- 1The United Graduate School of Veterinary Science, Yamaguchi University, Yamaguchi, Japan
- 2Department of Sciences, Graduate School of Sciences and Technology for Innovation, Yamaguchi University, Yamaguchi, Japan
- 3National BioResource Project, Japan Agency for Medical Research and Development, Tokyo, Japan
Legionella pneumophila is a facultative intracellular Gram-negative bacterium, which is a major causative agent of Legionnaires’ disease. In the environment, this bacterium survives in free-living protists such as amoebae and Tetrahymena. The association of L. pneumophila and protists leads to the replication and spread of this bacterium. Thus, from a public health perspective, their association can enhance the risk of L. pneumophila infection for humans. Paramecium spp. are candidates of natural hosts of L. pneumophila, but their detailed relationships remain unclear. In the present study, we used an environmental strain, L. pneumophila Ofk308 (Ofk308) and Paramecium tetraurelia st110-1a to reveal the relationship between L. pneumophila and Paramecium spp. Ofk308 was cytotoxic to P. tetraurelia in an infection-dependent manner. We focused on TolC, a component of the type I secretion system, which is a virulence factor of L. pneumophila toward protists and found that cytotoxicity was dependent on TolC but not on other T1SS components. Further, the number of bacteria in P. tetraurelia was not associated with cytotoxicity and TolC was not involved in the mechanism of resistance against the digestion of P. tetraurelia in Ofk308. We used a LysoTracker to evaluate the maturation process of P. tetraurelia phagosomes containing Ofk308. We found that there was no difference between Ofk308 and the tolC-deletion mutant. To assess the phagocytic activity of P. tetraurelia, Texas Red-conjugated dextran-uptake assays were performed. Ofk308 inhibited phagosome formation by P. tetraurelia through a TolC-dependent mechanism. Further, we evaluated the excretion of Legionella-containing vacuoles from P. tetraurelia. We found that P. tetraurelia failed to excrete undigested Ofk308 and that Ofk308 remained within cells through a TolC-dependent mechanism. Our results suggest that TolC is essential for L. pneumophila to remain within Paramecium cells and to show cytotoxicity. Because of the high mobility and high cell division rate of Paramecium spp., living with Paramecium spp. would be beneficial for L. pneumophila to expand its habitat. To control Legionaries’ disease, understanding the ecology of L. pneumophila in the environment is essential.
Introduction
Legionella pneumophila is a facultative intracellular Gram-negative bacterium that is the major causative agent of Legionnaires’ disease (Cunha et al., 2016), which is a severe pneumonia and a mild flu-like illness called Pontiac fever. The infection of humans occurs by inhalation of a Legionella-containing aerosol. Once L. pneumophila enters the lung, it invades alveolar macrophages and replicates within Legionella-containing vacuoles (LCVs) derived from host phagosomes (Isberg et al., 2009) that serve as a suitable niche for bacterial replication.
Legionella pneumophila normally inhabits natural or artificial aquatic environments where it can survive for long periods as free-living forms as well as in biofilms (Fliermans et al., 1981; Paszko-Kolva et al., 1992; Declerck et al., 2007). Further, L. pneumophila can survive in free-living protists. Amoebae are protistan hosts of L. pneumophila in bodies of fresh water (Rowbotham, 1980), and evidence indicates that Tetrahymena serves as its host (Fields et al., 1984). In the environment, the association of L. pneumophila and protists leads to the replication and spread of this bacterium as well as to the development of antibiotic resistance (Barker et al., 1995; Winiecka-Krusnell and Linder, 1999). Thus, from a public health perspective, their association can enhance the risk of L. pneumophila infection for humans.
Many bacteria other than L. pneumophila are resistant to protists (Greub and Raoult, 2004), including human pathogenic intracellular bacteria such as Coxiella burnetii and Listeria monocytogenes as well as nonpathogenic bacteria. Protistan hosts are thought to represent a primary evolutionary factor for the acquisition and maintenance of virulence toward humans (Al-Quadan et al., 2012). Because of the similarities of the digestive process between macrophages and protists, certain mechanisms to resist protist digestion help bacteria to survive in macrophages. Thus, association between bacteria and protists can cause the emergence of new pathogenic bacteria, because nonpathogenic bacteria may acquire a pathogenic phenotype within protistan hosts. To control current and future pathogenic bacteria as well as L. pneumophila, it is extremely important to understand and identify the association between environmental bacteria and protistan hosts.
Paramecium spp. are ciliates that exist widely in freshwater environments and are appreciated as model organisms for the analysis of cellular and molecular biology, including phagocytosis and exocytosis (Steinman et al., 1983; Plattner, 2010). Further, Paramecium spp. are used to study endosymbiosis, because they possess several endosymbionts (Görtz and Fokin, 2009), and symbionts of Paramecium spp. affect the host’s phenotype. For example, Paramecium caudatum can acquire resistance to salinity (Duncan et al., 2010) and heat-shock (Hori and Fujishima, 2003; Fujishima et al., 2005) if infected with Holospora spp. Previously, we found that Paramecium spp. are candidates of a natural host of L. pneumophila (Watanabe et al., 2016). L. pneumophila have resistance to Paramecium digestion and several L. pneumophila strains show cytotoxicity toward Paramecium spp. Paramecium spp. may increase the risk of L. pneumophila infection for humans as well as other protistan hosts such as amoebae and Tetrahymena. However, it is still unclear whether L. pneumophila establishes endosymbiosis with Paramecium spp. in the environment.
Type I secretion systems (T1SSs) are widespread in pathogenic Gram-negative bacteria such as Escherichia coli, Vibrio cholerae, and Bordetella pertussis (Thomas et al., 2014). This secretion machinery comprises three components that transport substrates to the exterior in one step across both lipid bilayers. The first reported protein secreted through T1SS is the hemolysin A (HlyA), which is produced by certain E. coli strains (Goebel and Hedgpeth, 1982). HlyA is a member of the repeats-in-toxin (RTX) family that is the most extensively studied. Further, L. pneumophila employs T1SS, and RtxA serves as its substrate in L. pneumophila. The association of T1SS and RtxA with virulence of L. pneumophila such as invasion, intracellular growth, and pore-forming activities toward amoebae has been revealed (Fuche et al., 2015). In addition, TolC, the outer membrane protein of T1SS, plays roles in virulence and multidrug resistance (Ferhat et al., 2009). TolC is a trimeric membrane protein and forms a long channel that protrudes deeply into the periplasm. TolC is composed of a short β-barrel (outer membrane) and a long α-helical (periplasmic) structure (Koronakis et al., 2000). TolC forms the T1SS by interacting with an ATP-binding cassette transporter and a membrane fusion protein that resides in the inner membrane. TolC couples with numerous inner membrane or periplasmic proteins and forms efflux systems such as the AcrAB-TolC efflux pump (Symmons et al., 2015). These efflux systems transport diverse molecules such as virulence-associated proteins, antibiotics, and detergents (Zgurskaya et al., 2011). However, we are unaware of published studies on the role of TolC in the relationship between L. pneumophila and Paramecium spp.
In the present study, we focused on the role of TolC in the association between L. pneumophila and Paramecium spp. We found that TolC was essential for L. pneumophila to remain within Paramecium cells. Our results suggest that L. pneumophila employs a TolC-dependent mechanism to survive within Paramecium spp. in the environment.
