituted 85% of the sequences found in DBM. In MOM the most abundant genus were Halomonas, Staphylococcus, Shewanella, Corynebacterium, Enterobacteriaceae genus, Acinetobacter, unclassified Methylobacteriaceae genus, unclassified Enterobacteriaceae genus, Bacteroides, Stenotrophomonas and Lactobacillus. The statistical analyses showed that Halomonas (p < 0.01) and Shewanella (p < 0.01) were more abundant in MOM than in DBM samples. Similarly, Staphylococcus, Corynebacterium, and Lactobacillus were more abundant in MOM, yet did not reach statistical significance (p < 0.1) when compared to that of DBM. On the contrary, Acinetobacter, an unclassified Enterobacteriaceae genus, and Serratia, showed a significantly higher relative abundance (p < 0.05) in DBM.
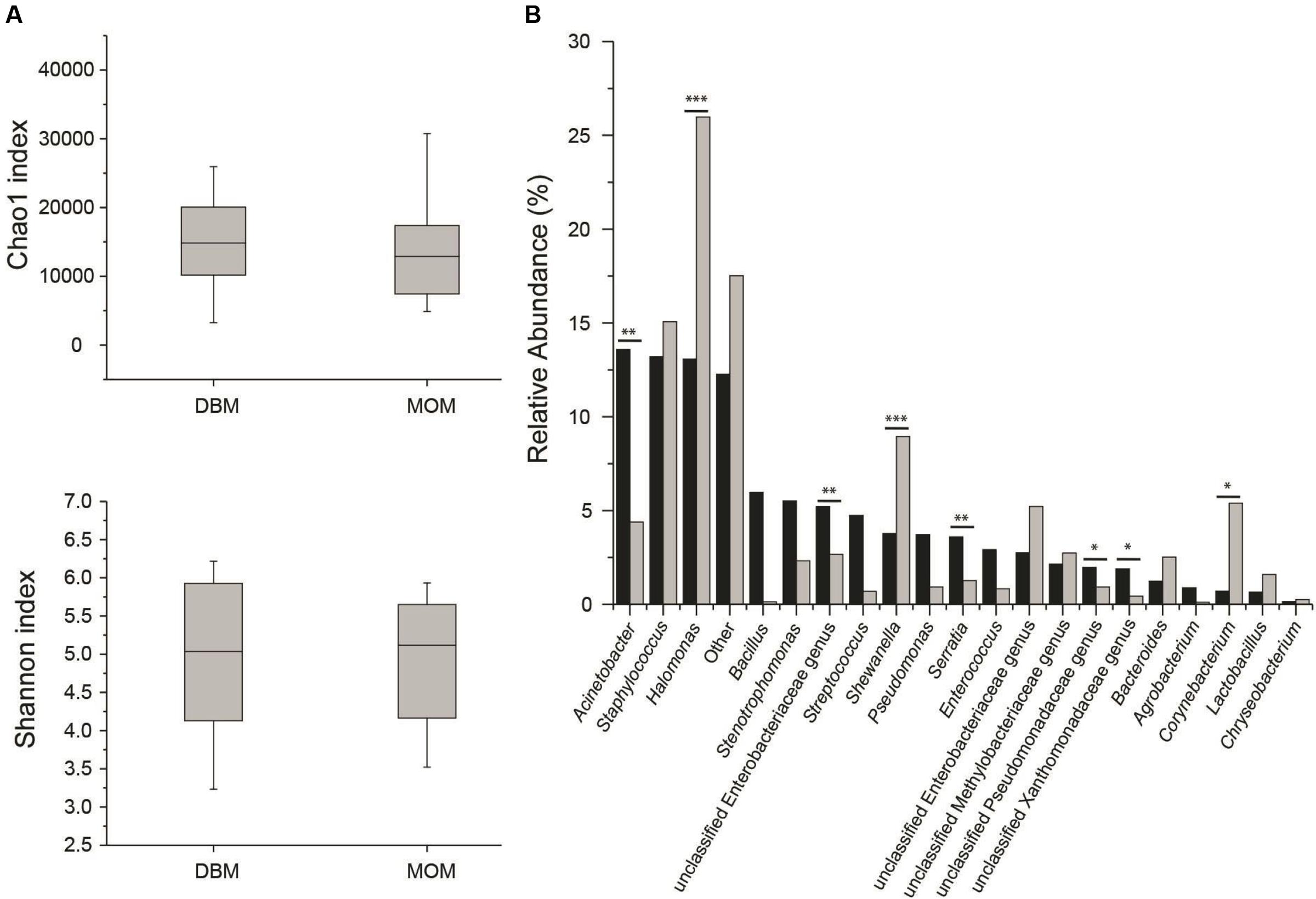
FIGURE 2. (A) Alpha diversity determinations of DBM and MOM at T0 using Chao1 and Shannon indexes. (B) Summary of the most abundant genera found in DBM (black) and MOM (gray) samples. Statistically significant values are indicated ∗∗p < 0.05 and ∗∗∗p < 0.01. Differences at ∗p < 0.1 are also indicated.
Fluctuations in the Alpha Diversity of RM Samples during Microbial Restoration
Next, the fluctuations in the microbial community as a result of the restoration process was determined. As described earlier, the working hypothesis is that the restoration process will result in the expansion of the microbial population without loss of diversity. The expansion of MOM was used as a positive control to determine microbial populations that will be able to expand in vitro. The Shannon index was utilized to determine the species richness across time between DBM, MOM, and the RM samples (Figure 3A). It was found that after 4 h of incubation the MOM samples had a significant decrease (p < 0.01) in alpha diversity only when compared to DBM and RM-1. In contrast, after 8 h of incubation, there is a significant decrease (p < 0.001) in diversity when comparing MOM to all other samples (DBM and RM). However, even when a decrease in diversity was observed in RM-10 and RM-30, they did not reach statistical significance (p = 0.07 and p = 0.14, respectively). The change in diversity of MOM and RM-10 or RM-30 suggests the differential growth of few bacterial species.