Materials and Methods
Bacterial Strains and Culture Conditions
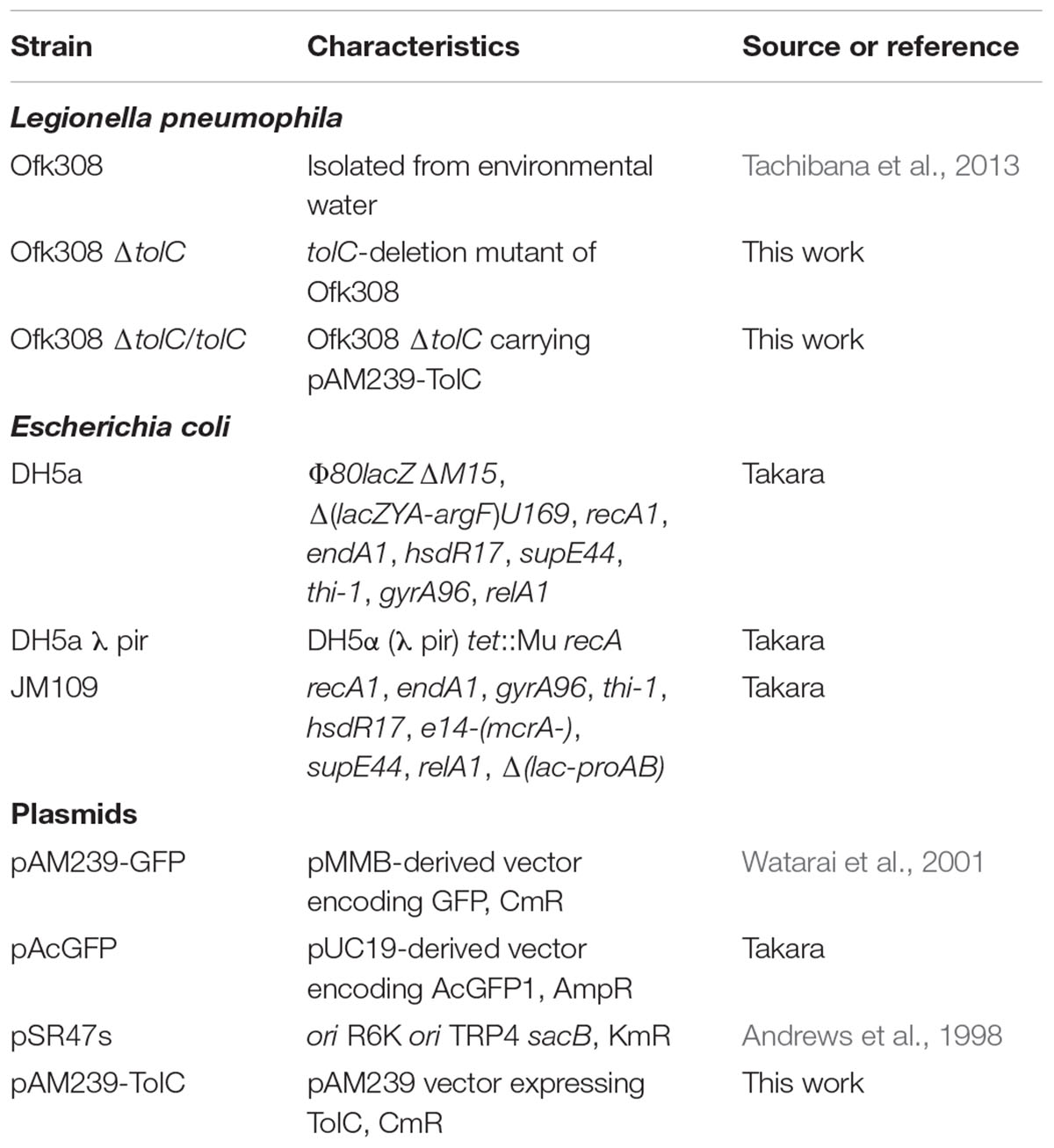
All bacterial strains used in this study are listed in Table 1. L. pneumophila strains were cultured at 37°C on N-(2-acetamido)-2-aminoethanesulfonic acid-buffered charcoal yeast extract agar (BCYE) or in the same medium without agar and charcoal (AYE; Nishida et al., 2017). E. coli strains were cultured in Lysogeny Broth (LB; Nacalai Tesuque, Kyoto, Japan) or on LB containing 1.5% agar (Wako, Tokyo, Japan). Media were supplemented with chloramphenicol (10 μg/mL, Wako), kanamycin (30 μg/mL, Wako), and ampicillin (250 μg/mL, Wako) as required.
P. tetraurelia and Culture Conditions
Paramecium tetraurelia st110-1a (ID: PT041001A) was provided by the Symbiosis Laboratory, Yamaguchi University, with support, in part, from the NBRP. Culture and maintenance were previously described (Fujishima et al., 1990). Briefly, the culture medium used was 2.5% (w/v) fresh lettuce juice in Dryl’s solution (Dryl, 1959) inoculated with a nonpathogenic strain of Klebsiella pneumoniae the day before use. The cultivation was performed at 25°C. Cells at the stationary phase of growth (20–24 h after the last feeding) were used for the experiments.
Construction of Deletion Mutants and Complementary Strains
Each deletion mutant was constructed using the homologous-recombination method. Briefly, two PCR fragments were cloned into SalI/NotI or BamHI-cleaved pSR47s (Andrews et al., 1998) using an In-Fusion HD Cloning Kit (Takara, Tokyo, Japan). Fragment 1 was a 1,500 or 2,000 bp fragment spanning a site located upstream of the 5’ end of each target gene. Fragment 2 was a 1,500 or 2,000 bp fragment spanning a site located downstream of the 3’ end of each target gene. These fragments were amplified using PCR. Each plasmid was introduced into E. coli DH5a λ pir and subsequently transferred into Ofk308 using electroporation with a Gene Pulser electroporator (Bio-Rad Laboratories, Hercules, CA, United States) in 10% glycerol at 2.5 kV/25 μF. Isolation of in-frame deletion mutants by positive selection for sucrose resistance has been described (Andrews et al., 1998). The tolC complementary strain was constructed by cloning a PCR fragment of tolC into PstI/EcoRI-cleaved pAM239-GFP (green fluorescence protein) using a DNA Ligation Kit (Takara). This tolC-inserted plasmid, pAM239-TolC, was introduced into E. coli DH5a and subsequently transferred into L. pneumophila using electroporation. GFP or TolC expression in L. pneumophila was induced by adding isopropyl-β-D-thiogalactopyranoside (1 mM, Wako) to AYE.
All primers and plasmids used in this work are listed in Tables 1, 2. Plasmid DNA from E. coli DH5a was prepared using a QIAGEN Plasmid Mini Kit (QIAGEN, Hilden, Germany). Restriction enzymes (Takara) were used according to the manufacturers’ protocols.
Cytotoxicity Assay and Determination of the Bacterial Loads in P. tetraurelia
Cytotoxicity assays and determination of bacterial loads were performed as previously described (Watanabe et al., 2016). Briefly, Ofk308 or each mutant strain was added to P. tetraurelia in 1.5-mL tubes and then incubated at 25°C. After incubation, viable P. tetraurelia was counted using microscopy. To determine the bacterial load, P. tetraurelia was infected with each strain of L. pneumophila at multiplicity of infection [MOI] = 20,000. After incubation at 25°C, P. tetraurelia was washed five times with 5 mL of fresh lettuce juice in Dryl’s solution through a 15-μm pore nylon mesh to remove extracellular bacteria. Samples were further treated at 50°C for 30 min to purge the K. pneumoniae fed to P. tetraurelia. Colony-forming units were determined using serial dilution on BCYE. In infections with E. coli JM109, incubation at 50°C was omitted, and LB-containing ampicillin (250 μg/mL) was used.
Fluorescence Microscopy
GFP-expressing bacteria were added to P. tetraurelia at MOI = 20,000, which were then incubated at 25°C for 30 min to 48 h. P. tetraurelia was fixed with 4% paraformaldehyde in PBS for 10 min at room temperature. Images of fluorescence were obtained using a FluoView FV100 confocal laser scanning microscope (Olympus, Tokyo, Japan).
When LysoTracker (Life Technologies, Carlsbad, CA, United States) was used, P. tetraurelia was fixed 30 min after infection. After fixation, samples were washed twice with PBS and then incubated with LysoTracker (50 nM) for 30 min. LysoTracker-positive LCVs were counted using microscopy, and the data are shown as an average of three fields.
When a Texas Red-conjugated dextran (TRDx, Thermo Fisher Scientific, MA, United States) was used, samples were washed 1 h after infection to remove extracellular bacteria using a nylon mesh as described above. TRDx (50 μg/mL) was added to P. tetraurelia. At each sampling time, P. tetraurelia was fixed and washed twice with PBS. The number of TRDx-containing vacuoles in individual P. tetraurelia was counted and expressed as the average of 30 P. tetraurelia cells.
Observations of Individual P. tetraurelia Under NiCl2-Induced Paralysis
Paramecium tetraurelia was infected with L. pneumophila at MOI = 20,000 and then incubated at 25°C for 1 h. After washing as described above, NiCl2 (2 mM) was added and the cells were incubated for 10 min at room temperature. After incubation, the P. tetraurelia cells were collected and transferred into new media supplemented with 0.2 mM NiCl2. Samples were immediately transferred to a 48-well plate. An IX75 inverted fluorescence microscope was used to observe and photograph the cells at 5 min intervals. The number of LCV-containing P. tetraurelia in 50 cells was counted at 0 and 40 min.
Statistical Analyses
Statistical analyses were performed using the Tukey–Kramer test or the Student’s t-test. Statistically significant differences between groups were accepted at P < 0.05 or P < 0.01. Data are presented as the average of three identical experiments, and the error bars shown in the figures represent SDs.