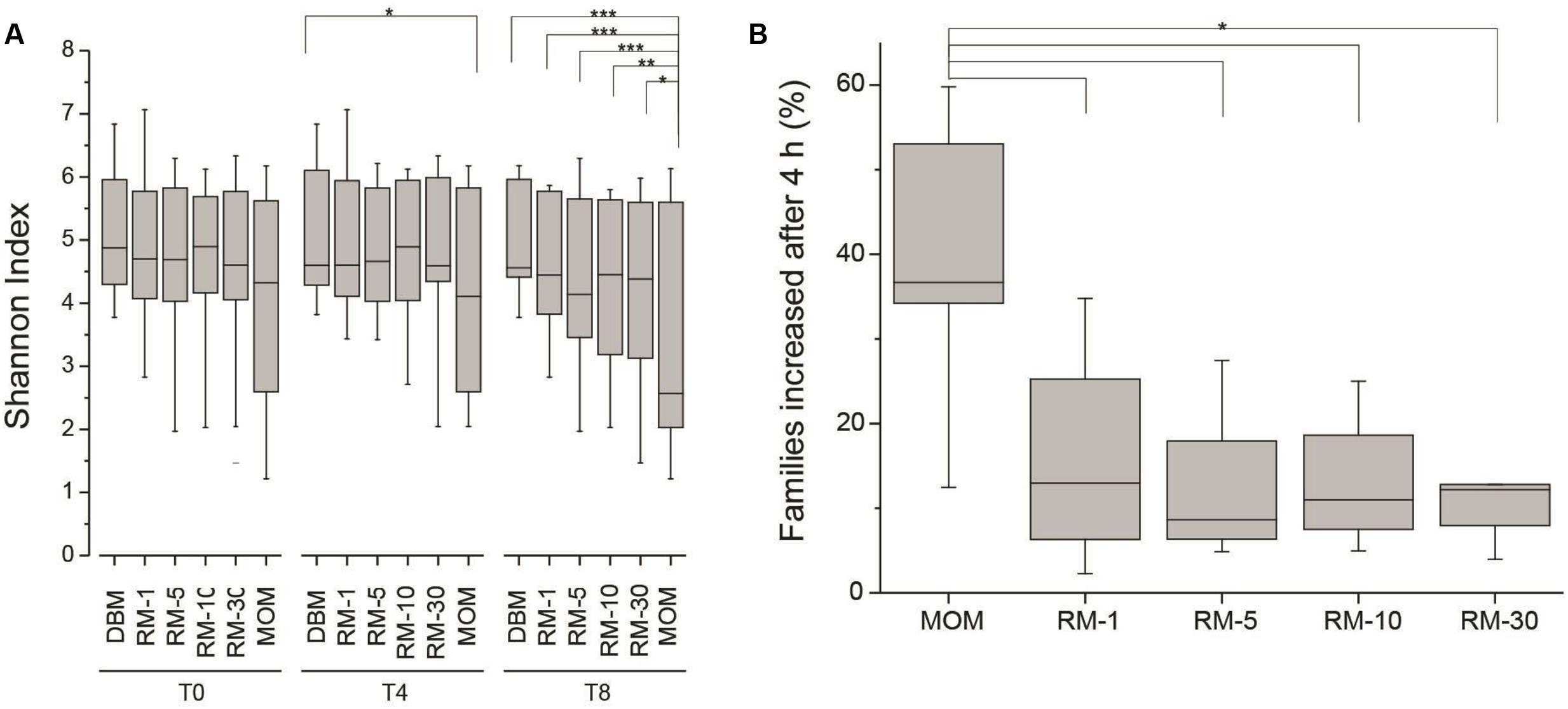
FIGURE 3. (A) Alpha diversity assessment in RM samples (RM-1, RM-5, RM-10 and RM-30, across time points (0, 4, and 8 h). DBM and MOM samples were included in the analyses. It was found that DBM and MOM are significantly different from each other at T4 and T8. The alpha diversity of the 30% dilution of MOM at T4 resembles that of MOM at T0. Statistically significant values are indicated as ∗p < 0.05, ∗∗p < 0.01, and ∗∗∗p < 0.001. (B) Summary of the expansion of bacterial families after 4 h of incubation. The expansion of bacterial families was determined for each sample by substracting the relative abundancy at T4 from the relative abundancy at T0. Statistically significant values are indicated as ∗p < 0.001.
The analysis of the bacterial richness of the RM allowed for the assessment of the expanding microbiome to determine if the RM samples, over time, become similar to MOM at T0. A multidimensional scaling (MDS) plot was generated to visualize the variability of the microbial community in each MOM and their derived RM samples (Figure 4). It was expected that the DBM samples of each individual set should remain clustered together since DBM is pasteurized and should have negative to little bacterial growth. In most cases, the DBM samples remained clustered together, such as MOM sample 4, 5, 6, 7, 8, 10, 11, and 12 (shown as circles in Figure 4) whereas, in DBM used for MOM samples 2 and 9 has a small difference in bacterial richness over time. Overall, these MDS plots allowed the visualization of fluctuations of the microbial population in the different RM samples over time. In some mothers, it was observed that the different RM samples migrated toward MOM at T0, suggesting that the RM sample became more similar to MOM at T0 (Figure 4). However, some exceptions were observed. For example, all samples derived from MOM 3 cluster together, indicative that the restoration process was not successful. Mothers 11 and 12 show migration of all the dilutions moving away from MOM at all time-points and toward DBM instead. Mothers 11 and 12 shared the same sample of DBM so it is possible that live bacteria still found within this DBM sample are hindering the expansion of MOM-derived microbiota.
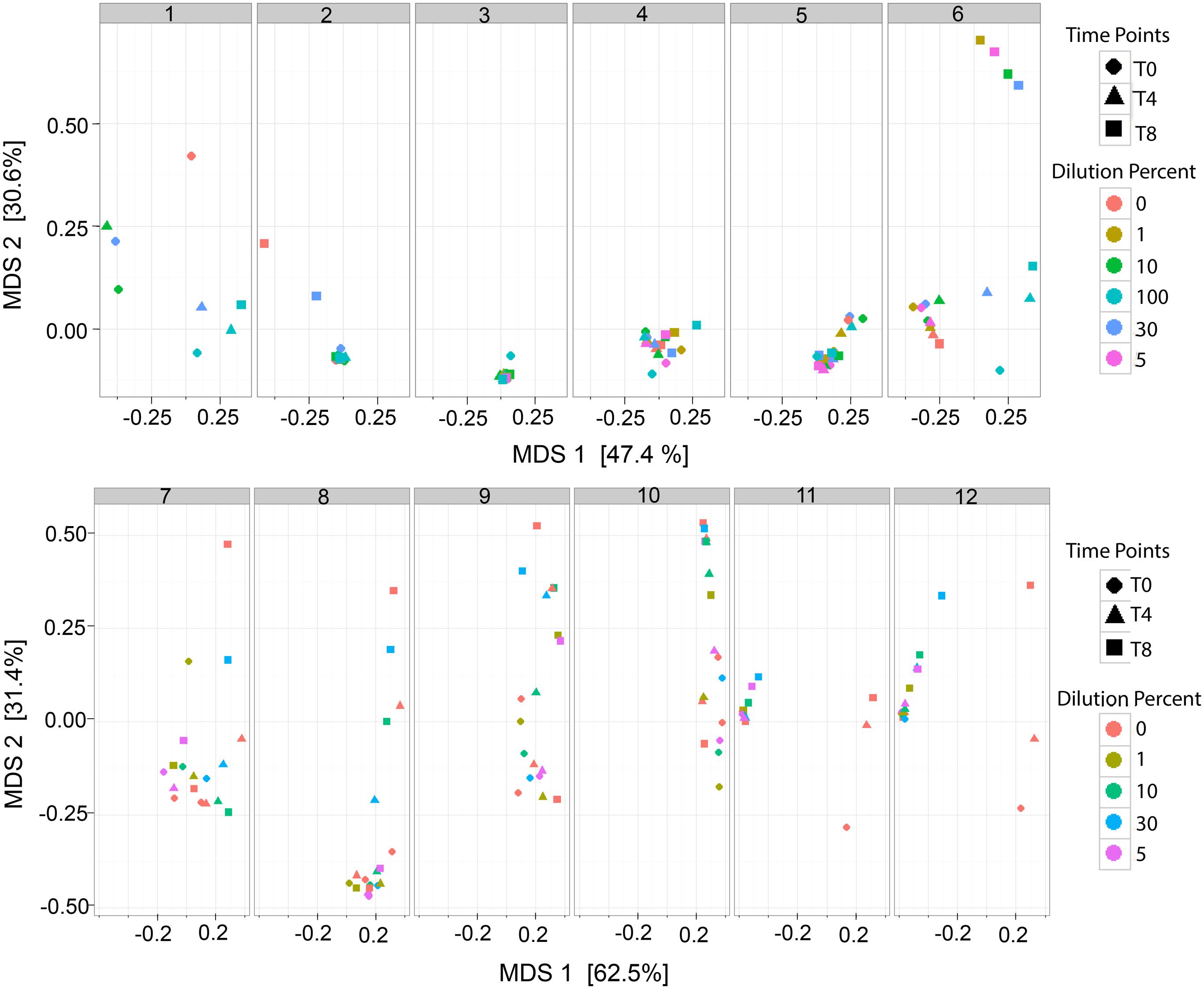
FIGURE 4. Multidimensional scaling (MDS) plots of RM samples. Two dimensional clustering determined by the distance of each sample to show individual clustering in each MOM derived subset. Each time point (T0, T4, and T8) is signified by a shape (Circle, Triangle, and Square, respectively). Each dilution is signified as a color. For MOM 1 and MOM2 samples RM-1 and RM-5 were not available. For MOM1, only DBM at T0 (DBM = 0 and T0 = circle) is shown.
RM Samples Do Not Share a Common Expanding Microbiome
Next, we determined if a core microbiome was expanding in all the RM samples. Based on the culture dependent results where the target bacterial load (of the culturable population) was reached after 4 h and a decrease in diversity was observed in the microbiome after 8 h of incubation, the next series of analyses were performed on all the samples (DBM, RM, and MOM) only after 4 h of incubation.
For these analyses the relative abundance of each bacterial family was obtained. To identify families that increase in abundance after 4 h, the bacterial load from T0 was subtracted from the bacterial load from T4 across all samples. If a family increased in concentration within the negative control (DBM), it was excluded in the results. It was found that out of the 120 ± 40 families identified within MOM samples, an average of 37 ± 12 were able to increase concentration after 4 h of incubation. Interestingly, in all RM samples, 23–38% of the families were able to increase in concentration (Figure 3B).
All bacterial genera within the families that increased in concentration after 4 h of incubation were analyzed. Some genera showed some statistical trends like Gemella, unclassified Gemellales genus, Salinicoccus, Gallionella, unclassified Proteobacteria genus 1, unclassified Proteobacteria genus 2, Shewanella, Alicyclobacillus, Pediococcus, and Lactobacillus, however, no significant differences (p < 0.2) were observed (data not shown). In contrast, Staphylococcus showed a significant increase in concentrations (p < 0.01) (data not shown). In summary, a common core of microbes that expand in RM samples was not identified, with the exception of Staphylococcus.
Mother’s Own Milk Samples Cluster by the Birth Mode of Delivery
Unable to determine a common expanding microbiome among RM samples, the data was stratified and analyzed considering the birth mode of delivery. The principal component analysis using MOM samples at T0, clustered into c-section and vaginal delivery with the exceptions of mothers 6 and 7 (Supplementary Figure S1). The distribution of the 15 most abundant genera among samples is shown in Supplementary Figure S2.
The microbiome of MOM samples divided by mode of delivery (c-section n = 5 and vaginal n = 7) were analyzed using a Welch’s t-test (Supplementary Table S1). It was found that Erwinia (p = 0.03) and Pseudomonas (p = 0.03) were more prevalent among c-section samples while Halomonas (p = 0.02), Lactobacillus (p = 0.04), Prevotella (p = 0.04), Ruminococcus (p = 0.04), unclassified Clostridiaceae genus (p = 0.03), and unclassified Enterococcaceae genus (p = 0.02) were found at higher concentrations in vaginal delivery samples (Supplementary Table S1). Other genera that had p-values greater than 0.05 and less than 0.13 were also taken into consideration for further analyses. These results indicate that the differences in genera observed between the mode of delivery of MOM might explain the inability to identify a core microbiome that expand in RM samples.