Results
L. pneumophila Ofk308 Exhibits Cytotoxicity Toward P. tetraurelia
Paramecium spp. feed on bacteria and bacteria uptaken by Paramecium are normally digested. On the other hand, L. pneumophila Ofk308 (Ofk308) showed cytotoxicity toward several Paramecium strains (Watanabe et al., 2016). However, the bacterial properties in Paramecium were not evaluated. Then, we evaluated intracellular localization of Ofk308. As a result of infection assays in which GFP-expressing Ofk308 was added to 93 strains of Paramecium spp. (MOI = 10,000), the intracellular localization of Ofk308 was clearly observed in P. tetraurelia strains (data not shown). Therefore, we decided to use P. tetraurelia st110-1a as a model to analyze the relationship between Ofk308 and Paramecium spp. in the present study.
Ofk308 exhibits cytotoxicity toward several Paramecium strains in an MOI-dependent manner (Watanabe et al., 2016). Therefore, the cytotoxicity of Ofk308 toward P. tetraurelia was assessed. P. tetraurelia was infected with Ofk308 at different MOIs, and viable P. tetraurelia was counted 48 h after infection. At MOI ≤ 5,000, the number of viable P. tetraurelia was the same as that of uninfected P. tetraurelia. However, at MOI = 10,000, the number of viable P. tetraurelia tended to decrease, and at MOI ≥ 20,000, the number of viable P. tetraurelia decreased significantly compared with uninfected P. tetraurelia (Figure 1). These results suggest that Ofk308 is cytotoxic to P. tetraurelia in an MOI-dependent manner.
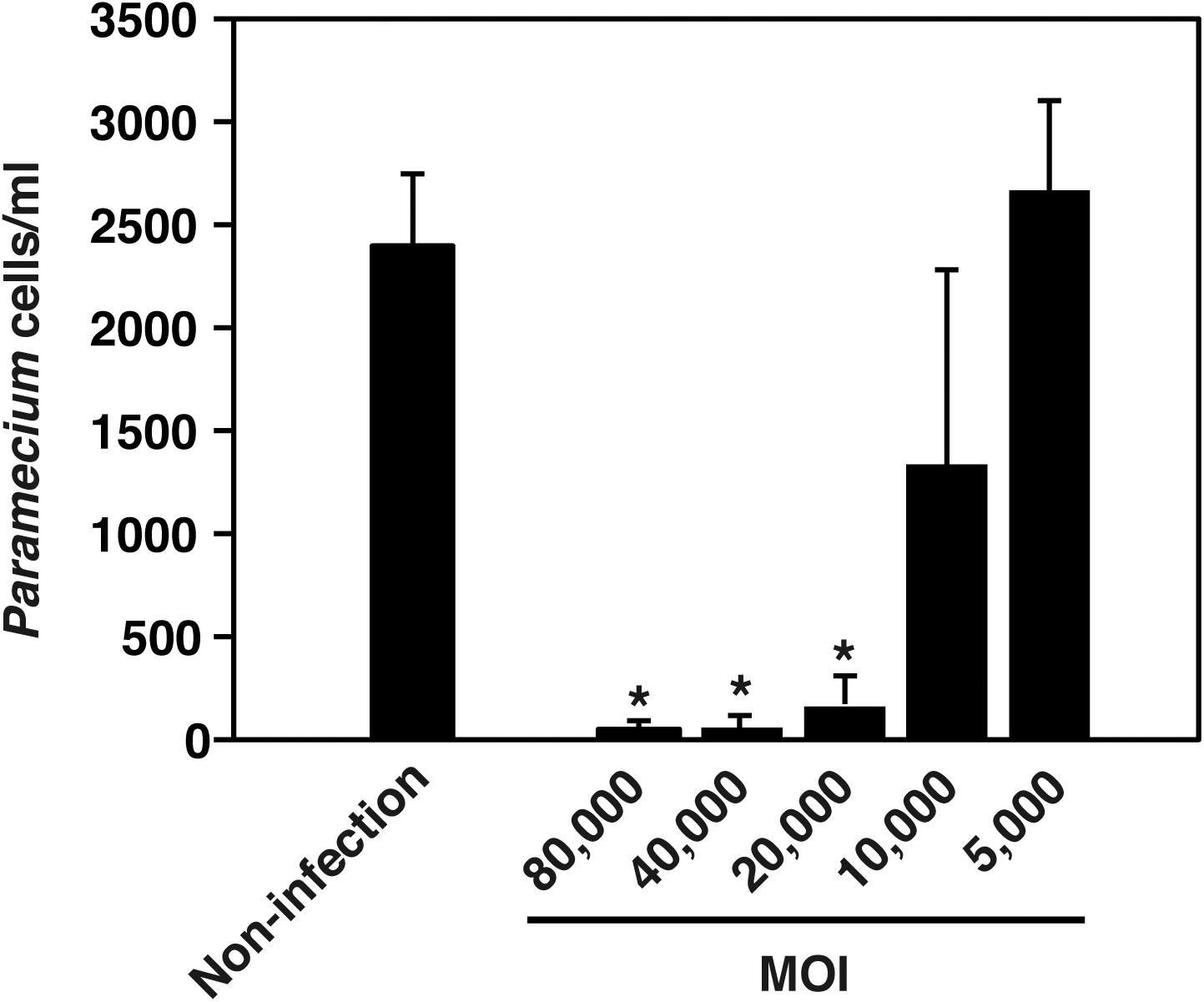
FIGURE 1. L. pneumophila Ofk308 exhibits cytotoxicity toward P. tetraurelia in an MOI-dependent manner. The number of P. tetraurelia 48 h after infection. P. tetraurelia was infected with Ofk308 at MOIs = 5,000, 10,000, 20,000 40,000, and 80,000. Data are expressed as the average of three identical experiments, and the error bars represent SDs. Asterisks indicate statistically significant differences compared with non-infection (∗P < 0.05).
TolC Plays a Crucial Role in the Cytotoxicity of L. pneumophila Ofk308
Type-IV secretion systems of L. pneumophila play a major role in its replication within mammalian macrophages and amoebae (Al-Quadan et al., 2012). However, the lack of Type-IV secretion systems does not affect the cytotoxicity of Ofk308 toward P. caudatum (Watanabe et al., 2016). Therefore, we focused on T1SS, which is a virulence factor of L. pneumophila to amoebae (Fuche et al., 2015). For this purpose, we constructed deletion mutants of tolC, lssBD, and rtxA. TolC is an outer membrane component of T1SS. LssBD serves as the inner membrane and periplasmic components of T1SS, and RtxA is a substrate of T1SS (Fuche et al., 2015). Using these strains, we examined the cytotoxicity toward P. tetraurelia at MOI = 20,000. The tolC-deletion mutant lost its cytotoxicity toward P. tetraurelia, and its complementary strain recovered cytotoxicity comparable to that of the parental strain Ofk308 (Figure 2). However, the number of viable P. tetraurelia decreased to the same degree as Ofk308 when the lssBD- and rtxA-deletion mutants were infected. These results indicate that the cytotoxicity of Ofk308 is dependent on TolC but not on T1SS.