The data from RM samples was stratified by mode of delivery and the genera that were differentially found between c-section and vaginal delivery were analyzed (Supplementary Table S1). The relative increase in abundace after 4 h of incubation was tested using ANOVA and Tukey’s post hoc tests. It was found that most genera did not change significantly. Agrobacter showed a significant difference (p < 0.05) between vaginal RM-10 and c-section RM-10. Interestingly, 3 unclassified genera (unclassified Clostridiaceae genus, unclassified Enterococcaceae genus 2, and unclassified Methylophilaceae genus) increased in relative abundace in all c-section RM samples but not in vaginal delivery birth RM samples (data not shown). These results suggest that each MOM has a unique diverse bacterial load, having no core microbes.
Discussion
In this work we show that by using a small amount of MOM to inoculate pasteurized DBM, it is possible to reestablish the potentially beneficial naturally occurring microbes. MOM contains irreplaceable immune modulating factors including commensal bacteria (Wold and Adlerberth, 2002; Jost et al., 2013). Feeding preterm infants MOM has been shown to decrease NEC and sepsis with even small amounts of MOM providing some protection (Furman et al., 2003; Schanler, 2005; Meinzen-Derr et al., 2009; Corpeleijn et al., 2012, 2016; Abrams et al., 2014; Chowning et al., 2016). The microbiome found in MOM may provide short term and long term benefits to infants including preventing colonization by pathogens, stimulating production of cross-reactive antibiodies, and possibly establishing a healthy intestinal microbiome which may prevent long term morbidities including obesity, type 2 diabetes, chronic intestinal inflammation, autoimmune disorders, allergy, irritable bowel syndrome, and allergic gastroenteritis (Goulet, 2015; Lemas et al., 2016; Wallace et al., 2016). Unfortunately, many mothers of preterm infants are unable to produce sufficient amounts of breast milk to sustain 100% of their infant’s nutritional needs (Smith et al., 2003; Lee and Gould, 2009). The RM may offer a personalized and beneficial alternative to DBM when MOM is limited. Since MOM contains a unique and unchanging microbiome (Hunt et al., 2011; Cacho and Neu, 2014), providing infants their own mother’s milk may be beneficial, especially for infants born preterm, at risk for infection and other premature specific morbidities.
Our culture dependent approach indicated that the main bacterial groups that were tested, namely Staphylococcus, lactic acid bacteria (and other that can grow in MRS agar), along with other facultative aerobes (including Streptococcus), can be propagated into DBM, however, a large variability in bacterial load was observed between mothers. Previous studies examining bacteria present in breast milk have required aseptic sample collection of milk by mothers to limit skin flora and potential contaminants (Martín et al., 2003, 2007; Collado et al., 2009; Hunt et al., 2011; Jost et al., 2013; Khodayar-Pardo et al., 2014). In contrast, we aimed to examine the typical microbes that the preterm babies consistently received from their mother’s milk during routine NICU expression practices and therefore, mothers performed routine NICU protocols for hand hygiene techniques and equipment cleaning. Consistent with Heikkilä and Saris (2003) and Martín et al. (2003), our results reveal that Staphylococcus, facultative aerobes, lactic acid bacteria, and few Gram negatives can be cultured from breast milk. We were not able to isolate Bifidobacterium in culture which may be due to the sensitive anaerobic nature of Bifidobacterium, having not provided the proper anaerobic environment or handling techniques to facilitate growth. In addition, it is possible that Bifidobaceterium is negligible in breast milk from mothers delivering preterm infants as indicated from the study by Khodayar-Pardo et al. (2014), which showed less Bifidobacterium compared to milk from mothers of term infants. Small numbers of Gram negative bacteria grew on MacConkey agar (results not shown), which is consistent with previous research indicating that approximately 30–70% of breast milk samples contained Gram negative bacteria (Botsford et al., 1986; Landers and Updegrove, 2010; Keim et al., 2013). Small amounts of Gram negative bacteria in breast milk may be important in preparing the immune system to utilize toll-like receptor mediated tolerizing mechanisms to prevent an exaggerated response to future Gram negative bacteria (Madara, 2004).
Interestingly, pasteurized DBM obtained from HMBANA showed that 102 CFU/mL of bacteria were still present in nutrient broth (facultative anaerobes including Staphylococcus and Streptococcus) in 44% of the DBM samples. This is consistent with previous studies describing the presence of staphylococcal species in DBM as well as spore forming bacteria such as Bacillus cereus (Crielly et al., 1994; Landers and Updegrove, 2010; Decousser et al., 2013; Akindolire et al., 2015; Dewitte et al., 2015). Although microbial diversity indexes were similar between DBM and MOM, the most abundant genera differed. The most prevalent genera in MOM were Halomonas, Shewanella, Corynebacterium, Staphylococcus, and Lactobacillus, while the most common genera in DBM were Acinetobacter, unclassified Enterobacteriaceae, and Serratia. These differences may reflect variances in the mothers who provided breast milk samples and the sanitation or method used for collection. DBM is typically obtained from mothers who are breastfeeding term infants who are often more than 6 months old. In contrast, the MOM in this study was obtained from mothers of preterm, hospitalized infants who because of their infant’s prematurity, were unable to breastfeed and were thus dependent on mechanical breast milk expression to obtain milk for their infants.
We found that Halomonas was present in greater abundance than Staphylococcus in comparison with other studies in MOM. Although Halomonas has not been previously described in breast milk from mothers delivering preterm, Staphylococcus is a well-known predominant phyla in breast milk (Hunt et al., 2011; Urbaniak et al., 2012, 2016; Jost et al., 2013). We also found Shewanellaceae was high in MOM (p < 0.05) compared to DBM. The main genera found to be of great abundance in MOM across different studies are Staphylococcus, Streptococcus, Proteobacteria groups, and Propionibacteria (Hunt et al., 2011; Kumar et al., 2016). However, Bifidobacterium, Bacteroides, Parabacteroides and Clostridia groups have also been identified to be a part of the breast milk microbiota (Sinkiewicz and Ljunggren, 2008; Kumar et al., 2016). Although Bifidobacterium was identified from our sequencing results, it was not shown to be prevalent. These differences in the predominant genera being Halomonas and Shewanellaceae as opposed to Staphylococcus and Streptococcus may be due to the fact that (except for rare occasions in three patients) mothers in our study did not breastfeed their infants. They were dependent on mechanical breast pumps for milk removal, thus the mother’s breasts were not routinely exposed to oral microbes from the infant’s mouth. Other reasons for these variations may be attributed to younger gestational age at delivery, stage of lactation, milk collection strategies and geographic variations in our study compared to others. In DBM, the most abundant genera were Acinetobacter, unclassified Enterobacteriaceae genus, and Serratia.