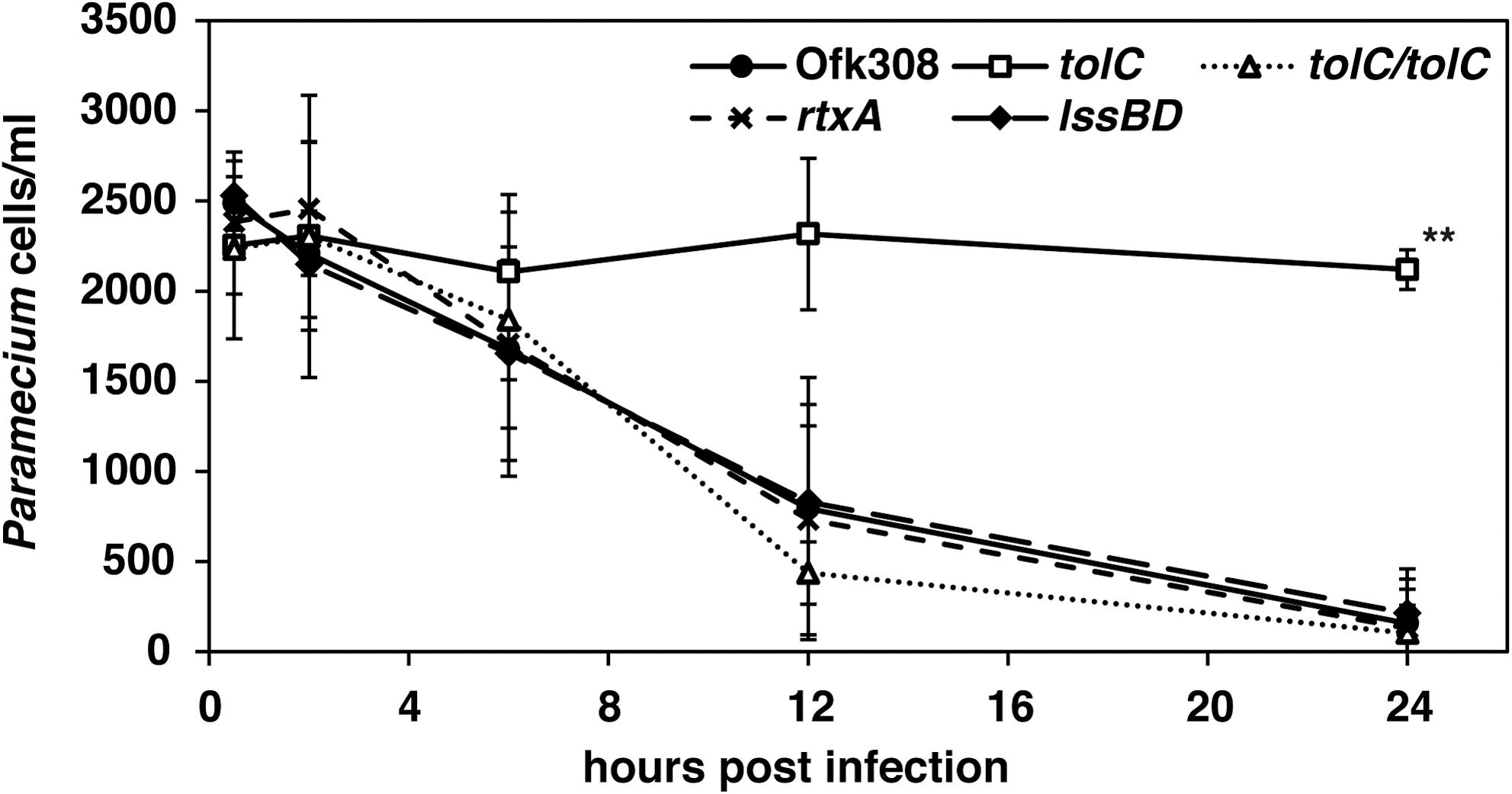
FIGURE 2. The tolC-deletion mutant of L. pneumophila Ofk308 is defective for cytotoxicity. P. tetraurelia was infected with Ofk308, the tolC-deletion mutant (tolC), the lssBD-deletion mutant (lssBD), the rtxA-deletion mutant (rtxA), or the tolC-complementary strain (tolC/tolC) at MOI = 20,000. Data are presented as the average of three identical experiments, and the error bars represent SDs. Asterisks indicate statistically significant differences compared with Ofk308 (∗∗P < 0.01).
The Cytotoxicity of L. pneumophila Ofk308 Is Independent on the Number of Bacteria Within P. tetraurelia
To investigate how Ofk308 exhibits cytotoxicity toward P. tetraurelia through a TolC dependent mechanism, we compared bacterial numbers in P. tetraurelia using Ofk308, the tolC-deletion mutant, and E. coli as a control. From 2 to 48 h after infection at MOI = 20,000, the numbers of E. coli decreased but the numbers of Ofk308 and those of the tolC-deletion mutant were unchanged (Figure 3). Comparable results were observed at lower MOIs (data not shown). In cells infected with Ofk308, the shape of P. tetraurelia changed unnaturally, although damage was not observed in cells infected with the tolC-deletion mutant. These results suggest that the number of bacteria in P. tetraurelia is not associated with cytotoxicity and that TolC is not involved in the mechanism of resistance against the digestion of P. tetraurelia in Ofk308.
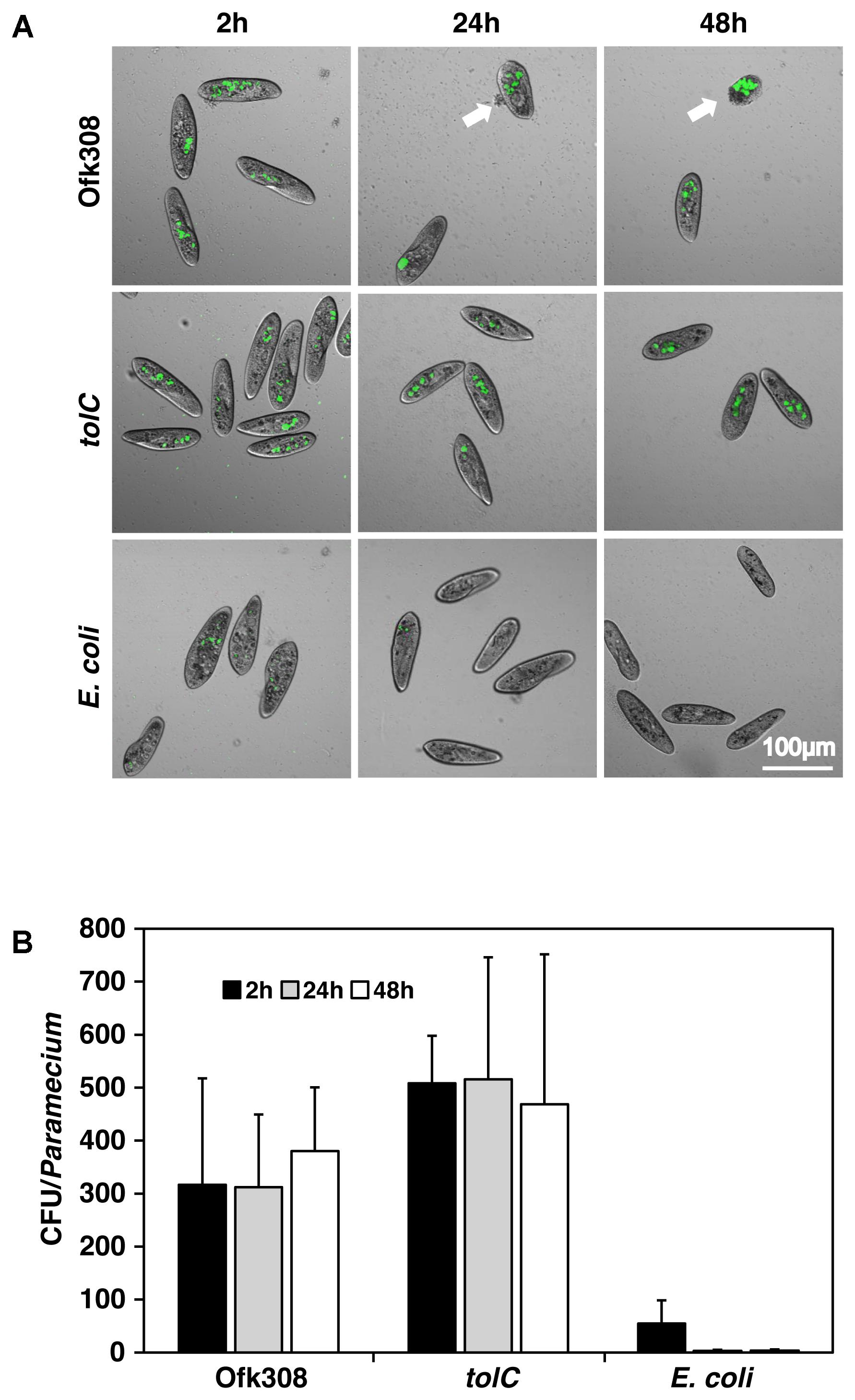
FIGURE 3. The number of L. pneumophila in P. tetraurelia is constant. (A) GFP-expressing bacteria, Ofk308, the tolC-deletion mutant (tolC), and E. coli JM109 in P. tetraurelia 2, 24, and 48 h after infection at MOI = 20,000. Arrows point to unnaturally shaped P. tetraurelia. (B) Number of bacteria per P. tetraurelia. Data are expressed as the average of three identical experiments, and the error bars represent SDs.
L. pneumophila Ofk308 Inhibits the Phagocytic Activity of P. tetraurelia Through a TolC-Dependent Mechanism
Next, we focused on phagocytic activity of P. tetraurelia after mixing with Ofk308 because we hypothesized that the inhibition of phagocytic activity could cause cytotoxicity. P. tetraurelia exhibits high phagocytic activity (Plattner and Kissmehl, 2003), and L. pneumophila modulates host phagosomes to survive within them (Isberg et al., 2009). In infection of P. caudatum, Ofk308 inhibits phagosome–lysosome fusion (PL-fusion; Watanabe et al., 2016). Therefore, we used a LysoTracker to evaluate the maturation of P. tetraurelia phagosomes containing Ofk308. We found that there was no difference between Ofk308 and the tolC-deletion mutant; 97.1% of LCVs were LysoTracker positive in cells infected with Ofk308 and 97.4% of LCVs were LysoTracker positive in cells infected with the tolC-deletion mutant (Figure 4).