Results of this study suggest that the optimal restoration strategy to reach a microbial content most similar to MOM was a mixture of RM-10 incubated for 4 h. Using the bacterial load and microbial content of MOM at T0 as the target, we were able to successfully restore the microbiome in the RM-10 and RM-30 mixtures after 4 h of incubation, whereas larger dilutions of RM-1 and RM-5 reached the target level after 8 h. Although it did not reach the microbial content of MOM, RM-10 was able to reach 60% of the bacterial load of MOM. In contrast, RM-30 exceeded the target goal of MOM in the majority of incubated samples. This may be clinically undesireable since potentially pathogenic bacterial strains may grow to possibly harmful levels. Overall, our results demonstrate that inoculation with an amount of MOM as small as 1% can populate DBM with the mother’s potentially beneficial bacteria.
The alpha diversity of MOM and the larger dilutions of RM (RM-10 and RM-30) decreased as incubation time increased. In contrast, the diversity of DBM and smaller dilutions of RM (RM-1 and RM-5) remained similar to their original levels as incubation time increased. This trend of MOM toward decreased diversity suggests replication of only a few microbial species. Breast milk studies show that microbial diversity is associated with a healthy lactating milk microbiome as opposed to the milk microbiome of a mother with mastitis where Staphylococcus or Streptococcus species predominate (Delgado et al., 2008). This decreased diversity indicates that in order to preserve microbial diversity similar to that found in MOM, a 4 h incubation time compared to an 8 h incubation time may be optimal, which confirms the culture based results favoring a 4 h incubation time over 8 h for the RM.
Birth mode of delivery has been shown to affect the microbiota of breast milk in the majority of studies, suggesting a difference in the milk microbiome between infants born via c-section and those born vaginally (Azad et al., 2013; Gregory et al., 2015; Liu et al., 2015; Brumbaugh et al., 2016; Dominguez-Bello et al., 2016; Lee et al., 2016; Nagpal et al., 2016; Rutayisire et al., 2016). In our study, we observed a differential clustering of microbiomes from the breast milk of c-sections versus vaginally delivering mothers. Although it is clear each individual mother’s milk microbiota has bacterial variability, c-section and vaginal deliveries cluster with one another. Further analysis illustrated that bacterial genera most prevalent in breast milk from vaginal deliveries were Halomonas, Lactobacillus, Prevotella, unclassified Clostridiaceae genus, Clostridium, Comamonas, and Dorea. Those genera most prevalent in breast milk from cesarean deliveries were Erwinia, Pseudomonas, Ruminococcus, unclassified Enterococcaceae genus, Agrobacterium, Citrobacter, Enterococcus, Klebsiella, unclassified Bacilli genus, unclassified Bradyrhizobiaceae genus, and unclassified Methylophilaceae genus. Further statistical analysis does not show a common microbiota across breast milk regardless of c-section or vaginal delivery, concluding that the microbiota of breast milk is variable between each mother.
Based on our results, restoration of the live microbiome of DBM with MOM appears to be a promising and innovative method to provide preterm infants with beneficial breast milk bacteria. It is well known that breast milk changes over time to meet specific needs of infants which are attributed to stages of lactation, gestational age, infant feeding, and the health status of the breastfeeding dyad (Daly et al., 1996; Kent, 2006; Hassiotou et al., 2013). If mothers of preterm infants are able to express even minimal amounts of breast milk, restoration of the microbiome in DBM may allow their infants to receive milk more specific to their individual needs based on the stage of lactation and gestational age, thereby potentially improving their overall health. We used non-culture based techniques to take a snapshot of the full range of bacteria present in fresh preterm milk and pasteurized donor milk. The main limitation of our study is its small sample size and the use of antibiotics during the peripartum period. Nevertheless, it addresses the concept that the live microbiota donor human milk can be effectively restablished by MOM. In addition, mothers were not required to clean their breasts prior to breast milk sampling so their samples may have contained a higher level of skin colonizing microbes. Another limitation of the study is that safety parameters were not assessed. Future studies will need to include the analyses of potential pathogenic bacterial groups that may proliferate in the RM samples in the NICU environment.
In summary, we have shown that each mother has a unique milk microbiota and that the live microbiome in DBM can be restored with these unique bacteria using small amounts of MOM. This is a novel approach to possibly improving the bioactivity of DBM by adding specific MOM microbes in small quantities to personalize her own infant’s milk. Personalizing DBM may benefit the mother–infant dyad and contribute to a more robust infant intestinal microbiome. The agreement between the results obtained from the viable bacterial counts and the microbiome analyses indicate that DBM incubated with 10 percent of the MOM for 4 h is a reasonable restoration strategy. Future studies should include larger samples sizes, activity of the microbes in RM in comparison to DBM and MOM samples, and clinical evaluation of the safety and efficacy.
Ethics Statement
This pilot study was carried out in accordance with the recommendations of the Institutional Review Board (# 201400527) at the University of Florida with written informed consent from all subjects. All subjects gave written informed consent in accordance with the Declaration of Helsinki. The protocol was approved by the Institutional Review Board at University of Florida.
Author Contributions
The authors’ responsibilities were as follows: NC, LP, JN, and GL designed the research; NC, NH, KP, GM, LC, and NL performed the research; DL contributed new reagents/analytic tools; NC, NH, LC, LP, and GL analyzed the data; NC, NH, LP, JN, and GL evaluated the data; NC, NH, LP, JN, and GL wrote the manuscript; NC, NH, LP, JN, and GL had primary responsibility for the final content; All authors read and approved the final manuscript. Medela, AG was not involved in the implementation, data collection, statistical analysis, interpretation of date, or manuscript preparation and writing.
Funding
Funding for this study was provided by Medela, AG.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors would like to thank the mothers who generously provided milk samples for the study. We acknowledge our dedicated neonatal nurses for taking excellent care our patients. We also acknowledge Christopher Gardner and Tara Miller for their technical help. Finally, we acknowledge Medela Company for supplying the new sterile pumping kits used in this study.