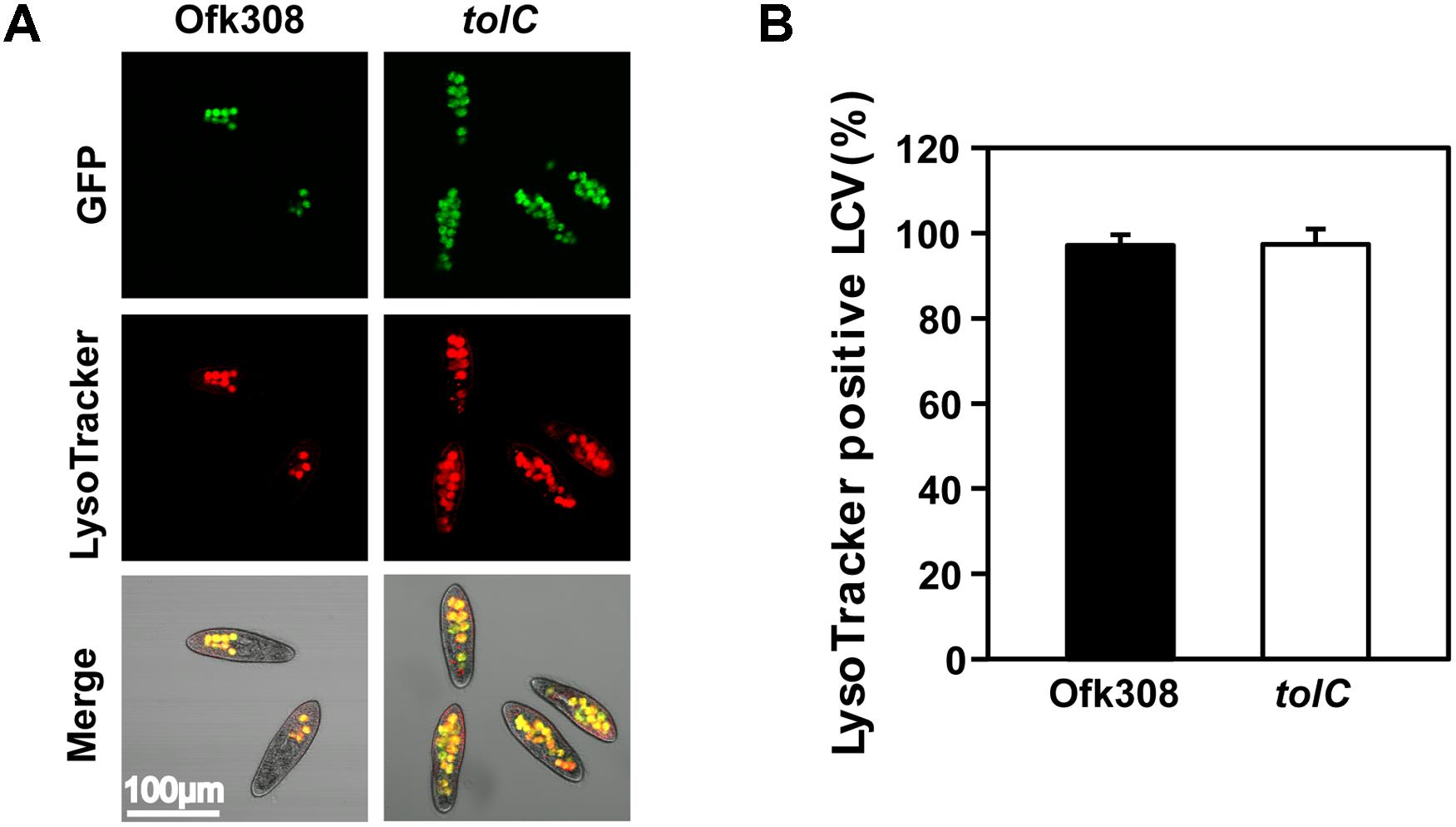
FIGURE 4. The maturation of host LCVs containing L. pneumophila Ofk308 is not inhibited. (A) LCV maturation 30 min after infection was evaluated using LysoTracker. Ofk308 or the tolC-deletion mutant (tolC) was added to P. tetraurelia at MOI = 20,000. (B) The percentage of LysoTracker-positive LCVs. Data are expressed as the average of three identical experiments, and the error bars represent SDs.
Next, to assess the formation of phagosomes, TRDx-uptake assay was performed (according to the schedule shown in Figure 5A). Dextran-containing vacuoles (DCVs) represent vacuoles that formed after adding TRDx. In cells infected with Ofk308, 5, 10, and 15 min after adding dextran, the average numbers of DCVs per cell were 2.5, 4.1, and 4.2, respectively. In contrast, in cells infected with the tolC-deletion mutant, the average numbers of DCVs were 5.8, 7.0, and 8.3, respectively (Figures 5B,C). There were significant differences between Ofk308 and the tolC-deletion mutant. These results suggest that Ofk308 inhibits phagosome formation by P. tetraurelia through a TolC-dependent mechanism.
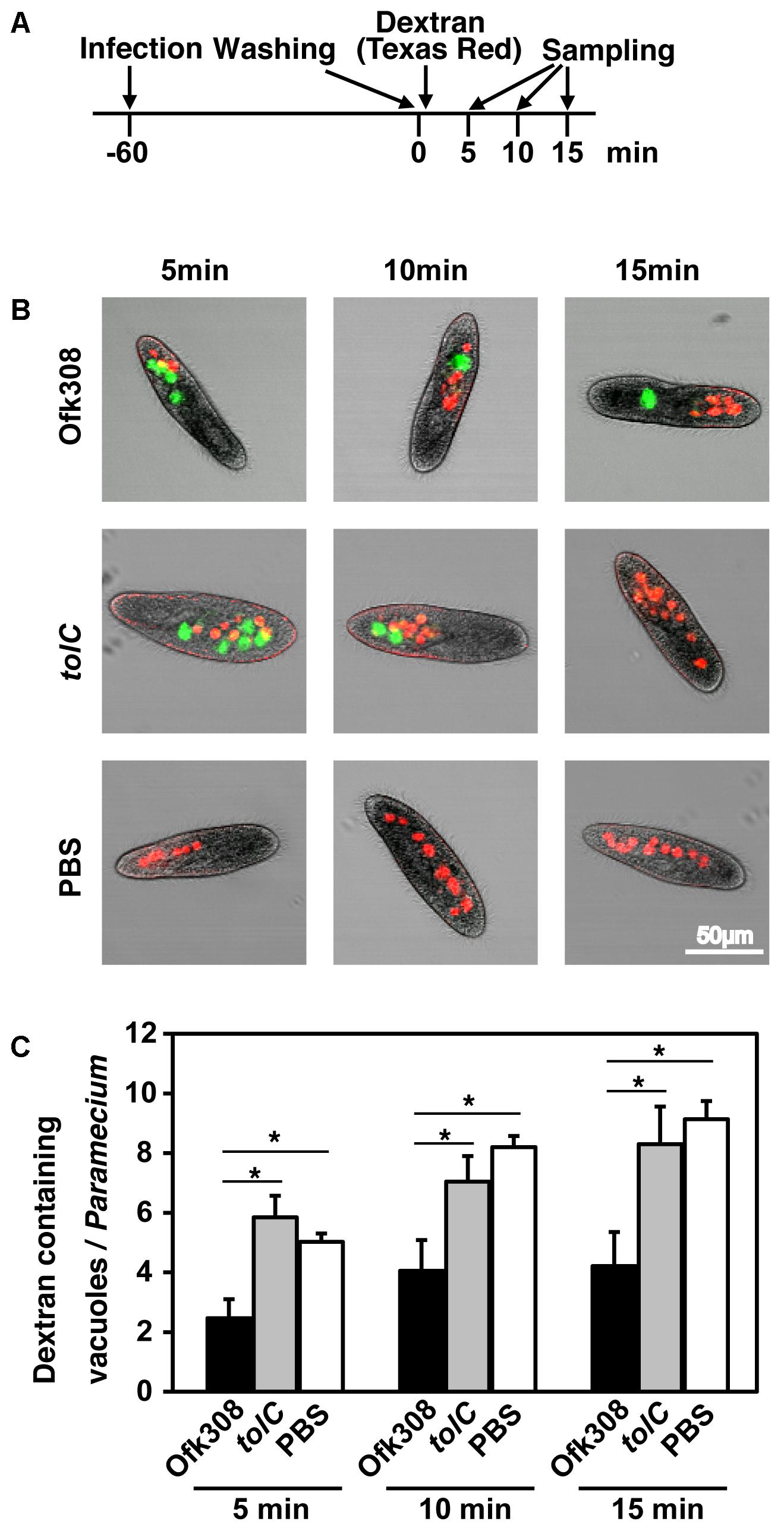
FIGURE 5. L. pneumophila Ofk308 inhibits phagosome formation by P. tetraurelia through a TolC-dependent mechanism. (A) Bacteria and TRDx were added to P. tetraurelia according to this schedule. The number of DCVs was counted at each indicated time point. (B) DCVs in P. tetraurelia infected with Ofk308 and tolC at each indicated time point. DCVs are red, and Legionella-containing vacuoles are green. (C) The number of DCVs in individual P. tetraurelia. Data are expressed as the average of three identical experiments, and error bars represent SDs. Asterisks indicate statistically significant differences (∗P < 0.05).
L. pneumophila Ofk308 Remains Within P. tetraurelia Through a TolC-Dependent Mechanism
Surviving in free-living protists is beneficial for L. pneumophila. L. pneumophila must inhibit the exocytosis of Paramecium to remain within them, because Paramecium exhibits high exocytic activity (Plattner and Kissmehl, 2003). We assumed that Ofk308 terminates the exocytic activities of P. tetraurelia. However, because of high mobility of P. tetraurelia, evaluation of the exocytic activities in individual P. tetraurelia is difficult. Therefore, the digestion vacuolar cycles of individual P. tetraurelia was observed under NiCl2-induced paralysis (according to the schedule shown in Figure 6A). By treated with NiCl2, P. tetraurelia stopped swimming but cytoplasmic streaming was observed; 93.6% of P. tetraurelia possessed Ofk308-containing vacuoles for at least 40 min (Figures 6B,C). In contrast, the tolC-deletion mutant-containing vacuoles were gradually excreted. Finally, most tolC-deletion mutants were excreted until 40 min and only 18.0% of P. tetraurelia possessed LCVs. These results suggest that P. tetraurelia failed to excrete undigested Ofk308 and that Ofk308 remained within cells through a TolC-dependent mechanism.
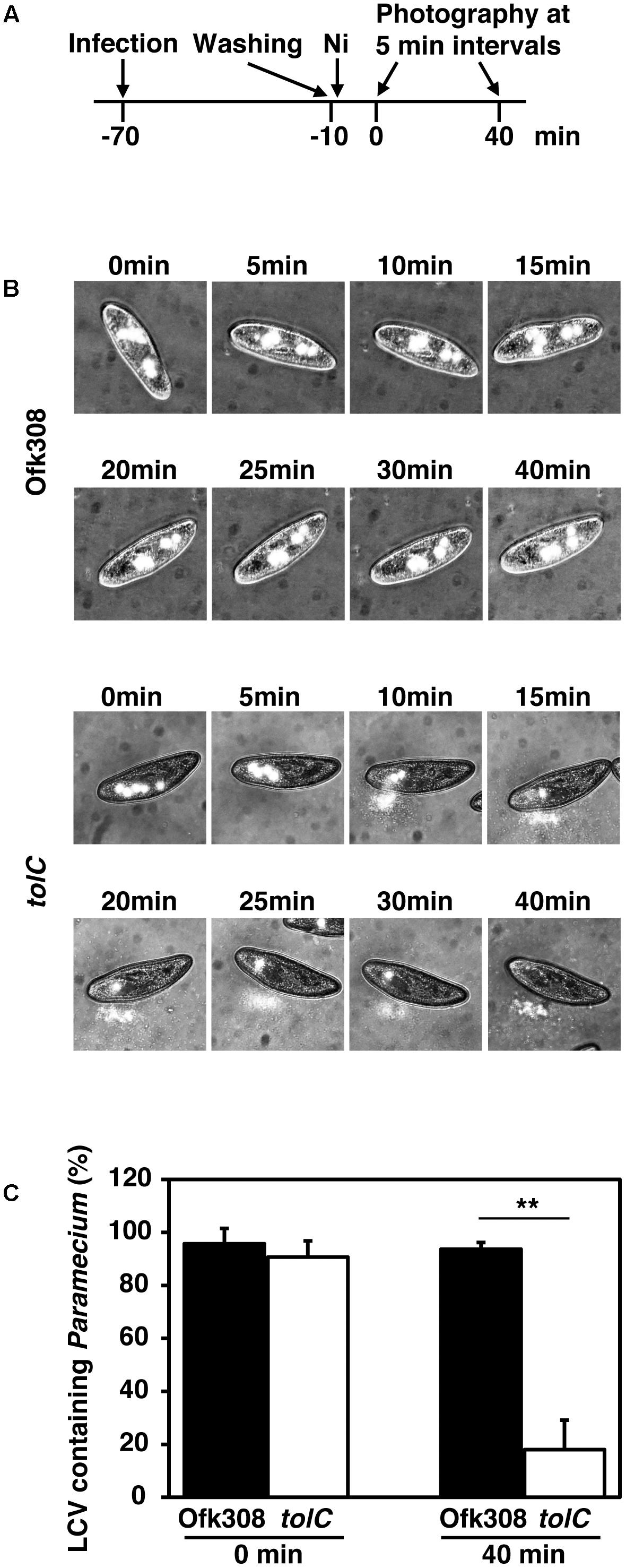
FIGURE 6. L. pneumophila Ofk308 inhibits excretion of LCVs from P. tetraurelia through a TolC-dependent mechanism. (A) Images of individual P. tetraurelia under NiCl2-induced paralysis were taken according to this schedule. (B) LCVs in individual P. tetraurelia at each indicated time point. LCVs are white. (C) The percentages of LCV-containing P. tetraurelia at 0 and 40 min. Data are expressed as the average of three identical experiments, and the error bars represent SDs. Asterisks indicate statistically significant differences (∗∗P < 0.01).
Discussion
In the environment, L. pneumophila replicates within a protistan host. Previous reports reveal the mechanisms of infection and replication in protists such as amoeba and Tetrahymena (Al-Khodor et al., 2009; Richards et al., 2013). However, in these reports, the temperatures used for most infection procedures ranged from 30 to 37°C, because these temperatures are appropriate for the culture of L. pneumophila and amoebae. Thus, it is possible that there are other mechanisms, which function at lower temperatures. In the present study, we used P. tetraurelia as a model protistan host of L. pneumophila. Paramecium spp. are candidates of natural host of L. pneumophila (Watanabe et al., 2016). Paramecium spp. are widely present in fresh water environments but their resistance to high temperature is lower compared with amoeba or Tetrahymena (Thatcher and Gorovsky, 1993; Hori and Fujishima, 2003). Thus, all infection assays in our present work were performed at 25°C, which is typical for Paramecium spp. culture conditions. Although the cytotoxicity of L. pneumophila toward amoeba decreases at lower temperature (Ohno et al., 2008), cytotoxicity of Ofk308 toward P. tetraurelia was clearly observed at 25°C in an MOI-dependent manner (Figure 1). The association of L. pneumophila and Paramecium spp. can lead to the replication and spread of L. pneumophila in natural aquatic environments. For this reason, our assay likely reflects the natural environmental conditions that are conducive to the survival of L. pneumophila and can be helpful to reveal the ecology of L. pneumophila in the environment.
The Dot/Icm system of L. pneumophila contributes to intracellular survival and replication in amoeba (Richards et al., 2013). However, in our previous report, the Dot/Icm system had no effect on cytotoxicity toward P. caudatum (Watanabe et al., 2016). Therefore, we focused on another secretion system, T1SS. Many Gram-negative pathogenic bacteria such as E. coli and B. pertussis employ T1SS, whose association with bacterial virulence is established (Goebel and Hedgpeth, 1982; Glaser et al., 1988). The role of T1SS in intracellular bacteria has been investigated as well. Several T1SS substrates are present in Orientia tsutsugamushi and Ehrlichia chaffeensis (Wakeel et al., 2011; VieBrock et al., 2015). T1SS in L. pneumophila was investigated, and the roles of T1SS in pore-forming activity and intracellular replication are established (Fuche et al., 2015). In the present study, we evaluated the effects of T1SS on cytotoxicity toward P. tetraurelia using a deletion-mutant of T1SS. As a result, in infection with the tolC-deletion mutant, decreased cytotoxicity was observed (Figure 2). However, other components of T1SS, including the T1SS substrate RtxA, did not affect cytotoxicity. These results suggest that TolC of L. pneumophila possesses another function separate from T1SS and that this function may be important in cytotoxicity of L. pneumophila toward P. tetraurelia. TolC forms several multidrug efflux pumps (Symmons et al., 2015), and TolC of L. pneumophila contributes to multidrug resistance (Ferhat et al., 2009). Multidrug efflux pumps are associated with the virulence of Salmonella enterica serovar Typhimurium (Buckley et al., 2006; Nishino et al., 2006). Therefore, an efflux pump composed of TolC may provide an important function in cytotoxicity of L. pneumophila toward P. tetraurelia. Further, Rickettsia typhi may employ another TolC-dependent secretion mechanism (Kaur et al., 2012). In this model, an ankyrin repeat-containing protein translocates to the periplasm via the Sec translocon. This protein is secreted via TolC. How TolC recognizes this protein in the periplasm is unknown, and it is unclear if L. pneumophila employs a similar secretion mechanism. Further study is therefore required to determine the function of TolC in L. pneumophila.