Abbreviations
DBM, donor breast milk; MOM, mother’s own milk; NICU, Neonatal Intensive Care Unit; RM, restored microbiota pasteurized donor milk.
Supplementary Material
The Supplementary Material for this article can be found online at: http://journal.frontiersin.org/article/10.3389/fmicb.2017.01470/full#supplementary-material
FIGURE S1 | This is a principal component analysis (PCA) of the microbiota of MOM samples at T0. The circles represent moms that delivered through C-Section and the squares are those that delivered through vaginal births.
FIGURE S2 | Heatmap showing the top 15 genera of each DBM sample, with the exception of replicates DBM 1, 2 and 11 (DBM 1 and 2 being a replicates of 3 and DBM 11 being a replicate of 12), and each MOM sample.
References
Abrams, S. A., Schanler, R. J., Lee, M. L., and Rechtman, D. J. (2014).Greater mortality and morbidity in extremely preterm infants fed a diet containing cow milk protein products. Breastfeed. Med. 9, 281–285. doi: 10.1089/bfm.2014.0024
Akindolire, M., Babalola, O., and Ateba, C. (2015). Detection of antibiotic resistant Staphylococcus aureus from milk: a public health implication. Int. J. Environ. Res. Public Health 12, 10254–10275. doi: 10.3390/ijerph120910254
Azad, M. B., Konya, T., Maughan, H., Guttman, D. S., Field, C. J., Chari, R. S., et al. (2013). Gut microbiota of healthy Canadian infants: profiles by mode of delivery and infant diet at 4 months. Can. Med. Assoc. J. 185, 385–394. doi: 10.1503/cmaj.121189
Botsford, K. B., Weinstein, R. A., Boyer, K. M., Nathan, C., Carman, M., and Paton, J. B. (1986). Gram-negative bacilli in human milk feedings: quantitation and clinical consequences for premature infants. J. Pediatr. 109, 707–710. doi: 10.1016/S0022-3476(86)80246-3
Brumbaugh, D. E., Arruda, J., Robbins, K., Ir, D., Santorico, S. A., Robertson, C. E., et al. (2016). Mode of delivery determines neonatal pharyngeal bacterial composition and early intestinal colonization. J. Pediatr. Gastroenterol. Nutr. 63, 320–328. doi: 10.1097/MPG.0000000000001124
Cacho, N., and Neu, J. (2014). Manipulation of the intestinal microbiome in newborn infants. Adv. Nutr. 5, 114–118. doi: 10.3945/an.113.004820
Caporaso, J. G., Lauber, C. L., Walters, W. A., Berg-Lyons, D., Huntley, J., Fierer, N., et al. (2012). Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624. doi: 10.1038/ismej.2012.8
Chowning, R., Radmacher, P., Lewis, S., Serke, L., Pettit, N., and Adamkin, D. H. (2016). A retrospective analysis of the effect of human milk on prevention of necrotizing enterocolitis and postnatal growth. J. Perinatol. 36, 221–224. doi: 10.1038/jp.2015.179
Collado, M. C., Delgado, S., Maldonado, A., and Rodríguez, J. M. (2009). Assessment of the bacterial diversity of breast milk of healthy women by quantitative real-time PCR. Lett. Appl. Microbiol. 48, 523–528. doi: 10.1111/j.1472-765X.2009.02567.x
Corpeleijn, W. E., de Waard, M., Christmann, V., van Goudoever, J. B., Jansen-van der Weide, M. C., Kooi, E. M. W., et al. (2016). Effect of donor milk on severe infections and mortality in very low-birth-weight infants: the early nutrition study randomized clinical trial. JAMA Pediatr. 170, 654–661. doi: 10.1001/jamapediatrics.2016.0183
Corpeleijn, W. E., Kouwenhoven, S. M. P., Paap, M. C., van Vliet, I., Scheerder, I., Muizer, Y., et al. (2012). Intake of own mother’s milk during the first days of life is associated with decreased morbidity and mortality in very low birth weight infants during the first 60 days of life. Neonatology 102, 276–281. doi: 10.1159/000341335
Crielly, E. M., Logan, N. A., and Anderton, A. (1994). Studies on the Bacillus flora of milk and milk products. J. Appl. Bacteriol. 77, 256–263. doi: 10.1111/j.1365-2672.1994.tb03072.x
Daly, S. E., Kent, J. C., Owens, R. A., and Hartmann, P. E. (1996). Frequency and degree of milk removal and the short-term control of human milk synthesis. Exp. Physiol. 81, 861–875. doi: 10.1113/expphysiol.1996.sp003982
Decousser, J.-W., Ramarao, N., Duport, C., Dorval, M., Bourgeois-Nicolaos, N., Guinebretière, M.-H., et al. (2013). Bacillus cereus and severe intestinal infections in preterm neonates: Putative role of pooled breast milk. Am. J. Infect. Control 41, 918–921. doi: 10.1016/j.ajic.2013.01.043
Delgado, S., Arroyo, R., Martín, R., and Rodríguez, J. M. (2008). PCR-DGGE assessment of the bacterial diversity of breast milk in women with lactational infectious mastitis. BMC Infect. Dis. 8:51. doi: 10.1186/1471-2334-8-51
DeSantis, T. Z., Hugenholtz, P., Larsen, N., Rojas, M., Brodie, E. L., Keller, K., et al. (2006). Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072. doi: 10.1128/AEM.03006-05
Dewitte, C., Courdent, P., Charlet, C., Dumoulin, D., Courcol, R., and Pierrat, V. (2015). Contamination du lait maternel par une flore aérobie: évaluation des pertes pour un lactarium. Arch. Pédiatrie 22, 461–467. doi: 10.1016/j.arcped.2015.02.011
Dominguez-Bello, M. G., De Jesus-Laboy, K. M., Shen, N., Cox, L. M., Amir, A., Gonzalez, A., et al. (2016). Partial restoration of the microbiota of cesarean-born infants via vaginal microbial transfer. Nat. Med. 22, 250–253. doi: 10.1038/nm.4039