Legionella-endosymbiosis modulating factor A (LefA) regulates the relationship between L. pneumophila and Paramecium spp. (Watanabe et al., 2016). LefA is associated with intracellular replication and inhibition of PL-fusions in P. caudatum. In mammalian macrophages or amoebae, L. pneumophila modifies host phagosomes and avoids PL-fusions (Richards et al., 2013; Xu and Luo, 2013). Therefore, we examined the effects of TolC on intracellular replication and PL-fusions of P. tetraurelia with Ofk308. As a result, the same levels of intracellular localization and PL-fusions were observed in infections with Ofk308 and the tolC-deletion mutant (Figures 3, 4). These results indicate that Ofk308 avoids P. tetraurelia digestion independently of TolC. The mechanism of the resistance to digestion is unclear, but temperature may serve as a key factor, because all experiments were performed at 25°C.
Paramecium spp. possess high phagocytic and exocytic activities (Plattner and Kissmehl, 2003). In contrast, Chlorella, which is known as an endosymbiont of P. bursaria, remains for at least 72 h (Kodama and Fujishima, 2005). The symbiotic relationship between P. bursaria and Chlorella was well investigated. Then, endosymbionts of Paramecium may inhibit phagocytic and exocytic activities. We show here that TolC played a role in the inhibition of the excretion of LCVs (Figure 6). This inhibition follows that L. pneumophila remains within the host Paramecium. Therefore, TolC may represent a crucial factor required for Ofk308 to remain within P. tetraurelia and to establish symbiosis.
High concentrations of Ofk308 exhibited cytotoxicity toward P. tetraurelia through a TolC-dependent mechanism (Figures 1, 2). In infections with P. caudatum, significant intracellular replication of Ofk308 occurs and causes the death of the host P. caudatum (Watanabe et al., 2016). Thus, intracellular replication can represent a major cause of death. However, in the present study, such intracellular replication of Ofk308 was not observed in P. tetraurelia (Figure 2). Therefore, we reasoned that the inhibition of phagosome formation can cause the death of P. tetraurelia. After infection with P. tetraurelia, Ofk308 inhibited new phagosome formation through a TolC-dependent mechanism (Figure 5). In Paramecium spp., the membrane used to form new phagosomes is provided by recruitment of cytoplasmic discoidal vesicles that originate, in part, at the cytoproct (Steinman et al., 1983; Guerrier et al., 2017). Thus, excess Ofk308 remaining within P. tetraurelia cells can stop the recycling of the phagosome membrane to form new phagosomes. As a result, P. tetraurelia may starve and subsequently die because of poor nutrition. Further, the inhibition of phagosome formation can result in inhibition of invasion of other microbes to Paramecium. In natural condition, L. pneumophila may inhibit phagocytic activities by TolC-dependent mechanism to occupy host Paramecium.
Conclusion
We show here that TolC is essential for L. pneumophila to remain within Paramecium cells and to show cytotoxicity. The association between L. pneumophila and Paramecium spp. in the environment can enhance the risk of infection by L. pneumophila. Because of the high mobility and high cell division rate of Paramecium spp., remaining within Paramecium spp. would be beneficial for L. pneumophila to expand its habitat. To control Legionaries’ disease, understanding the ecology of L. pneumophila in the environment is essential. Our work therefore promises to facilitate further studies focused on the ecology of L. pneumophila in the environment.
Author Contributions
TN and MW conceived and designed the research. TN, NH, KW, and TS performed the experiments and analyzed the data. KW and MF offered advice and technical assistance for carrying out the studies on protists. TN, KW, and MW wrote the paper.
Funding
This work was supported by JSPS KAKENHI Grant No. 15K18782 to KW.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Al-Khodor, S., Kalachikov, S., Morozova, I., Price, C. T., and Abu Kwaik, Y. (2009). The PmrA/PmrB two-component system of Legionella pneumophila is a global regulator required for intracellular replication within macrophages and protozoa. Infect. Immun. 77, 374–386. doi: 10.1128/IAI.01081-08
Al-Quadan, T., Price, C. T., and Abu Kwaik, Y. (2012). Exploitation of evolutionarily conserved amoeba and mammalian processes by Legionella. Trends Microbiol. 20, 299–306. doi: 10.1016/j.tim.2012.03.005
Andrews, H. L., Vogel, J. P., and Isberg, R. R. (1998). Identification of linked Legionella pneumophila genes essential for intracellular growth and evasion of the endocytic pathway. Infect. Immun. 66, 950–958.
Barker, J., Scaife, H., and Brown, M. R. (1995). Intraphagocytic growth induces an antibiotic-resistant phenotype of Legionella pneumophila. Antimicrob. Agents Chemother. 39, 2684–2688. doi: 10.1128/AAC.39.12.2684
Buckley, A. M., Webber, M. A., Cooles, S., Randall, L. P., La Ragione, R. M., Woodward, M. J., et al. (2006). The AcrAB-TolC efflux system of Salmonella enterica serovar Typhimurium plays a role in pathogenesis. Cell. Microbiol. 8, 847–856. doi: 10.1111/j.1462-5822.2005.00671.x
Cunha, B. A., Burillo, A., and Bouza, E. (2016). Legionnaires’ disease. Lancet 387, 376–385. doi: 10.1016/S0140-6736(15)60078-2
Declerck, P., Behets, J., van Hoef, V., and Ollevier, F. (2007). Detection of Legionella spp. and some of their amoeba hosts in floating biofilms from anthropogenic and natural aquatic environments. Water Res. 41, 3159–3167. doi: 10.1016/j.watres.2007.04.011
Dryl, S. (1959). Antigenic transformation in Paramecium aurelia after homologous antiserum treatment during autogamy and conjugation. J. Protozool. 6:25.
Duncan, A. B., Fellous, S., Accot, R., Alart, M., Chantung Sobandi, K., Cosiaux, A., et al. (2010). Parasite-mediated protection against osmotic stress for Paramecium caudatum infected by Holospora undulata is host genotype specific. FEMS Microbiol. Ecol. 74, 353–360. doi: 10.1111/j.1574-6941.2010.00952.x
Ferhat, M., Atlan, D., Vianney, A., Lazzaroni, J. C., Doublet, P., and Gilbert, C. (2009). The TolC protein of Legionella pneumophila plays a major role in multi-drug resistance and the early steps of host invasion. PLoS One 4:e7732. doi: 10.1371/journal.pone.0007732
Fields, B. S., Shotts, E. B. Jr., Feeley, J. C., Gorman, G. W., and Martin, W. T. (1984). Proliferation of Legionella pneumophila as an intracellular parasite of the ciliated protozoan Tetrahymena pyriformis. Appl. Environ. Microbiol. 47, 467–471.
Fliermans, C. B., Cherry, W. B., Orrison, L. H., Smith, S. J., Tison, D. L., and Pope, D. H. (1981). Ecological distribution of Legionella pneumophila. Appl. Environ. Microbiol. 41, 9–16.
Fuche, F., Vianney, A., Andrea, C., Doublet, P., and Gilbert, C. (2015). Functional type 1 secretion system involved in Legionella pneumophila virulence. J. Bacteriol. 197, 563–571. doi: 10.1128/JB.02164-14
Fujishima, M., Kawai, M., and Yamamoto, R. (2005). Paramecium caudatum acquires heat-shock resistance in ciliary movement by infection with the endonuclear symbiotic bacterium Holospora obtusa. FEMS Microbiol. Lett. 243, 101–105. doi: 10.1016/j.femsle.2004.11.053
Fujishima, M., Sawabe, H., and Iwatsuki, K. (1990). Scanning electron microscopic observation of differentiation from the reproductive short form to the infectious long form of Holospora obtusa. J. Protozool. 37, 123–128. doi: 10.1111/j.1550-7408.1990.tb05881.x
Glaser, P., Sakamoto, H., Bellalou, J., Ullmann, A., and Danchin, A. (1988). Secretion of cyclolysin, the calmodulin-sensitive adenylate cyclase-haemolysin bifunctional protein of Bordetella pertussis. EMBO J. 7, 3997–4004.
Goebel, W., and Hedgpeth, J. (1982). Cloning and functional characterization of the plasmid-encoded hemolysin determinant of Escherichia coli. J. Bacteriol. 151, 1290–1298.
Görtz, H. D., and Fokin, S. I. (2009). Diversity of endosymbiotic bacteria in Paramecium. Microbiol. Monogr. 12, 131–160. doi: 10.1007/978-3-540-92677-1_6
Greub, G., and Raoult, D. (2004). Microorganisms resistant to free-living amoebae. Clin. Microbiol. Rev. 17, 413–433. doi: 10.1128/CMR.17.2.413-433.2004
Guerrier, S., Plattner, H., Richardson, E., Dacks, J. B., and Turkewitz, A. P. (2017). An evolutionary balance: conservation vs innovation in ciliate membrane trafficking. Traffic 18, 18–28. doi: 10.1111/tra.12450
Hori, M., and Fujishima, M. (2003). The endosymbiotic bacterium Holospora obtusa enhances heat-shock gene expression of the host Paramecium caudatum. J. Eukaryot. Microbiol. 50, 293–298. doi: 10.1111/j.1550-7408.2003.tb00137.x
Isberg, R. R., O’Connor, T. J., and Heidtman, M. (2009). The Legionella pneumophila replication vacuole: making a cosy niche inside host cells. Nat. Rev. Microbiol. 7, 13–24. doi: 10.1038/nrmicro1967
Kaur, S. J., Rahman, M. S., Ammerman, N. C., Beier-Sexton, M., Ceraul, S. M., Gillespie, J. J., et al. (2012). TolC-dependent secretion of an ankyrin repeat-containing protein of Rickettsia typhi. J. Bacteriol. 194, 4920–4932. doi: 10.1128/JB.00793-12
Kodama, Y., and Fujishima, M. (2005). Symbiotic Chlorella sp. of the ciliate Paramecium bursaria do not prevent acidification and lysosomal fusion of host digestive vacuoles during infection. Protoplasma 225, 191–203. doi: 10.1007/s00709-005-0087-5
Koronakis, V., Sharff, A., Koronakis, E., Luisi, B., and Hughes, C. (2000). Crystal structure of the bacterial membrane protein TolC central to multidrug efflux and protein export. Nature 405, 914–919. doi: 10.1038/35016007
Nishida, T., Watanabe, K., Tachibana, M., Shimizu, T., and Watarai, M. (2017). Characterization of the cryptic plasmid pOfk55 from Legionella pneumophila and construction of a pOfk55-derived shuttle vector. Plasmid 90, 30–37. doi: 10.1016/j.plasmid.2017.02.004
Nishino, K., Latifi, T., and Groisman, E. A. (2006). Virulence and drug resistance roles of multidrug efflux systems of Salmonella enterica serovar Typhimurium. Mol. Microbiol. 59, 126–141. doi: 10.1111/j.1365-2958.2005.04940.x
Ohno, A., Kato, N., Sakamoto, R., Kimura, S., and Yamaguchi, K. (2008). Temperature-dependent parasitic relationship between Legionella pneumophila and a free-living amoeba (Acanthamoeba castellanii). Appl. Environ. Microbiol. 74, 4585–4588. doi: 10.1128/AEM.00083-08
Paszko-Kolva, C., Shahamat, M., and Colwell, R. R. (1992). Long-term survival of Legionella pneumophila serogroup 1 under low-nutrient conditions and associated morphological changes. FEMS Microbiol. Lett. 102, 45–55. doi: 10.1111/j.1574-6968.1992.tb05794.x
Plattner, H. (2010). Membrane trafficking in protozoa SNARE proteins, H+-ATPase, actin, and other key players in ciliates. Int. Rev. Cell Mol. Biol. 280, 79–184. doi: 10.1016/S1937-6448(10)80003-6
Plattner, H., and Kissmehl, R. (2003). Molecular aspects of membrane trafficking in Paramecium. Int. Rev. Cytol. 232, 185–216. doi: 10.1016/S0074-7696(03)32005-4
Richards, A. M., Von Dwingelo, J. E., Price, C. T., and Abu Kwaik, Y. (2013). Cellular microbiology and molecular ecology of Legionella-amoeba interaction. Virulence 4, 307–314. doi: 10.4161/viru.24290
Rowbotham, T. J. (1980). Preliminary report on the pathogenicity of Legionella pneumophila for freshwater and soil amoebae. J. Clin. Pathol. 33, 1179–1183. doi: 10.1136/jcp.33.12.1179
Steinman, R. M., Mellman, I. S., Muller, W. A., and Cohn, Z. A. (1983). Endocytosis and the recycling of plasma membrane. J. Cell Biol. 96, 1–27. doi: 10.1083/jcb.96.1.1
Symmons, M. F., Marshall, R. L., and Bavro, V. N. (2015). Architecture and roles of periplasmic adaptor proteins in tripartite efflux assemblies. Front. Microbiol. 6:513. doi: 10.3389/fmicb.2015.00513
Tachibana, M., Nakamoto, M., Kimura, Y., Shimizu, T., and Watarai, M. (2013). Characterization of Legionella pneumophila isolated from environmental water and ashiyu foot spa. Biomed Res. Int. 2013:514395. doi: 10.1155/2013/514395
Thatcher, T. H., and Gorovsky, M. A. (1993). A temperature-sensitive cell cycle arrest mutation affecting H1 phosphorylation and nuclear localization of a small heat shock protein in Tetrahymena thermophila. Exp. Cell Res. 209, 261–270. doi: 10.1006/excr.1993.1310
Thomas, S., Holland, I. B., and Schmitt, L. (2014). The Type 1 secretion pathway - the hemolysin system and beyond. Biochim. Biophys. Acta 1843, 1629–1641. doi: 10.1016/j.bbamcr.2013.09.017
VieBrock, L., Evans, S. M., Beyer, A. R., Larson, C. L., Beare, P. A., Ge, H., et al. (2015). Orientia tsutsugamushi ankyrin repeat-containing protein family members are Type 1 secretion system substrates that traffic to the host cell endoplasmic reticulum. Front. Cell. Infect. Microbiol. 4:186. doi: 10.3389/fcimb.2014.00186
Wakeel, A., den Dulk-Ras, A., Hooykaas, P. J., and McBride, J. W. (2011). Ehrlichia chaffeensis tandem repeat proteins and Ank200 are type 1 secretion system substrates related to the repeats-in-toxin exoprotein family. Front. Cell. Infect. Microbiol. 1:22. doi: 10.3389/fcimb.2011.00022
Watanabe, K., Nakao, R., Fujishima, M., Tachibana, M., Shimizu, T., and Watarai, M. (2016). Ciliate Paramecium is a natural reservoir of Legionella pneumophila. Sci. Rep. 6:24322. doi: 10.1038/srep24322
Watarai, M., Derre, I., Kirby, J., Growney, J. D., Dietrich, W. F., and Isberg, R. R. (2001). Legionella pneumophila is internalized by a macropinocytotic uptake pathway controlled by the Dot/Icm system and the mouse Lgn1 locus. J. Exp. Med. 194, 1081–1096. doi: 10.1084/jem.194.8.1081
Winiecka-Krusnell, J., and Linder, E. (1999). Free-living amoebae protecting Legionella in water: the tip of an iceberg? Scand. J. Infect. Dis. 31, 383–385. doi: 10.1080/00365549950163833
Xu, L., and Luo, Z. Q. (2013). Cell biology of infection by Legionella pneumophila. Microbes Infect. 15, 157–167. doi: 10.1016/j.micinf.2012.11.001
Keywords: Legionella pneumophila, Paramecium, TolC, cellular trafficking, symbiosis
Citation: Nishida T, Hara N, Watanabe K, Shimizu T, Fujishima M and Watarai M (2018) Crucial Role of Legionella pneumophila TolC in the Inhibition of Cellular Trafficking in the Protistan Host Paramecium tetraurelia. Front. Microbiol. 9:800. doi: 10.3389/fmicb.2018.00800
Received: 05 February 2018; Accepted: 10 April 2018;
Published: 25 April 2018.
Edited by:
Suhelen Egan, University of New South Wales, AustraliaReviewed by:
Helen Zgurskaya, The University of Oklahoma, United StatesM. Victoria Delpino, National Scientific and Technical Research Council (CONICET), Argentina
Copyright © 2018 Nishida, Hara, Watanabe, Shimizu, Fujishima and Watarai. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Masahisa Watarai, d2F0YXJhaUB5YW1hZ3VjaGktdS5hYy5qcA==
 Takashi Nishida
Takashi Nishida Naho Hara1
Naho Hara1 Kenta Watanabe
Kenta Watanabe Takashi Shimizu
Takashi Shimizu Masahiro Fujishima
Masahiro Fujishima Masahisa Watarai
Masahisa Watarai