Edgar, R. C. (2010). Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461. doi: 10.1093/bioinformatics/btq461
Furman, L., Taylor, G., Minich, N., and Hack, M. (2003). The effect of maternal milk on neonatal morbidity of very low-birth-weight infants. Arch. Pediatr. Adolesc. Med. 157, 66–71. doi: 10.1001/archpedi.157.1.66
Goulet, O. (2015). Potential role of the intestinal microbiota in programming health and disease. Nutr. Rev. 73, 32–40. doi: 10.1093/nutrit/nuv039
Gregory, K. E., LaPlante, R. D., Shan, G., Kumar, D. V., and Gregas, M. (2015). Mode of birth influences preterm infant intestinal colonization with Bacteroides over the early neonatal period. Adv. Neonatal Care 15, 386–393. doi: 10.1097/ANC.0000000000000237
Hammer,Ø., Harper, D. A. T., and Ryan, P. D. (2001). PAST: paleontological statistics software package for education and data analysis. Palaeontol. Electron 4, 1–9. doi: 10.1016/j.bcp.2008.05.025
Hassiotou, F., Hepworth, A. R., Metzger, P., Tat Lai, C., Trengove, N., Hartmann, P. E., et al. (2013). Maternal and infant infections stimulate a rapid leukocyte response in breastmilk. Clin. Transl. Immunol. 2:e3. doi: 10.1038/cti.2013.1
Heikkilä, M. P., and Saris, P. E. J. (2003). Inhibition of Staphylococcus aureus by the commensal bacteria of human milk. J. Appl. Microbiol. 95, 471–478. doi: 10.1046/j.1365-2672.2003.02002.x
Hunt, K. M., Foster, J. A., Forney, L. J., Schütte, U. M. E., Beck, D. L., Abdo, Z., et al. (2011). Characterization of the diversity and temporal stability of bacterial communities in human milk. PLoS ONE 6:e21313. doi: 10.1371/journal.pone.0021313
Jeurink, P. V., van Bergenhenegouwen, J., Jiménez, E., Knippels, L. M. J., Fernández, L., Garssen, J., et al. (2013). Human milk: a source of more life than we imagine. Benef. Microbes 4, 17–30. doi: 10.3920/BM2012.0040
Jost, T., Lacroix, C., Braegger, C., and Chassard, C. (2013). Assessment of bacterial diversity in breast milk using culture-dependent and culture-independent approaches. Br. J. Nutr. 110, 1253–1262. doi: 10.1017/S0007114513000597
Jost, T., Lacroix, C., Braegger, C., and Chassard, C. (2015). Impact of human milk bacteria and oligosaccharides on neonatal gut microbiota establishment and gut health. Nutr. Rev. 73, 426–437. doi: 10.1093/nutrit/nuu016
Keim, S. A., Hogan, J. S., McNamara, K. A., Gudimetla, V., Dillon, C. E., Kwiek, J. J., et al. (2013). Microbial contamination of human milk purchased via the internet. Pediatrics 132, e1227–e1235. doi: 10.1542/peds.2013-1687
Kent, J. C. (2006). Volume and frequency of breastfeedings and fat content of breast milk throughout the day. Pediatrics 117, e387–e395. doi: 10.1542/peds.2005-1417
Khodayar-Pardo, P., Mira-Pascual, L., Collado, M. C., and Martínez-Costa, C. (2014). Impact of lactation stage, gestational age and mode of delivery on breast milk microbiota. J. Perinatol. 34, 599–605. doi: 10.1038/jp.2014.47
Kumar, H., du Toit, E., Kulkarni, A., Aakko, J., Linderborg, K. M., Zhang, Y., et al. (2016). Distinct patterns in human milk microbiota and fatty acid profiles across specific geographic locations. Front. Microbiol. 7:1619. doi: 10.3389/fmicb.2016.01619
Landers, S., and Updegrove, K. (2010). Bacteriological screening of donor human milk before and after holder pasteurization. Breastfeed. Med. 5, 117–121. doi: 10.1089/bfm.2009.0032
Lee, E., Kim, B.-J., Kang, M.-J., Choi, K. Y., Cho, H.-J., Kim, Y., et al. (2016). Dynamics of gut microbiota according to the delivery mode in healthy korean infants. Allergy Asthma Immunol. Res. 8, 471–477. doi: 10.4168/aair.2016.8.5.471
Lee, H. C., and Gould, J. B. (2009). Factors influencing breast milk versus formula feeding at discharge for very low birth weight infants in California. J. Pediatr. 155, 657.e–662.e. doi: 10.1016/j.jpeds.2009.04.064
Lemas, D. J., Yee, S., Cacho, N., Miller, D., Cardel, M., Gurka, M., et al. (2016). Exploring the contribution of maternal antibiotics and breastfeeding to development of the infant microbiome and pediatric obesity. Semin. Fetal Neonatal Med. 21, 406–409. doi: 10.1016/j.siny.2016.04.013
Liu, D., Yu, J., Li, L., Ai, Q., Feng, J., Song, C., et al. (2015). Bacterial community structure associated with elective cesarean section versus vaginal delivery in Chinese newborns. J. Pediatr. Gastroenterol. Nutr. 60, 240–246. doi: 10.1097/MPG.0000000000000606
Madara, J. (2004). Building an intestine — architectural contributions of commensal bacteria. N. Engl. J. Med. 351, 1685–1686. doi: 10.1056/NEJMcibr042621
Marcial, G. E., Ford, A. L., Haller, M. J., Gezan, S. A., Harrison, N. A., Cai, D., et al. (2017). Lactobacillus johnsonii N6.2 modulates the host immune responses: a double-blind, randomized trial in healthy adults. Front. Immunol. 8:655. doi: 10.3389/fimmu.2017.00655
Martín, R., Heilig, H. G. H. J., Zoetendal, E. G., Jiménez, E., Fernández, L., Smidt, H., et al. (2007). Cultivation-independent assessment of the bacterial diversity of breast milk among healthy women. Res. Microbiol. 158, 31–37. doi: 10.1016/j.resmic.2006.11.004
Martín, R., Langa, S., Reviriego, C., Jimínez, E., Marín, M. L., Xaus, J., et al. (2003). Human milk is a source of lactic acid bacteria for the infant gut. J. Pediatr. 143, 754–758. doi: 10.1016/j.jpeds.2003.09.028
McMurdie, P. J., and Holmes, S. (2013). Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8:e61217. doi: 10.1371/journal.pone.0061217
Meinzen-Derr, J., Poindexter, B., Wrage, L., Morrow, A. L., Stoll, B., and Donovan, E. F. (2009). Role of human milk in extremely low birth weight infants’ risk of necrotizing enterocolitis or death. J. Perinatol. 29, 57–62. doi: 10.1038/jp.2008.117
Menon, G., and Williams, T. C. (2013). Human milk for preterm infants: why, what, when and how? Arch. Dis. Child Fetal Neonatal Ed. 98, F559–F562. doi: 10.1136/archdischild-2012-303582
Moretti, M. (2012). Breastfeeding and the use of antidepressants. J. Popul. Ther. Clin. Pharmacol. 19, e387–e390. doi: 10.1542/peds.2011-3552
Nagpal, R., Tsuji, H., Takahashi, T., Kawashima, K., Nagata, S., Nomoto, K., et al. (2016). Sensitive quantitative analysis of the meconium bacterial microbiota in healthy term infants born vaginally or by cesarean section. Front. Microbiol. 7:1997. doi: 10.3389/fmicb.2016.01997
Newburg, D. S., Ruiz-Palacios, G. M., and Morrow, A. L. (2005). Human milk glycans protect infants against enteric pathogens. Annu. Rev. Nutr. 25, 37–58. doi: 10.1146/annurev.nutr.25.050304.092553
Parks, D. H., Tyson, G. W., Hugenholtz, P., and Beiko, R. G. (2014). STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30, 3123–3124. doi: 10.1093/bioinformatics/btu494
Reviriego, C., Eaton, T., Martín, R., Jiménez, E., Fernández, L., Gasson, M. J., et al. (2005). Screening of virulence determinants in Enterococcus faecium strains isolated from breast milk. J. Hum. Lact. 21, 131–137. doi: 10.1177/0890334405275394
Rideout, J. R., He, Y., Navas-Molina, J. A., Walters, W. A., Ursell, L. K., Gibbons, S. M., et al. (2014). Subsampled open-reference clustering creates consistent, comprehensive OTU definitions and scales to billions of sequences. PeerJ 2:e545. doi: 10.7717/peerj.545
Rutayisire, E., Huang, K., Liu, Y., and Tao, F. (2016). The mode of delivery affects the diversity and colonization pattern of the gut microbiota during the first year of infants’ life: a systematic review. BMC Gastroenterol. 16:86. doi: 10.1186/s12876-016-0498-0
Schanler, R. J. (2005). Randomized trial of donor human milk versus preterm formula as substitutes for mothers’ own milk in the feeding of extremely premature infants. Pediatrics 116, 400–406. doi: 10.1542/peds.2004-1974
Sinkiewicz, G., and Ljunggren, L. (2008). Occurrence of Lactobacillus reuteri in human breast milk. Microb. Ecol. Health Dis. 20, 122–126. doi: 10.1080/08910600802341007
Smith, M. M., Durkin, M., Hinton, V. J., Bellinger, D., and Kuhn, L. (2003). Initiation of breastfeeding among mothers of very low birth weight infants. Pediatrics 111, 1337–1342. doi: 10.1542/peds.111.6.1337
Urbaniak, C., Angelini, M., Gloor, G. B., and Reid, G. (2016). Human milk microbiota profiles in relation to birthing method, gestation and infant gender. Microbiome 4:1. doi: 10.1186/s40168-015-0145-y
Urbaniak, C., Burton, J. P., and Reid, G. (2012). Breast, milk and microbes: a complex relationship that does not end with lactation. Womens Heal. 8, 385–398. doi: 10.2217/whe.12.23
Wallace, J. G., Gohir, W., and Sloboda, D. M. (2016). The impact of early life gut colonization on metabolic and obesogenic outcomes: what have animal models shown us? J. Dev. Orig. Health Dis. 7, 15–24. doi: 10.1017/S2040174415001518
Wickham, H. (2009). ggplot2: Elegant Graphics for Data Analysis. New York, NY: Springer. doi: 10.1007/978-0-387-98141-3
Wight, N. E. (2001). Donor human milk for preterm infants. J. Perinatol. 21, 249–254. doi: 10.1038/sj.jp.7200533
Wold, A. E., and Adlerberth, I. (2002). “Breast feeding and the intestinal microflora of the infant — implications for protection against infectious diseases,” in Short and Long Term Effects of Breast Feeding on Child Health, eds B. Koletzko, K. F. Michaelsen, and O. Hernell (Boston, MA: Kluwer Academic Publishers), 77–93. doi: 10.1007/0-306-46830-1_7
Keywords: mother’s own milk, restoration, personalization, donor breast milk, bacterial load, human milk microbiome
Citation: Cacho NT, Harrison NA, Parker LA, Padgett KA, Lemas DJ, Marcial GE, Li N, Carr LE, Neu J and Lorca GL (2017) Personalization of the Microbiota of Donor Human Milk with Mother’s Own Milk. Front. Microbiol. 8:1470. doi: 10.3389/fmicb.2017.01470
Received: 21 March 2017; Accepted: 20 July 2017;
Published: 03 August 2017.
Edited by:
Christophe Lacroix, ETH Zurich, SwitzerlandReviewed by:
M. Luisa De Garnica, Universidad de León, SpainJuan Miguel Rodriguez, Complutense University of Madrid, Spain
Copyright © 2017 Cacho, Harrison, Parker, Padgett, Lemas, Marcial, Li, Carr, Neu and Lorca. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Josef Neu, bmV1akBwZWRzLnVmbC5lZHU= Graciela L. Lorca, Z2xvcmNhQHVmbC5lZHU=
†These authors have contributed equally to this work.
 Nicole T. Cacho
Nicole T. Cacho Natalie A. Harrison
Natalie A. Harrison Leslie A. Parker
Leslie A. Parker Kaylie A. Padgett
Kaylie A. Padgett Dominick J. Lemas
Dominick J. Lemas Guillermo E. Marcial
Guillermo E. Marcial Nan Li
Nan Li Laura E. Carr
Laura E. Carr Josef Neu
Josef Neu Graciela L. Lorca
Graciela L. Lorca